Abstract
Idiosyncratic drug reactions (IDRs) are rare, somewhat dose-independent, patient-specific and hard to predict. Human leukocyte antigens (HLAs) are the major histocompatibility complex (MHC) in humans, are highly polymorphic and are associated with specific IDRs. Therefore, it is important to identify potential drug-HLA associations so that individuals who would develop IDRs can be identified before drug exposure. We harvested the associations between drugs and class I HLAs from the literature. Molecular docking was used to explore the known associations. From the analysis of docking scores between the 17 drugs and 74 class I HLAs, it was observed that the significantly associated drug-HLA pairs had statistically lower docking scores than those not reported to be significantly associated (t-test p < 0.05). This indicates that molecular docking can be utilized for screening drug-HLA interactions and predicting potential IDRs. Examining the binding modes of drugs in the docked HLAs suggested several distinct binding sites inside class I HLAs, expanding our knowledge of the underlying interaction mechanisms between drugs and HLAs.
Keywords: idiosyncratic drug reaction, human leukocyte antigen, docking, personalized medicine, adverse drug reaction, modelling
Introduction
Idiosyncratic drug reactions (IDRs), known as type B adverse drug reactions (ADRs) [1, 2], occur in only a small number of people within the normal clinical dose range. They are patient-specific and thus hard to predict [2, 3]. Due to their rare incidence, they are usually identified after the drug is approved and taken by a large number of patients. However, such IDRs can lead to significant morbidity and mortality, which may result in drug withdrawal or restricted use. These occurrences of IDRs generally do not involve the drug’s pharmacologic effects and most of them are immune-related. Patient-specific immunological factors such as human leukocyte antigens (HLAs) and T-cell receptors (TCRs) have been shown to be associated with IDRs occurrence [4].
Human leukocyte antigens (HLAs) are the major histocompatibility complex (MHC) in humans. They are a set of genes located at the short arm of chromosome 6, are highly polymorphic and have different frequencies across countries and ethnicities [5, 6]. There are three HLA classes: I, II and III. Class I HLA genes, including HLA A, B and C alleles, are expressed on the surface of all somatic cells [7] which can present antigens to helper CD8+ T cell and activate natural killer cell for cytotoxicity [8]. Class II HLA genes, including HLA-D alleles, are found on the surface of antigen-presenting cells such as B cells and activated T cells. They can present antigens to helper CD4+ T cells to activate B cells for antibody synthesis [9]. Class III HLA genes are a part of the complement system which helps to clear pathogens out of the body [10].
The class I and II HLA proteins have similar structures, both containing a long binding groove which can present a peptide to the T-cell receptors (Supplementary Figure S1) [11, 12]. Drug molecules may bind in this groove via different mechanisms and alter the peptide specificity [13, 14]. Although different HLA alleles may have only 2–3 amino acid differences in their protein sequences, single mutations inside the binding groove can dramatically change the drug effect [15]. Different from class I HLA molecules, the binding groove of class II HLA proteins contains two chains, and usually one chain is detected and recorded [5, 16]. Therefore, in order to ensure the accuracy of the HLA protein models, only class I HLAs molecules were considered in this study.
Some HLA molecules are reported to associate with specific drugs to cause IDRs. HLA allele B*57:01 has been identified as an important biomarker for flucloxacillin-induced liver injury and abacavir-induced hypersensitivity [16, 17]. Studies also have showed the association between HLA-B*15:02 and carbamazepine-induced Stevens-Johnson Syndrome, an adverse cutaneous drug reaction causing the detachment of the epidermis from the dermis [18]. Carbamazepine may also lead to hypersensitivity when HLA-A*31:01 is present [19]. According to the IMGT/HLA database [5], there are more than 9,000 HLA alleles. Compared to the total number of marketed drugs and HLA molecules, the number of studied associations is very small. Since the drug-HLA associations can lead to life-threating adverse events, it is hoped that one can take advantage of a docking method to screen potential drug-HLA interactions for further wet-lab validations and use the results of such analyses to guide drug candidate selection, improve clinical trial design and post-market surveillance.
Materials and Methods
Literature collection
All published studies on the associations between HLAs and IDRs were manually collected from PubMed as of 03/24/2014. The studies were collected according to the following criteria: for each drug-HLA association (1) the drug is a single component, (2) the HLA is a single Class I allele with available protein sequence from IMGT/HLA database [5], (3) the drug-HLA association, either significant or non-significant, has a p-value which is provided or can be inferred from the report, and (4) the study investigated more than one individual.
This was followed by a curation of 140 distinct drug-HLA associations including 17 drugs and 74 HLAs and a collection of their lowest reported association p-values. For those drug-HLA pairs without a given association p-value, 2 by 2 contingency tables based on the case-control studies were created and then the association p-values using Fisher’s exact test were calculated.
Docking preparation
SMILES codes of the 17 drugs were obtained from DrugBank [20]. Their 3D structures were generated using Corina online demo (http://www.molecular-networks.com) [21]. Corina generates one conformation for each molecule with coordinate optimization [22]. In order to get the accurate structures of the 74 HLA proteins, a search was made of the entire Protein Data Bank (PDB) [23] using keywords “MHC” and “HLA” which obtained 353 Class I HLA crystal structures. Their sequences were compared with the 74 HLA sequences from the IMGT/HLA database [5] by BLAST+ 2.2.28 [24] and this resulted in the 18 crystal structures listed in Table 1 with the following criteria: (1) the crystal structure has 100% matched sequence identity towards the HLA alleles and (2) the structure of best resolution is selected if there is redundancy. The remaining 56 HLAs shown in Table 2 do not have available crystal structures from Protein Data Bank. Therefore, we aligned each of their protein sequences with the sequences of 353 Class I HLA crystal structures by BLAST+ 2.2.28 to identify their templates with highest identity (Table 2). Then we implemented automodel class in MODELLER 9.12 [25] to automatically build one homology model for each of the 56 HLAs with default parameters. The automodel class follows the comparative modeling procedure including model optimizations by the variable target function method (VTFM) and simulated annealing with molecular dynamics. To evaluate the homology models, Ramachandran plots [26] were generated by PROCHECK [27].
Table 1.
The list of 18 available HLA crystal structures selected from Protein Data Bank. All the crystal structures were matched at Chain A with 100% identity.
| HLA allele | PDB ID | Matched length | Resolution |
|---|---|---|---|
| A*02:01 | 3D25 | 274 | 1.3 |
| A*02:03 | 3OX8 | 275 | 2.1 |
| A*02:06 | 3OXR | 275 | 1.7 |
| A*02:07 | 3OXS | 275 | 1.7 |
| A*11:01 | 1X7Q | 275 | 1.5 |
| A*24:02 | 4F7T | 275 | 1.7 |
| A*68:01 | 1HSB | 270 | 1.9 |
| B*07:02 | 3VCL | 275 | 1.7 |
| B*15:01 | 1XR9 | 276 | 1.8 |
| B*18:01 | 4JQV | 278 | 1.5 |
| B*35:01 | 1XH3 | 276 | 1.5 |
| B*44:02 | 1M6O | 276 | 1.6 |
| B*44:03 | 3DX7 | 276 | 1.6 |
| B*51:01 | 1E27 | 276 | 2.2 |
| B*52:01 | 3W39 | 277 | 3.1 |
| B*57:01 | 3VRI | 276 | 1.6 |
| C*03:04 | 1EFX | 278 | 3.0 |
| C*04:01 | 1IM9 | 275 | 2.8 |
Table 2.
The list of 56 modeled HLA alleles and their modelling information. Only Chain A is considered. The statistics of residues in allowed regions from the Ramachandran plots are compared between the templates and homology models.
| HLA allele | Template | Homology model | |||
|---|---|---|---|---|---|
| PDB ID | Residues in allowed regions | Identity | Matched length | Residues in allowed regions | |
| A*02:10 | 1OGA | 99.6% | 99% | 273 | 99.6% |
| A*24:20 | 2BCK | 100.0% | 99% | 276 | 99.5% |
| A*26:01 | 1X7Q | 100.0% | 94% | 258 | 99.6% |
| A*26:02 | 4HX1 | 100.0% | 94% | 257 | 100.0% |
| A*30:01 | 3RL1 | 100.0% | 97% | 265 | 100.0% |
| A*31:01 | 3RL1 | 100.0% | 95% | 259 | 100.0% |
| A*32:01 | 2BCK | 100.0% | 91% | 254 | 100.0% |
| A*33:03 | 4HWZ | 100.0% | 95% | 261 | 100.0% |
| B*07:05 | 3VCL | 100.0% | 99% | 274 | 100.0% |
| B*13:01 | 3DX7 | 100.0% | 96% | 265 | 100.0% |
| B*13:02 | 3W39 | 100.0% | 96% | 264 | 100.0% |
| B*13:13 | 1SYV | 99.9% | 96% | 263 | 99.6% |
| B*15:02 | 1XR9 | 100.0% | 98% | 271 | 100.0% |
| B*15:10 | 1A9E | 100.0% | 97% | 268 | 100.0% |
| B*15:11 | 1XR9 | 100.0% | 99% | 274 | 99.9% |
| B*15:18 | 1A9E | 100.0% | 97% | 269 | 100.0% |
| B*15:25 | 1XR9 | 100.0% | 99% | 272 | 99.5% |
| B*15:27 | 1XR9 | 100.0% | 99% | 275 | 100.0% |
| B*15:42 | 1XR9 | 100.0% | 97% | 269 | 100.0% |
| B*15:58 | 1XR9 | 100.0% | 99% | 274 | 100.0% |
| B*18:02 | 4JQV | 100.0% | 99% | 275 | 100.0% |
| B*27:04 | 2A83 | 99.6% | 99% | 273 | 100.0% |
| B*27:07 | 1K5N | 99.9% | 98% | 271 | 99.9% |
| B*35:05 | 1A9E | 100.0% | 99% | 274 | 99.9% |
| B*35:31 | 1A9E | 100.0% | 97% | 268 | 100.0% |
| B*38:01 | 1E27 | 100.0% | 95% | 263 | 99.5% |
| B*38:02 | 1E27 | 100.0% | 95% | 262 | 99.9% |
| B*39:01 | 3BVN | 100.0% | 98% | 271 | 99.6% |
| B*40:01 | 3LN4 | 99.9% | 98% | 269 | 99.6% |
| B*40:02 | 3LN4 | 99.9% | 97% | 267 | 99.9% |
| B*40:03 | 3LN5 | 100.0% | 97% | 266 | 100.0% |
| B*40:06 | 3LN4 | 99.9% | 97% | 266 | 100.0% |
| B*41:01 | 3LN4 | 99.9% | 99% | 273 | 100.0% |
| B*46:01 | 1XR9 | 100.0% | 97% | 269 | 99.9% |
| B*48:01 | 1MI5 | 99.7% | 96% | 265 | 99.6% |
| B*53:01 | 1A1O | 100.0% | 99% | 275 | 99.5% |
| B*54:01 | 1A9E | 100.0% | 95% | 263 | 100.0% |
| B*55:02 | 1MI5 | 99.7% | 95% | 263 | 99.6% |
| B*56:01 | 1A9E | 100.0% | 96% | 265 | 100.0% |
| B*58:01 | 3VH8 | 99.9% | 98% | 270 | 99.6% |
| B*58:02 | 3VH8 | 99.9% | 97% | 268 | 100.0% |
| B*59:01 | 1E27 | 100.0% | 97% | 269 | 99.9% |
| B*73:01 | 3BVN | 100.0% | 92% | 256 | 100.0% |
| C*01:02 | 1EFX | 99.1% | 95% | 263 | 99.6% |
| C*02:10 | 1EFX | 99.1% | 96% | 266 | 99.6% |
| C*03:02 | 1EFX | 99.1% | 99% | 276 | 99.6% |
| C*03:03 | 1EFX | 99.1% | 99% | 277 | 99.5% |
| C*06:02 | 1IM9 | 100.0% | 96% | 262 | 100.0% |
| C*07:02 | 1EFX | 99.1% | 92% | 256 | 99.6% |
| C*07:18 | 1EFX | 99.1% | 92% | 256 | 99.6% |
| C*08:01 | 1EFX | 99.1% | 95% | 264 | 99.1% |
| C*08:03 | 1EFX | 99.1% | 95% | 263 | 99.6% |
| C*12:02 | 1EFX | 99.1% | 96% | 267 | 99.1% |
| C*12:03 | 1EFX | 99.1% | 96% | 266 | 99.5% |
| C*14:02 | 1EFX | 99.1% | 96% | 265 | 99.1% |
| C*14:03 | 1EFX | 99.1% | 96% | 266 | 99.5% |
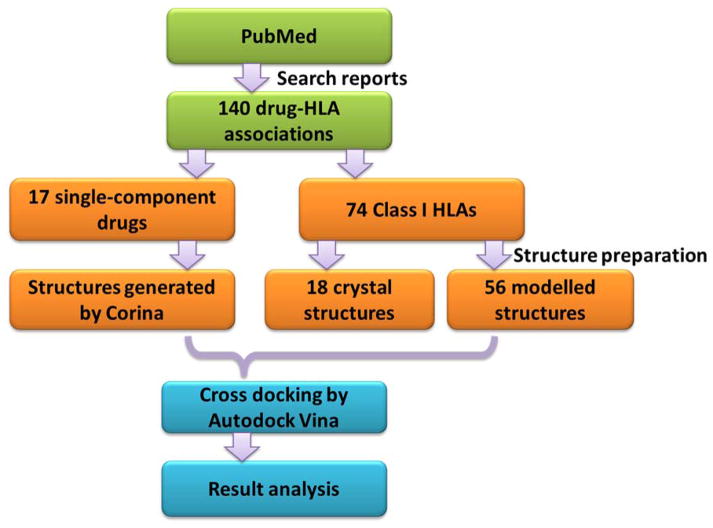
To help in the visualization of docking results, all 74 HLA structures were aligned to crystalline structure 3VRI with a root-mean-square deviation (RMSD) between 0.376 Å and 1.834 Å. An identical 36×34×36 Å3 binding box containing the entire binding groove was applied to all 74 HLA structures. To simply the problem, the embedded specific peptides inside the HLA binding grooves were not considered. The 17 drug molecules and the 74 HLA structures were prepared using AutoDock Tools 1.5.6 [28] with Gasteiger charges added and then cross-docked by AutoDock Vina 1.1.2 [29] with random seed set to 10,000 and other default parameters on Linux clusters. For each drug-HLA pair, the lowest docking score with its corresponding docking pose was adopted from 8 or more predicted poses which resulted in a matrix of 17×74 docking scores for subsequent analyses. The study design and overview of the workflow are depicted in Figure 1.
Figure 1.
The workflow. 140 drug-HLA associations including 17 single-component drugs and 74 Class I HLAs were collected from PubMed using 3 criteria. The drug structures were harvested from DrugBank and HLA structures were either crystal structures directly from Protein Data Bank by selecting the best resolutions or modelled from related crystal structures with highest identities. The entire HLA binding groove was set as the binding pocket and the cross-docking between 17 drugs and 74 HLAs was performed using AutoDock Vina.
Results and discussion
Drug-HLA associations
One hundred and forty drug-HLA associations among 17 drugs and 74 class I HLA alleles were identified and collected from the literature. The drugs, HLA alleles and p-values that were used for association significance estimation are summarized in Supplementary Table S1. Of the starting 140 associations, 63 are statistically significant with p-value < 0.05 from the literature, while the remaining 77 drug-HLA pairs are reported to not be significantly associated. This table was used for evaluation of the molecular docking approaches proposed here for identifying drug-HLA associations.
Homology modeling
The 56 modelled HLAs along with their templates are listed in Table 2. Since Chain A of Class I HLAs covers the entire binding groove (Supplementary Figure S1) and IMGT/HLA database only contains Chain A’s exact sequences, it is the only chain considered for homology modelling. We see the 56 HLAs share 91% or higher identities with their templates, indicating we can get reliable structures for them via homology modelling. The Ramachandran plots [26] of the 56 HLA structures which analyze the phi-psi torsion angles for each residue were generated (Supplementary Figure S2) and statistics of residues in allowed regions [30] for both the modelled structures and their templates are shown in Table 2. Since all the modelled structures contain 99.1% or higher residues in the allowed regions in the Ramachandran plots which are comparable to their templates, we believe the 56 homology models meet the quality for molecular docking.
Molecular docking
Each of the 17 drugs was docked into each of the 74 HLAs. The work flow is shown in Figure 1 with the details described in the Methods section. The docking scores were used to estimate the goodness of interaction between the drug and HLA allele, and the results (17×74=1,258 docking scores) are listed in Supplementary Table S2. Here, a lower docking score denotes a prediction of higher binding affinity. The 1,258 drug-HLA pairs were categorized into three groups according to the previous drug-HLA association table collected from the literature. The case group had 63 drug-HLA pairs that are significantly associated. The control group consisted of 77 pairs that are reported to have no significant associations, which mean the specific HLA allele doesn’t significantly affect the chance to develop the IDR with the specific drug. The remaining 1,118 drug-HLA pairs had no report to evaluate whether they are significantly associated or not and were termed as an unknown group. The docking results of the three groups of drug-HLAs were compared using the boxplot plotted in Figure 2.
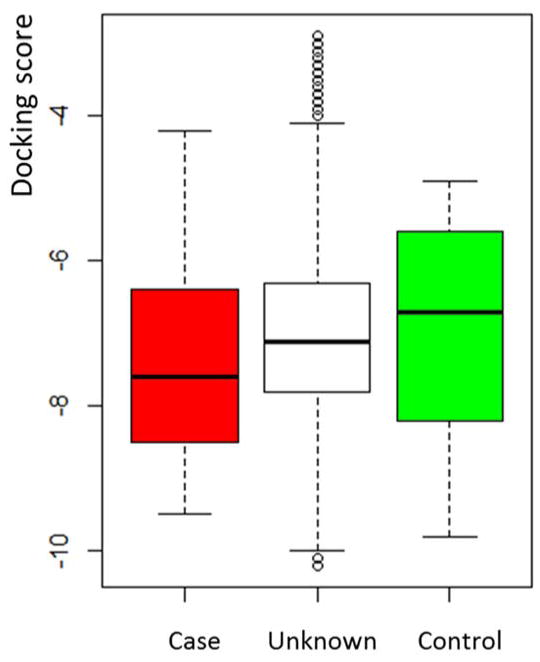
Figure 2.
The boxplot of docking scores in three groups. Case group/red: drug-HLA pairs with evidence showing the associations are significant (association p < 0.05); control group/green: drug-HLA pairs with evidence showing there are no significant associations (association p ≥ 0.05); unknown group/white: drug-HLA pairs without association studies. The docking scores are significantly different between the case and control group (t-test p = 0.045 < 0.05).
The case group (the red box) has lower docking scores (median score = −7.6) than the control group (median score = −6.7) (the green box) as shown in Figure 2, indicating molecular docking could differentiate the significant drug-HLA associations (case group) from the non-significant group (control group). In addition, the docking scores between the case and control group are significantly different with t-test p-value = 0.045 < 0.05 (Mann-Whitney U test p-value = 0.048 < 0.05). Interestingly, the unknown group has docking scores (median score = −7.1) between the case and control groups. There are no clinical data available for estimating associations of this group of drug-HLA pairs, so the speculation is that this group might be a mixture of both significant and non-significant associations based on these docking results.
In addition to p-values, the drug-HLA associations can also be evaluated by odds ratios (ORs). If the OR is larger than 1, the drug-HLA association increases the probability of IDR occurrence. In contrast, if the OR is smaller than 1, the drug-HLA association is protective towards the specific IDR [31], indicating patients carrying the specific HLA allele is less likely to get the IDR by taking the specific drug. Of all 63 significant drug-HLA associations reported in the literature, 19 have OR < 1 (see Supplementary Table S1). The molecular docking results of these 19 associations, as an independent group, were compared with the rest of the pairs in the case group as well as the control and unknown groups. The comparative analysis is shown in Supplementary Figure S3 and summarized in Supplementary Table S3. The drug-HLA associations with OR < 1 have lower median (−7.7) of docking scores than those with OR > 1 (−7.4). This observation led to our speculation that drugs in the drug-HLA pairs with OR > 1 may interact with the HLA as agonists to induce/enhance IDRs signaling pathways, while drugs in the drug-HLA pairs with OR < 1 may interact with the HLA as antagonists to reduce/inhibit IDRs signaling pathways. However, it is noted that 9 out of the 19 pairs with OR < 1 contain carbamazepine and, thus, this speculation must be met with caution.
Docking pose of abacavir
Abacavir appears to be the only IDR-inducing drug that has been co-crystalized with its corresponding HLA allele, B*57:01. However, in addition to the abacavir molecules, the existing crystal structures (PDB IDs: 3UPR [32], 3VRI and 3VRJ [13]) also have drug-specific peptides inside the HLA binding groove to stabilize the entire complexes. Because there are many possible peptides that could co-bind HLA molecules, peptides were not included in this molecular docking. To estimate the influence of neglecting co-binding peptides in molecular docking, abacavir binding positions were compared between the X-ray structures and the docked complex. As a control, we docked abacavir into HLA-B*57:01 (3VRI) keeping the original peptide. The docking results of abacavir with the crystal structures were superimposed and shown in Figure 3. Although the binding site was not specified and the drug-specific peptides were not included during the docking preparation, abacavir was successfully docked in the F pocket, the same pocket with those crystal structures [12]. However, the orientation of the drug molecule was reversed from those in the crystal structures and the predicted position had RMSD values from 7.33 Å to 7.50 Å compared to the crystal structures (Figure 3A). When keeping the peptide, the predicted position had much less RMSD values from 0.97 Å to 1.29 Å with a correct orientation compared to the crystal structures (Figure 3B). It could be that the difference was caused by neglecting the co-binding peptides yet this did not affect the prediction of binding sites of drugs in HLAs using molecular docking. Although peptides may improve docking results, we weren’t able to consider the peptides due to little knowledge about peptides that can specifically co-bind with each drug towards each HLA. However, these results still suggest that molecular docking could be a useful technique for predicting potential binding sites of drug-HLA interactions.
Figure 3.
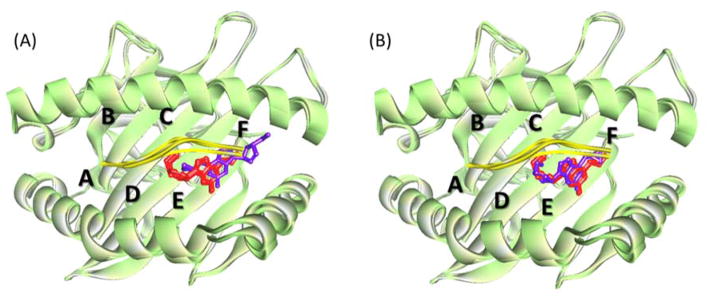
Superimposition of 3 abacavir-B*57:01 crystal structures 3UPR, 3VRI and 3VRJ (HLA: green, peptide: yellow and embedded abacavir: red) together with the docking predictions (purple) by Discovery Studio Visualizer 4.0. HLA pockets A–F are marked in the picture. (A) is the docking prediction without peptide, and (B) is the prediction considering peptide. From (A) we know although (1) the entire binding groove (pockets A–F) was selected as the docking pocket and (2) there was no consideration given to drug-specific peptides during the docking process, abacavir was successfully predicted to bind around the F pocket.
Distinct binding sites for drug-HLA interactions
To examine binding sites of drugs in different drug-HLA pairs, we superimposed all docked complexes for the 63 significantly associated drug-HLA pairs (see Figure 4). Although the same drug may appear in different sites within different HLA alleles, most carbamazepine molecules were bound among A–D pockets, which is consistent with the previous prediction [13]. Since researchers proposed that carbamazepine can directly interact with the HLA molecule and T-cell receptors without covalently binding to peptides (p-i concept) [33], the docking results additionally suggested that the HLA residues among A–D pockets may be important to the observed carbamazepine interaction with HLA molecules via the p-i concept. Meanwhile, two HLA-B*57:01 associated drugs, abacavir and flucloxacillin, both tended to bind around the F pocket, suggesting they could share some mechanistic similarities. Although the structural binding pattern of flucloxacillin is poorly understood, the latest studies showed that the immune reponse triggered by either abacavir or flucloxacillin is antigen-processing dependent [15, 34], which differs from the mechnism of carbamazepine [33]. Thus the predictions of the binding sites may serve as a preliminary screening tool for further validations and as a method to help identify possible mechanisms to such bindings.
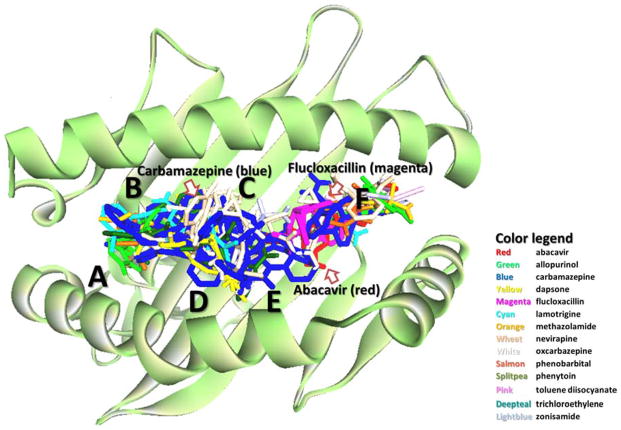
Figure 4.
Superimposition of all 14 drugs that have significant associations towards HLAs. One drug may associate with multiple HLAs and apprear more than one time. Only one HLA backbone (HLA-B*57:01) was shown. Pockets A–F were marked. B*57:01-associated drugs abacavir and flucloxacillin both bound around F pocket, while carbamazepine tended to bind among A–D pockets.
Limitations of molecular docking
There are several factors limiting the application of molecular docking and the interpretation of docking results. First, the reported drug-HLA associations are very few and thus one cannot get enough samples to differentiate associations with a high level of significance using the docking approach. There are a lot of binding assay studies exploring HLA binding preferences but those tests only consider peptides instead of drugs [35]. Therefore, it is not possible to find sufficient binding affinity data to validate the docking results. Second, the docking approach is a computational method to simulate drug-HLA interactions. Although this study demonstrated a predictive model, the real scenario might not be so trivial; the drug-HLA interactions happening inside a complex organism such as the human body are more complicated [14]. Third, the docking approach can be a fast and high through-put screening approach but it does not work for deciphering details of the interactions. Molecular docking followed by molecular dynamics/quantum dynamics might improve prediction accuracy and enhance elucidation of drug-HLA interactions. Furthermore, the unreported drug-HLA associations predicted by docking will need to be confirmed in the wet lab.
Conclusion
This work identified and curated the associations between drugs and class I HLAs from the literature and demonstrated that molecular docking could differentiate the significant drug-HLA associations that lead to IDRs from those non-significant ones. Future work will explore drug-HLA combinations identified by molecular docking for which there have been no reports of such interactions; such endeavors will include validations using wet-lab approaches. If confirmed, such results could set the stage for specific examinations of patients with IDRs caused by the drugs in question. Such in silico approaches might improve multiple facets of drug discovery starting with drug selection wherein a company might identify certain genetic susceptibilities for a new compound. This could enable better decision making based on the expected efficacy and need for the drug versus the potential for IDRs in a subset of patients. Such predictions might help to improve clinical trial design, the development of a needed companion diagnostic and postmarket surveillance. In any case, such tools would move the field of personalized medicine forward and help to protect public health.
Supplementary Material
The association table between 17 drugs and 74 Class I HLAs with lowest association p-values collected from the literature.
Lowest docking scores between 17 drugs and 74 HLAs
Acknowledgments
We thank Dr. Cesar Compadre from University of Arkansas for Medical Sciences for his help to this project. This research was supported in part by an appointment to the Research Participation Program at the National Center for Toxicological Research (Heng Luo, Hui Wen Ng and Wenqian Zhang) administered by the Oak Ridge Institute for Science and Education through an interagency agreement between the U.S. Department of Energy and the U.S. Food and Drug Administration. This project was partially supported by grants from the National Center for Research Resources (P20RR016460) and the National Institute of General Medical Sciences (P20GM103429) from the National Institutes of Health.
List of abbreviations
- IDR
idiosyncratic drug reactions
- ADR
adverse drug reactions
- HLA
human leukocyte antigen
- MHC
major histocompatibility complex
- TCR
T-cell receptors
- PDB
Protein Data Bank
- RMSD
root-mean-square deviation
- OR
odds ratio
Footnotes
Authors’ contributions
HL, LS, WT, DM and HH designed the project. HL, TD, PZ and LY collected the data. HL, HN, WZ, MS, LS, WT, and HH analyzed the results. HL, LS, WT, DM and HH wrote the paper.
Conflict of Interest
Not declared.
Supplementary information contains Supplementary Figure S1–S3, Supplementary Table S3 and the explanations.
Disclaimer
The findings and conclusions in this article have not been formally disseminated by the US Food and Drug Administration (FDA) and should not be construed to represent the FDA.
Supportive/Supplementary Material
The data sets supporting the results of this article are included within the article and its supplementary information.
References
- 1.Pirmohamed M, Park BK. Genetic susceptibility to adverse drug reactions. Trends Pharmacol Sci. 2001;22(6):298–305. doi: 10.1016/s0165-6147(00)01717-x. [DOI] [PubMed] [Google Scholar]
- 2.Wilke RA, Lin DW, Roden DM, Watkins PB, Flockhart D, Zineh I, Giacomini KM, Krauss RM. Identifying genetic risk factors for serious adverse drug reactions: current progress and challenges. Nat Rev Drug Discov. 2007;6(11):904–16. doi: 10.1038/nrd2423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Uetrecht J. Idiosyncratic drug reactions: current understanding. Annu Rev Pharmacol Toxicol. 2007;47:513–39. doi: 10.1146/annurev.pharmtox.47.120505.105150. [DOI] [PubMed] [Google Scholar]; Uetrecht J. Prediction of a new drug’s potential to cause idiosyncratic reactions. Curr Opin Drug Discov Devel. 2001;4(1):55–9. [PubMed] [Google Scholar]
- 4.Uetrecht J, Naisbitt DJ. Idiosyncratic adverse drug reactions: current concepts. Pharmacol Rev. 2013;65(2):779–808. doi: 10.1124/pr.113.007450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Robinson J, Halliwell JA, McWilliam H, Lopez R, Parham P, Marsh SG. The IMGT/HLA database. Nucleic Acids Res. 2013;41(Database issue):D1222–7. doi: 10.1093/nar/gks949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Illing PT, Vivian JP, Purcell AW, Rossjohn J, McCluskey J. Human leukocyte antigen-associated drug hypersensitivity. Curr Opin Immunol. 2013;25(1):81–9. doi: 10.1016/j.coi.2012.10.002. [DOI] [PubMed] [Google Scholar]; Hassen E, Nahla G, Bouaouina N, Chouchane L. The human leukocyte antigen class I genes in nasopharyngeal carcinoma risk. Mol Biol Rep. 2010;37(1):119–26. doi: 10.1007/s11033-009-9548-9. [DOI] [PubMed] [Google Scholar]; Gonzalez-Galarza FF, Christmas S, Middleton D, Jones AR. Allele frequency net: a database and online repository for immune gene frequencies in worldwide populations. Nucleic Acids Res. 2011;39(Database issue):D913–9. doi: 10.1093/nar/gkq1128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Complete sequence and gene map of a human major histocompatibility complex. The MHC sequencing consortium. Nature. 1999;401(6756):921–3. doi: 10.1038/44853. [DOI] [PubMed] [Google Scholar]
- 8.Bushkin Y, Demaria S, Le JM, Schwab R. Physical association between the CD8 and HLA class I molecules on the surface of activated human T lymphocytes. Proc Natl Acad Sci USA. 1988;85(11):3985–9. doi: 10.1073/pnas.85.11.3985. [DOI] [PMC free article] [PubMed] [Google Scholar]; Spaggiari GM, Contini P, Carosio R, Arvigo M, Ghio M, Oddone D, Dondero A, Zocchi MR, Puppo F, Indiveri F, Poggi A. Soluble HLA class I molecules induce natural killer cell apoptosis through the engagement of CD8: evidence for a negative regulation exerted by members of the inhibitory receptor superfamily. Blood. 2002;99(5):1706–14. doi: 10.1182/blood.v99.5.1706. [DOI] [PubMed] [Google Scholar]
- 9.Mangalam A, Rodriguez M, David C. Role of MHC class II expressing CD4+ T cells in proteolipid protein(91-110)-induced EAE in HLA-DR3 transgenic mice. Eur J Immunol. 2006;36(12):3356–70. doi: 10.1002/eji.200636217. [DOI] [PubMed] [Google Scholar]; Poncet P, Arock M, David B. MHC class II-dependent activation of CD4+ T cell hybridomas by human mast cells through superantigen presentation. J Leukoc Biol. 1999;66(1):105–12. doi: 10.1002/jlb.66.1.105. [DOI] [PubMed] [Google Scholar]
- 10.Sim E, Cross SJ. Phenotyping of human complement component C4, a class-III HLA antigen. Biochem J. 1986;239(3):763–7. doi: 10.1042/bj2390763. [DOI] [PMC free article] [PubMed] [Google Scholar]; Holland MC, Lambris JD. The complement system in teleosts. Fish Shellfish Immunol. 2002;12(5):399–420. doi: 10.1006/fsim.2001.0408. [DOI] [PubMed] [Google Scholar]
- 11.Chelvanayagam G. A roadmap for HLA-A, HLA-B, and HLA-C peptide binding specificities. Immunogenetics. 1996;45(1):15–26. doi: 10.1007/s002510050162. [DOI] [PubMed] [Google Scholar]; Stern LJ, Brown JH, Jardetzky TS, Gorga JC, Urban RG, Strominger JL, Wiley DC. Crystal structure of the human class II MHC protein HLA-DR1 complexed with an influenza virus peptide. Nature. 1994;368(6468):215–21. doi: 10.1038/368215a0. [DOI] [PubMed] [Google Scholar]
- 12.Saper MA, Bjorkman PJ, Wiley DC. Refined structure of the human histocompatibility antigen HLA-A2 at 2.6 A resolution. J Mol Biol. 1991;219(2):277–319. doi: 10.1016/0022-2836(91)90567-p. [DOI] [PubMed] [Google Scholar]
- 13.Illing PT, Vivian JP, Dudek NL, Kostenko L, Chen Z, Bharadwaj M, Miles JJ, Kjer-Nielsen L, Gras S, Williamson NA, Burrows SR, Purcell AW, Rossjohn J, McCluskey J. Immune self-reactivity triggered by drug-modified HLA-peptide repertoire. Nature. 2012;486(7404):554–8. doi: 10.1038/nature11147. [DOI] [PubMed] [Google Scholar]
- 14.Bharadwaj M, Illing P, Theodossis A, Purcell AW, Rossjohn J, McCluskey J. Drug hypersensitivity and human leukocyte antigens of the major histocompatibility complex. Annu Rev Pharmacol Toxicol. 2012;52:401–31. doi: 10.1146/annurev-pharmtox-010611-134701. [DOI] [PubMed] [Google Scholar]
- 15.Chessman D, Kostenko L, Lethborg T, Purcell AW, Williamson NA, Chen Z, Kjer-Nielsen L, Mifsud NA, Tait BD, Holdsworth R, Almeida CA, Nolan D, Macdonald WA, Archbold JK, Kellerher AD, Marriott D, Mallal S, Bharadwaj M, Rossjohn J, McCluskey J. Human leukocyte antigen class I-restricted activation of CD8+ T cells provides the immunogenetic basis of a systemic drug hypersensitivity. Immunity. 2008;28(6):822–32. doi: 10.1016/j.immuni.2008.04.020. [DOI] [PubMed] [Google Scholar]
- 16.Mallal S, Nolan D, Witt C, Masel G, Martin AM, Moore C, Sayer D, Castley A, Mamotte C, Maxwell D, James I, Christiansen FT. Association between presence of HLA-B*5701, HLA-DR7, and HLA-DQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet. 2002;359(9308):727–32. doi: 10.1016/s0140-6736(02)07873-x. [DOI] [PubMed] [Google Scholar]
- 17.Daly AK, Donaldson PT, Bhatnagar P, Shen Y, Pe’er I, Floratos A, Daly MJ, Goldstein DB, John S, Nelson MR, Graham J, Park BK, Dillon JF, Bernal W, Cordell HJ, Pirmohamed M, Aithal GP, Day CP. HLA-B*5701 genotype is a major determinant of drug-induced liver injury due to flucloxacillin. Nat Genet. 2009;41(7):816–9. doi: 10.1038/ng.379. [DOI] [PubMed] [Google Scholar]; Mallal S, Phillips E, Carosi G, Molina JM, Workman C, Tomazic J, Jägel-Guedes E, Rugina S, Kozyrev O, Cid JF, Hay P, Nolan D, Hughes S, Hughes A, Ryan S, Fitch N, Thorborn D, Benbow A, Team P-S. HLA-B*5701 screening for hypersensitivity to abacavir. N Engl J Med. 2008;358(6):568–79. doi: 10.1056/NEJMoa0706135. [DOI] [PubMed] [Google Scholar]; Martin AM, Nolan D, Gaudieri S, Almeida CA, Nolan R, James I, Carvalho F, Phillips E, Christiansen FT, Purcell AW, McCluskey J, Mallal S. Predisposition to abacavir hypersensitivity conferred by HLA-B*5701 and a haplotypic Hsp70-Hom variant. Proc Natl Acad Sci USA. 2004;101(12):4180–5. doi: 10.1073/pnas.0307067101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chung WH, Hung SI, Hong HS, Hsih MS, Yang LC, Ho HC, Wu JY, Chen YT. Medical genetics: a marker for Stevens-Johnson syndrome. Nature. 2004;428(6982):486. doi: 10.1038/428486a. [DOI] [PubMed] [Google Scholar]
- 19.McCormack M, Alfirevic A, Bourgeois S, Farrell JJ, Kasperavičiūtė D, Carrington M, Sills GJ, Marson T, Jia X, de Bakker PI, Chinthapalli K, Molokhia M, Johnson MR, O’Connor GD, Chaila E, Alhusaini S, Shianna KV, Radtke RA, Heinzen EL, Walley N, Pandolfo M, Pichler W, Park BK, Depondt C, Sisodiya SM, Goldstein DB, Deloukas P, Delanty N, Cavalleri GL, Pirmohamed M. HLA-A*3101 and carbamazepine-induced hypersensitivity reactions in Europeans. N Engl J Med. 2011;364(12):1134–43. doi: 10.1056/NEJMoa1013297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, Maciejewski A, Arndt D, Wilson M, Neveu V, Tang A, Gabriel G, Ly C, Adamjee S, Dame ZT, Han B, Zhou Y, Wishart DS. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 2014;42:D1091–7. doi: 10.1093/nar/gkt1068. Database issue. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sadowski J, Gasteiger J, Klebe G. Comparison of automatic three-dimensional model builders using 639 X-ray structures. J Chem Inf Comput Sci. 1994;34(4):1000–1008. [Google Scholar]
- 22.Gasteiger J, Rudolph C, Sadowski J. Automatic generation of 3D-atomic coordinates for organic molecules. Tetrahedron Computer Methodology. 1990;3(6):537–547. [Google Scholar]
- 23.Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE. The Protein Data Bank. Nucleic Acids Res. 2000;28(1):235–42. doi: 10.1093/nar/28.1.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic Local Alignment Search Tool. J Mol Biol. 1990;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 25.Eswar N, Webb B, Marti-Renom MA, Madhusudhan MS, Eramian D, Shen MY, Pieper U, Sali A. Comparative protein structure modeling using Modeller. Curr Protoc Bioinformatics. 2006;Chapter 5(Unit 5):6. doi: 10.1002/0471250953.bi0506s15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ramachandran GN, Ramakrishnan C, Sasisekharan V. Stereochemistry of polypeptide chain configurations. Journal of molecular biology. 1963;7:95–9. doi: 10.1016/s0022-2836(63)80023-6. [DOI] [PubMed] [Google Scholar]
- 27.Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. Journal of applied crystallography. 1993;26(2):283–291. [Google Scholar]
- 28.Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem. 2009;30(16):2785–91. doi: 10.1002/jcc.21256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Trott O, Olson AJ. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem. 2010;31(2):455–61. doi: 10.1002/jcc.21334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Morris AL, MacArthur MW, Hutchinson EG, Thornton JM. Stereochemical quality of protein structure coordinates. Proteins. 1992;12(4):345–64. doi: 10.1002/prot.340120407. [DOI] [PubMed] [Google Scholar]
- 31.Kang HR, Jee YK, Kim YS, Lee CH, Jung JW, Kim SH, Park HW, Chang YS, Jang IJ, Cho SH, Min KU, Lee KW. Positive and negative associations of HLA class I alleles with allopurinol-induced SCARs in Koreans. Pharmacogenet Genomics. 2011;21(5):303–7. doi: 10.1097/FPC.0b013e32834282b8. [DOI] [PubMed] [Google Scholar]
- 32.Ostrov DA, Grant BJ, Pompeu YA, Sidney J, Harndahl M, Southwood S, Oseroff C, Lu S, Jakoncic J, de Oliveira CA, Yang L, Mei H, Shi L, Shabanowitz J, English AM, Wriston A, Lucas A, Phillips E, Mallal S, Grey HM, Sette A, Hunt DF, Buus S, Peters B. Drug hypersensitivity caused by alteration of the MHC-presented self-peptide repertoire. Proc Natl Acad Sci US A. 2012;109(25):9959–64. doi: 10.1073/pnas.1207934109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wei CY, Chung WH, Huang HW, Chen YT, Hung SI. Direct interaction between HLA-B and carbamazepine activates T cells in patients with Stevens-Johnson syndrome. J Allergy Clin Immunol. 2012;129(6):1562–9. e5. doi: 10.1016/j.jaci.2011.12.990. [DOI] [PubMed] [Google Scholar]
- 34.Monshi MM, Faulkner L, Gibson A, Jenkins RE, Farrell J, Earnshaw CJ, Alfirevic A, Cederbrant K, Daly AK, French N, Pirmohamed M, Park BK, Naisbitt DJ. Human leukocyte antigen (HLA)-B*57:01-restricted activation of drug-specific T cells provides the immunological basis for flucloxacillin-induced liver injury. Hepatology. 2013;57(2):727–39. doi: 10.1002/hep.26077. [DOI] [PubMed] [Google Scholar]
- 35.Vita R, Zarebski L, Greenbaum JA, Emami H, Hoof I, Salimi N, Damle R, Sette A, Peters B. The immune epitope database 2.0. Nucleic Acids Res. 2010;38(Database issue):D854–62. doi: 10.1093/nar/gkp1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The association table between 17 drugs and 74 Class I HLAs with lowest association p-values collected from the literature.
Lowest docking scores between 17 drugs and 74 HLAs