Abstract
Endothelial cells are often present at inflammation sites. This is the case of endothelial cells of the blood-brain barrier (BBB) of patients afflicted with neurodegenerative disorders such as Alzheimer's, Parkinson's, or multiple sclerosis, as well as in cases of bacterial meningitis, trauma, or tumor-associated ischemia. Inflammation is a known modulator of gene expression through the activation of transcription factors, mostly NF-κB. RLIP76 (a.k.a. RALBP1), an ATP-dependent transporter of electrophile-glutathione conjugates, modulates BBB permeability through the regulation of tight junction function, cell adhesion, and exocytosis. Genes and pathways regulated by RLIP76 are transcriptional targets of tumor necrosis factor alpha (TNF-α) pro-inflammatory molecule, suggesting that RLIP76 may also be an inflammation target. To assess the effects of TNF-α on RLIP76, we faced the problem of choosing reference genes impervious to TNF-α. Since such genes were not known in human BBB endothelial cells, we subjected these to TNF-α, and measured by quantitative RT-PCR the expression of housekeeping genes commonly used as reference genes. We find most to be modulated, and analysis of several inflammation datasets as well as a metaanalysis of more than 5000 human tissue samples encompassing more than 300 cell types and diseases show that no single housekeeping gene may be used as a reference gene. Using three different algorithms, however, we uncovered a reference geneset impervious to TNF-α, and show for the first time that RLIP76 expression is induced by TNF-α and follows the induction kinetics of inflammation markers, suggesting that inflammation can influence RLIP76 expression at the BBB. We also show that MRP1 (a.k.a. ABCC1), another electrophile-glutathione transporter, is not modulated in the same cells and conditions, indicating that RLIP76 regulation by TNF-α is not a general property of glutathione transporters. The reference geneset uncovered herein should aid in future gene expression studies in inflammatory conditions of the BBB.
Introduction
Endothelial cells line the lumen of the entire circulatory system including all blood and lymphatic vessels, myocardium, renal glomeruli, and the blood-brain barrier (BBB). Endothelial cells also infiltrate tumors during the process of neoangiogenesis, putting them at the center stage of virtually all sites of inflammation. Endothelial cells have thus been reported to be exposed to the pro-inflammatory tumor necrosis factor α (TNF-α) cytokine in acute systemic conditions such as inflammation in response to bacteremia [1, 2], acute localized inflammation following myocardial infarction [1, 3], ischemic stroke [4], angioplasty [5], or post-ischemic reperfusion injury [6, 7]. Endothelial cells are also subjected to pro-inflammatory signals in chronic conditions such as atherosclerosis [8–10], Crohn's disease and ulcerative colitis [11–13], multiple sclerosis [14], periodontitis [15–18], rheumatoid arthritis [19, 20], type 2 diabetes mellitus [21, 22], and several cancers [23, 24]. At the blood-brain barrier (BBB), endothelial cells can be exposed to inflammation associated with brain-specific injuries and disorders such as Alzheimer's disease [25], bacterial meningitis [26], brain edema due to head trauma [27], ischemia and hypoxia [28, 29], tumors [30], epilepsy [31, 32], multiple sclerosis [28, 29], or Parkinson's disease [33, 34]. Inflammation modulates gene expression in many tissues, including endothelial cells [24, 35–38], via mechanisms that involve activation of transcription factors, most importantly NF-κB [39–43].
Quantification of gene expression modulation is usually carried out by means of quantitative real-time reverse transcription-coupled PCR (qRT-PCR), and amplification data are normalized to one or more reference genes to account for intra- and inter-experimental variability. A reference gene needs to be expressed in the tissue considered, and ought not to be modulated by the physiological or pathological variables studied. Examples of good reference human genes include; ACTB and GAPDH in prostate cancer [44]; GAPDH and YWHAZ in idiopathic pregnancy-derived placenta [45]; and HPRT1 and SDHA in sepsis [46]. As may be noticed from the above examples, these reference genes are housekeeping genes. Not all housekeeping genes, however, are suitable as reference genes as exemplified by the analysis of 81 ribosomal protein (Rpl) genes in 22 different tissues [47]. Although all Rpl genes were expressed in all tissues tested, and as such qualify as housekeeping genes, none could be used as a reference gene owing to high expression variability. Thus, and as may be surmised from the list of tissues/reference genes above, a gene or set of genes may serve as a reference in one tissue but may be inadequate for another. It may be inferred from these studies, therefore, that reference genes and genesets need to be determined for each tissue and in each physiological or pathological setting. This is particularly important in inflammatory conditions whereby the expression of many housekeeping genes can be modulated [46, 48, 49].
Due to the importance of endothelial cell inflammation in the physiopathology of the BBB, and owing to the widespread use of human ECV304 endothelial cells in BBB models (e.g. [50–61]), we set out to identify reference genes in these cells in a BBB model following treatment with TNF-α. We also probed more than 300 cell and disease types for universal reference genes, and specifically queried inflammation datasets for inflammation-impervious reference genes. Finally, we researched the impact of inflammation on the RLIP76 and MRP1 electrophile-glutathione conjugate transporters in BBB endothelial cells and discuss the implications of the observed modulation in BBB inflammatory disorders.
Materials & Methods
Reagents
Iscove's modified Dulbecco's medium (IMDM), Ham's F-12, new born calf serum (NCS), L-glutamine, penicillin/streptomycin, and trypsin were obtained from Invitrogen Life Technologies (Gibco™, Carlsbad, CA, USA). Heparin was purchased from MP Biomedicals (Irvine, CA, USA). TNF-α (hBA-158, sc-4564) was obtained from Santa Cruz Biotechnology, Inc. (Santa Cruz, CA, USA). Primers for qRT-PCR experiments were obtained from Metabion (Martinsried, Germany), and Brilliant SYBR Green QPCR Master Mix was from Stratagene (La Jolla, CA, USA). All other materials, including amphotericin B, transferring, and gelatin were obtained from Sigma (St. Louis, MO, USA) unless otherwise specified.
Cell Culture
Culture of human endothelial ECV304 cells in a BBB model was previously reported [53–56, 62]. For the purpose of this work, ECV304 cells were obtained from the European Collection of Cell Cultures (ECACC, Wiltshire, UK) and were cultured in C6 glial cells conditioned medium (C6CM). C6 cells were obtained from the German Cancer Research Center (DKFZ, Heidelberg, Germany) and were grown at 5% CO2 in 175 cm2 gelatin-coated tissue flasks at 37C in IF medium (1:1 mixture of IMDM and Ham's F-12), supplemented with 7.5% (v/v) NCS, 7 mM L-glutamine, 5 μg/ml transferrin, 0.5 U/ml heparin, 100 U/ml penicillin, 100 μg/ml streptomycin and 0.25 μg/ml amphotericin B. The supernatant of C6 cultures was collected every other day to yield the C6CM medium used to supplement the ECV304 cells. ECV304 cells were grown in IF medium/C6CM (1:1; v/v) at 37C in 5% CO2 for two weeks in six-well plates, then challenged with 5 ng/ml of TNF-α for 0, 2, 6, 24 or 48 hours prior to mRNA extraction.
RNA isolation and cDNA synthesis
Total RNA extraction including on-column DNase I digestion was performed using the NucleoSpin RNA II Kit (Macherey-Nagel, Dueren, Germany). To evaluate RNA purity and yield, all RNA samples were assessed on the Agilent Bioanalyzer 2100 using Nano LabChip analysis (Agilent Technologies, Palo Alto, California). Only those samples with an RNA integrity number exceeding 7 were used for reverse transcription (RT). Reverse transcription to cDNA was performed using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, USA) following manufacturer's instructions. Briefly, 1 μg of total RNA was reverse-transcribed in a 20 μl reaction using oligo(dT) primers and 2 U/μl RNase inhibitor (Invitrogen, Carlsbad, USA). All cDNA preparations were diluted at a ratio of 1:5 with RNase-free water prior to quantitative PCR.
Reference gene selection
To avoid any possible gene or species confusion [63], we list herein the NCBI Gene IDs of all tested genes. Nine housekeeping genes commonly used as reference genes were selected to evaluate their expression stability following treatment with TNF-α. These were ACTB (NCBI GeneID 60), B2M (NCBI GeneID 567), GAPDH (NCBI GeneID 2597), HPRT1 (NCBI GeneID 3251), PMM1 (NCBI GeneID 5372), PSMB6 (NCBI GeneID 5694), SDHA (NCBI GeneID 6389), TUBA1B (NCBI GeneID 10376), and YWHAZ (NCBI GeneID 7534). In addition, we researched the expression of the 76-kDa Ral-interacting protein (RLIP76 a.k.a. RALBP1; NCBI GeneID 10928) and multidrug resistance protein 1 (MRP1 a.k.a. ABCC1, NCBI GeneID 4363) glutathione conjugate transporters, as well as the following inflammation markers: intercellular adhesion molecule 1 (ICAM1; NCBI GeneID 3383), vascular cell adhesion molecule 1 (VCAM1; NCBI GeneID 7412), and monocyte chemoattractant protein-1 (MCP1 a.k.a. CCL2; NCBI GeneID 6347).
Quantitative real-time PCR of reverse transcription products (qRT-PCR)
Owing to the importance of transparent and comprehensive reporting of essential technical information in qPCR [64], we have followed the guidelines for the design and documentation of qPCR experiments as recently outlined [65], and provide a checklist detailing all information pertaining to the technical adequacy of used qPCR protocols (S1 Table).
All qRT-PCR reactions were performed in reaction mixtures containing 1 μl cDNA, 12.5 μl Brilliant SYBR Green QPCR Master Mix, primers, and nuclease-free water to 25 μl. Biological replicates were run in duplicate on an Mx3000P QPCR system (Stratagene La Jolla, CA, USA). Thermocycling conditions consisted of an initial polymerase activation step at 95°C for 10 min, followed by 45 cycles at 95°C for 30 sec, 55°C for 1 min, and 72°C for 1 min. Melting curves were generated in the range of 55 to 95°C to confirm single gene-specific peaks and to detect potential primer-dimer formation. No-template controls were included in each run to control for potential sample contaminations, and data were analyzed using the MxPro software v.4 (Stratagene). Baselines and thresholds were automatically set by the software and further tested through manual inspection. The crossing point of the amplification curve through the threshold represented the cycle threshold (Ct). Ct values were transformed to quantities based on the comparative Ct method that takes into consideration respective amplification efficiencies. Following appropriate formatting, data were imported into geNorm [66], NormFinder [67], and Bestkeeper [68] VBA algorithms.
Bioinformatic and statistical analyses
To query the modulation of gene expression by inflammation in patients, we searched the GEO database (http://www.ncbi.nlm.nih.gov/gds) for inflammation-related datasets. Three datasets encompassed genes of interest and included probes that passed quality criteria as previously established [69, 70]. These allowed us to probe for gene expression differences between healthy individuals and patients suffering of inflammatory bowel diseases (ulcerative colitis and Crohn's disease; GDS1615), idiopathic inflammatory myopathy (dermatomyositis, polymyositis, and inclusion body myositis; GDS2153), or rheumatoid arthritis (GDS3192). To test for gene expression variability in a wide array of human tissues, diseases, and drug treatments, we run an expression profiling metaanalysis for these genes in 5372 human tissues (ArrayExpress; E-MTAB-62; [71]). Samples were clustered in 15 or 96 metagroups and median expression of genes within each metagroup was compared to median expression across all samples. Statistical analyses were conducted using two-tailed unpaired t tests or one-way ANOVA with post-hoc Tukey tests cross-comparing all study groups, as appropriate. Data were considered significant at p < 0.05.
Results & Discussion
To identify suitable reference genes in endothelial cells of a human BBB model subjected to pro-inflammatory stimuli, we researched the gene expression stability, or lack thereof, of nine frequently used reference genes following treatment with TNF-α. These included β-actin (ACTB), β2-microglobulin (B2M), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), hypoxanthine phosphoribosyltransferase 1 (HPRT1), phosphomannomutase 1 (PMM1), proteasome subunit Y (PSMB6), succinate dehydrogenase complex subunit A (SDHA), α-tubulin (TUBA1B) and tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide (YWHAZ). The amplification profiles of each housekeeping gene were analyzed using three different algorithms implemented in the BestKeeper, geNorm, and NormFinder VBA applets. Once we identified a reference geneset, we used it as a Normalization Factor to assess expression modulation by TNF-α of additional test genes.
TNF-α induces the expression of inflammation markers in BBB endothelial cells
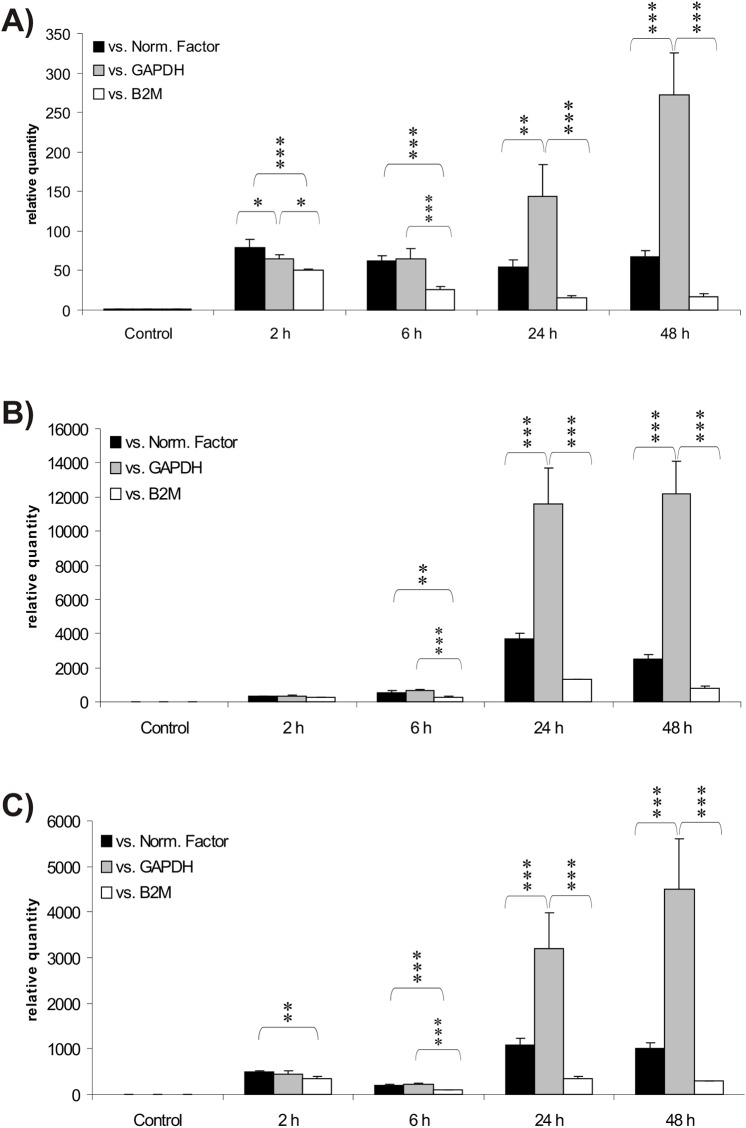
To test whether TNF-α treatment mimics inflammation in BBB endothelial cells, we carried out qRT-PCR experiments to follow the gene expression kinetics of endothelial/vascular inflammation markers. All three inflammation markers tested were strongly induced as early as 2 hours post-TNF-α treatment (Fig 1). Using the reference geneset identified below as a Normalization Factor, we find ICAM1 induction to be maximal at 2 hours (Fig 1A), whereas both VCAM1 and MCP1 were maximally induced at 24 hours (Fig 1B and 1C). All three markers remained highly expressed for the duration of the experiment, indicating that the experimental conditions used were commensurate with an inflammation model. As shown in Fig 1A–1C, fold induction of the three genes was overly exaggerated, or instead underestimated, by use of single housekeeping genes as reference genes. The large variations observed in ICAM1, VCAM1 and MCP1 gene expression levels following normalization with single housekeeping genes underscore the need for a stably expressed reference geneset to normalizing gene expression data.
Fig 1. TNF-α induces the expression of inflammation markers in blood-brain barrier endothelial cells.
Expression levels of ICAM1 (Panel A), VCAM1 (Panel B) and MCP1 (Panel C) were monitored by qRT-PCR prior (control) and 2 to 48 hours following TNF-α treatment. Expression levels were normalized against PSMB6 / HPRT1 as a reference geneset (Normalization Factor; black graphs), or against single genes, here GAPDH (grey graphs) or B2M (white graphs). Results are presented as arithmetic means ± standard deviations. Statistical significance: * signifies p < 0.05, ** p < 0.01, and *** p < 0.001; lack of asterisks denotes lack of a statistically significant difference.
Evaluation of candidate reference genes using BestKeeper
BestKeeper determines reference gene stability by employing pairwise correlation analysis of all candidate gene pairs and combines the best suited standards into a "BestKeeper index" [68]. A Pearson correlation coefficient (r) and probability value (p) are ascribed to each gene to describe the consistency between each candidate gene and the BestKeeper index. Using this algorithm, B2M exhibited a regulation inverse to that of the index, indicated by the negative correlation coefficient (r = -0.164). The other candidate genes showed a positive correlation with the index (Table 1). Descriptive statistics of the Ct values under the present experimental conditions, as computed by BestKeeper, are presented in Table 1. All Ct values were compared over the entire dataset, including control and treatment groups. An estimation of reference gene expression stability is provided by a comparison of standard deviations (SD values). Candidate reference genes ranking from the most to the least stable were SDHA ≈ HPRT1 > PMM1 ≥ PSMB6 > TUBA1B > YWHAZ ≥ ACTB > B2M > GAPDH. In this setup, B2M and GAPDH expression was found to be highly variable, arguing against using them as reference genes.
Table 1. Statistical output of the Bestkeeper analysis.
Candidate reference genes are listed according to increasing standard deviations (SD). Consistency between each candidate gene and the Bestkeeper index is described by the Pearson correlation coefficient (r) and the probability value (p), based on all candidate genes (n = 9) or after exclusion of GAPDH and B2M (n = 7). Abbreviations: Ar. Mean: arithmetic mean; Geo. Mean: geometric mean; BK: Bestkeeper index.
| Variable | SDHA | HPRT1 | TUBA1B | PSMB6 | ACTB | PPMM1 | YWHAZ | B2M | GAPDH | BK |
|---|---|---|---|---|---|---|---|---|---|---|
| n | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 |
| Geo M. | 20.39 | 20.59 | 15.79 | 20.00 | 14.90 | 22.89 | 17.44 | 17.52 | 16.01 | 18.22 |
| Ar. M. | 20.39 | 20.59 | 15.79 | 20.00 | 14.91 | 22.90 | 17.45 | 17.54 | 16.04 | 18.22 |
| Min Ct | 19.88 | 20.13 | 15.27 | 19.19 | 13.91 | 22.35 | 16.55 | 16.49 | 14.64 | 17.63 |
| Max Ct | 20.83 | 21.23 | 16.56 | 20.91 | 15.56 | 23.62 | 18.52 | 18.97 | 18.01 | 18.77 |
| SD ± Ct | 0.21 | 0.22 | 0.31 | 0.33 | 0.36 | 0.36 | 0.41 | 0.67 | 0.93 | 0.26 |
| CV % Ct | 1.03 | 1.06 | 1.99 | 1.65 | 2.43 | 1.56 | 2.35 | 3.79 | 5.80 | 1.45 |
| r (n = 9) | 0.305 | 0.919 | 0.654 | 0.862 | 0.869 | 0.745 | 0.925 | -0.164 | 0.854 | - |
| p (n = 9) | 0.191 | 0.001 | 0.002 | 0.001 | 0.001 | 0.001 | 0.001 | 0.487 | 0.001 | - |
| r (n = 7) | 0.258 | 0.935 | 0.732 | 0.875 | 0.827 | 0.781 | 0.924 | - | - | - |
| p (n = 7) | 0.273 | 0.001 | 0.001 | 0.001 | 0.001 | 0.001 | 0.001 | - | - | - |
Assessment of candidate reference genes using geNorm
geNorm is based on the principle that the expression ratio of two ideal reference genes is constant in all samples, regardless of experimental conditions [66]. Table 2 shows the ranking of all candidate reference genes according to their stability measure M (average pairwise variations). Genes with the lowest M values are characterized by the most stable expression. According to the geNorm analysis, the three most stable reference genes across all samples and experimental conditions were SDHA, PSMB6, and HPRT1. The order of the top two genes could not be further ranked by geNorm due to the intrinsic requirement of gene ratios for gene stability ranking in this methodology. Again, in agreement with BestKeeper analysis, B2M and GAPDH proved to be the most variably expressed genes under the experimental conditions tested (Table 2).
Table 2. Candidate reference genes ranked according to geNorm.
M values were calculated as average expression stability of reference genes during stepwise exclusion of the least stable control. V values were calculated as the pairwise variation between two sequential Normalization Factors.
| Ranking | Gene | M value | V value |
|---|---|---|---|
| 1/2 | SDHA/PSMB6 | 0.24 | - |
| 3 | HPRT1 | 0.27 | 0.086 |
| 4 | PPMM1 | 0.29 | 0.069 |
| 5 | TUBA1B | 0.32 | 0.067 |
| 6 | ACTB | 0.35 | 0.061 |
| 7 | YWHAZ | 0.39 | 0.06 |
| 8 | GAPDH | 0.51 | 0.109 |
| 9 | B2M | 0.62 | 0.107 |
Choosing the appropriate number of reference genes for qRT-PCR normalization involves a compromise between practical considerations and accuracy. geNorm permits to calculate the pairwise variations Vn/Vn+1 between two sequential Normalization Factors (NF-n and NF-n+1) to determine the added benefit of including additional reference genes for reliable normalization. geNorm thus allows to determine the minimum number of genes required to achieve the commonly accepted V-value threshold of 0.15, the maximum allowed for reference gene stability. In our study, the inclusion of a third reference gene yields a V-value V2/3 of 0.086 (significantly below the 0.15 cut-off value), indicating that two of the top three genes (SDHA, PSMB6, HPRT1) suffice as a Normalization Factor.
Ranking of candidate reference genes using NormFinder
The ranking of candidate genes provided by geNorm was further tested by a NormFinder analysis (Table 3). The most notable differences in the NormFinder test were the ranking of ACTB as the most stable gene, and the ranking of SDHA near the bottom. Again, as in the previous two analyses, GAPDH and B2M proved to be the least stably expressed of all tested genes. An analysis of promoter sequences of all housekeeping genes revealed the presence of NF-κB binding sites in GAPDH and B2M, providing a possible molecular basis for their modulation by TNF-α (~2-fold overrepresentation compared to the average random promoter, z-scores > 1; Table 4). Interestingly, the ranking of genes based on the significance of NF-κB binding sites within promoter sequences (Table 4) was very similar to the ranking of gene expression stability under TNF-α obtained with the NormFinder algorithm (Table 3). Thus, based on the NormFinder ranking, gene loci not modulated by TNF-α had no significant hits for NF-κB binding sites, whereas those modulated by TNF-α had the most hits. The genome-wide comparative analysis of NF-κB binding sites summarized in Table 4 further shows that NF-κB binding sites are underrepresented in PSMB6 and HPRT1 gene promoters, both when compared to the average of all genome promoters, or to the other housekeeping genes tested individually. While these data are consistent with the lack of modulation of PSMB6 and HPRT1 by TNF-α, they may or not reflect the activity of NF-κB on these two genes in vivo, which remains to be ascertained. These considerations notwithstanding, since PSMB6 and HPRT1 were the two housekeeping genes most consistently impervious to TNF-α, we used their average expression as a Normalization Factor for the remaining of this study.
Table 3. Candidate reference genes ranked according to NormFinder.
In this analysis, genes ranked 1–4 were deemed highly stable, whereas genes ranked 8–9 were deemed highly unstable.
| Ranking | Gene | Stability value |
|---|---|---|
| 1 | ACTB | 0.083 |
| 2 | HPRT1 | 0.092 |
| 3 | PSMB6 | 0.094 |
| 4 | TUBA1B | 0.107 |
| 5 | PPMM1 | 0.175 |
| 6 | YWHAZ | 0.186 |
| 7 | SDHA | 0.209 |
| 8 | B2M | 0.730 |
| 9 | GAPDH | 0.845 |
Table 4. NF-κB binding sites within housekeeping gene promoters.
2000 bp of 5' and 500 bp of 3' flanking sequences were searched for NF-κB binding sites (GGGRNNYYCC; whereby R is a purine and Y is a pyrimidine) and compared to the average binding site distribution of similarly-sized random genomic or promoter sequences. z-statistics show that B2M and GAPDH promoters contain significantly more NF-κB binding sites than the other housekeeping genes or the average genomic or promoter sequence, whereas NF-κB binding sites are underrepresented in PSMB6 and HPRT1.
| Reference Gene | # input sequences with match | # of matches in input | Expected in genome | SD | Over-representation vs genome | Z-score vs genome | Expected in promoters | SD | Over-representation vs promoters | Z-score vs promoters |
|---|---|---|---|---|---|---|---|---|---|---|
| PSMB6 | 1 | 1 | 2 | 1.41 | 0.5 | -1.06 | 3.06 | 1.75 | 0.33 | -1.46 |
| HPRT1 | 1 | 2 | 1.96 | 1.4 | 1.02 | -0.33 | 3 | 1.73 | 0.67 | -0.87 |
| ACTB | 1 | 4 | 2.06 | 1.44 | 1.94 | 1.06 | 3.16 | 1.78 | 1.27 | 0.19 |
| TUBA1B | 1 | 4 | 2.03 | 1.41 | 1.97 | 1.06 | 3.12 | 1.73 | 1.28 | 0.21 |
| PMM1 | 1 | 4 | 2.01 | 1.42 | 1.99 | 1.05 | 3.08 | 1.75 | 1.3 | 0.24 |
| YWHAZ | 1 | 5 | 2.05 | 1.43 | 2.44 | 1.71 | 3.13 | 1.77 | 1.6 | 0.77 |
| SDHA | 1 | 6 | 2.07 | 1.44 | 2.9 | 2.39 | 3.28 | 1.78 | 1.83 | 0.82 |
| GAPDH | 1 | 6 | 2.54 | 1.59 | 2.36 | 1.86 | 3.16 | 1.97 | 1.9 | 1.32 |
| B2M | 1 | 7 | 1.98 | 1.41 | 3.53 | 3.21 | 3.03 | 1.74 | 2.31 | 1.99 |
Comparison of the reference genes identified herein to those of other studies
To the best of our knowledge, there are only a few reported studies wherein reference genes were investigated in model systems of human endothelial cells undergoing inflammation. Similarly to our finding that HPRT1 is stably expressed in BBB endothelial cells, HPRT1 is also the gene most commonly reported to be impervious to pro-inflammatory stimuli in endothelial cells, as recently reported for human endothelial cells of the umbilical vein, choroid, and retina [72–74]. PSMB6, our second component of the Normalization Factor was also proposed as a reference gene in umbilical vein endothelial cells [75]. Other genes found here to be expressed at relatively stable levels after TNF-α treatment were also proposed as inflammation-stable reference genes in endothelial cells of the umbilical vein (e.g. ACTB [75], and YWHAZ [72]), or in endothelial cells of the lungs (e.g. TUBA1B [76]). However, contrary to this last study which also proposed GAPDH as a reference gene, we find GAPDH to be one of the least reliable reference genes. The discrepancy between the results of the studies may be due to the difference of tissues tested or may be due to the different stimuli used, which among other things can determine tissue-specific chromatin accessibility and ensuing gene transcriptional permissiveness [77, 78]. This calls for a cautionary note concerning the use of genes reported as good references in studies using similar cells from different tissues, or that were subjected to different physical or humoral environments. This further argues in favor of using the average of several genes as a reference set, preferably in comparable cells and tissues subjected to similar stimuli.
Lack of universal inflammation-impervious reference genes
To test whether genes uncovered here to be either impervious (e.g. PSMB6 and HPRT1) or sensitive to inflammation stimuli (e.g. B2M and GAPDH) behave similarly in other inflammatory conditions, we analyzed gene expression in healthy individuals and patient populations suffering from one of several inflammatory conditions. Similarly to our findings in TNF-α treated endothelial cells, PSMB6 expression was not modulated in rheumatoid arthritis (Fig 2A), dermatomyositis (Fig 2B), polymyositis (Fig 2B), and ulcerative colitis (Fig 2C), and HPRT1 expression was stable in rheumatoid arthritis (Fig 2A), all tested idiopathic inflammatory myopathies (Fig 2B), and in ulcerative colitis (Fig 2C). Also akin to findings herein, B2M expression was highly modulated in rheumatoid arthritis (Fig 2A) and in all tested idiopathic inflammatory myopathies (Fig 2B), and GAPDH expression was modulated in rheumatoid arthritis and dermatomyositis (Fig 2C). However, in contrast to our findings for these genes in TNF-α treated endothelial cells, PSMB6 was significantly modulated in inclusion body myositis (Fig 2B) and both PSMB6 and HPRT1 were modulated in Crohn's disease (Fig 2C). Moreover both B2M and GAPDH were stably expressed in inflammatory bowel diseases and GAPDH was in addition stably expressed in polymyositis and in inclusion body myositis (Fig 2C). These data show the limited "portability" of reference gene status, or lack thereof, from one inflammatory condition to another, which, therefore, ought to be independently determined for each tissue and pro-inflammatory stimulus involved.
Fig 2. Modulation of housekeeping gene expression in several inflammatory conditions.
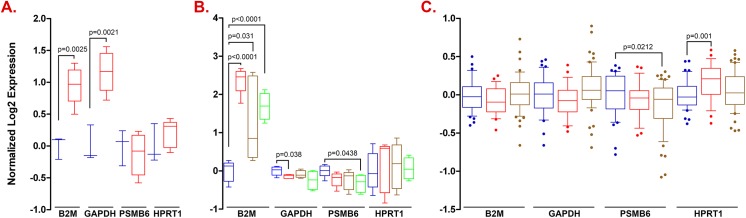
A. Expression was monitored in synovial fluid macrophages from healthy individuals (blue) or patients suffering from rheumatoid arthritis (red); B. Expression was monitored in skeletal muscle fibers from healthy individuals (blue), dermatomyositis patients (red), polymyositis patients (gold), and inclusion body myositis patients (green); C. Expression was monitored in peripheral blood mononuclear cells from healthy individuals (blue), patients suffering from ulcerative colitis (red), or of Crohn's disease (gold).
Lack of evidence supporting the concept of a universal reference gene or geneset
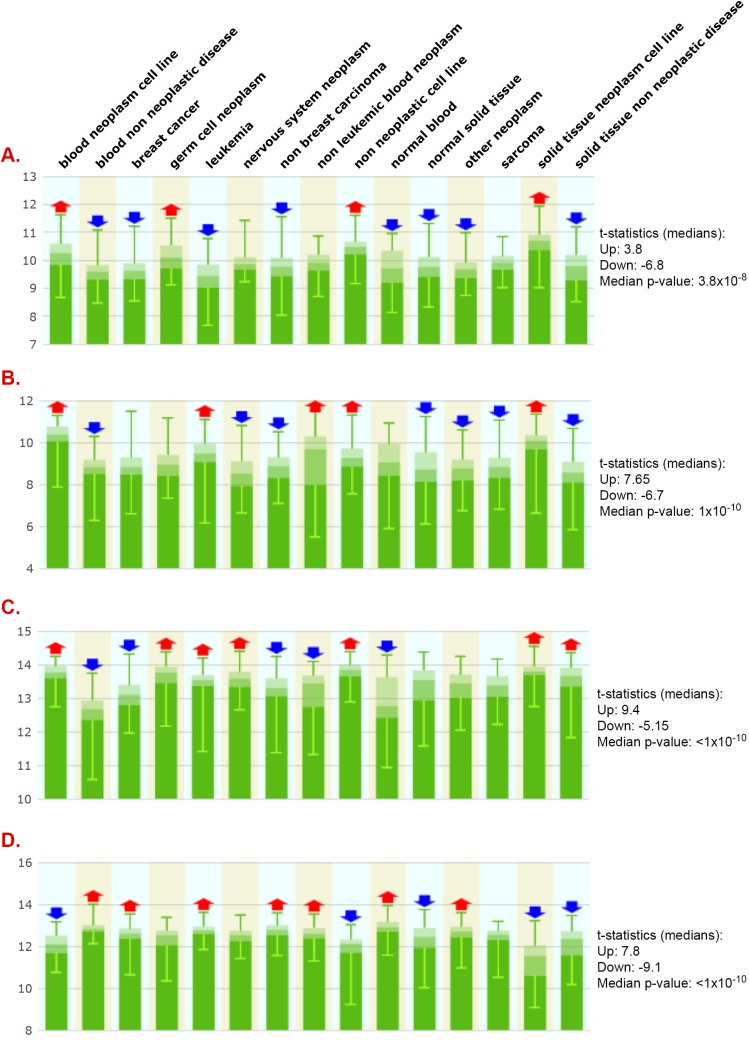
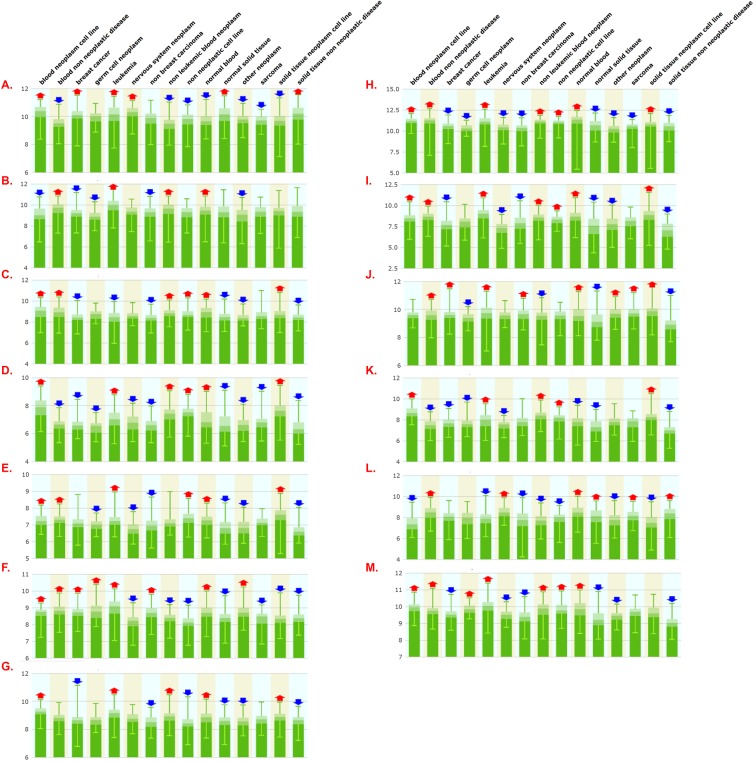
To further assess PSMB6, HPRT1, B2M, and GAPDH gene expression stability, we run a multivariate expression analysis encompassing 5,372 human tissue samples and 369 tissue and disease types. As shown in Fig 3, PSMB6 was again the least variable gene, followed by HPRT1 and B2M/GAPDH. However, as shown in Fig 3 and detailed in S2–S5 Tables, expression of all genes was highly variable in most metagroups, indicating that no gene is expressed stably enough to serve as a single reference gene in most cell types and conditions. To further test this premise, we also analyzed expression of 13 previously reported reference genes proposed to be stably expressed across different tissues based on in silico and molecular analyses [79]. As shown in Fig 4, all 13 genes show high gene expression variability in most 15 metagroups. Detailed analysis of the number of experiments wherein gene expression varies across the 369 tissue, cell, and disease types shows HPRT1 and SPG21 to be the least variably expressed genes (Table 5). However, cluster analysis in 96 metagroups shows SPG21 and HPRT1 expression to nonetheless significantly vary in 70 and 81 out of 96 metagroups, respectively (S6 and S7 Tables). Together, these data strongly support the need to validate reference genes in most tissues and conditions.
Fig 3. Gene expression variability of the PSMB6, HPRT1, B2M and GAPDH genes in human cells and diseases.
Gene expression of PSMB6 (A), HPRT1 (B), GAPDH (C), and B2M (D) was assessed across 5372 human tissue samples representing 369 tissues and cell types or diseases clustered into 15 meta-groups (indicated on top of figure). Red and blue arrows indicate expression significantly above (Up) or below (Down) median expression across 15 meta-groups, respectively. Medians of t-statistics and associated p-values are indicated. Ordinate axis: Log2 expression.
Fig 4. Gene expression variability of 'universal reference genes' in human cells and diseases.
Gene expression of previously reported universally stably expressed reference genes was monitored across 5372 human tissues samples representing 369 tissues and cell types or diseases clustered into 15 meta-groups (indicated on top of figure). Red and blue arrows indicate expression significantly above (Up) or below (Down) median expression within meta-group, respectively. p-values ranged from 10−2 to less than 10−10. Panels: A. ARL8B; B. CTBP1; C. CUL1; D. DIMT1L; E. FBXW2; F. GPBP1; G. LUC7L2; H. OAZ1; I. PAPOLA; J. SPG21; K. TRIM27; L. UBQLN1; M. ZNF-207.
Table 5. Modulation of gene expression by drugs or disease state across several tissue and cell types.
Expression of 13 previously reported 'universal reference genes' and of the most and least stably expressed genes uncovered in this study was monitored across 5372 human samples representing 369 different cell and tissue types, disease states and cell lines. Data show the number of studies documenting gene expression modulation for the considered parameters.
| Gene | Experiments | Cell types | Cell lines | Diseases | Compound treatment |
|---|---|---|---|---|---|
| ARL8B | 306 | 85 | 307 | 84 | 168 |
| CTBP1 | 343 | 90 | 301 | 99 | 315 |
| CUL1 | 297 | 71 | 213 | 77 | 114 |
| DIMT1L | 364 | 96 | 318 | 102 | 208 |
| FBXW2 | 242 | 67 | 256 | 51 | 180 |
| GPBP1 | 302 | 87 | 272 | 87 | 180 |
| LUC7L2 | 268 | 80 | 295 | 74 | 77 |
| OAZ1 | 284 | 93 | 194 | 82 | 199 |
| PAPOLA | 427 | 125 | 392 | 135 | 315 |
| SPG21 | 243 | 72 | 226 | 67 | 101 |
| TRIM27 | 383 | 107 | 329 | 105 | 129 |
| UBQLN1 | 296 | 85 | 217 | 74 | 160 |
| ZNF207 | 420 | 124 | 365 | 136 | 310 |
| HPRT1 | 334 | 94 | 264 | 83 | 115 |
| PSMB6 | 267 | 68 | 222 | 63 | 155 |
| B2M | 319 | 102 | 285 | 94 | 175 |
| GAPDH | 334 | 108 | 227 | 102 | 93 |
Expression of RLIP76, but not of MRP1, is modulated by TNF-α
RLIP76 is a tightly regulated gene expressed in all human tissues and cell lines so far examined. The RLIP76 gene encodes a GTPase-activating protein and a downstream effector of the RALA and RALB ras-like GTP-binding proteins, thus modulating the mitotic spindle, clathrin-dependent endocytosis, tight junctions, cell-to-cell adhesion, targeting of proteins to basolateral plasma membranes, and the assembly of exocyst complexes and secretagogue-dependent exocytosis [80–82]. RLIP76 was also recently shown to interact with ARNO (a.k.a. Cytohesin 2), a guanine-nucleotide exchange Sec7 domain-containing protein that works in concert with ARF6 (ADP-ribosylation factor 6), a small guanine nucleotide-binding protein of the RAS superfamily [83]. RLIP76 activates RAC1 in an ARF6- and PI3K-dependent manner, and regulates also cell migration and spreading through the activities of ARNO and ARF6 [83]. In addition, RLIP76 regulates HIF1α (Hypoxia Inducible Factor 1, alpha subunit) and VEGF (Vascular Endothelial Growth Factor), and is essential to proper endothelial cell function, normal angiogenesis, and to tumor-associated neoangiogenesis [84, 85]. Finally, RLIP76 is also a non-ATP binding cassette drug transporter that effects the efflux of glutathione conjugates of electrophiles including those of xenobiotics, thus mediating drug resistance (e.g. of doxorubicin, vinorelbine, unitinib, sorafenib) in several mouse models bearing human cancer xenografts of the lungs, kidney, skin, colon, and prostate [86–93]. Owing to its expression at the BBB, RLIP76 can also lead to drug resistance of brain tumors, as well as in neurological disorders such as epilepsy (e.g. to phenytoin and carbamazepine) [86, 94].
Since the BBB is often subjected to inflammation in brain disorders [32], and TNF-α is a known modulator of several RLIP76 downstream targets through the NF-κB transcription factor [81, 95–97], we asked whether TNF-α might also regulate RLIP76. A comparative analysis of human, murine and rat RLIP76 5' flanking sequences uncovered several evolutionarily conserved and non-conserved putative NF-κB transcription factor binding sites (Fig 5). Non-conserved binding sites mapped mostly to transcription start site (TSS)-distant upstream sequences and TSS-downstream sequences, whereas evolutionarily conserved NF-κB binding clustered within less than a kilobase upstream of the TSS. Since conservation of transcription factor binding sites across divergent species is an indication of regulatory element functionality [98, 99], the data suggest that RLIP76 can be a target of pro-inflammatory molecules via a cluster of evolutionarily conserved NF-κB binding sites upstream of the TSS (Fig 5).
Fig 5. Comparative analysis of human, murine and rat RLIP76 promoter sequences unveils conserved NF-κB binding sites.
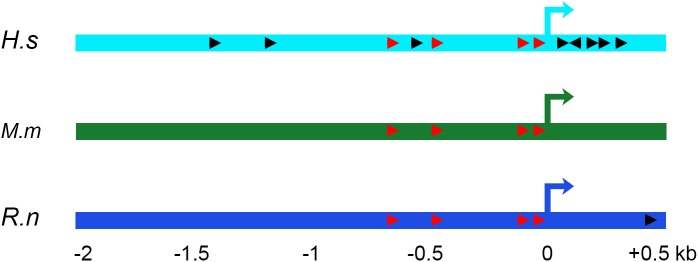
Human (Homo sapiens: H.s), murine (Mus musculus: M.m) and rat (Rattus norvegicus: R.n) DNA sequences spanning -2 to +0.5 kb relative to the transcription start site (+0 kb) were retrieved using the Cold Spring Harbor Laboratory promoter database (http://rulai.cshl.edu/cgi-bin/CSHLmpd2/promExtract.pl?). Sequence alignments were performed using NCBI Blast platform (http://blast.ncbi.nlm.nih.gov/Blast.cgi), and GGGRNNYYCC (whereby R is a purine and Y is a pyrimidine) was used as the consensus NF-κB binding site. Orientation of the binding sites is shown with arrow heads, with read arrowheads corresponding to evolutionarily conserved and black arrowheads non-conserved binding sites.
We therefore sought to explore the effects of TNF-α on RLIP76 gene expression in BBB endothelial cells. As shown in Fig 6A, TNF-α led to a gradual increase in RLIP76 expression, reaching a maximum at 24 hours. To assess whether TNF-α modulation of RLIP76 extends to other glutathione transporter-encoding genes, we also analyzed expression of the MRP1 gene. As shown in Fig 6B, TNF-α had no measurable effect on MRP1, supporting the specificity of RLIP76 modulation by TNF-α.
Fig 6. TNF-α induces RLIP76 but not MRP1 gene expression.
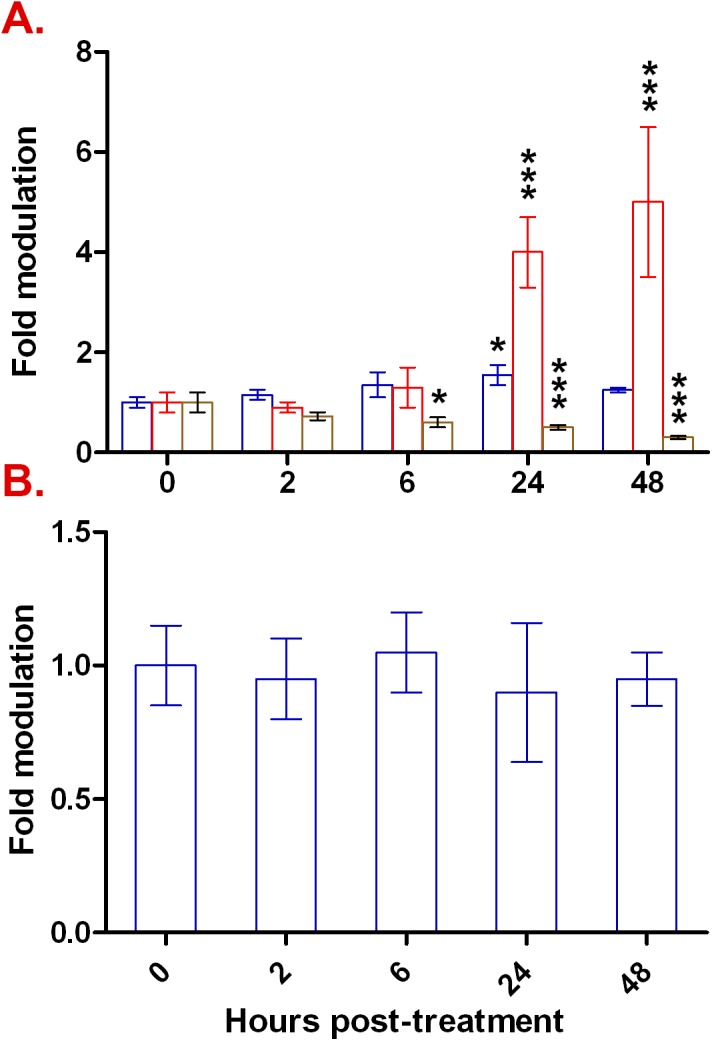
Expression levels of RLIP76 (Panel A) and MRP1 (Panel B) were monitored by qRT-PCR prior and after TNF-α treatment for 2–48 hours relative to the PSMB6/HPRT1 reference geneset (blue bars), GAPDH (red bars), or to B2M (brown bars). Results are presented as arithmetic means ± standard deviations. Statistical significance: * signifies p < 0.05, *** signifies p < 0.001; lack of asterisks denotes lack of a statistically significant difference.
Normalization of the RLIP76 expression data with single housekeeping genes, instead of the reference geneset, yielded very different and contrasting pictures, with either a maximal 500% induction of RLIP76 (GAPDH as reference gene; Fig 6A), or 70% suppression of its expression (B2M as a reference gene; Fig 6A). Although not as dramatic, one may also see the effect of using inappropriate single housekeeping genes in the normalization of highly inducible genes. For instance, normalization of ICAM1 using GAPDH would make its expression appear to gradually increase over the duration of the experiment by an additional 4–5 fold after the initial induction at 2h, thus giving the impression that ICAM1 maximal induction is reached only 48h post-treatment (Fig 1A). On the opposite end of the spectrum, normalization against B2M expression would intimate the notion of an escape mechanism from TNF-α induction taking place sometime between 2 and 6 hours post-induction (Fig 1A). Using the Normalization Factor, however, we show ICAM1 expression to be maximally induced 2h post TNF-α-treatment, and that the TNF-α-mediated induction is maintained at this level for the duration of the experiment (Fig 1A), consistent with previously reported ICAM1 induction kinetics in endothelial cells subjected to pro-inflammatory stimuli [100, 101]. Together, and as may be surmised from a comparison of data in Figs 1 and 6 in which TNF-α either appeared to cause profound effects (Fig 1A–1C) or exerted modest gene modulation (Fig 6A), data normalization skewing was more pronounced in datasets in which gene expression modulation was moderate (for example here RLIP76). We submit, therefore, that it is particularly important to use appropriate reference genesets instead of single untested housekeeping genes when quantifying moderate gene expression variations.
The fact that RLIP76 is induced by inflammation in BBB endothelial cells can be relevant to central nervous system diseases with an inflammatory component. This is for instance the case of epilepsy [102–105], and drugs used to treat epilepsy such as carbamazepine and phenytoin [106] are RLIP76 substrates [86]. A corollary to these notions is whether inflammation can modulate anti-epileptic drug efficacy, and importantly, whether non-steroidal anti-inflammatory drugs (NSAIDs) can improve epilepsy treatment. While data in patients are still lacking, several FDA-approved NSAIDs including acetylsalicylic acid, ibuprofen, indomethacin, metamizole, paracetamol, and piroxicam were reported to augment the antiepileptic effectiveness of phenytoin in a mouse seizure model [107]. Assuming this model can be translated in a clinical setting in patients, it suggests that NSAIDs may be useful not only in the treatment of epilepsy-associated inflammation, but may in addition potentiate the efficacy of anti-epileptic drugs. Since RLIP76 expression is increased in the brain of carbamazepine- and phenytoin-refractory epileptic patients wherein it extrudes these drugs [86], and since inflammation often associates with epilepsy [102–105] and may increase RLIP76 expression (this report), it may be pertinent to explore whether NSAIDs ought to systematically be co-administered with anti-epileptic drugs in cases wherein drug resistance is documented. Finally, from a mechanistic perspective, it would be of interest to explore whether the potentiating effect of NSAIDs on anti-epileptic drugs is due to the down-modulation of RLIP76 gene expression.
Conclusions
Treatment of human endothelial cells in a BBB model with TNF-α led to the induction of several inflammation markers indicating an inflammatory response. Under these conditions, the expression of housekeeping genes such as HPRT1 and PSMB6 remained relatively unaffected, whereas others showed time-dependent induction or suppression. Using the appropriate reference geneset, we show the RLIP76 drug transporter to be induced by TNF-α. This induction is not a general property of all glutathione transporter-encoding genes as expression of MRP1 was unchanged in the same conditions. Modulation of RLIP76 by TNF-α might be relevant to the treatment outcome of brain disorders and tumors wherein the blood-brain barrier is inflamed, as it may alter drug transport across the barrier. Finally, we show that no gene or geneset may act as a universal reference gene, which therefore ought to be established for each cell type and associated pathophysiological condition.
Supporting Information
Listed are the in silico and experimental variables pertaining to the design and conducting of qRT-PCR procedures on all genes tested.
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
Each metagroup contained at least 10 biological replicates. UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│ > 2 was found to be significant (p<0.05).
(PDF)
Each metagroup contained at least 10 biological replicates. UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│ > 2 was found to be significant (p<0.05).
(PDF)
Acknowledgments
The authors wish to thank Dr. Sanjay Awasthi (City of Hope, Comprehensive Cancer Center) and Dr. Lawrence E. Goldfinger (Temple University School of Medicine, and Cancer Biology Program at Fox Chase Cancer Center) for their valuable time and comments.
Data Availability
All relevant data are within the paper and its Supporting Information files.
Funding Statement
The authors have no support or funding to report.
References
- 1. Golias C, Tsoutsi E, Matziridis A, Makridis P, Batistatou A, Charalabopoulos K. Review. Leukocyte and endothelial cell adhesion molecules in inflammation focusing on inflammatory heart disease. In Vivo. 2007;21(5):757–69. . [PubMed] [Google Scholar]
- 2. Schmid MC, Schulein R, Dehio M, Denecker G, Carena I, Dehio C. The VirB type IV secretion system of Bartonella henselae mediates invasion, proinflammatory activation and antiapoptotic protection of endothelial cells. Mol Microbiol. 2004;52(1):81–92. . [DOI] [PubMed] [Google Scholar]
- 3. Vallance P, Collier J, Bhagat K. Infection, inflammation, and infarction: does acute endothelial dysfunction provide a link? Lancet. 1997;349(9062):1391–2. . [DOI] [PubMed] [Google Scholar]
- 4. Silva Y, Leira R, Tejada J, Lainez JM, Castillo J, Davalos A. Molecular signatures of vascular injury are associated with early growth of intracerebral hemorrhage. Stroke. 2005;36(1):86–91. . [DOI] [PubMed] [Google Scholar]
- 5. Marboeuf P, Corseaux D, Mouquet F, Van Belle E, Jude B, Susen S. Inflammation triggers colony forming endothelial cell mobilization after angioplasty in chronic lower limb ischemia. J Thromb Haemost. 2008;6(1):195–7. . [DOI] [PubMed] [Google Scholar]
- 6. Beekhuizen H, van de Gevel JS. Endothelial cell adhesion molecules in inflammation and postischemic reperfusion injury. Transplant Proc. 1998;30(8):4251–6. . [DOI] [PubMed] [Google Scholar]
- 7. Kupatt C, Habazettl H, Becker BF, Boekstegers P. Endothelial activation—a strategic event during postischemic myocardial inflammation. Z Kardiol. 2000;89 Suppl 9:IX/96–100. . [DOI] [PubMed] [Google Scholar]
- 8. Arnaud C, Mach F. Pleiotropic effects of statins in atherosclerosis: role on endothelial function, inflammation and immunomodulation. Arch Mal Coeur Vaiss. 2005;98(6):661–6. . [PubMed] [Google Scholar]
- 9. Gimbrone MA Jr., Bevilacqua MP, Cybulsky MI. Endothelial-dependent mechanisms of leukocyte adhesion in inflammation and atherosclerosis. Ann N Y Acad Sci. 1990;598:77–85. . [DOI] [PubMed] [Google Scholar]
- 10. Lahera V, Goicoechea M, de Vinuesa SG, Miana M, de las Heras N, Cachofeiro V, et al. Endothelial dysfunction, oxidative stress and inflammation in atherosclerosis: beneficial effects of statins. Curr Med Chem. 2007;14(2):243–8. . [DOI] [PubMed] [Google Scholar]
- 11. Deban L, Correale C, Vetrano S, Malesci A, Danese S. Multiple pathogenic roles of microvasculature in inflammatory bowel disease: a Jack of all trades. Am J Pathol. 2008;172(6):1457–66. 10.2353/ajpath.2008.070593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Horowitz S, Binion DG, Nelson VM, Kanaa Y, Javadi P, Lazarova Z, et al. Increased arginase activity and endothelial dysfunction in human inflammatory bowel disease. Am J Physiol Gastrointest Liver Physiol. 2007;292(5):G1323–36. . [DOI] [PubMed] [Google Scholar]
- 13. Schinzari F, Armuzzi A, De Pascalis B, Mores N, Tesauro M, Melina D, et al. Tumor necrosis factor-alpha antagonism improves endothelial dysfunction in patients with Crohn's disease. Clin Pharmacol Ther. 2008;83(1):70–6. . [DOI] [PubMed] [Google Scholar]
- 14. Minagar A, Jy W, Jimenez JJ, Alexander JS. Multiple sclerosis as a vascular disease. Neurol Res. 2006;28(3):230–5. . [DOI] [PubMed] [Google Scholar]
- 15. Higashi Y, Goto C, Jitsuiki D, Umemura T, Nishioka K, Hidaka T, et al. Periodontal infection is associated with endothelial dysfunction in healthy subjects and hypertensive patients. Hypertension. 2008;51(2):446–53. . [DOI] [PubMed] [Google Scholar]
- 16. Moutsopoulos NM, Madianos PN. Low-grade inflammation in chronic infectious diseases: paradigm of periodontal infections. Ann N Y Acad Sci. 2006;1088:251–64. . [DOI] [PubMed] [Google Scholar]
- 17. Seinost G, Wimmer G, Skerget M, Thaller E, Brodmann M, Gasser R, et al. Periodontal treatment improves endothelial dysfunction in patients with severe periodontitis. Am Heart J. 2005;149(6):1050–4. . [DOI] [PubMed] [Google Scholar]
- 18. Tonetti MS, D'Aiuto F, Nibali L, Donald A, Storry C, Parkar M, et al. Treatment of periodontitis and endothelial function. N Engl J Med. 2007;356(9):911–20. . [DOI] [PubMed] [Google Scholar]
- 19. Gonzalez-Gay MA, Gonzalez-Juanatey C, Martin J. Inflammation and endothelial dysfunction in rheumatoid arthritis. Clin Exp Rheumatol. 2006;24(2):115–7. . [PubMed] [Google Scholar]
- 20. Kerekes G, Szekanecz Z, Der H, Sandor Z, Lakos G, Muszbek L, et al. Endothelial dysfunction and atherosclerosis in rheumatoid arthritis: a multiparametric analysis using imaging techniques and laboratory markers of inflammation and autoimmunity. J Rheumatol. 2008;35(3):398–406. . [PubMed] [Google Scholar]
- 21. Antoniades C, Tousoulis D, Marinou K, Papageorgiou N, Bosinakou E, Tsioufis C, et al. Effects of insulin dependence on inflammatory process, thrombotic mechanisms and endothelial function, in patients with type 2 diabetes mellitus and coronary atherosclerosis. Clin Cardiol. 2007;30(6):295–300. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Leinonen ES, Hiukka A, Hurt-Camejo E, Wiklund O, Sarna SS, Mattson Hulten L, et al. Low-grade inflammation, endothelial activation and carotid intima-media thickness in type 2 diabetes. J Intern Med. 2004;256(2):119–27. . [DOI] [PubMed] [Google Scholar]
- 23. Mantovani A, Allavena P, Sica A, Balkwill F. Cancer-related inflammation. Nature. 2008;454(7203):436–44. Epub 2008/07/25. nature07205 [pii] 10.1038/nature07205 . [DOI] [PubMed] [Google Scholar]
- 24. Pitroda SP, Zhou T, Sweis RF, Filippo M, Labay E, Beckett MA, et al. Tumor endothelial inflammation predicts clinical outcome in diverse human cancers. PLoS One. 2012;7(10):e46104 Epub 2012/10/12. doi: 10.1371/journal.pone.0046104 PONE-D-12-07623 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Kalinowska A, Losy J. PECAM-1, a key player in neuroinflammation. Eur J Neurol. 2006;13(12):1284–90. . [DOI] [PubMed] [Google Scholar]
- 26. Sharief MK, Ciardi M, Thompson EJ. Blood-brain barrier damage in patients with bacterial meningitis: association with tumor necrosis factor-alpha but not interleukin-1 beta. J Infect Dis. 1992;166(2):350–8. . [DOI] [PubMed] [Google Scholar]
- 27. Beck KD, Nguyen HX, Galvan MD, Salazar DL, Woodruff TM, Anderson AJ. Quantitative analysis of cellular inflammation after traumatic spinal cord injury: evidence for a multiphasic inflammatory response in the acute to chronic environment. Brain. 2010;133(Pt 2):433–47. Epub 2010/01/21. doi: 10.1093/brain/awp322 awp322 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Chapouly C, Tadesse Argaw A, Horng S, Castro K, Zhang J, Asp L, et al. Astrocytic TYMP and VEGFA drive blood-brain barrier opening in inflammatory central nervous system lesions. Brain. 2015;138(Pt 6):1548–67. Epub 2015/03/26. doi: 10.1093/brain/awv077 awv077 [pii]. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Minten C, Alt C, Gentner M, Frei E, Deutsch U, Lyck R, et al. DARC shuttles inflammatory chemokines across the blood-brain barrier during autoimmune central nervous system inflammation. Brain. 2014;137(Pt 5):1454–69. Epub 2014/03/15. doi: 10.1093/brain/awu045 awu045 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Stamatovic SM, Dimitrijevic OB, Keep RF, Andjelkovic AV. Inflammation and brain edema: new insights into the role of chemokines and their receptors. Acta Neurochir Suppl. 2006;96:444–50. . [DOI] [PubMed] [Google Scholar]
- 31. Liu JY, Thom M, Catarino CB, Martinian L, Figarella-Branger D, Bartolomei F, et al. Neuropathology of the blood-brain barrier and pharmaco-resistance in human epilepsy. Brain. 2012;135(Pt 10):3115–33. Epub 2012/07/04. doi: 10.1093/brain/aws147 aws147 [pii]. . [DOI] [PubMed] [Google Scholar]
- 32. Vezzani A, Granata T. Brain inflammation in epilepsy: experimental and clinical evidence. Epilepsia. 2005;46(11):1724–43. . [DOI] [PubMed] [Google Scholar]
- 33. Carvey PM, Zhao CH, Hendey B, Lum H, Trachtenberg J, Desai BS, et al. 6-Hydroxydopamine-induced alterations in blood-brain barrier permeability. Eur J Neurosci. 2005;22(5):1158–68. . [DOI] [PubMed] [Google Scholar]
- 34. Wilms H, Zecca L, Rosenstiel P, Sievers J, Deuschl G, Lucius R. Inflammation in Parkinson's diseases and other neurodegenerative diseases: cause and therapeutic implications. Curr Pharm Des. 2007;13(18):1925–8. . [DOI] [PubMed] [Google Scholar]
- 35. Franscini N, Bachli EB, Blau N, Leikauf MS, Schaffner A, Schoedon G. Gene expression profiling of inflamed human endothelial cells and influence of activated protein C. Circulation. 2004;110(18):2903–9. . [DOI] [PubMed] [Google Scholar]
- 36. Gu J, Chen Y, Li S, Li Y. Identification of responsive gene modules by network-based gene clustering and extending: application to inflammation and angiogenesis. BMC Syst Biol. 2010;4:47 Epub 2010/04/22. doi: 10.1186/1752-0509-4-47 1752-0509-4-47 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Lok CA, Boing AN, Reitsma PH, van der Post JA, van Bavel E, Boer K, et al. Expression of inflammation-related genes in endothelial cells is not directly affected by microparticles from preeclamptic patients. J Lab Clin Med. 2006;147(6):310–20. . [DOI] [PubMed] [Google Scholar]
- 38. Zhou J, Jin Y, Gao Y, Wang H, Hu G, Huang Y, et al. Genomic-scale analysis of gene expression profiles in TNF-alpha treated human umbilical vein endothelial cells. Inflamm Res. 2002;51(7):332–41. . [DOI] [PubMed] [Google Scholar]
- 39. De Martin R, Hoeth M, Hofer-Warbinek R, Schmid JA. The transcription factor NF-kappa B and the regulation of vascular cell function. Arterioscler Thromb Vasc Biol. 2000;20(11):E83–8. . [DOI] [PubMed] [Google Scholar]
- 40. Kracht M, Saklatvala J. Transcriptional and post-transcriptional control of gene expression in inflammation. Cytokine. 2002;20(3):91–106. . [DOI] [PubMed] [Google Scholar]
- 41. Lee YW, Eum SY, Chen KC, Hennig B, Toborek M. Gene expression profile in interleukin-4-stimulated human vascular endothelial cells. Mol Med. 2004;10(1–6):19–27. . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Rahman I. Oxidative stress, chromatin remodeling and gene transcription in inflammation and chronic lung diseases. J Biochem Mol Biol. 2003;36(1):95–109. . [DOI] [PubMed] [Google Scholar]
- 43. Ruminy P, Gangneux C, Claeyssens S, Scotte M, Daveau M, Salier JP. Gene transcription in hepatocytes during the acute phase of a systemic inflammation: from transcription factors to target genes. Inflamm Res. 2001;50(8):383–90. . [DOI] [PubMed] [Google Scholar]
- 44. Mori R, Wang Q, Danenberg KD, Pinski JK, Danenberg PV. Both beta-actin and GAPDH are useful reference genes for normalization of quantitative RT-PCR in human FFPE tissue samples of prostate cancer. Prostate. 2008;68(14):1555–60. 10.1002/pros.20815 [DOI] [PubMed] [Google Scholar]
- 45. Murthi P, Fitzpatrick E, Borg AJ, Donath S, Brennecke SP, Kalionis B. GAPDH, 18S rRNA and YWHAZ are suitable endogenous reference genes for relative gene expression studies in placental tissues from human idiopathic fetal growth restriction. Placenta. 2008;29(9):798–801. 10.1016/j.placenta.2008.06.007 [DOI] [PubMed] [Google Scholar]
- 46. Cummings M, Sarveswaran J, Homer-Vanniasinkam S, Burke D, Orsi NM. Glyceraldehyde-3-phosphate Dehydrogenase is an Inappropriate Housekeeping Gene for Normalising Gene Expression in Sepsis. Inflammation. 2014;37(5):1889–94. Epub 2014/05/27. 10.1007/s10753-014-9920-3 . [DOI] [PubMed] [Google Scholar]
- 47. Thorrez L, Van Deun K, Tranchevent LC, Van Lommel L, Engelen K, Marchal K, et al. Using ribosomal protein genes as reference: a tale of caution. PLoS ONE. 2008;3(3):e1854 10.1371/journal.pone.0001854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Bamias G, Goukos D, Laoudi E, Balla IG, Siakavellas SI, Daikos GL, et al. Comparative study of candidate housekeeping genes for quantification of target gene messenger RNA expression by real-time PCR in patients with inflammatory bowel disease. Inflamm Bowel Dis. 2013;19(13):2840–7. Epub 2013/10/22. 10.1097/01.MIB.0000435440.22484.e8 . [DOI] [PubMed] [Google Scholar]
- 49. Montero-Melendez T, Perretti M. Gapdh gene expression is modulated by inflammatory arthritis and is not suitable for qPCR normalization. Inflammation. 2014;37(4):1059–69. Epub 2014/02/05. 10.1007/s10753-014-9829-x . [DOI] [PubMed] [Google Scholar]
- 50. Barar J, Gumbleton M, Asadi M, Omidi Y. Barrier functionality and transport machineries of human ECV304 cells. Med Sci Monit. 2010;16(1):BR52–60. Epub 2009/12/29. 878321 [pii]. . [PubMed] [Google Scholar]
- 51. Gonzalez-Burgos E, Carretero ME, Gomez-Serranillos MP. In vitro permeability study of CNS-active diterpenes from Sideritis spp. using cellular models of blood-brain barrier. Planta Med. 2013;79(16):1545–51. Epub 2013/09/14. 10.1055/s-0033-1350797 . [DOI] [PubMed] [Google Scholar]
- 52. Liverani E, Paul C. Glucocorticoids alter adrenomedullin receptor expression and secretion in endothelial-like cells and astrocytes. Int J Biochem Cell Biol. 2013;45(12):2715–23. Epub 2013/10/08. doi: 10.1016/j.biocel.2013.09.009 S1357-2725(13)00298-7 [pii]. . [DOI] [PubMed] [Google Scholar]
- 53. Neuhaus W, Freidl M, Szkokan P, Berger M, Wirth M, Winkler J, et al. Effects of NMDA receptor modulators on a blood-brain barrier in vitro model. Brain Res. 2011;1394:49–61. Epub 2011/05/10. doi: 10.1016/j.brainres.2011.04.003 S0006-8993(11)00670-6 [pii]. . [DOI] [PubMed] [Google Scholar]
- 54. Neuhaus W, Mandikova J, Pawlowitsch R, Linz B, Bennani-Baiti B, Lauer R, et al. Blood-brain barrier in vitro models as tools in drug discovery: assessment of the transport ranking of antihistaminic drugs. Pharmazie. 2012;67(5):432–9. Epub 2012/07/07. . [PubMed] [Google Scholar]
- 55. Novakova I, Subileau EA, Toegel S, Gruber D, Lachmann B, Urban E, et al. Transport rankings of non-steroidal antiinflammatory drugs across blood-brain barrier in vitro models. PLoS One. 2014;9(1):e86806 Epub 2014/01/28. doi: 10.1371/journal.pone.0086806 PONE-D-13-36709 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Poetsch V, Bennani-Baiti B, Neuhaus W, Muchitsch EM, Noe CR. Serum-derived immunoglobulins alter amyloid beta transport across a blood-brain barrier in vitro model. Pharmazie. 2010;65(4):267–73. Epub 2010/05/04. . [PubMed] [Google Scholar]
- 57. Rodriguez-Gaztelumendi A, Alvehus M, Andersson T, Jacobsson SO. Comparison of the effects of nicotine upon the transcellular electrical resistance and sucrose permeability of human ECV304/rat C6 co-cultures and human CaCo(2) cells. Toxicol Lett. 2011;207(1):1–6. Epub 2011/09/06. doi: 10.1016/j.toxlet.2011.08.014 S0378-4274(11)01500-1 [pii]. . [DOI] [PubMed] [Google Scholar]
- 58. Untucht C, Rasch J, Fuchs E, Rohde M, Bergmann S, Steinert M. An optimized in vitro blood-brain barrier model reveals bidirectional transmigration of African trypanosome strains. Microbiology. 2011;157(Pt 10):2933–41. Epub 2011/07/09. doi: 10.1099/mic.0.049106–0 mic.0.049106–0 [pii]. . [DOI] [PubMed] [Google Scholar]
- 59. Wang GY, Wang N, Liao HN. Effects of Muscone on the Expression of P-gp, MMP-9 on Blood-Brain Barrier Model In Vitro. Cell Mol Neurobiol. in press. Epub 2015/05/16. 10.1007/s10571-015-0204-8 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Wang LF, Li X, Gao YB, Wang SM, Zhao L, Dong J, et al. Activation of VEGF/Flk-1-ERK Pathway Induced Blood-Brain Barrier Injury After Microwave Exposure. Mol Neurobiol. 2014. Epub 2014/09/10. 10.1007/s12035-014-8848-9 . [DOI] [PubMed] [Google Scholar]
- 61. Wang Q, Luo W, Zhang W, Liu M, Song H, Chen J. Involvement of DMT1 +IRE in the transport of lead in an in vitro BBB model. Toxicol In Vitro. 2011;25(4):991–8. Epub 2009/11/17. doi: 10.1016/j.tiv.2009.11.006 S0887-2333(09)00335-X [pii]. . [DOI] [PubMed] [Google Scholar]
- 62. Plattner VE, Germann B, Neuhaus W, Noe CR, Gabor F, Wirth M. Characterization of two blood-brain barrier mimicking cell lines: distribution of lectin-binding sites and perspectives for drug delivery. Int J Pharm. 2010;387(1–2):34–41. Epub 2009/12/08. doi: 10.1016/j.ijpharm.2009.11.030 S0378-5173(09)00851-5 [pii]. . [DOI] [PubMed] [Google Scholar]
- 63. Bennani-Baiti B, Bennani-Baiti IM. Gene symbol precision. Gene. 2012;491(2):103–9. Epub 2011/10/25. S0378-1119(11)00562-2 [pii] 10.1016/j.gene.2011.09.035 . [DOI] [PubMed] [Google Scholar]
- 64. Bustin SA, Benes V, Garson J, Hellemans J, Huggett J, Kubista M, et al. The need for transparency and good practices in the qPCR literature. Nat Methods. 2013;10(11):1063–7. Epub 2013/11/01. doi: 10.1038/nmeth.2697 nmeth.2697 [pii]. . [DOI] [PubMed] [Google Scholar]
- 65. Bustin SA, Beaulieu JF, Huggett J, Jaggi R, Kibenge FS, Olsvik PA, et al. MIQE precis: Practical implementation of minimum standard guidelines for fluorescence-based quantitative real-time PCR experiments. BMC Mol Biol. 2010;11:74 Epub 2010/09/23. doi: 10.1186/1471-2199-11-74 1471-2199-11-74 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3(7):RESEARCH0034. Epub 2002/08/20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Andersen CL, Jensen JL, Orntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004;64(15):5245–50. . [DOI] [PubMed] [Google Scholar]
- 68. Pfaffl MW, Tichopad A, Prgomet C, Neuvians TP. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations. Biotechnol Lett. 2004;26(6):509–15. . [DOI] [PubMed] [Google Scholar]
- 69. Bennani-Baiti IM, Cooper A, Lawlor ER, Kauer M, Ban J, Aryee DN, et al. Intercohort gene expression co-analysis reveals chemokine receptors as prognostic indicators in Ewing's sarcoma. Clin Cancer Res. 2010;16(14):3769–78. Epub 2010/06/08. doi: 10.1158/1078-0432.CCR-10-0558 1078-0432.CCR-10-0558 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Bennani-Baiti IM, Aryee DN, Ban J, Machado I, Kauer M, Muhlbacher K, et al. Notch signalling is off and is uncoupled from HES1 expression in Ewing's sarcoma. J Pathol. 2011;225(3):353–63. Epub 2011/10/11. 10.1002/path.2966 . [DOI] [PubMed] [Google Scholar]
- 71. Lukk M, Kapushesky M, Nikkila J, Parkinson H, Goncalves A, Huber W, et al. A global map of human gene expression. Nat Biotechnol. 2010;28(4):322–4. Epub 2010/04/10. doi: 10.1038/nbt0410-322 nbt0410-322 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Chen G, Zhao L, Feng J, You G, Sun Q, Li P, et al. Validation of reliable reference genes for real-time PCR in human umbilical vein endothelial cells on substrates with different stiffness. PLoS One. 2013;8(6):e67360 Epub 2013/07/11. doi: 10.1371/journal.pone.0067360 PONE-D-13-02338 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Wei R, Stewart EA, Amoaku WM. Suitability of endogenous reference genes for gene expression studies with human intraocular endothelial cells. BMC Res Notes. 2013;6:46 Epub 2013/02/06. doi: 10.1186/1756-0500-6-46 1756-0500-6-46 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Zyzynska-Granica B, Koziak K. Identification of suitable reference genes for real-time PCR analysis of statin-treated human umbilical vein endothelial cells. PLoS One. 2012;7(12):e51547 Epub 2012/12/20. doi: 10.1371/journal.pone.0051547 PONE-D-12-17286 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Garcia-Vallejo JJ, Van Het Hof B, Robben J, Van Wijk JA, Van Die I, Joziasse DH, et al. Approach for defining endogenous reference genes in gene expression experiments. Anal Biochem. 2004;329(2):293–9. . [DOI] [PubMed] [Google Scholar]
- 76. Pinhu L, Park JE, Yao W, Griffiths MJ. Reference gene selection for real-time polymerase chain reaction in human lung cells subjected to cyclic mechanical strain. Respirology. 2008;13(7):990–9. 10.1111/j.1440-1843.2008.01396.x [DOI] [PubMed] [Google Scholar]
- 77. Liu PQ, Rebar EJ, Zhang L, Liu Q, Jamieson AC, Liang Y, et al. Regulation of an endogenous locus using a panel of designed zinc finger proteins targeted to accessible chromatin regions. Activation of vascular endothelial growth factor A. J Biol Chem. 2001;276(14):11323–34. Epub 2001/01/19. doi: 10.1074/jbc.M011172200 M011172200 [pii]. . [DOI] [PubMed] [Google Scholar]
- 78. Bennani-Baiti IM, Asa SL, Song D, Iratni R, Liebhaber SA, Cooke NE. DNase I-hypersensitive sites I and II of the human growth hormone locus control region are a major developmental activator of somatotrope gene expression. Proc Natl Acad Sci U S A. 1998;95(18):10655–60. Epub 1998/09/02. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Kwon MJ, Oh E, Lee S, Roh MR, Kim SE, Lee Y, et al. Identification of novel reference genes using multiplatform expression data and their validation for quantitative gene expression analysis. PLoS One. 2009;4(7):e6162 Epub 2009/07/09. 10.1371/journal.pone.0006162 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Moskalenko S, Henry DO, Rosse C, Mirey G, Camonis JH, White MA. The exocyst is a Ral effector complex. Nat Cell Biol. 2002;4(1):66–72. Epub 2001/12/12. doi: 10.1038/ncb728 ncb728 [pii]. . [DOI] [PubMed] [Google Scholar]
- 81. Sugihara K, Asano S, Tanaka K, Iwamatsu A, Okawa K, Ohta Y. The exocyst complex binds the small GTPase RalA to mediate filopodia formation. Nat Cell Biol. 2002;4(1):73–8. Epub 2001/12/18. doi: 10.1038/ncb720 ncb720 [pii]. . [DOI] [PubMed] [Google Scholar]
- 82. Vandenbroucke E, Mehta D, Minshall R, Malik AB. Regulation of endothelial junctional permeability. Ann N Y Acad Sci. 2008;1123:134–45. Epub 2008/04/01. 1123/1/134 [pii] 10.1196/annals.1420.016 . [DOI] [PubMed] [Google Scholar]
- 83. Lee S, Wurtzel JG, Goldfinger LE. The RLIP76 N-terminus binds ARNO to regulate PI 3-kinase, Arf6 and Rac signaling, cell spreading and migration. Biochem Biophys Res Commun. 2014;454(4):560–5. Epub 2014/12/03. S0006-291X(14)01934-2 [pii] 10.1016/j.bbrc.2014.10.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Lee S, Goldfinger LE. RLIP76 regulates HIF-1 activity, VEGF expression and secretion in tumor cells, and secretome transactivation of endothelial cells. FASEB J. 2014;28(9):4158–68. Epub 2014/06/15. doi: 10.1096/fj.14-255711 fj.14-255711 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Lee S, Wurtzel JG, Singhal SS, Awasthi S, Goldfinger LE. RALBP1/RLIP76 depletion in mice suppresses tumor growth by inhibiting tumor neovascularization. Cancer Res. 2012;72(20):5165–73. Epub 2012/08/21. doi: 10.1158/0008-5472.CAN-12-0468 0008-5472.CAN-12-0468 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Awasthi S, Hallene KL, Fazio V, Singhal SS, Cucullo L, Awasthi YC, et al. RLIP76, a non-ABC transporter, and drug resistance in epilepsy. BMC Neurosci. 2005;6:61 Epub 2005/09/29. 1471-2202-6-61 [pii] 10.1186/1471-2202-6-61 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Awasthi S, Singhal SS, Awasthi YC, Martin B, Woo JH, Cunningham CC, et al. RLIP76 and Cancer. Clin Cancer Res. 2008;14(14):4372–7. Epub 2008/07/17. 14/14/4372 [pii] 10.1158/1078-0432.CCR-08-0145 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Lim KH, Baines AT, Fiordalisi JJ, Shipitsin M, Feig LA, Cox AD, et al. Activation of RalA is critical for Ras-induced tumorigenesis of human cells. Cancer Cell. 2005;7(6):533–45. Epub 2005/06/14. S1535-6108(05)00157-1 [pii] 10.1016/j.ccr.2005.04.030 . [DOI] [PubMed] [Google Scholar]
- 89. Singhal SS, Sehrawat A, Sahu M, Singhal P, Vatsyayan R, Rao Lelsani PC, et al. Rlip76 transports sunitinib and sorafenib and mediates drug resistance in kidney cancer. Int J Cancer. 2010;126(6):1327–38. Epub 2009/07/25. 10.1002/ijc.24767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Singhal SS, Yadav S, Roth C, Singhal J. RLIP76: A novel glutathione-conjugate and multi-drug transporter. Biochem Pharmacol. 2009;77(5):761–9. Epub 2008/11/06. S0006-2952(08)00715-6 [pii] 10.1016/j.bcp.2008.10.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Singhal SS, Yadav S, Singhal J, Zajac E, Awasthi YC, Awasthi S. Depletion of RLIP76 sensitizes lung cancer cells to doxorubicin. Biochem Pharmacol. 2005;70(3):481–8. Epub 2005/06/14. S0006-2952(05)00312-6 [pii] 10.1016/j.bcp.2005.05.005 . [DOI] [PubMed] [Google Scholar]
- 92. Vatsyayan R, Chaudhary P, Lelsani PC, Singhal P, Awasthi YC, Awasthi S, et al. Role of RLIP76 in doxorubicin resistance in lung cancer. Int J Oncol. 2009;34(6):1505–11. Epub 2009/05/09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Vatsyayan R, Lelsani PC, Awasthi S, Singhal SS. RLIP76: a versatile transporter and an emerging target for cancer therapy. Biochem Pharmacol. 2010;79(12):1699–705. Epub 2010/01/26. doi: 10.1016/j.bcp.2010.01.016 S0006-2952(10)00027-4 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Bae YS, Chung W, Han K, Park KY, Kim H, Kim E, et al. Down-regulation of RalBP1 expression reduces seizure threshold and synaptic inhibition in mice. Biochem Biophys Res Commun. 2013;433(2):175–80. Epub 2013/03/15. doi: 10.1016/j.bbrc.2013.02.056 S0006-291X(13)00316-1 [pii]. . [DOI] [PubMed] [Google Scholar]
- 95. Birukov KG, Bochkov VN, Birukova AA, Kawkitinarong K, Rios A, Leitner A, et al. Epoxycyclopentenone-containing oxidized phospholipids restore endothelial barrier function via Cdc42 and Rac. Circ Res. 2004;95(9):892–901. Epub 2004/10/09. 01.RES.0000147310.18962.06 [pii] 10.1161/01.RES.0000147310.18962.06 . [DOI] [PubMed] [Google Scholar]
- 96. Zhao D, Kuhnt-Moore S, Zeng H, Pan A, Wu JS, Simeonidis S, et al. Substance P-stimulated interleukin-8 expression in human colonic epithelial cells involves Rho family small GTPases. Biochem J. 2002;368(Pt 2):665–72. Epub 2002/08/10. doi: 10.1042/BJ20020950 BJ20020950 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Zhao D, Kuhnt-Moore S, Zeng H, Wu JS, Moyer MP, Pothoulakis C. Neurotensin stimulates IL-8 expression in human colonic epithelial cells through Rho GTPase-mediated NF-kappa B pathways. Am J Physiol Cell Physiol. 2003;284(6):C1397–404. Epub 2003/02/14. doi: 10.1152/ajpcell.00328.2002 00328.2002 [pii]. . [DOI] [PubMed] [Google Scholar]
- 98. Hunt C, Morimoto RI. Conserved features of eukaryotic hsp70 genes revealed by comparison with the nucleotide sequence of human hsp70. Proc Natl Acad Sci U S A. 1985;82(19):6455–9. Epub 1985/10/01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Duret L, Bucher P. Searching for regulatory elements in human noncoding sequences. Curr Opin Struct Biol. 1997;7(3):399–406. Epub 1997/06/01. S0959-440X(97)80058-9 [pii]. . [DOI] [PubMed] [Google Scholar]
- 100. Tian H, Cao L, Tan Y, Williams S, Chen L, Matray T, et al. Multiplex mRNA assay using electrophoretic tags for high-throughput gene expression analysis. Nucleic Acids Res. 2004;32(16):e126 Epub 2004/09/10. doi: 10.1093/nar/gnh119 32/16/e126 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Ammar Z, Plazolles N, Baltz T, Coustou V. Identification of trans-sialidases as a common mediator of endothelial cell activation by African trypanosomes. PLoS Pathog. 2013;9(10):e1003710 Epub 2013/10/17. doi: 10.1371/journal.ppat.1003710 PPATHOGENS-D-13-01000 [pii]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Walker L, Sills GJ. Inflammation and epilepsy: the foundations for a new therapeutic approach in epilepsy? Epilepsy Curr. 2012;12(1):8–12. Epub 2012/03/01. 10.5698/1535-7511-12.1.8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Gorter JA, van Vliet EA, Aronica E. Status epilepticus, blood-brain barrier disruption, inflammation, and epileptogenesis. Epilepsy Behav. 2015. Epub 2015/05/11. S1525-5050(15)00216-4 [pii] 10.1016/j.yebeh.2015.04.047 . [DOI] [PubMed] [Google Scholar]
- 104. de Souza Bernardino TC, Teixeira AL, Miranda AS, Guidine PM, Rezende G, Doretto MC, et al. Wistar Audiogenic Rats (WAR) exhibit altered levels of cytokines and brain-derived neurotrophic factor following audiogenic seizures. Neurosci Lett. 2015;597:154–8. Epub 2015/05/06. doi: 10.1016/j.neulet.2015.04.046 S0304-3940(15)00345-6 [pii]. . [DOI] [PubMed] [Google Scholar]
- 105. Dupuis N, Auvin S. Inflammation and epilepsy in the developing brain: clinical and experimental evidence. CNS Neurosci Ther. 2015;21(2):141–51. Epub 2015/01/22. 10.1111/cns.12371 . [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Tudur Smith C, Marson AG, Clough HE, Williamson PR. Carbamazepine versus phenytoin monotherapy for epilepsy. Cochrane Database Syst Rev. 2002;(2):CD001911. Epub 2002/06/22. CD001911 [pii] 10.1002/14651858.CD001911 . [DOI] [PubMed] [Google Scholar]
- 107. Kaminski R, Kozicka M, Parada-turska J, Dziki M, Kleinrok Z, Turski WA, et al. Effect of non-steroidal anti-inflammatory drugs on the anticonvulsive activity of valproate and diphenylhydantoin against maximal electroshock-induced seizures in mice. Pharmacol Res. 1998;37(5):375–81. Epub 1998/06/27. S1043-6618(98)90309-7 [pii] 10.1006/phrs.1998.0309 . [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Listed are the in silico and experimental variables pertaining to the design and conducting of qRT-PCR procedures on all genes tested.
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│> 2 were found to be significant (p<0.05).
(PDF)
Each metagroup contained at least 10 biological replicates. UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│ > 2 was found to be significant (p<0.05).
(PDF)
Each metagroup contained at least 10 biological replicates. UP: up-regulated; DOWN: down-regulated; NONDE: non-differentially expressed. │t-statistic│ > 2 was found to be significant (p<0.05).
(PDF)
Data Availability Statement
All relevant data are within the paper and its Supporting Information files.