Abstract
A new virulent phage belonging to the Siphoviridae family and able to infect Lactococcus garvieae strains was isolated from compost soil. Phage GE1 has a prolate capsid (56 by 38 nm) and a long noncontractile tail (123 nm). It had a burst size of 139 and a latent period of 31 min. Its host range was limited to only two L. garvieae strains out of 73 tested. Phage GE1 has a double-stranded DNA genome of 24,847 bp containing 48 predicted open reading frames (ORFs). Putative functions could be assigned to only 14 ORFs, and significant matches in public databases were found for only 17 ORFs, indicating that GE1 is a novel phage and its genome contains several new viral genes and encodes several new viral proteins. Of these 17 ORFs, 16 were homologous to deduced proteins of virulent phages infecting the dairy bacterium Lactococcus lactis, including previously characterized prolate-headed phages. Comparative genome analysis confirmed the relatedness of L. garvieae phage GE1 to L. lactis phages c2 (22,172 bp) and Q54 (26,537 bp), although its genome organization was closer to that of phage c2. Phage GE1 did not infect any of the 58 L. lactis strains tested. This study suggests that phages infecting different lactococcal species may have a common ancestor.
INTRODUCTION
Lactococcus garvieae is the etiological agent of lactococcosis, a pathology affecting a variety of fish species that is responsible for significant economic losses in both marine and freshwater aquacultures (1). In addition, L. garvieae has been associated with subclinical intramammary infections in cows (2). More recently, L. garvieae was associated with infections in humans and is now considered an opportunistic and potentially zoonotic pathogen (3, 4). In an effort to find its ecological niches, genetic investigations have revealed its presence, and sometimes dominance, in several habitats: river and sewage waters and raw and processed foods, including milk, meats, and vegetables (5–7).
Despite growing attention, genomic characterization of L. garvieae progresses slowly, yet the evolutionary history and global diversity of this emerging pathogen are particularly interesting. The population structure and diversity of L. garvieae strains have been studied using comparative genome analysis, which led to the development of a multilocus sequence type (MLST) scheme to identify specific strains (8). These studies also revealed that L. garvieae is related to Lactococcus lactis, a nonpathogenic bacterium used for the manufacture of several fermented dairy products (9). Divergent genomic lineages were observed, with some ecotypes representing evolutionary intermediates between L. lactis and L. garvieae (8). The presence of insertion sequences (ISs) was also highly variable among L. garvieae strains (10). Analysis of these ISs within individual genomes revealed limited intragenomic sequence diversity, suggesting that many ISs in L. garvieae are evolutionarily young and may have been only recently acquired. Of note, the similarity of most L. garvieae ISs to L. lactis ISs suggests possible genetic exchange between these two species.
Little is known about the presence of other mobile genetic elements such as phages in L. garvieae. Phages are usually found in the same niches as their bacterial hosts and can have two main life cycles, lytic or lysogenic. During the lytic cycle, the phage redirects the host metabolism to produce new virions, resulting in host cell lysis and the release of numerous virulent phages that can infect other targeted cells. During the lysogenic cycle, the phage genome integrates into the host genome, remains in the host as a prophage, and replicates along with the host genome, until the lytic cycle is induced by stress factors. Phages play key roles in bacterial evolution, influencing the dynamics of bacterial populations and their adaptability through gene exchange (11, 12).
The emergence of multidrug-resistant bacteria has sparked renewed interest in virulent phages for potential use in therapy or in sanitation strategies in the agricultural, food processing, and fishing industries as well as in humans and animals (13, 14). However, phages can also destroy bacteria that play key roles in production or fermentation processes, particularly in the food and biotechnology industries (15, 16).
Very few lytic phages of L. garvieae have been identified to date (17–19). One L. garvieae phage belonging to the Siphoviridae family (double-stranded DNA [dsDNA] genome, long noncontractile tail) was investigated for controlling L. garvieae infections in fish (19). To our knowledge, only one L. garvieae-virulent phage has been investigated at the genomic level (20). L. garvieae phage WP-2 belongs to the Podoviridae family (dsDNA genome, short noncontractile tail) of the order Caudovirales (12). It was classified as a member of the Ahjdlikevirus genus in the Picovirinae subfamily because it has a small genome (18,899 bp, 24 open reading frames [ORFs]) and a protein-primed DNA polymerase (20).
Here, we report the molecular characterization of the novel L. garvieae phage GE1. This virulent phage was isolated from a Canadian compost soil in 2014 and belongs to the Siphoviridae family. Comparative genome analyses suggest that this new siphophage is related to virulent phages infecting L. lactis.
MATERIALS AND METHODS
Bacterial host and phage isolation.
A total of 73 L. garvieae strains previously isolated from different ecological niches (fish and dairy farms, vegetables, meat, and cereals) (21) and 58 industrial L. lactis strains were used to determine the host range of the phage. Lactococcal strains were grown statically at 30°C in M17 broth (Oxoid) supplemented with 0.5% glucose (GM17) or lactose (LM17). We tested Canadian samples of raw milk, wastewater, soil compost, and aquaculture water to isolate phages infecting L. garvieae. The soil compost from which we isolated phage GE1 was made from green waste (domestic compost) obtained from the Compo-Recycle Center at Chertsey (Québec, Canada), and the samples used for phage isolation were taken from the top.
All samples were kept at 4°C until used, and the same phage isolation protocol was used (22). Five milliliters of each environmental sample (for the solid compost soil, 5 g was resuspended in 5 ml of sterile water) was added to 5 ml of double-strength GM17 supplemented with 10 mM CaCl2 and inoculated with 100 μl of an overnight culture of an L. garvieae strain. Nineteen representative strains of previously studied L. garvieae (8) were used for lytic phage isolation. The mixtures were incubated overnight at 30°C. The cultures were centrifuged at 7,000 rpm for 10 min using an IEC clinical centrifuge (USA), and the supernatants were filtered (0.45-μm cellulose acetate filter). Five milliliters of the filtered supernatant was added to double-strength GM17 inoculated as described above. This second amplification procedure was repeated three more times. The presence of phages in the supernatants was tested by spot test, depositing 5 μl of the last amplification on a GM17 plate with 10 mM CaCl2 containing a single L. garvieae strain. Clear plaques were picked, purified three times, and amplified as described previously (23). To obtain concentrated phage preparations, 1 liter of lysate was mixed with polyethylene glycol 8000 (10%) and separated on a discontinuous CsCl gradient followed by a continuous CsCl gradient, as previously reported (24).
Microbiological assays.
The host range of phage GE1 was assessed by spotting 5 μl of a 10−2 dilution of a phage lysate on top agar containing one lactococcal strain. One-step growth assays were performed in triplicate with a multiplicity of infection (MOI) of 0.05 and the host strain L. garvieae INS1 (21). The burst size was calculated by dividing the average phage titer after the exponential phase by average titer before the infected cells began to release virions (25).
Electron microscopy.
Ten microliters of uranyl acetate (2%) was deposited on a Formvar carbon-coated grid (200 mesh; Pelco International). Cesium chloride-purified phages (10 μl) were mixed with the stain by pipetting up and down. After 2 min, the grid was deposited on blotting paper and dried for at least 5 min. Phages were observed at 80 kV using a JEOL1230 transmission electron microscope at the Plateforme d'Imagerie Moléculaire et Microscopie of the Université Laval. Capsid size and tail length were determined by measuring at least 10 phage specimens.
Phage DNA extraction, sequencing, and genome analysis.
Phage DNA was extracted from 500 ml of lysate using the Qiagen Lambda Maxi DNA purification kit with modifications as described previously (26). To determine the genome sequence of phage GE1, a sequencing library was first prepared with the Nextera XT DNA sample preparation kit (Illumina) according to the manufacturer's instructions. The library was sequenced using a MiSeq reagent kit (v2) on a MiSeq system (Illumina). De novo assembly was performed with the Ray assembler (27). Genome extremities were amplified using converging primers, and the PCR product was sequenced with an ABI 3730xl sequencer at the sequencing platform of the Centre Hospitalier de l'Université Laval. The cos site was also confirmed by direct sequencing of the phage DNA using forward (5′-GCAAGGAGGTAATCAGATGCA-3′) and reverse (5′-GAACGCATTCTGTGAGCTTG-3′) primers.
Open reading frame (ORF) prediction was carried out using ORF Finder (http://www.ncbi.nlm.nih.gov/gorf/gorf.html) and RAST Server (28). An ORF was considered valid only if its starting codon was AUG, UUG, or GUG and it was at least 30 amino acids (aa) in length. Ribosomal binding sites (RBSs) were also identified. Functions and domains were attributed by comparison of the translated products using BLASTp (29). PSI-BLAST and InterProScan (http://www.ebi.ac.uk/) were used to search for more distant homologous proteins and conserved domains, respectively, when significant similarity was found by BLAST searches. Theoretical molecular masses (MM) and isoelectric points (pI) of the phage proteins were obtained using ProtParam (http://web.expasy.org/protparam/). tRNAs were identified using the tRNAscan-SE server (30) and confirmed using ARAGORN (31). Virulence Factor Databases (32) together with DBETH (33) were used to identify virulence factors.
Structural protein profile.
Purified GE1 and c2 phages (∼1011 PFU/ml) were dialyzed (0.02 M Tris-HCl, pH 7.4, 0.1 M NaCl, 0.1 M MgSO4), mixed with 4× loading buffer (0.25 M Tris-HCl, pH 6.8, 40% [vol/vol] glycerol, 8% [wt/vol] SDS, 20% [vol/vol] β-mercaptoethanol, and 0.1% [wt/vol] bromophenol blue), and boiled for 5 min. Proteins were separated on an Any kD Mini-Protean TGX precast protein gel (Bio-Rad) and stained with Coomassie blue.
Proteomic tree design.
The predicted proteome of L. garvieae phage GE1 was compared with other lactococcal phage proteomes, including the L. garvieae phage WP-2 (20) and members of the 10 L. lactis phage groups (34). The genomes were downloaded from NCBI, and the proteins were extracted using the published annotations. The terminase was used as the starting point to generate a long phage protein concatemer for each phage (35 phages in total). The order of the proteins was identical to the gene order presented in the genome. For the two L. lactis phage genomes (KSY1 and P087) in which a terminase was not found, the protein concatemers were constructed according to the gene order as deposited in GenBank. MEGA5 (35) software and a BLOSUM matrix were used for multiple alignments. An unrooted phylogenetic tree was constructed using the neighbor-joining algorithm (36). The phylogeny was tested with 100 bootstrap replicates.
Nucleotide sequence accession number.
The annotated genomic sequence of phage GE1 was deposited in GenBank under accession number KT339177.
RESULTS
Characterization of phage GE1.
L. garvieae phage GE1, a virulent phage isolated from a compost soil sample, belongs to the Siphoviridae family (Fig. 1). It has a prolate capsid of 56 nm ± 3 nm by 38 nm ± 3 nm and a noncontractile tail of 123 nm ± 12 nm in length and 11 nm ± 2 nm in width. The phage morphology is highly reminiscent of phages c2 and Q54, members of two distinct L. lactis phage groups that bear their names (34). However, the tail length of phage GE1 is slightly greater (34).
FIG 1.
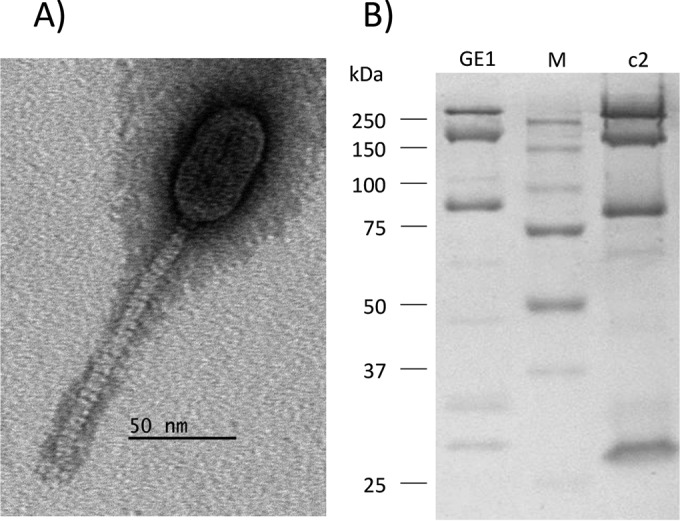
Morphology and structural proteome of phage GE1. (A) Electron micrograph of phage GE1. (B) Structural protein profiles of phages GE1 and c2. Numbers on the left indicate molecular mass of the markers (Bio-Rad) (lane M).
The burst size of phage GE1 was calculated to be 139 ± 13 virions per infected cell, and its latency period was 31 ± 4 min, similar to that of other lactococcal phages (36). The host range of phage GE1 was assessed using a collection of 73 L. garvieae strains and 58 industrial L. lactis strains. Phage GE1 was highly specific as it infected only two L. garvieae strains, its host INS1 and strain INS2, which was isolated from a similar ecological niche (vegetables) (21). Phage GE1 did not infect any of the 58 L. lactis strains tested.
Genome of phage GE1 and its cos site.
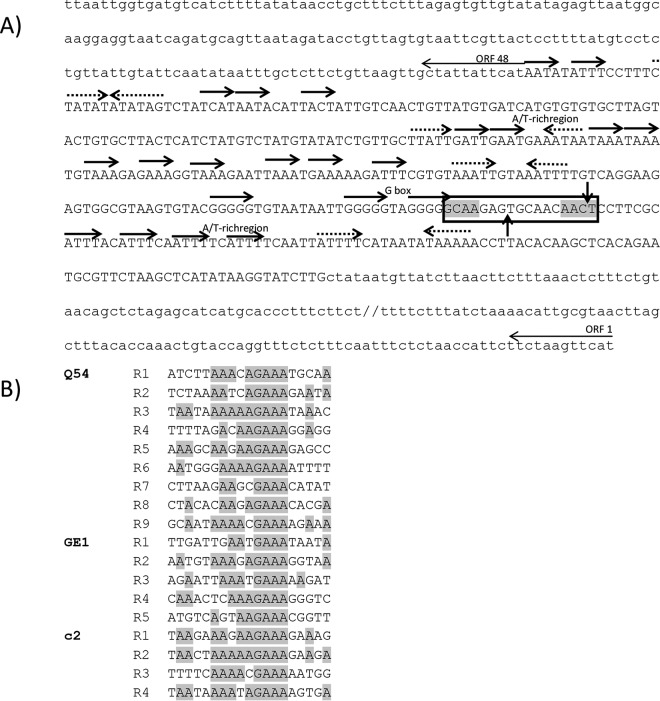
Phage GE1 has a 24,847-bp linear double-stranded DNA genome with a GC content of 37.8%, similar to the GC content of its host genome (8) but slightly higher than that of L. lactis phage genomes (35%) (36). Analysis of the phage GE1 genome did not identify any tRNA or recognizable virulence factors. The genome has cohesive ends consisting of single-stranded 9-bp 3′ overhangs (Fig. 2A). Analysis of the DNA region surrounding the cos site revealed characteristics similar to L. lactis phages belonging to the c2 group (37). This region is characterized by inverted repeat sequences as well as 22 direct repeats. Overall, this region is AT-rich (69%) compared to the rest of the GE1 genome (62%). Three G boxes and no C clusters were found near the cos site (Fig. 2A).
FIG 2.
Analysis of GE1 cos region. (A) Analysis of the cos site and the flanking regions. Direct repeats (bold arrows), inverted repeats (bold, dashed arrows), A/T-rich regions, and G-rich segments are indicated. Details of the cohesive termini (rectangle) are also shown together with the site of cleavage (vertical arrows). Shading indicates nucleotides that belong to the consensus sequence found in all lactococcal phages for which the cos site has been determined. (B) Multiple alignments of λ-like R consensus sequences from Q54, GE1, and c2. Conserved nucleotides are shaded.
The inverted repeat found in the cos site of phage GE1 (GCAA) lacks a conserved T in one of the consensus sequences [TCAN(N4–7)NACT], typically found within a 15-bp segment spanning the cos site of many L. lactis phages (38). The absence of this conserved base in the consensus cos site is reminiscent of the cos site of lactococcal phage Q54, which has a CCAA inverted repeat.
The cos region of GE1 also contains five λ-like R consensus sequences, which are usually involved in terminase recognition and binding, as well as in packaging termination (39). The sequences were compared with those previously reported for L. lactis phages c2, bIL67 (c2-like), and Q54. As reported for phage Q54 (37), phage GE1's five λ-R sequences are distributed unevenly but across a smaller genomic region (500 bp compared to 900 bp of phage Q54). This contrasts with coliphage λ and lactococcal phage c2, whose λ-R sequences are regularly spaced. Alignment analysis of the R sites from GE1 (Fig. 2B) found bases in common involved in the packaging process, similar to those observed for c2 and Q54.
Analysis of phage GE1 genes.
A total of 48 ORFs longer than 30 codons were predicted from the genome sequence (Table 1). The sizes of the gene products range from 44 (ORF17 and ORF26) to 839 (ORF37) amino acids, with an average predicted protein size of 153 amino acids. Coding regions represent approximately 88% of the genome, and the longest noncoding region is 657 bp, occurring between orf27 and orf28. As for c2-like lactococcal phages, the GE1 genome is organized in two main gene clusters, grouping the presumed early- and late-expressed genes (40, 41). The late-expressed genes code for proteins involved in packaging (orf40), cell lysis (orf29 and orf38), and phage morphology (orf32, orf34, and orf37). No lysogeny module was found in the genome, confirming the lytic nature of phage GE1.
TABLE 1.
Features of the ORFs of L. garvieae phage GE1b
| ORF | Stranda | Position |
Size (aa) | MM (kDa) | pI | Putative RBS and start codonc | Predicted functiond | Best-match BLASTp result | No. of aa shared with best match/total no. of aa in best match (% identity) | E value | Sizee (aa) | Accession no. | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Start | Stop | ||||||||||||
| 1 | − | 529 | 104 | 141 | 16.9 | 6.8 | AGGAGActagaATG | — | |||||
| 2 | − | 678 | 526 | 50 | 6.1 | 9.8 | aAGGAGtgcaagctATG | — | |||||
| 3 | − | 854 | 675 | 59 | 7.2 | 9.3 | AAGAAAGgcaatgtagtgaaagaATG | — | |||||
| 4 | − | 1039 | 842 | 65 | 7.7 | 9.5 | AAGAAAGAggtagaaaATG | — | |||||
| 5 | − | 1295 | 1029 | 88 | 10.1 | 9.8 | AAGAAAGaaaacatctaaaATG | — | |||||
| 6 | − | 1650 | 1477 | 57 | 6.8 | 8.7 | gAAGAGGtatagaaATG | — | |||||
| 7 | − | 1949 | 1653 | 98 | 11.1 | 4.5 | AGAGGAttgagATG | — | |||||
| 8 | − | 2256 | 1951 | 101 | 11.5 | 4.5 | GAGgtGAAaagagcATG | — | |||||
| 9 | − | 2678 | 2253 | 141 | 16.2 | 10.1 | AGGAGtttatagaccaGTG | HNH endonuclease | ORF31 Lactococcus phage 712 | 66/137 (48) | 3e−32 | 141 | YP_764291.1 |
| 10 | − | 2947 | 2675 | 90 | 10.4 | 7.9 | GAAAGGGGAtatatATG | — | |||||
| 11 | − | 3588 | 3073 | 171 | 20.0 | 9.4 | AGGAGAactttaATG | — | ORF41 Lactococcus phage 191 | 54/162 (33) | 5e−11 | 177 | AFE86775.1 |
| 12 | − | 3770 | 3588 | 60 | 7.1 | 4.7 | GAAAGGccattATG | tRNA synthetase | ORF6 Lactococcus phage c2 | 24/52 (46) | 0.15 | 54 | NP_043532.1 |
| 13 | − | 4179 | 3754 | 141 | 16.4 | 9.8 | ctgacttgccattttaacATG | HNH endonuclease | ORF48 Lactococcus phage P475 | 74/142 (52) | 3e−36 | 145 | AGI11121.1 |
| 14 | − | 4510 | 4181 | 109 | 11.9 | 4.7 | AAGAGAatataaattATG | ssDNA-binding protein | ORF40 Lactococcus phage 7 | 49/117 (42) | 2e−18 | 119 | YP_008318242.1 |
| 15 | − | 5031 | 4531 | 166 | 18.3 | 7.8 | AGGAAAttgaattcgcATG | ERF recombinase | Rec, S. pseudopneumoniae | 58/123 (47) | 1e−29 | 207 | WP_000163473.1 |
| 16 | − | 5231 | 5028 | 67 | 7.8 | 5.2 | AGGAGGacaagaagaaATG | — | |||||
| 17 | − | 5402 | 5268 | 44 | 5.4 | 6.7 | atacAGAGGtggatagATG | — | |||||
| 18 | − | 5637 | 5389 | 82 | 10.0 | 9.5 | AtgAGGGAtttaagttagATG | — | ORF10 Lactococcus phage c2 | 31/77 (40) | 2e−05 | 83 | NP_043536.1 |
| 19 | − | 5920 | 5621 | 99 | 11.9 | 4.9 | CGTAGAGGActttggcaagtATG | — | |||||
| 20 | − | 6176 | 5913 | 87 | 10.3 | 4.6 | AGGAGAgtaatATG | — | gp25 Lactococcus phage KSY1 | 27/60 (45) | 3e−07 | 99 | YP_001469023.1 |
| 21 | − | 6531 | 6352 | 59 | 7.1 | 7.9 | AGGttGAGAtattagaATG | — | |||||
| 22 | − | 7478 | 6528 | 316 | 36.1 | 5.4 | AAGGAGtAaatacaGTG | DNA polymerase | ORF42 Lactococcus phage jm3 | 166/317 (52) | 6e−106 | 315 | YP_008318192.1 |
| 23 | − | 7737 | 7465 | 90 | 10.5 | 8.9 | GGAAAGGctttggattATG | — | |||||
| 24 | − | 8148 | 7921 | 75 | 9.0 | 8.9 | AGGAGAaaacaaaATG | — | |||||
| 25 | − | 8355 | 8218 | 45 | 5.4 | 9.4 | AAAGGAcggtgttattATG | — | |||||
| 26 | − | 8476 | 8342 | 44 | 5.1 | 9.7 | GAGAGGtttagaaaATG | — | |||||
| 27 | − | 8651 | 8466 | 61 | 7.3 | 4.7 | AGGAGActattatATG | — | |||||
| 28 | + | 9309 | 9461 | 50 | 5.8 | 4.5 | AGGAGAaaggtttATG | — | |||||
| 29 | + | 9463 | 9951 | 162 | 18.2 | 5.2 | tactcattacctatttaatATG | Holin | ORF24 Lactococcus phage c2 | 92/159 (58) | 1e−59 | 161 | NP_043550.1 |
| 30 | + | 10496 | 11167 | 223 | 24.9 | 9.0 | AAAGGAAAtaattaaTTG | — | |||||
| 31 | + | 11177 | 12109 | 310 | 34.7 | 5.2 | AAAtAGAGGtaacaaATG | — | |||||
| 32 | + | 12096 | 13667 | 523 | 57.6 | 4.9 | GAAGAGtgaggtagaagATG | MCP | MCP Lactococcus phage CB17 | 130/535 (24) | 9e−22 | 489 | AAZ95019.1 |
| 33 | + | 13684 | 14484 | 266 | 28.9 | 5.2 | AAAGAGgtaacttATG | — | |||||
| 34 | + | 14486 | 15100 | 204 | 22.4 | 4.7 | AAGAAAGGtataatttaatATG | Major tail protein | ORF23 Lactococcus phage Q54 | 65/200 (33) | 1e−21 | 212 | YP_762592.1 |
| 35 | + | 15151 | 15342 | 63 | 6.7 | 6.0 | AGGGGGAAggTTG | Ig-like domain | |||||
| 36 | + | 15366 | 15617 | 83 | 9.6 | 5.6 | AGGGGActtaatATG | — | |||||
| 37 | + | 15798 | 18317 | 839 | 87.8 | 5.5 | AGGAAAttaatTTG | Tail protein | Pl10 Lactococcus phage 5447 | 123/471 (26) | 3e−22 | 622 | AAT73602.1 |
| 38 | + | 18331 | 18606 | 91 | 10.2 | 9.3 | AGGGAAaataATG | Holin | Gl17 Lactococcus phage 923 | 54/85 (65) | 1e−27 | 96 | AAT81366.1 |
| 39 | + | 18684 | 18968 | 94 | 10.9 | 8.3 | AGcttgtGGGGGtAtcATG | HNH endonuclease | Gp27 Lactococcus phage Q54 | 44/101 (44) | 4e−16 | 103 | YP_762596.1 |
| 40 | + | 18968 | 20605 | 545 | 61.3 | 6.6 | AtGGGGtGGtaATG | Terminase | ORF28 Lactococcus phage Q54 | 191/507 (38) | 2e−96 | 514 | YP_762597.1 |
| 41 | + | 20605 | 20901 | 98 | 11.4 | 9.1 | AGGGAAattaatctaATG | — | ORF35 Lactococcus phage c2 | 20/65 (31) | 0.005 | 99 | NP_043561.1 |
| 42 | + | 20901 | 22100 | 399 | 46.6 | 5.5 | AAGGAGtttatgtaATG | — | |||||
| 43 | + | 22100 | 22759 | 219 | 24.9 | 5.8 | GAAAGGttacgggtataatttaATG | — | |||||
| 44 | + | 22759 | 23526 | 255 | 27.5 | 4.8 | GAAGGGttatgttggtggattaATG | — | |||||
| 45 | − | 23708 | 23568 | 46 | 4.9 | 9.5 | AGGAGAacatcATG | — | |||||
| 46 | − | 24237 | 23797 | 146 | 16.4 | 10.2 | GAAAGGtggttatatgtaATG | — | |||||
| 47 | − | 24410 | 24237 | 57 | 6.6 | 4.0 | AAGcAGGttatataaaagATG | — | |||||
| 48 | − | 24576 | 24394 | 60 | 6.8 | 4.7 | AGGAAAtatattATG | — | |||||
Orientation of the gene in the genome.
Abbreviations: MM, molecular mass; RBS, ribosomal binding site; ssDNA, single-stranded DNA; MCP, major capsid protein; Rec, recombinase.
Uppercase letters represent hypothetical RBS sequences (also underlined); bold letters represent starting codons.
—, no significant matches.
Total size of the aligned proteins.
Seven ORFs (1, 11, 14, 20, 24, 27, and 28) possess a typical RBS domain (AGGAGA). Of these, orf27 and orf28 are located at the ends of the 657-bp intergenic region between early- and late-expressed genes. Moreover, orf27 (early gene) is also preceded by the consensus −10 and −35 promoter sequences (TTGACA-17 bp-TATAAT), whereas upstream of orf28 (late gene) only a −10 consensus sequence was found. These characteristics suggest that the region between orf27 and orf28 could act as an origin of replication, as already reported for phage c2 (42).
Proteome of phage GE1.
The functions of phage GE1 deduced proteins were determined by comparison (BLASTp) with the GenBank database. When low similarity was found between GE1 proteins and other phage proteins, putative functions were reinforced by identifying conserved domains and/or position-specific iterative comparisons. Only ORFs with a significant protein hit in GenBank are presented in Table 1. Predicted functions were attributed to 14 ORFs (out of 48 ORFs; 29%), and only 17 ORFs (35%) matched proteins in the GenBank database, indicating that new viral genes/proteins were identified by characterizing this novel phage. Details are given below for some ORFs.
Homing endonucleases and recombination genes.
Three ORFs (ORF9, ORF13, and ORF39) were predicted to be homing endonucleases. These ORFs had similarity with deduced proteins found in Lactococcus phages 712 and P745 (43), both 936/Sk1-like phages, as well as with L. lactis phage Q54 (37). Since homing endonucleases promote genetic exchange and horizontal transfer (44), this part of the phage GE1 genome may have arisen through the acquisition of mobile DNA, as already proposed for phage 712 (45). ORF15 and ORF22 had similarity to proteins with putative recombination functions, found in Streptococcus pseudopneumoniae and 936/Sk1-like phage jm3 (43).
Replication.
ORF12 and ORF14 have homology with a glutamyl-tRNA synthase protein found in Lactococcus phage c2 and with a single-stranded binding protein of lactococcal 936/Sk1-like phages, such as phi7 and P113G (43). Glutamate tRNA synthase catalyzes the attachment of the amino acid glutamine to its cognate tRNA molecule in a highly specific two-step reaction (46). Single-stranded binding proteins are required during DNA replication, repair, and recombination (40, 47, 48).
Morphogenesis and lysis.
Phage GE1 structural proteins (ORF32/major capsid protein, ORF34/major tail protein, and ORF37/minor tail protein) and proteins involved in packaging (ORF40/terminase) and lysis (ORF29/holin and ORF38/holin) showed similarities to proteins found in other phages possessing the same morphological features, particularly c2, Q54, CB17, 5447, and 923 (34, 49). ORF35 possesses an immunoglobulin-like domain but did not match with other proteins in databases. Immunoglobulin-like proteins may be involved in host recognition (50). The structural protein profile of phage GE1 was determined by SDS-PAGE (Fig. 1B) and found to be highly related to that of phage c2 (37).
Comparison of prolate-headed phages.
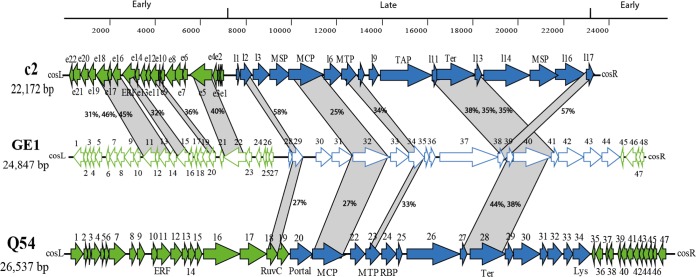
As demonstrated above, phage GE1 has similarities to L. lactis phages c2 and Q54, both prolate-headed phages. The genome organization and proteome of phage GE1 were specifically compared to those of L. lactis phages c2 and Q54. Figure 3 clearly confirms the relatedness of phage GE1 to c2 and Q54 phages. GE1 possesses 13 ORFs that align with those of phage c2 (amino acid identity of 25 to 58%). They are principally involved in replication (six ORFs) and morphogenesis (seven ORFs), and five of these seven proteins were also common to phage Q54 (amino acid identity of 27 to 44%). However, the genome organization of phage GE1 is closer to that of phage c2 than Q54, as illustrated by gene orientation (Fig. 3).
FIG 3.
Genomic comparison of GE1, Q54, and c2 phages. ORFs from GE1 showing similarity with other phages are linked by gray shading, and the percent amino acid identity indicated is representative of the aligned region.
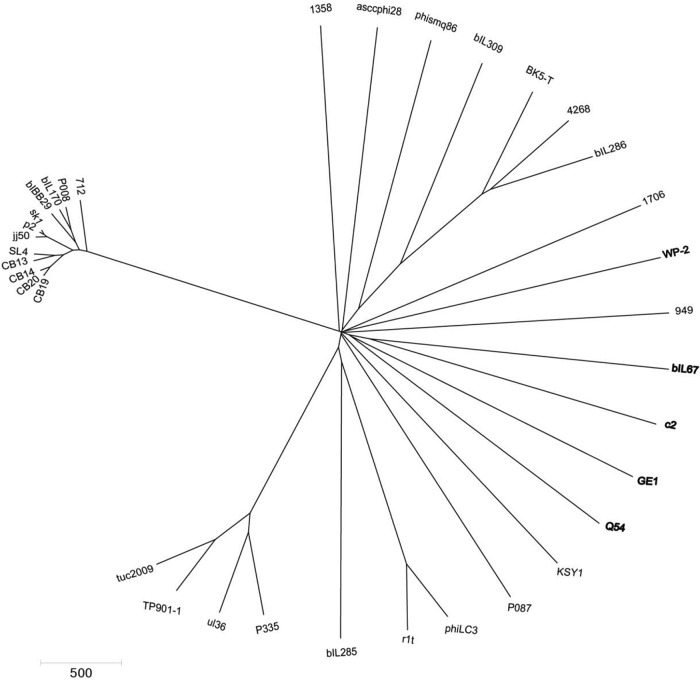
Phylogenetic studies of L. garvieae phages.
Finally, a phylogenetic analysis was performed using L. garvieae phages (GE1 and WP-2) as well as members of the 10 currently recognized groups of L. lactis phages to confirm the relationships described above. Figure 4 shows a proteomic phylogenetic tree constructed using MEGA5 software and the neighbor-joining method. This global comparison revealed that the two L. garvieae phages are distributed among L. lactis phages, indicating that they probably evolved from a common ancestor. The genomic comparison also demonstrated that phage GE1 originates at the same point as other prolate-headed phages, diverging, together with phage Q54, in a separate branch.
FIG 4.
Phylogenetic tree of different lactococcal phage proteomes determined using the neighbor-joining algorithm. The bar shows the difference between the phages in amino acids.
DISCUSSION
To date, L. garvieae phages have been poorly characterized. GE1 is the first L. garvieae siphophage to be characterized at the genomic level. Phage GE1 shares some characteristics with other lactococcal phages, specifically those infecting the food-grade bacterial species L. lactis. Phage GE1 has a prolate capsid (morphotype B2), a narrow host spectrum, and a burst size and latency period similar to those of other L. lactis phages.
Significant research attention has focused on phages infecting L. lactis in recent decades because these phages pose a significant risk to milk fermentation processes (51). The main interest in studying these phages is to understand their diversity and evolution to enable development of efficient phage control tools (52). L. lactis phages are currently grouped into 10 taxonomic groups based on morphology and genome analysis (17), and of these, phages 936/Sk1, c2, and P335 (all Siphoviridae phages) are the most frequently encountered in the dairy industry (53). Over 80 L. lactis phage genome sequences are now publicly available.
L. garvieae strains are found in raw milk and in a range of aquatic and terrestrial environments (21). Some L. garvieae strains may also be major components of the autochthonous microbial populations of certain artisanal cheeses (5). In fact, it is believed that the activity of L. garvieae strains may contribute to the final sensory characteristics of some dairy products (54). The adaptation of L. garvieae to the milk environment was likely enabled by the acquisition of certain plasmids (55). Not surprisingly, L. garvieae phages have similarities to L. lactis phages, also found in dairy environments.
Comparative genome and proteome analyses clearly demonstrated that phage GE1 is related to L. lactis phage c2 and Q54 groups (Fig. 3 and 4). Its genome size of 24,847 bp is also intermediate between those of c2 (22,172 bp) and Q54 (26,537 bp). The GE1 genome has cohesive extremities containing a cos site [GCAA(N9)AACT], differing from the c2 [TCAA(N4)AACT] and general lactococcal phage [TCAN(N4)NACT] consensus cos sites (38) but similar to the phage Q54 cos site [CCAA(N10)AACT]. The regions flanking the cos site contain several features common to other lactococcal phages such as the presence of direct and inverted repeats and an A/T-rich region.
A global analysis of the genomes of phages GE1, c2, and Q54 shows that c2 and GE1 are similarly organized, with two large modules in opposite orientations starting at the probable origin of replication. In phage GE1, an extra module (4 genes) was detected at the end of the late gene cluster, as in phage Q54. The predicted proteome of phage GE1, however, was closer to that of phage c2, with 13 ORFs sharing similarity. In the phage GE1 early-expressed module, five genes (ORFs 9, 11, 13, 14, and 22) code for proteins homologous to those found in L. lactis 936/Sk1-like phages. These latter phages, which also belong to the Siphoviridae family but have a small isometric capsid, are highly abundant in dairy environments (56), suggesting possible recombination events. Members of the rarely isolated Q54 group were proposed to be the result of past recombination events between 936/Sk1- and c2-like phages (37). Another striking feature of phage GE1 is the low level of similarity of its proteome with proteins in existing databases, confirming its novelty.
ACKNOWLEDGMENTS
We thank Barbara-Ann Conway (medical writer and editor) for editorial assistance. We also thank Alex Hynes for comments on the manuscript. We are grateful to Laetitia Bonifait and Caroline Duchaine for compost soil samples. We are grateful to Frederic Raymond and Jacques Corbeil for initial genome assembly.
S.M. acknowledges funding from the Natural Sciences and Engineering Research Council of Canada. S.M. holds a Tier 1 Canada Research Chair in Bacteriophages.
REFERENCES
- 1.Vendrell D, Balcázar JL, Ruiz-Zarzuela I, de Blas I, Gironés O, Múzquiz JL. 2006. Lactococcus garvieae in fish: a review. Comp Immunol Microbiol Infect Dis 29:177–198. doi: 10.1016/j.cimid.2006.06.003. [DOI] [PubMed] [Google Scholar]
- 2.Devriese LA, Hommez J, Laevens H, Pot B, Vandamme P, Haesebrouck F. 1999. Identification of aesculin-hydrolyzing streptococci, lactococci, aerococci and enterococci from subclinical intramammary infections in dairy cows. Vet Microbiol 70:87–94. doi: 10.1016/S0378-1135(99)00124-8. [DOI] [PubMed] [Google Scholar]
- 3.Aubin GG, Bémer P, Guillouzouic A, Crémet L, Touchais S, Fraquet N, Boutoille D, Reynaud A, Lepelletier D, Corvec S. 2011. First report of a hip prosthetic and joint infection caused by Lactococcus garvieae in a woman fishmonger. J Clin Microbiol 49:2074–2076. doi: 10.1128/JCM.00065-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Russo G, Iannetta M, D'Abramo A, Mascellino MT, Pantosti A, Erario L, Tebano G, Oliva A, D'Agostino C, Trinchieri V, Vullo V. 2012. Lactococcus garvieae endocarditis in a patient with colonic diverticulosis: first case report in Italy and review of the literature. New Microbiol 35:495–501. [PubMed] [Google Scholar]
- 5.Fortina MG, Ricci G, Acquati A, Zeppa G, Gandini A, Manachini PL. 2003. Genetic characterization of some lactic acid bacteria occurring in an artisanal protected denomination origin (PDO) Italian cheese, the Toma piemontese. Food Microbiol 20:397–404. doi: 10.1016/S0740-0020(02)00149-1. [DOI] [Google Scholar]
- 6.Fortina MG, Ricci G, Borgo F. 2009. A study of lactose metabolism in Lactococcus garvieae reveals a genetic marker for distinguishing between dairy and fish biotypes. J Food Prot 72:1248–1254. [DOI] [PubMed] [Google Scholar]
- 7.Aguado-Urda M, Cutuli MT, Blanco MM, Aspiroz C, Tejedor JL, Fernández-Garayzábal JF, Gibello A. 2010. Utilization of lactose and presence of the phospho-β-galactosidase (lacG) gene in Lactococcus garvieae isolates from different sources. Int Microbiol 13:189–193. [DOI] [PubMed] [Google Scholar]
- 8.Ferrario C, Ricci G, Milani C, Lugli GA, Ventura M, Eraclio G, Borgo F, Fortina MG. 2013. Lactococcus garvieae: where is it from? A first approach to explore the evolutionary history of this emerging pathogen. PLoS One 8:e84796. doi: 10.1371/journal.pone.0084796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Casalta E, Montel MC. 2008. Safety assessment of dairy microorganisms: the Lactococcus genus. Int J Food Microbiol 126:271–273. doi: 10.1016/j.ijfoodmicro.2007.08.013. [DOI] [PubMed] [Google Scholar]
- 10.Eraclio G, Ricci G, Fortina MG. 2015. Insertion sequence elements in Lactococcus garvieae. Gene 555:291–296. doi: 10.1016/j.gene.2014.11.019. [DOI] [PubMed] [Google Scholar]
- 11.Weinbauer MG. 2004. Ecology of prokaryotic viruses. FEMS Microbiol Rev 28:127–181. doi: 10.1016/j.femsre.2003.08.001. [DOI] [PubMed] [Google Scholar]
- 12.Veesler D, Cambillau C. 2011. A common evolutionary origin for tailed-bacteriophage functional modules and bacterial machineries. Microbiol Mol Biol Rev 75:423–433. doi: 10.1128/MMBR.00014-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Oliveira I, Almeida RCC, Hofer E, Almeida P. 2012. Bacteriophage amplification assay for detection of Listeria spp. using virucidal laser. Braz J Microbiol 43:1128–1136. doi: 10.1590/S1517-83822012000300040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hanlon GW. 2007. Bacteriophages: an appraisal of their role in the treatment of bacterial infections. Int J Antimicrob Agents 30:118–128. doi: 10.1016/j.ijantimicag.2007.04.006. [DOI] [PubMed] [Google Scholar]
- 15.Garneau JE, Moineau S. 2011. Bacteriophages of lactic acid bacteria and their impact on milk fermentations. Microb Cell Fact 10(Suppl 1):S20. doi: 10.1186/1475-2859-10-S1-S20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Moineau S, Lévesque C. 2005. Control of bacteriophages in industrial fermentations, p 285–286. In Kutter E, Sulakvelidze A (ed), Bacteriophages: biology and applications. CRC Press, Boca Raton, FL. [Google Scholar]
- 17.Park KH, Matsuoka S, Nakai T Muroga K. 1997. A virulent bacteriophage of L. garvieae (formerly Enterococcus seriolicida) isolated from yellowtail Seriola quinqueradiata. Dis Aquat Organ 29:145–149. doi: 10.3354/dao029145. [DOI] [Google Scholar]
- 18.Ghasemi SM, Bouzari M, Shaykh Baygloo N, Chang H-I. 2014. Insights into new bacteriophages of Lactococcus garvieae belonging to the family Podoviridae. Arch Virol 159:2909–2915. doi: 10.1007/s00705-014-2142-z. [DOI] [PubMed] [Google Scholar]
- 19.Nakai T, Sugimoto R, Park KH, Matsuoka S, Mori K, Nishioka T, Maruyama K. 1999. Protective effects of bacteriophage on experimental Lactococcus garvieae infection in yellowtail. Dis Aquat Organ 37:33–41. doi: 10.3354/dao037033. [DOI] [PubMed] [Google Scholar]
- 20.Ghasemi SM, Bouzari M, Yoon BH, Chang H-I. 2014. Comparative genomic analysis of Lactococcus garvieae phage WP-2, a new member of Picovirinae subfamily of Podoviridae. Gene 551:222–229. doi: 10.1016/j.gene.2014.08.060. [DOI] [PubMed] [Google Scholar]
- 21.Ferrario C, Ricci G, Borgo F, Rollando A, Fortina MG. 2012. Genetic investigation within Lactococcus garvieae revealed two genomic lineages. FEMS Microbiol Lett 332:153–161. doi: 10.1111/j.1574-6968.2012.02591.x. [DOI] [PubMed] [Google Scholar]
- 22.El Haddad L, Moineau S. 2013. Characterization of a novel Panton-Valentine leukocidin (PVL)-encoding staphylococcal phage and its naturally PVL-lacking variant. Appl Environ Microbiol 79:2828–2832. doi: 10.1128/AEM.03852-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Émond É, Holler BJ, Boucher I, Vandenbergh PA, Vedamuthu ER, Kondo JK, Moineau S. 1997. Phenotypic and genetic characterization of the bacteriophage abortive infection mechanism AbiK from Lactococcus lactis. Appl Environ Microbiol 63:1274–1283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Samson JE, Moineau S. 2010. Characterization of Lactococcus lactis phage 949 and comparison with other lactococcal phages. Appl Environ Microbiol 76:6843–6852. doi: 10.1128/AEM.00796-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Moineau S, Durmaz E, Pandian S, Klaenhammer TR. 1993. Differentiation of two abortive mechanisms by using monoclonal antibodies directed toward lactococcal bacteriophage capsid proteins. Appl Environ Microbiol 59:208–212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Deveau H, Van Calsteren MR, Moineau S. 2002. Effect of exopolysaccharides on phage-host interactions in Lactococcus lactis. Appl Environ Microbiol 68:4364–4369. doi: 10.1128/AEM.68.9.4364-4369.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Boisvert S, Laviolette F, Corbeil J. 2010. Ray: simultaneous assembly of reads from a mix of high-throughput sequencing technologies. J Comput Biol 17:1519–1533. doi: 10.1089/cmb.2009.0238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O. 2008. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lowe TM, Eddy SR. 1997. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. doi: 10.1093/nar/25.5.0955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Laslett D, Canback B. 2004. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res 32:11–16. doi: 10.1093/nar/gkh152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chen L, Xiong Z, Sun L, Yang J, Jin Q. 2012. VFDB 2012 update: toward the genetic diversity and molecular evolution of bacterial virulence factors. Nucleic Acids Res 40:D641–D645. doi: 10.1093/nar/gkr989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chakraborty A, Ghosh S, Chowdhary G, Maulik U, Chakrabarti S. 2012. DBETH: a database of bacterial exotoxins for human. Nucleic Acids Res 40:D615–D620. doi: 10.1093/nar/gkr942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Deveau H, Labrie SJ, Chopin M-C, Moineau S. 2006. Biodiversity and classification of lactococcal phages. Appl Environ Microbiol 72:4338–4346. doi: 10.1128/AEM.02517-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Dupuis ME, Moineau S. 2010. Genome organization and characterization of the virulent lactococcal phage 1358 and its similarities to Listeria phages. Appl Environ Microbiol 76:1623–1632. doi: 10.1128/AEM.02173-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Fortier LC, Bransi A, Moineau S. 2006. Genome sequence and global gene expression of Q54, a new phage species linking the 936 and c2 phage species of Lactococcus lactis. J Bacteriol 188:6101–6114. doi: 10.1128/JB.00581-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Perrin R, Billard P, Branlant C. 1997. Comparative analysis of the genomic DNA terminal regions of the lactococcal bacteriophages from species c2. Res Microbiol 148:573–583. doi: 10.1016/S0923-2508(97)88081-3. [DOI] [PubMed] [Google Scholar]
- 39.Cue D, Feiss M. 1993. The role of cosB, the binding site for terminase, the DNA packaging enzyme of bacteriophage lambda, in the nicking reaction. J Mol Biol 234:594–609. doi: 10.1006/jmbi.1993.1614. [DOI] [PubMed] [Google Scholar]
- 40.Brussow H. 2001. Phages of dairy bacteria. Annu Rev Microbiol 55:283–303. doi: 10.1146/annurev.micro.55.1.283. [DOI] [PubMed] [Google Scholar]
- 41.Lubbers MW, Waterfield NR, Beresford TP, Le Page RW, Jarvis AW. 1995. Sequencing and analysis of the prolate-headed lactococcal bacteriophage c2 genome and identification of the structural genes. Appl Environ Microbiol 61:4348–4356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Lubbers MW, Ward LJ, Beresford TP, Jarvis BD, Jarvis AW. 1994. Sequencing and analysis of the cos region of the lactococcal bacteriophage c2. Mol Gen Genet 245:160–166. [DOI] [PubMed] [Google Scholar]
- 43.Mahony J, Kot W, Murphy J, Ainsworth S, Neve H, Hansen LH, Heller KJ, Sørensen SJ, Hammer K, Cambillau C, Vogensen FK, Van Sinderen D. 2013. Investigation of the relationship between lactococcal host cell wall polysaccharide genotype and 936 phage receptor binding protein phylogeny. Appl Environ Microbiol 79:4385–4392. doi: 10.1128/AEM.00653-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Belfort M, Bonocora RP. 2014. Homing endonucleases: from genetic anomalies to programmable genomic clippers. Methods Mol Biol 1123:1–26. doi: 10.1007/978-1-62703-968-0_1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Mahony J, Deveau H, Mc Grath S, Ventura M, Canchaya C, Moineau S, Fitzgerald GF, Van Sinderen D. 2006. Sequence and comparative genomic analysis of lactococcal bacteriophages jj50, 712 and P008: evolutionary insights into the 936 phage species. FEMS Microbiol Lett 261:253–261. doi: 10.1111/j.1574-6968.2006.00372.x. [DOI] [PubMed] [Google Scholar]
- 46.Freist W, Gauss DH, Soll D, Lapointe J. 1997. Glutamyl-tRNA synthetase. Biol Chem 378:1313–1329. [PubMed] [Google Scholar]
- 47.Scaltriti E, Polverini E, Grolli S, Eufemi E, Moineau S, Cambillau C, Ramoni R. 2013. The DNA binding mechanism of a SSB protein from Lactococcus lactis siphophage p2. Biochim Biophys Acta 1834:1070–1076. doi: 10.1016/j.bbapap.2013.02.014. [DOI] [PubMed] [Google Scholar]
- 48.Scaltriti E, Tegoni M, Rivetti C, Launay H, Masson J-Y, Magadan AH, Tremblay D, Moineau S, Ramoni R, Lichière J, Campanacci V, Cambillau C, Ortiz-Lombardía M. 2009. Structure and function of phage p2 ORF34(p2), a new type of single-stranded DNA binding protein. Mol Microbiol 73:1156–1170. doi: 10.1111/j.1365-2958.2009.06844.x. [DOI] [PubMed] [Google Scholar]
- 49.Rakonjac J, O'Toole PW, Lubbers M. 2005. Isolation of lactococcal prolate phage-phage recombinants by an enrichment strategy reveals two novel host range determinants. J Bacteriol 187:3110–3121. doi: 10.1128/JB.187.9.3110-3121.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Fraser JS, Maxwell KL, Davidson AR. 2007. Immunoglobulin-like domains on bacteriophage: weapons of modest damage? Curr Opin Microbiol 10:382–387. doi: 10.1016/j.mib.2007.05.018. [DOI] [PubMed] [Google Scholar]
- 51.Mahony J, van Sinderen D. 2014. Current taxonomy of phages infecting lactic acid bacteria. Front Microbiol 5:7. doi: 10.3389/fmicb.2014.00007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Samson JE, Moineau S. 2013. Bacteriophages in food fermentations: new frontiers in a continuous arms race. Annu Rev Food Sci Technol 4:347–368. doi: 10.1146/annurev-food-030212-182541. [DOI] [PubMed] [Google Scholar]
- 53.Mahony J, Bottacini F, van Sinderen D, Fitzgerald GF. 2014. Progress in lactic acid bacterial phage research. Microb Cell Fact 13(Suppl 1):S1. doi: 10.1186/1475-2859-13-S1-S1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Fernández E, Alegría Á, Delgado S, Mayo B. 2010. Phenotypic, genetic and technological characterization of Lactococcus garvieae strains isolated from a raw milk cheese. Int Dairy J 20:142–148. doi: 10.1016/j.idairyj.2009.11.004. [DOI] [Google Scholar]
- 55.Flórez AB, Mayo B. 2015. The plasmid complement of the cheese isolate Lactococcus garvieae IPLA 31405 revealed adaptation to the dairy environment. PLoS One 10:e0126101. doi: 10.1371/journal.pone.0126101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Mahony J, Murphy J, Van Sinderen D. 2012. Lactococcal 936-type phages and dairy fermentation problems: from detection to evolution and prevention. Front Microbiol 3:335. doi: 10.3389/fmicb.2012.00335. [DOI] [PMC free article] [PubMed] [Google Scholar]