Abstract
The agreed biological function of the casein micelles in milk is to carry minerals (calcium, magnesium, and phosphorus) from mother to young along with amino acids for growth and development. Recently, native and modified casein micelles were used as encapsulating and delivery agents for various hydrophobic low-molecular-weight probes. The ability of modified casein micelles to bind certain probes may derive from the binding affinity of native casein micelles. Hence, a study with milk from single cows was conducted to further elucidate the association of hydrophobic molecules into native casein micelles and further understand their biological function. Hydrophobic and hydrophilic extraction followed by ultraperformance liquid chromatography-high resolution mass spectrometry analysis were performed over protein fractions obtained from size exclusion fractionation of raw skim milk. Hydrophobic compounds, including phosphatidylcholine, lyso-phosphatidylcholine, phosphatidylethanolamine, and sphingomyelin, showed strong association exclusively to casein micelles as compared with whey proteins, whereas hydrophilic compounds did not display any preference for their association among milk proteins. Further analysis using liquid chromatography-tandem mass spectrometry detected 42 compounds associated solely with the casein-micelles fraction. Mass fragments in tandem mass spectrometry identified 4 of these compounds as phosphatidylcholine with fatty acid composition of 16:0/18:1, 14:0/16:0, 16:0/16:0, and 18:1/18:0. These results support that transporting low-molecular-weight hydrophobic molecules is also a biological function of the casein micelles in milk.
Keywords: casein micelle, liquid chromatography-mass spectrometry, binding, hydrophobicity, sphingomyelin
INTRODUCTION
Bovine milk is a good source of protein comprising ca 3.6% of the total composition of the milk. About 80% of the total protein content in bovine milk consist of 4 types of casein, which are αs1-, αs2-, β-, and κ-CN in the approximate ratio of 4:1:3.5:1.5 (by weight). Caseins are phosphoproteins that exist as colloidal aggregates known as casein micelles (Horne, 1998; De Kruif and Holt, 2003; Dalgleish, 2011). Although the casein micelle structure is still under investigation, certain models have been proposed (De Kruif and Holt, 2003; Horne, 2006). The current consensus is that proteins in casein micelles are stabilized by various hydrophobic and electrostatic interactions, whereas calcium phosphate clusters bind to the phosphoseryl residues found on αs- and β-CN. The localization of κ-CN to the surface of the casein micelle to act as negatively charged hairs plays an important role in stabilizing adjacent casein micelles against aggregation. Recent electron microscopy observations suggest a sponge-like structure with native casein micelles containing internal serum-filled channels and voids that allow a dynamic interaction between the micelles and their surroundings (Trejo et al., 2011).
The agreed biological function of the casein micelle is to transport minerals (calcium, phosphorous) as well as amino acids from mother to newborns for their growth and development (De Kruif and Holt, 2003). This hypothesis is supported by the fact that milk is supersaturated with calcium, and phospho-serine residues are conserved in mammalian species (Oftedal, 2013).
Evidence exists in the literature for the use of casein micelles as delivery agents for hydrophobic molecules. For instance, reformed casein micelles formed by exposing commercially available skim milk to high-pressure homogenization were employed to encapsulate hydrophobic, antimicrobial triclosan (Roach and Harte, 2008). Reassembled casein micelles formed by treatment with potassium citrate bound hydrophobic vitamin D2 (Semo et al., 2007). Alternatively, native casein micelles separated from raw skim milk by ultracentrifugation were used as encapsulating agents for carrying and delivering curcumin, a natural spice with potential therapeutic properties, to cancer cells (Sahu et al., 2008). The formation of the complex between bovine casein micelles and curcumin was attributed to hydrophobic interactions. Our previous work conducted on native casein micelles in ultrapasteurized skim milk purified by size-exclusion chromatography (SEC) provides further evidence for the natural ability of casein to associate with hydrophobic vitamin A as compared with whey proteins (Mohan et al., 2013). It was also found that the amount of hydrophobic phospholipids associated with casein proteins was higher as compared with the whole milk (Cerbulis and Zittle, 1965).
Despite increasing evidence, the affinity of native casein micelles toward hydrophobic low-molecular-weight molecules has not yet been investigated thoroughly, most likely because of the belief that the milk fat globule is the appropriate system to transport hydrophobic compounds from mother to young. In addressing this knowledge gap, the objective of this study was to investigate and compare the capacity of casein micelles relative to other significant proteins in raw bovine milk (i.e., BSA, α-LA, β-LG) to associate with hydrophobic and hydrophilic compounds. Milk protein fractions purified by SEC were analyzed by ultraperformance liquid chromatography-high-resolution mass spectrometry (UPLC-HRMS) to compare the association of hydrophobic and hydrophilic compounds with each major milk protein fraction (i.e., casein micelle, BSA, and whey protein). Further analysis using liquid chromatography-tandem mass spectrometry (LC-MS/MS) was performed to identify the hydrophobic compounds associated exclusively with casein micelles by analyzing the corresponding mass fragments of parent ions. This study provides detailed information on associations between the casein micelle and hydrophobic molecules. Understanding fundamental structure-function properties of the casein micelle in milk is key to the development of future applications, including encapsulation and delivery systems.
MATERIALS AND METHODS
Milk Sample Collection and Preparation
Raw milk samples were collected from 6 Holstein cows of first parity at mid-lactation stage from the University of Tennessee Dairy Research and Education Center. The raw milk samples were collected directly from each cow in the milking apparatus without any mixing followed by transportation in ice and storage at 4°C. The fat removal from the raw milk was done by centrifugation at 4°C (6,414 × g for 20 min) followed by storage at 4°C. Tangential flow ultrafiltration was employed to generate protein-free serum (PFS) from the raw milk of corresponding cow using a 3KDa MWCO cellulose filter (PLBC Prep scale TFF Cartridge, Millipore, Billerica, MA). Protein-free serum was preserved in 0.07% NaN3 to prevent microbial growth and was stored at 4°C until SEC was performed to separate the proteins in the milk sample.
SEC
Casein micelle, BSA, β-LG, and α-LA fractions from raw milk were separated by SEC using a Superdex column (Superdex 200 prep grade, GE Life Sciences, Piscataway, NJ) connected to an AKTA fast-performance liquid chromatography unit (GE Life Sciences). The raw skim milk samples were loaded on the column and run using ice-cold PFS generated from the milk samples of the corresponding cow as mobile phase. The use of PFS during protein separation ensured that proteins were kept in their native state. The proteins were eluted using a 1.0 mL/min flow rate for 2 column volumes (240 mL total) and collecting 4-mL fractions. Protein elution was monitored using absorbance with a UV detector at 280 nm wavelength. Fractions were collected based on elution peaks corresponding to casein micelles, BSA, and β-LG and α-LA (based on SDS-PAGE analysis, see below), yielding 4 samples along with PFS as control. These samples were freeze-dried and stored at −40°C until liquid chromatography-mass spectrometry analysis.
SDS-PAGE
Sodium dodecyl sulfate-PAGE was performed on all 4 SEC fractions (CN, BSA, β-LG and α-LG, PFS) to verify the purity of the samples and also to confirm that PFS did not contain proteins (Figure 1). Electrophoresis was run under denaturing and reducing conditions. The sample loading buffer was prepared using 0.5 M Tris-HCl at pH 6.8, glycerol, 10% (wt/vol) SDS, β-mercaptoethanol, and 0.5% (wt/vol) bromophenol blue in water (Fisher Scientific, Fair Lawn, NJ). The samples were suspended in loading buffer and heat denatured for 5 min at 95°C before electrophoresis in 15% acrylamide SDS-PAGE ready gels (Bio-Rad, Hercules, CA). Sample volumes of 15 µL were loaded into each well, and electrophoresis was run at constant 200 V for 35 min. After electrophoretic separation, proteins were detected in gels using Coomassie blue staining.
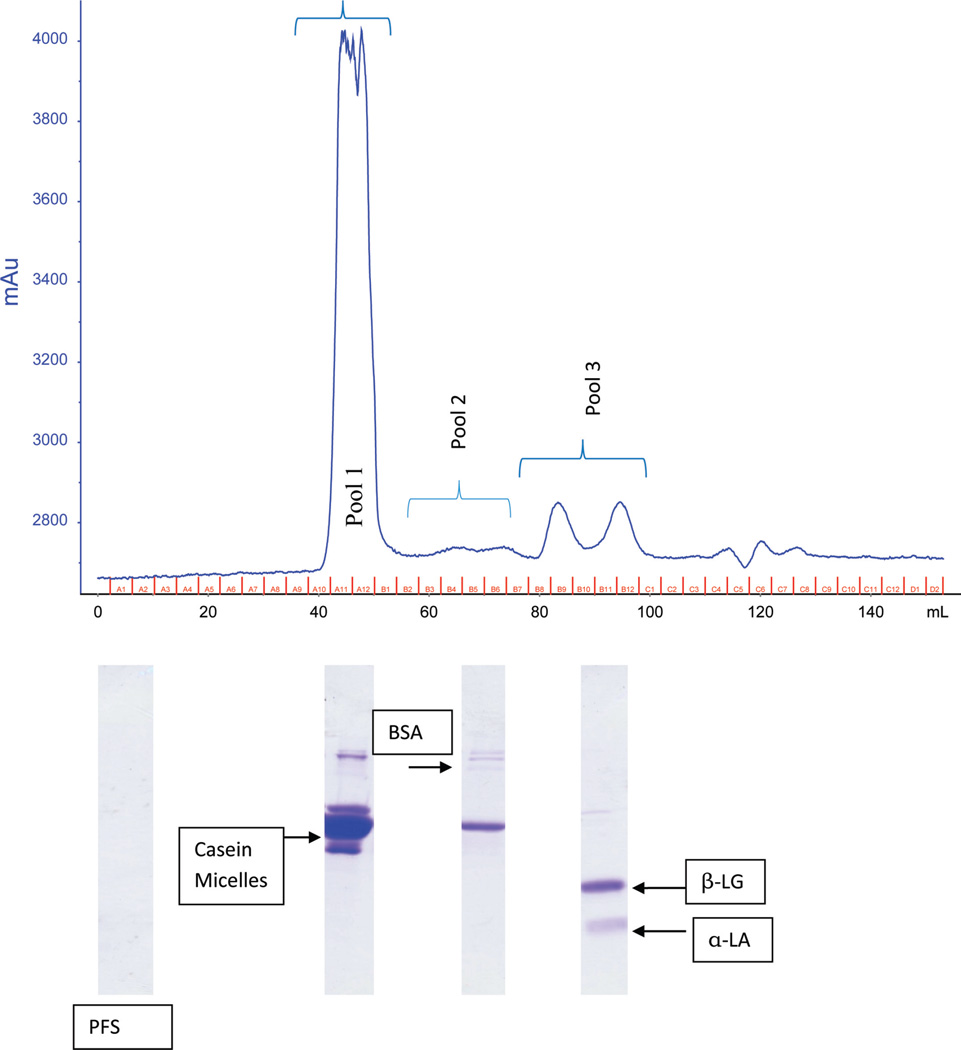
Figure 1.
Representative size-exclusion chromatogram and SDS-PAGE of protein fractions from bovine raw skim milk. PFS = protein-free serum. Letters A1–D2 indicate the FPLC fractions (2 mL each). Color version available online.
Sample Extraction for Hydrophobic Compounds and Preparation
The extraction of hydrophobic compounds was done following the method used by Bligh and Dyer (De Kruif, 1999). Briefly, 5 mg of the freeze-dried samples (by total weight) was extracted using 1 mL of the extraction solvent (chloroform-methanol-water, 1:2:0.8) followed by vortexing and centrifugation at 16,000 × g for 5 min at 4°C. The supernatant was transferred into a separate glass vial followed by repetition of the extraction procedure twice more. Each time, the resultant supernatant was transferred to the same glass vial. The combined supernatant was dried under nitrogen stream and finally dissolved in 3 mL of methanol to be analyzed using liquid chromatography-mass spectrometry.
UPLC-HRMS Analysis for Hydrophobic Compounds
Analysis by UPLC-HRMS was performed using a hydrophilic interaction chromatography column (150 × 2.1 mm, 2.6 µm; Phenomenex, Torrance, CA) equipped with ultimate 3000, Orbitrap Exactive plus LC pump (Thermo Fisher Scientific, Waltham, MA). All solvents and ammonium formate were purchased from Fisher Scientific (Atlanta, GA). The mobile phases were solvent A (10 mM aqueous ammonium formate pH 3) and solvent B (93/7 vol/vol: acetonitrile/142.86 mM ammonium formate pH 3). The gradient dilution used was 0 to 1 min 100% B, 1 to 15 min 100 to 81% B, 15.1 to 25 min 48% B, 25.1 to 35 min 100% B with a flow rate of 0.2 mL/min and 25°C column temperature. The injection volume used was 10 µL. The electrospray ionization mode was used along with Exactive Plus Orbitrap quadrupole mass spectrometer (Thermo Fisher Scientific). The analysis for hydrophobic compounds was performed in positive mode.
LC-MS/MS Analysis for Hydrophobic Compounds
Tandem mass spectrometry analysis was performed using a HPLC-QTOFMS Prominence UFLC XR (Shimadzu, Columbia, MD) and an AB Sciex 5600 quadrupole TOF mass spectrometer (Framingham, MA). The samples (5 µL) were separated using C18 column (100 × 2.1 mm, 1.7 µm (Waters, Milford, MA). The mobile phases were solvent A (60% acetonitrile with 40% water containing 10 mM ammonium formate, 0.1% formic acid) and solvent B (90% isopropanol, 10% acetonitrile 10 mM ammonium formate, 0.1% formic acid). The gradient dilution used was (min:s) 0:00 to 0:01 min 40% B, 0:01 to 2:00 min 40 to 43% B, 2:00 to 2:10 min 43 to 50% B, 2:10 to 12:00 min 50 to 54% B, 12:00 to 12:10 min 54 to 70% B, 12:10 to 18:00 min 70 to 99% B, 18:00 to 20:00 min 99% B, 20:00 to 20:10 min 99 to 40% B with a flow rate of 0.225 mL/min. Positive ion electrospray ionization mass spectra was acquired over the mass range 100 to 1,200 m/z in information-dependent-acquisition mode with one 100-ms survey scan and up to twenty 100-ms MS/MS product ion scans per duty cycle.
Sample Extraction for Hydrophilic Compounds and Preparation
The extraction of hydrophilic compounds was done by adding 0.5 mL of extraction solvent (acetonitrile:methanol:water 40:40:20 with 0.1 M formic acid) to 5 mg of freeze-dried sample (by total weight) followed by vortexing and centrifugation at 16,000 × g for 5 min at 4°C. The resultant supernatant was transferred to a glass vial. The extraction was repeated twice followed by a second extraction with 0.5 mL of extraction solvent (methanol:water 80:20), which was also repeated twice. The combined supernatant from both extractions was dried under nitrogen stream. The dried sample was dissolved in 3 mL of methanol to be analyzed using liquid chromatography-mass spectrometry.
UPLC-HRMS Analysis for Hydrophilic Compounds
A 10-µL aliquot was injected through a Synergi 2.5 µm Hydro-RP 100, 100 × 2.00 mm LC column (Phenomenex) kept at 25°C. The eluent was introduced into the mass spectrometer via an electrospray ionization source conjoined to a Thermo Scientific Exactive plus Orbitrap mass spectrometer (Thermo Fisher Scientific) through a 0.1-mm internal diameter fused silica capillary tube. The mass spectrometer was run in full scan mode with negative ionization with a method (Lu et al., 2010). The samples were analyzed with a resolution of 140,000 and a scan window of 85 to 800 m/z for from 0 to 9 min and 110 to 1,000 m/z from 9 to 25 min. Solvent A consisted of 97:3 water:methanol, 10 mM tributylamine, and 15 mM acetic acid. Solvent B was methanol. The gradient from 0 to 5 min is 0% B, from 5 to 13 min is 20% B, from 13 to 15.5 min is 55% B, from 15.5 to 19 min is 95% B, and from 19 to 25 min is 0% B with a flow rate of 200 µL/min.
Data Analysis
Files generated by the Xcalibur software (Thermo Fisher Scientific) were converted to the open-source mzML format (Martens et al., 2011) via the open-source mass spectrometer–convert software as part of the ProteoWizard package (Chambers et al., 2012). Maven (mzroll) software (Clasquin et al., 2002; Melamud et al., 2010) was used to automatically correct the total ion chromatograms based on the retention times for each sample. Unknown peaks were automatically selected via Maven’s automated peak detection algorithms. The data were transformed and clustered using Cluster software (Eisen Lab, Berkeley, CA). Hierarchical clustering was used to assemble a set of mass/charge ion in terms of similarity using Pearson correlation. Heat maps were then generated using Java Treeview (Saldanha, 2004) software. Peak view software (AB Sciex) was used to identify the fragments from MS/MS data, and Marker view software (AB Sciex) was used to perform statistical analysis; that is, pair-wise Student t-test to compare peak intensities of hydrophobic compounds associated with casein fractions versus whey proteins (BSA, β-LG and α-LA).
Enumeration of Microorganisms
Freeze-dried casein protein fraction and PFS [5 mg each (by total weight)] from all 6 cows were dissolved in 1 mL of 0.1% peptone water and plated onto trypticase soy agar (Thermo Fisher Scientific) plates, using 0.1 mL of suspension per plate and testing in duplicate. After incubation at 37°C for 48 h, the number of colonies observed was low, and no serial dilutions were needed. The numbers of colonies per plate were counted and are reported as log cfu/5 mg of the casein and PFS samples for each of the 6 cows.
RESULTS AND DISCUSSION
SEC
A representative chromatogram obtained from SEC purification of raw skim milk showed the presence of 4 major peaks eluted in order of decreasing size (Figure 1). The fractions included in each peak were pooled together, freeze-dried, and stored at −40°C for further analysis. Sodium dodecyl sulfate-PAGE gel electrophoresis was performed on these fractions to verify the presence of corresponding proteins and also most importantly to confirm the absence of any proteins in PFS, obtained from 3-kDa molecular weight cut-off filtration of raw skim milk. Based on the molecular weight of the protein bands and previous reports (Mohan et al., 2013), proteins in fraction pool 1 were identified as casein proteins, pool 2 contained BSA protein, and the third pool consisted of the 2 major whey proteins (β-LG and α-LG).
Enumeration of Microorganisms
Microbial counts were done on casein and PFS fractions to ensure absence of microbial growth in the fast-performance liquid chromatography column and SEC fractions, because significant microbial growth could lead to false peak intensities in subsequent liquid chromatograph-mass spectrometry metabolomics studies. The plate counts showed nondetectable levels of microbial growth in samples from all 6 cows in the various protein and PFS fractions tested, confirming that the peak areas for hydrophobic and hydrophilic compounds detected in liquid chromatograph-mass spectrometry would correspond to molecules and not to microbes present in the fractions.
Analytical UPLC-HRMS Analysis and Generation of Heat Maps
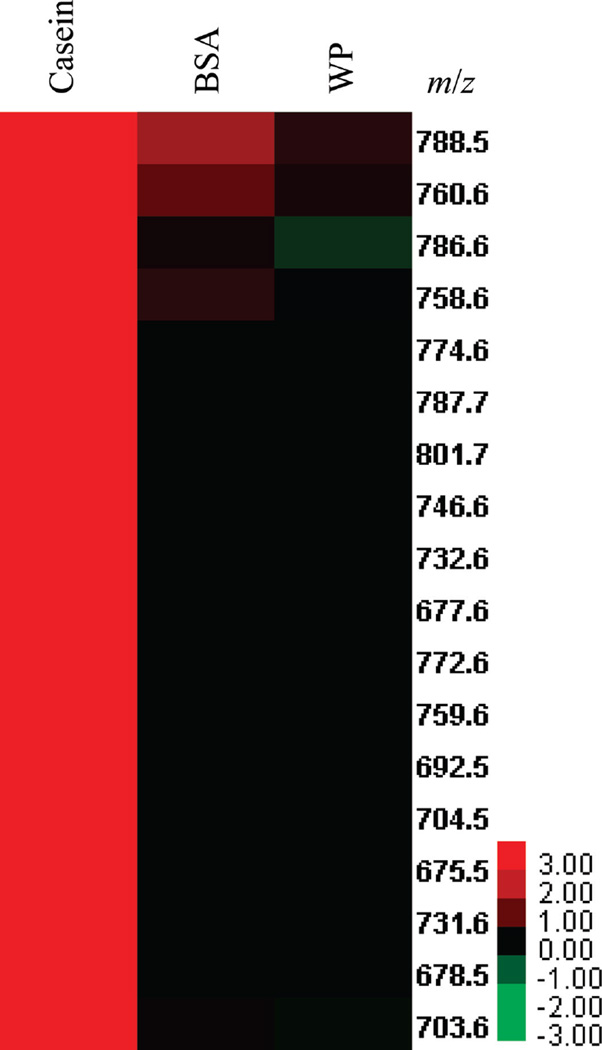
The UPLC-HRMS analysis for hydrophobic compounds was performed with the 3 fraction pools obtained from SEC purification of raw skim milk from 6 cows of the same lactation period. Analyzed pooled fractions were casein (pool 1), BSA (pool 2), whey protein (β-LG and α-LG; pool 3), and PFS (control). To understand the affinities of hydrophobic compounds for casein proteins, BSA, whey proteins, and PFS, heat maps were generated corresponding to the peak areas obtained for various hydrophobic compounds in each corresponding SEC fraction pool. The heat map was generated using peak areas calculated by taking the averages over cows 1 to 6 corresponding to mass/charge ratios (m/z), as shown in Figures 2 and 3. A wide range of m/z for hydrophobic species with 100- to 1,000-fold increase in peak areas was observed for casein fraction versus whey protein or control sample (Figure 2), supporting the hypothesis that native casein micelles bind hydrophobic compounds in raw bovine milk. Certain compounds showing high enrichment in casein compared with other pooled fractions were phospholipids tentatively identified based on literature (Sanchez-Juanes et al., 2009; Donato et al., 2011; Zhao et al., 2011) and the online database LIPID MAPS (http://www.lipidmaps.org/data/structure/index.html) as phosphatidylcholine (PC), phosphatidylethanolamines, and sphingomyelins (SM), as shown in Table 1.
Figure 2.
Mass/charge ratio of hydrophobic molecules detected (heat map) in the various protein pooled fractions obtained from size-exclusion chromatography of raw skim milk. Color code indicates log fold of average peak signal from fraction versus the average peak signal from protein-free serum, e.g., 3.00 (red) = log 1,000 indicates a 1,000-times stronger average peak signal for the casein fraction versus protein-free serum. WP = whey protein (β-LG + α-LA). Color version available online.
Figure 3.
Mass/charge ratio of hydrophilic molecules detected (heat map) in the various protein pooled fractions obtained from sizeexclusion chromatography of raw skim milk. Color code indicates log fold of average peak signal from fraction versus the average peak signal from protein-free serum, e.g., 3.00 (red) = log 1,000 indicates a 1,000-times stronger average peak signal for the casein fraction versus protein-free serum. WP is whey protein (β-LG + α-LA). Color version available online.
Table 1.
Hydrophobic compounds tentatively identified as phospholipids, extracted exclusively from casein micelles
| Phospholipid class | [M+H]+ | Molecular formula |
Tentative molecular species |
|---|---|---|---|
| Phosphatidylcholine (PC) | 788.5 | C44H86NPO8 | PC(18:0/18:1) |
| 786.6 | C44H84NO8P | PC(18:1/18:1) | |
| 760.6 | C42H82NO8P | PC(16:0/18:1) | |
| 758.6 | C42H80NPO8 | PC(16:0/18:2) | |
| 732.6 | C40H78NPO8 | PC(16:0/16:1) | |
| 678.5 | C36H72NO8P | PC(14:0/14:0) | |
| 772.6 | C37H74NO8P | PC(13:0/22.2) | |
| Sphingomyelin (SM) | 787.7 | C45H91N2O6P | SM(d18:1/22:0) |
| 786.6 | C45H91N2O6P | SM(d18:0/22:1) | |
| 784.7 | C45H89N2O6P | SM(d18:1/22:1) | |
| 759.6 | C43H89N2O6P | SM(d18:1/20:0) | |
| 703.6 | C39H79N2O6P | SM(d18:0/16:1) | |
| 677.6 | C37H77N2O6P | SM(d18:0/14:0) | |
| 801.7 | C46H93N2O6P | SM(d18:1/23:0) | |
| 675.5 | C37H75N2O6P | SM(d18:1/14:0) | |
| Phosphatidylethanolamine (PE) | 746.6 | C41H80NO8P | PE(18:1/18:0) |
| 704.5 | C38H74NO8P | PE(18:1/15:0) | |
| 692.5 | C37H74NO8P | PE(16:0/16:0) | |
| 774.6 | C43H84NO8P | PE(18:1/20:0) | |
| 772.6 | C43H82NO8P | PE(24:1/14:1) |
The presence of hydrophobic compounds such as PC and SM associated with casein in raw bovine milk is in agreement with higher amounts of PC and SM in skim milk compared with whole milk (Zeisel et al., 2003). Also in support of associations between casein and phospholipids, Cerbulis and Zittle (1965) reported that the ratio of phospholipid to total lipid was greater in the casein protein fraction compared with whole milk. Considering these observations together with our results, we propose that hydrophobic compounds such as PC and SM present in skim milk preferably associate with casein micelles. This hypothesis is also supported by reports on the use of modified casein micelles to bind various hydrophobic probes.
Pooled fractions containing casein micelles contained similar levels of hydrophilic compounds as compared with whey proteins and control pools, as depicted by the hydrophilic compound heat map in Figure 3. Although ~3,000 unknown hydrophilic compounds were detected, heat maps generated from mean values of replications using samples from 6 cows did not detect higher levels of any specific hydrophilic compound in casein micelles compared with whey protein or PFS samples.
LC-MS/MS Analysis of Hydrophobic Compounds
To investigate the association of the casein micelles with hydrophobic compounds, the SEC-purified pooled fractions of casein and whey proteins were analyzed using LC-MS/MS. The results were averaged over milk samples obtained from 6 cows. The purpose was to analyze and identify hydrophobic compounds associated exclusively with the casein micelle fractions of the raw bovine milk.
Statistical analyses of the MS data showed 42 compounds exclusively associated with the casein protein pooled fraction (P < 0.05). The mass fragments of the monoisotopic m/z values were further investigated using peak view software (AB Sciex). Some of the compounds associated with casein micelles were identified as PC based on MS/MS analysis of the fragments (Table 2). The likely fatty acid composition was assigned based on the molecular [M+H]+ product ions observed in the MS/MS and compared with the LIPID MAPS online database (http://www.lipidmaps.org/data/structure/index.html) and the literature (Donato et al., 2011; Zhao et al., 2011). The protonated molecular ion m/z 760.6 was found to be PC with fatty acid composition of 16:0/18:1, with dominant ion m/z 184, the polar phosphocholine head group. The product ions at m/z 478 corresponded to [M+H-18:1], indicating the 18:1 fatty acid (m/z 282). Also the m/z 504 corresponded to loss of m/z 256, i.e., [M+H-16:0], indicating the presence of a 16:0 fatty acyl group. The protonated molecular ion m/z 734.6 with the fatty acyl composition 16:0/16:0 showed the main fragment ion m/z 184, and the product ions at m/z 478 corresponded to [M+H-16:0], indicating the presence of 16:0 fatty acyl groups. Also, the molecular ion m/z 706.5 [M+H]+ with the fatty acid composition 16:0/14:0, with base peak corresponding to dominant ion at m/z 184 and product ions at m/z 478 and m/z 450, corresponded to [M+H-14:0]+ and [M+H-16:0]+ species, respectively. The m/z 788.5 was identified as PC due to dominant m/z 184 with other fragment ions m/z 506 and 504 corresponding to 18:1 and 18:0 fatty acyl groups. The m/z 506 and 504 corresponded to loss of m/z of 282 [M+H-18:1]+ and loss of m/z 284 [M+H-18:0]+ fragments, respectively. There are other m/z values in Table 3 identified as PC based on the dominant ions m/z 184 and loss of m/z 59 corresponding to −N (CH3)3 groups, for example m/z 744.5 and 772.6, which show fragments of m/z 685 and 713 with loss of the [M+H−N (CH3)3]+ group from the phosphocholine head group. The fragments were in correlation with the values reported in literature (Zhao et al., 2011). The reported PC are known to be highly hydrophobic compounds with high logarithmic values of partition coefficient, i.e., log P-values greater than 5.00 (octanol/water; http://www.lipidmaps.org/data/structure/index.html). The hydrophobic compounds of m/z values of 760.6, 788.5, and 772.6 were identified as PC using the MS/MS. These results further identify highly hydrophobic compounds (such as PC) that are preferentially associated with casein micelles rather than to whey proteins in raw bovine milk.
Table 2.
Compounds identified associated exclusively with casein fractions
| Ion (m/z), PC [M+H]+1 |
Fatty acid | Molecular formula | Product ions in tandem mass spectrometry |
|---|---|---|---|
| 760.6 | PC(16:0/18:1) | C42H82NO8P | 184, 478, 504, 577, 760 |
| 706.5 | PC(16:0/14:0) | C38H76NO8P | 184, 706, 478, 450 |
| 734.6 | PC(16:0/16:0) | C40H80NO8P | 184, 478 |
| 788.5 | PC(18:1/18:0) | C44H86NO8P | 184, 788, 506, 504 |
| 744.5 | — | — | 184, 685, 744 |
| 772.6 | — | — | 184, 713, 772 |
| 798.5 | — | — | 184, 739, 798 |
m/z = mass/charge; PC = phosphatidylcholine.
Table 3.
The mass/charge (m/z) values of hydrophobic compounds exclusively associated with casein micelles (fraction 1) as compared with whey proteins (fraction 2 and 3)
|
m/z [M+H]+ |
P-value |
m/z [M+H]+ |
P-value |
|---|---|---|---|
| 760.6 | 2.37E-18 | 563.5511 | 0.0097 |
| 734.6 | 4.65E-06 | 230.2452 | 0.01 |
| 706.5 | 1.87E-05 | 284.3337 | 0.0102 |
| 744.5 | 0.00014 | 468.414 | 0.0111 |
| 741.5 | 0.00021 | 725.5547 | 0.0111 |
| 219.0 | 0.00058 | 381.3762 | 0.0118 |
| 782.6 | 0.00068 | 424.4352 | 0.0132 |
| 772.6 | 0.00251 | 103.1004 | 0.014 |
| 796.5 | 0.00435 | 798.544 | 0.016 |
| 761.6 | 0.00474 | 687.5487 | 0.0179 |
| 324.3 | 0.00886 | 297.1687 | 0.018 |
| 398.1 | 0.00888 | 465.2993 | 0.0196 |
| 235.1 | 0.00892 | 351.0981 | 0.021 |
| 659.9 | 0.00905 | 980.287 | 0.0246 |
| 371.3 | 0.00907 | 701.5095 | 0.0265 |
| 735.6 | 0.00913 | 174.1476 | 0.0318 |
| 788.5 | 0.00924 | 186.1837 | 0.032 |
| 366.3 | 0.00935 | 527.3993 | 0.0329 |
| 609.5 | 0.00938 | 214.0976 | 0.0355 |
| 589.6 | 0.00947 | 531.4139 | 0.0425 |
| 786.6 | 0.00956 | 355.077 | 0.0483 |
CONCLUSIONS
The main contribution of this study lies in the identification of hydrophobic compounds associated with casein micelles in their native form. As a consequence of using native conditions, emphasis was put on qualitative rather than quantitative analysis and identification of the compounds associated with the micelles. Our data demonstrate that hydrophobic compounds are preferentially associated with native casein micelles in raw bovine milk, rather than to whey protein and PFS fractions. In contrast, hydrophilic compounds were found to be uniformly distributed among the casein micelles, whey protein, and PFS pooled fractions. These observations support the hypothesis that casein micelles have natural affinity toward binding of hydrophobic compounds in their native state as compared with hydrophilic compounds. Further investigations resulted in identification of compounds with high hydrophobicity (log P > 5) such as PC in the casein micelle pooled fractions. Our data support the hypothesis that casein micelles may play an important role as hydrophobic bioactive compound carriers in addition to their other biological functions. These findings also support the use of casein micelles as encapsulation and delivery agents for hydrophobic bioactive agents in pharmaceutical and food-industry applications. In addition, this study lays the foundation for further analysis of structure-activity relationship studies in casein micelles.
ACKNOWLEDGMENTS
The authors thank the University of Tennessee Dairy Research and Education Center (Lewisburg) for providing us with the raw bovine milk samples. We also thank Abigail Tester and Stephen Dearth, graduate students at Department of Chemistry, University of Tennessee, Knoxville, for their assistance in liquid chromatograph-mass spectrometry method development and data analysis. We thank Philip Smith from the metabolomics core facility of the Pennsylvania State University for performing LC-MS/MS analysis supported by grant NSF MRI 1126373. Research reported in this publication was supported by the Eunice Kennedy Shriver National Institute of Child Health and Human Development (Rockville, MD) of the National Institutes of Health under award number R21HD065170. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
REFERENCES
- Cerbulis J, Zittle C. Lipids associated with acid-precipitated casein. J. Dairy Sci. 1965;48:1154–1156. doi: 10.3168/jds.S0022-0302(65)88418-1. [DOI] [PubMed] [Google Scholar]
- Chambers MC, Maclean B, Burke R, Amodei D, Ruderman DL, Neumann S, Gatto L, Fischer B, Pratt B, Egertson J, Hoff K, Kessner D, Tasman N, Shulman N, Frewen B, Baker TA, Brusniak M-Y, Paulse C, Creasy D, Flashner L, Kani K, Moulding C, Seymour SL, Nuwaysir LM, Lefebvre B, Kuhlmann F, Roark J, Rainer P, Detlev S, Hemenway T, Huhmer A, Langridge J, Connolly B, Chadick T, Holly K, Eckels J, Deutsch EW, Moritz RL, Katz JE, Agus DB, MacCoss M, Tabb DL, Mallick P. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 2012;30:918–920. doi: 10.1038/nbt.2377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clasquin MF, Melamud E, Rabinowitz JD. Pages 14.11.1–14.11.23 in Current Protocols in Bioinformatics. Hoboken, NJ.: John Wiley Sons Inc.; 2002. LC-MS data processing with MAVEN: A metabolomic analysis and visualization engine. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dalgleish DG. On the structural models of bovine casein micelles— Review and possible improvements. Soft Matter. 2011;7:2265–2272. [Google Scholar]
- De Kruif C. Casein micelle interactions. Int. Dairy J. 1999;9:183–188. [Google Scholar]
- De Kruif C, Holt C. Pages 233–276 in Advanced Dairy Chemistry—1 Proteins. New York, NY.: Springer; 2003. Casein micelle structure, functions and interactions. [Google Scholar]
- Donato P, Cacciola F, Cichello F, Russo M, Dugo P, Mondello L. Determination of phospholipids in milk samples by means of hydrophilic interaction liquid chromatography coupled to evaporative light scattering and mass spectrometry detection. J. Chromatogr. A. 2011;1218:6476–6482. doi: 10.1016/j.chroma.2011.07.036. [DOI] [PubMed] [Google Scholar]
- Horne DS. Casein interactions: Casting light on the black boxes, the structure in dairy products. Int. Dairy J. 1998;8:171–177. [Google Scholar]
- Horne DS. Casein micelle structure: Models and muddles. Curr. Opin. Colloid Interface Sci. 2006;11:148–153. [Google Scholar]
- Lu W, Clasquin MF, Melamud E, Amador-Noguez D, Caudy AA, Rabinowitz JD. Metabolomic analysis via reversed-phase ion-pairing liquid chromatography coupled to a stand alone orbitrap mass spectrometer. Anal. Chem. 2010;82:3212–3221. doi: 10.1021/ac902837x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martens L, Chambers M, Sturm M, Kessner D, Levander F, Shofstahl J, Tang WH, Römpp A, Neumann S, Pizarro AD, Montecchi-Palazzi L, Tasman N, Coleman M, Reisinger F, Souda P, Hermjakob H, Binz P-A, Deutsch EW. mzML—A community standard for mass spectrometry data. Mol. Cell. Proteomics. 2011;10:R110.000133. doi: 10.1074/mcp.R110.000133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melamud E, Vastag L, Rabinowitz JD. Metabolomic analysis and visualization engine for LC-MS data. Anal. Chem. 2010;82:9818–9826. doi: 10.1021/ac1021166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohan M, Jurat-Fuentes J, Harte F. Binding of vitamin A by casein micelles in commercial skim milk. J. Dairy Sci. 2013;96:790–798. doi: 10.3168/jds.2012-5777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oftedal OT. Pages 1–42 in Advanced Dairy Chemistry. New York, NY.: Springer; 2013. Origin and evolution of the major constituents of milk. [Google Scholar]
- Roach A, Harte F. Disruption and sedimentation of casein micelles and casein micelle isolates under high-pressure homogenization. Innov. Food Sci. Emerg. Technol. 2008;9:1–8. [Google Scholar]
- Sahu A, Kasoju N, Bora U. Fluorescence study of the curcumin− casein micelle complexation and its application as a drug nanocarrier to cancer cells. Biomacromolecules. 2008;9:2905–2912. doi: 10.1021/bm800683f. [DOI] [PubMed] [Google Scholar]
- Saldanha AJ. Java Treeview—Extensible visualization of microarray data. Bioinformatics. 2004;20:3246–3248. doi: 10.1093/bioinformatics/bth349. [DOI] [PubMed] [Google Scholar]
- Sanchez-Juanes F, Alonso J, Zancada L, Hueso P. Distribution and fatty acid content of phospholipids from bovine milk and bovine milk fat globule membranes. Int. Dairy J. 2009;19:273–278. [Google Scholar]
- Semo E, Kesselman E, Danino D, Livney YD. Casein micelle as a natural nano-capsular vehicle for nutraceuticals. Food Hydrocoll. 2007;21:936–942. [Google Scholar]
- Trejo R, Dokland T, Jurat-Fuentes J, Harte F. Cryotransmission electron tomography of native casein micelles from bovine milk. J. Dairy Sci. 2011;94:5770–5775. doi: 10.3168/jds.2011-4368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeisel S, Mar HM-H, Howe JC, Holden JM. Concentrations of choline-containing compounds and betaine in common foods. J. Nutr. 2003;133:1302–1307. doi: 10.1093/jn/133.5.1302. [DOI] [PubMed] [Google Scholar]
- Zhao Y-Y, Xiong Y, Curtis JM. Measurement of phospholipids by hydrophilic interaction liquid chromatography coupled to tandem mass spectrometry: The determination of choline containing compounds in foods. J. Chromatogr. A. 2011;1218:5470–5479. doi: 10.1016/j.chroma.2011.06.025. [DOI] [PubMed] [Google Scholar]