Abstract
Nucleoside analogues, such as penciclovir, ganciclovir, acyclovir, and their fluoro-substituted derivatives, have wide utility as antivirals. Among these analogues, FHBG (18F-Fluorohydroxybutylguanine) is a well-validated PET (positron emission tomography) probe for monitoring reporter gene expression. To evaluate, whether or not imposing rigidity into the flexible side chain of FHBG 4 could also impact its interaction, with amino acid residues within the binding site of HSV1-TK (Herpes Simplex Virus-1 Thymidine Kinase), thus influencing its cytotoxic activity. Herein, the synthesis of a new fluorinated nucleoside analogue 6 (conceived via ligand-docking studies) is reported. Agent 6 demonstrates selective activity against HeLa cells stably transfected with mutant HSV1-sr39TK, and is also 47-fold more potent than FHBG.
Humans are the natural host to several herpes viruses, such as herpes simplex virus type I (HSV-1), herpes simplex virus type II (HSV-2), cytomegalovirus (CMV), Epstein Barr virus (EBV), Varicella-Zoster virus (chicken pox), and HHV-6 (roseola infantum) 1. Among these viruses, HSV-1 and HSV-2 remain dormant within the body, but upon activation, cause cold sores (vesicular lesions of the oral mucosa) 2, genital skin lesions 3, corneal infections 4, and encephalitis 5. Antiviral molecules, such as, acyclovir 1, ganciclovir 2, and penciclovir 3, are known to demonstrate potent antiherpetic activity via targeting thymidine kinase (TK) 6. Unlike the mammalian TK which is highly specific for thymidine, HSV1-TK and related mutants have broad levels of substrate specificity. This characteristic has been exploited in antiviral therapy as well as suicide gene therapy protocols involving HSV1-TK activated prodrug therapy wherein the targeted tissue is transduced with the HSV1-tk gene 7. Upon exposure to compounds, such as acyclovir 1 and ganciclovir 2, host cells expressing HSV1-TK selectively retain drug, leading to apoptosis and destruction of target tissues8. Additionally, the HSV1-tk gene has also been commonly employed as the reporter gene for imaging gene regulation and expression using positron emission tomography (PET) 9,10. Several PET reporter probes [18F]-(2’-deoxy-2’-fluoro-5-methyl-1-β-D-arabinofuranosyluracil, 18FMAU 11; [124I]-2’-deoxy-2’-fluoro-1-β-D-arabinofuranosyl-5-iodo-uracil, 124I-FIAU 10; 18FHBG 12-14; and [18F]-9-[(3-fluoro-1-hydroxy-2-propoxy)methyl]guanine, 18FHPG 15) have also been evaluated as molecular imaging probes. Our laboratories and others have demonstrated that 18FHBG is selectively accumulated in HeLa cells transfected with HSV1-sr39TK compared with their WT type counterparts 13,16,17. The effectiveness of these molecules to either act as a substrate (mono- or diphosphates) or an inhibitor (triphosphates for DNA polymerase) is likely impacted by the ability of the side chain to mimic the interaction of the glycosyl entity of the natural substrate with the enzyme. It is noteworthy that flexibility in the side chain could either allow these molecules to adopt an unfavorable conformation in solution or represent a population of potential low-energy rotamers. Consequently, imposing restrictions into the side chain to achieve a conformation optimal for interaction with the targeted enzyme could lead to enhanced biological-activity and target-specificity. While acyclovir 1, ganciclovir 2, and penciclovir 3, and FHBG 4 possess flexible side chains, the cyclopropylpenciclovir A5021 5 (Figure 1), which incorporates a side chain containing a methylene spacer between the base and carbocyclic ring, has been shown to be 20-fold more potent than acyclovir 1 against HSV-1 18,19. Additionally, compared with acyclovir 1 and penciclovir 3, the selectivity index of 5 for HSV-1 has also been found to be superior, implying that the imposed restrictions arising from the presence of the cyclopropyl ring in the side chain leads to an optimal conformation for interaction with the enzyme 20,21. To further evaluate whether or not upon imposition of rigidity into the flexible side chain of FHBG 4 via incorporation of a fluoromethylcyclopropyl ring could also impact its efficacy, a new derivative 6 was synthesized, characterized, and evaluated for its efficacy in HeLa (WT) and HeLa cells stably transfected with HSV1-mNLS-sr39TK13. Successful execution of this strategy could also be beneficial for designing a surrogate PET reporter probe for comparative analysis of target-sensitivity and - specificity compared with 18FHBG, an agent undergoing clinical validation for monitoring the gene expression in cancer.
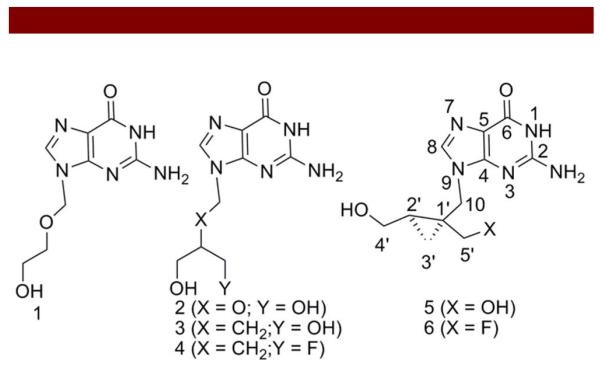
Figure 1.
Chemical structures of antiviral molecules.
We opted to substitute a fluorine atom as an isopolar mimic of the hydroxyl group into the design of 6 due to: a) fluorine is the smallest atom that closely mimics hydrogen and is capable of producing significant electronic changes with minimum steric perturbation within the overall geometry of bioactive molecules 22,23; b) fluorine can serve as an isosteric mimic of the hydroxyl group since the C-F bond length (1.39Å) is fairly close to that of C-O (1.43Å) 24; c) fluorine is also a hydrogen acceptor 25,26; and d) replacement of hydroxyl with the fluorine has been known to increase biological half-life (stability) of compounds, thus providing improved therapeutic effects 27. Finally, it has been demonstrated that the 2’-OH position of A5021 5 is preferentially phosphorylated by thymidine kinase 20.
For accomplishing the synthesis of 6, we also opted to preserve the 2’-OH position in the side chain for phosphorylation thus performed selective fluorination of the 1’-OH. Prior to performing chemical synthesis, to test the hypothesis whether or not imposing rigidity into the flexible side chain of FHBG 4 could also impact its binding characteristics with certain amino residues of the HSV1-TK, we docked both 4 and 6 (containing cyclopropyl linker, (Figure 2) into the known active site of the protein, employing parental compounds 3 and 5 as controls (Supporting Information, Figure SI 1-2).
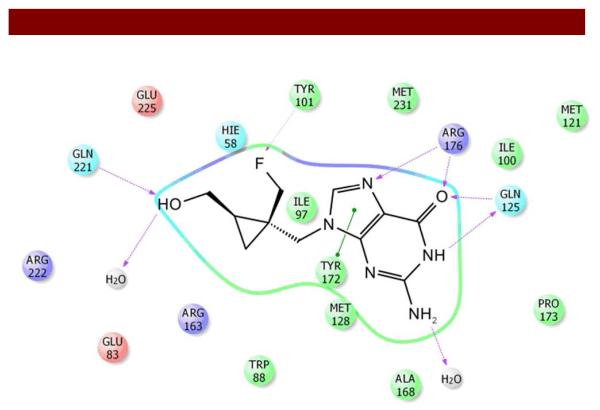
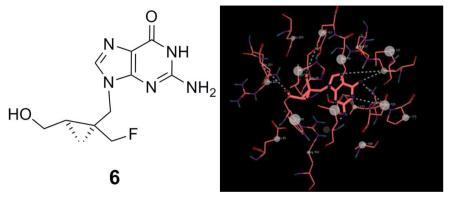
Figure 2.
Post-Docking view for the bindings of 6 to thymidine kinase (TK) showing active site residues, intermolecular hydrogen bonding, and pi-pi interaction.
To accomplish this process, numerous docking programs, such as GOLD 28, FlexX 29, DOCK 30, and Glide 31 have been extensively employed in pharmaceutical industry. We opted to use Glide for docking because it allows a search of potential binding sites of the ligand in the active site-region of the protein. Thus, the shape and properties of the binding site region could be represented by a grid using different sets of fields providing fairly accurate scoring system for the ligand (position and orientation relative to the active site, core confirmation, and rotamer-group confirmation). Following generation of the grid for the protein, compounds 3-6 were docked into the binding site of HSV1-TK using a docking option with standard precision and the rank order (GlideScore) was assigned on the basis of binding energy. Docking of compound 4 demonstrated binding of guanine with the amino acid residues consistent with patterns observed for 3 (Penciclovir, control) 32. Compared with 3 (Figure SI 1), the 2-amino group of 4 was not involved in hydrogen bonding to Gln-125. Importantly, while the fluorine atom showed interactions with Tyr-101, the hydroxyl group did not show interactions with Glu-83 rather it was hydrogen bonded to only a satellite water molecule. Furthermore, 5 (Figure SI 2; control for 6) retained overall binding characteristics visualized for 3 (Figure SI 1) while displaying additional pi-pi interactions of guanine with Tyr-172 and Tyr-101. Additionally, while 1’-OH showed binding with Glu-225 and Tyr-101, the 2’-OH indicated H-bonding to Gln-221 and a satellite water molecule. Finally and importantly, docking of 6 retained binding pattern of residues visualized for 5 with the exception of pi-pi interactions of guanine with only Tyr-172 and interaction of fluorine with Tyr-101 (Figure 2). Overall docking data from GlideScore indicated a rank order of 5, 3, 6, and 4 for preferential interaction with the binding site of HSV1-TK, thus indicating that rigidity within the flexible side chain of 3 and 4 results in a favorable orientation of 5 and 6 for mapping amino acid residues in the binding site of the protein.
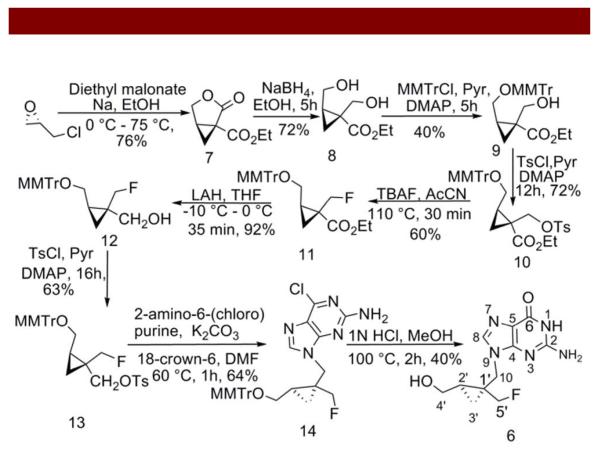
For synthesis (Scheme 1), the lactone ester 7 was obtained from (R)-(−)epichlorohydrin and further reduced using sodium borohydride to yield the diol 8 employing a known methodology, under modified conditions 18. Thereafter, the diol derivative 8 was selectively protected using methoxymethyl-tritylchloride and DMAP to obtain 9 which in turn was reacted with p-toluenesulfonyl chloride to yield 10 (Scheme 1). Further, the fluorinated side chain 11 was obtained via nucleophilic substitution by reacting tosylated precursor 10 with tetrabutyl-ammonium fluoride. Upon reduction using LAH, the ester was converted into the alcohol 12 which was further converted into tosylmethylcyclo-propyl derivative 13 using p-toluenesulfonyl chloride. Thereafter, the purine analogue 14 was obtained by alkylation of 6-chloropurine using 13 in DMF. Furthermore, the desired bioactive fluoro analogue 6 was obtained by deprotection of methoxymethyl-trityl and conversion of the purine into guanine using 1 N HCl. Finally, 6 was purified on a C-18 column using water and methanol as the gradient mixture. Fractions eluting around Rt = 22 min were collected, combined, lyophilized, resuspended in water or DMSO for analytical characterization. All proton (Figure SI 3) and carbon chemical shifts for 6 were assigned (SI: Table1) by analysis of COSY, and HMQC (Figure SI 4) as well as HMBC (Figure SI 5) spectra, respectively. Furthermore, the observed doublet attributed to C5 resonance at 84.4 ppm (Figure SI 5) and triplet assigned to F-19 signal at −214.5 ppm (Figure SI 6) indicated that fluorine was attached to a neighboring methylene carbon at C5’ position. Finally, the proposed formulation of 6 was also ascertained via HRMS (Figure SI 7) and the compound was evaluated for its cytotoxic activity in HeLa cells.
Scheme 1.
To evaluate the effect of incorporation of a fluorine on activity of A5021 5, we performed cytotoxicity assays of 6 in HeLa cells stably transfected with plasmid encoding HSV1-mNLS-sr39TK 13. For comparative analysis, penciclovir 3, FHBG 4, and A5021 5 were also evaluated under identical conditions. For determining efficacy, cells in monolayer culture in 96-well plates were exposed to penciclovir 3, FHBG 4, A5021 5, and 6 over a range of pharmacologically relevant concentrations, and cell survivals were determined by the MTS method following incubation at 37°C for 5 d in culture 33,34. For analyses, cells grown in the presence of drug vehicle alone (DMSO, 0.1%) served as control preparations and cell survival was determined as a percentage of control. Cytotoxic activity was evaluated by cell survival curves and LC50 values were determined for comparative analysis. Compared to penciclovir 3, FHBG 4 was found to be less potent (LC50: 3, HeLa HSV-1-mNLS-sr39TK, 32 nM, HeLa >> 100 μM; 4, HeLa HSV-1-mNLS-sr39TK; 3.5 μM, HeLa >> 100 μM) indicating that incorporation of fluorine into the hydroxybutyl side chain of penciclovir 3 decreased the efficacy of the molecule. Compared to 3 and 4, overall higher activities were observed in A5021 5 and 6 (LC50: 5, HeLa HSV-1-mNLS-sr39TK, 1 nM, HeLa >> 100 μM; 6, HeLa HSV-1-mNLS-sr39TK; 74 nM, HeLa, >> 100 μM, Figure 3) thus displaying potency profiles consistent with the replacement of hydroxyl group by fluorine. Importantly, derivative 6 was found to be 47-fold more potent compared to 4. Overall results suggest that incorporation of conformational rigidity within 6 dramatically enhanced its potency compared with its flexible counterpart 4, thus offering a strategic template scaffold to design and develop highly sensitive and specific reporter molecules. Finally, 6 is less hydrophobic (NMR in D2O) than its counterpart 4 (NMR in DMSO-d6) with a flexible linker, a critical characteristic that would potentially facilitate renal mode of excretion compared to both hepatobiliary and renal modes of excretion observed with the F-18 labeled counterpart of 4 (18FHBG). Further comparative analysis of radiolabeled PET reporter probes to image mutant HSV1-tk expression in vivo is under investigation.
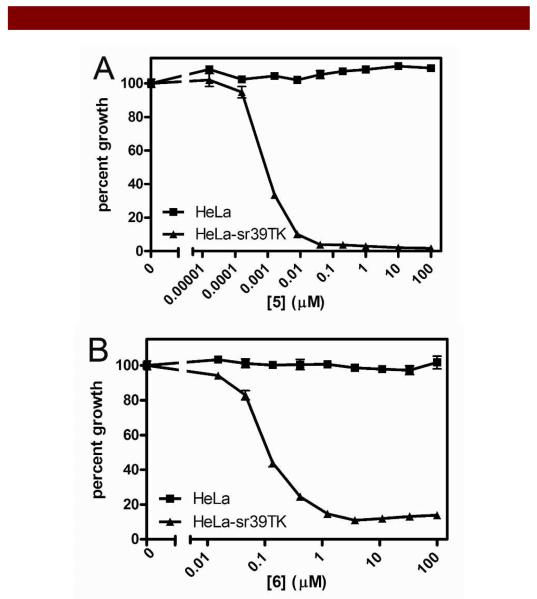
Figure 3.
Cell survival studies and LC50 determination. Survival of Hela and HeLa HSV1-mNLS-sr39TK cells grown in the presence of increasing amounts of either 5 (3A, top) or 6 (3B, bottom). Cells grown in the presence of vehicle alone served as negative control; data for cell survival in the presence of either 5 or 6 was plotted as a percent of vehicle control. LC50 (μM): 3A. 5, HeLa: >> 100 μM, HeLa HSV-1-mNLS-sr39TK: 0.00078 μM; 3B. 6, HeLa: >> 100 μM, HeLa HSV-1-mNLS-sr39TK: 0.074 μM). Each point represents the mean value of sextuplicate determinations; bars represent ± standard deviation when larger than symbol.
In summary, a new fluorinated nucleoside analogue 6 was synthesized and analytically characterized. Compared to penciclovir 3 and FHBG 4, the derivative 6 demonstrated 2-fold lower and 47-fold higher activity against sr39TK transfected HeLa cells, respectively. Furthermore, these data indicate that potency profiles for molecules were in accord with the rank order assigned via docking studies (SI Table 2). Finally, these data offer a template scaffold for developing potent antiviral molecules and the next generation of highly sensitive reporter probes for imaging HSV1-tk gene expression in vivo.
Supplementary Material
Acknowledgements
The authors thank Prof. David Piwnica-Worms, Washington University School of Medicine for his suggestions. Financial assistance for this work was provided in part by grants from the National Institutes of Health (P50 CA94056, AG033328, and AI45640).
Footnotes
Supporting Information Available.
Full experimental details, ligand-docking figures, and NMR spectra are provided. This information is available free of charge from the internet at http//:pubs.acs.org.
References
- (1).Roismann B, Sears A. In: Herpes simplex viruses and their replication In “Fundamental Virology”. Fields BN, Knipe DM, Howley PM, editors. Vol. 1996. Lippincott-Raven; New York: 1996. pp. 1043–1107. [Google Scholar]
- (2).Hull CM, Brunton S. Postgrad Med. 2010;122:1–6. doi: 10.3810/pgm.2010.09.2216. [DOI] [PubMed] [Google Scholar]
- (3).Sauerbrei A, Wutzler P. Med Microbiol Immunol. 2007;196:89–94. doi: 10.1007/s00430-006-0031-0. [DOI] [PubMed] [Google Scholar]
- (4).Satpathy G, Mishra AK, Tandon R, Sharma MK, Sharma A, Nayak N, Titiyal JS, Sharma N. Br J Ophthalmol. 2011;95:415–8. doi: 10.1136/bjo.2010.191049. [DOI] [PubMed] [Google Scholar]
- (5).Steiner I, Kennedy PG, Pachner AR. Lancet Neurol. 2007;6:1015–28. doi: 10.1016/S1474-4422(07)70267-3. [DOI] [PubMed] [Google Scholar]
- (6).Sharma V, Luker G, Piwnica-Worms D. J Mag Reson Imaging. 2002;16:336–351. doi: 10.1002/jmri.10182. [DOI] [PubMed] [Google Scholar]
- (7).Sandmair AM, Loimas S, Puranen P, Immonen A, Kossila M, Puranen M, Hurskainen H, Tyynela K, Turunen M, Vanninen R, Lehtolainen P, Paljarvi L, Johansson R, Vapalahti M, Yla-Herttuala S. Hum Gene Ther. 2000;11:2197–205. doi: 10.1089/104303400750035726. [DOI] [PubMed] [Google Scholar]
- (8).Ram Z, Culver KW, Oshiro EM, Viola JJ, DeVroom HL, Otto E, Long Z, Chiang Y, McGarrity GJ, Muul LM, Katz D, Blaese RM, Oldfield EH. Nat Med. 1997;3:1354–61. doi: 10.1038/nm1297-1354. [DOI] [PubMed] [Google Scholar]
- (9).Tjuvajev JG, Stockhammer G, Desai R, Uehara H, Watanabe K, Gansbacher B, Blasberg RG. Cancer Res. 1995;55:6126–6132. [PubMed] [Google Scholar]
- (10).Tjuvajev J, Avril N, Oku T, Sasajima T, Miyagawa T, Joshi R, Safer M, Beattie B, DiResta G, Daghighian F, Augensen F, Koutcher J, Zweit J, Humm J, Larson S, Finn R, Blasberg R. Cancer Res. 1998;58:4333–4341. [PubMed] [Google Scholar]
- (11).Conti PS, Alauddin MM, Fissekis JR, Schmall B, Watanabe KA. Nucl Med Biol. 1995;22:783–9. doi: 10.1016/0969-8051(95)00017-r. [DOI] [PubMed] [Google Scholar]
- (12).Alauddin M, Conti P. Nucl Med Biol. 1998;25:175–180. doi: 10.1016/s0969-8051(97)00160-1. [DOI] [PubMed] [Google Scholar]
- (13).Luker GD, Sharma V, Pica CM, Dahlheimer JL, Li W, Ochesky J, Ryan CE, Piwnica-Worms H, Piwnica-Worms D. Proc Natl Acad Sci U S A. 2002;99:6961–6966. doi: 10.1073/pnas.092022399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (14).Luker G, Sharma V, Piwnica-Worms D. Methods. 2003;29:110–122. doi: 10.1016/s1046-2023(02)00285-2. [DOI] [PubMed] [Google Scholar]
- (15).Alauddin MM, Shahinian A, Gordon EM, Conti PS. Mol Imaging. 2004;3:76–84. doi: 10.1162/15353500200403160. [DOI] [PubMed] [Google Scholar]
- (16).Kesarwala AH, Prior JL, Sun J, Harpstrite SE, Sharma V, Piwnica-Worms D. Mol Imaging. 2006;5:465–74. [PubMed] [Google Scholar]
- (17).Gambhir S, Barrio J, Phelps M, Iyer M, Namavari M, N S, Wu L, Green A, Bauer E, Maclaren D, Nguyen K, Berk A, Cherry D, Herschman H. Proc Natl Acad Sci (USA) 1999;96:2333–2338. doi: 10.1073/pnas.96.5.2333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (18).Sekiyama T, Hatsuya S, Tanaka Y, Uchiyama M, Ono N, Iwayama S, Oikawa M, Suzuki K, Okunishi M, Tsuji T. J Med Chem. 1998;41:1284–1298. doi: 10.1021/jm9705869. [DOI] [PubMed] [Google Scholar]
- (19).Onishi T, Matsuzawa T, Nishi S, Tsuji T. Tetrahedron Lett. 1999;40:8845–8847. [Google Scholar]
- (20).Ono N, Iwayama S, Suzuki K, Sekiyama T, Nakazawa H, Tsuji T, Okunishi M, Daikoku T, Nishiyama Y. Antimicrob Agents Chemother. 1998;42:2095–102. doi: 10.1128/aac.42.8.2095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (21).Neyts J, De Clercq E. Antiviral Res. 2001;49:121–7. doi: 10.1016/s0166-3542(00)00145-5. [DOI] [PubMed] [Google Scholar]
- (22).Rowley M, Hallett DJ, Goodacre S, Moyes C, Crawforth J, Sparey TJ, Patel S, Marwood R, Patel S, Thomas S, Hitzel L, O’Connor D, Szeto N, Castro JL, Hutson PH, MacLeod AM. J Med Chem. 2001;44:1603–14. doi: 10.1021/jm0004998. [DOI] [PubMed] [Google Scholar]
- (23).Wu YJ, Davis CD, Dworetzky S, Fitzpatrick WC, Harden D, He H, Knox RJ, Newton AE, Philip T, Polson C, Sivarao DV, Sun LQ, Tertyshnikova S, Weaver D, Yeola S, Zoeckler M, Sinz MW. J Med Chem. 2003;46:3778–81. doi: 10.1021/jm034111v. [DOI] [PubMed] [Google Scholar]
- (24).Husstedt W, Wiehle S, Stillig C, Bergander K, Grimme S, Haufe G. Eur J Org Chem. 2011;2011:355–363. [Google Scholar]
- (25).West R, Powell D, Whatley L, Lee M, Schleyer P. J Am Chem Soc. 1962;84:3221–3222. [Google Scholar]
- (26).Carosati E, Sciabola S, Cruciani G. J Med Chem. 2004;47:5114–25. doi: 10.1021/jm0498349. [DOI] [PubMed] [Google Scholar]
- (27).Dugar S, Yumibe N, Calder J, Vizzinao M, Huie K, VanHeek M, Compton D, Davis H., Jr. Biorg Med Chem Lett. 1996;6:1271–1274. [Google Scholar]
- (28).Jones G, Willett P, Glen RC, Leach AR, Taylor R. J Mol Biol. 1997;267:727–48. doi: 10.1006/jmbi.1996.0897. [DOI] [PubMed] [Google Scholar]
- (29).Rarey M, Kramer B, Lengauer T, Klebe G. J Mol Biol. 1996;261:470–89. doi: 10.1006/jmbi.1996.0477. [DOI] [PubMed] [Google Scholar]
- (30).Ewing TJ, Makino S, Skillman AG, Kuntz ID. J Comput Aided Mol Des. 2001;15:411–28. doi: 10.1023/a:1011115820450. [DOI] [PubMed] [Google Scholar]
- (31).Piccagli L, Fabbri E, Borgatti M, Bezzerri V, Mancini I, Nicolis E, Dechecchi MC, Lampronti I, Cabrini G, Gambari R. BMC Struct Biol. 2008;8:38. doi: 10.1186/1472-6807-8-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (32).Champness JN, Bennett MS, Wien F, Visse R, Summers WC, Herdewijn P, de Clerq E, Ostrowski T, Jarvest RL, Sanderson MR. Proteins. 1998;32:350–61. doi: 10.1002/(sici)1097-0134(19980815)32:3<350::aid-prot10>3.0.co;2-8. [DOI] [PubMed] [Google Scholar]
- (33).Skehan P, Storeng R, Scudiero D, Monks A, McMahon J, Vistica D, Warren JT, Bokesch H, Kenney S, Boyd MR. J. Natl. Cancer Inst. 1990;82:1107–1112. doi: 10.1093/jnci/82.13.1107. [DOI] [PubMed] [Google Scholar]
- (34).Fazlina N, Maha A, Jamal R, Zarina AL, Cheong SK, Hamidah H, Ainoon O, Zulkifli SZ, Hamidah NH. Hematology. 2007;12:33–7. doi: 10.1080/10245330600940030. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.