Abstract
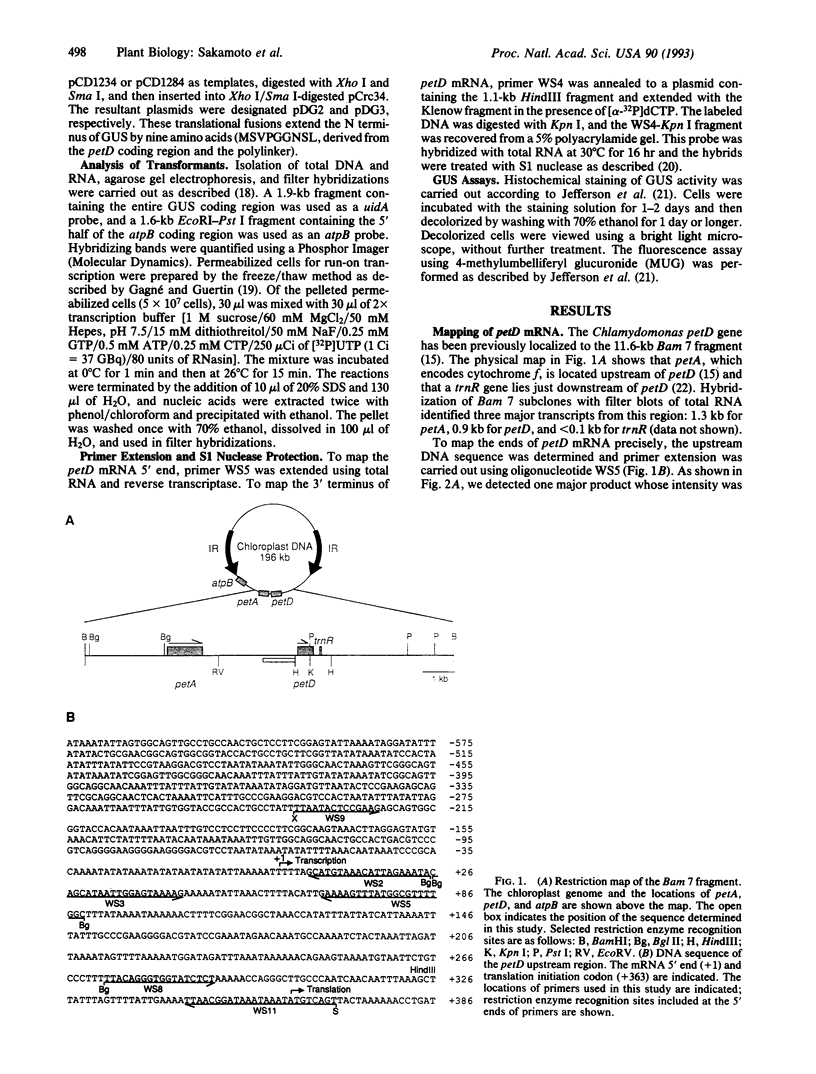
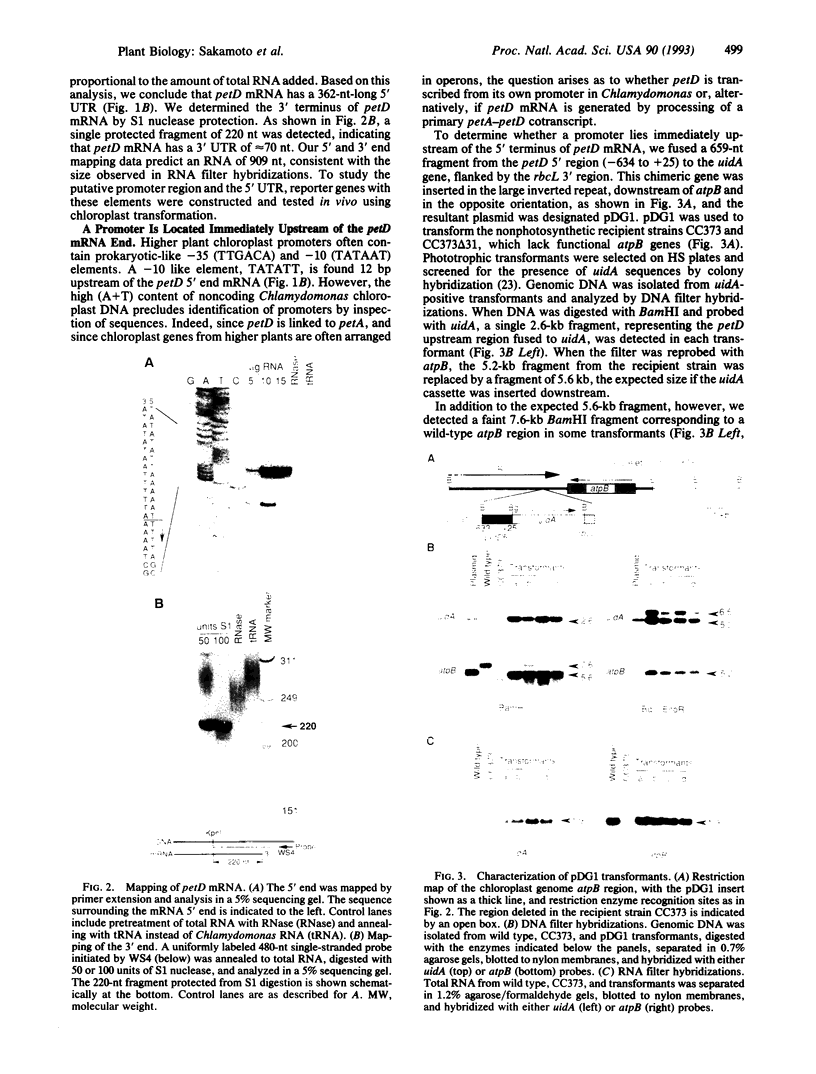
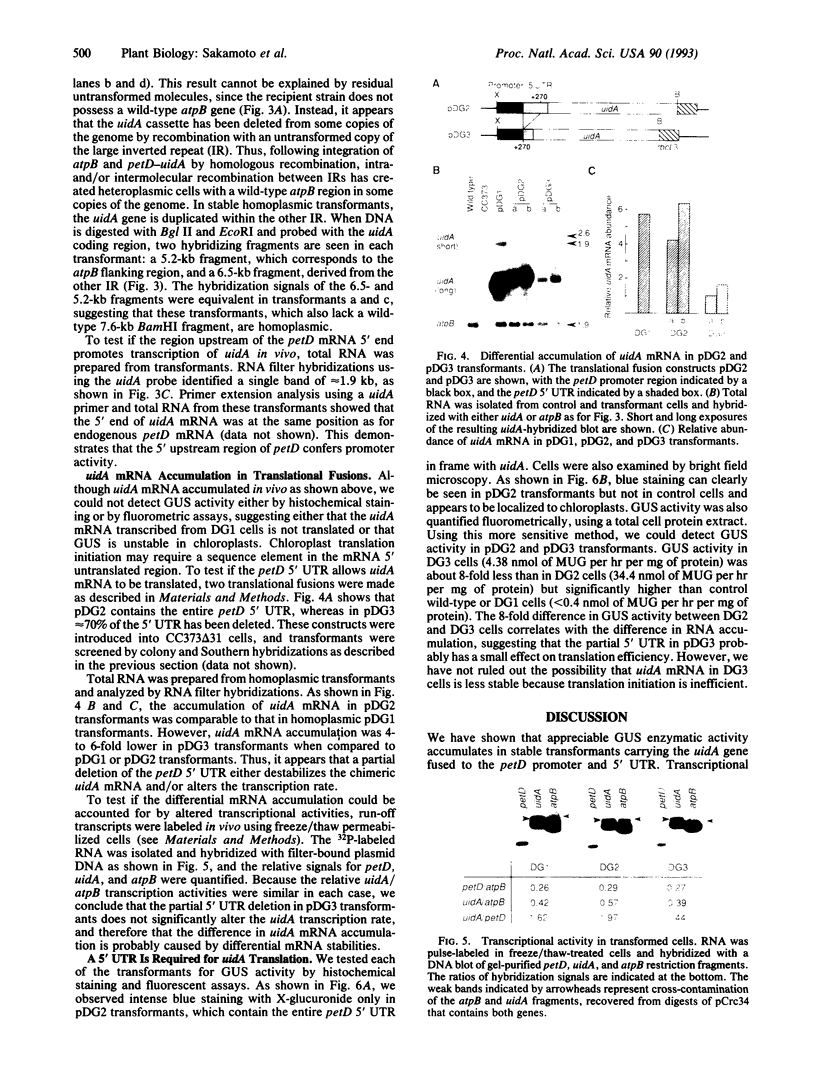
We have used the Escherichia coli beta-glucuronidase (uidA) gene as a reporter gene to localize the promoter and analyze the function of the 5' untranslated region (UTR) of the Chlamydomonas chloroplast petD gene. Using particle bombardment, petD-uidA transcriptional and translational fusion genes were introduced into the chloroplast genome in the large inverted repeat flanking the atpB gene. In transformants carrying a petD-uidA transcriptional fusion, uidA mRNA accumulated but was not translated. However, in a translational fusion that included the entire petD 5' UTR, uidA mRNA accumulated and a high level of beta-glucuronidase activity was detected. When approximately 70% of the petD 5' UTR was deleted from the translational fusion, uidA mRNA accumulation and beta-glucuronidase activity decreased 4- to 6-fold and 8-fold, respectively. Run-on transcription assays demonstrated that all strains transcribe the uidA gene at equivalent rates. Our results show that sequences essential for translation reside in the petD 5' UTR and also that sequences within the 5' UTR directly or indirectly affect mRNA stability. The expression of beta-glucuronidase under the control of chloroplast transcriptional and translational signals will facilitate further studies of chloroplast gene regulatory mechanisms.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Blowers A. D., Ellmore G. S., Klein U., Bogorad L. Transcriptional analysis of endogenous and foreign genes in chloroplast transformants of Chlamydomonas. Plant Cell. 1990 Nov;2(11):1059–1070. doi: 10.1105/tpc.2.11.1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boynton J. E., Gillham N. W., Harris E. H., Hosler J. P., Johnson A. M., Jones A. R., Randolph-Anderson B. L., Robertson D., Klein T. M., Shark K. B. Chloroplast transformation in Chlamydomonas with high velocity microprojectiles. Science. 1988 Jun 10;240(4858):1534–1538. doi: 10.1126/science.2897716. [DOI] [PubMed] [Google Scholar]
- Büschlen S., Choquet Y., Kuras R., Wollman F. A. Nucleotide sequences of the continuous and separated petA, petB and petD chloroplast genes in Chlamydomonas reinhardtii. FEBS Lett. 1991 Jun 24;284(2):257–262. doi: 10.1016/0014-5793(91)80698-3. [DOI] [PubMed] [Google Scholar]
- Dieckmann C. L., Mittelmeier T. M. Nuclearly-encoded CBP1 interacts with the 5' end of mitochondrial cytochrome b pre-mRNA. Curr Genet. 1987;12(6):391–397. doi: 10.1007/BF00434815. [DOI] [PubMed] [Google Scholar]
- Drapier D., Girard-Bascou J., Wollman F. A. Evidence for Nuclear Control of the Expression of the atpA and atpB Chloroplast Genes in Chlamydomonas. Plant Cell. 1992 Mar;4(3):283–295. doi: 10.1105/tpc.4.3.283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gagné G., Guertin M. The early genetic response to light in the green unicellular alga Chlamydomonas eugametos grown under light/dark cycles involves genes that represent direct responses to light and photosynthesis. Plant Mol Biol. 1992 Feb;18(3):429–445. doi: 10.1007/BF00040659. [DOI] [PubMed] [Google Scholar]
- Gay D. A., Sisodia S. S., Cleveland D. W. Autoregulatory control of beta-tubulin mRNA stability is linked to translation elongation. Proc Natl Acad Sci U S A. 1989 Aug;86(15):5763–5767. doi: 10.1073/pnas.86.15.5763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldschmidt-Clermont M. Transgenic expression of aminoglycoside adenine transferase in the chloroplast: a selectable marker of site-directed transformation of chlamydomonas. Nucleic Acids Res. 1991 Aug 11;19(15):4083–4089. doi: 10.1093/nar/19.15.4083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Jefferson R. A., Kavanagh T. A., Bevan M. W. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987 Dec 20;6(13):3901–3907. doi: 10.1002/j.1460-2075.1987.tb02730.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kindle K. L. High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A. 1990 Feb;87(3):1228–1232. doi: 10.1073/pnas.87.3.1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kindle K. L., Richards K. L., Stern D. B. Engineering the chloroplast genome: techniques and capabilities for chloroplast transformation in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A. 1991 Mar 1;88(5):1721–1725. doi: 10.1073/pnas.88.5.1721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klaff P., Gruissem W. Changes in Chloroplast mRNA Stability during Leaf Development. Plant Cell. 1991 May;3(5):517–529. doi: 10.1105/tpc.3.5.517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein U., De Camp J. D., Bogorad L. Two types of chloroplast gene promoters in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A. 1992 Apr 15;89(8):3453–3457. doi: 10.1073/pnas.89.8.3453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuchka M. R., Goldschmidt-Clermont M., van Dillewijn J., Rochaix J. D. Mutation at the Chlamydomonas nuclear NAC2 locus specifically affects stability of the chloroplast psbD transcript encoding polypeptide D2 of PS II. Cell. 1989 Sep 8;58(5):869–876. doi: 10.1016/0092-8674(89)90939-2. [DOI] [PubMed] [Google Scholar]
- Marsh J. L., Erfle M., Wykes E. J. The pIC plasmid and phage vectors with versatile cloning sites for recombinant selection by insertional inactivation. Gene. 1984 Dec;32(3):481–485. doi: 10.1016/0378-1119(84)90022-2. [DOI] [PubMed] [Google Scholar]
- Monod C., Goldschmidt-Clermont M., Rochaix J. D. Accumulation of chloroplast psbB RNA requires a nuclear factor in Chlamydomonas reinhardtii. Mol Gen Genet. 1992 Feb;231(3):449–459. doi: 10.1007/BF00292715. [DOI] [PubMed] [Google Scholar]
- Newman S. M., Gillham N. W., Harris E. H., Johnson A. M., Boynton J. E. Targeted disruption of chloroplast genes in Chlamydomonas reinhardtii. Mol Gen Genet. 1991 Nov;230(1-2):65–74. doi: 10.1007/BF00290652. [DOI] [PubMed] [Google Scholar]
- Rochaix J. D., Kuchka M., Mayfield S., Schirmer-Rahire M., Girard-Bascou J., Bennoun P. Nuclear and chloroplast mutations affect the synthesis or stability of the chloroplast psbC gene product in Chlamydomonas reinhardtii. EMBO J. 1989 Apr;8(4):1013–1021. doi: 10.1002/j.1460-2075.1989.tb03468.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shepherd H. S., Boynton J. E., Gillham N. W. Mutations in nine chloroplast loci of Chlamydomonas affecting different photosynthetic functions. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1353–1357. doi: 10.1073/pnas.76.3.1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sieburth L. E., Berry-Lowe S., Schmidt G. W. Chloroplast RNA Stability in Chlamydomonas: Rapid Degradation of psbB and psbC Transcripts in Two Nuclear Mutants. Plant Cell. 1991 Feb;3(2):175–189. doi: 10.1105/tpc.3.2.175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stern D. B., Radwanski E. R., Kindle K. L. A 3' stem/loop structure of the Chlamydomonas chloroplast atpB gene regulates mRNA accumulation in vivo. Plant Cell. 1991 Mar;3(3):285–297. doi: 10.1105/tpc.3.3.285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takahashi Y., Goldschmidt-Clermont M., Soen S. Y., Franzén L. G., Rochaix J. D. Directed chloroplast transformation in Chlamydomonas reinhardtii: insertional inactivation of the psaC gene encoding the iron sulfur protein destabilizes photosystem I. EMBO J. 1991 Aug;10(8):2033–2040. doi: 10.1002/j.1460-2075.1991.tb07733.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turmel M., Boulanger J., Bergeron A. Nucleotide sequence of the chloroplast petD gene of Chlamydomonas eugametos. Nucleic Acids Res. 1989 May 11;17(9):3593–3593. doi: 10.1093/nar/17.9.3593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu W., Spreitzer R. J. Sequences of trnR-ACG and petD that contain a tRNA-like element within the chloroplast genome of Chlamydomonas reinhardtii. Nucleic Acids Res. 1991 Feb 25;19(4):957–957. doi: 10.1093/nar/19.4.957. [DOI] [PMC free article] [PubMed] [Google Scholar]