Coexpression of the receptor and ligand that function in crucifer self-incompatibility inhibits receptor signaling and abrogates the ability of stigma epidermal cells to arrest self pollen.
Abstract
The inhibition of self-pollination in self-incompatible Brassicaceae is based on allele-specific trans-activation of the highly polymorphic S-locus receptor kinase (SRK), which is displayed at the surface of stigma epidermal cells, by its even more polymorphic pollen coat-localized ligand, the S-locus cysteine-rich (SCR) protein. In an attempt to achieve constitutive activation of SRK and thus facilitate analysis of self-incompatibility (SI) signaling, we coexpressed an Arabidopsis lyrata SCR variant with its cognate SRK receptor in the stigma epidermal cells of Arabidopsis (Arabidopsis thaliana) plants belonging to the C24 accession, in which expression of SRK and SCR had been shown to exhibit a robust SI response. Contrary to expectation, however, coexpression of SRK and SCR was found to inhibit SRK-mediated signaling and to disrupt the SI response. This phenomenon, called cis-inhibition, is well documented in metazoans but has not as yet been reported for plant receptor kinases. We demonstrate that cis-inhibition of SRK, like its trans-activation, is based on allele-specific interaction between receptor and ligand. We also show that stigma-expressed SCR causes entrapment of its SRK receptor in the endoplasmic reticulum, thus disrupting the proper targeting of SRK to the plasma membrane, where the receptor would be available for productive interaction with its pollen coat-derived SCR ligand. Although based on an artificial cis-inhibition system, the results suggest novel strategies of pollination control for the generation of hybrid cultivars and large-scale seed production from hybrid plants in Brassicaceae seed crops and, more generally, for inhibiting cell surface receptor function and manipulating signaling pathways in plants.
Ligand receptor signaling plays important roles in cell-cell communication between neighboring cells in a variety of developmental and physiological processes. This communication typically relies on the interaction of transmembrane receptors displayed on the surface of signal-receiving cells with their cognate ligands derived from signal-sending neighboring cells, which, in turn, leads to the activation of receptor-mediated signaling cascades that modify intracellular activities of the signal-receiving cell. Such is the case with communication between pollen grains and stigma epidermal cells, a process that has an important role in directing reproductive success and determining pollination modes (i.e. selfing or outcrossing) in the Brassicaceae. In this family, outcrossing is enforced by self-incompatibility (SI), a mechanism controlled by haplotypes of the S locus, by which the stigma epidermal cells of a plant recognize and reject self pollen grains (i.e. those derived from the same flower, the same plant, or plants expressing the same S-locus haplotype), thus preventing self-pollination, while allowing the growth of tubes from nonself pollen grains (i.e. those derived from plants expressing a different S-locus haplotype; Nasrallah and Nasrallah, 2014a). Inhibition of self pollen in the SI response is initiated by allele-specific interaction between two highly polymorphic proteins encoded at the S locus: the S-locus receptor kinase (SRK), which is localized at the plasma membrane of stigma epidermal cells (Stein et al., 1991, 1996), and its ligand, the S-locus cysteine-rich protein (SCR), which accumulates in the pollen coat and diffuses onto the stigma surface upon pollen-stigma contact (Schopfer et al., 1999; Takayama et al., 2000; Shiba et al., 2001). The interaction of the SRK extracellular domain, or S domain, with its cognate SCR ligand is thought to activate downstream signaling cascades in stigma epidermal cells, which lead to inhibition of pollen germination on the stigma surface and/or pollen tube penetration through the stigma epidermal cell wall. The SRK and SCR genes are the primary determinants of the transition between the outcrossing and selfing modes of mating in the Brassicaceae, as demonstrated by the observation that transformation of SRK and SCR gene pairs derived from self-incompatible Arabidopsis lyrata or Capsella grandiflora restored SI in several accessions of the normally self-fertile Arabidopsis (Arabidopsis thaliana; Nasrallah et al., 2002, 2004; Boggs et al., 2009).
Tight regulation of the SI response is critical for ensuring reproductive success in self-incompatible plants. Activation of SRK signaling must be triggered only by pollen-derived cognate SCR ligand upon interaction of stigma epidermal cells with self pollen grains, because constitutive activation of SI signaling in stigma epidermal cells would result in inhibition of nonself as well as self pollen grains and would result in female sterility. This adverse outcome is averted by tight regulation of the SCR gene, which is expressed exclusively in the anthers of self-incompatible plants and whose protein products are localized exclusively in the pollen coat (Schopfer and Nasrallah, 2000; Shiba et al., 2001). For experimental studies of SI, however, constitutive activation of SRK-mediated signaling in stigma epidermal cells would be useful, as it might provide a convenient means of identifying components of the poorly understood SRK-mediated signaling pathway.
A reaction that resembles a constitutive SI response, in which stigma epidermal cells inhibit both self and nonself pollen grains, has been obtained by manual application of purified recombinant SCR proteins produced in bacteria (Kachroo et al., 2001; Chookajorn et al., 2004) or synthetic SCR (Takayama et al., 2001) to stigmas that express their cognate SRK receptors. Unlike the highly localized activation induced by pollen-derived SCR at the site of pollen-stigma contact, treatment of the stigma surface with SCR protein can clearly cause global activation of SRK in most, if not all, epidermal cells of a stigma. However, treating stigmas in the numbers required for analysis of SRK signaling is extremely laborious, can damage stigmas, and produces inconsistent results. Therefore, a method that circumvents these problems would be advantageous. In metazoans, constitutive activation of receptor kinases has been shown to result not only from receptor mutations that cause constitutive kinase activity (Webster and Donoghue, 1996; Hirota et al., 1998) and mutations in signaling components that cause ligand-independent activation of downstream cascades (Wang et al., 2012, 2014; Roberts et al., 2013; Han, 2014), but also from ectopic expression of ligands within the same cells as their receptors, as occurs in several pathological conditions (Sporn and Roberts, 1985; Castellano et al., 2006; Krasagakis et al., 2011).
Accordingly, an attempt was made to generate Arabidopsis plants having a stable constitutive stigma SI response by coexpressing an SRK variant and its cognate SCR in stigma epidermal cells, which should, in principle, constitutively activate the SI response in these cells. This report shows that, while pollen-derived SCR trans-activates the SRK-mediated SI response, stigma-expressed SCR inhibits the activity of its cognate SRK by causing entrapment of the receptor in the endoplasmic reticulum (ER). This phenomenon is similar in its outcome to the ligand-mediated cis-inhibition phenomenon that had previously been observed in metazoans for some signaling systems that use transmembrane proteins as ligands (Yaron and Sprinzak, 2012) but had not been described for plant receptor-like kinases. The results suggest novel strategies for control of receptor-like kinase activity and manipulation of signaling pathways in plants and for pollination control in hybrid breeding programs and seed production from hybrid plants in the Brassicaceae.
RESULTS
Stigma-Expressed SCR Ligand Inhibits the Activity of Its Cognate SRK Receptor
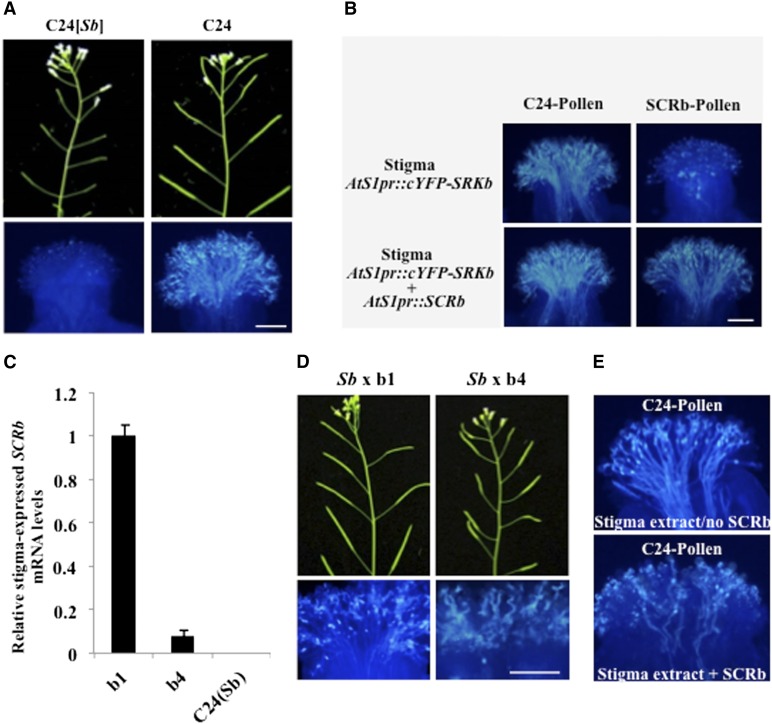
To coexpress an SRK variant with its cognate SCR in stigma epidermal cells, we used the SRKb and SCRb alleles, which are derived from the A. lyrata S-locus haplotype b (Sb) haplotype (Kusaba et al., 2001), and the C24 accession of Arabidopsis, which was previously shown to exhibit a robust and developmentally SI response upon transformation with the SRKb-SCRb gene pair (Nasrallah et al., 2004; Boggs et al., 2009; Nasrallah and Nasrallah, 2014b). As shown in Figure 1A, the self-incompatible phenotype of C24[SRKb-SCRb] plants is manifested by the production of unexpanded siliques lacking seed and severe inhibition of pollen grains at the surface of self-pollinated stigmas. These phenotypes, which faithfully recapitulate the phenotypes observed in naturally self-incompatible Brassicaceae, reflect strong trans-activation of SRKb by its SCRb ligand. By contrast, untransformed C24 plants are characterized by the production of abundant seed in fully expanded siliques and by prolific pollen tube growth in self-pollinated pistils (Fig. 1A).
Figure 1.
The SRK-mediated SI response in self-incompatible Arabidopsis transgenic plants and its inhibition by stigma-expressed SCRb. A, The SI response exhibited by C24[SRKb-SCRb] is manifested by severe inhibition of self pollen at the stigma surface, resulting in unexpanded siliques lacking seeds (left sections). By contrast, self-pollinated stigmas of a self-compatible untransformed C24 plant allow profuse pollen tube growth, resulting in the production of seed-filled siliques (right sections). B, Effect of the AtS1pr:SCRb transgene on the SI response in AtS1pr:cYFP-SRKb transgenic plants. The top sections show the stigmas of an AtS1pr:cYFP-SRKb plant exhibiting a compatible response toward pollen grains from an untransformed C24 plant (left) and an incompatible response toward SCRb-pollen grains (right). The bottom sections show the stigmas of an AtS1pr:cYFP-SRKb + AtS1pr:SCRb plant exhibiting compatible responses toward both untransformed C24 pollen grains and SCRb-pollen grains. C, Expression levels of stigma-expressed SCRb. The histogram shows the results of quantitative RT-PCR analysis of the relative levels of stigma-expressed SCRb transcripts in two independent AtS1pr:SCRb transgenic lines, b1 and b4, and untransformed C24(Sb) control. D, Concentration-dependent effects of stigma-expressed SCRb on the growth of SCRb-pollen tubes and on seed production in SRKb-SCRb transformants. Note that F1 plants produced by crossing SRKb-SCRb with AtS1pr:SCRb(b1; Sb × b1) exhibited complete breakdown of SI (seed-filled, fully expanded siliques and profuse SCRb-pollen tube growth at the stigma surface), while F1 plants produced by crossing SRKb-SCRb with AtS1pr:SCRb(b4; Sb × b4) exhibited partial breakdown of SI (partially elongated siliques and a relatively small number of SCRb-pollen tubes at the stigma surface). Whole-stigma images of the pollinated stigmas in this section are shown in Supplemental Figure S1B. E, Biological activity of stigma-expressed SCRb. A pollination assay of stigmas that express SRKb showing that application of a stigma extract from AtS1pr:SCRb plants induces the SI response, resulting in the inhibition of the majority of pollen grains from untransformed C24 plants, with only a few pollen tubes extending into the pistil (top). By contrast, a control treatment of SRKb-expressing stigmas with a stigma extract from untransformed C24 plants does not activate the SI response, as demonstrated by the numerous wild-type C24 pollen grains that germinate and invade the walls of stigma epidermal cells (bottom). Bars = 0.1 mm.
To express SCRb in stigma epidermal cells, the SCRb transcriptional unit was inserted downstream of the Arabidopsis S locus-related1 promoter, which is active specifically in stigma epidermal cells (Dwyer et al., 1994). Transformation of C24 plants with the AtS1pr:SCRb transgene produced 16 independent transformants. From these, three independent AtS1pr:SCRb homozygous lines, each of which contained transfer DNA fragments that had integrated at a single locus and exhibited appreciable SCRb expression (as determined by reverse transcription [RT]-PCR; Supplemental Fig. S1A), were selected for further analysis. Each of the three lines, designated AtS1pr:SCRb(b1), AtS1pr:SCRb(b2), and AtS1pr:SCRb(b3), was crossed with the previously described C24[AtS1pr:cYFP-SRKb] line (Kitashiba et al., 2011; Rea and Nasrallah, 2015), which is homozygous for a single integration of a transgene designed to express a tagged version of SRKb carrying the Citrine variant of yellow fluorescent protein (cYFP) at its N terminus. As shown previously (Kitashiba et al., 2011; Rea and Nasrallah, 2015), the AtS1pr:cYFP-SRKb transgene conferred a strong SI response in C24 stigmas: in manual pollination assays, C24[AtS1pr:cYFP-SRKb] stigmas exhibited severe inhibition of the SCRb-expressing pollen derived from C24[SRKb-SCRb] plants, hereafter referred to as SCRb-pollen (Fig. 1B, top row). However, C24[AtS1pr:cYFP-SRKb] plants produced abundant seed because they lack the SCRb gene.
Stigma-expressed SCR protein will hereafter be designated cis-SCRb to distinguish it from SCR protein that originates from the pollen coat or recombinant SCR protein purified from bacteria, both of which will be referred to as trans-SCR. Due to the presence of an N-terminal signal sequence in SCR transcriptional units (Schopfer et al., 1999; Kusaba et al., 2001), cis-SCR proteins are expected to be secreted and to be localized in the stigma epidermal cell wall. Consequently, in stigmas expressing both SRKb and SCRb, SCRb might potentially interact with the extracellular domain of SRKb, possibly resulting in activation of SRK and SI signaling. We reasoned that if cis-SCRb is able to activate the SRKb receptor, F1 plants containing both the AtS1pr:cYFP-SRKb and AtS1pr:SCRb transgenes would not set seed, despite the fact that the pollen grains of these plants lack SCRb protein. A total of eight AtS1pr:cYFP-SRKb + AtS1pr:SCRb F1 plants derived from each of the AtS1pr:SCRb b1, b2, and b3 transgenic lines was analyzed. Contrary to our expectation, all F1 plants produced large amounts of seed, indicating that their stigma epidermal cells did not exhibit a constitutive SI response.
To understand why a constitutive SI response was not observed, F1 stigmas were pollinated with SCRb-pollen. As shown in Figure 1B, Supplemental Figure S1A, and Supplemental Table S1, F1 stigmas were found to be fully receptive to SCRb-pollen, indicating that the SI response was not operational in these plants. Importantly, the breakdown of SI cosegregated with the AtS1pr:SCRb transgene in F2 plants. As shown in Supplemental Table S2 for F2 plants derived from the AtS1pr:SCRb b1 and b2 transgenic lines, among F2 plants that inherited the AtS1pr:cYFP-SRKb transgene, the breakdown of SI was observed only in the plants that also inherited the AtS1pr:SCRb transgene. This result demonstrates that the disruption of SI in AtS1pr:cYFP-SRKb + AtS1pr:SCRb stigmas was not due to some defect in the SRKb transgene that might have been introduced during the generation of F1 plants but was rather caused by the AtS1pr:SCRb transgene.
Thus, instead of activating the SRKb-mediated SI response, the cis-SCRb protein disrupted this response. To determine if the inhibitory effect of cis-SCRb on SRKb-mediated SI signaling was correlated with the expression level of the AtS1pr:SCRb transgene, the AtS1pr:SCRb(b1) highly expressing line was compared to a fourth AtS1pr:SCRb line, designated AtS1pr:SCRb(b4), whose stigmas exhibited a 12-fold lower level of SCRb transcripts than AtS1pr:SCRb(b1; Fig. 1C). Each of the AtS1pr:SCRb(b1) and AtS1pr:SCRb(b4) lines was crossed with a self-incompatible C24 transgenic line that was homozygous for a single integration of the SRKb-SCRb gene pair, exhibited a strong and stable SI response, and, as a result, produced no seed (Fig. 1A; Nasrallah et al., 2004). Analysis of the SRKb-SCRb + AtS1pr:SCRb F1 plants showed that F1 plants derived from crossing the SRKb-SCRb line with AtS1pr:SCRb(b1), which exhibit high levels of cis-SCRb transcripts, produced fully elongated seed-filled siliques (35.6 ± 4.3 seed, n = 10), and their stigmas were fully compatible with SCRb-pollen (Fig. 1D; Supplemental Fig. S1B). By contrast, F1 plants derived from the SRKb-SCRb × AtS1pr:SCRb(b4) cross, which exhibit low levels of cis-SCRb transcripts, produced short siliques containing some seed (9.3 ± 5.6 seed, n = 10), and their stigmas showed partial breakdown of the SI response (Fig. 1D; Supplemental Fig. S1B). These phenotypes indicate that cis-SCRb protein exerts an inhibitory effect on the SI response in a concentration-dependent manner.
To confirm that the contrasting effects of pollen-derived SCRb and stigma-expressed SCRb on the SI response are not caused by aberrant modification of the SCRb proteins in stigma epidermal cells, the biological activity of stigma-expressed SCRb was investigated using a pollination bioassay modified from Kachroo et al. (2001). Total proteins were extracted from stigmas of an AtS1pr:SCRb highly expressing line and applied to the stigmas of SRKb-SCRb homozygous plants (see “Materials and Methods”). As shown in Figure 1E, this treatment induced the SI response and caused the stigma epidermal cells to inhibit the germination of the SCRb-lacking pollen grains of untransformed C24 plants, allowing on average only approximately six pollen tubes to penetrate through the stigma cell wall (6.7 ± 4.1; n = 18). By contrast, pretreatment of SRKb-SCRb stigmas with stigma extracts from C24 untransformed plants did not cause inhibition of SCRb-lacking pollen grains (Fig. 1E). This result indicates that stigma-expressed SCRb is biologically active and is as able to activate the SI response as pollen coat-derived SCRb.
Taken together, our results indicate that the SCR ligand can act as an activator or a repressor of SI signaling, depending on whether it originates from outside (i.e. trans-SCR) or from inside (i.e. cis-SCR) the stigma epidermal cell. The inhibitory effect of cis-SCR will be called cis-inhibition, a term that has been used for similar phenomena in metazoans (Yaron and Sprinzak, 2012).
SCR-Mediated cis-Inhibition of SI Signaling Depends on Allele-Specific Interaction between SRK and Its SCR Ligand
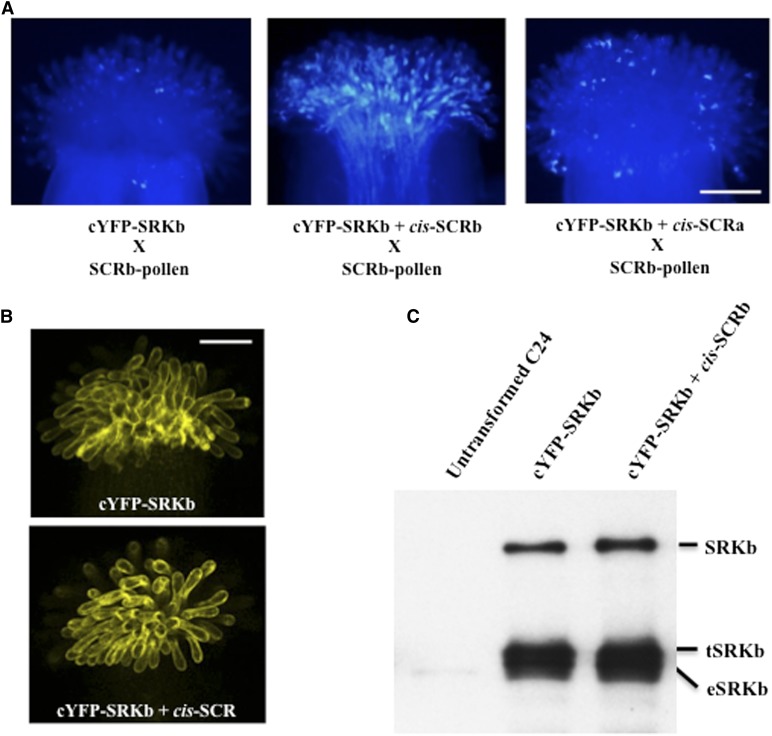
To determine if the inhibitory effect of cis-SCR is allele specific, transgenic lines were generated, which expressed the A. lyrata SCRa variant (Kusaba et al., 2001) under control of the AtS1 promoter. Three independent AtS1pr:SCRa transgenic lines exhibiting appreciable levels of stigma-expressed SCRa transcripts (Supplemental Fig. S1C) were each crossed with the AtS1pr:cYFP-SRKb transgenic line described above. Manual pollination of F1 stigmas with SCRb-pollen demonstrated that, unlike stigmas that expressed cYFP-SRKb and SCRb, stigmas that expressed cYFP-SRKb and SCRa exhibited an SI response toward SCRb-pollen that was as strong as that exhibited by stigmas that expressed only cYFP-SRKb (Fig. 2A; Supplemental Fig. S1C; Supplemental Table S3). This allelic specificity suggests that, like trans-activation of SRK-mediated SI signaling, the cis-inhibition phenomenon described here is based on allele-specific interaction between SRK and its cognate SCR.
Figure 2.
Allele-specific interaction of cis-SCRb with SRKb and the levels of SRKb proteins in SCRb-expressing stigmas. A, Effects of cis-SCRb and cis-SCRa on the SRKb-mediated SI response. Pollination assays using SCRb-pollen grains and stigmas expressing only cYFP-SRKb (left), cYFP-SRKb and cis-SCRb (middle), or cYFP-SRKb and cis-SCRa (right) demonstrate that the SRKb-mediated SI response is disrupted by expression of cis-SCRb but is not affected by expression of cis-SCRa. B, Confocal images of stigma epidermal cells expressing only cYFP-SRKb (top) or expressing both cYFP-SRKb and cis-SCRb (bottom). Note that the overall cYFP signal intensities, which report on the localization of all three SRKb protein products (full-length SRKb, eSRKb, and tSRKb), are equivalent in the two types of stigmas. C, Protein immunoblot analysis of extracts from untransformed C24 stigmas, stigmas expressing only cYFP-SRKb from an AtS1pr:cYFP-SRKb hemizygous plant, and stigmas expressing both cYFP-SRKb and cis-SCRb from an AtS1pr:cYFP-SRKb + AtS1pr:SCRb F1 plant derived from the AtS1pr:SCRb(b1) line. Each sample was obtained by extracting 25 stigmas from one plant. Note that the levels of full-length SRKb, eSRKb, and tSRKb proteins are not affected by expression of cis-SCRb. Bars = 0.1 mm.
SCR-Mediated cis-Inhibition Is Not Caused by Degradation of SRK
Several transmembrane receptors undergo endocytosis followed by protein degradation after interacting with their ligands (Sorkin and Goh, 2008; Bar and Avni, 2014). For example, in plants, the plasma membrane-localized pattern recognition receptor FLAGELLIN SENSITIVE2 (FLS2) is degraded after prolonged incubation of cells with its flagellin22 (flg22) ligand (Robatzek et al., 2006). To determine if disruption of the SI response in the presence of cis-SCRb might be caused by degradation of SRKb protein, the levels of cYFP-SRKb proteins in stigmas that express cis-SCRb were compared to those that lacked cis-SCRb. To avoid differences in cYFP-SRKb expression levels resulting from different AtS1pr:cYFP-SRKb transgene dosage, protein levels were compared in F1 plants containing one copy of the AtS1pr:cYFP-SRKb transgene that were generated either by crossing an AtS1pr:cYFP-SRKb homozygote with the AtS1pr:SCRb(b1) homozygous line or by crossing the same AtS1pr:cYFP-SRKb homozygote with a C24 untransformed plant.
It should be noted that SRK genes produce three types of proteins: (1) the full-length transmembrane SRK signaling receptor, which consists of the extracellular S domain, a single-pass transmembrane domain, and a cytoplasmic kinase domain (Stein et al., 1991); (2) a soluble protein (designated eSRK), which consists of only the S domain and is produced from a short SRK transcript (Stein et al., 1991; Giranton et al., 1995); and (3) a membrane-anchored truncated form of SRK (designated tSRK), which lacks the kinase domain and is generated by proteolytic cleavage of the full-length receptor (Shimosato et al., 2007). Because of their identical N termini, the three SRKb protein species produced in AtS1pr:cYFP-SRKb stigmas carry the cYFP tag and are therefore detectable both by confocal microscopy of live stigma epidermal cells and by protein immunoblot analysis using anti-GFP antibodies. As shown in Figure 2, B and C, the stigmas of AtS1pr:cYFP-SRKb hemizygous plants were found to exhibit the same overall cYFP-SRKb signal by live imaging of epidermal cells (Fig. 2B) and to accumulate equivalent levels of the three SRKb protein products by protein gel-blot analysis (Fig. 2C), irrespective of the presence or absence of cis-SCRb. Thus, the SCR-mediated cis-inhibition of SRK signaling is not caused by degradation of the full-length SRKb protein or its eSRK and tSRK isoforms.
cis-SCR Causes Mislocalization of the SRKb Protein
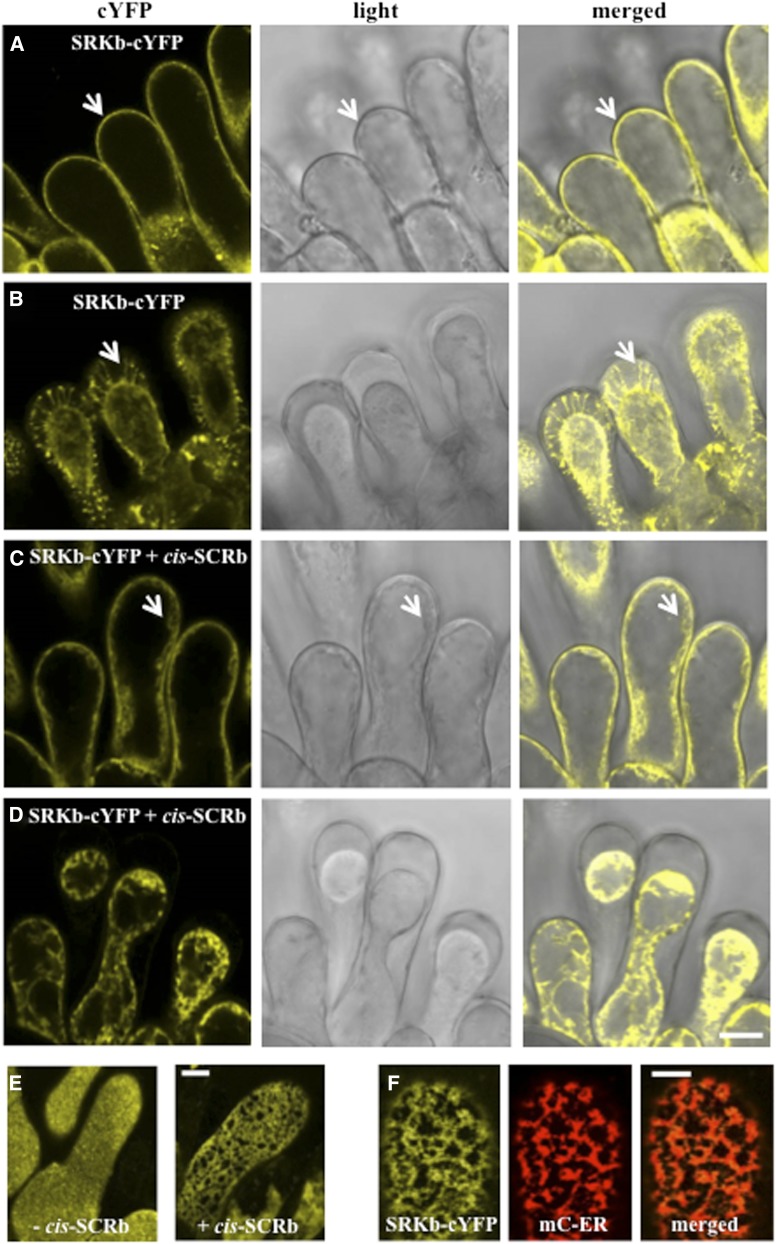
To investigate further the mechanism by which cis-SCR disrupts the SRK-mediated SI response, the possible effect of cis-SCRb on the subcellular localization of SRKb was investigated. To visualize only the full-length SRKb signaling receptor without interference from the eSRKb and tSRKb proteins, a version of SRKb carrying a C-terminal cYFP tag (designated SRKb-cYFP), which confers a strong incompatible response toward SCRb-pollen (Yamamoto et al., 2014; Rea and Nasrallah, 2015) was used. As shown in Figure 3A, imaging of stigma epidermal cells lacking cis-SCRb showed the SRKb-cYFP signal to be localized primarily at the cell periphery, consistent with the previously demonstrated plasma membrane localization of the receptor (Yamamoto et al., 2014; Rea and Nasrallah, 2015). Moreover, as is typical of plasma membrane-localized proteins in plants and previously shown for SRKb-cYFP (Rea and Nasrallah, 2015), upon plasmolysis of stigma epidermal cells, the SRKb-cYFP signal was observed in the stretched portions of the plasma membrane called Hechtian strands, which form upon retraction of the protoplast from the cell wall following plasmolysis (Fig. 3B). By contrast, in the presence of cis-SCRb, the SRKb-cYFP signal accumulated primarily within the protoplast (Fig. 3C) rather than at the plasma membrane. Consistent with this conclusion, and despite the roughly equivalent overall SRKb-cYFP signal intensities observed in the presence and absence of cis-SCRb, cYFP-labeled Hechtian strands were not observed in plasmolyzed stigma epidermal cells under our standard visualization parameters (Fig. 3D). It was only when the confocal images were manipulated by increasing overall brightness that cYFP-labeled positive Hechtian strands became faintly visible (Supplemental Fig. S2A).
Figure 3.
Aberrant localization of the full-length SRKb receptor in stigmas expressing cis-SCRb. A and B, Distribution of SRKb-cYFP in stigma epidermal cells lacking cis-SCRb. The SRKb-cYFP signal, which reports exclusively on the full-length SRKb receptor, is detected primarily at the periphery (arrow) of stigma epidermal cells (A) and in the Hechtian strands (arrow) of plasmolyzed stigma epidermal cells (B), consistent with plasma membrane localization (Rea and Nasrallah, 2015). C to F, Distribution of SRKb-cYFP in stigma epidermal cells expressing cis-SCRb. The SRKb-cYFP signal is observed in cytoplasmic structures (C, arrow) but is not detected in the Hechtian strands of plasmolyzed cells (D), except when image brightness is increased (Supplemental Fig. S2A). E, Cell surface images of stigma epidermal cells lacking cis-SCRb showing uniform distribution of full-length SRKb at the cell surface (left) and of stigma epidermal cells expressing cis-SCRb showing a reticulate pattern of SRKb-cYFP distribution (right). F, Colocalization of SRKb-cYFP and an mC-labeled ER marker (mC-ER) in the presence of cis-SCRb. The images show stigma epidermal cells of plants containing the AtS1pr:SRKb-cYFP, AtS1pr:SCRb, and AtS1pr:mC-HDEL transgenes. Note the perfect overlap of the SRKb-cYFP signal with the mC-HDEL signal. Bars = 10 µm (A–E) and 5 µm (F).
The different distribution of SRKb-cYFP in the absence and presence of cis-SCRb was particularly evident in surface views of stigma epidermal cells, in which the cYFP signal exhibited a relatively uniform distribution at the cell surface in the absence of cis-SCRb but a reticulate pattern in the presence of cis-SCRb (Fig. 3E). Because a reticulate pattern of localization is characteristic of proteins localized in the cortical ER, the possibility that SRKb-cYFP might be primarily localized in the ER of stigmas expressing cis-SCRb was investigated. Toward this goal, the AtS1pr:SRKb-cYFP homozygous line was retransformed with an AtS1pr:mCherry (mC)-HDEL (His-Asp-Glu-Leu) transgene designed for stigma epidermal cell expression of the mC variant of red fluorescent protein fused at its C terminus to the ER retention signal HDEL (see “Materials and Methods;” Nelson et al., 2007; Rea and Nasrallah, 2015). As shown in Figure 3F, in the presence of cis-SCRb, the bulk of the SRKb-cYFP signal colocalized with the ER marker. This result indicates that cis-SCRb disrupts the proper localization of SRKb by causing the receptor to be retained in the ER.
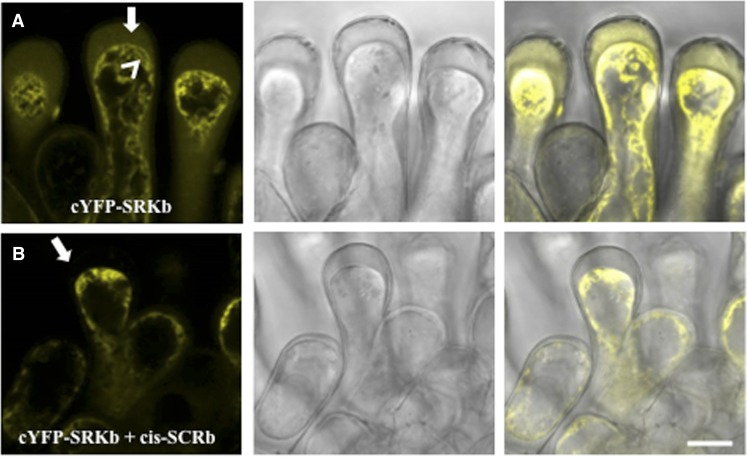
To determine if cis-SCRb also affects the localization of other SRKb isoforms, plants homozygous for the AtS1pr:cYFP-SRKb transgene described above were used for confocal microscopy. In plasmolyzed stigma epidermal cells, the cYFP-SRKb signal was detected in the apoplast (i.e. the expected location of eSRK proteins) in the absence (Fig. 4A) but not in the presence of cis-SCRb (Fig. 4B). Therefore, cis-SCRb affects the distribution of the eSRK isoform in addition to that of the full-length SRKb receptor, suggesting that cis-SCRb interacts with both eSRKb and full-length SRKb in the ER. Because tSRK has been shown to accumulate in the ER of stigma epidermal cells that lack cis-SCRb (Rea and Nasrallah, 2015), this study cannot determine if cis-SCRb affects tSRK localization.
Figure 4.
Effect of cis-SCRb on the localization of SRKb isoforms in stigma epidermal cells. The confocal images show plasmolyzed stigma epidermal cells expressing only cYFP-SRKb (A) or both cYFP-SRKb and cis-SCRb (B). A, In the absence of cis-SCRb, the cYFP-SRKb signal observed in the apoplast (arrow) likely represents eSRKb, while the cYFP-SRKb signal detected in cytoplasmic structures (arrowhead) represents the localization of the eSRK/tSRK within the ER, as previously demonstrated (Rea and Nasrallah, 2015). Note that cYFP-positive Hechtian strands are obscured by the apoplastic signal emanating from cYFP-eSRKb proteins but are visible when image brightness is increased (Supplemental Fig. S2B). B, In the presence of cis-SCRb, the cYFP-SRKb signal is observed exclusively in intracellular compartments but not in the apoplast (arrow). Bars = 10 µm.
DISCUSSION
The SRK/SCR-mediated signaling that occurs upon contact between a stigma epidermal cell and a pollen grain conforms to the classical view of cell contact-dependent receptor-based cell-to-cell signaling, in which the directionality of signaling is defined by expression of a receptor in the signal-receiving cell and of its cognate ligand in a signal-sending adjacent cell. The coexpression of SRK and SCR in stigma epidermal cells and the resulting cis-inhibition of SRK signaling described in this study simulate the situation observed for some contact-dependent receptor-ligand systems in metazoans. The phenomenon of cis-inhibition of a receptor by its coexpressed ligand has been well documented for several signaling pathways in metazoans, such as the Notch signaling pathway that functions in cell fate specification (Shaya and Sprinzak, 2011; Yaron and Sprinzak, 2012), the semaphorin and ephrin signaling pathways that function in axon pathfinding/guidance (Yaron and Sprinzak, 2012), and the Major Histocompatibility Complex signaling cascades that underlie immune responses (Held and Mariuzza, 2008).
Mechanism of cis-Inhibition of SRK by Its Cognate SCR
Despite similar outcomes, cis-inhibition of SRK by its soluble SCR ligand differs mechanistically from cis-inhibition of the Notch, semaphorin, and ephrin receptors. In the case of these metazoan receptors, all of which have membrane-bound ligands (Yaron and Sprinzak, 2012), it has been suggested that cis-interactions, like trans-interactions, of the receptors with their ligands occur at the cell surface (Carvalho et al., 2006; Glittenberg et al., 2006; Haklai-Topper et al., 2010). For example, in cis-inhibition of the Notch receptor, the amount of receptor at the cell surface is not affected by coexpression of ligand; rather, cis- and trans-interactions utilize different combinations of ligand-interacting domains, resulting in different effects on signaling outcomes (Glittenberg et al., 2006; Fleming et al., 2013). By contrast, cis-SCRb was found to interfere with proper targeting of the SRKb protein to the plasma membrane by causing the receptor to be trapped in the ER. Because the severity of cis-inhibition was positively correlated with the level of SCRb expression in stigma epidermal cells, the disruption of the SI response by cis-SCRb may be ascribed to the reduced number of SRKb molecules at the cell surface and therefore to their reduced availability for productive interaction with pollen-derived trans-SCRb.
The entrapment of SRKb in the ER, together with the observation that SCR exerts its cis-inhibitory effect in an allele-specific manner (i.e. resulting from coexpression of self SCRb but not of nonself SCRa), suggests that export of SRKb from the ER is prevented by interaction of the receptor with cis-SCRb in the secretory pathway, likely via its extracellular domain, which is the SCR-binding domain (Kachroo et al., 2001). In this respect, cis-inhibition of SRK is similar to cis-inhibition of the low-density lipoprotein receptor-related protein1 (LRP1) by secreted forms of its ligands (Willnow et al., 1996; Sakamoto et al., 2011). It has been shown that, in the absence of the specific chaperone receptor-associated protein, cis-inhibition of LRP1 is caused by premature binding of the receptor to ligand in the secretory pathway, resulting in the formation of receptor aggregates that are trapped in the ER (Willnow et al., 1996; Sakamoto et al., 2011). However, unlike cis-inhibition of LRP1, which causes reductions of approximately 75% in receptor abundance (Willnow et al., 1995), SCRb-mediated cis-inhibition did not cause a discernible change in the overall level of SRKb in any of its three isoforms. Further analysis is required to determine if entrapment of SRKb in the ER is caused by cis-SCRb-induced receptor aggregation and to understand why ER-trapped SRKb molecules are not subject to ER-associated degradation. In any case, the mechanistic similarity of cis-inhibition of SRKb and LRP1 on the one hand and of the Notch, ephrin, and semaphorin receptors on the other hand suggests that the site of receptor-ligand cis-interactions (i.e. in the ER or at the cell surface) and the mechanism of cis-inhibition might depend on whether the ligand is secreted or membrane bound.
Ligand-mediated cis-inhibition of SRK signaling also differs from cis-inhibition of signaling in metazoans with respect to the importance of these interactions and of their interplay with trans-interactions for normal developmental or physiological processes. While ligand-mediated cis-inhibition of signaling pathways in metazoans was first observed by overexpression of a ligand in the same cell as its receptor (Klein et al., 1997), this phenomenon is now recognized as an endogenous regulatory mechanism that is important for robust signaling and proper orchestration of cellular functions (Shaya and Sprinzak, 2011). By contrast, cis-inhibition of SRK by SCR has no physiological role in fine-tuning the response of the stigma epidermal cell to incoming pollen grains. Rather, it must be avoided for the SI system to function properly. Clearly, the integrity of the SI response must depend on tight regulation of the SCR gene and its exclusive expression in anthers, because even a relatively low-level expression of SCRb transcripts in stigma epidermal cell was found to be sufficient for weakening the SI response and allowing the growth of SCRb-pollen tubes. Therefore, cis-inhibition might provide a safeguard against the sterility that would be caused by constitutive activation of the SI response in stigmas, should the tight developmental regulation of the SCR gene be disrupted by mutation.
Ligand-Mediated cis-Inhibition as a Strategy for Inhibiting Cell Surface Receptor Function and Manipulating Signaling Pathways in Plants
By providing, to our knowledge, the first example of cis-inhibition of a plant receptor, our results suggest that at least some plant receptors that are trans-activated by protein ligands may also be subject to ligand-mediated cis-inhibition. The Arabidopsis genome contains more than 1,000 genes encoding putative peptide ligands (Lease and Walker, 2006), most of which have no known function. The most common approach used to assess the potential roles of these peptides in signaling has been to overexpress the peptide (Strabala et al., 2006; Jun et al., 2010), an approach that avoids complications stemming from possible functional redundancy of closely related peptide ligands. The cis-inhibition phenomenon described in this study suggests that any phenotypic changes that might be observed in plants that overexpress a given peptide ligand may not reflect gain-of-function phenotypes as is typically assumed but rather may be caused by cis-inhibition of the receptor by its coexpressed ligand. It should be noted, however, that ligand-mediated cis-inhibition of receptors is unlikely to apply to all receptor-ligand pairs in plants. In metazoans, the LRP receptor, which interacts with several ligands, is trapped in the ER by only a few of these ligands (Willnow et al., 1996). Moreover, overexpression of a ligand in receptor-expressing cells and exogenous application of ligand to these cells produced the same result for some ligands but opposite effects for other ligands (Strabala, 2008), likely due to the trans-activation of signaling pathways in the former case and cis-inhibition of signaling in the latter case.
Should cis-inhibition be observed in plant receptor-ligand systems other than the SRK-SCR system described here, it will be important to determine on a case-by-case basis if this phenomenon might have a physiological role in plants. In any case, the discovery of SCR-mediated cis-inhibition of SRK suggests a novel and potentially powerful strategy for inhibiting cell surface receptor function and manipulating signaling pathways in plants. Such a strategy would entail coexpressing a ligand in the same cell as its receptor and, in some cases, may also require addition of an ER retention signal to the ligand to ensure entrapment of the receptor in the ER. Inactivation of receptors by ligand-mediated cis-inhibition may be used as an alternative to conventional gene silencing methods, which target transcript production or stability. Like gene silencing methods, ligand-mediated cis-inhibition can result in phenotypes of varying strengths, as demonstrated by our observation that the severity of SRK cis-inhibition depends on the amount of SCR produced in stigma epidermal cells. However, a cis-inhibition strategy has several advantages over conventional gene silencing approaches, which can suffer from off-target effects (Jackson et al., 2003; Scacheri, et al., 2004) and the generation of epigenetic marks that may persist into subsequent generations (Cuzin et al., 2008; Paszkowski and Grossniklaus, 2011; Borges and Martienssen, 2013). By contrast, in cis-inhibition, the interaction between a receptor and its cognate ligand is highly specific, the distribution of the ligand may be controlled by use of cell type-specific promoters, and the suppression of signaling is easily reversed by segregation of the ligand-encoding transgene in transgenic progenies.
Ligand-Mediated cis-Inhibition of SRK as a Strategy for Hybrid Seed Production
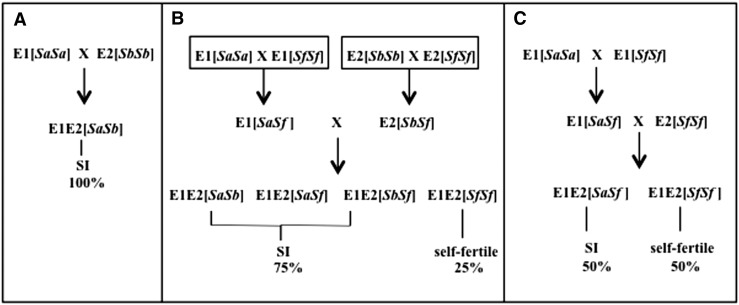
In the particular case of SI, cis-inhibition of SRK signaling has potential for use in SI-based schemes designed for production of hybrid cultivars and seeds. These hybrids are commercially desirable because of their superior performance relative to their nonhybrid counterparts, but their efficient production requires the use of genetic pollination control systems that prevent selfed seed production while allowing outcrossed seed production. Although SI clearly fulfills this requirement, several issues complicate its utilization in practice. All SI-based hybrid-seed production schemes that have been implemented or proposed to date employ one or more self-incompatible inbred lines as parental lines (Fig. 5), the maintenance of which requires the use of some method to overcome SI. However, the strategies that are currently available for overcoming SI, including manual self-pollination in young floral buds before the onset of SI in developing stigmas, treatments with sodium chloride or carbon dioxide, and micropropagation, are all laborious and inefficient (Gray, 1993; Ockendon and Smith, 1993; Nasrallah, 2004). Furthermore, a large proportion of hybrid plants generated by currently used SI-based breeding programs remain self-incompatible (Fig. 5). This fact is a major impediment for the use of SI-based hybridization schemes in Brassicaceae seed crops such as oilseed rape (Brassica napus), because seed production in self-incompatible hybrids can only occur when pollinators transfer pollen from other self-incompatible plants carrying different SI alleles or pollen from self-fertile plants. Consequently, seed production in these crops is adversely impacted by environmental conditions that are not suitable for insect pollination and by the reduced pollinator populations that are increasingly observed in the field.
Figure 5.
Examples of conventional SI-based methods for the generation of hybrids in the Brassicaceae. Three strategies for SI-based hybrid breeding are shown (modified from Nasrallah, 2004). A, A strategy that uses two different elite self-incompatible inbred lines (E1[SaSa] and E2[SbSb]). This method requires intensive labor for the maintenance of parental inbred lines, and it generates only self-incompatible hybrid plants, which is an undesirable outcome for seed crops. B and C, Two strategies that incorporate isogenic elite self-fertile lines (E[SfSf]) into the crossing scheme. These strategies were developed to reduce the labor associated with the maintenance of self-incompatible inbred lines and to produce populations of hybrid plants that contain some self-fertile plants. In B, two elite self-incompatible inbred lines (E1[SaSa] and E2[SbSb]) are each pollinated with an isogenic self-fertile line (E1[SfSf] and E2[SfSf], respectively), and the self-incompatible progenies from each cross are then used to generate hybrid plants. In C, an elite self-incompatible inbred line (E1[SaSa]) is pollinated with an isogenic self-fertile line (E1[SfSf]). The self-incompatible progenies are then pollinated with another elite self-fertile inbred line (E2[SfSf]) for hybrid production. Sa refers to the Sa haplotype containing the SRKa and SCRa genes, Sb refers to the Sb haplotype containing the SRKb and SCRb genes, and Sf refers to a nonfunctional S haplotype.
These problems may be overcome by the development of breeding and seed production schemes in which transgenes designed for stigma expression of specific SCR variant proteins are used as suppressors of the SI response. The use of cis-SCR transgenes, or SRK off switches, is expected to provide a robust method for both maintenance of self-incompatible parental lines and large-scale production of hybrids and their seed. In the following, two cis-SCR-based strategies are outlined, which may be used separately or in combination depending on whether the crop is a vegetable or root crop, for which only an efficient strategy for the maintenance of self-incompatible parental lines is required, or a seed crop, which additionally requires a strategy for large-scale seed production from hybrid plants in the field. In all cases, SRK off switches would be tailored to the particular S haplotypes harbored by elite inbred lines already used in current hybrid breeding programs (compare Figs. 5 and 6B). Moreover, the proposed cis-SCRb-based schemes are applicable to both naturally self-incompatible inbred lines and transgenic self-incompatible plants that are generated by transformation with matched alleles of the SRK and SCR genes.
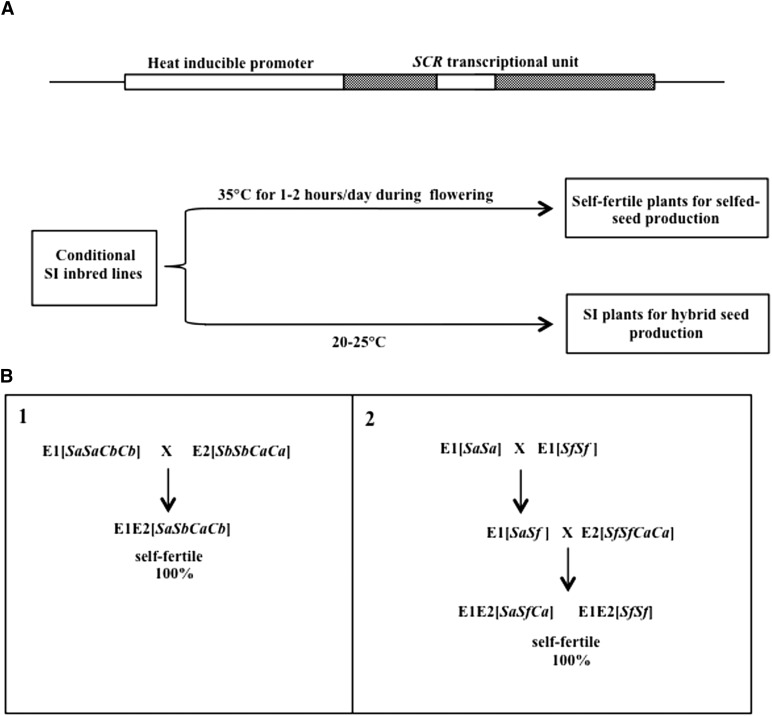
Figure 6.
Novel cis-SCR-based strategies for the maintenance of self-incompatible inbred lines and generation of self-fertile hybrids. A, An inducible cis-SCR strategy using a heat-inducible promoter for the maintenance of self-incompatible inbred lines. The diagram shows a heat-inducible cis-SCR-encoding transgene (top) and seed production in conditionally self-incompatible plants under inducing and noninducing conditions (bottom). B, Novel schemes for the generation of self-fertile hybrids that may be incorporated into current breeding programs. 1, A scheme modified from the conventional hybrid breeding method illustrated in Figure 5A: A transgene containing the SRKb off switch, AtS1pr:SCRb (Cb), is introduced into an elite self-incompatible inbred line carrying the Sa haplotype (E1[SaSa]) to generate E1[SaSaCbCb], and a transgene containing the SRKa off switch, AtS1pr:SCRa (Ca), is introduced into an elite self-incompatible inbred line carrying the Sb haplotype (E2[SbSb]) to generate E2[SbSbCaCa]. All hybrid progenies derived from a cross between these two modified parental lines are self-fertile. 2, A scheme modified from the conventional method illustrated in Figure 5C: A transgene containing the SRKa off switch (Ca) is introduced into an elite self-fertile inbred line (E2[SfSf]) to generate E2[SfSfCaCa]. All hybrid progenies derived from crossing the self-incompatible E1[SaSa]) line with the E2[SfSfCaCa] line are self-fertile. Both schemes produce self-fertile hybrids that set seed autonomously and do not require pollinator activity. Sa refers to the Sa haplotype containing the SRKa and SCRa genes, Sb refers to the Sb haplotype containing the SRKb and SCRb genes, and Sf refers to a nonfunctional S haplotype.
For the maintenance of elite self-incompatible inbred parental lines in vegetable and seed Brassicaceae crops, a practical approach is to control expression of cis-SCR using a conditional promoter, such as a heat-inducible promoter (Fig. 6A). The heat-inducible cis-SCR transgene would be introduced into these elite inbred lines and the resulting transformants would be grown in polythene tunnels or greenhouses, as is common practice for hybrid production in Brassica spp. crops. The transformants would express SI in normal ambient temperatures but would become self-fertile when expression of the SRK off switch is induced by exposure to higher temperatures during seed production (Fig. 6A). Because the process of pollen germination and pollen tube penetration through the wall of stigma epidermal cells occurs within a few minutes of pollen-stigma contact during compatible pollinations in the Brassicaceae, this strategy would require only brief daily heat treatments during the flowering phase and is therefore not expected to have negative effects on flower and seed development.
For large-scale seed production from hybrid plants in seed crops, the SRK off switch must function only in F1 hybrids and not in their parental inbred lines. Therefore, the SRK off switch for a particular SRK variant would not be introduced directly into the parental line that expresses this SRK variant but into the other parental inbred line used in the cross, whether the cross involves only self-incompatible parental lines or a combination of self-incompatible and self-fertile inbred lines (Fig. 6B). For example, in a cross between the self-incompatible inbred parental lines SaSa and SbSb, the SRKa off switch would be introduced into the SbSb inbred and the SRKb off switch would be introduced into the SaSa inbred (Fig. 6B). These two modified inbred lines would still express SI because they carry an SRK off switch that does not inactivate the function of the resident SRK variant. However, the hybrid progenies produced by crossing these two modified parental lines would be self-fertile because the SRKa off switch suppresses SRKa function and the SRKb off switch suppresses SRKb function (Fig. 6B). Moreover, a major advantage of these schemes is that all produced hybrid plants will be self-fertile, unlike conventional SI-based hybrid breeding programs, in which a large proportion of hybrid plants retains SI (compare Figs. 5 and 6B). As a result, the cis-inhibition approach will produce crops that exhibit increased seed production and a reduced requirement for high pollinator activity.
MATERIALS AND METHODS
Plant Materials, Growth Conditions, Construction of Transgenes, and Plant Transformation
All experiments described in this study were performed using plants of the Arabidopsis (Arabidopsis thaliana) C24 accession. Plants were grown at 22°C and a photoperiod of 16 h. The self-incompatible C24[SRKb-SCRb] transgenic plants were described previously (Nasrallah et al., 2004).
The AtS1pr:SCRb and AtS1pr:SCRa transformation plasmids were generated as follows. The SCRb and SCRa exons with intervening intron and 3′ regulatory sequences were amplified from previously described bacterial artificial chromosome clones of the Arabidopsis lyrata Sb and Sa haplotypes (Kusaba et al., 2001), using forward primers located immediately upstream of the start codon and containing KpnI restriction sites together with reverse primers containing Sma1 restriction sites. The following gene-specific primers were used: for SCRb, 5′-GGGGTACCATTTTTTCTTGATTCATGA-3′ and 5′-CCCCCCGGGACAGATTTGATTGGTTAGTT-3′, and for SCRa, 5′-GGGGTACCAAGAAAATCATGAGGTGTAGTG-3′ and 5′-CCCCCCGGGAGTTTCATAACCTAGTAAT-3′. PCR products were then ligated into the pGEMTeasy vector (Promega). The absence of PCR-generated sequence errors in the SCRb and SCRa genomic fragments in these pGEMTeasy derivatives were confirmed by sequencing. The AtS1pr:SCRb transgene was assembled in the pCAMBIA1300 binary vector by inserting a 0.55-kb KpnI-EcoRI fragment containing the SCRb genomic region excised from the pGEMTeasy derivative and subsequently inserting a 1.2-kb BamHI-KpnI fragment containing the AtS1 promoter into the BamHI-KpnI sites located at the 5′ end of the SCRb genomic fragment. To generate the AtS1pr:SCRa construct, the SCRb genomic fragment in the AtS1pr:SCRb construct was excised as a KpnI-XmaI fragment and replaced with a KpnI-SmaI fragment containing the SCRa sequence.
The construction of the AtS1pr:cYFP-SRKb and AtS1pr:SRKb-cYFP chimeric genes was described previously (Rea and Nasrallah, 2015). In brief, the cYFP coding sequence together with a linker sequence, which serves to minimize improper folding of the cYFP-tagged SRKb protein, was inserted using a recombinant PCR strategy into a pCAMBIA1300 derivative containing the AtS1pr:SRKb gene. The cYFP coding sequence was inserted either directly upstream of the stop codon to express C-terminally tagged SRKb or directly downstream of the signal sequence to express N-terminally tagged SRKb (Yamamoto et al., 2014; Rea and Nasrallah, 2015). To generate the AtS1pr:mC-HDEL construct, the previously described 35Spr:mC-HDEL construct (Nelson et al., 2007) was modified by replacing the double Cauliflower mosaic virus 35S promoter with the AtS1 promoter by recombinant PCR (Rea and Nasrallah, 2015).
All plasmid constructs were introduced into Agrobacterium tumefaciens strain GV3101, and transgenic plants were generated in the C24 accession using the floral dip method (Clough and Bent, 1998). Seeds were germinated on Murashige and Skoog medium (Sigma) containing 25 µg mL–1 kanamycin sulfate for selection of plants carrying the SRKb-SCRb transgene, 50 µg mL–1 hygromycin for selection of plants carrying the AtS1pr:SCRb, AtS1pr:cYFP-SRKb, or AtS1pr:SRKb-cYFP transgenes, and 10 µg mL–1 glufosinate-ammonium for selection of plants carrying the AtS1pr:mC-HDEL transgene.
For each construct, several independent transformants were generated, and unless specifically noted, all transgenic plants used for the analyses described herein were homozygous plants in which the transfer DNA fragment had integrated at a single chromosomal location as determined by segregation analysis in transgenic progenies. To identify the presence of SRKb, SCRb, and SCRa in transgenic plants, genomic DNA was subjected to PCR using the following gene-specific primers: SRKb, 5′-TGGGTTGGGATGTCAAGAAAG-3′ and 5′-CAACTTCATCTTTCTCAGGCACAA-3′; SCRb, 5′-TGCTACTTTCTTCATTGTTTTT-3′ and 5′-CGTCTCCTCTCCGTTGTT-3′; and SCRa, 5′-CAGACAAGAAAATCATGAGGTGTAGTG-3′ and 5′-ACATCCCAAAAACTTCGCAGTAAATAC-3′.
Pollination Assays and Bioassay of Stigma-Expressed SCRb
Pollination assays were performed using the stigmas of stage-12 flower buds (Smyth et al., 1990), which is the final bud stage before anthesis. These stigmas and their epidermal cells are fully developed, as shown by the confocal images in Figures 3 and 4. Stigmas were manually pollinated with wild-type pollen or with SCRb-pollen and were left for 2 to 4 h before fixation in a solution consisting of three parts ethanol and one part acetic acid for at least 30 min. The fixed stigmas were then softened in 1 n NaOH at 65°C for 15 min, washed in water, and stained in decolorized aniline blue before observing pollen tube growth by epifluorescence microscopy (Kho and Baër, 1968). For each plant tested, the pollination assay was performed on at least three stigmas and was repeated on at least two different dates.
To assess the biological activity of stigma-expressed SCRb, 50 stigmas of plants homozygous for the AtS1pr:SCRb transgene were ground in 5 µL of water containing 0.05% (v/v) Tween 20 (Research Organics). After centrifugation, the clarified stigma extract was applied to stage-12 flower buds of C24[SRKb-SCRb] homozygotes. The treated stigmas were allowed to dry for 1 h, after which they were pollinated with wild-type pollen grains. Stigma extracts prepared from C24 wild-type plants were used as control in the assays. For each assay, six stigmas from C24[SRKb-SCRb] plants were used per treatment (i.e. six stigmas for the cis-SCRb extract and six stigmas for the C24 wild-type extract), and three independent repeats were performed.
RNA Analysis
Total RNA was isolated from 40 stigmas of stage-12 flower buds using the Trizol reagent (Invitrogen), and 0.5 µg of total RNA was treated with DNase I (Invitrogen) according to the manufacturer’s instructions. RT-PCR was carried out using the SuperScript III One-Step RT-PCR System (Invitrogen) and the following gene-specific primers: SCRb, 5′-TGCTACTTTCTTCATTGTTTTT-3′ and 5′-CGTCTCCTCTCCGTTGTT-3′, and SCRa, 5′-GTTTTGTTTGTGGTTTCTTATGTT-3′ and 5′-GTCCCGTTTGCACTGTTCG-3′.
For the quantification of SCRb transcripts by real-time quantitative PCR, DNase-treated total RNA was reverse transcribed with oligo(dT) and the First-Strand cDNA Synthesis Kit for real-time PCR (Affymetrix). The following gene-specific primers were used: SCRb, 5′-TGCTACTTTCTTCATTGTTTTT-3′ and 5′-CGTCTCCTCTCCGTTGTT-3′, and ACTIN2, 5′-GCACCCTGTTCTTCTTACCG-3′ and 5′-AACCCTCGTAGATTGGCACA-3′. Real-time quantitative PCR was performed using the HotStart-IT SYBR Green qPCR Master Mix (Affymetrix) in an Applied Biosystems ViiA 7 Real-Time PCR System at 55°C annealing temperature for 40 cycles. Relative transcript levels in three replicates for each sample were calculated by the comparative threshold cycle method using the ViiA software.
Protein Gel-Blot Analysis
For analysis of cYFP-tagged SRKb proteins, total protein was extracted from 25 stigmas of stage-12 flower buds, subjected to SDS-PAGE using a 7.5% (w/v) polyacrylamide gel (Laemmli, 1970), and transferred to an Immobilon polyvinylidene fluoride membrane as described (Towbin et al., 1979). The membranes were treated with α-GFP mouse monoclonal primary antibody (1:3,000 dilution; Covance) followed by peroxidase-conjugated α-mouse monoclonal secondary antibody (1:6,000 dilution; Sigma-Aldrich). The cYFP-tagged SRKb proteins were detected using the Amersham ECL Plus Western Blotting Detection Kit (GE Healthcare Life Science).
Laser Scanning Confocal Microscopy
Stigmas from stage-12 flower buds were imaged at the Plant Cell Imaging Center of the Boyce Thompson Institute using a Leica TCS SP5 Laser Scanning Confocal Microscope equipped with a hybrid detection system. Stigmas were observed with 10× or 63× objectives. cYFP was excited with the argon laser (15% 514 nm), and emitted fluorescence was collected from 522 to 550 nm, while mCherry was excited with a diode-pumped, solid-state laser (15% 561 nm), and emitted fluorescence was collected from 598 to 644 nm. For plasmolysis experiments, stigmas were placed in 5% (w/v) NaCl for at least 5 min before imaging.
Sequence data from this article can be found in GenBank (http://www.ncbi.nlm.nih.gov) under accession numbers AlSCRa (BAB40984.1), AlSCRb (BAB40985), and AlSRKb (BAB40987).
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. Expression levels of stigma-expressed SCRb (cis-SCRb) and stigma-expressed SCRa (cis-SCRa) and their effects on the SRKb-mediated SI response.
Supplemental Figure S2. Confocal images of cYFP-labeled SRKb proteins in plasmolyzed stigma epidermal cells.
Supplemental Table S1. Pollination assays of F1 plants derived by crossing AtS1pr:SCRb transformants with an AtS1pr:cYFP-SRKb homozygote.
Supplemental Table S2. Pollination assays of AtS1pr:cYFP-SRKb-containing F2 plants.
Supplemental Table S3. Pollination assays of F1 plants derived by crossing AtS1pr:SCRa transformants with an AtS1pr:cYFP-SRKb homozygote.
Supplementary Material
Acknowledgments
We thank Pei Liu for pGEMTeasy derivatives containing SCRb and SCRa genomic fragments lacking promoter regions.
Glossary
- SI
self-incompatibility
- RT
reverse transcription
- ER
endoplasmic reticulum
Footnotes
This work was supported by the National Science Foundation (grant no. IOS–1146725).
Articles can be viewed without a subscription.
References
- Bar M, Avni A (2014) Endosomal trafficking and signaling in plant defense responses. Curr Opin Plant Biol 22: 86–92 [DOI] [PubMed] [Google Scholar]
- Boggs NA, Dwyer KG, Shah P, McCulloch AA, Bechsgaard J, Schierup MH, Nasrallah ME, Nasrallah JB (2009) Expression of distinct self-incompatibility specificities in Arabidopsis thaliana. Genetics 182: 1313–1321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borges F, Martienssen RA (2013) Establishing epigenetic variation during genome reprogramming. RNA Biol 10: 490–494 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carvalho RF, Beutler M, Marler KJ, Knöll B, Becker-Barroso E, Heintzmann R, Ng T, Drescher U (2006) Silencing of EphA3 through a cis interaction with ephrinA5. Nat Neurosci 9: 322–330 [DOI] [PubMed] [Google Scholar]
- Castellano G, Reid JF, Alberti P, Carcangiu ML, Tomassetti A, Canevari S (2006) New potential ligand-receptor signaling loops in ovarian cancer identified in multiple gene expression studies. Cancer Res 66: 10709–10719 [DOI] [PubMed] [Google Scholar]
- Chookajorn T, Kachroo A, Ripoll DR, Clark AG, Nasrallah JB (2004) Specificity determinants and diversification of the Brassica self-incompatibility pollen ligand. Proc Natl Acad Sci USA 101: 911–917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16: 735–743 [DOI] [PubMed] [Google Scholar]
- Cuzin F, Grandjean V, Rassoulzadegan M (2008) Inherited variation at the epigenetic level: paramutation from the plant to the mouse. Curr Opin Genet Dev 18: 193–196 [DOI] [PubMed] [Google Scholar]
- Dwyer KG, Kandasamy MK, Mahosky DI, Acciai J, Kudish BI, Miller JE, Nasrallah ME, Nasrallah JB (1994) A superfamily of S locus-related sequences in Arabidopsis: diverse structures and expression patterns. Plant Cell 6: 1829–1843 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleming RJ, Hori K, Sen A, Filloramo GV, Langer JM, Obar RA, Artavanis-Tsakonas S, Maharaj-Best AC (2013) An extracellular region of Serrate is essential for ligand-induced cis-inhibition of Notch signaling. Development 140: 2039–2049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giranton JL, Ariza MJ, Dumas C, Cock JM, Gaude T (1995) The S locus receptor kinase gene encodes a soluble glycoprotein corresponding to the SKR extracellular domain in Brassica oleracea. Plant J 8: 827–834 [DOI] [PubMed] [Google Scholar]
- Glittenberg M, Pitsouli C, Garvey C, Delidakis C, Bray S (2006) Role of conserved intracellular motifs in Serrate signalling, cis-inhibition and endocytosis. EMBO J 25: 4697–4706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gray AR. (1993) Broccoli Brassica oleracea L. (Italica group). In Kalloo G, Bergh BO, eds, Genetic Improvement of Vegetable Crops. Pergamon Press, New York, pp 61–86 [Google Scholar]
- Haklai-Topper L, Mlechkovich G, Savariego D, Gokhman I, Yaron A (2010) cis-Interaction between Semaphorin6A and Plexin-A4 modulates the repulsive response to Sema6A. EMBO J 29: 2635–2645 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han X. (2014) Constitutively active chemokine CXC receptors. Adv Pharmacol 70: 265–301 [DOI] [PubMed] [Google Scholar]
- Held W, Mariuzza RA (2008) cis-Interactions of immunoreceptors with MHC and non-MHC ligands. Nat Rev Immunol 8: 269–278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirota S, Isozaki K, Moriyama Y, Hashimoto K, Nishida T, Ishiguro S, Kawano K, Hanada M, Kurata A, Takeda M, et al. (1998) Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 279: 577–580 [DOI] [PubMed] [Google Scholar]
- Jackson AL, Bartz SR, Schelter J, Kobayashi SV, Burchard J, Mao M, Li B, Cavet G, Linsley PS (2003) Expression profiling reveals off-target gene regulation by RNAi. Nat Biotechnol 21: 635–637 [DOI] [PubMed] [Google Scholar]
- Jun J, Fiume E, Roeder AH, Meng L, Sharma VK, Osmont KS, Baker C, Ha CM, Meyerowitz EM, Feldman LJ, et al. (2010) Comprehensive analysis of CLE polypeptide signaling gene expression and overexpression activity in Arabidopsis. Plant Physiol 154: 1721–1736 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kachroo A, Schopfer CR, Nasrallah ME, Nasrallah JB (2001) Allele-specific receptor-ligand interactions in Brassica self-incompatibility. Science 293: 1824–1826 [DOI] [PubMed] [Google Scholar]
- Kho YO, Baër J (1968) Observing pollen tubes by means of fluorescence. Euphytica 17: 298–302 [Google Scholar]
- Kitashiba H, Liu P, Nishio T, Nasrallah JB, Nasrallah ME (2011) Functional test of Brassica self-incompatibility modifiers in Arabidopsis thaliana. Proc Natl Acad Sci USA 108: 18173–18178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein T, Brennan K, Arias AM (1997) An intrinsic dominant negative activity of serrate that is modulated during wing development in Drosophila. Dev Biol 189: 123–134 [DOI] [PubMed] [Google Scholar]
- Krasagakis K, Fragiadaki I, Metaxari M, Krüger-Krasagakis S, Tzanakakis GN, Stathopoulos EN, Eberle J, Tavernarakis N, Tosca AD (2011) KIT receptor activation by autocrine and paracrine stem cell factor stimulates growth of merkel cell carcinoma in vitro. J Cell Physiol 226: 1099–1109 [DOI] [PubMed] [Google Scholar]
- Kusaba M, Dwyer K, Hendershot J, Vrebalov J, Nasrallah JB, Nasrallah ME (2001) Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 13: 627–643 [PMC free article] [PubMed] [Google Scholar]
- Laemmli UK. (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685 [DOI] [PubMed] [Google Scholar]
- Lease KA, Walker JC (2006) The Arabidopsis unannotated secreted peptide database, a resource for plant peptidomics. Plant Physiol 142: 831–838 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasrallah JB, Nasrallah ME (2014a) S-locus receptor kinase signalling. Biochem Soc Trans 42: 313–319 [DOI] [PubMed] [Google Scholar]
- Nasrallah JB, Nasrallah ME (2014b) Robust self-incompatibility in the absence of a functional ARC1 gene in Arabidopsis thaliana. Plant Cell 26: 3838–3841 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasrallah ME. (2004) Biotechnology for pollination control in crucifer oilseed crops. In EC Pua, CJ Douglas, eds, Biotechnology in Agriculture and Forestry, Vol 54. Springer-Verlag, Heidelberg, Germany, pp 65–80 [Google Scholar]
- Nasrallah ME, Liu P, Nasrallah JB (2002) Generation of self-incompatible Arabidopsis thaliana by transfer of two S locus genes from A. lyrata. Science 297: 247–249 [DOI] [PubMed] [Google Scholar]
- Nasrallah ME, Liu P, Sherman-Broyles S, Boggs NA, Nasrallah JB (2004) Natural variation in expression of self-incompatibility in Arabidopsis thaliana: implications for the evolution of selfing. Proc Natl Acad Sci USA 101: 16070–16074 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson BK, Cai X, Nebenführ A (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51: 1126–1136 [DOI] [PubMed] [Google Scholar]
- Ockendon DJ, Smith BM (1993) Brussels sprouts Brassica oleracea var. gemmifera DC. In Kalloo G, Bergh BO, eds. Genetic Improvement of Vegetable Crops. Pergamon Press, New York, pp 87–110 [Google Scholar]
- Paszkowski J, Grossniklaus U (2011) Selected aspects of transgenerational epigenetic inheritance and resetting in plants. Curr Opin Plant Biol 14: 195–203 [DOI] [PubMed] [Google Scholar]
- Rea AC, Nasrallah JB (2015) In vivo imaging of the S-locus receptor kinase, the female specificity determinant of self-incompatibility, in transgenic self-incompatible Arabidopsis thaliana. Ann Bot (Lond) 115: 789–805 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robatzek S, Chinchilla D, Boller T (2006) Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev 20: 537–542 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts M, Tang S, Stallmann A, Dangl JL, Bonardi V (2013) Genetic requirements for signaling from an autoactive plant NB-LRR intracellular innate immune receptor. PLoS Genet 9: e1003465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakamoto K, Bu G, Chen S, Takei Y, Hibi K, Kodera Y, McCormick LM, Nakao A, Noda M, Muramatsu T, et al. (2011) Premature ligand-receptor interaction during biosynthesis limits the production of growth factor midkine and its receptor LDL receptor-related protein 1. J Biol Chem 286: 8405–8413 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scacheri PC, Rozenblatt-Rosen O, Caplen NJ, Wolfsberg TG, Umayam L, Lee JC, Hughes CM, Shanmugam KS, Bhattacharjee A, Meyerson M, et al. (2004) Short interfering RNAs can induce unexpected and divergent changes in the levels of untargeted proteins in mammalian cells. Proc Natl Acad Sci USA 101: 1892–1897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schopfer CR, Nasrallah JB (2000) Self-incompatibility. Prospects for a novel putative peptide-signaling molecule. Plant Physiol 124: 935–940 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schopfer CR, Nasrallah ME, Nasrallah JB (1999) The male determinant of self-incompatibility in Brassica. Science 286: 1697–1700 [DOI] [PubMed] [Google Scholar]
- Shaya O, Sprinzak D (2011) From Notch signaling to fine-grained patterning: Modeling meets experiments. Curr Opin Genet Dev 21: 732–739 [DOI] [PubMed] [Google Scholar]
- Shiba H, Takayama S, Iwano M, Shimosato H, Funato M, Nakagawa T, Che FS, Suzuki G, Watanabe M, Hinata K, et al. (2001) A pollen coat protein, SP11/SCR, determines the pollen S-specificity in the self-incompatibility of Brassica species. Plant Physiol 125: 2095–2103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimosato H, Yokota N, Shiba H, Iwano M, Entani T, Che FS, Watanabe M, Isogai A, Takayama S (2007) Characterization of the SP11/SCR high-affinity binding site involved in self/nonself recognition in Brassica self-incompatibility. Plant Cell 19: 107–117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smyth DR, Bowman JL, Meyerowitz EM (1990) Early flower development in Arabidopsis. Plant Cell 2: 755–767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sorkin A, Goh LK (2008) Endocytosis and intracellular trafficking of ErbBs. Exp Cell Res 314: 3093–3106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sporn MB, Roberts AB (1985) Autocrine growth factors and cancer. Nature 313: 745–747 [DOI] [PubMed] [Google Scholar]
- Stein JC, Dixit R, Nasrallah ME, Nasrallah JB (1996) SRK, the stigma-specific S locus receptor kinase of Brassica, is targeted to the plasma membrane in transgenic tobacco. Plant Cell 8: 429–445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stein JC, Howlett B, Boyes DC, Nasrallah ME, Nasrallah JB (1991) Molecular cloning of a putative receptor protein kinase gene encoded at the self-incompatibility locus of Brassica oleracea. Proc Natl Acad Sci USA 88: 8816–8820 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strabala TJ. (2008) CLE genes in plant development: gain-of-function analyses, pleiotropy, hypermorphy and neomorphy. Plant Signal Behav 3: 457–459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strabala TJ, O’donnell PJ, Smit AM, Ampomah-Dwamena C, Martin EJ, Netzler N, Nieuwenhuizen NJ, Quinn BD, Foote HC, Hudson KR (2006) Gain-of-function phenotypes of many CLAVATA3/ESR genes, including four new family members, correlate with tandem variations in the conserved CLAVATA3/ESR domain. Plant Physiol 140: 1331–1344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takayama S, Shiba H, Iwano M, Shimosato H, Che FS, Kai N, Watanabe M, Suzuki G, Hinata K, Isogai A (2000) The pollen determinant of self-incompatibility in Brassica campestris. Proc Natl Acad Sci USA 97: 1920–1925 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takayama S, Shimosato H, Shiba H, Funato M, Che FS, Watanabe M, Iwano M, Isogai A (2001) Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature 413: 534–538 [DOI] [PubMed] [Google Scholar]
- Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA 76: 4350–4354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang JQ, Jeelall YS, Ferguson LL, Horikawa K (2014) Toll-like receptors and cancer: MYD88 mutation and inflammation. Front Immunol 5: 367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang ZY, Bai MY, Oh E, Zhu JY (2012) Brassinosteroid signaling network and regulation of photomorphogenesis. Annu Rev Genet 46: 701–724 [DOI] [PubMed] [Google Scholar]
- Webster MK, Donoghue DJ (1996) Constitutive activation of fibroblast growth factor receptor 3 by the transmembrane domain point mutation found in achondroplasia. EMBO J 15: 520–527 [PMC free article] [PubMed] [Google Scholar]
- Willnow TE, Armstrong SA, Hammer RE, Herz J (1995) Functional expression of low-density lipoprotein receptor-related protein is controlled by receptor-associated protein in vivo. Proc Natl Acad Sci USA 92: 4537–4541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willnow TE, Rohlmann A, Horton J, Otani H, Braun JR, Hammer RE, Herz J (1996) RAP, a specialized chaperone, prevents ligand-induced ER retention and degradation of LDL receptor-related endocytic receptors. EMBO J 15: 2632–2639 [PMC free article] [PubMed] [Google Scholar]
- Yamamoto M, Tantikanjana T, Nishio T, Nasrallah ME, Nasrallah JB (2014) Site-specific N-glycosylation of the S-locus receptor kinase and its role in the self-incompatibility response of the Brassicaceae. Plant Cell 26: 4749–4762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yaron A, Sprinzak D (2012) The cis side of juxtacrine signaling: a new role in the development of the nervous system. Trends Neurosci 35: 230–239 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.