Abstract
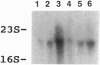
We have tested the effect of increased ribosomal fidelity on a modified version of the programmed release factor 2 (RF2) translational frameshift. In the constructs tested, the original UGA codon at the site of the shift was replaced by either of two sense codons, UGG (tryptophan), which allows a frameshift of approximately 13%, or CUG (leucine), which allows a frameshift of only approximately 2%. We confirmed the results of Curran and Yarus [Curran, J. F. & Yarus, M. (1989) J. Mol. Biol. 209, 65-77] in a wild-type ribosomal host, including a reduction of the UGG shift following induction of tRNA(Trp) from a plasmid copy of the tRNA gene. But to our surprise, in a hyperaccurate streptomycin pseudo-dependent host, the UGG frameshift increased to more than 50%. When we added a tRNA(Trp) plasmid to these cells, induction of the tRNA(Trp) gene reduced the shift back to approximately 7%. Messenger RNA levels did not vary greatly under these different induced conditions. Other increased accuracy alleles also showed increased frameshifting with UGG at the frameshift site. All increased accuracy alleles led to slower translation rates, and there appeared to be a proportionality between the extent of reduction of synthesis for the in-frame reporter and the extent of UGG frameshift for the out-of-frame reporter. There were little effects of increased accuracy on the lower level CUG frameshift. However, over-production of the cognate tRNA(1Leu) dramatically reduced even this lower level of shift, despite the fact that tRNA(1Leu) is already the most abundant isoacceptor in Escherichia coli. These results can be rationalized by following the hypothesis of Curran and Yarus as follows: with wild-type ribosomes, limited availability of tRNA(Trp) (about 1% of total tRNA) facilitates a pause at the UGG codon (due to the vacant A site), allowing increased opportunity for ribosome realignment. Excess tRNA(Trp) reduces the time the A site is vacant and thus reduces the frameshift. The slower hyperaccurate ribosomes increase the pause time and thus increase the opportunity for shifting, a process again reversed by increasing the in-frame cognate tRNA(Trp). These data provide strong support for a model in which the extent of ribosome pause time at a programmed frameshift site is a major determinant in the efficiency of the frameshift and in which tRNA availability can be a major influence on this process.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andersson D. I., Andersson S. G., Kurland C. G. Functional interactions between mutated forms of ribosomal proteins S4, S5 and S12. Biochimie. 1986 May;68(5):705–713. doi: 10.1016/s0300-9084(86)80164-x. [DOI] [PubMed] [Google Scholar]
- Andersson D. I., van Verseveld H. W., Stouthamer A. H., Kurland C. G. Suboptimal growth with hyper-accurate ribosomes. Arch Microbiol. 1986 Feb;144(1):96–101. doi: 10.1007/BF00454963. [DOI] [PubMed] [Google Scholar]
- Atkins J. F., Elseviers D., Gorini L. Low activity of -galactosidase in frameshift mutants of Escherichia coli. Proc Natl Acad Sci U S A. 1972 May;69(5):1192–1195. doi: 10.1073/pnas.69.5.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Atkins J. F., Weiss R. B., Gesteland R. F. Ribosome gymnastics--degree of difficulty 9.5, style 10.0. Cell. 1990 Aug 10;62(3):413–423. doi: 10.1016/0092-8674(90)90007-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belcourt M. F., Farabaugh P. J. Ribosomal frameshifting in the yeast retrotransposon Ty: tRNAs induce slippage on a 7 nucleotide minimal site. Cell. 1990 Jul 27;62(2):339–352. doi: 10.1016/0092-8674(90)90371-K. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bilgin N., Claesens F., Pahverk H., Ehrenberg M. Kinetic properties of Escherichia coli ribosomes with altered forms of S12. J Mol Biol. 1992 Apr 20;224(4):1011–1027. doi: 10.1016/0022-2836(92)90466-w. [DOI] [PubMed] [Google Scholar]
- Blinkowa A. L., Walker J. R. Programmed ribosomal frameshifting generates the Escherichia coli DNA polymerase III gamma subunit from within the tau subunit reading frame. Nucleic Acids Res. 1990 Apr 11;18(7):1725–1729. doi: 10.1093/nar/18.7.1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohman K., Ruusala T., Jelenc P. C., Kurland C. G. Kinetic impairment of restrictive streptomycin-resistant ribosomes. Mol Gen Genet. 1984;198(2):90–99. doi: 10.1007/BF00328706. [DOI] [PubMed] [Google Scholar]
- Bredenbeek P. J., Pachuk C. J., Noten A. F., Charité J., Luytjes W., Weiss S. R., Spaan W. J. The primary structure and expression of the second open reading frame of the polymerase gene of the coronavirus MHV-A59; a highly conserved polymerase is expressed by an efficient ribosomal frameshifting mechanism. Nucleic Acids Res. 1990 Apr 11;18(7):1825–1832. doi: 10.1093/nar/18.7.1825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brierley I., Digard P., Inglis S. C. Characterization of an efficient coronavirus ribosomal frameshifting signal: requirement for an RNA pseudoknot. Cell. 1989 May 19;57(4):537–547. doi: 10.1016/0092-8674(89)90124-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Condron B. G., Atkins J. F., Gesteland R. F. Frameshifting in gene 10 of bacteriophage T7. J Bacteriol. 1991 Nov;173(21):6998–7003. doi: 10.1128/jb.173.21.6998-7003.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coulondre C., Miller J. H. Genetic studies of the lac repressor. III. Additional correlation of mutational sites with specific amino acid residues. J Mol Biol. 1977 Dec 15;117(3):525–567. doi: 10.1016/0022-2836(77)90056-0. [DOI] [PubMed] [Google Scholar]
- Craigen W. J., Caskey C. T. Expression of peptide chain release factor 2 requires high-efficiency frameshift. Nature. 1986 Jul 17;322(6076):273–275. doi: 10.1038/322273a0. [DOI] [PubMed] [Google Scholar]
- Curran J. F., Yarus M. Base substitutions in the tRNA anticodon arm do not degrade the accuracy of reading frame maintenance. Proc Natl Acad Sci U S A. 1986 Sep;83(17):6538–6542. doi: 10.1073/pnas.83.17.6538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Curran J. F., Yarus M. Rates of aminoacyl-tRNA selection at 29 sense codons in vivo. J Mol Biol. 1989 Sep 5;209(1):65–77. doi: 10.1016/0022-2836(89)90170-8. [DOI] [PubMed] [Google Scholar]
- Curran J. F., Yarus M. Use of tRNA suppressors to probe regulation of Escherichia coli release factor 2. J Mol Biol. 1988 Sep 5;203(1):75–83. doi: 10.1016/0022-2836(88)90092-7. [DOI] [PubMed] [Google Scholar]
- Curran J. F., Yarus M. Use of tRNA suppressors to probe regulation of Escherichia coli release factor 2. J Mol Biol. 1988 Sep 5;203(1):75–83. doi: 10.1016/0022-2836(88)90092-7. [DOI] [PubMed] [Google Scholar]
- Dinman J. D., Icho T., Wickner R. B. A -1 ribosomal frameshift in a double-stranded RNA virus of yeast forms a gag-pol fusion protein. Proc Natl Acad Sci U S A. 1991 Jan 1;88(1):174–178. doi: 10.1073/pnas.88.1.174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faxén M., Kirsebom L. A., Isaksson L. A. Is efficiency of suppressor tRNAs controlled at the level of ribosomal proofreading in vivo? J Bacteriol. 1988 Aug;170(8):3756–3760. doi: 10.1128/jb.170.8.3756-3760.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flower A. M., McHenry C. S. The gamma subunit of DNA polymerase III holoenzyme of Escherichia coli is produced by ribosomal frameshifting. Proc Natl Acad Sci U S A. 1990 May;87(10):3713–3717. doi: 10.1073/pnas.87.10.3713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldman E., Holmes W. M., Hatfield G. W. Specificity of codon recognition by Escherichia coli tRNALeu isoaccepting species determined by protein synthesis in vitro directed by phage RNA. J Mol Biol. 1979 Apr 25;129(4):567–585. doi: 10.1016/0022-2836(79)90469-8. [DOI] [PubMed] [Google Scholar]
- Goldman E., Jakubowski H. Uncharged tRNA, protein synthesis, and the bacterial stringent response. Mol Microbiol. 1990 Dec;4(12):2035–2040. doi: 10.1111/j.1365-2958.1990.tb00563.x. [DOI] [PubMed] [Google Scholar]
- Gorini L. Ribosomal discrimination of tRNAs. Nat New Biol. 1971 Dec 29;234(52):261–264. doi: 10.1038/newbio234261a0. [DOI] [PubMed] [Google Scholar]
- Holmes W. M., Goldman E., Miner T. A., Hatfield G. W. Differential utilization of leucyl-tRNAs by Escherichia coli. Proc Natl Acad Sci U S A. 1977 Apr;74(4):1393–1397. doi: 10.1073/pnas.74.4.1393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacks T., Madhani H. D., Masiarz F. R., Varmus H. E. Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell. 1988 Nov 4;55(3):447–458. doi: 10.1016/0092-8674(88)90031-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker J. Errors and alternatives in reading the universal genetic code. Microbiol Rev. 1989 Sep;53(3):273–298. doi: 10.1128/mr.53.3.273-298.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petrullo L. A., Gallagher P. J., Elseviers D. The role of 2-methylthio-N6-isopentenyladenosine in readthrough and suppression of nonsense codons in Escherichia coli. Mol Gen Genet. 1983;190(2):289–294. doi: 10.1007/BF00330653. [DOI] [PubMed] [Google Scholar]
- Rojiani M. V., Jakubowski H., Goldman E. Effect of variation of charged and uncharged tRNA(Trp) levels on ppGpp synthesis in Escherichia coli. J Bacteriol. 1989 Dec;171(12):6493–6502. doi: 10.1128/jb.171.12.6493-6502.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rojiani M. V., Jakubowski H., Goldman E. Relationship between protein synthesis and concentrations of charged and uncharged tRNATrp in Escherichia coli. Proc Natl Acad Sci U S A. 1990 Feb;87(4):1511–1515. doi: 10.1073/pnas.87.4.1511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruusala T., Andersson D., Ehrenberg M., Kurland C. G. Hyper-accurate ribosomes inhibit growth. EMBO J. 1984 Nov;3(11):2575–2580. doi: 10.1002/j.1460-2075.1984.tb02176.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sipley J., Dunn J., Goldman E. Bacteriophage T7 morphogenesis and gene 10 frameshifting in Escherichia coli showing different degrees of ribosomal fidelity. Mol Gen Genet. 1991 Dec;230(3):376–384. doi: 10.1007/BF00280294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sipley J., Stassi D., Dunn J., Goldman E. Analysis of bacteriophage T7 gene 10A and frameshifted 10B proteins. Gene Expr. 1991 May;1(2):127–136. [PMC free article] [PubMed] [Google Scholar]
- Spanjaard R. A., Chen K., Walker J. R., van Duin J. Frameshift suppression at tandem AGA and AGG codons by cloned tRNA genes: assigning a codon to argU tRNA and T4 tRNA(Arg). Nucleic Acids Res. 1990 Sep 11;18(17):5031–5036. doi: 10.1093/nar/18.17.5031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spanjaard R. A., van Duin J. Translation of the sequence AGG-AGG yields 50% ribosomal frameshift. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7967–7971. doi: 10.1073/pnas.85.21.7967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z., Kornberg A. Translational frameshifting generates the gamma subunit of DNA polymerase III holoenzyme. Proc Natl Acad Sci U S A. 1990 Apr;87(7):2516–2520. doi: 10.1073/pnas.87.7.2516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z. Translational frameshifting in the Escherichia coli dnaX gene in vitro. Nucleic Acids Res. 1991 May 11;19(9):2457–2462. doi: 10.1093/nar/19.9.2457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss R. B., Dunn D. M., Atkins J. F., Gesteland R. F. Slippery runs, shifty stops, backward steps, and forward hops: -2, -1, +1, +2, +5, and +6 ribosomal frameshifting. Cold Spring Harb Symp Quant Biol. 1987;52:687–693. doi: 10.1101/sqb.1987.052.01.078. [DOI] [PubMed] [Google Scholar]
- Weiss R. B., Dunn D. M., Dahlberg A. E., Atkins J. F., Gesteland R. F. Reading frame switch caused by base-pair formation between the 3' end of 16S rRNA and the mRNA during elongation of protein synthesis in Escherichia coli. EMBO J. 1988 May;7(5):1503–1507. doi: 10.1002/j.1460-2075.1988.tb02969.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss R., Gallant J. Mechanism of ribosome frameshifting during translation of the genetic code. 1983 Mar 31-Apr 6Nature. 302(5907):389–393. doi: 10.1038/302389a0. [DOI] [PubMed] [Google Scholar]
- Xu H., Boeke J. D. Host genes that influence transposition in yeast: the abundance of a rare tRNA regulates Ty1 transposition frequency. Proc Natl Acad Sci U S A. 1990 Nov;87(21):8360–8364. doi: 10.1073/pnas.87.21.8360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yates J. L. Role of ribosomal protein S12 in discrimination of aminoacyl-tRNA. J Biol Chem. 1979 Nov 25;254(22):11550–11554. [PubMed] [Google Scholar]
- Zhang S. P., Zubay G., Goldman E. Low-usage codons in Escherichia coli, yeast, fruit fly and primates. Gene. 1991 Aug 30;105(1):61–72. doi: 10.1016/0378-1119(91)90514-c. [DOI] [PubMed] [Google Scholar]