Abstract
Background
Identification of biological specimens is a requirement for a range of applications. Reference-free methods analyse unprocessed sequencing data without relying on prior knowledge, but generally do not scale to arbitrarily large genomes and arbitrarily large phylogenetic distances.
Results
We present Cnidaria, a practical tool for clustering genomic and transcriptomic data with no limitation on genome size or phylogenetic distances. We successfully simultaneously clustered 169 genomic and transcriptomic datasets from 4 kingdoms, achieving 100 % identification accuracy at supra-species level and 78 % accuracy at the species level.
Conclusion
CNIDARIA allows for fast, resource-efficient comparison and identification of both raw and assembled genome and transcriptome data. This can help answer both fundamental (e.g. in phylogeny, ecological diversity analysis) and practical questions (e.g. sequencing quality control, primer design).
Electronic supplementary material
The online version of this article (doi:10.1186/s12859-015-0806-7) contains supplementary material, which is available to authorized users.
Keywords: Clustering, k-mer, NGS, RNA-seq, Phylogeny, Species identification
Background
Unequivocal identification of biological specimens is a major requirement for reliable and reproducible (bio)medical research, control of intellectual property by biological patent holders, regulating the flow of biological specimen across national borders, enforcing the Nagoya protocol [1] and verifying the authenticity of claims of the biological source of products by customs authority.
Several methods for species identification have been developed based on DNA analysis, that can be classified as probe-based and nucleotide sequencing based methods. Probe-based technologies include microarrays, PCR probes, DNA fingerprinting and immunoassays involving the hybridization of DNA samples with predetermined sets of probes or primers. Such methods are cheap and allow precise identification, but may fail in cases where target DNA is not precisely matched by the probes or primers. Alternatively, nucleotide sequencing methods have been developed to increase accuracy, flexibility and throughput. These can be separated into complete or targeted approaches. Targeted identification of short and highly variable genomic regions by exome capture, Expressed Sequence Tag (EST), DNA barcoding and ribosomal DNA (rDNA) sequencing has been used for many years. Targeted DNA sequencing can be done iteratively for taxonomic identification at subspecies, accession and cultivar levels. Whole Genome Sequencing (WGS) and RNA-seq using Next Generation Sequencing (NGS) technology, examples of complete sequencing methods, have the highest information content of all methods, although its high cost has prevented it from being adopted massively. However, with the recent reduction of costs and increase in throughput, NGS starts to become more prevalent, making it a feasible alternative method for species identification. This calls for the creation of a new a set of tools to comprehensively analyse the deluge of data.
Methods for species identification based on NGS data can be separated into two main classes: reference-based and reference-free methods (reviewed in [2]). Reference-based methods usually map the sequence reads to the genome of a close relative and infer the phylogeny by aligning the observed polymorphisms. This technology requires quality control (cleaning) of the data, mapping the data to the genomic sequence of a close relative, and detection and comparison of polymorphisms [3]. In contrast, reference-free methods (RFMs) are designed to analyse unprocessed sequencing data without any previous knowledge of its identity. The data can be compared against other datasets of unknown samples, in the case of metagenomics comparing population structures [4–13] or against a panel of known species. In the latter case, it can identify a previously unknown sample, giving it an approximate position relative to the known species.
RFMs can be based on the Discrete Fourier Transform (DFT), compression and k-mers. DFT methods, such as in [14], transform nucleotide sequences into frequency statistics and compare these for species classification. Although remarkably fast, these methods are not able to store the differences between the genomes for further enquiry, yielding no insight into sequence composition. Compression based methods calculate the distance between pairs of sequences by analysing the reduction in computer memory usage when both sequences are compressed together [15]. However, compression-based methods are time and resource intensive for large genomes or large datasets.
Given a set of samples S = {s 1, s 2, …, s n}, represented either by assembled genome or transcriptome sequences (.fasta files) or by unprocessed sequencing data (.fastq files), k-mer based methods split the nucleotide sequences into all constituent substrings of length k. The presence/absence or counts of these k-mers are then used to calculate a dissimilarity D(s i, s j) between each sample pair (s i, s j), which should be minimal for samples with identical sequence composition. Several implementations of k-mer based RFMs exists, such as FFP [16], CO-PHYLOG [17], NEXTABP [18], MULTIALIGNFREE [19], KSNP [20] and SPACED WORDS/KMACS [21]. Although their underlying principles are generally useful for the analysis of large data collections, most implementations are designed for either analysis of a limited portion of the data, such as organelles or ribosomal DNA, or analysis of closely related species (such as bacteria, in metagenomics applications). As a consequence, it is not feasible to apply these tools on large amounts of whole-genome sequencing data or to analyse data that spans large phylogenetic distances. Two exceptions are the AAF [22] and REFERENCEFREE [23]. In AAF, the authors successfully clustered infra-family plant species using whole genome sequencing data; in REFERENCEFREE, it was demonstrated that it is possible to find polymorphisms shared by subsets of samples by counting and merging sets of k-mers. This latter method was effectively applied in [24] to compare 174 chloroplast genomes. As this approach is similar to ours, we compare our tool with their software.
Here we present CNIDARIA, an algorithm that employs a novel RFM strategy for species identification based on k-mer counting, designed from the ground up to allow analysis of very large collections of genome, transcriptome and raw NGS data using minimal resources. CNIDARIA improves over previous methods and overcomes their limitations on size and phylogenetic distance by allowing fast analysis of complete NGS data. To this end, it can export a database with pre-processed data so that new samples can be quickly compared against a large database of references, without the need to re-process all the data. In contrast to the method proposed by REFERENCEFREE, CNIDARIA is much faster, produces smaller files, is able to produce phylogenetic trees and uses the popular and fast k-mer count software JELLYFISH [25], allowing for easy integration in existing NGS quality checking pipelines. We demonstrate the performance and capabilities of CNIDARIA by analysing 169 samples, achieving excellent identification accuracy.
Implementation
CNIDARIA works with both raw sequencing data and assembled data, both from WGS and RNA-seq sources, in any combination. It uses k-mers extracted by JELLYFISH [25], a fast k-mer counting tool that produces a database of all k-mers present in a query sequence. The advantage of JELLYFISH over comparable software is its ability to create a sparse, compressed database in which the k-mers are ordered according to a deterministic hashing algorithm, thus allowing for the parallel and efficient merging/processing of the databases since all k-mers are in the same predictable order across different databases. CNIDARIA performs a parallel merge of the sorted sparse databases created by JELLYFISH, creating another sparse database containing, for each k-mer, its sequence and a fixed size binary array indicating its presence/absence in each sample. For parallelization, as the number of possible k-mers is 4k, where k is the k-mer size, each instance of CNIDARIA processes all k-mers corresponding to the range for p = 1, …, n, with n equal to the total number of instances. The partial databases created by each CNIDARIA instance can then simply be concatenated to create a full database containing all k-mers.
While merging the JELLYFISH databases into a single database, CNIDARIA extracts the number of k-mers shared between each pair of samples and then uses this information to calculate the distance between the samples. For that we used, by default, the Jaccard distance as described in CO-PHYLOG [17]:
Here, V ij is the number of k-mers shared by both samples s i and s j, V i is the number of k-mers in sample s i and V j is the number of k-mers in sample s j. When s i is equal to s j, the distance is 0. In our implementation, we use the number of valid k-mers in a sample, i.e. k-mers shared with at least one other sample, to filter out uninformative and possibly erroneous k-mers. Please notice that the k-mer frequency of each sample is ignored and only their presence/absence used, allowing us to compare divergent sequencing coverage, assembly statuses (from raw data to fully assembled) and sources (DNA or RNA). Besides the Jaccard metric, 70 other distance measures are also implemented in the package.
The resulting distance matrix is then processed by PYCOGENT v.1.5.3 [26], which clusters the data using Neighbour-Joining and creates a phylogenetic tree in NEWICK format. For easy visualization of the data, the summary database can also be converted to a standalone HTML page for (dynamic) display of the phylogenetic tree and plotting any statistics of the analysis directly in the tree. A graphical representation of these steps can be found in Fig. 1.
Fig. 1.
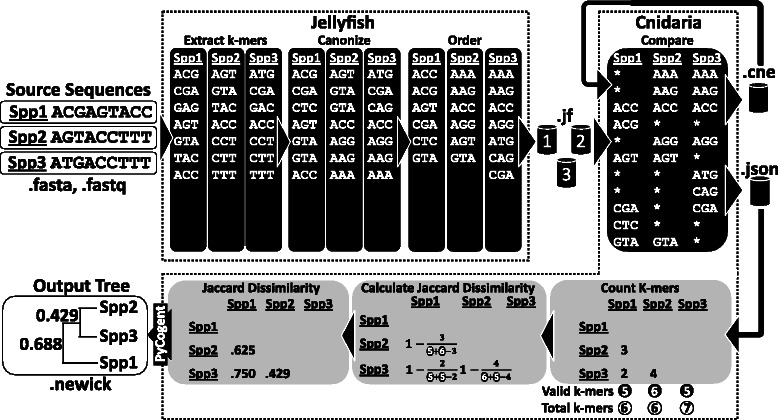
Cnidaria analysis summary. The JELLYFISH software reads each of the source sequence files (in Fasta or Fastq formats), extracts their k-mers (k = 3 in this example), canonizes them (by generating the reverse complement of each k-mer and storing only the k-mer which appears first lexicographically), orders them according to a deterministic hashing algorithm (in this example, alphabetically) and then saves each dataset in a separated database file (.jf). CNIDARIA subsequently reads these databases and compares them, side-by-side, by counting the total number of k-mers (white circles), the number of valid k-mers (k-mers shared by at least two samples, black circles) and the number of shared k-mers for each pair of samples as a matrix. Those values are exported to a Cnidaria Summary Database (CSD, a .json file) that is then used to construct a matrix of, by default Jaccard, distances between the samples (Formula 1). This dissimilarity matrix is then used for Neighbour-Joining clustering and exported as a NEWICK tree. Alternatively, Cnidaria can export a Cnidaria Complete Database (CCD, a .cne file) containing all k-mers and a linked list describing their presence/absence in the samples. This second database can be used as an input dataset together with other .cne or .jf files for new analysis
CNIDARIA can be run in two modes: Sample Analysis Mode and Database Creation Mode. Sample Analysis Mode generates a Cnidaria Summary Database (CSD) containing the total number of k-mers for each sample, the number of k-mers shared by at least two samples (valid k-mers), and the pairwise number of shared k-mers. Database Creation Mode is an order of magnitude slower than the Sample Analysis Mode but, besides generating the same CSD file, it also exports a Cnidaria Complete Database (CCD). The CCD file contains all k-mers present in the datasets analysed, stored in using a two bits per nucleotide encoding (same as JELLYFISH), and their respective presence/absence list. The CCD can be used as an input to CNIDARIA itself in both modes, allowing new samples to be directly compared against a pre-calculated larger dataset, speeding up the analysis significantly since the speed of CNIDARIA is directly correlated to the number and size of the input files. Hence, the software permits a shorter run time for the comparison of a new sample, using Sample Analysis Mode, against a large reference panel stored in a single larger CCD file.
Results and discussion
Data set
To validate the performance of CNIDARIA, we gathered a collection of 135 genomic, transcriptomic and raw NGS datasets covering a wide range of organisms. A list of all samples can be found in Additional file 1: Table S1 [27–79]. All datasets were analysed using JELLYFISH with canonized k-mers. Canonization is the process of storing the lexicographically smallest k-mer between a k-mer and its reverse complement. This step is required as both molecules are technically the same: the existence of one implies the existence of the other on the complementary DNA strand. The datasets were then split in 50 pieces and divided over 20 threads on an 80 core Intel(R) Xeon(R) CPU E7- 4850 @ 2.00 GHz machine, speeding up the analysis approximately 40 times compared to single-thread analysis on the same machine. We then created a Cnidaria Complete Database (CCD) containing all 135 samples. K-mer counts, k-mer statistics and Jaccard distances can be found in Additional file 2: Table S2, Additional file 3: Table S3 and Additional file 4: Table S4, respectively.
Identification accuracy
To verify the accuracy of the clustering of the samples, we used the 1-nearest neighbour algorithm on 30 samples for supra-species level analysis (8 genus, 7 families, 7 orders, 4 phylum and 3 kingdoms, described in Additional file 5: Table S5) and on 33 samples for species level analysis (11 species of the Solanum clade, described in Additional file 6: Table S6). The 1-nearest neighbour classifier reports the percentage of samples for which the sample with the smallest distance belongs to the same rank at each phylogenetic level (species, genus, family, order, phylum and kingdom). We report the percentage of samples correctly classified in Fig. 2 and Additional file 7: Table S7.
Fig. 2.
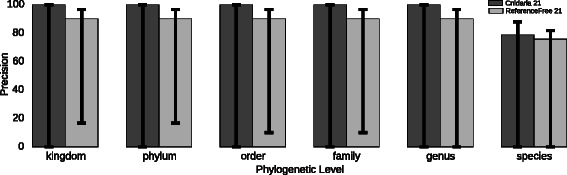
1-nearest-neighbour analysis for species and supra-species levels at each taxonomic level for CNIDARIA and REFERENCEFREE using 21-mers and Jaccard distance. Supra-species level analysis contains 30 samples (Additional file 5: Table S5) from 8 genus, 7 families, 7 orders, 4 phylum and 3 kingdoms. Species level analysis contains 33 samples (Additional file 5: Table S5) from 11 species of the Solanum clade. Classification reports the Leave-One-Out Cross-Validation error estimate (LOOCV) for 21-mers. Error bars indicate the minimum and maximum performance found across the 71 distance metrics tested
Influence of k-mer size
To investigate the influence of the k-mer size on the accuracy of the phylogenetic inference of CNIDARIA, we analysed the panel of 135 samples with k = 11, 15, 17, 21 and 31 (predefined hash sizes of 128 million, 256 million, 512 million, 1 billion and 4 billion, respectively). The resulting statistics can also be found in Additional file 1: Table S1.
Due to the low complexity of 11-mers, all possible k-mer of this size were found in the datasets and all k-mers were valid, i.e. shared by at least two samples (Table 1). This carries little clustering information and generates many zero distances (minimum dissimilarity) as shown in Additional file 8: Figure S1, Additional file 9: Figure S2, Additional file 10: Figure S3, Additional file 11: Figure S4, Additional file 12: Figure S5, Additional file 13: Figure S6 to Additional file 14: Figure S7 and Additional file 4: Table S4. Phylogenetic distances increase with k-mer size and 31-mers have most distances equal to 1, i.e. maximum dissimilarity (except for highly related species), which does not allow clustering of distant species. Therefore we chose 21-mers as the default k-mer size as it showed the best trade-off between speed and discriminating power (consistent with [23]).
Table 1.
Summary of search space per k-mer size and number of k-mers found in datasets
| k-mer size | # Canonical k-mer combinations | % of k-mers found per sample | % of k-mers found per sample, shared by at least two samples | ||
|---|---|---|---|---|---|
| Median | MAD | Median | MAD | ||
| 11-mer | 2.10 × 1006 | 100.00 % | 1.58 % | 100.00 % | 0.00 % |
| 15-mer | 5.40 × 1008 | 53.59 % | 17.07 % | 100.00 % | 0.00 % |
| 17-mer | 8.60 × 1009 | 8.90 % | 4.03 % | 98.37 % | 0.99 % |
| 21-mer | 2.20 × 1012 | 0.05 % | 0.03 % | 81.45 % | 20.55 % |
| 31-mer | 2.30 × 1018 | 0.000000061 % | 0.000000032 % | 67.05 % | 24.14 % |
The second column contains the total number of possible k-mers, calculated as (4k-mer size/2), where the division by two is due to canonization. The third column is the median and the Median Absolute Deviation (MAD) of the total number of k-mers found in the samples (Additional file 3: Table S3) divided by the number of possible k-mers, showing the percentage of combinations actually found and, consequently, the saturation of the search space; the fourth column gives the median and MAD of the percentage of valid k-mers (k-mers shared between at least two samples, Additional file 3: Table S3)
15-mers and 17-mers yielded, at the supra-species level, accuracy above 70 and 90 %, respectively, but below 75 % at the species level. Both 21- and 31-mers allowed us to correctly classify 100 % of the samples at the supra-species level and 78 % at the species level (Additional file 7: Table S7). The lower accuracy for species level classification in the tomato clade can be attributed to introgressions and sympatric speciation in tomato and is in agreement with the clustering obtained by [27], which used whole genome SNP analysis to construct trees. Compared to using 21-mers, the use of 31-mers resulted in an increased run time and disk usage without yielding a discernibly higher discriminative power. This suggests 21 is a good k-mer size for general purpose clustering. However, 31-mers are frequently used for NGS data quality checking (reviewed in [80]) and the same JELLYFISH database created for quality checking can be used for species identification.
Influence of distance measure
In order to identify the best distance measure to apply, 71 binary distances measures were implemented in CNIDARIA according to [81] and the results can be found in Additional file 7: Table S7. Some measures gave a better sensitivity than the Jaccard distance at shorter k-mer lengths, but in these cases the accuracy was below 100 %. At k = 21, Jaccard distance presented an overall high accuracy, although other methods achieved similar results. We decided to use Jaccard as the default measure due to its simplicity and equally high accuracy as other methods.
Joint analysis of DNA and RNA-seq data
Next, we expanded the 135 sample dataset (built using Database Creation Mode) with 34 extra samples, 26 genomic and 8 RNA-seq (Additional file 1: Table S1), using 21-mers and the faster Sample Analysis Mode. RNA-seq samples were added to verify whether transcriptome data would cluster with their genomic NGS counterparts, despite their small coverage of the genome length. Results are shown in Fig. 3 and Additional file 15: Figure S8. The clustering of the original 135 samples is not changed and new samples cluster correctly according to their phylogeny. The consistent clustering observed for the RNA-seq dataset illustrates the ability of CNIDARIA to use such data for accurate species identification.
Fig. 3.
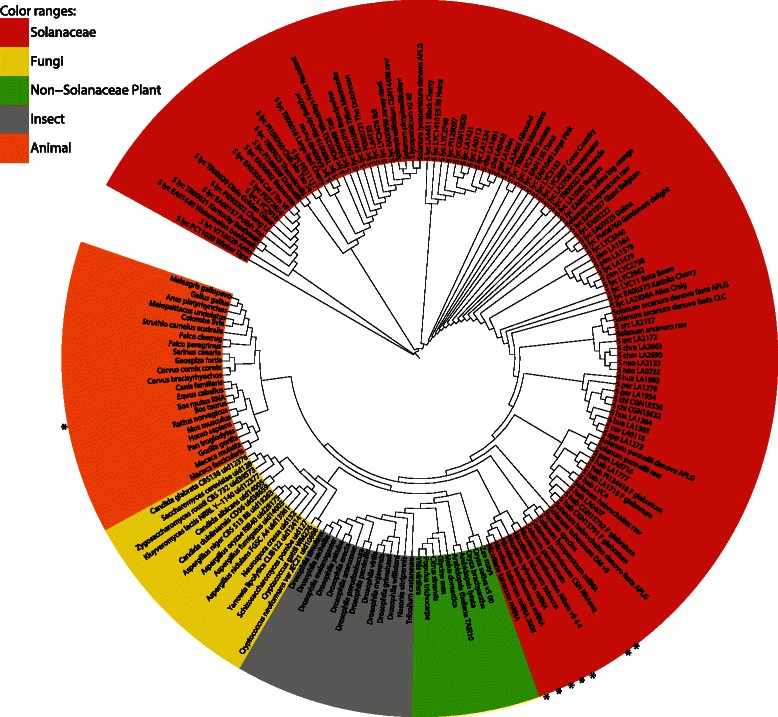
Results for the 21-mer dataset of 169 individuals using the Jaccard distance and Neighbour-Joining. The phylogenetic tree shows the clustering of the samples without displaying branch lengths (plotted using iTOL, [85]). RNAseq samples are highlighted with a * in the outer rim of the tree
Speedup by subsampling
To test the influence of data set size (and possibility of speedup) we sample 2 % of the 21-mer dataset, by analysing just 1 of the 50 pieces the data was originally split into. Additional file 16: Figure S9 shows the phylogenetic placement of species in the trees constructed using this dataset and Additional file 7: Table S7 shows the classification accuracy. The tree is indistinguishable from the one generated on the full dataset, illustrating the ability of CNIDARIA to correctly classify samples even at very low sequencing coverage. This suggests that CNIDARIA should be able to correctly cluster and identify samples using small and affordable NGS sequencing technology such as Illumina MiSeq nano runs (500 Mbp in 2 × 250 bp reads, [82]).
Comparison with REFERENCEFREE
To demonstrate the advantages of CNIDARIA, we compare it to a state-of-the-art tool called REFERENCEFREE [23]. Its latest version (1.1.3) was downloaded and run in conjunction with ABYSS [83] version 1.3.3. We run this older version rather than the latest version (1.9.0) since that was the version REFERENCEFREE was designed to work with. REFERENCEFREE was run single threaded on an Intel(R) Xeon(R) CPU E7- 4850 @ 2.00 GHz with a k-mer size of 21, a minimum frequency of 0 (i.e. using all k-mers appearing 1 or more times), no complexity filter and no sampling of k-mers. The list of shared k-mers generated was then parsed using the CNIDARIA scripts in order to generate a comparable phylogenetic tree, since REFERENCEFREE does not provide a method for phylogenetic analysis.
Using a subset of our data (41 assembled genomes, Additional file 1: Table S1) containing 40 Gbp and 20 billion k-mers, REFERENCEFREE (Additional file 1: Table S1) and JELLYFISH have a comparable speed for k-mer counting, taking 4 h to count 445 million k-mers (2 % of the total; Additional file 17: Table S8). REFERENCEFREE then took 60 % more time than CNIDARIA in single threaded Sample Analysis Mode for merging and summarizing the results (70 h vs. 44 h, respectively). Note that the databases created by CNIDARIA can be re-used in subsequent comparisons, whereas REFERENCEFREE requires all the k-mer count files to be merged again when re-run. Moreover, CNIDARIA has the important advantage of being highly parallelizable while REFERENCEFREE can only be run single threaded.
Regarding accuracy, Fig. 2 shows that REFERENCEFREE, using Jaccard distance and 21-mers, was slightly less accurate than CNIDARIA, although it can achieve comparable results with different distance measures (Additional file 7: Table S7). Besides speed, CNIDARIA (and JELLYFISH) use significantly less disk space due to their binary formats. The files generated are smaller than the equivalent files created by REFERENCEFREE, with median sizes of 9.2 Gb vs. 42.2 Gb (and median absolute deviations of 2.5 Gb and 11.0 Gb, respectively) for the k-mer count file and 227 Gb vs. 2.1 Tb for the merged k-mer count file, despite the merged k-mer count file created by REFERENCEFREE containing only 2 % of the total number of k-mers, all of which are present in CNIDARIA.
Conclusions
We have introduced CNIDARIA, a tool to quickly and reliably analyse WGS and RNA-seq samples from both assembled and unassembled NGS data, offering significant advantages in terms of time and space requirements compared to a state-of-the-art tool. By clustering in total 169 eukaryotic samples from 78 species (42 genus, 32 families, 27 orders, 5 phyla, 6 divisions and 3 kingdoms from the Eukaryota superkingdom) we have demonstrated that CNIDARIA can handle a large number of samples from very distant phylogenetic origins, producing a reliable tree with up to 100 % classification accuracy at the supra species level and 78 % accuracy at the species level, the later value being low mostly due to interspecific crossings. As CNIDARIA is also able to analyse RNA-seq data, researchers can acquire, besides the species information, physiological state information such as pathogenicity and stress response of the sample for downstream analysis.
A database created in Database Creation Mode allows querying directly for k-mers shared by a specified set of samples, enabling comparisons useful in several applications. Examples include identifying and quantifying polymorphisms between closely related samples, quantifying sequence diversity in the setup phase of large sequencing projects for sample selection, and ecological diversity analysis. In addition, k-mers shared exclusively by a set of samples can be used for diagnostic primer design, supporting the detection of target genes. Furthermore, mismatching k-mers between a sample and a close relative can be used to identify the source of contamination or introgressions, as performed by [84].
Availability and requirements
Project name: Cnidaria
Project home page: http://www.ab.wur.nl/cnidaria; https://github.com/sauloal/cnidaria/wiki
Operating system(s): 64-bit Linux
Programming language: C++ ×11 and Python 2.7
Other requirements: None to run; GCC 4.8 or higher for compiling
License: MIT
Any restrictions to use by nonacademics: No
Acknowledgements
This project was funded by Centre for BioSystems Genomics (CBSG) under the grant number TO09.
Abbreviations
- CCD
Cnidaria Complete Database
- CSD
Cnidaria Summary Database
- CSV
Comma-Separated Values
- DFT
Discrete Fourier Transform
- EST
Expressed Sequence Tag
- K-mer
Substring of size k
- MAD
Median Absolute Deviation
- NGS
Next Generation Sequencing
- PNG
Portable Network Graphics
- rDNA
Ribosomal DNA
- RFM
Reference-Free Methods
- RNA-seq
RNA sequencing
- WGS
Whole Genome Sequencing
- .fastq
Raw assembly file
- .fasta
Assembled sequence
- .newick
Phylogenetic tree file format
- HTML
HyperText Markup Language
- Read
Contiguous sequence outputted by sequencing machine
Additional files
Sample description. Intermediate headers show database size and analysis time running with 1 thread (1×) or 20 threads (20×) for each CNIDARIA database group. Each line contains a list of the names of the samples used, sequence ID, source type, source name, reference, size of JELLYFISH database, size of input data, GC content, percentage of Ns, number of sequences in the input data and list of samples used in the REFERENCEFREE comparison. For each k-mer size (11, 15, 17, 21 and 31 bp): number of distinct k-mers, total number of k-mers, number of k-mers occurring only once, number of shared k-mers and percentage of k-mers shared. Input data is in the form of assembled genome (genomic - fasta files), raw genomic data (raw - fastq or BAM), filtered genomic data (raw filtered - BAM) or RNA-seq. The 34 samples of the extended dataset were used exclusively against the 21-mer dataset. Analysis time and database sizes are calculated for each dataset and do not correspond to the sum of the partial times and sizes. (XLS 87 kb)
Matrix containing the pairwise number of shared k-mers. The diagonal contains the number of valid k-mers (k-mers shared with at least 1 other sample) of a given sample. 11, 15, 17, 21 and 31-mers are shown as well as the 21-mer dataset downsampled to 2 % of its original size and 21-mer dataset with 34 extra samples. (XLS 1217 kb)
Statistics of k-mer counting. Total number of k-mers in each sample, total number of valid k-mers in each sample (k-mers shared by at least two samples) and the percentage of valid k-mers in each sample. 11, 15, 17, 21 and 31-mers are shown as well as the 21-mer dataset downsampled to 2 % of its original size and 21-mer dataset with 34 extra samples. (XLS 104 kb)
Matrix containing pairwise Jaccard distance between samples. The diagonal is blanked but contains zeroes, meaning identity; 11, 15, 17, 21 and 31-mers are shown, as well as the 21-mer dataset downsampled to 2 % of its original size and 21-mer dataset with 34 extra samples. (XLS 2617 kb)
List of samples used for the 1-nearest-neighbour analysis for supra-species classification and their respective taxonomic ranks. (XLS 29 kb)
List of samples used for the 1-nearest-neighbour analysis for species classification and their respective taxonomic ranks. (XLS 26 kb)
1-nearest-neighbour accuracy results for all k-mer sizes, distance measures and programs. CNIDARIA is tested against all k-mer sizes. REFERENCEFREE is tested using 21-mers. Cnidaria 21 2 % is the dataset containing only 2 % of the data. (XLS 91 kb)
Histogram of Jaccard distances for each k-mer size of the 135 samples. A distance of 0 means identity while a distance of 1 means no similarity. Using 11-mers most samples are identical to each other. For 31-mers, most samples share no similarity with any other sample except for phylogenetically closely related samples. 17 and 21-mers show higher similarity between groups. (PDF 97 kb)
Heatmaps of Jaccard distance and phylogenetic trees of 135 samples using 11-mers. Here, 0 (red) means identity between samples while 1 (blue) means no identity. Generally, closely related species show high similarity with closely related species and no similarity with outgroups. This leads to strong clustering inside groups but loose coupling between groups. Trees in the left shows phylogenetic distances while trees in the right ignores the distances, showing the clustering more clearly; trees plotted using iTOL [85]. (PDF 1668 kb)
Heatmaps of Jaccard distance and phylogenetic trees from 135 samples using 15-mers. Here, 0 (red) means identity between samples while 1 (blue) means no identity. Generally, closely related species show high similarity with closely related species and no similarity with outgroups. This leads to strong clustering inside groups but loose coupling between groups. Trees on the left shows phylogenetic distances while trees on the right ignores the distances, showing the clustering more clearly. Trees were plotted using iTOL [85]. (PDF 1499 kb)
Heatmaps of Jaccard distance and phylogenetic trees from 135 samples using 17-mers. Here, 0 (red) means identity between samples while 1 (blue) means no identity. Generally, closely related species show high similarity with closely related species and no similarity with outgroups. This leads to strong clustering inside groups but loose coupling between groups. Trees on the left shows phylogenetic distances while trees on the right ignores the distances, showing the clustering more clearly. Trees were plotted using iTOL [85]. (PDF 1363 kb)
Heatmaps of Jaccard distance and phylogenetic trees from 135 samples using 21-mers. Here, 0 (red) means identity between samples while 1 (blue) means no identity. Generally, closely related species show high similarity with closely related species and no similarity with outgroups. This leads to strong clustering inside groups but loose coupling between groups. Trees in the left shows phylogenetic distances while trees in the right ignores the distances, showing the clustering more clearly; trees plotted using iTOL [85]. (PDF 1303 kb)
Heatmaps of Jaccard distance and phylogenetic trees from 135 samples using 31-mers. Here, 0 (red) means identity between samples while 1 (blue) means no identity. Generally, closely related species show high similarity with closely related species and no similarity with outgroups. This leads to strong clustering inside groups but loose coupling between groups. Trees on the left shows phylogenetic distances while trees on the right ignores the distances, showing the clustering more clearly. Trees were plotted using iTOL [85] (PDF 1292 kb)
Phylogenetic tress with and without branch lengths of 98 Solanum taxa from 13 species. The Lycopersicon group (comprised of Solanum lycopersicum, S pimpinellifolium, S. cheesmaniae and S. galapagense) clusters as a monophyletic group. Sometimes the non-S. lycopersicum species cluster inside the S. lycopersicum clade. We speculate these are S. lycopersicum varieties containing introgression clustering with the donor species, consistently with the findings of [27]. The Arcanum group (comprised of S. arcanum, S. chmielewskii and S. neorikii) also clusters monophyletically, closer to the Eriopersicon group, its sister group. The North Eriopersicon group (comprised of S. huaylasense, S. chilense, S. peruvianum and S. corneliomulleri) groups with the South Eriopersicon group (comprised of S. habrochaites, its only member) and its sister group, Neolycopersicon (comprised of S. pennelli, its only member). S. tuberosum and Nicotiana were added as outgroups. Sample names ending in RAW are raw genomic data; names ending in APLG and CLC are assembled genomes. Trees were plotted using iTOL [85] (PDF 1862 kb)
Results for the 21-mer dataset of 169 individuals using Jaccard distance and Neighbour-Joining. (A) phylogenetic tree with distance; (B) phylogenetic tree without distance (tree branch length); (C) heatmap of phylogenetic distances showing low inter-group similarity and high intra-group similarity; (D) histogram of Jaccard distances showing the same feature of low inter-group similarity and high intra-group similarity. Sample names ending in RAW are raw genomic data; names ending in APLG and CLC are assembled genomes; names ending in RNA, RNAseq and mRNA are RNA-seq datasets. Trees were plotted using iTOL [85] (PDF 2029 kb)
Results for 2 % of the 21-mer dataset. (A) phylogenetic tree with distance; (B) phylogenetic tree without distance; (C) heatmap of phylogenetic distances showing low inter-group similarity and high intra-group similarity; (D) histogram of Jaccard distances showing the same feature of low inter-group similarity and high intra-group similarity. Trees were plotted using iTOL [85] (PDF 1709 kb)
REFERENCEFREE datasets and statistics. Datasets used in the REFERENCEFREE analysis with the respective number of sequences, number of k-mers, number of valid k-mers (present in at least two samples) and percentage of k-mers considered valid for each dataset. On average, 0.016 ± 0.023 % of the data is used. (XLS 34 kb)
Footnotes
Competing interests
The author(s) declare that they have no competing interests.
Authors’ contributions
SAA designed, wrote and tested the software; DR participated in the design of the software and validated the algorithm; ES, GSP, SP and HJ participated in the design of the software; All authors contributed in the confection of the manuscript, read it and approved the final manuscript.
Authors’ information
Saulo Alves Aflitos: PhD candidate in Bioinformatics
Edouard Severing: PhD in Bioinformatics - Post-doc in Max-Planck-Institut für Pflanzenzüchtungsforschung - Köln
Gabino Sanchez-Perez: PhD in Bioinformatics - Senior Researcher Bioinformatics at Wageningen University Cluster leader Cluster Bioinformatics – Plant Research International – Wageningen University
Sander Peters: PhD in Bioinformatics - Senior scientist/Bioinformatician at Plant Research International – Wageningen University
Hans de Jong: Professor of cytogenetics at Wageningen University
Dick de Ridder: Professor of bioinformatics at Wageningen University
Contributor Information
Saulo Alves Aflitos, Email: sauloalves.aflitos@wur.nl.
Edouard Severing, Email: severing@mpipz.mpg.de.
Gabino Sanchez-Perez, Email: gabino.sanchezperez@wur.nl.
Sander Peters, Email: sander.peters@wur.nl.
Hans de Jong, Email: hans.deJong@wur.nl.
Dick de Ridder, Email: dick.deridder@wur.nl.
References
- 1.Nagoya Protocol on Access to Genetic Resources and the Fair and Equitable Sharing of Benefits Arising from their Utilization to the Convention on Biological Diversity www.cbd.int, accessed at 15 Sept 2015.
- 2.Pettengill JB, Luo Y, Davis S, Chen Y, Gonzalez-Escalona N, Ottesen A, Rand H, Allard MW, Strain E. An evaluation of alternative methods for constructing phylogenies from whole genome sequence data: a case study with Salmonella. PeerJ. 2014;2 doi: 10.7717/peerj.620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bertels F, Silander OK, Pachkov M, Rainey PB, van Nimwegen E. Automated reconstruction of whole-genome phylogenies from short-sequence reads. Mol Biol Evol. 2014;31(5):1077–1088. doi: 10.1093/molbev/msu088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chan CK, Hsu AL, Halgamuge SK, Tang SL. Binning sequences using very sparse labels within a metagenome. BMC Bioinformatics. 2008;9:215. doi: 10.1186/1471-2105-9-215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chan CK, Hsu AL, Tang SL, Halgamuge SK. Using growing self-organising maps to improve the binning process in environmental whole-genome shotgun sequencing. J Biomed Biotechnol. 2008;2008:513701. doi: 10.1155/2008/513701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Diaz NN, Krause L, Goesmann A, Niehaus K, Nattkemper TW. TACOA: taxonomic classification of environmental genomic fragments using a kernelized nearest neighbor approach. BMC Bioinformatics. 2009;10:56. doi: 10.1186/1471-2105-10-56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Greenblum S, Carr R, Borenstein E. Extensive strain-level copy-number variation across human gut microbiome species. Cell. 2015;160(4):583–594. doi: 10.1016/j.cell.2014.12.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hurwitz BL, Westveld AH, Brum JR, Sullivan MB. Modeling ecological drivers in marine viral communities using comparative metagenomics and network analyses. Proc Natl Acad Sci U S A. 2014;111(29):10714–10719. doi: 10.1073/pnas.1319778111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.McHardy AC, Rigoutsos I. What’s in the mix: phylogenetic classification of metagenome sequence samples. Curr Opin Microbiol. 2007;10(5):499–503. doi: 10.1016/j.mib.2007.08.004. [DOI] [PubMed] [Google Scholar]
- 10.Schloissnig S, Arumugam M, Sunagawa S, Mitreva M, Tap J, Zhu A, Waller A, Mende DR, Kultima JR, Martin J, et al. Genomic variation landscape of the human gut microbiome. Nature. 2013;493(7430):45–50. doi: 10.1038/nature11711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Smits SL, Bodewes R, Ruiz-Gonzalez A, Baumgartner W, Koopmans MP, Osterhaus AD, Schurch AC. Assembly of viral genomes from metagenomes. Front Microbiol. 2014;5:714. doi: 10.3389/fmicb.2014.00714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wood DE, Salzberg SL. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014;15(3):R46. doi: 10.1186/gb-2014-15-3-r46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yang B, Peng Y, Leung HC, Yiu SM, Chen JC, Chin FY. Unsupervised binning of environmental genomic fragments based on an error robust selection of l-mers. BMC Bioinformatics. 2010;11 Suppl 2:S5. doi: 10.1186/1471-2105-11-S2-S5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hoang T, Yin C, Zheng H, Yu C, Lucy He R, Yau SS. A new method to cluster DNA sequences using Fourier power spectrum. J Theor Biol. 2015;372:135–145. doi: 10.1016/j.jtbi.2015.02.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tran NH, Chen X. Comparison of next-generation sequencing samples using compression-based distances and its application to phylogenetic reconstruction. BMC Res Notes. 2014;7:320. doi: 10.1186/1756-0500-7-320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Sims GE, Jun SR, Wu GA, Kim SH. Alignment-free genome comparison with feature frequency profiles (FFP) and optimal resolutions. Proc Natl Acad Sci U S A. 2009;106(8):2677–2682. doi: 10.1073/pnas.0813249106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yi H, Jin L. Co-phylog: an assembly-free phylogenomic approach for closely related organisms. Nucleic Acids Res. 2013;41(7) doi: 10.1093/nar/gkt003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Roychowdhury T, Vishnoi A, Bhattacharya A. Next-Generation Anchor Based Phylogeny (NexABP): constructing phylogeny from next-generation sequencing data. Sci Rep. 2013;3:2634. doi: 10.1038/srep02634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ren J, Song K, Sun F, Deng M, Reinert G. Multiple alignment-free sequence comparison. Bioinformatics. 2013;29(21):2690–2698. doi: 10.1093/bioinformatics/btt462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gardner SN, Hall BG. When whole-genome alignments just won’t work: kSNP v2 software for alignment-free SNP discovery and phylogenetics of hundreds of microbial genomes. PLoS One. 2013;8(12) doi: 10.1371/journal.pone.0081760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Horwege S, Lindner S, Boden M, Hatje K, Kollmar M, Leimeister CA, Morgenstern B. Spaced words and kmacs: fast alignment-free sequence comparison based on inexact word matches. Nucleic Acids Res. 2014;42(Web Server issue):W7–W11. doi: 10.1093/nar/gku398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fan H, Ives AR, Surget-Groba Y, Cannon CH. An assembly and alignment-free method of phylogeny reconstruction from next-generation sequencing data. BMC Genomics. 2015;16:522. doi: 10.1186/s12864-015-1647-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cannon CH, Kua CS, Zhang D, Harting JR. Assembly free comparative genomics of short-read sequence data discovers the needles in the haystack. Mol Ecol. 2010;19(Suppl 1):147–161. doi: 10.1111/j.1365-294X.2009.04484.x. [DOI] [PubMed] [Google Scholar]
- 24.Kua CS, Ruan J, Harting J, Ye CX, Helmus MR, Yu J, Cannon CH. Reference-free comparative genomics of 174 chloroplasts. PLoS One. 2012;7(11) doi: 10.1371/journal.pone.0048995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Marcais G, Kingsford C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics. 2011;27(6):764–770. doi: 10.1093/bioinformatics/btr011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Knight R, Maxwell P, Birmingham A, Carnes J, Caporaso JG, Easton BC, Eaton M, Hamady M, Lindsay H, Liu Z, et al. PyCogent: a toolkit for making sense from sequence. Genome Biol. 2007;8(8):R171. doi: 10.1186/gb-2007-8-8-r171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Aflitos S, Schijlen E, de Jong H, de Ridder D, Smit S, Finkers R, Wang J, Zhang G, Li N, Mao L, et al. Exploring genetic variation in the tomato (Solanum section Lycopersicon) clade by whole-genome sequencing. Plant J. 2014;80(1):136–148. doi: 10.1111/tpj.12616. [DOI] [PubMed] [Google Scholar]
- 28.Zimin AV, Delcher AL, Florea L, Kelley DR, Schatz MC, Puiu D, Hanrahan F, Pertea G, Van Tassell CP, Sonstegard TS, et al. A whole-genome assembly of the domestic cow, Bos taurus. Genome Biol. 2009;10(4):R42. doi: 10.1186/gb-2009-10-4-r42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lindblad-Toh K, Wade CM, Mikkelsen TS, Karlsson EK, Jaffe DB, Kamal M, Clamp M, Chang JL, Kulbokas EJ, 3rd, Zody MC, et al. Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature. 2005;438(7069):803–819. doi: 10.1038/nature04338. [DOI] [PubMed] [Google Scholar]
- 30.Wade CM, Giulotto E, Sigurdsson S, Zoli M, Gnerre S, Imsland F, Lear TL, Adelson DL, Bailey E, Bellone RR, et al. Genome sequence, comparative analysis, and population genetics of the domestic horse. Science. 2009;326(5954):865–867. doi: 10.1126/science.1178158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Scally A, Dutheil JY, Hillier LW, Jordan GE, Goodhead I, Herrero J, Hobolth A, Lappalainen T, Mailund T, Marques-Bonet T, et al. Insights into hominid evolution from the gorilla genome sequence. Nature. 2012;483(7388):169–175. doi: 10.1038/nature10842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 33.Ebeling M, Kung E, See A, Broger C, Steiner G, Berrera M, Heckel T, Iniguez L, Albert T, Schmucki R, et al. Genome-based analysis of the nonhuman primate Macaca fascicularis as a model for drug safety assessment. Genome Res. 2011;21(10):1746–1756. doi: 10.1101/gr.123117.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gibbs RA, Rogers J, Katze MG, Bumgarner R, Weinstock GM, Mardis ER, Remington KA, Strausberg RL, Venter JC, Wilson RK, et al. Evolutionary and biomedical insights from the rhesus macaque genome. Science. 2007;316(5822):222–234. doi: 10.1126/science.1139247. [DOI] [PubMed] [Google Scholar]
- 35.Church DM, Goodstadt L, Hillier LW, Zody MC, Goldstein S, She X, Bult CJ, Agarwala R, Cherry JL, DiCuccio M, et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 2009;7(5) doi: 10.1371/journal.pbio.1000112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Consortium TCSaA Initial sequence of the chimpanzee genome and comparison with the human genome. Nature. 2005;437(7055):69–87. doi: 10.1038/nature04072. [DOI] [PubMed] [Google Scholar]
- 37.Gibbs RA, Weinstock GM, Metzker ML, Muzny DM, Sodergren EJ, Scherer S, Scott G, Steffen D, Worley KC, Burch PE, et al. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature. 2004;428(6982):493–521. doi: 10.1038/nature02426. [DOI] [PubMed] [Google Scholar]
- 38.Nierman WC, Pain A, Anderson MJ, Wortman JR, Kim HS, Arroyo J, Berriman M, Abe K, Archer DB, Bermejo C, et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature. 2005;438(7071):1151–1156. doi: 10.1038/nature04332. [DOI] [PubMed] [Google Scholar]
- 39.Galagan JE, Calvo SE, Cuomo C, Ma LJ, Wortman JR, Batzoglou S, Lee SI, Basturkmen M, Spevak CC, Clutterbuck J, et al. Sequencing of Aspergillus nidulans and comparative analysis with A. fumigatus and A. oryzae. Nature. 2005;438(7071):1105–1115. doi: 10.1038/nature04341. [DOI] [PubMed] [Google Scholar]
- 40.Pel HJ, de Winde JH, Archer DB, Dyer PS, Hofmann G, Schaap PJ, Turner G, de Vries RP, Albang R, Albermann K, et al. Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat Biotechnol. 2007;25(2):221–231. doi: 10.1038/nbt1282. [DOI] [PubMed] [Google Scholar]
- 41.Machida M, Asai K, Sano M, Tanaka T, Kumagai T, Terai G, Kusumoto K, Arima T, Akita O, Kashiwagi Y, et al. Genome sequencing and analysis of Aspergillus oryzae. Nature. 2005;438(7071):1157–1161. doi: 10.1038/nature04300. [DOI] [PubMed] [Google Scholar]
- 42.Chibana H, Oka N, Nakayama H, Aoyama T, Magee BB, Magee PT, Mikami Y. Sequence finishing and gene mapping for Candida albicans chromosome 7 and syntenic analysis against the Saccharomyces cerevisiae genome. Genetics. 2005;170(4):1525–1537. doi: 10.1534/genetics.104.034652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jackson AP, Gamble JA, Yeomans T, Moran GP, Saunders D, Harris D, Aslett M, Barrell JF, Butler G, Citiulo F, et al. Comparative genomics of the fungal pathogens Candida dubliniensis and Candida albicans. Genome Res. 2009;19(12):2231–2244. doi: 10.1101/gr.097501.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, Lafontaine I, De Montigny J, Marck C, Neuveglise C, Talla E, et al. Genome evolution in yeasts. Nature. 2004;430(6995):35–44. doi: 10.1038/nature02579. [DOI] [PubMed] [Google Scholar]
- 45.D’Souza CA, Kronstad JW, Taylor G, Warren R, Yuen M, Hu G, Jung WH, Sham A, Kidd SE, Tangen K, et al. Genome variation in Cryptococcus gattii, an emerging pathogen of immunocompetent hosts. mBio. 2011;2(1):e00342–00310. doi: 10.1128/mBio.00342-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Loftus BJ, Fung E, Roncaglia P, Rowley D, Amedeo P, Bruno D, Vamathevan J, Miranda M, Anderson IJ, Fraser JA, et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science. 2005;307(5713):1321–1324. doi: 10.1126/science.1103773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Galagan JE, Calvo SE, Borkovich KA, Selker EU, Read ND, Jaffe D, FitzHugh W, Ma LJ, Smirnov S, Purcell S, et al. The genome sequence of the filamentous fungus Neurospora crassa. Nature. 2003;422(6934):859–868. doi: 10.1038/nature01554. [DOI] [PubMed] [Google Scholar]
- 48.Goffeau A, Barrell BG, Bussey H, Davis RW, Dujon B, Feldmann H, Galibert F, Hoheisel JD, Jacq C, Johnston M, et al. Life with 6000 genes. Science 1996, 274(5287):546, 563–47. [DOI] [PubMed]
- 49.Wood V, Gwilliam R, Rajandream MA, Lyne M, Lyne R, Stewart A, Sgouros J, Peat N, Hayles J, Baker S, et al. The genome sequence of Schizosaccharomyces pombe. Nature. 2002;415(6874):871–880. doi: 10.1038/nature724. [DOI] [PubMed] [Google Scholar]
- 50.Souciet JL, Dujon B, Gaillardin C, Johnston M, Baret PV, Cliften P, Sherman DJ, Weissenbach J, Westhof E, Wincker P, et al. Comparative genomics of protoploid Saccharomycetaceae. Genome Res. 2009;19(10):1696–1709. doi: 10.1101/gr.091546.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hu TT, Pattyn P, Bakker EG, Cao J, Cheng JF, Clark RM, Fahlgren N, Fawcett JA, Grimwood J, Gundlach H, et al. The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nat Genet. 2011;43(5):476–481. doi: 10.1038/ng.807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Tabata S, Kaneko T, Nakamura Y, Kotani H, Kato T, Asamizu E, Miyajima N, Sasamoto S, Kimura T, Hosouchi T, et al. Sequence and analysis of chromosome 5 of the plant Arabidopsis thaliana. Nature. 2000;408(6814):823–826. doi: 10.1038/35048507. [DOI] [PubMed] [Google Scholar]
- 53.Xu Q, Chen LL, Ruan X, Chen D, Zhu A, Chen C, Bertrand D, Jiao WB, Hao BH, Lyon MP, et al. The draft genome of sweet orange (Citrus sinensis) Nat Genet. 2013;45(1):59–66. doi: 10.1038/ng.2472. [DOI] [PubMed] [Google Scholar]
- 54.Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, et al. Genome sequence of the palaeopolyploid soybean. Nature. 2010;463(7278):178–183. doi: 10.1038/nature08670. [DOI] [PubMed] [Google Scholar]
- 55.Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D, et al. The genome of the domesticated apple (Malus x domestica Borkh.) Nat Genet. 2010;42(10):833–839. doi: 10.1038/ng.654. [DOI] [PubMed] [Google Scholar]
- 56.Bombarely A, Rosli HG, Vrebalov J, Moffett P, Mueller LA, Martin GB. A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol Plant Microbe Interact. 2012;25(12):1523–1530. doi: 10.1094/MPMI-06-12-0148-TA. [DOI] [PubMed] [Google Scholar]
- 57.Sierro N, Battey JN, Ouadi S, Bakaher N, Bovet L, Willig A, Goepfert S, Peitsch MC, Ivanov NV. The tobacco genome sequence and its comparison with those of tomato and potato. Nat Commun. 2014;5:3833. doi: 10.1038/ncomms4833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chen J, Huang Q, Gao D, Wang J, Lang Y, Liu T, Li B, Bai Z, Luis Goicoechea J, Liang C, et al. Whole-genome sequencing of Oryza brachyantha reveals mechanisms underlying Oryza genome evolution. Nat Commun. 2013;4:1595. doi: 10.1038/ncomms2596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yamamoto T, Nagasaki H, Yonemaru J, Ebana K, Nakajima M, Shibaya T, Yano M. Fine definition of the pedigree haplotypes of closely related rice cultivars by means of genome-wide discovery of single-nucleotide polymorphisms. BMC Genomics. 2010;11:267. doi: 10.1186/1471-2164-11-267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A, et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray) Science. 2006;313(5793):1596–1604. doi: 10.1126/science.1128691. [DOI] [PubMed] [Google Scholar]
- 61.Tomato Genome C. The tomato genome sequence provides insights into fleshy fruit evolution. Nature. 2012;485(7400):635–641. doi: 10.1038/nature11119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Park SJ, Jiang K, Schatz MC, Lippman ZB. Rate of meristem maturation determines inflorescence architecture in tomato. Proc Natl Acad Sci U S A. 2012;109(2):639–644. doi: 10.1073/pnas.1114963109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Xu X, Pan S, Cheng S, Zhang B, Mu D, Ni P, Zhang G, Yang S, Li R, Wang J, et al. Genome sequence and analysis of the tuber crop potato. Nature. 2011;475(7355):189–195. doi: 10.1038/nature10158. [DOI] [PubMed] [Google Scholar]
- 64.Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature. 2007;449(7161):463–467. doi: 10.1038/nature06148. [DOI] [PubMed] [Google Scholar]
- 65.Schnable PS, Ware D, Fulton RS, Stein JC, Wei F, Pasternak S, Liang C, Zhang J, Fulton L, Graves TA, et al. The B73 maize genome: complexity, diversity, and dynamics. Science. 2009;326(5956):1112–1115. doi: 10.1126/science.1178534. [DOI] [PubMed] [Google Scholar]
- 66.Aflitos SA, Sanchez-Perez G, de Ridder D, Fransz P, Schranz ME, de Jong H, Peters SA. Introgression browser: high-throughput whole-genome SNP visualization. Plant J. 2015;82(1):174–182. doi: 10.1111/tpj.12800. [DOI] [PubMed] [Google Scholar]
- 67.Huang Y, Li Y, Burt DW, Chen H, Zhang Y, Qian W, Kim H, Gan S, Zhao Y, Li J, et al. The duck genome and transcriptome provide insight into an avian influenza virus reservoir species. Nat Genet. 2013;45(7):776–783. doi: 10.1038/ng.2657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Consortium* ICGS Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature. 2004;432(7018):695–716. doi: 10.1038/nature03154. [DOI] [PubMed] [Google Scholar]
- 69.Shapiro MD, Kronenberg Z, Li C, Domyan ET, Pan H, Campbell M, Tan H, Huff CD, Hu H, Vickrey AI, et al. Genomic diversity and evolution of the head crest in the rock pigeon. Science. 2013;339(6123):1063–1067. doi: 10.1126/science.1230422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Poelstra JW, Vijay N, Bossu CM, Lantz H, Ryll B, Muller I, Baglione V, Unneberg P, Wikelski M, Grabherr MG, et al. The genomic landscape underlying phenotypic integrity in the face of gene flow in crows. Science. 2014;344(6190):1410–1414. doi: 10.1126/science.1253226. [DOI] [PubMed] [Google Scholar]
- 71.Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B, Markow TA, Kaufman TC, Kellis M, Gelbart W, Iyer VN, et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature. 2007;450(7167):203–218. doi: 10.1038/nature06341. [DOI] [PubMed] [Google Scholar]
- 72.St Pierre SE, Ponting L, Stefancsik R, McQuilton P, FlyBase C. FlyBase 102--advanced approaches to interrogating FlyBase. Nucleic Acids Res. 2014;42(Database issue):D780–D788. doi: 10.1093/nar/gkt1092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Celniker SE, Wheeler DA, Kronmiller B, Carlson JW, Halpern A, Patel S, Adams M, Champe M, Dugan SP, Frise E, et al. Finishing a whole-genome shotgun: release 3 of the Drosophila melanogaster euchromatic genome sequence. Genome Biol. 2002;3(12):Research0079. doi: 10.1186/gb-2002-3-12-research0079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Zhan X, Pan S, Wang J, Dixon A, He J, Muller MG, Ni P, Hu L, Liu Y, Hou H, et al. Peregrine and saker falcon genome sequences provide insights into evolution of a predatory lifestyle. Nat Genet. 2013;45(5):563–566. doi: 10.1038/ng.2588. [DOI] [PubMed] [Google Scholar]
- 75.Zhang Y, Wiggins BE, Lawrence C, Petrick J, Ivashuta S, Heck G. Analysis of plant-derived miRNAs in animal small RNA datasets. BMC Genomics. 2012;13:381. doi: 10.1186/1471-2164-13-381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Dalloul RA, Long JA, Zimin AV, Aslam L, Beal K, Le Blomberg A, Bouffard P, Burt DW, Crasta O, Crooijmans RP, et al. Multi-platform next-generation sequencing of the domestic turkey (Meleagris gallopavo): genome assembly and analysis. PLoS Biol. 2010;8(9):e1000475. doi: 10.1371/journal.pbio.1000475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Werren JH, Richards S, Desjardins CA, Niehuis O, Gadau J, Colbourne JK, Werren JH, Richards S, Desjardins CA, Niehuis O, et al. Functional and evolutionary insights from the genomes of three parasitoid Nasonia species. Science. 2010;327(5963):343–348. doi: 10.1126/science.1178028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Richards S, Gibbs RA, Weinstock GM, Brown SJ, Denell R, Beeman RW, Gibbs R, Beeman RW, Brown SJ, Bucher G, et al. The genome of the model beetle and pest Tribolium castaneum. Nature. 2008;452(7190):949–955. doi: 10.1038/nature06784. [DOI] [PubMed] [Google Scholar]
- 79.Qiu Q, Zhang G, Ma T, Qian W, Wang J, Ye Z, Cao C, Hu Q, Kim J, Larkin DM, et al. The yak genome and adaptation to life at high altitude. Nat Genet. 2012;44(8):946–949. doi: 10.1038/ng.2343. [DOI] [PubMed] [Google Scholar]
- 80.Leggett RM, Ramirez-Gonzalez RH, Clavijo BJ, Waite D, Davey RP. Sequencing quality assessment tools to enable data-driven informatics for high throughput genomics. Front Genet. 2013;4:288. doi: 10.3389/fgene.2013.00288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Choi S-S, Cha S-H, Tappert CC. A survey of binary similarity and distance measures. J Syst Cybern Inf. 2010;8(1):43–48. [Google Scholar]
- 82.Illumina inc. http://www.illumina.com, accessed at 15 Sept 2015.
- 83.Simpson JT, Wong K, Jackman SD, Schein JE, Jones SJ, Birol I. ABySS: a parallel assembler for short read sequence data. Genome Res. 2009;19(6):1117–1123. doi: 10.1101/gr.089532.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Byrd AL, Perez-Rogers JF, Manimaran S, Castro-Nallar E, Toma I, McCaffrey T, Siegel M, Benson G, Crandall KA, Johnson WE. Clinical PathoScope: rapid alignment and filtration for accurate pathogen identification in clinical samples using unassembled sequencing data. BMC Bioinformatics. 2014;15:262. doi: 10.1186/1471-2105-15-262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Letunic I, Bork P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics. 2007;23(1):127–128. doi: 10.1093/bioinformatics/btl529. [DOI] [PubMed] [Google Scholar]


