Abstract
Magnetic resonance imaging (MRI) has become an important imaging technique for quantifying the spatial location and magnitude/direction of longitudinal cartilage morphology changes in patients with osteoarthritis (OA). Although several analytical methods, such as subregion-based analysis, have been developed to refine and improve quantitative cartilage analyses, they can be suboptimal due to two major issues: the lack of spatial correspondence across subjects and time and the spatial heterogeneity of cartilage progression across subjects. The aim of this paper is to present a statistical method for longitudinal cartilage quantification in OA patients, while addressing these two issues. The 3D knee image data is preprocessed to establish spatial correspondence across subjects and/or time. Then, a Gaussian hidden Markov model (GHMM) is proposed to deal with the spatial heterogeneity of cartilage progression across both time and OA subjects. To estimate unknown parameters in GHMM, we employ a pseudo-likelihood function and optimize it by using an expectation-maximization (EM) algorithm. The proposed model can effectively detect diseased regions in each OA subject and present a localized analysis of longitudinal cartilage thickness within each latent subpopulation. Our GHMM integrates the strengths of two standard statistical methods including the local subregion-based analysis and the ordered value approach. We use simulation studies and the Pfizer longitudinal knee MRI dataset to evaluate the finite sample performance of GHMM in the quantification of longitudinal cartilage morphology changes. Our results indicate that GHMM significantly outperforms several standard analytical methods.
Index Terms: Diseased regions detection, EM algorithm, Gaussian hidden Markov model, Longitudinal cartilage thickness, Pseudo-likelihood method
I. Introduction
Osteoarthritis (OA) known as degenerative arthritis represents a group of mechanical abnormalities involving degradation of joints, including articular cartilage and subchondral bone. Symptoms include swelling, pain, discomfort, locking, and other problems in mobility and are often characterized by the progressive loss of joint cartilage [1]. Bone may be exposed and damaged without the protection of cartilage. Figure 1 shows the anatomy of the human knee and illustrates the cartilage loss. OA is the most common form of arthritis and a major cause of chronic disability in the United States [2]. It is estimated that more than 16% of all adults 45 years or older suffer from symptomatic OA of the knee [1]. Cartilage loss [3] is believed to be the dominating factor in OA. Studying cartilage morphological changes can help to understand OA progression and drug development.
Fig. 1.

Anatomy of human knee and illustration of cartilage loss. (left) A sagittal slice of a healthy knee. Bones are annotated in blue, femoral cartilage in purple and tibial cartilage in orange. (middle) A coronal slice of the same healthy knee. (right) A coronal slice of an OA knee with cartilage loss indicated by the red arrow.
In recent years, magnetic resonance imaging (MRI) has increasingly been used in OA diagnosis [4], [5], since MRI offers the distinct advantage of detecting signal and morphologic changes in articular cartilage in 3D. MRI is the most sensitive and accurate test for evaluating the articular cartilage noninvasively [6]. Particularly, MRI can demonstrate reactive bone edema or soft tissue swelling as well as small cartilage or bone fragments in the joint. However, little is known about the development of such cartilage lesions and their natural progression.
Recently, several large MRI databases have been acquired for OA research in order to help understand the progression of cartilage loss across OA subjects and to identify risk factors (such as genetics, dietary intake, and muscle weakness) associated with such loss [7]–[10]. For example, the Pfizer A9001140 is a longitudinal study, including baseline, 3-month, 6-month, 1-year and 2-year follow-up images of a single knee from 155 subjects [7], [8]; the Osteoarthritis Initiative (OAI) is another longitudinal cohort study of risk factors for incidence and progression of OA, where 4796 subjects with or at high risk of knee OA were recruited and image data was collected annually during 4 years of follow-up [9], [10]. Moreover, analyzing such large MRI datasets is necessary to establish knowledge about the expected pace of progression in knees with different exposure to risk factors for structural progression (e.g. radiographic joint space narrowing [JSN]). Such knowledge can then be used for the design of interventional trials.
Statistical analysis of cartilage thickness changes faces two major challenges. The first one is the lack of spatial correspondence across subjects and/or time. Observational studies have shown that cartilage loss in OA may not be uniform throughout the cartilage [11], and advanced statistical analysis methods are necessary to understand localized cartilage thinning. However, the localized analysis requires establishing spatial correspondence across time and subjects, and such analysis is challenging due to the small volume of cartilage in relation to the rest of the knee. The second challenge is the spatial heterogeneity of cartilage progression across subjects. Cartilage thinning may happen at different locations in different subjects, which is usually caused by biomechanical factors [12] or local risk factors [13]. Cartilage thinning at a particular location may only be consistent for a subset of the full subject population. Thus, treating all OA subjects equally and using all of them for standard statistical methods (such as voxel-wise analysis and region-of-interest analysis) would be problematic. In clinical practice, some non-surgical treatments for OA in the knee (e.g., corticosteroid injections) are most likely administered via injection and act globally [14] since the exact locations of lesions are unknown. Therefore, it is of great interest to propose some methods to detect such locations and refine these treatments.
Subregion-based analysis and the ordered value approach are two major statistical methods for quantifying cartilage thickness changes. Subregion-based analysis is one of the most popular approaches. Here, average thickness changes in defined anatomic subregions of the femorotibial joint are studied. In [15], the tibial cartilage was divided into a central area of the total subchondral bone area, and anterior, posterior, internal, and external subregions surrounding it. However, this approach is problematic due to the spatial heterogeneity of cartilage progression discussed above. In another region-wise statistical analysis method [16], a region partitioning process is performed by using a supervised machine learning framework on anatomically aligned medial tibial and medial femoral cartilage thickness maps, and then a point distribution model is used to establish spatial correspondence and a common coordinate system. However, their machine learning procedures are only implemented on the thickness maps at baseline. Buck et al. [17] proposed an ordered value approach by sorting subregional changes within each knee by ascending order to focus on the magnitude of the thickness irregardless of the location, in which subregion the change occurs in each subject. The ordered value approach was demonstrated to provide improved discrimination between healthy subjects and OA participants longitudinally [17]. The key advantage of the ordered value approach is to remove the link between the spatial location and the magnitude/direction of the change in cartilage thickness. But the ordered value approach still relies on subdivisions of the cartilage and the average thickness within each subregion. The subdivisions of the cartilage are purely geometric, and small local changes (that happen to a smaller region than the size of a subregion) can be difficult to recover, since they may be weakened by averaging over a large subregion. Therefore, to fully understand the spatial pattern of OA progression, it is necessary to pinpoint localized cartilage thickness changes.
The paper is organized as follows. Section II summarizes the image preprocessing pipeline and the key contributions in this paper. Section III introduces a novel Gaussian hidden Markov model (GHMM) and its related EM algorithm. In Section IV, simulation studies are performed to show the advantages of our proposed model in diseased regions detection. Section V applies GHMM to the 2D thickness maps derived from the 3D MRI data of the Pfizer Longitudinal Study (PLS-A9001140).
II. Image Preprocessing and Main Contributions
In this paper, the 3D knee MR images are preprocessed via a set of four image analysis steps including (I.1) segmentation, (I.2) 3D thickness map computation, (I.3) registration, and (I.4) thickness map projection. These four methods summarized in Fig. 2 can successfully establish spatial correspondence across subjects and/or time and therefore address the first challenge in statistical analysis of cartilage thickness changes.
Fig. 2.
Path diagram of the 3D knee MR image data preprocessing toolbox: ➀ extracting the joint region and obtaining the cartilage segmentation; ➁ computing the 3D cartilage thickness map; ➂ registering all thickness maps to a common atlas space; and ➃ projecting 3D thickness map onto a 2D plane.
The main contributions of this paper include (i) a statistical method based on GHMM to appropriately address the spatial heterogeneity of cartilage progression across OA subjects and quantify their longitudinal cartilage morphology changes and (ii) a systematic evaluation of our analysis method by using simulations and real data analysis. Moreover, we have developed companion software for GHMM and will release it to the public through http://www.nitrc.org/ and http://www.bios.unc.edu/research/bias.
The path diagram of GHMM is presented in Fig. 3. Specifically, a novel hidden Markov random field (HMRF) [18]–[27] is proposed to specifically discover the diseased regions for each OA subject and to integrate cartilage progression in different diseased regions with different change magnitudes/directions across OA subjects. Moreover, based on the correspondence established in the preprocessing step, our GHMM allows us to carry out a localized analysis of longitudinal cartilage thickness at each voxel. Therefore, GHMM integrates the strengths of both the local subregion-based analysis and the ordered value approach in a single framework.
Fig. 3.

Path diagrams of GHMM component. (a) Path diagram of GHMM for four different subjects (one normal control and three OA subjects). The change of the map thickness at pixel site k includes three parts in the proposed model, i.e., (i) the common dynamic change in both normal controls and OA subjects, (ii) the change caused by random effects and measurement errors, and (iii) the change caused by different diseased regions (moderately diseased region (yellow) and severely diseased region (red)). (b) Path diagram of GHMM for one subject at two neighboring pixel sites across time. Following the convention of path diagrams, observed variables are framed with rectangles whereas latent variables are framed with ellipses. Regression associations are depicted as (solid) one-headed arrows (parent=predictor, child=response); correlations between diagnosis result, the diseased region label and the observed thickness are described as (dotted-dashed) one-headed arrows; and the correlation of diseased region labels in the Potts model is shown as (solid) double-headed arrow. For easier interpretation, the following color-codes are used: exogenous variables (light red), endogenous variables (purple), continuous latent variables (blue), discrete latent variables (green), and measurement errors (yellow).
III. Background and Methods
A. Cartilage Segmentation
Cartilage segmentation in the knee has been investigated for several decades. Many segmentation approaches have been proposed, e.g., region growing methods [28], [29], watershed methods [30], live wire methods [31], active contour methods [32], [33], and graph cut methods [34]. However, all the aforementioned methods are semi-automatic, which precludes their applications to large image databases. Recently, several automatic methods have been proposed for cartilage segmentation, including pattern recognition methods [35], model-based segmentation method [36], and graph-based method [37]. However, the spatial interactions between neighboring pixels are neglected in [35]; the model-based segmentation method [36] is prone to local minima in the fitting process; the graph-based method [37] generally suffers from grid bias, also known as metrication errors.
Here we applied the automatic cartilage segmentation method [38], which is an extension of the multi-atlas segmentation approach initially proposed by us for cartilage segmentation in [39]. The main difference in these two cartilage segmentation approaches is how label-fusion is achieved once multiple segmentation candidates (from the multiple atlases) are available. In [39], a simple locally weighted label-fusion strategy was used, whereas in [38] the locally weighted label-fusion was compared to the baseline majority-voting approach and a more advanced non-local patch-based label fusion approach was also investigated. The latter approach turned out to be superior to the other two label-fusion strategies. In general, the segmentation method [38] has the following advantages. First, the method is fully automatic and requires no user interaction (besides quality control). Secondly, the method is robust as it benefits from multi-atlas-based strategies. Thirdly, both spatial and local appearance information are utilized in the segmentation. Local tissue classification is probabilistic (unlike [35]) and combined with the spatial prior to generate the final segmentation within a segmentation model. Furthermore, the segmentation model is convex and thus allows for the computation of global optimal solutions which cannot be guaranteed by active contour models, active shape models or active appearance models. The segmentation model also allows for the incorporation of spatial and temporal regularization.
The approach that we use for segmenting cartilage and to compute local thicknesses works in a three-dimensional (3D) space. Specifically, our segmentation produces a 3D label map for femoral and tibial cartilage. Given a segmentation, we compute thickness using a Laplace-equation approach [40] through which we define a top and a bottom of the structure and correspondence lines between them. Each point on such a correspondence line is then assigned the thickness value (distance between beginning and end of this line) resulting in a 3D distance map. It should be noted that, all the segmentations are performed for each subject independently and in their own untransformed coordinate systems to avoid any bias introduced by resampling the image. For the same reason, the thickness maps are computed in the native image spaces obtained from cartilage segmentations.
Once these measures have been computed, population comparisons require a common coordinate system for the statistical analysis across time-points and subjects, which requires an additional registration step to this common space. Note that this may cause slight local distortions (depending on the quality of the registration), but it will not change the actual thickness measures as they are computed in the native image spaces. Here, we have used an affine transformation followed by a B-Spline transform based on the bone segmentations for each cartilage thickness volume. Since we are dealing with sheet-like structures, which we have approximately axis-aligned in atlas space, we simply compute the local median values of this 3D thickness map in the axial directions, where thicknesses are approximately constant.
B. Gaussian Hidden Markov Model
We consider a longitudinal thickness map dataset with thickness data {yij(sk) : k = 1, …, m} measured at time tij for j = 1, …, Ti and i = 1, …, n, where n is the total number of subjects, S0 = {s1, …, sm} is the set of m pixels in a common template, denoted by
 , and Ti is the total number of time points for the i-th subject. Let xi represent disease status of OA for each subject such that xi = 0 and 1, respectively, represent normal control and patient with OA.
, and Ti is the total number of time points for the i-th subject. Let xi represent disease status of OA for each subject such that xi = 0 and 1, respectively, represent normal control and patient with OA.
We introduce a latent partition of
 for each subject. Specifically, it is assumed that for the i–th subject,
for each subject. Specifically, it is assumed that for the i–th subject,
 can be decomposed into the union of normal region Ri0, moderately diseased region Ri1, and severely diseased region Ri2, that is
can be decomposed into the union of normal region Ri0, moderately diseased region Ri1, and severely diseased region Ri2, that is
Here we also assume that normal controls are expected to be perfectly healthy, i.e. to not have any cartilage loss or lesions. Then, both Ri1 and Ri2 should be the empty set for normal controls. For OA patients, the size and location of Ri1 and Ri2 may vary across subjects and play a critical role in establishing spatial correspondences of diseased regions at different locations across subjects. Statistically, we introduce an unobserved random effect bi(sk) ∈ L = {0, 1, 2} to label Ri0, Ri1, and Ri2 at each pixel sk of
 for the i–th subject. Thus, bi(sk)s are label configurations of three nonoverlapping regions {Ri0, Ri1, Ri2}. It is also possible to extend GHMM to the scenarios with more than two types of diseased regions.
for the i–th subject. Thus, bi(sk)s are label configurations of three nonoverlapping regions {Ri0, Ri1, Ri2}. It is also possible to extend GHMM to the scenarios with more than two types of diseased regions.
Our Gaussian hidden Markov model consists of a spatial random effect model and a Potts model [20], [21], [23]. Spatial random effects models (SREM), also called hidden Markov models, are very useful for image segmentation and restoration in image analysis [18], [19], [22], [24]–[27].
We first propose SREM to model the conditional distribution of the observed thickness maps given two sets of random effects including bi = {bi(sk) : sk ∈
 } and vi = {vi(sk) : sk ∈
} and vi = {vi(sk) : sk ∈
 }, where vi(sk) will be defined below. We present the path diagrams of SREM in Fig. 3. Given bi and vi, we propose a general spatial random effect model as
}, where vi(sk) will be defined below. We present the path diagrams of SREM in Fig. 3. Given bi and vi, we propose a general spatial random effect model as
| (1) |
where wij is a qw × 1 vector of covariates (e.g., time, gender, or genetic marker) that usually includes an intercept 1, but is independent of disease status. Moreover, β(sk) and vi(sk) are both qw × 1 vectors of regression coefficients. It should be noted that, if wij only comprises an intercept 1 and time, β(sk) represents both the average thickness at baseline and average thickness change rate at time point tij, while the two components in vi(sk) indicate the individual thickness difference relative to the average thickness at baseline and the individual difference of the thickness change rate relative to the average thickness change rate at time point tij respectively. The c(xi, wij, bi(sk), ᾱ) is defined as
where δ(·, ·) is the Kronecker function and ᾱ = (ᾱ(1)T, ᾱ(2)T)T is a 2qw × 1 vector of coefficients, in which ᾱ(1) and ᾱ(2) respectively characterizes the dynamic thickness differences in the diseased regions Ri1 and Ri2 in addition to the dynamic thickness changes at pixel sk in normal regions. Thus, c(xi, wij, bi(sk), ᾱ) equals zero for all pixels for normal controls and the pixels with bi(sk) = 0 for OA patients. For OA patients, pixels in different diseased regions may have different dynamic thickness changes. Throughout the paper, we assume wij = (1, tij)T, β(sk) = (β1(sk), β2(sk))T and set qw = 2 for simplicity. Generally, the time measurement tij in the first two terms of model (1) is not required to be the same as the one in the last item of model (1). In principle, age is used as tij in the first two terms of (1) to capture the average thickness changes across age, whereas time to diagnosis is used in the last term of (1) to character the relative thickness changes in the diseased regions caused by OA. However, neither the age nor time to diagnosis are available in this dataset. Instead, the scan date is the only time measurement in this dataset. Since the range of thickness changes is small compared to the thickness at baseline and the time range (2 years) is also small relative to the age of each subject (all subjects are middle aged [41]), it is reasonable to use the scan date as tij throughout the model (1). In this paper, for each subject, the fist scan date was set to 0 and the following scan dates were calculated in years, e.g., 0.5 means six months. Therefore, it is straightforward to consider some basis functions (such as spline or wavelet) for wij. Moreover, εij(sk)s are independent measurement errors across subjects, time, and pixels, following .
Second, we model the random effects bi and vi as follows. First, it is assumed that bi = (bi(s1), …, bi(sm))T and vi = {vi(sk) : k = 1, …, m} are mutually independent. Moreover, vi(sk) are assumed to be mutually independent across pixels and vi(sk) follows N(0, Σvsk) at pixel sk. It is assumed that bis are independent across subjects and each bi follows a Potts model [20], [21], [23], whose Gibbs form is given by
| (2) |
where U(bi) = −Σsk~sl δ(bi(sk), bi(sl)) and τ is introduced to encourage spatial smoothness in homogeneous regions. Moreover, C(τ) is the partition function such that p(bi|τ) is a probability function. The notation “Σsi~sj” means that si is a neighbor of sj and each neighboring pair enters the summation only once. Throughout the paper, we only consider the closest 4 neighbors of each pixel.
To illustrate the key ideas of GHMM, we present two path-diagrams of GHMM in Figure 3(a) and (b). Model (1) has two key novelties compared with the subregion-based analysis and the ordered value approach. As shown in the subcomponents (i) and (ii) of Figure 3(a), the first novelty is that allows us to utilize information in all normal regions Ri0 across all normal subjects and OA patients and then carry out a localized (or pixel-wise) analysis of longitudinal cartilage thickness at each pixel. As shown in the subcomponent (iii) of Figure 3(a), the second novelty is that the use of c(xi, wij, bi(sk), ᾱ) and ᾱ allows us to integrate information in all diseased regions Ri1 ∪ Ri2 across all subjects together and then carry out a joint analysis in the union of all moderately diseased regions (or all severely diseased regions ) across all OA patients.
C. Estimation Procedure
Our next problem of interest is to estimate all unknown parameters, denoted as θ, and random effects. Here θ is a vector of all unknown parameters consisting of τ, β(sk), ᾱ, , and the covariance matrices Σvsk for k = 1, …, m. We decompose the parameter θ into two parts: τ, and all others, denoted as θ̄. The maximum likelihood estimate (MLE) of θ̄ can be calculated by using the expectation-maximization (EM) algorithm [42], whereas τ can be estimated by using a pseudo-likelihood method [43] since the MLE of τ is generally difficult to compute due to the normalizing part of the probability function in (2).
To derive the EM algorithm, we need the complete-data log-likelihood function of GHMM as follows. Let yi(sk) = (yi1(sk), …, yiTi (sk))T, Wi = (wi1, …, wiTi), c̄(xi, Wi, bi(sk), ᾱ) = (c(xi, wi1, bi(sk), ᾱ), …, c(xi, wiTi, bi(sk), ᾱ))T, , where ITi is Ti × Ti identity matrix. Thus, the distribution of yi(sk) conditional on random effects is given by N(zi(sk), Σski), where zi(sk) is given by . Thus, the complete-data log-likelihood function is given by
| (3) |
where n0 = #{xi : xi ≠ 0} denotes the number of OA patients. Given the current estimate θ̃(r) at iteration r, the value is obtained via maximizing the following Q-function Qθ̃(r) (θ̄) ≐ Eθ̃(r) [log L̃(θ)|y, x] with respect to θ̄:
| (4) |
We consider the E-step and M-step of the EM algorithm as follows.
E-step
In the E-step, we need to calculate four conditional expectations: E[vi(sk)δ(bi(sk), l)|yi(sk), x, θ̃(r)], E[vi(sk)|yi(sk), x, θ̃(r)], , and E[δ(bi(sk), l)|yi(sk), x, θ̃(r)]. In order to calculate these conditional probabilities, the class labels b should be estimated first. Here we consider the maximum a posteriori on Markov random field (MRF-MAP) estimation, which is efficient and adopted in many existing papers, e.g., [23], [44], [45]. First, the conditional probability density function of yi(sk) given xi and bi(sk) is derived as
| (5) |
where and .
According to the MAP criterion, is defined as
| (6) |
To obtain the optimal solution to (6), in this paper, we adopt the iterated conditional modes (ICM) algorithm [20], which which uses a greedy iterative strategy for minimization. Convergence is achieved after only a few iterations. As a result, the conditional expectation E[δ(bi(sk), l)|yi(sk), x, θ̃(r)] can be calculated as
| (7) |
where
| (8) |
Here
 is the set of sites neighboring sk.
is the set of sites neighboring sk.
Given xi and is normally distributed as
Then, given yi(sk), x, and , we have
Finally, the desired expectations can be estimated as
| (9) |
M-step
Taking derivatives of (4) with respect to θ̄ and equating them to zeros, we find the updates of θ̄ as follows. For β(sk) and ᾱ(l), we have
| (10) |
and
| (11) |
For the covariance matrix Σvsk, we have
| (12) |
For in Σsk i,
| (13) |
where .
Since τ in model (2) is not the primary parameter of interest, we use an approximate, but computationally efficient method based on a pseudo-likelihood function. A key advantage of using the pseudo-likelihood function is its computational simplicity, since it does not involve the intractable partition function. The pseudo-likelihood at the r-th step is a simple product of the conditional likelihood
| (14) |
where ∂
 denotes the set of points at the boundaries of
denotes the set of points at the boundaries of
 , and
is given by
, and
is given by
Thus, the MPL estimate τ̃(r+1) can be obtained by solving
| (15) |
The E-step and M-step are repeated until the difference between log L(θ̃(r+1)) and log L(θ̃(r)) is smaller than a desired value (e.g., 10−4).
D. Inference Procedure
After all the parameters are estimated, we carry out formal statistical inference consisting of three different statistical tools below. First, we calculate the standard errors of computed MLEs, β̂(sk), at each voxel sk and in the moderately and severely diseased regions. As captures the information in all normal regions Ri0 across all normal subjects and OA patients, we calculate the standard errors of β̂(sk) via pixel-wise analysis of longitudinal cartilage thickness on all subjects with b̂i(sk) = 0, in which model (1) is simplified as follows:
| (16) |
Thus, the estimated covariance matrix of β̂(sk) can be approximated by
| (17) |
where ñ(sk) is the number of subjects with b̂i(sk) = 0 at voxel sk. Then the standard errors can be estimated by the positive square root of the diagonal elements in (17).
On the other hand, the standard error of α is estimated based on data in the voxels of all diseased regions in all OA patients, since α captures the information in all diseased regions Ri1 ∪ Ri2 across all subjects together. Since the information of these diseased regions has to be estimated from the data, the standard errors associated with α̂ are the standard errors conditional on the given diseased regions across subjects. Moreover, the pixel-wise analysis is not suitable here, and thus it is impossible to obtain explicit expressions of the estimated standard errors. In order to tackle this problem, a resampling approach, wild bootstrap [46], is considered here. The idea of wild bootstrap is to leave the regressors at their sample value, but to resample the response variable based on the residuals values. This method proceeds as follows:
Fit the model with the original data and retain the fitted values and the residuals ε̂i(sk) = yi(sk) − ŷi(sk) for i = 1, …, n and k = 1, …, m;
Create synthetic response variables yi(sk)* = ŷi(sk) + aiε̂i(sk), where ai is a random variable following standard normal distribution;
Refit the model using the synthetic response variables yi(sk)* and retain the estimates ;
Repeat Steps 1 and 2 independently B times to give B independent realizations of , denoted by ;
-
The bootstrap covariance matrix of is estimated by
(18) where for l = 1, 2. The standard error of can be estimated by the positive square root of the diagonal element of (18).
It has been shown that 50 to 100 bootstrap replications (B = 100 in this paper) are generally sufficient for standard error estimation [47].
Second, several hypothesis testing problems are quite interesting in real applications. Specifically, we are interested in testing (i) whether there is any cartilage progression across time at each pixel site in normal regions; (ii) whether there is any difference of cartilage thickness at baseline between normal regions and moderately (or severely) diseased regions; and (iii) whether there is any difference of cartilage progression across time between normal regions and moderately (or severely) diseased regions. They can written as
| (19) |
| (20) |
| (21) |
| (22) |
| (23) |
where ᾱl′(l) is the l′th component of ᾱ(l), l = 1, 2. A sequence of t tests can be used here. The standard errors of estimates β̂2(sk), k = 1, …, m, and , l = 1, 2 can be obtained as described above. The false discovery rate (FDR) adjustment method [48] is employed here to calculate the adjusted p-values corrected for the multiple comparison problem (19).
Third, after obtaining the diseased region labels across all pixel sites for each OA subject, we calculate the empirical probabilities of both the moderately diseased region and severely diseased region at each pixel across all OA subjects. The related formula is listed below:
| (24) |
where #A denotes the number of the elements in set A.
IV. Simulation Studies
We examine the finite sample performance of GHMM for diseased region detection. We first fitted the model (1) to the femoral cartilage thickness data of all the normal controls derived from the 3D MRI data of the Pfizer Longitudinal Study (PLS-A9001140) and then we used the obtained parameter estimators of β(sk), Σvsk, and as their true values for simulations. Moreover, ᾱ(1) and ᾱ(2) were set (−0.17, −0.19)T and (−0.26, −0.33)T, respectively. The covariate vector wi for each subject was generated according to the real data set analyzed in Section V. We generated 20 subjects from 3 groups of OA patients with 7 patients in group 1, 7 patients in group 2, and 6 patients in group 3. Without loss of generality, the location of all diseased regions is predetermined and does not vary for subjects in the same group, whereas it varies across groups. Fig. 4 presents different lesions that were manually designed and inserted into the thickness maps and the diseased regions associated with the three simulated groups.
Fig. 4.

Visualizations of simulated lesions in femoral cartilage sheets: normal region (blue); moderately diseased region (light blue); severely diseased region (orange).
We applied GHMM to detect the diseased regions in each subject. We randomly selected one subject from each group and presented its detection results in Fig. 5. For all selected subjects, the diseased regions can be successfully detected with few misclassifications in the results. To show the average detection performance among subjects in each group, the Jaccard index [49], Dice index [50], Rand index [51], and adjusted Rand index [52] were calculated to compare the detection results with the truth. Finally, we calculated the averages of all index values taken over all subjects in each subgroup. For all these indices, a higher index value indicates a more accurate detection result. Simulation results based on all the three groups are presented in Table I, where the related standard deviations are also shown in parentheses. For all these groups, the average of the four index values are all larger than 86 percent, and the related standard deviations are all less than 0.05. Figure 5 and Table I indicate that GHMM performs very well in diseased region detection of cartilage MRI data.
Fig. 5.
Diseased region detection for subjects from three groups: (a) subject with 5 time points in group 1; (b) subject with 3 time points in group 2; (c) subject with 4 time point in group 3. This is the simulated result and hence we have an imposed ground truth, which is shown in Fig. 4. In each group, the image with green background indicates the detection result, i.e., normal region (blue); moderately diseased region (light blue); severely diseased region (orange), the remaining ones are the observations for the time points, where the color bar for all the observations is presented on the right side of the fist raw in (a)–(c).
TABLE I.
Average detection performance among subjects in different groups
| Group 1 | Group 2 | Group 3 | |
|---|---|---|---|
| Jaccard index | 0.8509 (0.0045) | 0.9200 (0.0113) | 0.8683 (0.0040) |
| Dice index | 0.9142 (0.0026) | 0.9583 (0.0078) | 0.9295 (0.0023) |
| Rand index | 0.9794 (0.0114) | 0.9592 (0.0124) | 0.9721 (0.0097) |
| Adjusted Rand index | 0.9242 (0.0399) | 0.8749 (0.0500) | 0.8853 (0.0367) |
V. Real Data Analysis
We consider the MRI dataset from the Pfizer longitudinal non-treatment study on osteoarthritis (PLS-A9001140). This dataset contains T1-weighted (3D SPGR) images from 155 subjects imaged at baseline, 3, 6, 12, and 24 months at a resolution of 1.00 × 0.31 × 0.31mm3. All the subjects are females. Some of them have missing scans and thus we have 706 MRIs in total. The Kellgren-Lawrence grades (KLG)1 [53] were determined for all subjects from the baseline extended weight-bearing radiographs, classifying 81 subjects as normal control (KLG0), 1 subject as KLG1 (mild OA), 40 subjects as KLG2 (severe OA), and 33 subjects as KLG3 (severe OA). It seems that most of the OA patients in this dataset were selected to have predominantly medial OA.
We first computed the cartilage segmentations in the knee MRI data. We randomly chose two subjects with different KLG from the dataset, and presented the related 2D thickness maps for the chosen subjects at baseline in Fig. 6.
Fig. 6.
Local cartilage thickness (mm) for subjects with automatic segmentations at baseline with different KLGs: (top) femoral cartilage (FC) thickness map (left) and tibial cartilage (TC) thickness map (right) for subjects with KLG0; (bottom) femoral cartilage (FC) thickness map (left) and tibial cartilage (TC) thickness map (right) for subject with KLG3. The medial (M), lateral (L), anterior (A), and posterior (P) directions are annotated in the thickness map (top & left).
After extraction of the 2D thickness maps, we first applied the linear mixed model for voxel-based analysis to the thickness maps from the whole dataset and calculated the parameter estimation at each pixel site. The model is described as below
| (25) |
where wij, xi, vi(sk) and εij(sk) are all defined the same as those in (1). Compared with GHMM, the reduced model (25) cannot handle the spatial heterogeneity of cartilage progression across subjects, and thus it can only capture the diseased regions at population level instead of individual level. The estimates of α1(sk) across all the pixels in both femoral and tibial cartilages are presented in Fig. 7(a) and (d). We carried out a sequence of t tests on the hypotheses H0(sk) : α1(sk) = 0 versus H1(sk) : α1(sk) ≠ 0 across all pixels to identify the regions with substantial differences between OA patients and normal controls. Figure 7(b) and (e) present the FDR adjusted − log10(p)-value [48] at each pixel in both femoral and tibial cartilages. Based on the testing results, we are able to detect some significant subregions in the cartilage. Furthermore, for better visualization, we used the K-means clustering method [54] to cluster voxels into several sub-regions based on the FDR adjusted −log10(p)-map. We chose clustering −log10(p)-values, since the p-value indicates the statistical significance of the effect sizes. The clustering results are also presented in Fig. 7(c) and Fig. 7(f), where four subregions are discovered for both femoral and tibial cartilage. The mean values of FDR adjusted −log10(p)-value for all the pixel sites in each cluster are also summarized in Table II. Table II reveals that the first three clusters are diseased regions, whereas the fourth one is a normal region. Such results are averages across all the subjects. Since the cartilage thinning happens at different locations for different subjects, the voxel-based analysis based on model (25) fails to capture the diseased regions for individual OA subjects.
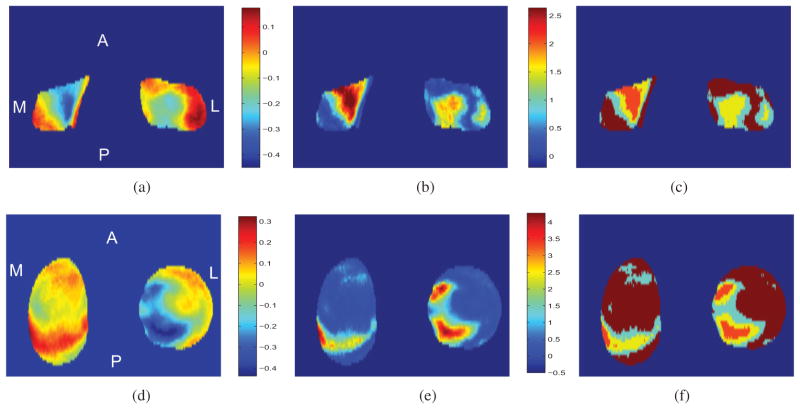
Fig. 7.
Voxel-wise analysis based on linear mixed model (25): (a) estimates of α1(sk) in femoral cartilage; (b) FDR adjusted −log10(p)-values in femoral cartilage; (c) clustering results based on FDR adjusted −log10(p)-values in femoral cartilage: cluster 1 (light blue), cluster 2 (yellow), cluster 3 (orange), cluster 4 (dark red); (d) estimates of α1(sk) in tibial cartilage; (e) FDR adjusted −log10(p)-values in tibial cartilage; and (f) clustering results based on FDR adjusted −log10(p)-values in tibial cartilage: cluster 1 (light blue), cluster 2 (yellow), cluster 3 (orange), and cluster 4 (dark red).
TABLE II.
Mean values of FDR adjusted −log10(p)-value for all the pixel locations in each cluster
| Cartilage | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 |
|---|---|---|---|---|
| femoral | 1.2565 | 2.3616 | 3.8733 | 0.3111 |
| tibial | 1.5198 | 3.0583 | 5.0483 | 0.3491 |
As a comparison, we applied GHMM to the whole dataset. As an illustration, the estimates of β(sk) across all the pixels in both femoral and tibial cartilages and their standard errors obtained via wild bootstrap are presented in Fig. 8(a)–(h). From Fig. 8(c)–(d) and (g)–(h), coefficients of most pixels show the range of cartilage thickness changes in normal regions similar to those in [17]. However, coefficients of some pixels show progression in normal regions. One of the reasons for these progressions may be that some regions may undergo tissue swelling, diluting, or even masking cartilage loss in adjacent pixels [15]. The FDR adjusted −log10(p)-value for test (19) at each pixel is calculated in Fig. 8(i) and (j). The inference results of α, including estimation, standard error and −log10(p)-value for tests (20)–(23), are presented in Table III as well. Fig. 8 and Table III reveal that (i) there is significant cartilage progression across time on only a few pixel sites in normal regions; (ii) the difference of cartilage progression across time between normal regions and moderately (or severely) diseased regions is highly significant with respect to their cartilage thickness at baseline; and (iii) both the difference of cartilage thickness at baseline and difference of cartilage progression across time between normal regions and severely diseased regions are larger than those between normal regions and moderately diseased regions. Then we chose two pixel sites from the femoral and tibial cartilages, respectively, and presented them in Fig. 9(a). Fig. 9(b)–(g) show the dynamic changes of cartilage thickness at two given pixel sites across time. We also included both the estimated curves and thickness changes of some chosen subjects. The estimated cartilage thicknesses at both two pixel sites follow a trend of decline across time for OA patients. Moreover, the estimated cartilage thickness in severely diseased regions decreases more quickly than that in moderately diseased regions. We randomly selected 3 subjects with different KLGs, i.e., 1 KLG1 subject, 1 KLG2 subject, and 1 KLG3 subject, and presented their diseased regions in Fig. 10. Fig. 11 presents the empirical probabilities of both moderately diseased region and severely diseased region. The locations of diseased regions change across subjects, and the detected diseased regions are more inclined to locate in some regions (e.g., center) of the cartilage. This indicates that the cartilage loss in OA may not be uniform throughout the cartilage, but OA may affect certain regions more frequently than others [11].
Fig. 8.
Inference results of β(sk) based on GHMM: (a) estimates of β1(sk) in femoral cartilage; (b) standard errors of estimates β̂1(sk) in femoral cartilage; (c) estimates of β2(sk) in femoral cartilage; (d) standard errors of estimates β̂2(sk) in femoral cartilage; (e) estimates of β1(sk) in tibial cartilage; (f) standard errors of estimates β̂1(sk) in tibial cartilage; (g) estimates of β2(sk) in tibial cartilage; (h) standard errors of estimates β̂2(sk) in tibial cartilage; (i) FDR adjusted −log10(p)-values for test (19) across all pixel sites in femoral cartilage; and (j) FDR adjusted −log10(p)-values for test (19) across all pixel sites in tibial cartilage.
TABLE III.
Inference results of α̂ based on GHMM
| Femoral cartilage | Tibial cartilage | |||||
|---|---|---|---|---|---|---|
| Parameter | Estimation | Standard Error | −log10(p)-value | Estimation | Standard Error | −log10(p)-value |
| ᾱ1(1) | −0.0104 | 0.0027 | 3.9721 | −0.0256 | 0.0080 | 2.8562 |
| ᾱ2(1) | −0.1374 | 0.0353 | 4.0128 | −0.1534 | 0.0405 | 3.8111 |
| ᾱ1(2) | −0.1253 | 0.0289 | 4.8366 | −0.1387 | 0.0362 | 3.9018 |
| ᾱ2(2) | −0.2619 | 0.0628 | 4.5107 | −0.2328 | 0.0584 | 4.1733 |
Fig. 9.
Dynamic changes of cartilage thickness at randomly selected pixel sites across time: (a) two randomly selected pixel sites on femoral cartilage and tibial cartilage; (b) dynamic changes of cartilage thickness at sk1 of subjects with bi(sk1) = 0 across time, i.e., estimated curve (solid line) and chosen subjects with bi(sk1) = 0 (dashed lines); (c) dynamic changes of cartilage thickness at sk2 of subjects with bi(sk2) = 0 across time, i.e., estimated curve (solid line) and chosen subjects with bi(sk2) = 0 (dashed lines); (d) dynamic changes of cartilage thickness at sk1 of OA patients with bi(sk1) = 1 across time, i.e., estimated curve (solid line) and chosen OA patients with bi(sk1) = 1 (dashed lines); (e) dynamic changes of cartilage thickness at sk2 of OA patients with bi(sk2) = 1 across time, i.e., estimated curve (solid line) and chosen OA patients with bi(sk2) = 1 (dashed lines); (f) dynamic changes of cartilage thickness at sk1 of OA patients with bi(sk1) = 2 across time, i.e., estimated curve (solid line) and chosen OA patients with bi(sk1) = 2 (dashed lines); and (g) dynamic changes of cartilage thickness at sk2 of OA patients with bi(sk2) = 2 across time, i.e., estimated curve (solid line) and chosen OA patients with bi(sk2) = 2 (dashed lines).
Fig. 10.

Diseased region detection for subjects with different KLGs, including 1 KLG1, 1 KLG2, and 1 KLG3. Detection results of femoral cartilage thickness map (top) and tibial cartilage thickness map (bottom) for every subject are shown in each row, i.e., normal region (light blue); moderately diseased region (yellow); severely diseased region (orange).
Fig. 11.
Empirical probabilities of both moderately diseased region and severely diseased region based on estimated diseased region labels: (a) empirical probability of moderately diseased region in femoral cartilage; (b) empirical probability of severely diseased region in femoral cartilage; (c) empirical probability of moderately diseased region in tibial cartilage; and (d) empirical probability of severely diseased region in tibial cartilage.
Furthermore, we investigate whether there are some subgroups among patients and delineate their associated diseased region pattern in each subgroup. To achieve this, we used a clustering method to cluster all estimated maps {b̂i}i≥1 at the subject level. Since the number of informative pixels is much larger than the number of OA subjects, e.g., m = 4484 and n = 74 for tibial cartilage, we adopted a sparse k-means clustering method and its associated R package sparcl (http://www.icesi.edu.co/CRAN/web/packages/sparcl/) [55], which combines a Lasso-type penalty with the K-means clustering method to cluster observations. We applied sparcl to the estimated maps {b̂i}i≥1. Two subgroups were detected for the femoral cartilage, whereas three subgroups were identified for the tibial cartilage. The KLG information of subjects in each subgroup is presented in Table IV. Inspecting Table IV reveals that for the femoral cartilage case, the group membership of most patients is consistent with their KLG information. Specifically, most KLG2 subjects belong to Group 1, whereas most KLG3 subjects fall into Group 2. Similar to the procedures in voxel-based analysis, the diseased region pattern of each group in the average sense can also be calculated based on the detection results. For every group, the diseased-region detection results of two randomly chosen subjects and their related pattern are presented in Fig. A1, which can be found in Supplementary Materials. It reveals the spatial heterogeneity of cartilage progression across OA subjects.
TABLE IV.
KLG information of patients in each group
| Femoral cartilage | ||||
|---|---|---|---|---|
| Group | KLG1 | KLG2 | KLG3 | Total |
| 1 | 0 | 36 | 8 | 44 |
| 2 | 1 | 4 | 25 | 30 |
|
| ||||
| Tibial cartilage | ||||
| Group | KLG1 | KLG2 | KLG3 | Total |
| 1 | 0 | 26 | 26 | 52 |
| 2 | 1 | 13 | 0 | 14 |
| 3 | 0 | 1 | 7 | 8 |
VI. Conclusion
We have developed a GHMM for longitudinal cartilage quantification in OA patients, while addressing these two issues: the lack of spatial correspondence across subjects and time and the spatial heterogeneity of cartilage progression across subjects. Our GHMM is designed to efficiently account for the spatial heterogeneity of cartilage progression across subjects. We have developed an efficient algorithm to estimate unknown parameters in the GHMM. The proposed model can effectively detect diseased regions in each OA subject and present a localized analysis of longitudinal cartilage thickness within each latent subpopulation. Our GHMM integrates the strengths of two standard statistical methods: the local subregion-based analysis and the ordered value approach. Simulation studies and real data analysis have demonstrated that GHMM can effectively detect diseased regions in localizing longitudinal cartilage thickness.
Several important issues need to be addressed in our future research. First, it is assumed that the disease effect bi is an ordinal variable at each location and follows a Potts model. However, if the disease effect is strongly associated with the continuous loss of cartilage thickness, then it may be more reasonable to treat them as continuous variables. In this case, computational and theoretical challenges may arise. For example, we may need to introduce another hidden Markov model for continuous disease effects, and thus both the estimation and inference procedures have to be rederived. Second, we plan to apply GHMM to other knee MRI datasets, such as OAI, to investigate other clinical questions of interest. Third, it is interesting to extend GHMM to the group analysis of various neuroimaging data sets, such as the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database [56]. Most group analysis methods (e.g., Statistical Parametric Mapping (SPM)) assume the spatial homogeneity and spatial correspondence of effect regions across subjects, which may not be valid for many neuroimaging studies. For instance, in Alzheimer’s disease (AD) studies, quantifying patterns of structural change during early stages of AD is a major challenge. Brain atrophy in the early stages of AD may be relatively subtle and spatially distributed over different brain regions, and the patterns of atrophy have been found spatially heterogeneous [57]. Moreover, the error associated with structural measurements can vary throughout the brain and across subjects and time points. In this case, our GHMM can be used to capture complex morphological characteristics that distinguish AD from normal controls, furthermore to determine which patterns are most informative from a diagnostic perspective.
Supplementary Material
Acknowledgments
We thank the Editor, the Associate Editor, and three anonymous referees for valuable suggestions, which greatly helped to improve our presentation.
This work was supported in part by NIH grants R025747-01, CA142538-01, MH086633, B005149-01, R01 MH091645-01A1, R21 AR059890, 5R21AR059890-02, and the Pfizer longitudinal study on osteoarthritis (PLS-A9001140).
Footnotes
According to the definition of KLG [53], the radiographic score is from 0–4, where KLG0 is healthy, KLG1 is considered borderline or mild OA, and KLG2-KLG4 are severe OA.
Contributor Information
Chao Huang, Department of Biostatistics, and Biomedical Research Imaging Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA.
Liang Shan, Department of Computer Sciences, and Biomedical Research Imaging Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA.
H. Cecil Charles, Department of Radiology, Duke University, Durham, NC, 27710 USA.
Wolfgang Wirth, Institute of Anatomy and Musculoskeletal Research, Paracelsus Medical University, Strubergasse 21, 5020, Salzburg, Austria, and Chondrometrics GmbH, Ainring, Germany.
Marc Niethammer, Department of Computer Sciences, and Biomedical Research Imaging Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA.
Hongtu Zhu, Department of Biostatistics, and Biomedical Research Imaging Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA.
References
- 1.Arthritis related statistics. www.cdc.gov/arthritis/datastatistics/arthritisrelatedstats.htm, download date: October 2013.
- 2.Woolf AD, Pfleger B. Burden of major musculoskeletal conditions. Bulletin of the World Health Organization. 2003;81(9):646–656. [PMC free article] [PubMed] [Google Scholar]
- 3.Felson DT, Lawrence RC, Dieppe PA, Hirsch R, Helmick CG, Jordan JM, Kington RS, Lane NE, Nevitt MC, Zhang Y, et al. Osteoarthritis: new insights. part 1: the disease and its risk factors. Annals of internal medicine. 2000;133(8):635–646. doi: 10.7326/0003-4819-133-8-200010170-00016. [DOI] [PubMed] [Google Scholar]
- 4.Cicuttini F, Hankin J, Jones G, Wluka A. Comparison of conventional standing knee radiographs and magnetic resonance imaging in assessing progression of tibiofemoral joint osteoarthritis. Osteoarthritis and cartilage. 2005;13(8):722–727. doi: 10.1016/j.joca.2005.04.009. [DOI] [PubMed] [Google Scholar]
- 5.Raynauld JP. Quantitative magnetic resonance imaging of articular cartilage in knee osteoarthritis. Current opinion in rheumatology. 2003;15(5):647–650. doi: 10.1097/00002281-200309000-00021. [DOI] [PubMed] [Google Scholar]
- 6.Biswal S, Hastie T, Andriacchi TP, Bergman GA, Dillingham MF, Lang P. Risk factors for progressive cartilage loss in the knee. Arthritis & Rheumatism. 2002;46(11):2884–2892. doi: 10.1002/art.10573. [DOI] [PubMed] [Google Scholar]
- 7.Buck RJ, Dreher D, Eckstein F. Femorotibial cartilage thickness change distributions for subjects without signs, symptoms, or risk factors of knee osteoarthritis. Cartilage. 2012;3(4):305–313. doi: 10.1177/1947603511430326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Eckstein F, Buck RJ, Wyman BT, Kotyk JJ, Graverand L, Hellio MP, Remmers AE, Evelhoch JL, Hudelmaier M, Charles HC. Quantitative imaging of cartilage morphology at 3.0 tesla in the presence of gadopentate dimeglumine (gd-dtpa) Magnetic Resonance in Medicine. 2007;58(2):402–406. doi: 10.1002/mrm.21290. [DOI] [PubMed] [Google Scholar]
- 9.Felson DT, Niu J, Yang T, Torner J, Lewis CE, Aliabadi P, Sack B, Sharma L, Guermazi A, Goggins J, et al. Physical activity, alignment and knee osteoarthritis: data from most and the oai. Osteoarthritis and Cartilage. 2013;21(6):789–795. doi: 10.1016/j.joca.2013.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Eckstein F, Wirth W, Nevitt MC. Recent advances in osteoarthritis imaginglthe osteoarthritis initiative. Nature Reviews Rheumatology. 2012;8(10):622–630. doi: 10.1038/nrrheum.2012.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Biswal S, Hastie T, Andriacchi TP, Bergman GA, Dillingham MF, Lang P. Risk factors for progressive cartilage loss in the knee. Arthritis & Rheumatism. 2002;46(11):2884–2892. doi: 10.1002/art.10573. [DOI] [PubMed] [Google Scholar]
- 12.Moisio K, Chang A, Eckstein F, Chmiel JS, Wirth W, Almagor O, Prasad P, Cahue S, Kothari A, Sharma L. Varus–valgus alignment: reduced risk of subsequent cartilage loss in the less loaded compartment. Arthritis & Rheumatism. 2011;63(4):1002–1009. doi: 10.1002/art.30216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chang A, Moisio K, Chmiel JS, Eckstein F, Guermazi A, Almagor O, Cahue S, Wirth W, Prasad P, Sharma L. Subregional effects of meniscal tears on cartilage loss over 2 years in knee osteoarthritis. Annals of the rheumatic diseases. 2011;70(1):74–79. doi: 10.1136/ard.2010.130278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Seo SS, Kim CW, Jung DW. Management of focal chondral lesion in the knee joint. Knee surgery & related research. 2011;23(4):185–196. doi: 10.5792/ksrr.2011.23.4.185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wirth W, Eckstein F. A technique for regional analysis of femorotibial cartilage thickness based on quantitative magnetic resonance imaging. Medical Imaging, IEEE Transactions on. 2008;27(6):737–744. doi: 10.1109/TMI.2007.907323. [DOI] [PubMed] [Google Scholar]
- 16.Jørgensen DR, Dam EB, Lillholm M. Predicting knee cartilage loss using adaptive partitioning of cartilage thickness maps. Computers in biology and medicine. 2013;43(8):1045–1052. doi: 10.1016/j.compbiomed.2013.05.012. [DOI] [PubMed] [Google Scholar]
- 17.Buck RJ, Wyman BT, Hudelmaier M, Wirth W, Eckstein F, et al. Does the use of ordered values of subregional change in cartilage thickness improve the detection of disease progression in longitudinal studies of osteoarthritis? Arthritis Care & Research. 2009;61(7):917–924. doi: 10.1002/art.24613. [DOI] [PubMed] [Google Scholar]
- 18.Besag J. Spatial interaction and the statistical analysis of lattice systems. Journal of the Royal Statistical Society. Series B (Methodological) 1974:192–236. [Google Scholar]
- 19.Geman S, Geman D. Stochastic relaxation, gibbs distributions, and the bayesian restoration of images. Pattern Analysis and Machine Intelligence, IEEE Transactions on. 1984;(6):721–741. doi: 10.1109/tpami.1984.4767596. [DOI] [PubMed] [Google Scholar]
- 20.Besag J. On the statistical analysis of dirty pictures. Journal of the Royal Statistical Society. Series B (Methodological) 1986:259–302. [Google Scholar]
- 21.Qian W, Titterington D. Estimation of parameters in hidden markov models. Philosophical Transactions of the Royal Society of London Series A: Physical and Engineering Sciences. 1991;337(1647):407–428. [Google Scholar]
- 22.Rajapakse JC, Giedd JN, Rapoport JL. Statistical approach to segmentation of single-channel cerebral mr images. Medical Imaging, IEEE Transactions on. 1997;16(2):176–186. doi: 10.1109/42.563663. [DOI] [PubMed] [Google Scholar]
- 23.Zhang Y, Brady M, Smith S. Segmentation of brain mr images through a hidden markov random field model and the expectation-maximization algorithm. Medical Imaging, IEEE Transactions on. 2001;20(1):45–57. doi: 10.1109/42.906424. [DOI] [PubMed] [Google Scholar]
- 24.Marroquin JL, Santana EA, Botello S. Hidden markov measure field models for image segmentation. Pattern Analysis and Machine Intelligence, IEEE Transactions on. 2003;25(11):1380–1387. [Google Scholar]
- 25.Zhu H, Gu M, Peterson B. Maximum likelihood from spatial random effects models via the stochastic approximation expectation maximization algorithm. Statistics and computing. 2007;17(2):163–177. [Google Scholar]
- 26.Diggle P, Ribeiro PJ. Model-based geostatistics. Springer; 2007. [Google Scholar]
- 27.Li SZ, Singh S. Markov random field modeling in image analysis. Vol. 26 Springer; 2009. [Google Scholar]
- 28.Piplani MA, Disler DG, McCauley TR, Holmes TJ, Cousins JP. Articular cartilage volume in the knee: semiautomated determination from three-dimensional reformations of mr images. Radiology. 1996;198(3):855–859. doi: 10.1148/radiology.198.3.8628883. [DOI] [PubMed] [Google Scholar]
- 29.Eckstein F, Gavazzeni A, Sittek H, Haubner M, Lösch A, Milz S, Englmeier KH, Schulte E, Putz R, Reiser M. Determination of knee joint cartilage thickness using three-dimensional magnetic resonance chondrocrassometry (3d mr-ccm) Magnetic Resonance in Medicine. 1996;36(2):256–265. doi: 10.1002/mrm.1910360213. [DOI] [PubMed] [Google Scholar]
- 30.Grau V, Mewes A, Alcaniz M, Kikinis R, Warfield SK. Improved watershed transform for medical image segmentation using prior information. Medical Imaging, IEEE Transactions on. 2004;23(4):447–458. doi: 10.1109/TMI.2004.824224. [DOI] [PubMed] [Google Scholar]
- 31.Gougoutas AJ, Wheaton AJ, Borthakur A, Shapiro EM, Kneeland JB, Udupa JK, Reddy R. Cartilage volume quantification via live wire segmentation. Academic radiology. 2004;11(12):1389–1395. doi: 10.1016/j.acra.2004.09.003. [DOI] [PubMed] [Google Scholar]
- 32.Lynch JA, Zaim S, Zhao J, Stork A, Peterfy CG, Genant HK. Medical Imaging 2000. International Society for Optics and Photonics; 2000. Cartilage segmentation of 3d mri scans of the osteoarthritic knee combining user knowledge and active contours; pp. 925–935. [Google Scholar]
- 33.Brem M, Lang P, Neumann G, Schlechtweg P, Schneider E, Jackson R, Yu J, Eaton C, Hennig F, Yoshioka H, et al. Magnetic resonance image segmentation using semi-automated software for quantification of knee articular cartilagelinitial evaluation of a technique for paired scans. Skeletal radiology. 2009;38(5):505–511. doi: 10.1007/s00256-009-0658-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bae K, Shim H, Tao C, Chang S, Wang J, Boudreau R, Kwoh C. Intra-and inter-observer reproducibility of volume measurement of knee cartilage segmented from the oai mr image set using a novel semiautomated segmentation method. Osteoarthritis and Cartilage. 2009;17(12):1589–1597. doi: 10.1016/j.joca.2009.06.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Koo S, Hargreaves BA, Gold GE. uS Patent 8,706,188. Automatic segmentation of articular cartilage from mri. 2014 Apr 22;
- 36.Vincent G, Wolstenholme C, Scott I, Bowes M. Fully automatic segmentation of the knee joint using active appearance models. Medical Image Analysis for the Clinic: A Grand Challenge. 2010:224–230. [Google Scholar]
- 37.Yin Y, Zhang X, Williams R, Wu X, Anderson DD, Sonka M. Logismosllayered optimal graph image segmentation of multiple objects and surfaces: cartilage segmentation in the knee joint. Medical Imaging, IEEE Transactions on. 2010;29(12):2023–2037. doi: 10.1109/TMI.2010.2058861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Shan L, Zach C, Charles C, Niethammer M. Automatic atlas-based three-label cartilage segmentation from MR knee images. Medical Image Analysis. 2014;18:1233–1246. doi: 10.1016/j.media.2014.05.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Shan L, Charles C, Niethammer M. Automatic multi-atlas-based cartilage segmentation from knee mr images. Biomedical Imaging (ISBI), 2012 9th IEEE International Symposium on; IEEE; 2012. pp. 1028–1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yezzi AJ, Prince JL. An eulerian pde approach for computing tissue thickness. Medical Imaging, IEEE Transactions on. 2003;22(10):1332–1339. doi: 10.1109/TMI.2003.817775. [DOI] [PubMed] [Google Scholar]
- 41.Stahl R, Jain SK, Lutz J, Wyman BT, Le Graverand-Gastineau MPH, Vignon E, Majumdar S, Link TM. Osteoarthritis of the knee at 3.0 t: comparison of a quantitative and a semi-quantitative score for the assessment of the extent of cartilage lesion and bone marrow edema pattern in a 24-month longitudinal study. Skeletal radiology. 2011;40(10):1315–1327. doi: 10.1007/s00256-011-1156-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dempster AP, Laird NM, Rubin DB. Maximum likelihood from incomplete data via the em algorithm. Journal of the Royal Statistical Society. Series B (Methodological) 1977:1–38. [Google Scholar]
- 43.Geman S, Graffigne C. Proceedings of the International Congress of Mathematicians. Vol. 1. AMS; Providence, RI: 1986. Markov random field image models and their applications to computer vision; p. 2. [Google Scholar]
- 44.Marroquín JL, Vemuri BC, Botello S, Calderon E, Fernandez-Bouzas A. An accurate and efficient bayesian method for automatic segmentation of brain mri. Medical Imaging, IEEE Transactions on. 2002;21(8):934–945. doi: 10.1109/TMI.2002.803119. [DOI] [PubMed] [Google Scholar]
- 45.Nie J, Xue Z, Liu T, Young GS, Setayesh K, Guo L, Wong ST. Automated brain tumor segmentation using spatial accuracy-weighted hidden markov random field. Computerized Medical Imaging and Graphics. 2009;33(6):431–441. doi: 10.1016/j.compmedimag.2009.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wu C-FJ. Jackknife, bootstrap and other resampling methods in regression analysis. the Annals of Statistics. 1986:1261–1295. [Google Scholar]
- 47.Efron B, Tibshirani RJ. An introduction to the bootstrap. Vol. 57 CRC press; 1994. [Google Scholar]
- 48.Yekutieli D, Benjamini Y. Resampling-based false discovery rate controlling multiple test procedures for correlated test statistics. Journal of Statistical Planning and Inference. 1999;82(1):171–196. [Google Scholar]
- 49.Jaccard P. Etude comparative de la distribution florale dans une portion des Alpes et du Jura. Impr Corbaz. 1901 [Google Scholar]
- 50.Dice LR. Measures of the amount of ecologic association between species. Ecology. 1945;26(3):297–302. [Google Scholar]
- 51.Rand WM. Objective criteria for the evaluation of clustering methods. Journal of the American Statistical association. 1971;66(336):846–850. [Google Scholar]
- 52.Hubert L, Arabie P. Comparing partitions. Journal of classification. 1985;2(1):193–218. [Google Scholar]
- 53.Kellegren J, Lawrence J. Radiological assessment of osteoarthritis. Ann Rheum Dis. 1957;16:494–501. doi: 10.1136/ard.16.4.494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.MacQueen J, et al. Some methods for classification and analysis of multivariate observations. Proceedings of the fifth Berkeley symposium on mathematical statistics and probability; California, USA. 1967; pp. 281–297. [Google Scholar]
- 55.Witten DM, Tibshirani R. A framework for feature selection in clustering. Journal of the American Statistical Association. 2010;105(490) doi: 10.1198/jasa.2010.tm09415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Mueller SG, Weiner MW, Thal LJ, Petersen RC, Jack C, Jagust W, Trojanowski JQ, Toga AW, Beckett L. The alzheimer’s disease neuroimaging initiative. Neuroimaging Clinics of North America. 2005;15(4):869–877. doi: 10.1016/j.nic.2005.09.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Frisoni G, Sabattoli F, Lee A, Dutton R, Toga A, Thompson P. In vivo neuropathology of the hippocampal formation in AD: a radial mapping MR-based study. Neuroimage. 2006;32(1):104–110. doi: 10.1016/j.neuroimage.2006.03.015. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.