Abstract
A phenomenon of yield decline due to weak plant growth in strawberry was recently observed in non-chemo-fumigated soils, which was not associated with the soil fungal pathogen Verticillium dahliae, the main target of fumigation. Amplicon-based metagenomics was used to profile soil microbiota in order to identify microbial organisms that may have caused the yield decline. A total of 36 soil samples were obtained in 2013 and 2014 from four sites for metagenomic studies; two of the four sites had a yield-decline problem, the other two did not. More than 2000 fungal or bacterial operational taxonomy units (OTUs) were found in these samples. Relative abundance of individual OTUs was statistically compared for differences between samples from sites with or without yield decline. A total of 721 individual comparisons were statistically significant – involving 366 unique bacterial and 44 unique fungal OTUs. Based on further selection criteria, we focused on 34 bacterial and 17 fungal OTUs and found that yield decline resulted probably from one or more of the following four factors: (1) low abundance of Bacillus and Pseudomonas populations, which are well known for their ability of supressing pathogen development and/or promoting plant growth; (2) lack of the nematophagous fungus (Paecilomyces species); (3) a high level of two non-specific fungal root rot pathogens; and (4) wet soil conditions. This study demonstrated the usefulness of an amplicon-based metagenomics approach to profile soil microbiota and to detect differential abundance in microbes.
Introduction
Verticillium dahliae Kleb. is a soil-borne fungal pathogen, which penetrates the roots of a wide range of host plant species causing the disease Verticillium wilt1. The pathogen colonises the vascular system of the roots and crown depriving the leaves and stems of water. Also, it produces microsclerotia in the host as the infected tissues senesce, which are released into the soil as the plant decays and are the primary inoculum of V. dahliae in the soil for subsequent infection. Microsclerotia may survive for more than 10 years in the soil in the absence of its hosts1. Wilt incidence tends to be higher in soils infested with the root lesion nematode, Pratylenchus penetrans, which feed on the roots causing wounds, thus increasing entry sites for the pathogen and affecting fungal infection/colonisation of vascular tissue2,3, although the magnitude of this interaction may vary greatly with individual fungal strains4.
Chemical treatments, such as methyl bromide and chloropicrin, have been an indispensable tool for the past 40 years because of their excellent efficacy, effectively managing Verticillium wilt in strawberry; however, several of these treatments are already banned (e.g. methyl bromide) or face an uncertain future due to legislation (e.g. chloropicrin)5. With the loss of methyl bromide and other fumigants, strawberry (Fragaria × annanasa) production has come under increasing threat of losses due to wilt caused by V. dahliae. Some alternative measures have control efficacy similar to that achieved by commercial chemical fumigants, but others are not as good6. One of the alternatives being investigated is biofumigation, which uses decay products of green manures7–10. Control of V. dahliae through the use of Brassica species plants is believed to result from the toxic isothiocyanates (ITCs), released into the soil after incorporation of glucosinolate-containing plant tissues11. A recent study suggested that biofumigation based on the Brassica species cannot fully control wilt because of the limited amount of the ITC released9. Other plant species have also been studied for their biofumigation effects against V. dahliae. For example, biofumigation using Lavandula plant materials can result in large reductions in the numbers of viable microsclerotia recovered12. Recently, anaerobic soil disinfestation (ASD) methods have been studied for their effects against a range of soil pests and pathogens. Traditional ASD with grass was less effective than with organic materials; all materials proved to be effective at 16 °C against P. penetrans, Meloidogyne hapla, Globodera pallida and V. dahliae with V. dahliae being most difficult to control13. Control efficacy depends on many factors, including soil characteristics, types of organic material, temperature, dosage and exposure time. Soil organic amendments, especially combined with biocontrol agents, can satisfactorily suppress Verticillium wilt14, but the effect is often inconsistent and pathosystem specific, revealed by a meta-study15. Rotation with other crops, e.g. Brassica and lettuce, can reduce wilt on strawberry16,17, but this management strategy is generally not commercially viable.
Specific microbial organisms have been tested against soil-borne pathogens, including V. dahliae. Application of two biocontrol organisms (Paenibacillus alvei K165 and the nonpathogenic Fusarium oxysporum F2) at the transplant stage reduced Verticillium wilt symptom development in aubergine18. Both organisms induced the expression of the pathogenesis-related (PR) proteins PR1 and PR4 in the stem of aubergines. Many fungi and bacteria were isolated from rhizosphere of oilseed rape and strawberry and tested against V. dahliae19,20. Many bacterial (primarily Pseudomonas and Serratia spp.) and fungal species were found to be antagonistic against wilt. There was also some evidence to suggest that some fungi were specifically enriched in each rhizosphere, which is supported by a recent finding that rhizosphere communities are partially genetically controlled by hosts21. Dipping plants in a suspension of Serratia plymuthica prior to planting reduced Verticillium wilt of strawberry and increased yield22. Plant growth-promoting rhizobacteria (PGPR) have recently received much attention for their use to increase crop production, including their role in suppressing disease development23. Non-pathogenic strains of V. dahliae can also be used as a biological control agent to exploit the concept that preoccupation of the ecological niche rendered strawberry plants immune to infection with pathogenic V. dahliae. Inoculation of non-pathogenic strains of V. dahliae reduced wilt on 20% of treated plants but led to increased wilt development on 50% of treated plants24.
The monoterpenoids associated with the Lavandula spp., which are of lower volatility than the ITCs associated with brassica decomposition, were detected for more than one week after materials were incorporated in soil12. Three of these terpenoids were shown to reduce microsclerotium viability in microcosm tests. Recently, we have conducted field experiments to compare a new product based on microencapsulation of the three terpenes (cineole, camphor and borneol) and several other alternatives for their efficacy against wilt on strawberry with the commercial chemofumigant (chloropicrin) as a standard. Results from the trials showed that overall the terpene-based product reduced wilt development but did not increase fruit yield, compared to the un-treated control. However, at two of the eight trial sites, the chloropicrin treatment led to nearly 25% increase in yield, compared to all other treatments, but the level of wilt was similar among all treatments. Chloropicrin-treated plants had increased growth vigour compared to all other treatments; all other treatments had more or less uniformly stunted growth, which cannot be associated with any obvious biotic and abiotic factors (although strawberry roots were not sampled for assessment). We speculated that this yield decline resulted from interactions among a number of microbial organisms, similar to apple replant disease25. Involvement of other beneficial or pathogenic microbial organisms in strawberry was implicated in several other experiments on controlling Verticillium wilt13,22.
This paper reports the results from studies aiming to identify candidate microorganisms that are responsible for observed yield decline in strawberry of non-chloropicrin-treated plants. This study reports primarily data-driven research, aiming to generate hypotheses on the possible candidate organisms responsible for the observed yield decline, which can then be further tested in future. Specifically, we used an amplicon-based metagenomics approach to profile soils from different treatments at four sites – two with and two without yield-decline phenomenon. Through statistical comparison of individual microbial operational taxonomy units (OTUs), combined with several objective selection criteria, several candidate organisms were identified as candidates that may have played a role in the observed yield decline.
Materials and methods
Field sites and sample collection
Evaluation of the efficacy of a new microencapsulated terpene-based product and other alternatives to control Verticillium wilt was conducted at eight sites: three in 2011, three in 2012 and two in 2013. At two sites in 2012, a yield-decline phenomenon in strawberry was observed. Based on the physical distances between the trial sites, four sites (two in 2012: PV12 and HB12 and two in 2013: HB13 and EM13) were selected for microbial profiling to identify candidate microorganisms that may be involved in the yield-decline phenomenon. HB12 and HB13 were two fields at the same farm (about 1 km apart), about 15 km and 25 km from the PV12 and EM13 sites, respectively; PV12 was about 40 km from the EM13 site. At each site there were three blocks; within each block, there was one plot per treatment. The plot size was 15 m long (a single bed of double rows) with four metres between neighbouring plots. Immediately after soil treatment, all beds were covered with black polythene; beds were planted three weeks after treatment. All plots were automatically irrigated through drip tape.
Soil samples from the four sites were obtained in 2013 or 2014; details of samples are given in Table 1. Although there were six (2012) and eight (2013) treatments, only selected treatments were sampled for microbial profiling. For the two yield-decline sites, nearly two years had passed since the treatment when soil samples were taken in 2013. Thus, the chloropicrin effect in preventing yield decline may have significantly reduced – this had to be taken into account when trying to identify causal agents responsible for yield decline at these two sites. Although for the two 2012 sites, soil samples were taken long after the trial cropping was finished, the land was not used for other purposes and hence the plot (bed) structure was still intact at the time of sampling. For each plot, a composite soil sample was obtained, consisting of 10 core soils that were obtained with a sampler (2.5 cm in diameter) from a depth of 20 cm at randomly selected locations and then mixed by sieving (mesh size 2 mm). A subsample (approximately 2 g) of each composite sample was collected in a 2 ml Eppendorf tube and stored at −80 °C until DNA extraction.
Table 1. Summary of soils sampled for metagenomic profiling; there were three composite samples (one from each replicate plot) for each treatment at each site, giving a total of 36 samples.
| Treatments sampled |
Last harvest time | |||||
|---|---|---|---|---|---|---|
| Site | Yield decline? | Cultivar | Treatment applied | October 2013 | May 2014 | |
| EM13 | No | Elsanta | May 2013 | Not sampled | Chloropicrin, untreated | 14 July |
| HB13 | ||||||
| HB12 | Yes | Elsanta | May 2012 | Not sampled | Chloropicrin, untreated | 13 July |
| PV12 | Eves Delight | November 2011 | BFa, chloropicrin, LWa, untreated | 12 October | ||
Coded products – the exact identity is commercially confidential.
DNA extraction and next generation sequencing
Total genomic DNA was isolated in triplicate from each soil sample (0.25 g) using the PowerSoil DNA Isolation Kit (MoBio Laboratories) with minor modifications as described below. Before bead-beating, the samples were incubated in lysis solution at 65 °C for 10 min. Samples were homogenised by two 20-sec cycles at power setting of 5.0 in FastPrep instrument (FP120, Bio 101, Thermo Savant, Qbiogene), with 5 min on ice in between cycles. The DNA was further extracted according to the kit protocol. Triplicate samples were pooled after extraction and purified using GeneClean Turbo Kit (MP Biomedicals) using GNomic salt solution as protocol. After preliminary trial sequencing runs, two primer pairs were selected: one for bacteria (27F/534R 16S rDNA) and one for fungi (ITSI-F/Ek28-R 18S ITS). The two primer sets were modified at the 5′ end with adaptors, TCG TCG GCA GCG TCA GAT GTG TAT AAG AGA CAG – forward adaptor and GTC TCG TGG GCT CGG AGA TGT GTA TAA GAG ACA – reverse adaptor. The ITSI-F/Ek28-R primer set was chosen as it gave better amplification when combined with the barcode attachments used in the Illumina sequencing. PCR amplification using these primers gave a product of ∼750 bp, which was consistent with the target region of the rRNA genes including the end of the SSU, ITS1, 5.8S and the start of the LSU region plus the adaptor primers. All PCR reactions were carried out in triplicate 13.0 µl reactions with ×1 buffer basic (Molezym GmbH and Co. Bremen Germany), 2 mM MgCl2 (Qiagen, Hielden, Germany), 0.2 mM dNTP (Invitrogen, Life Technologies, USA), 0.25 U Mol Taq basic DNA polymerase (Molezym GmbH and Co. Bremen Germany), 0.2 µM forward and reverse primers each (Integrated DNA Technologies) and about 2 ng template DNA, was made up to 13 µl with molecular biology reagent water (Sigma, UK). Each reaction was performed in a Dyad thermocyler (MJ research), according to the following protocol, thermal cycling consisted of initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 55 °C for 45 s, and elongation at 72 °C for 60 s, reducing 0.5 °C per cycle until 50 °C, with a final extension at 72 °C for 5 min. Negative control samples were treated similarly with the exclusion of template DNA. PCR products were visualised by agarose gel electrophoresis.
Following PCR, DNA amplicons were purified using Agencourt AMPure XP beads (Beckman Coulter, USA), as per manufacturer’s instructions. The adapted amplicons were then modified by attaching indices and Illumina sequencing adapters using the Nextera XT Index Kit by PCR as described in the manufacturer’s protocol, enabling simultaneous sequencing of multiple samples, i.e. multiplexing. Following the index PCR clean-up step, using the Agencourt AMPure XP beads, as per manufacturer’s instructions, PCR products were qualitatively assessed using a Fragment Analyzer (Advanced Analytical, Ames, IA, USA) with the High Sensitivity NGS Fragment Analysis Kit (Advanced Analytical, Ames, IA, USA). PCR products were also quantitatively assessed using a Qubit 2.0 Fluorometer (Life Technologies, USA).
DNA from the different samples was then pooled so as to be analysed on the same Illumina run to avoid run-quality bias. The unique DNA barcode indices allowed sequences from all samples to be de-multiplexed in subsequent processing. Samples were pooled in such a way to ensure each of them was equimolar. The final concentration of the pooled library was 4 nM. The amplicon library was denatured using 1 mM NaOH and diluted to 30 pM as per manufacturer’s protocol. The diluted and denatured amplicon library was then combined with a denatured PhiX library at an equimolar concentration at a rate of 20% to increase heterogeneity of the sample. These samples were then run on an Illumina MiSeq with 300 bp paired end sequencing (version 3 chemistry). Samples from 2013 were sequenced separately from the 2014 samples; within each sequence run, there were 64 samples (which included samples from other studies).
Sequence processing
Raw sequences were automatically de-multiplexed by the Illumina MiSeq and then further quality-filtered by the QIIME analysis pipeline:26 (1) removing primers from sequences, (2) removing low-quality reads, (3) identifying an OTU for each sequence against two international databases: 16S (bacteria) – Silva27 and the UNITE fungal 18S ITS database28 at 97% similarity, (4) storing every unique sequence and its frequency in each sample. Finally, we wrote a small utility programme in Delphi to (1) produce summary for OTU frequency for each sample, (2) merge the OTU frequency data over all samples and (3) produce an overall OTU table in the BIOM format, enabling further downstream statistical analysis.
We also customised the UNITE fungal ITS database to include ITS sequences for oomycetes and vascular wilts; all oomycete ITS sequences were first obtained from international repositories – if multiple sequences were available for a single species then a consensus sequence was generated with Geneious version 6.1 (Biomatters Ltd). Custom Perl scripts were written to query the Index Fungorum database (http://www.indexfungorum.org/), using the Index Fungorum Fungus API to query the website to return taxonomic information on the additional species, absent from the UNITE database. Both the consensus sequences and the linked taxonomic information were then appended to the UNITE database files. Full scripts are available for download from (https://github.com/eastmallingresearch/metagenomics).
Statistical analysis
Two types of statistical analyses were performed: (1) initial exploratory analysis and (2) detecting differential OTU abundance between samples from different sites to identify candidates responsible for the yield-decline phenomenon.
Individual sample diversity (i.e., α diversity) was calculated: the number of distinct OTUs observed per sample (Sobs) and Shannon and Simpson indices (both related to the frequency of individual OTUs within a sample). To reduce biases in sequencing depth on these indices, a re-sampling (i.e. bootstrap) scheme was used to estimate α diversity indices for each sample. A bootstrap sample was obtained via randomly sampling a minimum number of sequences from the sequences in each sample (i.e., rarefying). All indices are calculated from each bootstrap sample at the rarefaction point – a total of 25 bootstraps were conducted. Mean indices were calculated from these bootstrap samples. Next, we calculated diversity indices among samples (i.e. β diversity): Morrison-Horn (MH), Bray-Curtis (BC) and ThetaYC (YC) indices. MH measures the similarity between two samples whereas the other two methods measure the dissimilarity between two samples. Both α and β diversity indices were calculated using the Win64 version of the Explicet software29. A principal component analysis was conducted to detect overall differences between samples using the STAMP programme30.
To assess differential OTU abundance among treatments, we used the DESeq2 statistical package in R31. DESeq2 was developed for comparing differential gene expressions but is equally applicable to analysis of metagenomic data. DESeq2 provides a new statistical fitting routine to account for variance heterogeneity often observed in sequence data; it uses the negative binomial distribution as an error distribution to compare abundance of each OTU between groups of samples in the framework of generalised linear modelling. This method was superior to other methods commonly used for this purpose32; the same study also showed that rarefying samples is inferior to not rarefying in identifying differences in OTU abundances but with correct distribution assumptions. Thus, in the present study, rarefying was not used when comparing OTU abundance. The median-of-ratios method33 was used to normalise the data to correct for unequal sequencing depth; this procedure was implemented as a default in DESeq2. To correct false discovery rate associated with multiple testing, DESeq2 uses the Benjamini-Hochberg (BH) adjustment34. In addition, DESeq2 also implemented an algorithm to adjust for large variability in log-fold changes for small counts. Candidate OTUs were selected at the 5% significance level (BH adjusted). Any OTU with the total number of reads across all the samples less than three was omitted from differential abundance testing.
In comparing OTU abundance, seven different comparisons were made, taking into account the potentially complex nature of candidate organisms, and the persistency of chloropicrin effect in relation to sampling time. The first five comparisons were between sites with and without the yield-decline phenomenon; only samples collected in 2014 were used for these five comparisons since they were sampled at the same time. In these comparisons, chloropicrin-treated soil samples from HB12 and PV12 were excluded because it is not certain whether the chloropicrin effect could last for more than two years. These five comparisons were:
EM13/HB13 (12 samples) vs. HB12/PV12 (six samples);
EM13 (six samples) vs. HB12 (three samples);
EM13 (six samples) vs. PV12 (three samples);
HB13 (six samples) vs. HB12 (three samples);
HB13 (six samples) vs. PV12 (six samples).
The four individual comparisons (b–e) are necessary since it is possible that yield decline may be caused by more than one organism; thus an overall comparison, as in (1), may not be able to reveal all possible candidates because of potential site-to-site variability in the abundance of candidate organisms. The final two comparisons were:
2013 chloropicrin (three samples) vs. other treatments (nine samples) at PV12;
2014 chloropicrin (three samples) vs. the untreated (three samples) at HB12.
For comparison (f), only 2013 samples were used because the sampling time elapsed was close to two years after the application of treatments at PV12, similar to the time elapsed for the 2014 samples at HB12.
Results
Sequencing quality
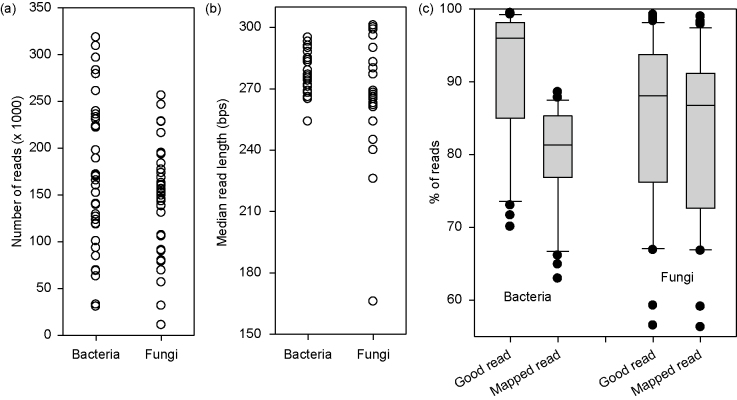
Samples were sequenced in two runs: in February (2013 samples) and July (2014 samples), 2014. Most samples had more than 50 000 high-quality sequences (Figure 1). The median read length of the first read (P1), after quality trimming in the QIIME pipeline, was greater than 250 bps for all examples except four samples for the fungal sequencing (Figure 1). About 91% and 85% of total reads were of good quality for bacteria and fungi, respectively.
Figure 1.
General information of sequencing data: (a) number of reads per sample, (b) median read length per sample and (c) % of good quality reads and % of reads mapped to OTUs.
The majority of these high-quality reads are mapped to OTUs in the two international databases. For bacteria, on average 88% of reads were mapped to OTUs (ranging from 83% to 94% for individual samples). For fungi, the percentage of good quality reads mapped to OTUs ranged from 85% to 100% with mean of 96%. The total number of distinct bacterial and fungal OTUs was 2142 and 2022, respectively. Of these OTUs, there were respective 306 and 326 cases where there was only a single read across all samples, i.e. only present in one sample.
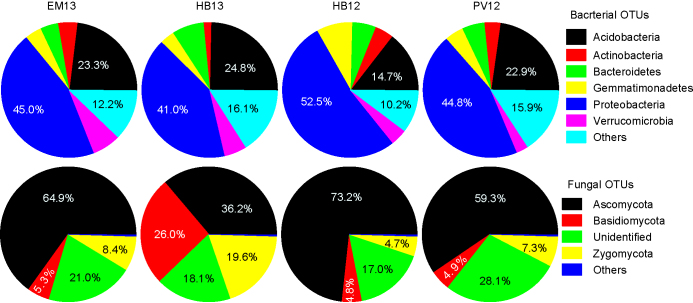
The two most common bacterial phyla are Proteobacteria and Acidobacteria, accounting for ca. 67% of the total mapped reads (Figure 2). For fungi, Ascomycota was the most common phylum, accounting for more than 50% of the mapped reads on average, and the next two common phyla were Zygomycota and Basidiomycota (Figure 2).
Figure 2.
Proportion of major bacterial and fungal phylum found in the three untreated soil samples at each of the four sites; soils were sampled in May 2014.
The relationship between number of OTUs observed (Sobs) and the sequencing depth is shown in Figure 3 for a few samples. Overall, the present sequencing depth appeared to be sufficient since the minimum sampling depth in our samples was not in the part of curve with steep increases – indicating a diminishing return of further sequencing in uncovering new OTUs.
Figure 3.
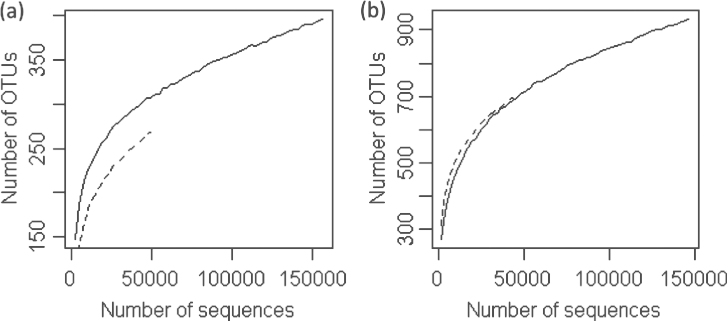
Four examples of rarefaction curves: two for fungi (a) and two for bacteria (b). All curves terminated at the number of sequences obtained in the examples.
α diversity
Within sample diversity measures varied greatly with samples (Table 2). Overall, there were more bacterial OTUs in individual samples than fungal OTUs (Table 3). At the rarefaction point (35 000 and 46 000 for bacteria and fungi, respectively), the number of bacterial OTUs ranged from 506 to 764 per sample, compared to the corresponding fungal value of 123–367. The α diversity of fungal OTUs varied more from sample to sample than bacterial OTUs. For instance, the Simpson index ranged from 0.054 to 0.50, compared to the corresponding value of 0.013–0.037 for bacteria. The effect of chloropicrin on the number of OTUs was inconsistent between sites. Only at the EMR site, chloropicrin appeared to consistently reduce the number of OTUs, compared to the control treatment (Table 2).
Table 2. Summary of α (within-group) diversity measures calculated at the rarefaction point (35 000 and 46 000 for the bacteria and fungi, respectively): number of OTUs (Sobs), Shannon H index (SH), Shannon E index (SE), Simpson Index (SI).
| Bacteria |
Fungi |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Year | Site | Treatment | Sobs | SH | SE | SI | Sobs | SH | SE | SI |
| 2013 | PV12 | BF | 587 | 6.570 | 0.714 | 0.033 | 266 | 3.620 | 0.449 | 0.228 |
| 2013 | PV12 | BF | 602 | 6.780 | 0.734 | 0.026 | 236 | 4.012 | 0.509 | 0.157 |
| 2013 | PV12 | BF | 599 | 6.697 | 0.725 | 0.024 | 231 | 3.464 | 0.442 | 0.207 |
| 2013 | PV12 | Untreated | 586 | 6.562 | 0.714 | 0.033 | 287 | 4.481 | 0.549 | 0.105 |
| 2013 | PV12 | Untreated | 551 | 6.273 | 0.689 | 0.037 | 269 | 4.653 | 0.576 | 0.088 |
| 2013 | PV12 | Untreated | 620 | 6.756 | 0.729 | 0.026 | 249 | 3.373 | 0.424 | 0.278 |
| 2013 | PV12 | Chloropicrin | 600 | 6.715 | 0.728 | 0.025 | 180 | 2.705 | 0.361 | 0.342 |
| 2013 | PV12 | Chloropicrin | 583 | 6.690 | 0.728 | 0.024 | 203 | 3.724 | 0.485 | 0.168 |
| 2013 | PV12 | Chloropicrin* | 237 | 3.840 | 0.487 | 0.202 | ||||
| 2013 | PV12 | LW | 635 | 6.898 | 0.740 | 0.023 | 205 | 4.581 | 0.596 | 0.080 |
| 2013 | PV12 | LW | 657 | 7.093 | 0.758 | 0.020 | 226 | 4.200 | 0.537 | 0.115 |
| 2013 | PV12 | LW* | ||||||||
| 2014 | EM13 | Untreated | 629 | 6.679 | 0.718 | 0.024 | 284 | 4.924 | 0.604 | 0.062 |
| 2014 | EM13 | Untreated | 654 | 6.800 | 0.727 | 0.023 | 349 | 4.959 | 0.587 | 0.070 |
| 2014 | EM13 | Untreated | 641 | 6.704 | 0.719 | 0.026 | 246 | 5.079 | 0.640 | 0.054 |
| 2014 | EM13 | Chloropicrin | 506 | 6.362 | 0.709 | 0.028 | 169 | 2.236 | 0.303 | 0.500 |
| 2014 | EM13 | Chloropicrin | 545 | 6.358 | 0.699 | 0.025 | 154 | 2.026 | 0.279 | 0.491 |
| 2014 | EM13 | Chloropicrin | 572 | 6.569 | 0.717 | 0.020 | 181 | 3.028 | 0.403 | 0.229 |
| 2014 | HB12 | Untreated | 664 | 7.006 | 0.747 | 0.016 | 326 | 4.951 | 0.593 | 0.073 |
| 2014 | HB12 | Untreated | 672 | 6.913 | 0.736 | 0.018 | 323 | 4.559 | 0.547 | 0.100 |
| 2014 | HB12 | Untreated | 632 | 6.831 | 0.734 | 0.019 | 350 | 4.956 | 0.587 | 0.078 |
| 2014 | HB12 | Chloropicrin | 639 | 6.900 | 0.741 | 0.017 | 208 | 3.893 | 0.505 | 0.186 |
| 2014 | HB12 | Chloropicrin | 649 | 7.044 | 0.754 | 0.014 | 246 | 4.243 | 0.535 | 0.128 |
| 2014 | HB12 | Chloropicrin | 576 | 7.101 | 0.775 | 0.013 | 304 | 5.026 | 0.610 | 0.076 |
| 2014 | HB13 | Untreated | 666 | 6.774 | 0.722 | 0.024 | 367 | 5.064 | 0.594 | 0.075 |
| 2014 | HB13 | Untreated | 576 | 6.804 | 0.743 | 0.022 | 289 | 4.763 | 0.583 | 0.093 |
| 2014 | HB13 | Untreated* | 656 | 6.761 | 0.723 | 0.025 | ||||
| 2014 | HB13 | Chloropicrin | 613 | 6.634 | 0.716 | 0.024 | 159 | 3.595 | 0.492 | 0.173 |
| 2014 | HB13 | Chloropicrin | 639 | 6.711 | 0.720 | 0.023 | 284 | 4.098 | 0.503 | 0.150 |
| 2014 | HB13 | Chloropicrin | 645 | 6.705 | 0.718 | 0.024 | 204 | 4.340 | 0.566 | 0.100 |
| 2014 | PV12 | Untreated | 664 | 6.981 | 0.745 | 0.024 | 261 | 3.396 | 0.423 | 0.208 |
| 2014 | PV12 | Untreated | 651 | 6.608 | 0.706 | 0.026 | 347 | 5.238 | 0.621 | 0.060 |
| 2014 | PV12 | Untreated | 654 | 6.767 | 0.724 | 0.027 | 202 | 5.024 | 0.656 | 0.054 |
| 2014 | PV12 | Chloropicrin | 713 | 7.091 | 0.748 | 0.016 | 303 | 4.794 | 0.582 | 0.081 |
| 2014 | PV12 | Chloropicrin | 735 | 6.861 | 0.721 | 0.025 | 349 | 5.032 | 0.596 | 0.072 |
| 2014 | PV12 | Chloropicrin | 764 | 7.197 | 0.751 | 0.017 | 339 | 5.309 | 0.631 | 0.059 |
Table 3. Estimates of three β diversity (between-group) measures for bacterial and fungal OTUs, describing the extent of similarity or dissimilarity among combinations of years and sites.
| Bacteria |
Fungi |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| EM13 | HB13 | HB12 | 2013 PV12 | 2014 PV12 | EM13 | HB13 | HB12 | 2013 PV12 | 2014 PV12 | |
| Morrista-Horn (similarity) | ||||||||||
| EM13 | 1 | 0.856 | 0.934 | 0.77 | 0.826 | 1 | 0.74 | 0.756 | 0.361 | 0.486 |
| HB13 | 1 | 0.834 | 0.886 | 0.89 | 1 | 0.806 | 0.71 | 0.633 | ||
| HB12 | 1 | 0.745 | 0.802 | 1 | 0.573 | 0.738 | ||||
| 2013 PV12 | 1 | 0.97 | 1 | 0.695 | ||||||
| 2014 PV12 | 1 | 1 | ||||||||
| ThetaYC (dissimilarity) | ||||||||||
| EM13 | 0.000 | 0.251 | 0.123 | 0.374 | 0.297 | 0.000 | 0.413 | 0.392 | 0.78 | 0.679 |
| HB13 | 0 | 0.285 | 0.205 | 0.198 | 0 | 0.325 | 0.45 | 0.537 | ||
| HB12 | 0 | 0.407 | 0.33 | 0 | 0.598 | 0.415 | ||||
| 2013 PV12 | 0 | 0.058 | 0 | 0.467 | ||||||
| 2014 PV12 | 0 | 0 | ||||||||
| Bray-Curtis (dissimilarity) | ||||||||||
| EM13 | 0 | 0.279 | 0.316 | 0.412 | 0.586 | 0 | 0.484 | 0.462 | 0.649 | 0.505 |
| HB13 | 0 | 0.358 | 0.302 | 0.564 | 0 | 0.439 | 0.492 | 0.609 | ||
| HB12 | 0 | 0.394 | 0.442 | 0 | 0.632 | 0.493 | ||||
| 2013 PV12 | 0.372 | 0 | 0.582 | |||||||
| 2014 PV12 | 0 | 0 | ||||||||
β diversity
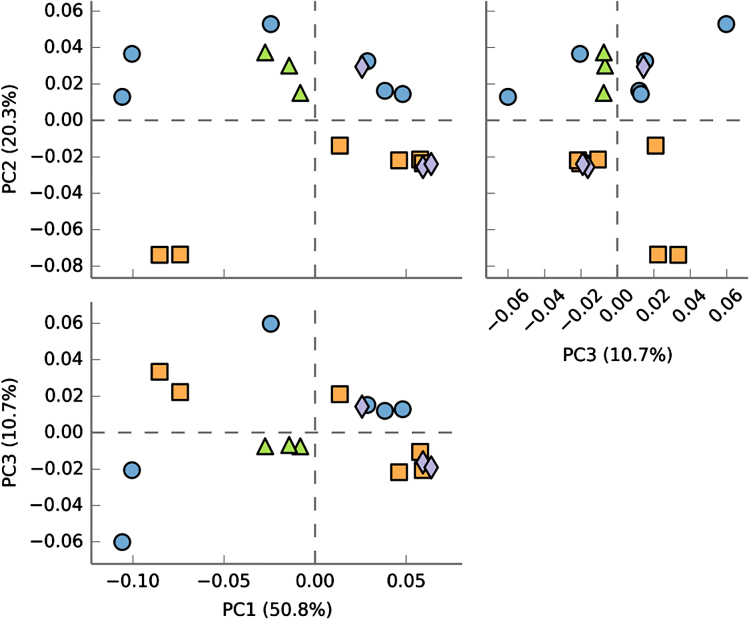
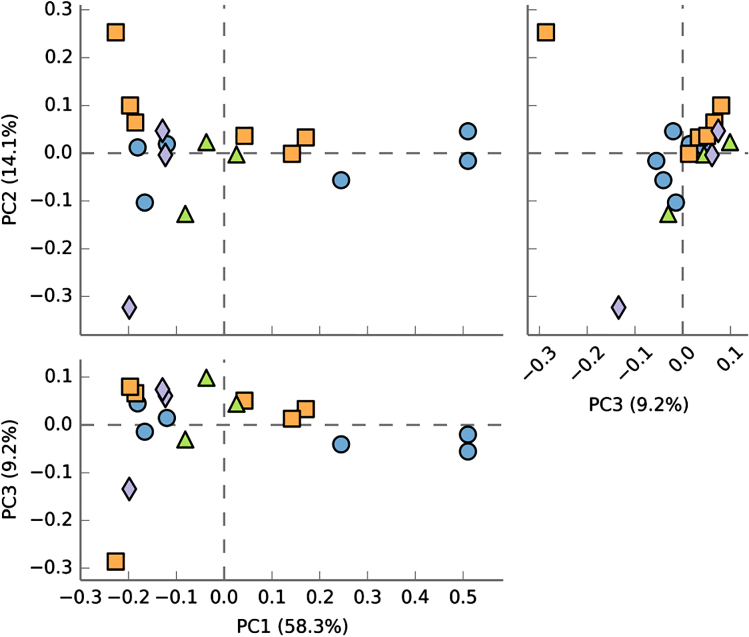
Table 3 shows the estimated β diversity measures for bacteria and fungi. In general, β diversity among samples was greater for fungi (i.e. lower similarity, high dissimilarity) than for bacteria. However, these diversity estimates did not show consistent patterns regarding their relationship with site and yield decline. For example, EMR site (no yield decline) showed the least similarity to the two sites with the yield-decline phenomenon (HB12 and PV12). However, HB12 was the least similar to PV12. Samples from HB12 and PV12, as a group, were not clearly separated from other samples based on principal component analysis of all bacterial OTUs (Figure 4) or fungal OTUs (Figure 5).
Figure 4.
Pairwise plots of the first three principal components from a principal component analysis of the bacterial OTU data (together with % variance accounted by each component) for samples taken in May 2014 from four sites: EM13 (circle), HB12 (triangle), HB13 (square) and PV12 (diamond). For PV12 and HB12 sites where yield decline was observed for non-chloropicrin-treated plants, samples for the chloropicrin-treated plots were not included.
Figure 5.
Pairwise plots of the first three principal components from a principal component analysis of the fungal OTU data (together with % variance accounted by each component) for samples taken in May 2014 from four sites: EM13 (circle), HB12 (triangle), HB13 (square) and PV12 (diamond). For PV12 and HB12 sites where yield decline was observed for non-chloropicrin-treated plants, samples for the chloropicrin-treated plots were not included.
Differential abundance
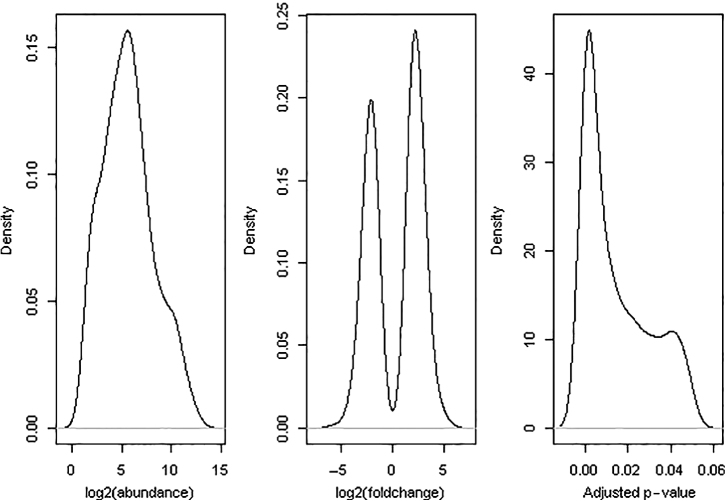
A total of 452 bacterial and 485 fungal OTUs were omitted from differential abundance testing because of extreme low counts across all samples (total counts ≤ 3), leaving 1690 bacterial and 1537 fungal OTUs for statistical testing. Of all pairwise comparisons, 715 comparisons were statistically significant at the 5% level. Figures 6 and 7 show the density plots of three statistics for these significant comparisons: average abundance for the OTUs, log2-fold change (no-yield-decline samples over yield-decline samples) and BH-adjusted p-values. Overall, more bacterial OTUs appeared to be more abundant in the non-yield-decline soils than in the yield-decline soils; the opposite was true for fungal OTUs.
Figure 6.
Density plots of three statistics for 591 cases for which there were significant differences in bacterial abundance between non-yield-decline and yield-decline samples: overall OTU abundance across all samples, log2-fold changes and BH-adjusted p-values.
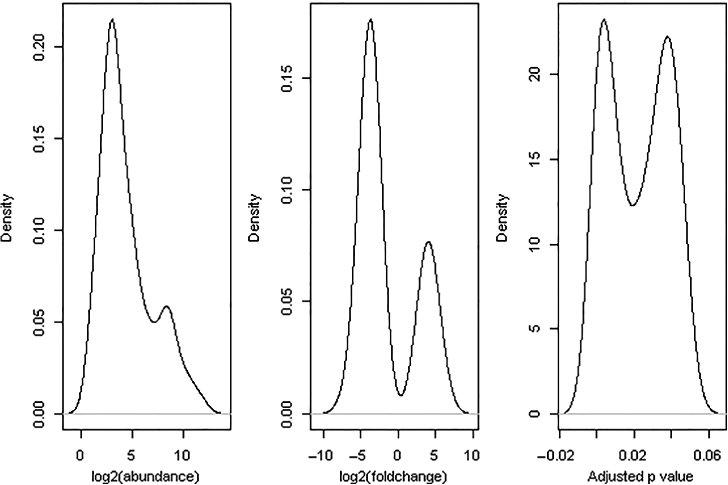
Figure 7.
Density plots of three statistics for 124 cases for which there were significant differences in fungal abundance between non-yield-decline and yield-decline samples: overall OTU abundance across all samples, log2-fold changes and BH-adjusted p-values.
There were 591 and 124 significant comparisons for bacterial and fungal OTUs, respectively. Excluding multiple significant comparisons for a single OTU, there were 366 and 88 unique bacterial and fungal OTUs for which there were significant differences in their abundance between the two types of samples (yield-decline vs. non-yield-decline). These fungal OTUs did not include common soil-borne pathogens, e.g. Fusarium, Verticillium, Phytophthora and Pythium. The log2-fold change was greater and less than zero for 326 and 325 bacterial comparisons, respectively; the corresponding values for fungal OTUs were 38 and 86. For bacteria, there were 203, 112, 35, 11 and 3 OTUs for which one, two, three, four and five out of the seven comparisons were statistically significant, respectively. For fungi, there were 63, 15, 9 and 1 OTUs for which one, two, three and four out of the seven comparisons were statistically significant, respectively.
To narrow down the number of OTUs for further interpretation of their possible roles in affecting yield decline, the following criteria were applied to these 454 OTUs (366 bacteria and 88 fungi):
average abundance should be over 10 and 6 across all samples for bacterial and fungal OTUs, respectively; this was used to exclude those OTUs with low counts since it is reasonable to assume that a high level of an OTU is needed if it was partially responsible for causing yield-decline;
the absolute log2-fold change is greater than 2.0 (i.e. the difference in abundance is at least four-fold);
there should be at least two statistically significant comparisons (out of the seven) for one single OTU;
for a single OTU, its effect sign (i.e. negative or positive) must be consistent among those significant comparisons involving the OTU.
In total, 32 bacterial and 17 fungal OTUs met these criteria. Of the 32 bacterial OTUs, log2-fold change was positive for 24 cases, i.e. no-yield-decline samples had greater abundance than the yield-decline samples. In 13 of the 17 fungal OTUs, no-yield-decline samples had lower abundance than yield-decline samples. Statistical test showed that there was significant differential abundance between fungal and bacterial OTUs between the two types of samples (p < 0.001).
Table 4 gives the taxonomical information of these 32 bacterial and 17 fungal OTUs. Of the 32 bacterial OTUs, 16 were from the phylum of Proteobacteria and six from Firmicutes. The remaining 10 OTUs were from eight phylum groups (Table 4). Ten fungal OTUs were from the Ascomycota and five from Basidiomycota. Further analysis of these OTUs in Table 4 (based on published research studies or online information) suggested 12 bacterial and 4 fungal OTUs (Table 5) could have played a role in the yield-decline phenomenon. Yield decline may result from one or more (and/or their interactions) of the following four factors: (1) lack of beneficial bacteria, (2) lack of nematode-parasitic fungi, (3) high levels of non-specific fungal root rot pathogens and (4) wet soil conditions.
Table 4. Bacterial and fungal OTUs that show significant differential abundance in soil samples from two types of soils: strawberry yield decline was or was not observed.
| Phylum | Class | Order | Family | Genus | Species | Effect sign |
|---|---|---|---|---|---|---|
|
Bacteria | ||||||
| Acidobacteria | Solibacteres | Solibacterales | (Bryobacteraceae) | Unidentified | Unidentified | + |
| ″ | Actinobacteria | Actinomycetales | Propionibacteriaceae | Unidentified | Unidentified | − |
| Bacteroidetes | (Saprospirae) | (Saprospirales) | Chitinophagaceae | Chitinophaga | Unidentified | + |
| ″ | Cytophagia | Cytophagales | Cytophagaceae | Hymenobacter | Unidentified | − |
| Chlorobi | Unidentified | Unidentified | Unidentified | Unidentified | Unidentified | + |
| Chloroflexi | Anaerolineae | Anaerolineales | Anaerolinaceae | Anaerolinea | Unidentified | − |
| Elusimicrobia | Elusimicrobia | FAC88 | Unidentified | Unidentified | Unidentified | + |
| Fibrobacteres | Fibrobacteria | 258ds10 | Unidentified | Unidentified | Unidentified | + |
| Firmicutes | Bacilli | Bacillales | (Exiguobacteraceae) | Unidentified | Unidentified | + |
| ″ | ″ | ″ | Bacillaceae | Bacillus | flexus | + |
| ″ | ″ | ″ | ″ | ″ | Unidentified | + |
| ″ | ″ | ″ | Planococcaceae | Solibacillus | Unidentified | + |
| ″ | ″ | ″ | ″ | Unidentified | Unidentified | + |
| ″ | Clostridia | Clostridiales | Ruminococcaceae | Unidentified | Unidentified | − |
| Gemmatimonadetes | Gemmatimonadetes | Gemmatimonadales | Ellin5301 | Unidentified | Unidentified | + |
| OD1 | ABY1 | Unidentified | Unidentified | Unidentified | Unidentified | − |
| Proteobacteria | Alphaproteobacteria | Caulobacterales | Caulobacteraceae | Asticcacaulis | biprosthecium | + |
| ″ | ″ | Caulobacterales | Caulobacteraceae | Unidentified | Unidentified | + |
| ″ | ″ | Rhizobiales | Bradyrhizobiaceae | Bosea | genosp. | + |
| ″ | ″ | Rhizobiales | Rhizobiaceae | Agrobacterium | sullae | + |
| ″ | ″ | Rhodospirillales | Rhodospirillaceae | Skermanella | Unidentified | − |
| ″ | ″ | Rickettsiales | mitochondria | Unidentified | Unidentified | + |
| ″ | ″ | Sphingomonadales | Sphingomonadaceae | Novosphingobium | Unidentified | + |
| ″ | Betaproteobacteria | Methylophilales | Methylophilaceae | Methylotenera | mobilis | + |
| ″ | ″ | Methylophilales | Methylophilaceae | Unidentified | Unidentified | + |
| ″ | ″ | Nitrosomonadales | Nitrosomonadaceae | Unidentified | Unidentified | + |
| ″ | Deltaproteobacteria | Desulfuromonadales | Geobacteraceae | Geobacter | Unidentified | − |
| ″ | ″ | Desulfuromonadales | Pelobacteraceae | Unidentified | Unidentified | − |
| ″ | Gammaproteobacteria | Alteromonadales | 211ds20 | Unidentified | Unidentified | + |
| ″ | ″ | Pseudomonadales | Moraxellaceae | Unidentified | Unidentified | + |
| ″ | ″ | ″ | Pseudomonadaceae | Pseudomonas | Unidentified | + |
| ″ | ″ | ″ | ″ | Unidentified | Unidentified | + |
|
Fungi | ||||||
| Ascomycota | Dothideomycetes | Capnodiales | Davidiellaceae | Cladosporium | C._sp_234B | − |
| ″ | ″ | ″ | Teratosphaeriaceae | Devriesia | fraseriae | + |
| ″ | ″ | Incertae_sedis | Pseudeurotiaceae | Pseudeurotium | bakeri | − |
| ″ | Eurotiomycetes | Eurotiales | Trichocomaceae | Paecilomyces | GZU_BCECYN4_3 | + |
| ″ | Leotiomycetes | Helotiales | Incertae_sedis | Tetracladium | T._sp_WMM_2012d | − |
| ″ | Sordariomycetes | Hypocreales | Incertae_sedis | Ilyonectria | coprosmae | − |
| ″ | ″ | ″ | ″ | ″ | robusta | − |
| ″ | ″ | Microascales | Microascaceae | Scedosporium | minutispora | + |
| ″ | ″ | Microascales | Unidentified | Unidentified | M._sp_MCJA12 | − |
| ″ | ″ | Unidentified | Unidentified | Unidentified | Sordariomycetes_sp | − |
| Basidiomycota | Agaricomycetes | Agaricales | Unidentified | Unidentified | uncultured_Agaricales | − |
| ″ | Microbotryomycetes | Leucosporidiales | Leucosporidiaceae | Leucosporidiella | fragaria | − |
| ″ | Tremellomycetes | Tremellales | Incertae_sedis | Cryptococcus | carnescens | + |
| ″ | ″ | ″ | ″ | ″ | macerans | − |
| ″ | Unidentified | Unidentified | Unidentified | Unidentified | Unidentified | − |
| Unidentified | Unidentified | Unidentified | Unidentified | Unidentified | fungal_sp_229c4c | − |
| Zygomycota | Incertae_sedis | Mortierellales | Mortierellaceae | Mortierella | Mortierella_sp_1_61D | − |
Table 5. Summary of differential abundance between non-yield-decline and yield-decline samples and had met several other selection criteria (see the text) for a number of bacterial and fungal OTUs, together with the log2-fold change (LFC) and possible roles they played in the yield decline of strawberry, which was inferred from published studies and/or from online resources.
| Significant comparisons |
|||||
|---|---|---|---|---|---|
| Family | Genus | Species | IDa | Mean LFC | Possible roles |
| Bacteria | |||||
| Bacillaceae | Bacillus | flexus | abc | 2.66 | Biocontrol, PGPRb |
| Unidentified | ade | 2.80 | Biocontrol, PGPR | ||
| Bradyrhizobiaceae | Bosea | genosp. | cfg | 2.17 | Nitrogen fixation |
| Rhizobiaceae | Agrobacterium | Sullae | abcde | 3.38 | Nitrogen fixation |
| Rhodospirillaceae | Skermanella | Unidentified | deg | −2.79 | Usually in anaerobic aquatic environments |
| Methylophilaceae | Methylotenera | mobilis | aceg | 3.26 | Nitrogen cycling in natural environments |
| Methylophilaceae | Unidentified | Unidentified | acef | 3.78 | Nitrogen cycling in natural environments |
| Nitrosomonadaceae | Unidentified | Unidentified | cfg | 2.22 | Oxidize ammonia into nitrite |
| Geobacteraceae | Geobacter | Unidentified | cfg | −2.21 | Anaerobic respiration |
| Pelobacteraceae | Unidentified | Unidentified | cef | −2.52 | Anaerobic respiration |
| Pseudomonadaceae | Pseudomonas | Unidentified | acdeg | 2.91 | PGPR |
| Unidentified | Unidentified | ace | 3.14 | PGPR | |
|
Fungi (Ascomycota) | |||||
| Trichocomaceae | Paecilomyces | GZU_BCECYN4_3 | de | 3.66 | Killing nematodes |
| Incertae_sedis | Tetracladium | WMM_2012d | bd | −2.91 | Water-logging rich fungus |
| Incertae_sedis | Ilyonectria | coprosmae | ace | −4.99 | Including Cylindrocarpon spp. or Cylindrocarpon-like fungi that can cause non-specific root rot |
| robusta | ae | −5.18 | |||
Identity of individual comparisons for which differences in microbial abundance were statistically different; a – EM13/HB13 vs. HB12/PV12; b – EM13 vs. HB12; c – EM13 vs. PV12; d – HB13 vs. HB12; e – HB13 vs. PV12; f – chloropicrin-treated vs. others at PV12 (2013 samples); and g – chloropicrin-treated vs. control at HB12.
PGPR, plant growth-promoting rhizobacteria.
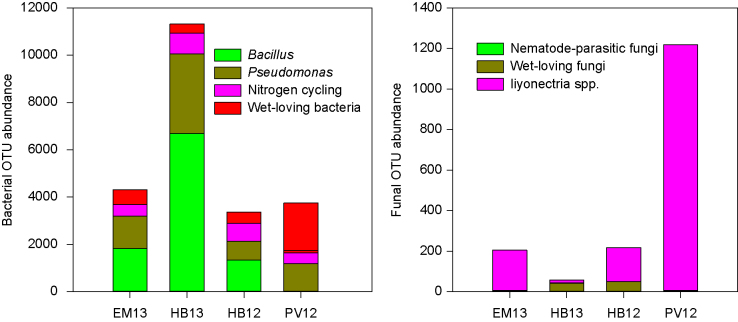
Figure 8 shows the relative abundance of these 16 candidate OTUs at the four sites. HB13 and EM13 had much higher levels of Bacillus and Pseudomonas OTUs than HB12 and PV12. The level of wet-loving bacterial OTUs was higher at PV12 than at the other sites; the opposite was true for the level of bacterial OTUs related to nitrogen cycling. For fungal OTUs, the level of two Ilyonectria species was the highest and lowest at PV12 and HB13, respectively. There was a higher level of wet-loving fungi at HB12 than at the other three sites. Nematode-parasitic fungi were nearly exclusively found at HB13.
Figure 8.
Relative abundance of the 16 (12 bacterial and 4 fungal OTUs – Table 7) organisms per sample that may have played a role in affecting strawberry yield decline observed at HB12 and PV12; counts data were proportionally adjusted to the total reads of 300 000 per sample. Samples from chloropicrin-treated plots at HB12 and PV12 were excluded.
Discussion
The amplicon-based metagenomic analysis of soil samples utilised in this study identified several groups of microbial organisms that may be involved in causing strawberry yield decline. Many OTUs differ in their abundance between samples from the yield decline and non-decline soils. Based on the comparisons of abundance of each individual OTU and several stringent criteria, we identified up to 51 (34 bacterial and 17 fungal OTUs) that were most likely to be involved in affecting yield decline of strawberry. Of these 51 OTUs, only for 12 bacterial OTUs and four fungal OTUs is there published information about their possible roles (or those of closely related species), which is biologically plausible to explain why the OTUs were implicated in strawberry yield decline.
There are two fungal OTUs that had very high abundance at one of the two yield-decline sites (PV12); these two OTUs are Ilyonectria robusta and I. coprosmae. Of these two OTUs, I. robusta is particularly abundant at PV12. Based on recent molecular taxonomy, these two IIyonectria species are closely related to Cylindrocarpon spp; indeed, IIyonectria contains many Cylindrocarpon-like species that have been commonly associated with root and decay of woody and herbaceous plants35. Recently, I. robusta has been shown to cause root diseases on grapevine (including vascular invasion)36–38. Cylindrocarpon-like and Ilyonectria cause diseases on Laurustinus39. Potential fungal pathogens other than V. dahliae were frequently recovered from strawberry roots, e.g. Rhizoctonia sp.40,41, and Cylindrocarpon destructans, Fusarium oxysporum, Fusarium solani, Pestalotia longiseta and Pythium spp42–44. Cylindrocarpon destructans can cause variable degrees of crown and root rot in strawberry45,46. Collectively these general, non-specific pathogens that cause a root disease are commonly referred to as black root rot, a name that is descriptive of the appearance of the roots47.
The importance of these non-specific root pathogens may have been masked by two factors. First, broad-spectrum chemo-fumigants (e.g., methyl bromide and chloropicrin) have been used to fumigate soil. Second, unlike V. dahliae these non-specific root pathogens do not usually lead to plant mortality but only lead to reduced plant growth. This reduced plant growth is often difficult to differentiate from nutrient deficiency or other abiotic factors. In non-fumigated soils, strawberry yield reduction between 20–25%43 and 46% (http://www.mbao.org/altmet00/32martin.pdf, accessed on 26 November 2014) was observed but was not attributable to V. dahliae, similar to the loss observed at HB12 and PV12 (ca. 25%). Based on root isolations, this yield decline was attributed to root rot caused by Pythium, binucleate Rhizoctonia and Cylindrocarpon spp.
Of the 12 bacterial OTUs, 2 were from the genus Bacillus and 2 from the genus Pseudomonas, which were more abundant in non-yield-decline soil samples. Many strains from Bacillus and Pseudomonas have been demonstrated to have anti-fungal effects and can promote plant growth48–56. There are already several commercial products based on the strains from these two genera marketed as biocontrol products against plant diseases and/or as PGPR products. Probably the most widely used biocontrol strain for plant pathogens is B. subtlis, e.g. one formulated commercial product is called Serenade®. Soil suppressiveness against non-specific root pathogens is associated with the high level of total fungi and fluorescent bacteria42; suppressiveness is reduced on sites where a strawberry monoculture without organic input has been grown for several years. Two Bacillus strains reduced ginseng root rot caused by C. destructans57. Several Pseudomonas strains isolated from the rhizosphere of oilseed rape and strawberry were antagonistic against V. dahliae19. Early and localised root surface and root endophytic colonisation by P. fluorescens PICF7 is needed to impair full progress of Verticillium wilt epidemics in olive58. However, these identified beneficial bacterial OTUs in the present study are not likely to have much antagonistic effect against V. dahliae since the wilt incidence was similar between yield-decline and non-yield-decline plots at HB12 and PV12.
At the HB13 site, an OTU from the genus of Paecilomyces was much greater in abundance than at the two yield-decline sites. This genus is known to contain nematophagous fungal species, killing nematodes by pathogenesis59–61. It is also generally accepted that nematodes may exacerbate wilt problems in strawberry by providing wounds as fungal entry sites. The presence of nematodes increased the rate of Verticillium wilt development on strawberry except when the fungal inoculum level was very low62. In the presence of V. dahliae inoculum, the tolerance of strawberry to lesion nematodes was reduced by 50%63. Wilt in potatoes is more severe in the presence of lesion nematodes64.
Five bacterial OTUs were more abundant in non-yield-decline soils than in yield-decline soils. They are reported to have been involved in global nitrogen cycles, e.g., Methylophilaceae – de-nitrification65, Rhizobiaceae – nitrogen fixation66 and Nitrosomonadaceae – nitrification (oxidising ammonia into nitrite)67. Thus, these organisms may improve plant growth due to increased availability of nitrogen in the soil to plants. Several other microbial OTUs were more abundant in yield-decline soils than in non-decline soils and they are usually abundant in water or wet conditions or involved in anaerobic respiration. These OTUs are not likely to directly influence strawberry growth. Rather their abundance may indicate water logging or high levels of soil moisture content, which in general would favour pathogen development and reduce root development in strawberry68. Soil physiochemical properties may also affect plant and pathogen development. It is not always clear whether such effects are through their direct influence on plant development or (partially) mediated through their effects on soil microbial populations.
The two yield-decline sites have been in continuous strawberry production for many years. In contrast, HB13 site was previously an apple orchard and EM13 site was planted with cereals in previous years. Rotation is one of the cultural practices used to manage diseases, particularly soil-borne pathogens69–72. Rotation was able to reduce the severity of strawberry root rot73–75. Continuous cropping of a single species for a long period of time often leads to reduced cropping potential in rosaceous species, e.g., apple25 and almond76. Further research is needed to understand which bacterial and fungal OTUs associated with strawberry yield decline are also associated with continuous cropping of strawberry and which of these OTUs are due to the nature of soils.
Soil sampling in the present work was unfortunately constrained by the limited number of sites with yield-decline and commercial horticulture. Thus it was not possible to adopt a more controlled sampling plan that could adequately consider other site-specific factors, such as soil type, previous crops and host genotypes. These site-specific factors undoubtedly affect plant health, either contributing to soil disease suppression and nutrient supply/uptake or predisposing plants to pathogens77,78. However, they are unlikely to be the determinant of strawberry yield decline because strawberry plants in plots treated with chloropicrin at the same sites did not suffer from the yield decline. The present study, based on the data-driven approach, was able to identify candidate microbes for further studies where some of these site-specific factors could be included. For instance, how do the two candidate fungal pathogens interact with beneficial soil microbiota in different types of soil in relation to strawberry yield?
It should be noted that the present approach (relative DNA abundance) will not be applicable to scenarios where a very low level of microbes could cause disproportional effects on plant development, e.g., by producing toxins. Such a problem (linking soil microbial functions to plant development) may potentially be dealt with metatranscriptomics (RNA-seq) or metabolomics-based approaches. Recent research reports on the transcriptomes of microbial community in soil79 and Arctic peat soil80 highlight the feasibility of applying metatranscriptomics to soil microbes (though still challenging).
In summary, this study demonstrated the usefulness of amplicon-based metagenomics to identify candidate organisms involved in affecting strawberry yield decline. Isolation and culture techniques are usually used to obtain microbial organisms from root systems for further pathological studies. But this time-consuming approach suffers from the fact that most microbial organisms cannot be cultured in artificial media. An amplicon-profiling approach provided an efficient way to profile microbiota and identify candidate organisms for further hypothesis testing and confirmatory studies.
Acknowledgments
This research is funded by the Innovate UK (project number: 100867) with matching funding from several commercial companies. Feng Wei gratefully acknowledges the financial assistance of the China Scholarship Council.
The authors declare no conflict of interest.
References
- Pegg GF, Brady BL. Verticillium wilts, New York: CABI Publishing, 2002. [Google Scholar]
- Rotenberg D, MacGuidwin AE, Saeed IAM, Rouse DI. Interaction of spatially separated Pratylenchus penetrans and Verticillium dahliae on potato measured by impaired photosynthesis. Plant Pathol 2004; 53: 294–302. [Google Scholar]
- Saeed IAM, Macguidwin AE, Rouse DI. Effect of initial nematode population density on the interaction of Pratylenchus penetrans and Verticillium dahliae on ‘Russet burbank’ potato. J Nematol 1998; 30: 100–107. [PMC free article] [PubMed] [Google Scholar]
- Botseas DD, Rowe RC. Development of potato early dying in response to infection by 2 pathotypes of Verticillium dahliae and coinfection by Pratylenchus penetrans. Phytopathol 1994; 84: 275–282. [Google Scholar]
- Martin FN. Development of alternative strategies for management of soilborne pathogens currently controlled with methyl bromide. Annu Rev Phytopathol 2003; 41: 325–350. [DOI] [PubMed] [Google Scholar]
- Korthals GW, Thoden TC, van den Berg W, Visser JHM. Long-term effects of eight soil health treatments to control plant-parasitic nematodes and Verticillium dahliae in agro-ecosystems. Appl Soil Ecol 2014; 76: 112–123. [Google Scholar]
- Easton GD, Nagle ME, Seymour MD. Potato production and incidence of Verticillium dahliae following rotation to nonhost crops and soil fumigation in the State of Washington. Am Potato J 1992; 69: 489–502. [Google Scholar]
- Matthiessen JN, Kirkegaard JA. Biofumigation and enhanced biodegradation: opportunity and challenge in soilborne pest and disease management. Crit Rev Plant Sci 2006; 25: 235–265. [Google Scholar]
- Neubauer C, Heitmann B, Müller C. Biofumigation potential of Brassicaceae cultivars to Verticillium dahliae. Eur J Plant Pathol 2014; 140: 341–352. [Google Scholar]
- Mercier J, Jimenez JI. Demonstration of the biofumigation activity of Muscodor albus against Rhizoctonia solani in soil and potting mix. Biocontrol 2009; 54: 797–805. [Google Scholar]
- Angus JF, Gardner PA, Kirkegaard JA, Desmarchelier JM. Biofumigation: Isothiocyanates released from brassica roots inhibit growth of the take-all fungus. Plant Soil 1994; 162: 107–112. [Google Scholar]
- Yohalem D, Passey T. Amendment of soils with fresh and post-extraction lavender (Lavandula angustifolia) and lavandin (Lavandula x intermedia) reduce inoculum of Verticillium dahliae and inhibit wilt in strawberry. Appl Soil Ecol 2011; 49: 187–196. [Google Scholar]
- Runia WT, Molendirk LP, Ludeking DJ, Schomaker CH. Improvement of anaerobic soil disinfestation. Commun Agric Appl Biol Sci 2012; 77: 753–62. [PubMed] [Google Scholar]
- Vitullo D, Altieri R, Esposito A et al. Suppressive biomasses and antagonist bacteria for an eco-compatible control of Verticillium dahliae on nursery-grown olive plants. Int J Env Sci Technol 2013; 10: 209–220. [Google Scholar]
- Bonanomi G, Antignani V, Capodilupo M, Scala F. Identifying the characteristics of organic soil amendments that suppress soilborne plant diseases. Soil Biol Biochem 2010; 42: 136–144. [Google Scholar]
- Njoroge SMC, Kabir Z, Martin FN, Koike ST, Subbarao KV. Comparison of crop rotation for Verticillium wilt management and effect on Pythium species in conventional and organic strawberry production. Plant Dis 2009; 93: 519–527. [DOI] [PubMed] [Google Scholar]
- Subbarao KV, Kabir Z, Martin FN, Koike ST. Management of soilborne diseases in strawberry using vegetable rotations. Plant Dis 2007; 91: 964–972. [DOI] [PubMed] [Google Scholar]
- Angelopoulou DJ, Naska EJ, Paplomatas EJ, Tjamos SE. Biological control agents (BCAs) of verticillium wilt: influence of application rates and delivery method on plant protection, triggering of host defence mechanisms and rhizosphere populations of BCAs. Plant Pathol 2014; 63: 1062–1069. [Google Scholar]
- Berg G, Fritze A, Roskot N, Smalla K. Evaluation of potential biocontrol rhizobacteria from different host plants of Verticillium dahliae Kleb. J Appl Microbiol 2001; 91: 963–971. [DOI] [PubMed] [Google Scholar]
- Berg G, Zachow C, Lottmann J et al. Impact of plant species and site on rhizosphere-associated fungi antagonistic to Verticillium dahliae Kleb. Appl Env Microbiol 2005; 71: 4203–4213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peiffer JA, Spor A, Koren O et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci U S A 2013; 110: 6548–6553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurze S, Bahl H, Dahl R, Berg G. Biological control of fungal strawberry diseases by Serratia plymuthica HRO-C48. Plant Dis 2001; 85: 529–534. [DOI] [PubMed] [Google Scholar]
- Lugtenberg B, Kamilova F. Plant-growth-promoting rhizobacteria. Ann Rev Microbiol 2009; 63: 541–556. [DOI] [PubMed] [Google Scholar]
- Diehl K, Rebensburg P, Lentzsch P. Field application of non-pathogenic Verticillium dahliae genotypes for regulation of wilt in strawberry plants. Am J Plant Sci 2013; 4: 24–32. doi:10.4236/ajps.2013.47A2004. [Google Scholar]
- Mazzola M, Manici L. Apple replant disease: role of microbial ecology in cause and control. Ann Rev Phytopathol 2012; 50: 45–65. [DOI] [PubMed] [Google Scholar]
- Caporaso JG, Kuczynski J, Stombaugh J et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 2010; 7: 335–336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quast C, Pruesse E, Yilmaz P et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 2013; 41: D590–D596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abarenkov K, Henrik Nilsson R, Larsson K-H et al. The UNITE database for molecular identification of fungi – recent updates and future perspectives. New Phytol 2010; 186: 281–285. [DOI] [PubMed] [Google Scholar]
- Robertson CE, Harris JK, Wagner BD et al. Explicet: graphical user interface software for metadata-driven management, analysis and visualization of microbiome data. Bioinformatics 2013; 29: 3100–3101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parks DH, Tyson GW, Hugenholtz P, Beiko RG. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 2014; 30: 3123–3124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 2014; 550. doi:10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMurdie PJ, Holmes S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 2014; 10: e1003531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol 2010; 11: R106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powderul approach to multple testing. J Royal Stat Soc Ser B 1995; 57: 289–300. [Google Scholar]
- Cabral A, Groenewald J, Rego C, Oliveira H, Crous P. Cylindrocarpon root rot: multi-gene analysis reveals novel species within the Ilyonectria radicicola species complex. Mycol Prog 2012; 11: 655–688. [Google Scholar]
- Reis P, Cabral A, Nascimento T, Olivera H, Rego C. Diversity of Ilyonectria species in a young vineyard affected by black foot disease. Phytopathologia Mediterranea 2013; 52: 335–346. [Google Scholar]
- dos Santos RF, Blume E, da Silva GBP et al. First report of Ilyonectria robusta associated with black foot disease of grapevine in Southern Brazil. Plant Dis 2013; 98: 845–845. [DOI] [PubMed] [Google Scholar]
- Úrbez-Torres JR, Haag P, Bowen P, O’Gorman DT. Grapevine trunk diseases in British Columbia: incidence and characterization of the fungal pathogens associated with black foot disease of grapevine. Plant Dis 2013; 98: 456–468. [DOI] [PubMed] [Google Scholar]
- Aiello D, Guarnaccia V, Epifani F, Perrone G, Polizzi G. ‘Cylindrocarpon’ and Ilyonectria species causing root and crown rot disease of potted Laurustinus plants in Italy. J Phytopathol 2014. doi:10.1111/jph.12306.
- Martin FN. Rhizoctonia spp. recovered from strawberry roots in Central Coastal California. Phytopathol 2000; 90: 345–353. [DOI] [PubMed] [Google Scholar]
- Botha A, Denman S, Lamprecht SC, Mazzola M, Crous PW. Characterisation and pathogenicity of Rhizoctonia isolates associated with black root rot of strawberries in the Western Cape Province, South Africa. Aust Plant Pathol 2003; 32: 195–201. [Google Scholar]
- Manici LM, Caputo F, Baruzzi G. Additional experiences to elucidate the microbial component of soil suppressiveness towards strawberry black root rot complex. Ann Appl Biol 2005; 146: 421–431. [Google Scholar]
- Yuen GY, Schroth MN, Weinhold AR, Hancock JG. Effects of soil fumigation with methyl bromide and chloropicrin on root health and yield of strawberry. Plant Dis 1991; 75: 416–420. [Google Scholar]
- Koike ST, Kirkpatrick SC, Gordon TR. Fusarium wilt of strawberry caused by Fusarium oxysporum in California. Plant Dis 2009; 93: 1077–1077. [DOI] [PubMed] [Google Scholar]
- Fang X, Phillips D, Li H, Sivasithamparam K, Barbetti MJ. Comparisons of virulence of pathogens associated with crown and root diseases of strawberry in Western Australia with special reference to the effect of temperature. Sci Horticult 2011; 131: 39–48. [Google Scholar]
- Fang XL, Phillips D, Li H, Sivasithamparam K, Barbetti MJ. Severity of crown and root diseases of strawberry and associated fungal and oomycete pathogens in Western Australia. Aust Plant Pathol 2011; 40: 109–119. [Google Scholar]
- Wing KB, Pritts MP, Wilcox. WF. Strawberry black root rot: a review. Adv Strawberry Res 1994; 13: 13–19. [Google Scholar]
- Arrebola E, Sivakumar D, Bacigalupo R, Korsten L. Combined application of antagonist Bacillus amyloliquefaciens and essential oils for the control of peach postharvest diseases. Crop Prot 2010; 29: 369–377. [Google Scholar]
- Arrebola E, Sivakumar D, Korsten L. Effect of volatile compounds produced by Bacillus strains on postharvest decay in citrus. Biol Control 2010; 53: 122–128. [Google Scholar]
- Casals C, Elmer PAG, Vinas I et al. The combination of curing with either chitosan or Bacillus subtilis CPA-8 to control brown rot infections caused by Monilinia fructicola. Postharv Biol Technol 2012; 64: 126–132. [Google Scholar]
- Dedej S, Delaphane KS, Scherm H. Effectiveness of honey bees in delivering the biocontrol agent Bacillus subtilis to blueberry flowers to suppress mummy berry disease. Biol Control 2004; 31: 422–427. [Google Scholar]
- Gilardi G, Manker DC, Garibaldi A, Gullino ML. Efficacy of the biocontrol agents Bacillus subtilis and Ampelomyces quisqualis applied in combination with fungicides against powdery mildew of zucchini. J Plant Dis Prot 2008; 115: 208–213. [Google Scholar]
- Gao G, Yin D, Chen S et al. Effect of biocontrol agent Pseudomonas fluorescens 2P24 on soil fungal community in cucumber rhizosphere using T-RFLP and DGGE. PLoS One 2012; 7: e31806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Latha P, Anand T, Prakasam V et al. Combining Pseudomonas, Bacillus and Trichoderma strains with organic amendments and micronutrient to enhance suppression of collar and root rot disease in physic nut. Appl Soil Ecol 2011; 49: 215–223. [Google Scholar]
- Dandurishvili N, Toklikishvili N, Ovadis M et al. Broad-range antagonistic rhizobacteria Pseudomonas fluorescens and Serratia plymuthica suppress Agrobacterium crown gall tumours on tomato plants. J Appl Microbiol 2011; 110: 341–352. [DOI] [PubMed] [Google Scholar]
- Couillerot O, Prigent-Combaret C, Caballero-Mellado J, Moënne-Loccoz Y. Pseudomonas fluorescens and closely-related fluorescent pseudomonads as biocontrol agents of soil-borne phytopathogens. Lett Appl Microbiol 2009; 48: 505–512. [DOI] [PubMed] [Google Scholar]
- Jang Y, Kim SG, Kim YH. Biocontrol efficacies of Bacillus species against Cylindrocarpon destructans causing ginseng root rot. Plant Pathol J 2011; 27: 333–341. [Google Scholar]
- Prieto P, Navarro-Raya C, Valverde-Corredor A et al. Colonization process of olive tissues by Verticillium dahliae and its in planta interaction with the biocontrol root endophyte Pseudomonas fluorescens PICF7. Microb Biotechnol 2009; 2: 499–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan A, Williams K, Nevalainen HM. Control of plant-parasitic nematodes by Paecilomyces lilacinus and Monacrosporium lysipagum in pot trials. Biocontrol 2006; 51: 643–658. [Google Scholar]
- Anastasiadis IA, Giannakou IO, Prophetou-Athanasiadou DA, Gowen SR. The combined effect of the application of a biocontrol agent Paecilomyces lilacinus, with various practices for the control of root-knot nematodes. Crop Prot 2008; 27: 352–361. [Google Scholar]
- Castillo JD, Lawrence KS, Kloepper JW. Biocontrol of the reniform nematode by Bacillus firmus GB-126 and Paecilomyces lilacinus 251 on cotton. Plant Dis 2013; 97. [DOI] [PubMed] [Google Scholar]
- McKinley R, Talboys PW. Effects of Practylenchus penetrans on development of strawberry wilt caused by Verticillium dahliae. Ann Appl Biol 1979; 92: 347–357. [Google Scholar]
- Szczygiel A. On the tolerance limit of strawberry plants to root lesion nematode Pratylenchus penetrans (Cobb). Fruit Sci Rep 1983; 10: 135–142. [Google Scholar]
- Powelson ML, Rowe RC. Biology and management of early dying of potatoes. Annu Rev Phytopathol 1993; 21: 111–126. [DOI] [PubMed] [Google Scholar]
- Kalyuhznaya MG, Martens-Habbena W, Wang T et al. Methylophilaceae link methanol oxidation to denitrification in freshwater lake sediment as suggested by stable isotope probing and pure culture analysis. Env Microbiol Rep 2009; 1: 385–392. [DOI] [PubMed] [Google Scholar]
- Dilworth J, James EK, Sprent JI, Newton WE (eds.). Nitrogen-fixing leguminous symbioses. Dordrecht: Springer, 2008. [Google Scholar]
- Garrity GM, Brenner DJ, Krieg NR, Staley JT (eds.). Bergey’s Manual of Systematic Bacteriology, Volume two: the proteobacteria, part C: the alpha-, beta-, delta-, and epsilonproteobacteria. New York: Springer, 2005. [Google Scholar]
- Maas J (ed.) Compendium of strawberry diseases. St. Paul: 128 American Phytopathological Society, 1998. [Google Scholar]
- Wheeler TA, Bordovsky JP, Keeling JW, Mullinix BG, Woodward JE. Effects of crop rotation, cultivar, and irrigation and nitrogen rate on Verticillium wilt in cotton. Plant Dis 2012; 96: 985–989. [DOI] [PubMed] [Google Scholar]
- Bernard E, Larkin RP, Tavantzis S et al. Compost, rapeseed rotation, and biocontrol agents significantly impact soil microbial communities in organic and conventional potato production systems. Appl Soil Ecol 2012; 52: 29–41. [Google Scholar]
- Ebihara Y, Uematsu S, Nomiya S. Control of Verticillium dahliae at a strawberry nursery by paddy-upland rotation. J Gen Plant Pathol 2010; 76: 7–20. [Google Scholar]
- Xing LJ, Westphal A. Effects of crop rotation of soybean with corn on severity of sudden death syndrome and population densities of Heterodera glycines in naturally infested soil. Field Crops Res 2009; 112: 107–117. [Google Scholar]
- Elmer WH, LaMondia JA. Influence of ammonium sulfate and rotation crops on strawberry black root rot. Plant Dis 1999; 83: 119–123. [DOI] [PubMed] [Google Scholar]
- Seigies AT, Pritts M. Cover crop rotations alter soil microbiology and reduce replant disorders in strawberry. Hortsci 2006; 41: 1303–1308. [Google Scholar]
- LaMondia JA. Influence of rotation crops on the strawberry pathogens Pratylenchus penetrans, Meloidogyne hapla, and Rhizoctonia fragariae. J Nematol 1999; 31: 650–655. [PMC free article] [PubMed] [Google Scholar]
- Browne GT, Connell JH, Schneider SM. Almond replant dsease and its management with alternative pre-plant soil fumigation treatments and rootstocks. Plant Dis 2006; 90: 869–876. [DOI] [PubMed] [Google Scholar]
- Janvier C, Villeneuve F, Alabouvette C et al. Soil health through soil disease suppression: which strategy from descriptors to indicators? Soil Biol Biochem 2007; 39: 1–23. [Google Scholar]
- Kinkel LL, Bakker MG, Schlatter DC. A coevolutionary framework for managing disease-suppressive soils. Annu Rev Phytopathol 2011; 49: 47–67. [DOI] [PubMed] [Google Scholar]
- Turner TR, Ramakrishnan K, Walshaw J et al. Comparative metatranscriptomics reveals kingdom level changes in the rhizosphere microbiome of plants. ISME J 2013; 7: 2248–2258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tveit A, Urich T, Svenning MM. Metatranscriptomic analysis of Arctic peat soil microbiota. Appl Env Microbiol 2014; 80: 5761–5772. [DOI] [PMC free article] [PubMed] [Google Scholar]