Abstract
Plant innate immune response against viruses utilizes intracellular Nucleotide Binding domain Leucine Rich Repeat (NLR) class of receptors. NLRs recognize different viral proteins termed elicitors and initiate diverse signaling processes that induce programmed cell death (PCD) in infected cells and restrict virus spread. In this review we describe the recent advances made in the study of plant NLRs that detect viruses. We describe some of the physical and functional interactions these NLRs undertake. We elaborate on the intra-molecular and homotypic association of NLRs that function in self-regulation and activation. Nuclear role for some viral NLRs is discussed as well as the emerging importance of the RNAi pathway in regulating the NLR family.
Introduction
The Dutch microbiologist Martinus Beijerinck described in 1898 a unique infectious agent, one much smaller than other known microbes of the time, that caused mosaic disease in tobacco. This agent, later named Tobacco mosaic virus (TMV), was the first virus to be discovered and laid the foundation for the new field of virology. Since this discovery, plant pathologists have identified more than 1000 viruses that infect almost all plant species, many of which can cause serious economic impact on crop production. Viruses are obligate pathogens that rely on plant proteins, processes, and organelles for successful infection. Therefore, one strategy plants commonly use to prevent virus infection is to avoid production of a host factor required for infection. This explains why many viruses have a very limited host range. Plants can also use non-specific RNAi-based surveillance to detect the presence of viral nucleic acids, which are targeted by the host RNAi machinery for degradation [1]. A third and highly specific plant response to viral infection requires the function of specialized intracellular immune receptors belonging to the Nucleotide Binding domain Leucine Rich Repeat (NLR) class [2, 3]. In this review we discuss recent discoveries made with respect to these plant NLRs that detect viruses. Specific emphasis is placed on the mechanistic insights on pathogen recognition, NLR activation, and the potential nuclear functions of NLRs.
Plant viral NLRs
The NLR family is the major class of intracellular immune receptors in plants and animals and functions to sense pathogen invasion and activate immune signaling [2]. NLRs universally contain a centrally located nucleotide-binding domain (NB) and a C-terminal leucine rich repeat (LRR) domain. The N-terminal regions of plant viral NLRs further possess either a Toll/Interleukin-1 Receptor homology domain (TIR) or Coil-Coiled (CC) domain (Table 1). The fact that mammalian and other eukaryotic immune receptors also possess these domains in different combinations suggests the utility of these domains as pathogen sensors and signal transmitters. Plant viral NLRs activate immune signaling after recognizing different viral-encoded proteins referred to as elicitors or effectors (Table 1). In contrast, animal NLRs recognizes Pathogen Associated Molecular Patterns (PAMPs) such as viral RNA [4]. In addition to intracellular NLRs, animals also use cell surface Pattern Recognition Receptors (PRRs) such as Toll-like receptors (TLRs) and cytoplasmic RIG-I-like Receptors (RLRs) to detect viral nucleic acid [4]. In plants, cell surface receptor-like PRRs detect bacterial and fungal PAMPs, but so far there is no evidence that these receptors function in viral pathogen detection. Instead, plants activate RNAi-mediated anti-viral defense against invading viral nucleic acid [1].
Table 1.
Cloned NLRs that confer resistance to different viruses
| NLR | Plant | NLR Class | Virus | Elicitor |
|---|---|---|---|---|
| N | Nicotiana | TIR-NB-LRR | Tobacco Mosaic Virus | Replicase |
| Y-1 | Potato | TIR-NB-LRR | Potato Virus Y | Nla Protease |
| Rx | Potato | CC-NB-LRR | Potato Virus X | Coat Protein |
| Rx2 | Potato | CC-NB-LRR | Potato Virus X | Coat Protein |
| HRT | Arabidopsis | CC-NB-LRR | Turnip Crinkle Virus | Coat Protein |
| RCY1 | Arabidopsis | CC-NB-LRR | Cucumber Mosaic Virus Y | Coat Protein |
| Tm-2 | Tomato | CC-NB-LRR | Tomato Mosaic Virus | Movement Protein |
| Tm-22 | Tomato | CC-NB-LRR | Tomato Mosaic Virus | Movement Protein |
| Sw-5 | Tomato | CC-NB-LRR | Tomato Spotted Wilt Virus | Unknown |
| RSV1 * | Soybean | CC-NB-LRR | Soybean Mosaic Virus | Hc-Pro and P3 |
RSV1 maps to a cluster of NLR genes. The exact gene(s) that confers resistance has/have not yet been identified.
For references to specific gene discoveries, see [56]
Recognition of plant viral elicitors by NLRs
The tobacco N gene was the first TIR-NLR cloned from plants and confers resistance to TMV [5]. N recognizes the helicase domain (referred to as p50) within the TMV replicase protein to activate successful immune signaling. Since the cloning of N, at least 9 viral NLRs and corresponding viral elicitors have been reported (Table 1). Evidence indicates that N recognizes TMV p50 indirectly through a chloroplast localized N Receptor Interacting Protein 1 (NRIP1) [6]. In the presence of TMV, a fraction of NRIP1 relocalizes from the chloroplast to the cytoplasm and nucleus. Cytoplasmic NRIP1 associates with the TMV replicase which then leads to N recruitment through interaction with its TIR domain. Yeast two-hybrid and in vitro pull down assays have shown that the LRR of N can directly associate with p50 [7]. However, in planta association has not been demonstrated. An additional confounding factor is that the p50 mutant (p50P149L) used in these studies and presumed to abolish N-mediated recognition, is naturally present in all TMV strains that are recognized by N [8].
NLR recognition of elicitors/effectors through an intermediary host protein has been documented for several plant bacterial NLRs [2, 3]. However, in the absence of in planta association data between other viral NLRs and corresponding viral elicitors, it is not possible to generalize on whether indirect recognition is a common phenomenon for viral recognition. The CC domain of the Rx NLR does form a stable complex with the host factor Ran GTPase Activating Protein 2 (RanGAP2) for complete activation of defense against Potato Virus X (PVX) [9]. However, since RanGAP2 has not been shown to associate with viral CP elicitor, it is still unclear if RanGAP2 mediates elicitor recognition or if it has other roles (see section on Potential Nuclear function of NLRs) [10].
Another example of potential indirect viral recognition is immunity against some strains of TMV including Tomato Mosaic Virus (ToMV) provided by the Tm-22 CC-NLR. Recently, MP Interacting Protein 1 (NbMIP1) was found to associate in vivo with the ToMV MP as well as the Tm-22 CC-NLR [11]. NbMIP1 is a J domain containing protein and has molecular chaperone functions. NbMIP1 associates with the LRR domain of Tm-22, which has been implicated in recognition of the ToMV MP elicitor. Intriguingly, NbMIP1 and its related members are required for both ToMV infection as well as Tm-22-mediated immunity. NbMIPs were shown to be necessary for maintaining protein stability of ToMV MP and Tm-22. One possibility is that NbMIP1 functions simply as a general co-chaperone. Alternately, it has been suggested that NbMIP1 could be an intermediary protein facilitating Tm-22 association with MP. While preliminary evidence suggests that Tm-22 and MP associate with NbMIP1, it is imperative to confirm whether these three exist in the same complex.
NLR domain(s) involved in elicitor recognition
Early models of viral recognition suggested a predominant role for the LRR domain in pathogen recognition and the N-terminal CC/TIR domain in signaling [12]. Domain swap analysis between the Arabidopsis RCY1 (Resistance to Cucumber Mosaic virus Y) CC-LRR, which recognizes Cucumber Mosaic Virus strain Y (CMV-Y), and RPP8 (Resistance to Hyaloperonospora arabidopsidis isolate Emco5) CC-LRR which is allelic to RCY1 but recognizes Hyaloperonospora arabidopsidis, showed that the RCY1 LRR domain is the primary determinant for detection of CMV-Y [13]. A similar observation was made in the case of Tm-22 CC-NLR and its ability to recognize the ToMV MP. Tm-2 and its allele Tm-22 differ by only 4 amino acids but elicit differential response to two mutant strains of ToMV. A single amino acid in the LRR is responsible for this differential recognition [14]. Although the above examples suggest that LRR may be involved in viral recognition, it is becoming evident that recognition of elicitors and subsequent activation of NLR is a multistep process involving multiple NLR domains. For example, some NLRs like Rx recognize viruses through complex interactions between the LRR and N-terminal domains. Programmed cell death (PCD) induction against PVX-CP can be reconstituted in a transient assay when CC-NB and LRR or CC and NB-LRR fragments of Rx are co-expressed [15], while the presence of the viral CP disrupts these intramolecular associations. In planta transient assays have shown that the Arabidopsis CC-NLR HRT (Hypersensitive Response to TCV) which detects Turnip Crinkle Virus (TCV), also exhibits intra-molecular interactions [15]. Together these findings suggest that intra-molecular associations between the different domains of some NLRs keep them in an inactive, resting state. The elicitor disrupts these associations leading to recognition of the pathogen and subsequent activation of NLR signaling (Figure 1).
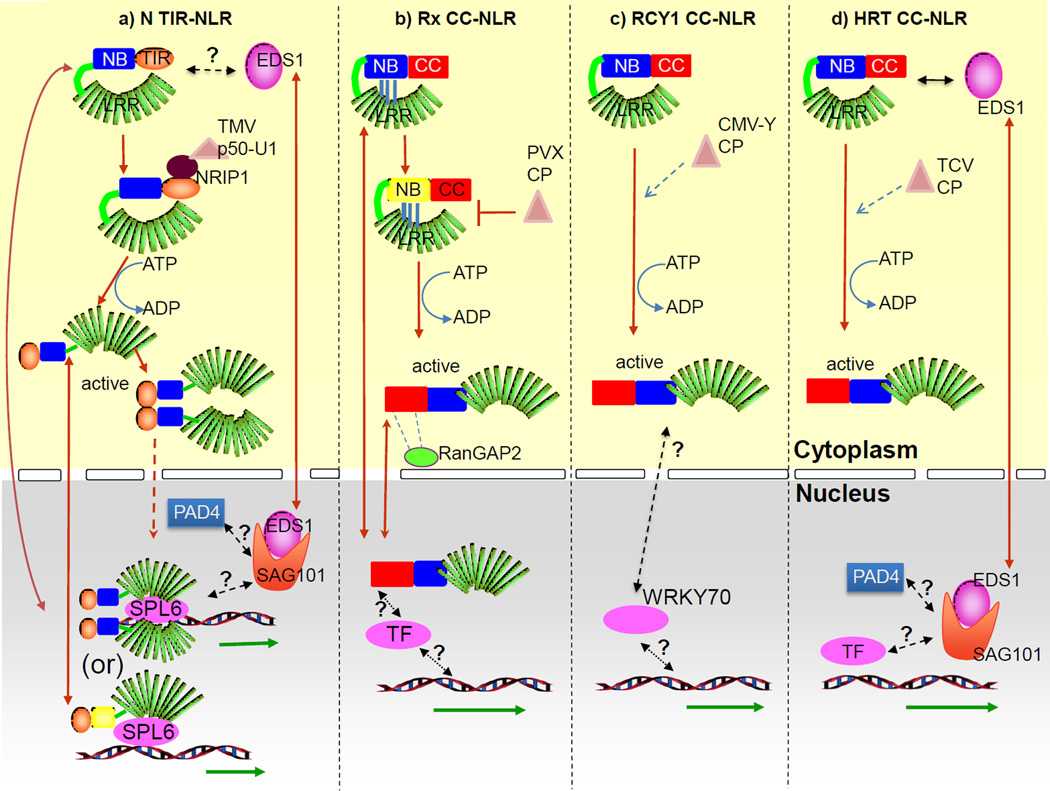
Figure 1. Activation and regulation of nuclear signaling by some viral NLRs.
In the absence of the viral elicitor, NLRs are in an inactive state. In the case of Rx NLR, the association between LRR and CC-NB domain achieves this. For others like N, the mode of inhibition is still unknown. Different NLRs sense the viral elicitor by different mechanisms but in all cases, it is shown or predicted that ATP hydrolysis at the NB domain, shifts its conformation into an open, signaling competent state. In this state the NLR can signal for the activation of defense gene transcription (green arrows) in the nucleus. This is achieved in different ways.
a) N is a nucleo-cytoplasmic protein. In the presence of the viral elicitor TMV p50-U1, N associates with the host protein NRIP1-p50 complex in the cytoplasm. Subsequent ATP hydrolysis and homotypic associations between TIR domains lead to N activation. Activated N (either as a monomer or a dimer/oligomer) associates with the SPL6 transcription factor in the nucleus to modulate defense gene transcription Nucleo-cytoplasmic immune modulator EDS1 is required for N function. EDS1 is known to associate with PAD4 and SAG101 in the nucleus. However, EDS1 interaction with N in the cytoplasm and/or its interaction with SPL6 in the nucleus in unknown.
b) Rx is also a nucleo-cytoplasmic protein whose localization is regulated by the RanGAP2. The CP of PVX disrupts Rx intramolecular associations thus initiating activation. The activated Rx may associate or regulate the function of unknown transcription factors/transcriptional modulators in the nucleus to promote defense gene transcription.
c) RCY1 may regulate nuclear gene transcription by modulating its association with the WRKY70 transcription factor. The location and dynamics of this association are unknown.
d) Activated HRT may regulate transcription by modulating the activity of immune modulators like EDS1 (with which it associates) or SAG101 and PAD4 which are required for HRT–mediated immunity.
Activation of the viral NLRs following elicitor recognition
NLR activation causes dramatic changes in the plant cell that ultimately lead to PCD at the site of infection. It is therefore imperative to place functional constraints on the NLR to avoid auto-activation in the absence of a pathogen. At the same time, the NLR also needs to display a hair-trigger response by sensing the presence of a pathogen. NLRs are predicted to achieve this because they can function as conformational switches: Their structural conformation is different in the absence and in the presence of the elicitor. Different NLRs may achieve this by mechanisms that show subtle variations.
For Rx CC-NLR, as mentioned above, intramolecular association between the LRR and the NB domains retain it in an inactive state. Association of the intermediary host proteins with the elicitor or direct association of the elicitor with the sensory domain(s) of the NLR leads to disruptions in the inactive conformation. For Rx, this disassociation is predicted to be a part of the activation mechanism, exposing the NB domain for an exchange of ADP for ATP leading to a conformational change that switches the NLR into a signaling competent ‘ON’ state (Figure 1). This may explain why overexpression of just the NB domain (which is no longer under the negative constraint of LRR) of Rx can cause elicitor independent PCD in transient assays [16]. Furthermore, mutations within the conserved P-loop of the NB domain inactivates most tested plant NLRs to date [17].
Analyses of chimeric exchanges between Rx and its homolog Gpa2 (which does not recognize PVX-CP) identified a minimal region of Rx required for controlled activation. The NB domain possesses three subunits – the NB, ARC1 and ARC2 that together form the NB pocket [18]. An acidic 68 amino acid loop within the ARC2 domain and a basic patch within the first LRR repeat are the critical interaction partners [19]. Furthermore, two closely spaced amino acids within the ARC2 domain can distinguish between auto-activation and elicitor-triggered activation. In-silico generated 3D-models of the Rx NB and LRR domains have suggested that the amino acids within the ARC2 domain and the first LRR are solvent exposed, strengthening the theory that they are involved in protein interaction. This analysis also predicted that the complementary-charged nature of the patches in these domains could promote rapid association and disassociation in the presence of the elicitor thus showing the potential to undergo multiple rounds of activation. These in-silico analyses predict structural determinants that co-ordinate NLR activation. Its tempting also to draw broad comparisons to the mammalian NLRC4 whose structure at the resting state was recently resolved [20]. Here too the LRR domain was found to associate at multiple points with the NB-ARC domain sterically hindering access of the NB domain for additional interactions.
For the N TIR-NLR, intra-molecular interactions have not been detected in vivo [21]. These interactions are either too weak to be detected or unknown protein(s) may associate with N to retain it in an inactive state. However, it is clear that N activation is also a multi-step process [6, 22]. Association of p50 with the N TIR domain via NRIP1 is not sufficient to activate N since the non-elicitor p50-Ob can also associate with N. Furthermore, functionally inactive NB domain mutant of N can still associate with both the eliciting and non-eliciting p50 [22]. Upon p50-U1 recognition, N undergoes homotypic dimerization/oligomerization via the TIR domain (Figure 1). Oligomerization occurs only during an active immune response and requires a functional NB domain [21]. The importance of homotypic and heterotypic TIR domain interactions has recently been reported for the TIR-NLRs RPS4 (Resistance to Pseudomonas syringae 4) and RRS1 (Resistance to Ralstonia solanacearum 1) [23]. These Arabidopsis NLRs recognize the effectors AvrRps4 from the bacteria P. syringae pv. pisi and PopP2 from the fungal pathogen R. solanacearum. Elegant structure-function studies show that TIR domain interactions between RPS4 and RRS1 keep the two proteins in an inactive pre-immune complex. Effector dependent reconfigurations in this complex result in stronger homotypic associations between the TIR domains of RPS4 which lead to the formation of an active complex. Self association of NLRs is reminiscent of the mammalian NODs that self-associate following PAMP recognition and this step is a prerequisite for the formation of functional signaling complexes [24]. For N, association of eliciting p50 (via NRIP1) could uniquely alter its structure to induce a conformational change that require ATP binding and/or hydrolysis. Although N is not fully activated, this step primes N for further interaction/signaling events such as homo-dimerization through TIR domain. It is possible that self-association enables the TIR domains to engage in further downstream signaling events that then lead to complete activation of N. For a few NLRs, including N, these downstream activities include direct involvement in the regulation of defense gene transcription in the nucleus.
Potential nuclear functions of NLRs
Emerging evidence indicates that some plant NLRs and immune modulators could participate in subcellular compartment specific function during immune signaling [2, 3]. Both Rx and N recognize their cognate viral elicitors in the cytoplasm but nuclear function is essential for activation of immune signaling [10, 25, 26]. However, neither receptor possesses a known Nuclear Localization Signal (NLS). For Rx, its association with RanGAP2 regulates its nucleo-cytoplasmic accumulation, with RanGAP2 primarily serving as a cytoplasmic retention protein while the CC domain is implicated in promoting nuclear localization along with the requirement of ATP binding (Figure 1) [10]. For N, no single domain has been implicated in nuclear localization [25]. The SGT1 co-chaperone positively influences nuclear accumulation of Rx and N [26, 27]. SGT1’s nucleo-cytoplasmic localization is in turn dynamically regulated by its phosphorylation status [27].
What is the function of NLRs in the nucleus? Evidence indicates that NLRs may directly control defense transcriptome reprogramming by associating with and regulating the activity of transcription factors [28]. N associates with the Squamosa Promoter Binding Protein Like 6 (SPL6) transcription factor in the nucleus only during an active immune response [22] (Figure 1). This association is essential for promoting complete resistance to TMV which suggests that SPL6 is a positive regulator of viral immunity. SPL6 is also required for RPS4 TIR-NLR that mediates resistance to P. syringae effector avrRps4 indicating an evolutionarily conserved role for SPL6 in defense. Similarly, recent studies indicate that RCY1 CC-NB domain, but not the full length protein, interacts with WRKY70 transcription factor [29]. wrky70 mutant inhibits CMV-Y viral replication but not PCD induction. The cellular location of RCY1-WRKY70 association and its dynamics prior to or after viral infection are as yet unknown. Also the subcellular localization of RCY1 has not yet been determined. These are required to establish the precise role of WRKY70-RCY1 association (Figure 1).
Non-nuclear localized CC-NLRs like HRT can also control nuclear function indirectly by associating with and regulating the function of important nucleo-cytoplasmic immune modulators like Enhanced Disease Susceptibility1 (EDS1) [30]. EDS1, along with nucleocytoplasmic localized Phytoalexin Deficient 4 (PAD4) and nuclear Senescence Associated Gene 101 (SAG101) form secondary and possibly ternary complexes that regulate the activation of a subset of defense responsive genes [31, 32]. Analysis of genetic mutants has shown that all of these genes are required for HRT-mediated defense against TCV [30] (Figure 1). Interestingly, EDS1 is also required for the function of N TIR-NLR [33, 34]. However, association between N and EDS1 or PAD4 and SAG101 have not been reported.
Crosstalk between NLR-mediated immunity and the antiviral RNAi pathway
Recent studies have discovered a potential role for the RNAi pathway in regulating viral NLR activity. In particular, a role of micro RNAs (miRNA) and phased, small, secondary interfering RNA (phasiRNA) has been shown [35]. miRNAs are short non-coding RNAs that regulate gene expression by binding to the complementary sequences in target mRNA leading to the silencing of the transcript or inhibition of translation. Some of the 22nt miRNAs are capable of triggering the synthesis of secondary phasiRNA from either non-coding or protein coding genes. This occurs when the initiator miRNA is acted upon by RNA dependent RNA polymerase 6 (RDR6) to synthesize a dsRNA which is then processed by Dicer-Like 4 (DCL4) to produce phasiRNAs [36]. In tobacco, microRNA 6019 (miR6019) and miR6020 have sequence complementarity to conserved residues that code for the N TIR domain and induce specific cleavage of N and its homologs [35]. The miR6019 also triggers the production of phasiRNA from the N coding sequence. Overexpression of the miRNAs substantially reduced N transcripts and also suppressed N-mediated resistance against TMV thus confirming a functional importance to their presence.
Networks of highly redundant miRNAs belonging to the miRN482/2118 superfamily could potentially target a large number of NLRs in Solanaceae species [37]. These 22 nt miRNAs target highly conserved sequences within multiple predicted NLRs and also trigger the production of phasiRNAs [35, 38]. Target homology analysis indicates that the highly conserved P-loop sequences within the NB domain of CC-NLRs are the main targets [35, 38].
Why do NLRs need to be regulated by miRNAs? The activity of NLRs comes with a known fitness cost so it is beneficial for the plant to keep their transcript levels low in the absence of pathogen infection. Upon infection, many pathogens and especially viruses produce potent suppressors of RNAi. These silencing suppressors also down regulate production or activity of the miR482/2118 family thereby inadvertently turning up the transcript levels of the targeted NLRs. Indeed, tomato plants infected with TCV (which has a potent silencing suppressor) and two other viruses showed significant reduction in miR482 levels and concomitant increase in the corresponding target CC-NLRs [37]. It therefore appears that the miR482/2118 family can finely tune the levels of corresponding target NLRs by maximizing their production only during times of need by exploiting the viral silencing suppressor activity which evolved to suppress plant defense. This is an excellent example of an evolving defense-counter defense strategy between pathogens and their hosts.
Another example has been elaborated with the Tobacco Bushy Stunt Virus p19 silencing suppressor [39]. In certain Nicotiana species, p19 activates an enhanced form of PCD termed Extreme Resistance (ER). This ER is dependent on the siRNA binding ability of p19. While the exact mechanism by which ER is triggered still remains elusive it has been proposed that the sequestration of miR482-like family members (and their related phasiRNA) by p19 might enhance the abundance of several NLRs, some of which may trigger ER. Alternately, siRNA bound p19 dimers can themselves serve as elicitors for an unknown NLR which monitors the status of RNAi pathway components.
Recent cloning of tomato Ty-1 and Ty-3 resistance genes against Tomato Yellow Leaf Curl Virus (TYLCV) provides another link between the RNAi pathway and virus resistance pathways. The resistance phenotype provided by Ty-1 and Ty-3 is atypical in that they appear to provide tolerance to TYLCV rather than immunity. Surprisingly, Ty-1 and Ty-3 encode for an RNA dependent RNA polymerase (RDR) belonging to the RDRγ class [40]. While a specific function has not been ascribed to this class of RDRs, RDR6 from the RDRα sub family has a well characterized role in the biogenesis of virus specific siRNAs [41]. Therefore it is tempting to speculate that Ty-1 and Ty-3 gene products may be involved in the biogenesis of dsRNA against TYLCV. RDR6, DCL4, and its associated interaction partner Double stranded RNA binding protein 4 (DRBP4) have been shown to be required for successful activation of resistance to TCV mediated by the HRT CC-NLR [42].
Conclusions and perspectives
Since the cloning of the first plant TIR-NLR that provided immunity against a plant virus [5], we have gained many insights into how plant NLRs recognize elicitors. However, many of the mechanistic details and post-effector association events that lead to activation of NLRs are still not well understood. One of the primary reasons for this is the lack of structural information as purifying full-length NLRs is technically challenging. Recently, crystal structures for plant NLR TIR and CC domains (including Rx) have been reported [9, 23, 43, 44]. These are beginning to shed light on the structural similarities between different plant NLRs. Similar structural studies, especially of NLRs in complex with their elicitors and/or intermediary accessory proteins will be invaluable in deciphering the mechanism of NLR activation. Continued research is also needed to better understand the spatial distribution, subcellular localization, and dynamics of NLRs during the immune response. Eventually such studies will accelerate the development of artificially engineered NLRs that can provide durable, broad spectrum resistance against economically important viral diseases [45].
Highlights.
NLRs are pattern recognition receptors that mediate immunity against pathogens
Intra-molecular associations retain some viral NLRs in an inactive state.
NLR activation is a multistep process. Pathogen recognition is just the first step.
Some NLRs associate with nuclear immune modulators to regulate transcription.
Components of the RNAi pathway regulate NLR activated immunity.
Acknowledgments
We thank Andrew Hayward for critical reading and thoughtful discussions of the manuscript. Innate immunity research in the Dinesh-Kumar laboratory is supported by NIH-GM097587 and University of California Davis Research Investments in Science and Engineering (UCD-RISE).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and recommended reading
Papers of particular interest, published within the period of review, have been highlighted as:
*of special interest
**of outstanding interest
- 1.Pumplin N, Voinnet O. RNA silencing suppression by plant pathogens: defence, counter-defence and counter-counter-defence. Nat Rev Microbiol. 2013;11:745–760. doi: 10.1038/nrmicro3120. [DOI] [PubMed] [Google Scholar]
- 2.Bonardi V, Cherkis K, Nishimura MT, Dangl JL. A new eye on NLR proteins: focused on clarity or diffused by complexity? Curr Opin Immunol. 2012;24:41–50. doi: 10.1016/j.coi.2011.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Qi D, Innes RW. Recent Advances in Plant NLR Structure, Function, Localization, and Signaling. Front Immunol. 2013;4:348. doi: 10.3389/fimmu.2013.00348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Iwasaki A. A virological view of innate immune recognition. Annu Rev Microbiol. 2012;66:177–196. doi: 10.1146/annurev-micro-092611-150203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Whitham S, Dinesh-Kumar SP, Choi D, Hehl R, Corr C, Baker B. The product of the tobacco mosaic virus resistance gene N: similarity to toll and the interleukin-1 receptor. Cell. 1994;78:1101–1115. doi: 10.1016/0092-8674(94)90283-6. [DOI] [PubMed] [Google Scholar]
- 6.Caplan JL, Mamillapalli P, Burch-Smith TM, Czymmek K, Dinesh-Kumar SP. Chloroplastic protein NRIP1 mediates innate immune receptor recognition of a viral effector. Cell. 2008;132:449–462. doi: 10.1016/j.cell.2007.12.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ueda H, Yamaguchi Y, Sano H. Direct interaction between the tobacco mosaic virus helicase domain and the ATP-bound resistance protein, N factor during the hypersensitive response in tobacco plants. Plant Mol Biol. 2006;61:31–45. doi: 10.1007/s11103-005-5817-8. [DOI] [PubMed] [Google Scholar]
- 8.Padgett HS, Beachy RN. Analysis of a tobacco mosaic virus strain capable of overcoming N gene-mediated resistance. Plant Cell. 1993;5:577–586. doi: 10.1105/tpc.5.5.577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Hao W, Collier SM, Moffett P, Chai J. Structural Basis for the Interaction between the Potato Virus X Resistance Protein (Rx) and Its Cofactor Ran GTPase-activating Protein 2 (RanGAP2) J Biol Chem. 2013;288:35868–35876. doi: 10.1074/jbc.M113.517417.. Report the crystal structure of the CC domain of Rx complexed with the C terminal region of its cofactor RanGAP2. Structure shows that association between the two proteins is strengthened via hydrophobic bonds. Rx CC structure shows distinct differences with respect to the previously identified structure of a CC domain from MLA10 NLR.
- 10.Tameling WI, Nooijen C, Ludwig N, Boter M, Slootweg E, Goverse A, Shirasu K, Joosten MH. RanGAP2 mediates nucleocytoplasmic partitioning of the NB-LRR immune receptor Rx in the Solanaceae, thereby dictating Rx function. Plant Cell. 2010;22:4176–4194. doi: 10.1105/tpc.110.077461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Du Y, Zhao J, Chen T, Liu Q, Zhang H, Wang Y, Hong Y, Xiao F, Zhang L, Shen Q, et al. Type I J-domain NbMIP1 proteins are required for both Tobacco mosaic virus infection and plant innate immunity. PLoS Pathog. 2013;9:e1003659. doi: 10.1371/journal.ppat.1003659.. The authors identify NbMIP proteins that associate with both the Tm-22 CC-NLR and its viral elicitor ToMV-MP. MIPs are co-chaperons that are required for both viral infectivity as well as NLR-mediated immunity hinting at the possibility that they may be indirectly involved in activation of Tm-22.
- 12.Ravensdale M, Nemri A, Thrall PH, Ellis JG, Dodds PN. Co-evolutionary interactions between host resistance and pathogen effector genes in flax rust disease. Mol Plant Pathol. 2011;12:93–102. doi: 10.1111/j.1364-3703.2010.00657.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Takahashi H, Shoji H, Ando S, Kanayama Y, Kusano T, Takeshita M, Suzuki M, Masuta C. RCY1-mediated resistance to Cucumber mosaic virus is regulated by LRR domain-mediated interaction with CMV(Y) following degradation of RCY1. Mol Plant Microbe Interact. 2012;25:1171–1185. doi: 10.1094/MPMI-04-12-0076-R. [DOI] [PubMed] [Google Scholar]
- 14.Kobayashi M, Yamamoto-Katou A, Katou S, Hirai K, Meshi T, Ohashi Y, Mitsuhara I. Identification of an amino acid residue required for differential recognition of a viral movement protein by the Tomato mosaic virus resistance gene Tm-2(2) J Plant Physiol. 2011;168:1142–1145. doi: 10.1016/j.jplph.2011.01.002. [DOI] [PubMed] [Google Scholar]
- 15.Rairdan GJ, Moffett P. Distinct domains in the ARC region of the potato resistance protein Rx mediate LRR binding and inhibition of activation. Plant Cell. 2006;18:2082–2093. doi: 10.1105/tpc.106.042747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Rairdan GJ, Collier SM, Sacco MA, Baldwin TT, Boettrich T, Moffett P. The coiled-coil and nucleotide binding domains of the Potato Rx disease resistance protein function in pathogen recognition and signaling. Plant Cell. 2008;20:739–751. doi: 10.1105/tpc.107.056036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Takken FL, Albrecht M, Tameling WI. Resistance proteins: molecular switches of plant defence. Curr Opin Plant Biol. 2006;9:383–390. doi: 10.1016/j.pbi.2006.05.009. [DOI] [PubMed] [Google Scholar]
- 18.Takken FL, Goverse A. How to build a pathogen detector: structural basis of NB-LRR function. Curr Opin Plant Biol. 2012;15:375–384. doi: 10.1016/j.pbi.2012.05.001. [DOI] [PubMed] [Google Scholar]
- 19. Slootweg EJ, Spiridon LN, Roosien J, Butterbach P, Pomp R, Westerhof L, Wilbers R, Bakker E, Bakker J, Petrescu AJ, et al. Structural determinants at the interface of the ARC2 and leucine-rich repeat domains control the activation of the plant immune receptors Rx1 and Gpa2. Plant Physiol. 2013;162:1510–1528. doi: 10.1104/pp.113.218842.. The authors characterize the molecular and structural determinants involved in co-operation between the NB-ARC and LRR domains. They identify the minimal fragments within these regions required for the proper activation of Rx. They present a mechanistic model on how Rx may be activated.
- 20. Hu Z, Yan C, Liu P, Huang Z, Ma R, Zhang C, Wang R, Zhang Y, Martinon F, Miao D, et al. Crystal structure of NLRC4 reveals its autoinhibition mechanism. Science. 2013;341:172–175. doi: 10.1126/science.1236381.. Reports the crystal structure of a mammalian NLR family member NLRC4. Reveals the domains that are critical for interaction with ADP and keeps NLRC4 in its inactive state. Also identifies the domains that promote oligomerization during activation.
- 21.Mestre P, Baulcombe DC. Elicitor-mediated oligomerization of the tobacco N disease resistance protein. Plant Cell. 2006;18:491–501. doi: 10.1105/tpc.105.037234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Padmanabhan MS, Ma S, Burch-Smith TM, Czymmek K, Huijser P, Dinesh-Kumar SP. Novel positive regulatory role for the SPL6 transcription factor in the N TIR-NB-LRR receptor-mediated plant innate immunity. PLoS Pathog. 2013;9:e1003235. doi: 10.1371/journal.ppat.1003235.. Identify and characterize an association between nuclear-localized N TIR-NLR and the transcription factor SPL6. Authors show that the two proteins associate only during an active immune response. SPL6 is a positive regulator of defense gene transcription. This is the first report of an SPL family member having a role in innate immunity.
- 23. Williams SJ, Sohn KH, Wan L, Bernoux M, Sarris PF, Segonzac C, Ve T, Ma Y, Saucet SB, Ericsson DJ, et al. Structural basis for assembly and function of a heterodimeric plant immune receptor. Science. 2014;344:299–303. doi: 10.1126/science.1247357.. Structure-function studies on the NLR pair, RPS4 and RRS1 reveal novel and essential roles for TIR-TIR interactions in defense. Heterodimerization between RRS1 and RPS4 prior to effector recognition helps to keep RPS4 in an inhibited state. Following effector recognition, homodimerization of RPS4 TIR domains switches it to a signaling competent state.
- 24.Philpott DJ, Sorbara MT, Robertson SJ, Croitoru K, Girardin SE. NOD proteins: regulators of inflammation in health and disease. Nat Rev Immunol. 2014;14:9–23. doi: 10.1038/nri3565. [DOI] [PubMed] [Google Scholar]
- 25.Burch-Smith TM, Schiff M, Caplan JL, Tsao J, Czymmek K, Dinesh-Kumar SP. A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol. 2007;5:e68. doi: 10.1371/journal.pbio.0050068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Slootweg E, Roosien J, Spiridon LN, Petrescu AJ, Tameling W, Joosten M, Pomp R, van Schaik C, Dees R, Borst JW, et al. Nucleocytoplasmic distribution is required for activation of resistance by the potato NB-LRR receptor Rx1 and is balanced by its functional domains. Plant Cell. 2010;22:4195–4215. doi: 10.1105/tpc.110.077537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Hoser R, Zurczak M, Lichocka M, Zuzga S, Dadlez M, Samuel MA, Ellis BE, Stuttmann J, Parker JE, Hennig J, et al. Nucleocytoplasmic partitioning of tobacco N receptor is modulated by SGT1. New Phytol. 2013 doi: 10.1111/nph.12347. [DOI] [PubMed] [Google Scholar]
- 28.Bhattacharjee S, Garner CM, Gassmann W. New clues in the nucleus: transcriptional reprogramming in effector-triggered immunity. Front Plant Sci. 2013;4:364. doi: 10.3389/fpls.2013.00364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ando S, Obinata A, Takahashi H. WRKY70 interacting with RCY1 disease resistance protein is required for resistance to Cucumber mosaic virus in Arabidopsis thaliana. Physiol Mol Plant Pathol. 2014;85:8–14. [Google Scholar]
- 30.Zhu S, Jeong RD, Venugopal SC, Lapchyk L, Navarre D, Kachroo A, Kachroo P. SAG101 forms a ternary complex with EDS1 and PAD4 and is required for resistance signaling against turnip crinkle virus. PLoS Pathog. 2011;7:e1002318. doi: 10.1371/journal.ppat.1002318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Garcia AV, Blanvillain-Baufume S, Huibers RP, Wiermer M, Li G, Gobbato E, Rietz S, Parker JE. Balanced nuclear and cytoplasmic activities of EDS1 are required for a complete plant innate immune response. PLoS Pathog. 2010;6:e1000970. doi: 10.1371/journal.ppat.1000970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wagner S, Stuttmann J, Rietz S, Guerois R, Brunstein E, Bautor J, Niefind K, Parker JE. Structural Basis for Signaling by Exclusive EDS1 Heteromeric Complexes with SAG101 or PAD4 in Plant Innate Immunity. Cell Host Microbe. 2013;14:619–630. doi: 10.1016/j.chom.2013.11.006. [DOI] [PubMed] [Google Scholar]
- 33.Peart JR, Cook G, Feys BJ, Parker JE, Baulcombe DC. An EDS1 orthologue is required for N-mediated resistance against tobacco mosaic virus. Plant J. 2002;29:569–579. doi: 10.1046/j.1365-313x.2002.029005569.x. [DOI] [PubMed] [Google Scholar]
- 34.Liu Y, Schiff M, Marathe R, Dinesh-Kumar SP. Tobacco Rar1, EDS1 and NPR1/NIM1 like genes are required for N-mediated resistance to tobacco mosaic virus. Plant J. 2002;30:415–429. doi: 10.1046/j.1365-313x.2002.01297.x. [DOI] [PubMed] [Google Scholar]
- 35. Li F, Pignatta D, Bendix C, Brunkard JO, Cohn MM, Tung J, Sun H, Kumar P, Baker B. MicroRNA regulation of plant innate immune receptors. Proc Natl Acad Sci U S A. 2012;109:1790–1795. doi: 10.1073/pnas.1118282109.. This paper along with Shivaprasad et al demonstrates the importance of miRNAs and phasiRNAs in regulating some NLR gene family members. They identified and characterized miR6019 and miR6020 as regulators of N gene transcripts and N-mediated resistance to TMV.
- 36.Fei Q, Xia R, Meyers BC. Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell. 2013;25:2400–2415. doi: 10.1105/tpc.113.114652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Shivaprasad PV, Chen HM, Patel K, Bond DM, Santos BA, Baulcombe DC. A microRNA superfamily regulates nucleotide binding site-leucine-rich repeats and other mRNAs. Plant Cell. 2012;24:859–874. doi: 10.1105/tpc.111.095380.. This paper along with Li et al demonstrates the importance of miRNAs and phasiRNAs in regulating some NLR gene family members. They characterize the miR482 superfamily and show that miR482 mediated silencing is repressed in virus infected tissue. They propose a model where viral suppressors of RNAi relieve the negative regulation mediated by miRNAs that target NLRs thereby inducing upregulation of certain NLRs. They propose that the target NLRs can promote broad non-race specific resistance against viruses.
- 38.Zhai J, Jeong DH, De Paoli E, Park S, Rosen BD, Li Y, Gonzalez AJ, Yan Z, Kitto SL, Grusak MA, et al. MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased, trans-acting siRNAs. Genes Dev. 2011;25:2540–2553. doi: 10.1101/gad.177527.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Sansregret R, Dufour V, Langlois M, Daayf F, Dunoyer P, Voinnet O, Bouarab K. Extreme resistance as a host counter-counter defense against viral suppression of RNA silencing. PLoS Pathog. 2013;9:e1003435. doi: 10.1371/journal.ppat.1003435. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 40. Verlaan MG, Hutton SF, Ibrahem RM, Kormelink R, Visser RG, Scott JW, Edwards JD, Bai Y. The Tomato Yellow Leaf Curl Virus resistance genes Ty-1 and Ty-3 are allelic and code for DFDGD-class RNA-dependent RNA polymerases. PLoS Genet. 2013;9:e1003399. doi: 10.1371/journal.pgen.1003399.. Report on the identification of the Ty-1 and Ty-3 resistance genes which are allelic. Made the interesting discovery that rather than being NLRs, these genes code for RNA-dependent RNA polymerases (RDR) belonging to the RDRγ type. They thus discover a completely new class of resistance genes.
- 41.Wang XB, Wu Q, Ito T, Cillo F, Li WX, Chen X, Yu JL, Ding SW. RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc Natl Acad Sci U S A. 2010;107:484–489. doi: 10.1073/pnas.0904086107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zhu S, Jeong RD, Lim GH, Yu K, Wang C, Chandra-Shekara AC, Navarre D, Klessig DF, Kachroo A, Kachroo P. Double-stranded RNA-binding protein 4 is required for resistance signaling against viral and bacterial pathogens. Cell Rep. 2013;4:1168–1184. doi: 10.1016/j.celrep.2013.08.018. [DOI] [PubMed] [Google Scholar]
- 43.Bernoux M, Ve T, Williams S, Warren C, Hatters D, Valkov E, Zhang X, Ellis JG, Kobe B, Dodds PN. Structural and functional analysis of a plant resistance protein TIR domain reveals interfaces for self-association, signaling, and autoregulation. Cell Host Microbe. 2011;9:200–211. doi: 10.1016/j.chom.2011.02.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Maekawa T, Cheng W, Spiridon LN, Toller A, Lukasik E, Saijo Y, Liu P, Shen QH, Micluta MA, Somssich IE, et al. Coiled-coil domain-dependent homodimerization of intracellular barley immune receptors defines a minimal functional module for triggering cell death. Cell Host Microbe. 2011;9:187–199. doi: 10.1016/j.chom.2011.02.008. [DOI] [PubMed] [Google Scholar]
- 45.Harris CJ, Slootweg EJ, Goverse A, Baulcombe DC. Stepwise artificial evolution of a plant disease resistance gene. Proc Natl Acad Sci U S A. 2013;110:21189–21194. doi: 10.1073/pnas.1311134110. [DOI] [PMC free article] [PubMed] [Google Scholar]