Background: The DNA mismatch repair protein MutS homolog 2 (MSH2) is an acetylated protein.
Results: Histone deacetylase 10 (HDAC10) interacts with MSH2 and deacetylates MSH2 at Lys-73, which might stimulate MSH2 activity.
Conclusion: HDAC10 promotes DNA mismatch repair activity and may involve the deacetylation of MSH2.
Significance: HDAC10 has an important role in regulating DNA mismatch repair.
Keywords: DNA mismatch repair, histone acetylase, histone deacetylase (HDAC), histone deacetylase inhibitor (HDAC inhibitor) (HDI), post-translational modification (PTM)
Abstract
MutS homolog 2 (MSH2) is an essential DNA mismatch repair (MMR) protein. It interacts with MSH6 or MSH3 to form the MutSα or MutSβ complex, respectively, which recognize base-base mispairs and insertions/deletions and initiate the repair process. Mutation or dysregulation of MSH2 causes genomic instability that can lead to cancer. MSH2 is acetylated at its C terminus, and histone deacetylase (HDAC6) deacetylates MSH2. However, whether other regions of MSH2 can be acetylated and whether other histone deacetylases (HDACs) and histone acetyltransferases (HATs) are involved in MSH2 deacetylation/acetylation is unknown. Here, we report that MSH2 can be acetylated at Lys-73 near the N terminus. Lys-73 is highly conserved across many species. Although several Class I and II HDACs interact with MSH2, HDAC10 is the major enzyme that deacetylates MSH2 at Lys-73. Histone acetyltransferase HBO1 might acetylate this residue. HDAC10 overexpression in HeLa cells stimulates cellular DNA MMR activity, whereas HDAC10 knockdown decreases DNA MMR activity. Thus, our study identifies an HDAC10-mediated regulatory mechanism controlling the DNA mismatch repair function of MSH2.
Introduction
The mismatch repair (MMR)4 system recognizes DNA mismatches that occur during DNA replication and corrects these defects to maintain genomic integrity. Failure of this mutation avoidance system can lead to microsatellite instability, elevated mutation rates, and predisposition for cancer. An example of the latter is hereditary nonpolyposis colorectal cancer kindred, where germ-line mutations in genes associated with mismatch repair have been identified (1, 2). In mammalian cells, MMR is initiated by MutS homolog 2 (MSH2)-containing heterodimeric complexes consisting of MSH2-MSH6 (MutSα) and MSH2-MSH3 (MutSβ). The MutSα complex recognizes base-base mispairs and single base insertions/deletions, whereas the MutSβ complex detects large insertions/deletions to initiate the repair process (3, 4). The MutSα and MutSβ complexes undergo ATP hydrolysis-dependent conformational transitions following binding to DNA mispairs or insertions/deletions and recruit a heterodimeric complex consisting of MLH1-PMS2 (MutLα). Interaction between the MutS and MutL complexes is essential for activating subsequent steps in the mismatch repair pathway (i.e. excision of mispairs and resynthesis of DNA strands) (5). Accumulating evidence indicates that MMR initiates the DNA mismatch repair pathway and induces G2 checkpoint activation and apoptosis (1, 6). Thus, the mismatch repair pathway plays a crucial role in mediating the DNA damage response and maintaining genomic integrity.
Histone deacetylases (HDACs) are enzymes that catalyze the removal of acetyl groups from lysine residues, whereas histone acetyltransferases (HATs) catalyze the addition of acetyl groups onto lysine residues. Both HDACs and HATs use core histones and non-histone proteins as substrates (7). Currently, there are 18 HDACs that fall into four classes in mammals (8). Class I HDACs (HDAC1, HDAC2, HDAC3, and HDAC8) share sequence similarity with the yeast RPD3 deacetylase. They exist in repressive complexes such as Sin3, NuRD, CoREST, PRC2, N-CoR, and SMRT complexes, which deacetylate histones and other nuclear proteins. Class II HDACs (HDAC4, HDAC5, HDAC6, HDAC7, HDAC9, and HDAC10) are homologous to the yeast Hda1 and exhibit tissue-specific expression. Class II HDACs are further subdivided into IIa (HDAC4, HDAC5, HDAC7, and HDAC9) and IIb (HDAC6 and HDAC10) subclasses. Class III HDACs include silent information regulator 2 (Sir2)-related NAD+-dependent deacetylases, a family of HDACs homologous to yeast Sir2. Class IV contains only one member, HDAC11, which is characterized by low sequence similarity with Class I and Class II members. Class I, IIa, and IV HDACs are sensitive to the inhibitors trichostatin A (TSA) and sodium butyrate (NaB), whereas Class IIb members are sensitive to TSA but insensitive to NaB (9). Class III HDACs, whose deacetylase activities require the coenzyme NAD+ as a cofactor, are specifically inhibited by nicotinamide (NIC) (8). HATs can be classified into three subfamilies: Gcn5-related N-acetyltransferases (GNATs), E1A-associated protein of 300 kDa (p300)/CREB-binding protein (CBP), and MYST family proteins (7).
Numerous studies show that HDACs and HATs have important roles in DNA repair and DNA damage responses. For example, Tip60, a MYST family member, cooperates with the Mre11-Rad50-Nbs1 complex to activate ataxia telangiectasia mutated protein (ATM) in response to DNA double-strand breaks (10). HDAC1 and HDAC2 are recruited to DNA damage sites to deacetylate histones H3K56 and H4K16 and facilitate non-homologous end-joining (11). HDAC1 also associates with other proteins involved in the DNA damage response, including PCNA, BRCA1, ATM, and ATR (ATM and Rad3-related) (12). HDAC4 associates with 53BP and is involved in repairing double-strand breaks (13). HDAC9 and HDAC10 are required for homologous recombination (HR) (14). Depletion of HDAC9 or HDAC10 inhibits HR and sensitizes cells to interstrand cross-linker mitomycin C treatment (14). We previously reported that SirT1 deacetylates NBS1 and regulates the cellular response to γ-IR (15). Two studies showed that depletion or inhibition of HDAC6 sensitizes cancer cells to compounds such as cisplatin, etoposide, and doxorubicin (16, 17). Recent evidence shows that HDAC6 deacetylates and ubiquitinates MSH2, and modulates the DNA damage response and MMR activity (18). However, whether MSH2 is regulated by other HDACs (in addition to HDAC6) and HATs is largely unknown.
Here, we identify Lys-73 as a novel acetylation site in MSH2. This site may be regulated through acetylation and deacetylation by HBO1, a MYST family HAT, and HDAC10, a Class IIb HDAC, respectively. HDAC10 might stimulate DNA mismatch repair activity by deacetylation of MSH2.
Experimental Procedures
Plasmids, Antibodies, Reagents, and Cell Lines
Plasmids expressing all Class I, II, and IV HDACs, including FLAG-HDAC1, -HDAC2, -HDAC3, -HDAC4, -HDAC5, -HDAC7, -HDAC8, -HDAC9, -HDAC10, and -HDAC11, were described previously (15). FLAG-HDAC10 plasmids were kindly provided to us by Tso-Pang Yao (Duke University) and Xiang-Jiao Yang (McGill University). HA-MSH2 and FLAG-MSH2 were constructed using the pcDNA3.1 vector. HA-CBP, HA-p300, HA-HBO1, HA-Tip60, and FLAG-PCAF were also described previously (15). HA-HBO1G485A (HA-HBO1Δ), a catalytically deficient mutant, was a generous gift from Dr. Mitchell Smith at the University of Virginia. The anti-HA antibody, anti-FLAG-M2 antibody, anti-HA-agarose beads, anti-FLAG M2-agarose beads, protein A and protein G beads, rabbit IgG-agarose beads, mouse IgG-agarose beads, trichostatin A, andnicotinamide were purchased from Sigma-Aldrich. The anti-MSH2 antibodies were purchased from EMD Millipore and Bethyl Laboratories. The anti-acetyl-lysine antibody was purchased from Cell Signaling Technology. The anti-MSH2Ac-K73 antibody was custom-made from Cell Signaling Technology. Antibody specificity was validated by ELISA using acetylated and non-acetylated peptides. NaB was obtained from Thermo Fisher Scientific.
293T and HeLa cells were cultured in DMEM. HeLaS3 cells were grown in Joklik medium supplemented with amphotericin B and sodium bicarbonate. All media were supplemented with 10% fetal calf serum, 100 μg/ml streptomycin, and 100 IU/ml penicillin. All cells were grown at 37 °C with 5% CO2. Transfections were performed using FuGENE 6 (Roche Applied Science).
Mass Spectrometry
For acetylation analysis, 293T cells were transfected with the HA-MSH2 construct. Forty hours after transfection, cells were treated with 1.3 μm TSA for an additional 8 h. Cells were collected and lysed following the standard procedure, and immunoprecipitation was performed with HA-agarose beads (Sigma). The MSH2 immunoprecipitate was resolved on an 8% SDS-PAGE gel and stained using the Colloidal Blue staining kit (Invitrogen). The prominent HA-MSH2 band was excised and divided into two equal parts for analysis. Each part was subjected to in-gel reduction and carboxyamidomethylation, followed by separate tryptic or chymotryptic digestion. Acetylated peptides from each digest were detected and sequenced using microcapillary reverse-phase high-performance LC-MS/MS on a Thermo Fisher LTQ-Orbitrap hybrid mass spectrometer. Data analysis was performed using SEQUEST and a customized version of the Proteomics Browser Suite software (Thermo Fisher Scientific).
Immunoprecipitation and Immunoblotting
Cells were incubated on ice for 30 min in 300 μl of NETN buffer (50 mm Tris-HCl, pH 8.0, 0.5 mm EDTA, 150 mm NaCl, 0.5% NP-40, and protease inhibitor cocktail). Cell lysates were precleared with protein A/G-agarose for 1 h and incubated overnight with primary antibody at 4 °C. The beads were washed five times with NETN buffer containing 150 mm NaCl, and the immunoprecipitated proteins were resolved by SDS-PAGE and subsequently transferred to nitrocellulose membranes. The blots were probed with the indicated antibodies. Bound antibodies were detected using a chemiluminescent detection kit (Amersham Biosciences).
Mismatch Repair Substrate Preparation and Mismatch Repair Assay
5′- and 3′-mismatch repair substrates were prepared as described previously (19, 20). Preparation of nuclear extracts for the mismatch repair assay was also described previously (20, 21).
The mismatch repair assay was performed as reported previously (20, 21). Briefly, the MMR assay was carried out in a 15-μl reaction mixture containing 23 fmol of heteroduplex DNA, ∼50–200 μg of nuclear lysate, 10 mm Tris-HCl (pH 7.6), 5 mm MgCl2, 100 mm KCl, 1.5 mm ATP, and 0.1 mm dNTPs. The reactions were incubated at 37 °C for 15 min, and DNA samples were recovered by phenol extraction and ethanol precipitation and then digested with restriction enzymes (New England Biolabs) according to the manufacturer's recommendation to score repair (20). The digested products were examined on 1% agarose gels. The band intensity was quantified, and the repair percentage was analyzed.
Results
Multiple HDACs Interact with MSH2
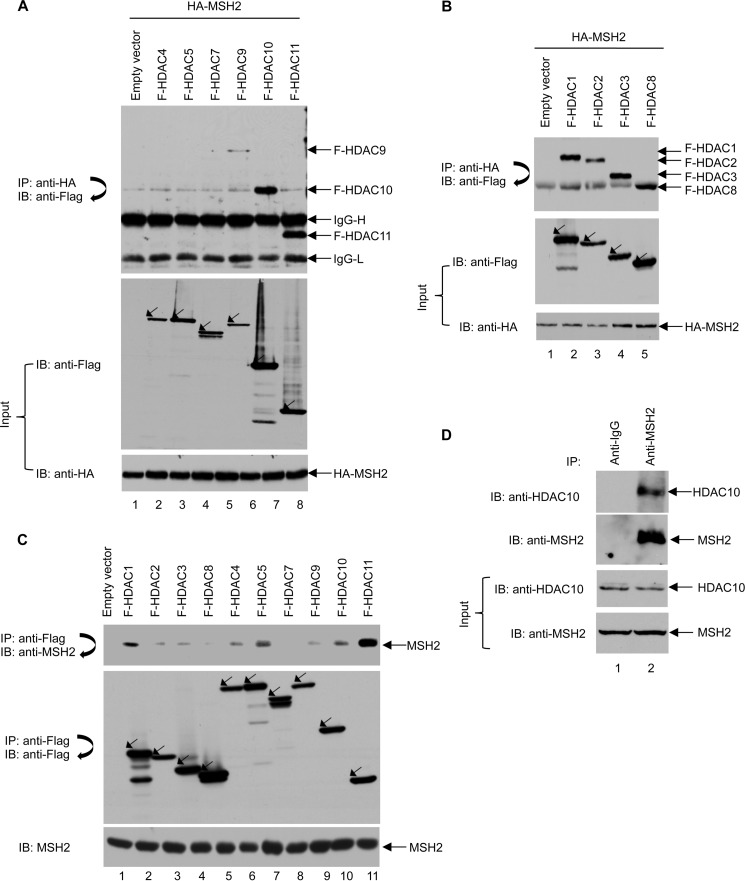
HDACs interact with and deacetylate core histones and non-histone proteins (7). Previous studies show that Hda1, a yeast Class II HDAC, interacts with MSH2 (22). Consistent with these studies, a recent study shows that a Class IIb HDAC, HDAC6, interacts with MSH2 (18). However, it remains unclear whether other HDACs also bind MSH2. To this end, FLAG-tagged HDAC11 and all Class II HDACs except HDAC6 were co-transfected with HA-MSH2 into 293T cells. As shown in Fig. 1A, anti-HA immunoprecipitation showed binding of FLAG-tagged HDAC9, HDAC10, and HDAC11 to MSH2. Similar results were obtained when the HDACs were immunoprecipitated with FLAG-conjugated agarose beads followed by immunoblotting with an anti-HA antibody to identify proteins that interact with MSH2 (data not shown). Overexpression of HDAC11 and Class II HDACs with MSH2 showed that HDAC9, HDAC10, and HDAC11 bind MSH2. As MSH2 and Class I HDACs are primarily nuclear proteins, we investigated whether Class I HDACs interact with MSH2. HA-MSH2 was co-transfected with FLAG-tagged Class I HDACs including HDAC1, HDAC2, HDAC3, and HDAC8 into 293T cells. The cell lysates were immunoprecipitated with anti-HA-conjugated agarose beads to pull down HA-MSH2 and its interacting partners. As shown in Fig. 1B, all Class I HDACs interacted with MSH2.
FIGURE 1.
Multiple HDACs interact with MSH2. A, several Class II HDACs interact with MSH2 in 293T cells. 293T cells were transiently transfected with plasmids expressing HA-MSH2 and FLAG (F)-tagged Class II HDACs as indicated. HA-MSH2 immunoprecipitation (IP) with anti-HA-agarose beads was performed followed by immunoblotting (IB) with an anti-FLAG antibody (upper panel). Total cell lysates were subjected to immunoblot analyses with anti-FLAG and anti-HA antibodies (middle and lower panels). B, all Class I HDACs interact with MSH2 in 293T cells. 293T cells were transiently transfected with plasmids expressing HA-MSH2 and FLAG-tagged Class I HDACs as indicated. HA-MSH2 immunoprecipitation with anti-HA-agarose beads was performed followed by immunoblotting with an anti-FLAG antibody (upper panel). Total cell lysates were subjected to immunoblot analyses with anti-FLAG and anti-HA antibodies (middle and lower panels). C, multiple HDACs interact with endogenous MSH2 in 293T cells. 293T cells were transiently transfected with plasmids expressing the indicated FLAG-tagged Class I and II HDACs. Immunoprecipitation of HDACs with anti-FLAG M2-agarose beads was performed followed by immunoblot analyses with anti-MSH2 antibody to detect endogenous MSH2 (upper panel). The blot was then stripped and reprobed with anti-FLAG antibody (middle panel). Levels of MSH2 were confirmed by Western blot analysis of the cell lysates using an anti-MSH2 antibody (lower panel). D, interaction between endogenous HDAC10 and MSH2 in HeLa cells. HeLa cell lysates were first immunoprecipitated with anti-IgG or anti-MSH2 antibody and then subjected to immunoblot analysis using an anti-HDAC10 antibody. The blot was then stripped and reprobed with an anti-MSH2 antibody.
We examined whether HDACs interacted with endogenous MSH2. For this purpose, all FLAG-tagged Class I, II, and IV HDACs except HDAC6 were overexpressed in 293T cells. Anti-FLAG immunoprecipitation was then performed to pull down endogenous MSH2. As shown in Fig. 1C, HDAC1 and HDAC11 interacted strongly with endogenous MSH2, whereas HDAC5 and HDAC10 moderately associated with MSH2. The remaining HDACs exhibited only weak interactions with MSH2. Fig. 1D shows that an interaction between endogenous HDAC10 and MSH2 could be identified in HeLa cells. These results demonstrate that other HDACs in addition to HDAC6 interact with MSH2, suggesting that it may be regulated by multiple HDACs.
MSH2 Is Acetylated at Lysine 73
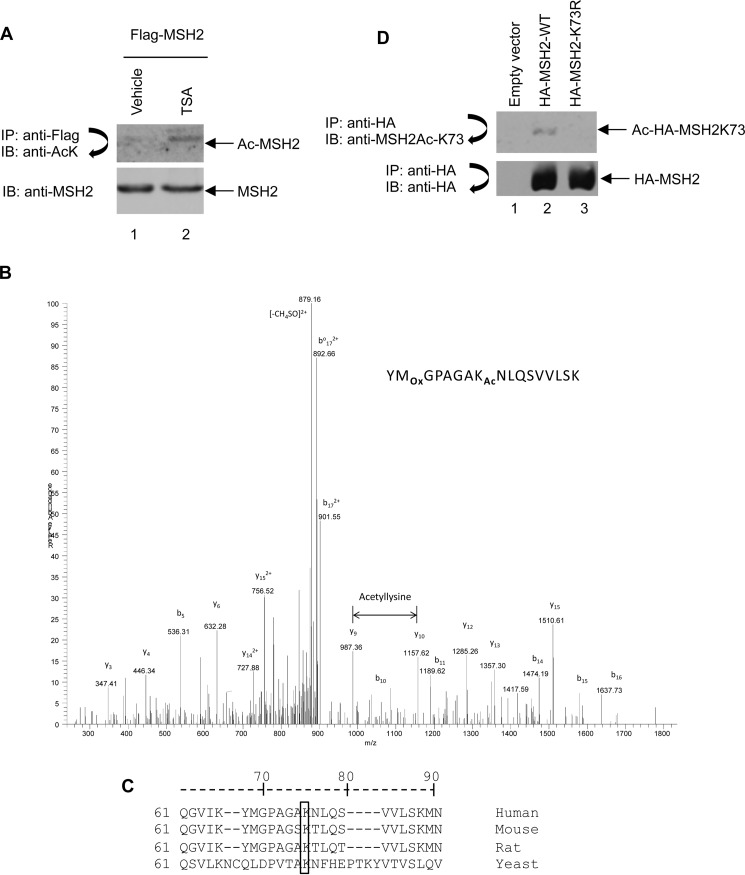
Four acetylation sites, Lys-845, Lys-847, Lys-871, and Lys-892, have been identified at the MSH2 C terminus by mass spectrometry (18). However, mutation of these four sites from lysine to arginine still allows for a modest level of acetylation when compared with that of wild type (18), suggesting that additional acetylation sites exist in MSH2. To confirm that MSH2 is acetylated and can be deacetylated by HDACs, we took an unbiased approach to inhibit HDACs in 293T cells using the pan HDAC inhibitor TSA, which inactivates Class I, II, and IV HDACs, and then assessed the acetylation level of overexpressed HA-MSH2. Cells were treated with either vehicle or TSA for 12 h before harvest. As shown in Fig. 2A, the level of MSH2 acetylation increased in TSA-treated cells, suggesting that Class I, II, and IV HDACs deacetylate MSH2.
FIGURE 2.
MSH2 is acetylated at Lys-73. A, MSH2 acetylation (Ac) level was increased by TSA treatment. 293T cells were transiently transfected with plasmid expressing FLAG-tagged MSH2. At 40 h after transfection, cells were treated with the HDAC inhibitor TSA as indicated. HA-MSH2 was immunoprecipitated (IP) with anti-FLAG M2-agarose beads followed by immunoblotting (IB) with anti-acetyl-lysine antibody (upper panel). Total cell lysates were subjected to immunoblot analysis with anti-MSH2 antibody (lower panel). B, lysine 73 is acetylated in MSH2. A doubly charged peptide was detected with a mass-to-charge (m/z) ratio of 910.9821, which represents an error of 0.3 ppm. The tandem mass spectrum matched the following sequence: YM[Ox]GPAGAK[Ac]73NLQSVVLSK, indicating that Lys-73 was acetylated (Ox, oxidation; Ac, acetylation). The assignment was made using the Proteomics Browser Suite (PBS) with a probability of 53 (= −10×log(P)). Acetylation was localized to Lys-73 by the PBS Graphmod software without requiring the modification to be specific to lysine. Detection of y9 and y10 is consistent with this localization. C, MSH2 amino acid sequence showing conservation of the Lys-73 site across yeast, mouse, rat, and human. D, the anti-MSH2Ac-K73 antibody specifically recognizes acetylated Lys-73 in MSH2. 293T cells were transfected with empty vector, HA-MSH2-WT, or HA-MSH2-K73R and incubated for 48 h. Cells were then harvested, lysed, and subjected to immunoprecipitation with anti-HA-agarose beads followed by immunoblotting with anti-MSH2Ac-K73 antibody. Finally, the blot was stripped and reprobed with anti-HA antibody.
We proceeded to identify novel acetylation sites in MSH2 by overexpressing HA-MSH2 in 293T cells, treating the cells with TSA, and then performing mass spectrometric analyses. As shown in Fig. 2B, Lys-73 was detected as an acetylation site. The total peptide coverage for proteomic analysis was ∼97% (data not shown). Sequence alignment demonstrated that Lys-73 is highly conserved across yeast, rat, mouse, and human (Fig. 2C). To examine Lys-73 acetylation, we used a custom-made purified antibody specifically recognizing MSH2 acetylation at residue Lys-73. We validated this antibody by ELISA using both the non-acetylated and the acetylated 17-mer peptides that encompass the MSH2 Lys-73 region and found that this antibody specifically binds to the acetylated peptide (data not shown). To further confirm the antibody specificity, we mutated Lys-73 to an arginine to generate the MSH2-K73R mutant and transfected 293T cells with either wild-type MSH2 or MSH2-K73R. As shown in Fig. 2D, the anti-MSH2Ac-K73 antibody recognized wild-type MSH2 but not the K73R mutant, suggesting that this antibody specifically recognizes the acetylated Lys-73 residue in MSH2. These results show for the first time that MSH2 is acetylated at residue Lys-73.
HDAC10 Is the Major Deacetylase Catalyzing Lys-73 Deacetylation in MSH2
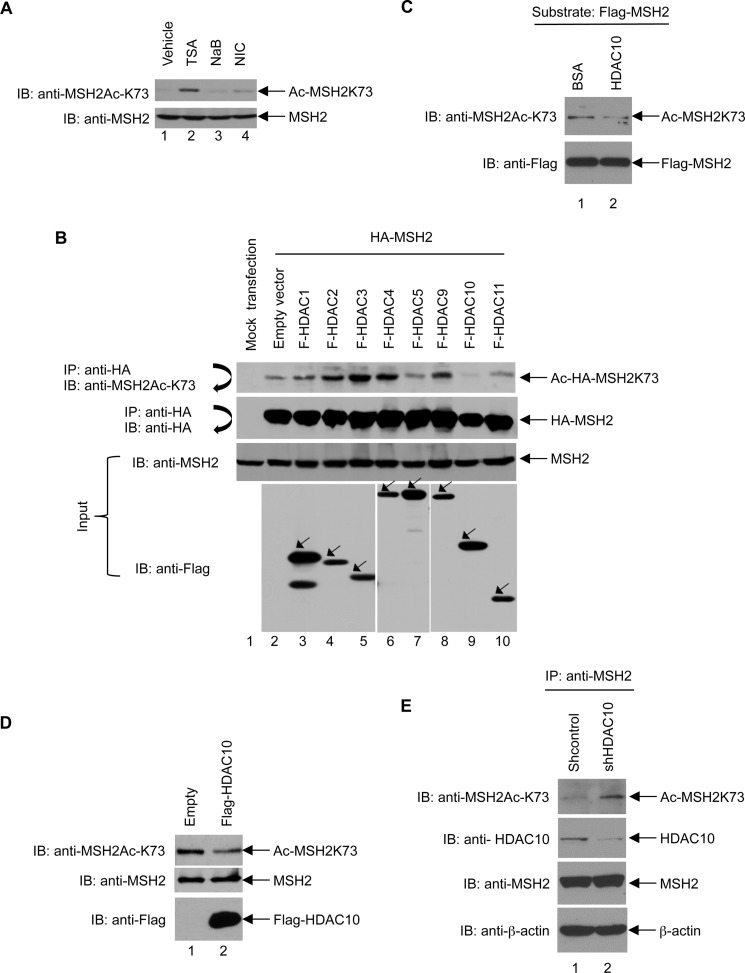
We investigated which HDAC deacetylates MSH2 Lys-73. HeLa cells were treated with TSA, NaB, or nicotinamide, and then Lys-73 acetylation was examined by immunoblot analysis using the anti-MSH2Ac-K73 antibody. As shown in Fig. 3A, TSA induces the highest level of MSH2-K73 acetylation (lane 2). By contrast, NaB (lane 3) and NIC (lane 4) treatment did not significantly increase Lys-73 acetylation. Because TSA inhibits Class I, II, and IV HDACs, whereas nicotinamide inhibits Class III HDACs, our results indicate that Lys-73 can be deacetylated by Class I, II, and IV, but not by Class III HDACs. Because Class IIb HDACs, including HDAC6 and HDAC10, are insensitive to NaB, our data suggest that Class IIb HDACs are involved in regulating Lys-73 deacetylation.
FIGURE 3.
HDAC10 deacetylates MSH2 at Lys-73. A, the level of Lys-73 acetylation (Ac) was dramatically increased by treatment with TSA but not NIC or NaB. HeLa cells were treated with HDAC inhibitors, including TSA, NaB, and NIC, for 12 h as indicated. Total lysates were subjected to immunoblot (IB) analysis using the anti-MSH2Ac-K73 antibody (upper panel). The blot was stripped and reprobed with anti-MSH2 antibody (lower panel). B, FLAG-HDAC10 deacetylates Lys-73 in MSH2. 293T cells were transiently transfected with plasmids expressing HA-MSH2 and FLAG (F)-tagged HDACs as indicated. Total cell lysates were immunoprecipitated (IP) with anti-HA-agarose beads followed by immunoblotting with anti-MSH2Ac-K73 antibody (first panel). The blot was then stripped and reprobed with anti-HA antibody (second panel). Total cell lysates were subjected to immunoblotting with anti-MSH2 (third panel) or anti-FLAG (fourth panel) antibody. C, HDAC10 deacetylates MSH2 at Lys-73 in vitro. 293T cells were transiently transfected with plasmids expressing FLAG-MSH2 and treated with TSA for 12 h. Highly acetylated MSH2 was immunoprecipitated by anti-FLAG-agarose beads and followed by incubating with 3 μg of BSA (as a control) or recombinant HDAC10 (BML-SE559-0050, Enzo Life Sciences) at 37 °C for 2 h. After washing the anti-FLAG-agarose beads with LS buffer (0.1% NP-40, 10% glycerol in PBS), the FLAG-MSH2 immunoprecipitates were subjected to the immunoblotting analysis with the anti-MSH2Ac-K73 antibody (upper panel). The blot was then stripped and reprobed with the anti-FLAG antibody (lower panel). D, HDAC10 overexpression reduces Lys-73 acetylation in HeLa cells. FLAG-HDAC10 was stably overexpressed in HeLa cells. Total lysates prepared from control and FLAG-HDAC10-overexpressing cells were subjected to immunoblot analysis with anti-MSH2Ac-K73, anti-MSH2, and anti-FLAG antibodies (upper, middle, and lower panels, respectively). E, HDAC10 knockdown increases Lys-73 acetylation in MSH2. The lentiviral system was used to transduce HDAC10-specific shRNA (TRCN0000004861, Thermo Fisher Scientific) or scramble shRNA into HeLa cells. Lentivirus-transduced cells were selected for puromycin resistance, and the shRNA stably expressing cells were lysed in NETN buffer (100 mm NaCl, 20 mm Tris-Cl (pH 8.0), 0.5 mm EDTA, 0.5% NP-40). The total cell lysates were immunoprecipitated with the anti-MSH2 antibody followed by immunoblotting analysis with the anti-MSH2Ac-K73 antibody (first panel), anti-HDAC10 antibody (second panel), anti-MSH2 antibody (third panel), or anti-β-actin antibody (fourth panel).
To further determine which HDAC is the major deacetylase for Lys-73 deacetylation in MSH2, HA-MSH2 was co-transfected with FLAG-tagged HDAC1, -HDAC2, -HDAC3, -HDAC4, -HDAC5, -HDAC9, -HDAC10, or -HDAC11 in 293T cells. The level of Lys-73 acetylation was assessed by immunoblot analysis using the anti-MSH2Ac-K73 antibody. These results show that, among all HDACs tested, only HDAC10 deacetylated Lys-73 dramatically (Fig. 3B, compare lane 9 with other lanes). In addition, recombinant HDAC10 was able to directly deacetylate MSH2 efficiently (Fig. 3C). We then examined the role of HDAC10 in deacetylating endogenous MSH2 by stably overexpressing HDAC10 in HeLa cells. As shown in Fig. 3D, HDAC10-overexpressing cells exhibited a reduction in MSH2 acetylation at Lys-73 when compared with that of control cells. Conversely, HDAC10-depleting cells exhibited an increase in MSH2 acetylation at Lys-73 when compared with that of control cells (Fig. 3E). These data unequivocally demonstrate that Lys-73 deacetylation is mediated by HDAC10.
HBO1 Acetylates MSH2 at Lys-73
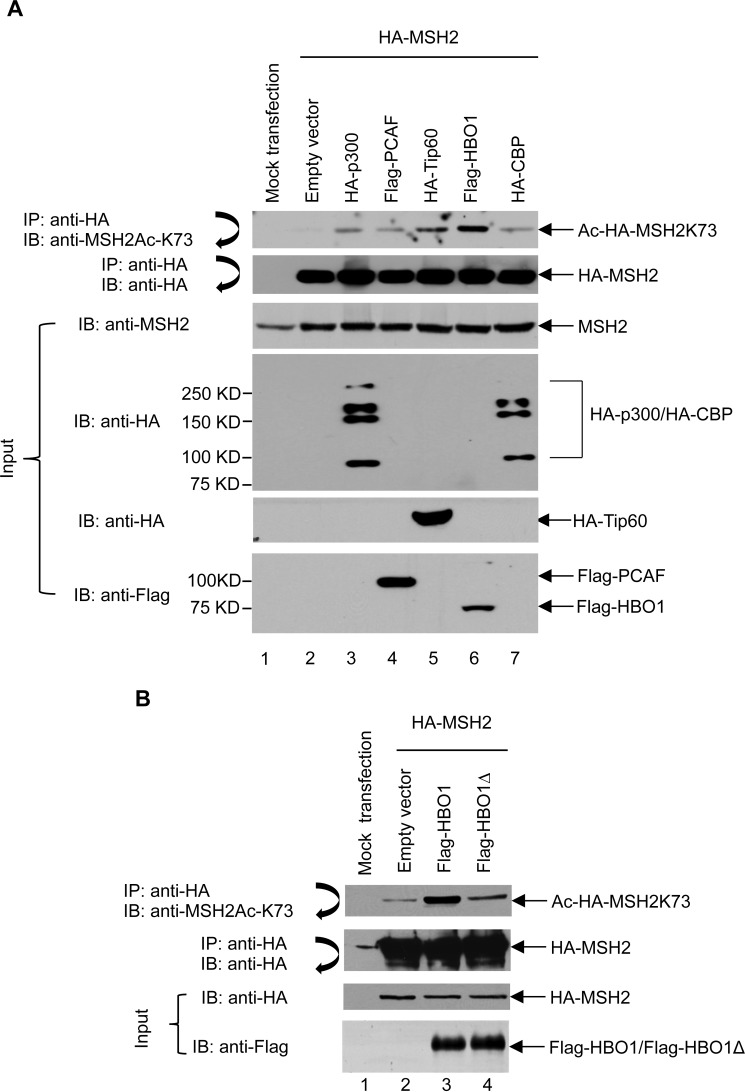
To determine which HAT acetylates MSH2 Lys-73, several HATs that have important roles in DNA repair (23), including CBP, PCAF, HBO1, Tip60, and p300, were co-transfected with HA-MSH2 into 293T cells. These HATs represent all three HAT subfamilies. PCAF belongs to the GNAT subfamily; CBP and p300 belong to the p300/CBP subfamily; and HBO1 and Tip60 belong to the MYST subfamily. Cells were then subjected to immunoprecipitation of HA-MSH2 followed by immunoblotting using the anti-MSH2Ac-K73 antibody. As shown in Fig. 4A, all tested HATs acetylated MSH2 at Lys-73. Among these, HBO1 enhanced MSH2 acetylation at Lys-73 most dramatically (lane 6). To further validate these results, HA-MSH2 was co-transfected with either HBO1-WT or HBO1Δ, a catalytically deficient mutant, in 293T cells, and then the level of Lys-73 acetylation in MSH2 was examined. As expected, HBO1-WT, but not HBO1Δ, strongly acetylated MSH2 at the Lys-73 site (Fig. 5B). Therefore, HBO1 is the major acetyltransferase catalyzing Lys-73 acetylation in MSH2.
FIGURE 4.
HBO1 acetylates MSH2 at Lys-73. A, HBO1 is the major acetyltransferase for Lys-73 acetylation in MSH2. 293T cells were transiently co-transfected with plasmids expressing HA-MSH2 and acetyltransferases such as FLAG-PCAF, HA-Tip60, FLAG-HBO1, and HA-CBP. Cells were harvested at 48 h after transfection. HA-MSH2 was then immunoprecipitated (IP) by anti-HA-agarose beads followed by immunoblotting (IB) with anti-MSH2Ac-K73 antibody (first panel). The blot was then stripped and reprobed with anti-HA antibody (second panel). Total cell lysates were subjected to immunoblot analyses with anti-MSH2 antibody, anti-HA antibody, and anti-FLAG antibody (third, fourth, fifth, and sixth panels, respectively). B, a catalytically deficient HBO1 mutant cannot acetylate MSH2 at Lys-73. 293T cells were co-transfected with plasmids expressing HA-MSH2 and FLAG-HBO1 or FLAG-HBO1Δ as indicated. Cells were harvested at 48 h after transfection. HA-MSH2 was then immunoprecipitated by anti-HA-agarose beads followed by immunoblotting using anti-MSH2Ac-K73 antibody (first panel). The blot was then stripped and reprobed with an anti-HA antibody (second panel). Total cell lysates were subjected to immunoblot analyses with anti-HA (third panel) and anti-FLAG (fourth panel) antibodies.
FIGURE 5.
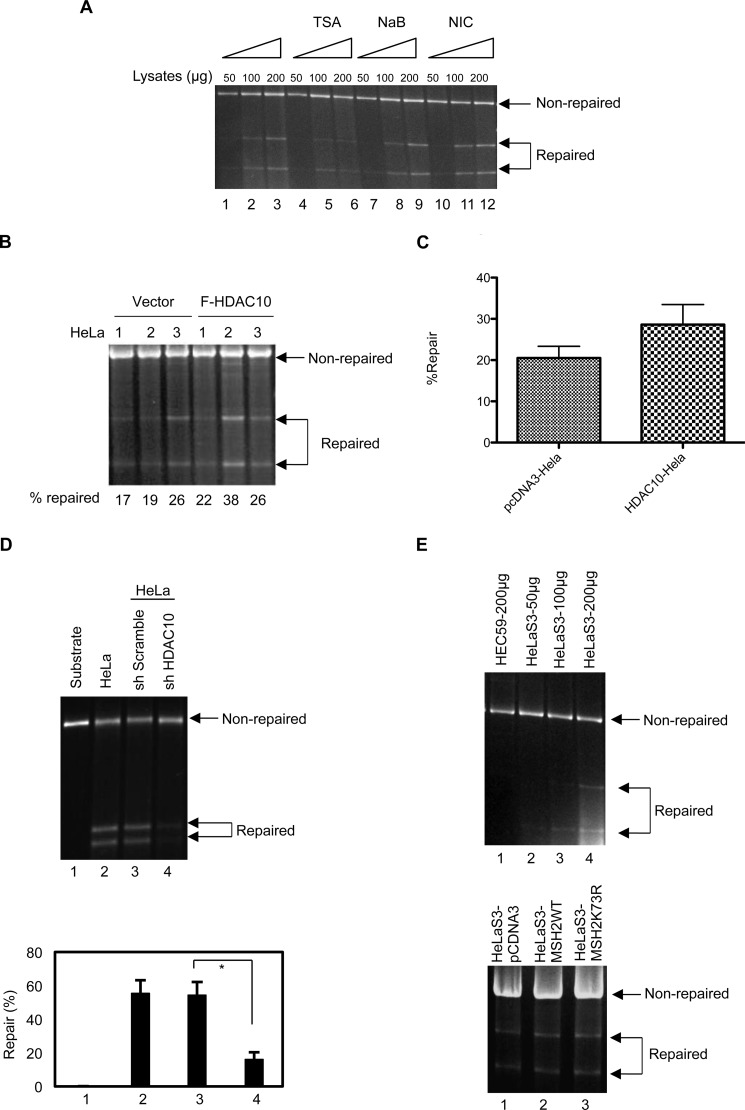
HDAC10 promotes DNA MMR activity. A, TSA, but not NaB or NIC, reduces DNA MMR activity. HeLaS3 cells were treated with 1.3 μm TSA, 10 mm NaB, or 10 mm NIC for 8 h. Nuclear extracts prepared from these cells (50, 100, or 200 μg) were used in the DNA MMR assay. B, HDAC10 promotes DNA MMR activity. FLAG-HDAC10 was stably overexpressed in HeLaS3 cells. Nuclear extracts of control and FLAG (F)-HDAC10-expressing stable cells were used in the DNA MMR assay. Data represent three independent experiments. C, repaired bands in panel B were quantified by densitometry and graphed. Values represent mean ± S.D. D, HDAC10 knockdown reduces DNA MMR activity. HDAC10 was stably knocked down in HeLa cells as in Fig. 3E. The substrate only (as a negative control) and the substrate with nuclear extracts of HeLa cells (as a positive control), sh Scramble-, or shHDAC10-transduced HeLa cells were used in the DNA MMR assay (upper panel). Data represent three independent experiments. The lower panel is the quantitative result of repair percentage in each group (lower panel). Values represent mean ± S.D. *, p < 0.05. E, MSH2K73R does not promote DNA MMR activity. MSH2WT and MSH2K73R were stably overexpressed in HeLaS3 cells. Nuclear extracts of control, MSH2WT-, and MSH2K73R-expressing stable cells were used in the DNA MMR assay (lower panel). For standardization of MMR assay, HelaS3 nuclear lysates were used for MMR assay at the indicated concentration. As a negative control for the assay, HEC59 cells that lack MSH2 were used (upper panel). The MMR assay was performed as outlined under “Experimental Procedures.”
HDAC10 Might Stimulate the DNA MMR Activity of MSH2
Recent work shows that HDAC6 promotes MSH2 degradation and leads to down-regulation of cellular DNA mismatch repair activity. We examined whether HDAC10 also has a role in DNA mismatch repair. The HDAC inhibitors TSA, NaB, and NIC were utilized to inhibit HDAC10 activity. Briefly, HeLaS3 cells were treated for 8 h with the HDAC inhibitors at the concentrations indicated in Fig. 5. Nuclear lysates were prepared from these cells, and a DNA mismatch repair assay was performed with the 3′-GT mismatch repair substrate. Our results show that TSA treatment inhibited mismatch repair activity in 100- and 200-μg nuclear lysate protein samples (Fig. 5A, compare lanes 4, 5, 6 and lanes 1, 2, 3). Neither NaB nor NIC (18) had any effect on mismatch repair activity (Fig. 5A, compare lanes 7–9 and 10–12 with lanes 1–3). These data indicate an important role for HDAC10 in DNA mismatch repair. We stably overexpressed HDAC10 in HeLa cells and performed DNA mismatch repair assays with nuclear lysates prepared from control cells and HDAC10-overexpressing cells. The results show that HDAC10 overexpression stimulated DNA mismatch repair activity (Fig. 5, B and C). Consistently, HDAC10-depleting HeLa cells exhibited more than 3-fold reduction in MMR activity with a 5′-GT mismatch substrate when compared with the control HeLa cells (Fig. 5D).
Discussion
The results in this study do not directly prove that HDAC10 regulates DNA mismatch repair through deacetylation of MSH2. They do, however, indirectly point to a novel functional role for HDAC10 in DNA mismatch repair. Emerging evidence shows that post-translational modifications such as acetylation/deacetylation have a critical role in DNA repair and DNA damage response. MSH2 phosphorylation is reported to promote the mismatch repair activity of MutSα and is required for nuclear translocation of the MutSα complex (24). Recent work shows that HDAC6 sequentially deacetylates and ubiquitinates MSH2 at Lys-845, Lys-847, Lys-871, and Lys-892, leading to MSH2 degradation (18). MSH2 deacetylation at these four sites by HDAC6 facilitates disassembly of the MutSα complex, reduces cellular MMR activity, and causes cellular tolerance to DNA-damaging agents such as 6-thioguanine and N-methyl-N′-nitro-N-nitrosoguanidine (18). In contrast to HDAC6 functional activity, we found that HDAC10 promotes the DNA MMR activity of MSH2 as evidenced by the data that knockdown of HDAC10 dramatically increases the cellular DNA MMR activity (Fig. 5D). Although overexpression of the deacetylation mimetic K73R of MSH2 did not alter the DNA MMR activity when compared with that of wild type (Fig. 5E), HDAC10 may regulate DNA MMR through deacetylation of other lysine sites. In fact, in addition to Lys-73, we have detected multiple lysine sites in MSH2, which may be deacetylated by HDAC10 (data not shown), and deacetylation of these sites could stimulate DNA MMR activity. It has also been reported that HDAC10 is required for HR, which is specifically inhibited by HDAC10 depletion (14). MMR machinery components such as MSH2 have been shown to suppress HR (23). Based on these studies, HDAC10 may affect HR through MSH2.
Protein post-translational modifications, including acetylation, usually influence protein stability and localization (25, 26). To test whether Lys-73 acetylation affects MSH2 stability, we examined the half-life of MSH2. No difference in half-life was detected among HA-MSH2-WT, HA-MSH2-K73R, and HA-MSH2-K73Q (data not shown), indicating that Lys-73 deacetylation/acetylation does not affect MSH2 stability. We also investigated whether MSH2 acetylation affects its subcellular localization. However, we did not detect significant changes in subcellular distribution among the HA-MSH2-K73R and HA-MSH2-K73Q mutants and HA-MSH2-WT (data not shown).
Although the K73R and K73Q mutants exhibited similar interactions with MSH6 and MSH3 as that of wild-type MSH2 (data not shown), K73Q, a mutant that mimics acetylation, exhibited reduced association with proliferating cell nuclear antigen (data not shown), an important component required for mismatch repair at a step preceding DNA resynthesis (27, 28). We used mass spectrometric analysis to identify a new MSH2-binding protein, designated as And-1. The K73Q mutant displayed reduced interaction with And-1 when compared with that of wild type (data not shown). And-1 is a replication initiation factor that brings together the MCM2–7 helicase and DNA polymerase α complex to initiate DNA replication (29). Therefore, Lys-73 acetylation may affect the mismatch repair function of MSH2 during DNA resynthesis. Future studies will elucidate the exact role of And-1 in DNA mismatch repair.
Author Contributions
E. S. conceived and coordinated the study. R. R., Y. L., S. X., F. Y., Z. Y., E. T., J. F., D. C., D. S., W. S. L., Y. Z., and X. Z. performed the experiments, contributed substantially to conception and design, acquisition of data, or analysis and interpretation of the data, or prepared the figures. The manuscript was written with inputs from all authors, and all authors reviewed the results and approved the final version of the manuscript.
This study was supported in part by National Institutes of Health grants R01CA169210 (to E. S.) and R01HL105631 (to Y. Z.), an endowment from the Kaul Foundation (to E. S.), and a postdoctoral fellowship from the Bankhead Coley Biomedical Research Program (to R. R.) The authors declare that they have no conflicts of interest with the contents of this article.
- MMR
- mismatch repair
- HDAC
- histone deacetylase
- HAT
- histone acetyltransferase
- TSA
- trichostatin A
- NaB
- sodium butyrate
- NIC
- nicotinamide
- CBP
- CREB-binding protein
- CREB
- cAMP-response element-binding protein
- PCAF
- P300/CBP-associated factor
- HR
- homologous recombination.
References
- 1. Modrich P. (1994) Mismatch repair, genetic stability, and cancer. Science 266, 1959–1960 [DOI] [PubMed] [Google Scholar]
- 2. Peltomäki P. (2003) Role of DNA mismatch repair defects in the pathogenesis of human cancer. J. Clin. Oncol. 21, 1174–1179 [DOI] [PubMed] [Google Scholar]
- 3. Drummond J. T., Li G. M., Longley M. J., Modrich P. (1995) Isolation of an hMSH2-p160 heterodimer that restores DNA mismatch repair to tumor cells. Science 268, 1909–1912 [DOI] [PubMed] [Google Scholar]
- 4. Genschel J., Littman S. J., Drummond J. T., Modrich P. (1998) Isolation of MutSβ from human cells and comparison of the mismatch repair specificities of MutSβ and MutSα. J. Biol. Chem. 273, 19895–19901 [DOI] [PubMed] [Google Scholar]
- 5. Li G. M. (2008) Mechanisms and functions of DNA mismatch repair. Cell Res. 18, 85–98 [DOI] [PubMed] [Google Scholar]
- 6. Fink D., Aebi S., Howell S. B. (1998) The role of DNA mismatch repair in drug resistance. Clin. Cancer Res. 4, 1–6 [PubMed] [Google Scholar]
- 7. Yang X. J., Seto E. (2008) Lysine acetylation: codified crosstalk with other posttranslational modifications. Mol. Cell 31, 449–461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Yang X. J., Seto E. (2008) The Rpd3/Hda1 family of lysine deacetylases: from bacteria and yeast to mice and men. Nat. Rev. Mol. Cell Biol. 9, 206–218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Guardiola A. R., Yao T. P. (2002) Molecular cloning and characterization of a novel histone deacetylase HDAC10. J. Biol. Chem. 277, 3350–3356 [DOI] [PubMed] [Google Scholar]
- 10. Sun Y., Jiang X., Price B. D. (2010) Tip60: connecting chromatin to DNA damage signaling. Cell Cycle 9, 930–936 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Miller K. M., Tjeertes J. V., Coates J., Legube G., Polo S. E., Britton S., Jackson S. P. (2010) Human HDAC1 and HDAC2 function in the DNA-damage response to promote DNA nonhomologous end-joining. Nat. Struct. Mol. Biol. 17, 1144–1151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Rajendran P., Ho E., Williams D. E., Dashwood R. H. (2011) Dietary phytochemicals, HDAC inhibition, and DNA damage/repair defects in cancer cells. Clin. Epigenetics 3, 4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Kao G. D., McKenna W. G., Guenther M. G., Muschel R. J., Lazar M. A., Yen T. J. (2003) Histone deacetylase 4 interacts with 53BP1 to mediate the DNA damage response. J. Cell Biol. 160, 1017–1027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Kotian S., Liyanarachchi S., Zelent A., Parvin J. D. (2011) Histone deacetylases 9 and 10 are required for homologous recombination. J. Biol. Chem. 286, 7722–7726 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Yuan Z., Zhang X., Sengupta N., Lane W. S., Seto E. (2007) SIRT1 regulates the function of the Nijmegen breakage syndrome protein. Mol. Cell 27, 149–162 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Namdar M., Perez G., Ngo L., Marks P. A. (2010) Selective inhibition of histone deacetylase 6 (HDAC6) induces DNA damage and sensitizes transformed cells to anticancer agents. Proc. Natl. Acad. Sci. U.S.A. 107, 20003–20008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Wang L., Xiang S., Williams K. A., Dong H., Bai W., Nicosia S. V., Khochbin S., Bepler G., Zhang X. (2012) Depletion of HDAC6 enhances cisplatin-induced DNA damage and apoptosis in non-small cell lung cancer cells. PLoS One 7, e44265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Zhang M., Xiang S., Joo H. Y., Wang L., Williams K. A., Liu W., Hu C., Tong D., Haakenson J., Wang C., Zhang S., Pavlovicz R. E., Jones A., Schmidt K. H., Tang J., Dong H., Shan B., Fang B., Radhakrishnan R., Glazer P. M., Matthias P., Koomen J., Seto E., Bepler G., Nicosia S. V., Chen J., Li C., Gu L., Li G. M., Bai W., Wang H., Zhang X. (2014) HDAC6 deacetylates and ubiquitinates MSH2 to maintain proper levels of MutSα. Mol. Cell 55, 31–46 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Gu L., Ensor C. M., Li G. M. (2012) In vitro DNA mismatch repair in human cells. Methods Mol. Biol. 920, 135–147 [DOI] [PubMed] [Google Scholar]
- 20. Yuan F., Song L., Liu F., Gu L., Zhang Y. (2012) Eukaryotic DNA mismatch repair in vitro. Methods Mol. Biol. 920, 149–162 [DOI] [PubMed] [Google Scholar]
- 21. Holmes J. Jr., Clark S., Modrich P. (1990) Strand-specific mismatch correction in nuclear extracts of human and Drosophila melanogaster cell lines. Proc. Natl. Acad. Sci. U.S.A. 87, 5837–5841 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Gavin A. C., Bösche M., Krause R., Grandi P., Marzioch M., Bauer A., Schultz J., Rick J. M., Michon A. M., Cruciat C. M., Remor M., Höfert C., Schelder M., Brajenovic M., Ruffner H., Merino A., Klein K., Hudak M., Dickson D., Rudi T., Gnau V., Bauch A., Bastuck S., Huhse B., Leutwein C., Heurtier M. A., Copley R. R., Edelmann A., Querfurth E., Rybin V., Drewes G., Raida M., Bouwmeester T., Bork P., Seraphin B., Kuster B., Neubauer G., Superti-Furga G. (2002) Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 [DOI] [PubMed] [Google Scholar]
- 23. Smith J. A., Bannister L. A., Bhattacharjee V., Wang Y., Waldman B. C., Waldman A. S. (2007) Accurate homologous recombination is a prominent double-strand break repair pathway in mammalian chromosomes and is modulated by mismatch repair protein Msh2. Mol. Cell. Biol. 27, 7816–7827 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Christmann M., Tomicic M. T., Kaina B. (2002) Phosphorylation of mismatch repair proteins MSH2 and MSH6 affecting MutSα mismatch-binding activity. Nucleic Acids Res. 30, 1959–1966 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Sadoul K., Boyault C., Pabion M., Khochbin S. (2008) Regulation of protein turnover by acetyltransferases and deacetylases. Biochimie 90, 306–312 [DOI] [PubMed] [Google Scholar]
- 26. Abdel-Hafiz H. A., Horwitz K. B. (2014) Post-translational modifications of the progesterone receptors. J. Steroid Biochem. Mol. Biol. 140, 80–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Masih P. J., Kunnev D., Melendy T. (2008) Mismatch Repair proteins are recruited to replicating DNA through interaction with Proliferating Cell Nuclear Antigen (PCNA). Nucleic Acids Res. 36, 67–75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Umar A., Buermeyer A. B., Simon J. A., Thomas D. C., Clark A. B., Liskay R. M., Kunkel T. A. (1996) Requirement for PCNA in DNA mismatch repair at a step preceding DNA resynthesis. Cell 87, 65–73 [DOI] [PubMed] [Google Scholar]
- 29. Zhu W., Ukomadu C., Jha S., Senga T., Dhar S. K., Wohlschlegel J. A., Nutt L. K., Kornbluth S., Dutta A. (2007) Mcm10 and And-1/CTF4 recruit DNA polymerase α to chromatin for initiation of DNA replication. Genes Dev. 21, 2288–2299 [DOI] [PMC free article] [PubMed] [Google Scholar]