In this study, Sienski et al. characterize CG9754/Silencio as an essential piRNA pathway factor that is required for Piwi's nuclear function in guiding the transcriptional silencing of transposons. These results provide novel insight into the transcriptional silencing process downstream from Piwi and the regulation of piRNA-guided heterochromatin formation.
Keywords: H3K9 methylation, Piwi, transposon silencing, heterochromatin formation, piRNA pathway, transcriptional silencing
Abstract
The repression of transposable elements in eukaryotes often involves their transcriptional silencing via targeted chromatin modifications. In animal gonads, nuclear Argonaute proteins of the PIWI clade complexed with small guide RNAs (piRNAs) serve as sequence specificity determinants in this process. How binding of nuclear PIWI–piRNA complexes to nascent transcripts orchestrates heterochromatin formation and transcriptional silencing is unknown. Here, we characterize CG9754/Silencio as an essential piRNA pathway factor that is required for Piwi-mediated transcriptional silencing in Drosophila. Ectopic targeting of Silencio to RNA or DNA is sufficient to elicit silencing independently of Piwi and known piRNA pathway factors. Instead, Silencio requires the H3K9 methyltransferase Eggless/SetDB1 for its silencing ability. In agreement with this, SetDB1, but not Su(var)3-9, is required for Piwi-mediated transcriptional silencing genome-wide. Due to its interaction with the target-engaged Piwi–piRNA complex, we suggest that Silencio acts as linker between the sequence specificity factor Piwi and the cellular heterochromatin machinery.
In fungi, plants, and animals, nuclear Argonaute proteins target nascent transcripts—RNAs that are still attached to their originating DNA locus via the transcribing RNA polymerase. This provides an opportunity for small RNA silencing pathways to connect to the various cellular chromatin-modifying activities for the sequence-specific formation of heterochromatin, typically at transposable element (TE) insertions (Castel and Martienssen 2013).
Pioneering work in fission yeast has identified methylation of histone H3 at Lys9 (H3K9me2/3) and histone deacetylation as two major hallmarks that are required for small RNA-guided silencing and heterochromatin formation (Nakayama et al. 2001; Hall et al. 2002; Volpe et al. 2002; Yamada et al. 2005). A sequential order of events downstream from the recruitment of the Schizosaccharomyces pombe Ago1/siRNA complex to nascent RNA of centromeric repeats has been described. According to this, the Ago1/siRNA complex recruits the H3K9 methyltransferase Clr4 to chromatin, which results in H3K9 methylation (Noma et al. 2004; Verdel et al. 2004). This is believed to be a binding platform for the chromodomain-containing protein Swi6/HP1, which in turn recruits the major histone deacetylase and remodeling complex SHREC (Sugiyama et al. 2007). The final outcome of these events is the establishment of a nucleosome-dense region with low histone turnover, which effectively prevents the recruitment of RNA polymerase II (Pol II) to transcription initiation sites and hence transcription as such (e.g., Aygun et al. 2013).
H3K9 methylation is also a hallmark of small RNA-guided heterochromatin formation in plants, ciliates, and multicellular animals. In animal gonads, many TE insertions are methylated at H3K9 residues in a PIWI–piRNA (PIWI clade Argonaute protein complexed with small guide RNA)-dependent manner, and loss of the piRNA pathway results in the selective loss of H3K9me3 at targeted TE insertions and in their desilencing (Wang and Elgin 2011; Sienski et al. 2012; Le Thomas et al. 2013; Rozhkov et al. 2013; Pezic et al. 2014). In analogy to heterochromatin formation in S. pombe, the PIWI–piRNA complex likely provides the sequence-specific information to target chromatin-modifying enzymes to TE loci for their silencing. However, how recruitment of nuclear PIWI clade proteins to nascent transcripts leads to heterochromatin formation is unknown.
In S. pombe, the adaptor protein Stc1 is reported to bridge the Ago1/siRNA complex to the methyltransferase Clr4 (Bayne et al. 2010), and the chromodomain protein Chp1 is implicated in tethering the target-bound Ago1/siRNA complex to chromatin (Ishida et al. 2012). However, neither of these factors is conserved in animals. In turn, two proteins with largely unknown functions that are required for piRNA-guided transcriptional silencing in animals (Gtsf1/Asterix and Maelstrom) (Sienski et al. 2012; Donertas et al. 2013; Muerdter et al. 2013; Ohtani et al. 2013) are absent in S. pombe, highlighting significant molecular differences in these two systems. Moreover, the Piwi–piRNA complex fails to mediate transcriptional silencing if recruited artificially (not involving a piRNA–target interaction) to a reporter RNA in a Drosophila cell culture model that expresses a fully functional piRNA pathway centered on nuclear Piwi (Post et al. 2014). Whether this reflects the necessity for a conformational change in Piwi upon piRNA–target binding or is due to the requirement of a minimum threshold of target-engaged Piwi molecules to initiate silencing is unclear. Similarly unclear is the identity of the factors that establish and maintain piRNA-specified heterochromatin. For example, evidence for an involvement of the two H3K9 methyltransferases Su(var)3-9 and Eggless/SetDB1 has been reported (Fritsch et al. 2010; Rangan et al. 2011; Huang et al. 2013; Bulut-Karslioglu et al. 2014), but it is not known which roles these enzymes have in piRNA-guided silencing and what they contribute to heterochromatin formation at TE insertions.
Here, we establish an in vivo assay to test the sufficiency of piRNA pathway factors in mediating transcriptional silencing and heterochromatin formation. This led to the discovery of CG9754/Silencio (Czech et al. 2013; Handler et al. 2013; Muerdter et al. 2013) as a key factor that links target-engaged Piwi to the cellular heterochromatin machinery. Using this assay and genome-wide profiling of the H3K9me3 mark, we demonstrate an integral role of Eggless/SetDB1 in piRNA-guided silencing and show that Su(var)3-9 contributes to spreading of the H3K9me3 mark to TE-flanking domains. Our work provides an entry point into the mechanistic dissection of piRNA-guided silencing and clarifies the roles of H3K9 methyltransferases in this process.
Results
Recruitment of CG9754 to RNA or DNA elicits potent transcriptional silencing
Several studies indicate that binding of the Piwi–piRNA complex to a nascent target RNA causes heterochromatin formation and transcriptional silencing (e.g., Sarot et al. 2004; Sienski et al. 2012; Le Thomas et al. 2013; Post et al. 2014). In agreement with the study by Lau and colleagues (Post et al. 2014), we found that recruitment of Piwi to a reporter construct via a sequence stretch complementary to endogenous piRNAs elicits potent reporter silencing in a Piwi-dependent and target orientation-dependent manner (Supplemental Fig. S1A). In contrast, artificial recruitment (tethering) of Piwi via the λN-boxB system (thereby bypassing the requirement for piRNA target sites) to a reporter RNA expressed from a transiently transfected plasmid does not affect reporter expression (Supplemental Fig. S1B; Post et al. 2014).
To determine whether the λN-boxB assay failed due to the reporter plasmid not being properly chromatinized, we established an in vivo RNA-tethering system in the Drosophila ovary (Fig. 1A; Supplemental Fig. S1C,D). This assay builds on the α-tubulin promoter driving the ubiquitous expression of a GFP reporter, which harbors five boxB sites in its 3′ untranslated region (UTR), ∼1.5 kb downstream from the transcriptional start site (TSS). Expression of λN-HA-tagged Gawky/GW182 (a cytoplasmic factor that elicits post-transcriptional gene silencing) (Jonas and Izaurralde 2015) with the germline-specific MTD-Gal4 driver leads to potent reporter silencing in germline but not somatic cells of the ovary (Fig. 1A,B). Tethering of Piwi to the reporter instead fails to induce silencing despite λN-tagged Piwi being abundant in the nucleus (Fig. 1A,B; Supplemental Fig. S1C). This indicates that Piwi is only able to signal to the downstream silencing machinery if engaged with a target via its bound piRNA.
Figure 1.
CG9754 elicits transcriptional silencing upon recruitment to nascent RNA or DNA. (A) The cartoon depicts the GFP-5xboxB reporter used for RNA tethering. It harbors the ubiquitous α-tubulin promoter, the GFP-coding sequence, and a 3′ UTR harboring five boxB sites. The confocal image below depicts GFP fluorescence signal in an egg chamber expressing the GFP-boxB reporter. Throughout this study, we converted GFP signal to monochrome images with an inverted black/white color scheme for displaying optimal contrast. Soma and germline tissue is indicated. Images at the right depict the expression of λN-HA fusion proteins under control of the germline-specific MTD-Gal4 driver (λN-GW182: cytoplasmic; λN-Piwi: nuclear-enriched). (B) GFP fluorescence in egg chambers expressing the ubiquitous GFP-boxB reporter and the indicated λN fusions in the germline. (C) Simplified cartoon of the linear piRNA pathway feeding into Piwi, also indicating key players involved at the indicated steps. (TGS) Transcriptional gene silencing. (D) GFP fluorescence in egg chambers expressing the ubiquitous GFP-boxB reporter and the indicated λN fusions in the germline. Expression of LacI-CG9754 served as a negative control. (E) Shown are changes in GFP-boxB steady-state RNA levels in ovaries expressing the indicated λN fusions (normalized to rp49). n = 3. Error bars indicate SD. (F) Shown is RNA Pol II occupancy at the indicated loci determined by chromatin immunoprecipitation (ChIP) followed by quantitative PCR using ovaries expressing the indicated λN fusions. n = 3. Error bars indicate SD. Signals were normalized to those at the TSS of the act 5C gene. arrestin is not expressed in ovaries. (G) The cartoon depicts the 8xlacO-GFP reporter used for DNA tethering. This reporter is expressed in germline and somatic cells of the ovary (nanos promoter coupled to intronic piwi enhancer) (Hayashi et al. 2014). The confocal image below depicts GFP fluorescence signal of the reporter. The middle image depicts the expression of LacI-CG9754 under control of the germline-specific MTD-Gal4 driver. Images at the right show GFP fluorescence signal of the reporter in egg chambers expressing the indicated LacI fusions in the germline. Note that LacI-Piwi accumulates mostly in the cytoplasm (potentially due to the reported oligomerization of LacI) (Supplemental Fig. S1E) and thus cannot be interpreted unambiguously. (H) Shown are GFP fluorescence signals in egg chambers expressing the GFP-boxB reporter, λN-CG9754, and the indicated shRNA constructs in the germline.
We reasoned that an adaptor protein might link target-engaged Piwi to the silencing machinery and therefore asked whether any factor that has been implicated in Piwi-mediated transcriptional silencing (Fig. 1C) triggers repression if recruited ectopically to the reporter. While Gtsf1/Asterix and Maelstrom were inert in this assay, tethering of CG9754, an uncharacterized protein with a suggested role in Piwi-mediated silencing (Czech et al. 2013; Handler et al. 2013; Muerdter et al. 2013), results in potent reporter repression that was dependent on the λN peptide on CG9754 (Fig. 1D).
Silencing induced by tethered CG9754, but not by GW182, results in strongly reduced steady-state reporter RNA levels (Fig. 1E) and an approximately threefold reduction of RNA Pol II occupancy at the α-tubulin promoter of the reporter construct. The endogenous α-tubulin promoter instead (identical in sequence except for a small stretch of SNPs) exhibits constant Pol II occupancy (Fig. 1F). These findings indicate that CG9754 is sufficient to induce transcriptional silencing if targeted to a nascent RNA. In support of this, ectopic recruitment of CG9754 to DNA upstream of the TSS of a different GFP reporter leads to similarly potent silencing (Fig. 1G; Supplemental Fig. S1E,F). Also in this assay, Piwi, Gtsf1, and Maelstrom are incapable of inducing silencing.
We next combined the λN-CG9754-mediated silencing assay with potent germline-specific RNAi to establish a genetic hierarchy between CG9754 and other piRNA pathway factors (Supplemental Fig. S1G). Depletion of λN-CG9754/CG9754 results in robust reporter derepression, demonstrating the feasibility of this approach (Fig. 1H). Depletion of Piwi, Zucchini (an essential piRNA biogenesis factor for Piwi), Gtsf1/Asterix, or Maelstrom instead had no impact (Fig. 1H). Considering that CG9754 is required for all aspects of Piwi-mediated silencing (see below), this factor acts most likely downstream from all known piRNA pathway factors.
Taken together, our data indicate that recruiting CG9754 to chromatin—directly to DNA or indirectly via a nascent RNA—elicits transcriptional silencing of the target locus that is reminiscent of piRNA-guided silencing yet does not require known piRNA pathway factors.
CG9754 is an essential piRNA pathway factor in the ovarian germline and soma
CG9754 was identified genetically as a factor required for TE silencing in the ovary using transgenic RNAi (Czech et al. 2013; Handler et al. 2013; Muerdter et al. 2013). We used CRISPR/Cas9 (Jinek et al. 2012; Gokcezade et al. 2014) to generate two mutant alleles of CG9754 that harbor nucleotide insertions/deletions in the coding sequence close to the N terminus. Both alleles result in frameshifts and are expected to be strong mutant alleles (Fig. 2A). In support of this, CG9754 protein is not detectable in ovarian lysates from homozygous mutants by Western blot analysis using a monoclonal antibody raised against CG9754 (Fig. 2B).
Figure 2.
Genetic characterization of CG9754. (A) Cartoon of the CG9754 protein showing the large unstructured N-terminal region, predicted secondary structure elements at the C terminus, and a predicted nuclear localization signal (NLS). Frameshift positions caused by the guide RNA-induced insertions/deletions and the molecular nature of two alleles (CG9754g1 and CG9754g2) are indicated below. (B) Western blot analysis showing protein levels of CG9754 (top) and Piwi (bottom) in white1118, CG9754g1/g1, CG9754g2/g2, and transheterozygous CG9754g1/g2 ovaries. Tubulin served as a loading control. The asterisk marks an unspecific band. Note that CG9754 migrates at 100 kDa instead of its predicted molecular weight of 61 kDa. (C) Shown are ovarian morphologies from flies of the indicated genotypes. Bar, 500 µm. (D) Depicted are egg-laying rates of females of the indicated genotype (right) and hatching rates of the corresponding laid eggs (left). (E) Bar plot showing fold changes in steady-state RNA levels of somatic (mdg1 and springer) and germline (HeT-A, burdock, and bel) TEs in total ovarian RNA from the indicated genotypes (normalized to rp49). n = 2. Error bars indicate SD. (F) Confocal sections of egg chambers from flies of the indicated genotypes stained simultaneously for CG9754 and Piwi.
Flies homozygous for either mutant allele as well as transheterozygous flies are viable yet exhibit oogenesis defects and severe fertility defects (Fig. 2C,D). In this, they resemble piwi mutant flies (Cox et al. 2000). In addition, patterns of TE derepression (for soma- and germline-specific TEs) are very similar between CG9754 mutants and piwi mutants (Fig. 2E). In support of previous findings (Handler et al. 2013), complete loss of CG9754 does not affect levels and nuclear accumulation of Piwi in germline and somatic cells (Fig. 2B,F), indicating that CG9754 is specifically required for Piwi-mediated transcriptional target silencing (also, localization and levels of Aub and Ago3 in germline cells are unaffected) (Supplemental Fig. S2).
CG9754/Silencio is required for H3K9 methylation and transcriptional silencing of Piwi-repressed TE insertions
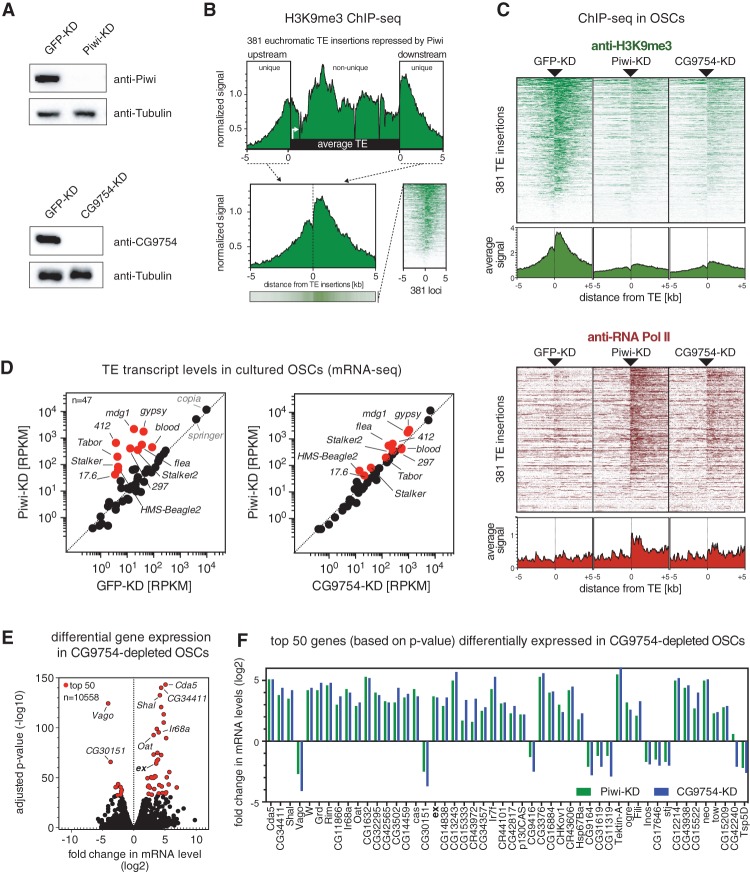
In order to directly probe a role of CG9754 in heterochromatin formation and transcriptional silencing downstream from Piwi, we used cultured ovarian somatic cells (OSCs) (Niki et al. 2006; Saito et al. 2009). The detailed knowledge of their TE insertion landscape (Sienski et al. 2012) and the ability to efficiently deplete factors of interest using RNAi (Fig. 3A) make these cells an ideal system for genome-wide studies. To determine changes in H3K9me3 patterns and RNA Pol II occupancy, we focused on 381 euchromatic insertions of the 11 TE families that are strongly repressed by the piRNA pathway in OSCs. The genomic regions flanking these TE insertions are genome-unique anchor points that allow extracting levels and spreading of the H3K9 methyl mark as well as the transcriptional activity of each individual TE insertion (Fig. 3B).
Figure 3.
CG9754 is a core piRNA pathway factor. (A) Western blots showing protein levels of Piwi and CG9754 in the respective siRNA transfected OSCs. (KD) Knockdown. (B) Shown is the metaprofile of H3K9me3 levels calculated from 381 euchromatic insertions of Piwi-repressed TEs in OSCs. Also shown are average signals within the upstream and downstream 5 kb flanking these insertions. For all analyses, we display exclusively the flanking regions, which serve as genome-unique identifications of these TE insertions. In the heat map at the right, signals at all 381 TE insertions were sorted for decreasing intensity. (C) Heat maps showing H3K9me3 levels (green) or RNA Pol II occupancy (red) within the 10 kb flanking the TE insertions (as described in B) in OSCs depleted for GFP (control), Piwi, or CG9754. All heat maps were sorted for decreasing H3K9me3 signal in GFP knockdown cells. The respective average signals are shown below each heat map (GFP knockdown and Piwi knockdown data from Sienski et al. 2012). (D) Scatter plot showing expression levels (reads per kilobase per million sequenced reads) of TEs in OSCs depleted for the indicated factors. TEs in red are those whose abundance changes more than fourfold in Piwi-depleted cells. (E) Scatter plot showing fold change (log2) in mRNA levels and corresponding P-values (adjusted for multiple testing with the Benjamini-Hochberg procedure; −log10) in OSCs depleted for CG9754. The top 50 differentially expressed genes are in red. (F) Shown are the fold changes in mRNA levels of the top 50 (according to P-value as in E) deregulated genes in CG9754-depleted OSCs in Piwi-depleted cells.
Depletion of CG9754 leads to a strong reduction in H3K9me3 levels at Piwi-repressed TE insertions and to directional transcriptional readthrough into flanking genomic regions (Fig. 3C). The resulting patterns are highly similar to those obtained from Piwi-depleted cells (Sienski et al. 2012). Similarly, patterns of TE derepression as well as changes in the expression of protein-coding genes are highly correlated between CG9754-depleted versus Piwi-depleted cells (Fig. 3D–F). As observed previously (Sienski et al. 2012), many of the up-regulated genes are flanked by Piwi-repressed TE insertions, leading to their silencing in wild-type cells by the spreading of heterochromatin (e.g., expanded) (Fig. 3F). Altogether, our data from flies (Fig. 2) and OSCs (Fig. 3) assign a specific and essential role to CG9754 in local heterochromatin formation and transcriptional silencing downstream from Piwi. We therefore named CG9754 “silencio.”
CG9754/Silencio associates with target-engaged Piwi–piRNA complexes
As Silencio is required for Piwi-mediated silencing and induces silencing if tethered ectopically to an RNA, we tested for an interaction between Piwi and Silencio. Reciprocal coimmunoprecipitation (co-IP) experiments indicate that a large part of the nuclear Silencio pool interacts with Piwi (Fig. 4A). Because of Silencio's silencing capacity, we wondered whether it interacts with a distinct subset of nuclear Piwi. We therefore sequenced small RNAs isolated from Silencio immunoprecipitations as well as from Piwi immunoprecipitations and compared both with the population of cellular piRNAs from total small RNA libraries. The majority of the small RNAs obtained from the Silencio immunoprecipitation resemble mature piRNAs, as evidenced by their average length of 26.7 nucleotides (nt) and their tendency for a terminal 5′ uridine (∼80%). Interestingly, piRNAs originating from 3′ UTRs of cellular mRNAs (Robine et al. 2009; Saito et al. 2009), but not from TEs, are specifically depleted in the Silencio immunoprecipitation library but not in the Piwi immunoprecipitation library (Fig. 4B; Supplemental Fig. S3A,B). As 3′ UTR-derived piRNAs have no widespread role in target identification due to the absence of highly complementary RNAs, these results were a first hint that target-engaged Piwi–piRNA complexes might be enriched in Silencio immunoprecipitations.
Figure 4.
CG9754/Silencio interacts with target-engaged Piwi–piRNA complexes. (A) Western blots showing reciprocal co-IP of endogenous CG9754 with endogenous Piwi from nuclear OSC lysate (relative amount loaded in immunoprecipitation lanes; 10×; mouse IgGs were used as negative control). (B) Scatter plot showing levels of 3′ UTR-derived piRNAs in Piwi (green) and CG9754 (blue) immunoprecipitates in comparison with levels in a total small RNA library (shown are the top 100 genic piRNA sources in OSCs; values were normalized to 1 million sequenced reads per library). Each point shows the average value of two biological replicates, and error bars represent SEM. (C) Bar graphs showing size distribution and first nucleotide composition of small RNAs mapping in sense (top) or antisense (bottom) orientation to TEs regulated by Piwi in OSCs (note the different scales for sense and antisense). Reads from total small RNA (left), Piwi immunoprecipitation (middle), and CG9754 immunoprecipitation (right) are compared. (D) Cartoon of a Piwi–piRNA complex bound to a target RNA. (Below) Sequence logos indicating the 5′ and 3′ nucleotide bias of TE antisense reads (piRNAs) and sense reads (target RNAs) are shown. (E) Bar graphs displaying the fold enrichment of sense reads in the indicated immunoprecipitation libraries over the total small RNA library on all annotated TEs. n = 102. Two independent experiments are shown; error bars represent SEM. The black bars show corresponding piRNA levels in OSCs. The heat map below displays the fold changes in TE RNA levels upon Piwi depletion in OSCs. (F) Shown are density plots of normalized small RNA reads from CG9754 immunoprecipitation (sense), Piwi immunoprecipitation (sense), and total small RNAs (antisense) mapping to the mdg1 element. The piRNA-free gap in mdg1 is depicted.
Upon closer inspection of the small RNA populations in the Silencio immunoprecipitation, we noticed a considerable portion of RNA reads that map in sense orientation to Piwi-repressed TE sequences. These TE sense reads are specifically enriched in Silencio immunoprecipitations but not in Piwi immunoprecipitations from the same nuclear OSC lysate (Fig. 4C). In this context, it is important to note that TE-derived piRNAs in OSCs are almost exclusively antisense to the active transposon transcript (e.g., mdg1, 99.9%; gypsy, 99.4%; 412, 99.99%).
Several observations suggest that the observed sense RNAs represent footprints of endogenous piRNA targets. First, their size profile mirrors that of mature piRNAs yet, on average, is 1–2 nt larger, suggesting ∼1-nt extensions on both sides of a small RNA duplex (Fig. 4C); whether this size profile of sense reads also exists in vivo or is generated by nucleases during the experiment is currently unclear. Second, the sense reads lack a uridine bias at the 5′ end, arguing that they are not mature piRNAs (Fig. 4C,D). Instead, their three terminal nucleotides at the 3′ end are enriched for adenosine residues, which would be expected due to the 5′ uridine bias of complementary piRNAs. Third, the sense reads map predominantly to TEs that are repressed by Piwi–piRNA complexes in OSCs (e.g., gypsy, mdg1, 412, and Tabor) (Fig. 4E). Finally, the sense read profiles are essentially a mirror image of antisense piRNA populations. piRNAs targeting mdg1, for example, exhibit a gap in the 5′ portion of the TE sequence, and this gap is also devoid of corresponding sense reads (Fig. 4F). These observations suggest that target-engaged Piwi–piRNA complexes interact with Silencio, further supporting a central role for this factor in the effector step of piRNA-guided transcriptional silencing.
CG9754/Silencio induces heterochromatin formation, and its transcriptional silencing capacity depends on SetDB1/Eggless
A hallmark of piRNA-guided transcriptional silencing in flies and mice is the local establishment of an H3K9me3 domain. To test whether experimental tethering of Silencio to the RNA of the GFP-boxB reporter also leads to H3K9me3, we performed chromatin immunoprecipitation (ChIP) followed by quantitative PCR (qPCR) from ovaries where λN alone, λN-GW182, λN-Piwi, or λN-Silencio was expressed in germline cells (Fig. 5A). Only tethering of Silencio results in H3K9me3 specifically at the reporter locus (TSS and internal) to levels that are comparable with those at the light locus, which maps to heterochromatin. In contrast, the endogenous α-tubulin promoter that is identical in sequence to the reporter except for a few SNPs does not change methylation status. Surprisingly, tethering of HP1a/Su(var)205, a factor that is known to induce potent transcriptional silencing and heterochromatin formation if targeted to DNA (Ayyanathan et al. 2003; Li et al. 2003), does not trigger silencing or H3K9me3 (Fig. 5A).
Figure 5.
Silencing and heterochromatin formation by CG9754/Silencio depends on the H3K9 methyltransferase SetDB1. (A) Shown are GFP fluorescence signals in egg chambers expressing the GFP-boxB reporter and the indicated λN fusions. The bar diagram shows corresponding H3K9me3 levels at the indicated loci, measured by ChIP followed by qPCR (values were normalized to a gene desert; light is a heterochromatic control locus). Displayed is the mean of three independent experiments. Error bars indicate SD. The cartoons detail the reporter locus and the endogenous α-tubulin locus in scale and depict the location of used primer pairs. (B) Everything is as in A except for the lacO-GFP reporter experiment. (C) Shown are GFP fluorescence signals in egg chambers expressing the GFP-boxB reporter, λN-CG9754, and the indicated shRNA constructs in the germline [two independent shRNA lines for Su(var)3-9 were used with similar results]. (D) As in C except for the lacO-GFP reporter.
We obtained very similar results when targeting Silencio to DNA upstream of the lacO-GFP reporter (Fig. 5B). In this assay, HP1a was equally potent to induce silencing and H3K9me3. This strongly indicates that Silencio not only orchestrates heterochromatin formation (like HP1a) but also integrates unknown functions that allow it to effectively signal to the underlying DNA locus when targeted to a nascent RNA, which is only transiently associated with chromatin via the transcribing polymerase.
The trimethylation of H3K9 triggered by Silencio recruitment to the GFP reporter prompted us to investigate which histone methyltransferase mediates this modification. To do so, we depleted the Drosophila H3K9 methyltransferases Su(var)3-9 and Eggless/SetDB1 in the germline using transgenic RNAi in the background of the GFP-boxB reporter silenced by λN-Silencio. Loss of SetDB1, but not of Su(var)3-9, results in derepression of the boxB reporter in the germline (Fig. 5C). Similar results were obtained when Silencio was targeted to the reporter at the DNA level (Fig. 5D). SetDB1 is also essential for HP1a-induced reporter silencing. This instead is independent of Silencio (Supplemental Fig. S4)—an expected result, as it is a piRNA pathway-specific factor, while HP1a is a general heterochromatin effector in all cells. Based on these findings, we conclude that SetDB1 is the central effector methyltransferase that acts downstream from Silencio and HP1a.
Distinct roles of SetDB1 and Su(var)3-9 at piRNA targeted TE insertions
Previous studies have implicated both Eggless/SetDB1 and Su(var)3-9 in the Drosophila piRNA pathway (Rangan et al. 2011; Huang et al. 2013). However, no systematic survey on the role of these two chromatin modifiers downstream from Piwi has been reported. We therefore used OSCs and established potent siRNA-mediated depletion conditions for Egg/SetDB1, Su(var)3-9, and G9a, an enzyme that might contribute to H3K9 methylation (Supplemental Fig. S5A,B; Stabell et al. 2006). Genome-wide profiling of the H3K9me3 mark as well as of RNA Pol II occupancy shows that only depletion of SetDB1 impairs heterochromatin formation and transcriptional silencing at piRNA targeted TE insertions (Fig. 6A). In contrast, H3K9me3 levels within constitutive heterochromatin, which are established independently of Piwi (Sienski et al. 2012), are insensitive to SetDB1 depletion and instead are moderately affected in Su(var)3-9-depleted cells (Fig. 6B), supporting a role of Su(var)3-9 in the maintenance of constitutive heterochromatin. The impact of SetDB1 loss on TE silencing in OSCs is not a result of reduced piRNA levels (Rangan et al. 2011), as depletion of SetDB1 does not impair piRNA production from flamenco or cluster 20A, the two dominant piRNA sources in OSCs (Fig. 6C). Thus, at least in somatic ovarian cells, SetDB1 acts specifically in target repression and is not required for piRNA cluster transcription.
Figure 6.
Distinct roles for Eggless/SetDB1 and Su(var)3-9 at piRNA-repressed TE insertions. (A) Heat maps showing H3K9me3 signal (green) and RNA Pol II occupancy (red) on the 10 kb flanking the 381 euchromatic TE insertions targeted by Piwi in OSCs. Signals from GFP-depleted cells (control) were compared with cells depleted for CG9754, SetDB1, Su(var)3-9, G9a, or HP1a. All heat maps were sorted for the average H3K9me3 intensity in the control experiment. Below each heat map, area charts represent the average signal. (B) Box plot showing changes in H3K9me3 levels at euchromatic H3K9me3 peaks (enriched in Piwi-repressed TE insertions) and heterochromatic H3K9me3 peaks in OSCs depleted for the indicated factors. The identities of analyzed H3K9me3 peaks are as in Sienski et al. (2012). (C) Bar graph showing mean levels of piRNAs derived from flamenco and cluster 20A (each bar is the average value of piRNA densities in 40 nonoverlapping 1-kb windows) in OSCs depleted for the indicated factors (values were normalized for those present in GFP knockdown control cells). Error bars correspond to SEM. (D) Jitter plot showing fold changes in mRNA levels of TEs in the indicated knockdowns. The top panel displays TEs regulated by Piwi. Fold change, >4; n = 11. (****) P-value < 0.0001, according to Mann-Whitney test. The bottom panel displays TEs not regulated by Piwi. n = 36. (E) Metaprofile showing HP1a occupancy at the genomic regions flanking the 381 euchromatic TE insertions that are repressed by Piwi in wild-type as well as Piwi-depleted OSCs (using two different HP1a antibodies). (F) Bar diagram showing the occupancy of HP1a and Su(var)3-9 at the indicated loci in wild-type OSCs (light and parp are loci within constitutive heterochromatin). n = 2. Error bars indicate SD. (G) Shown is the average H3K9me3 signal at genomic regions flanking the 146 euchromatic gypsy insertions in OSCs depleted for the indicated factors.
In addition, transcriptome profiling by RNA sequencing (RNA-seq) indicates that the steady-state transcript levels of Piwi-repressed TEs are only increased upon depletion of SetDB1 (Fig. 6D; Supplemental Fig. S5C). We note that TE desilencing in SetDB1-depleted cells is considerably weaker than in Piwi-depleted cells. As the simultaneous knockdown of SetDB1 and either of the two other methyltransferases does not lead to higher TE derepression (Fig. 6D; Supplemental Fig. S5C), this most likely does not reflect redundancies among the methyltransferases. Instead, it could be due to the different kinetics of H3K9me3 loss in SetDB1 versus Piwi-depleted cells.
Further support for the integral role of the H3K9me3 mark in piRNA-guided TE silencing stems from the observation that depletion of HP1a, a major reader of the H3K9me3 mark, leads genome-wide to strong transcriptional increases at piRNA targeted TE insertions—again, without affecting global piRNA levels (Fig. 6A,C; Supplemental Fig. S5D; Ohtani et al. 2013). Consistent with this, HP1a is present at piRNA-targeted TE insertions as well as at the flanking genomic regions in a Piwi-dependent manner (Fig. 6E,F).
Although loss of Su(var)3-9 does not impair Piwi-mediated TE silencing, a closer inspection of the H3K9me3 profiles at piRNA-repressed TE insertions reveals impaired spreading of the H3K9me3 mark to flanking genomic regions in the absence of this enzyme (Fig. 6A,G; Supplemental Fig. S5F). A calculation of the spreading index (distance from TE insertion at which the signal is 50% of the peak) indicates nearly twofold-reduced H3K9me3 spreading in Su(var)3-9-depleted cells (upstream, 1.1 kb vs. 1.8 kb; downstream, 1.6 kb vs. 2.8 kb). This is most likely a direct effect, as Su(var)3-9 is enriched at piRNA targeted TE insertions (including their immediately flanking genomic regions) to levels that are comparable with those found in constitutive heterochromatin (e.g., light and parp) (Fig. 6F). Similar roles in heterochromatin formation and spreading have been attributed to SetDB1 and Su(var)3-9 in mouse embryonic stem cells (Fritsch et al. 2010; Bulut-Karslioglu et al. 2014), indicating a conserved division of labor between these two H3K9 methyltransferases in heterochromatin biology.
Our data support a model in which Piwi–piRNA complexes target SetDB1 in a sequence-specific manner to TE insertions in order to initiate a repressive heterochromatin state, while Su(var)3-9 assists in local bidirectional spreading of the methyl mark, probably by binding to pre-existing H3K9me3 marks via its own chromodomain or HP1a (Lachner et al. 2001; Nakayama et al. 2001).
Discussion
The formation of heterochromatin (with the resulting transcriptional silencing of the locus) downstream from nuclear Argonaute proteins is an intriguing process. On the one hand, this is because the target RNA, which is essential for Argonaute targeting, will be suppressed in its own production. On the other hand, the process must deal with the fact that transcripts are only transiently attached to their originating chromatin locus; transcript maturation or transcription termination should in principle lead to the release of the target RNA, including bound Argonaute complexes. A wealth of pioneering work in S. pombe on centromeric heterochromatin formation downstream from Ago1 has provided significant insight into these questions (see above for references). However, how these findings relate to the molecular steps that are involved in transcriptional target silencing downstream from animal PIWI clade proteins is only poorly understood. In fact, the involvement of several nonconserved proteins between yeast and animals suggests significant differences.
Here, we characterize CG9754/Silencio as an essential piRNA pathway factor required for nuclear Piwi–piRNA complexes to mediate transcriptional silencing and heterochromatin formation. The central conclusions from our work are as follows: (1) Silencio is the first protein in the piRNA pathway that triggers heterochromatin formation and silencing if tethered ectopically to RNA or DNA. In this, our work provides a powerful entry point into the mechanistic dissection of the silencing process in a controlled setting. (2) We show that the two central H3K9 methyltransferases, Su(var)3-9 and Eggless/SetDB1, are acting downstream from Piwi yet in different and nonredundant manners: While SetDB1 is required for heterochromatin formation downstream from Piwi, Su(var)3-9 appears to be involved in spreading the H3K9me3 to flanking genomic areas (see also Brower-Toland et al. 2009).
The molecular function of Silencio
The primary sequence of Silencio does not reveal insights into its molecular roles in the piRNA-guided silencing process. Silencio does not harbor any domain of known function. Also, orthologs of Silencio are not identifiable outside Drosophilids. However, considering that heterochromatin formation downstream from nuclear PIWI proteins also occurs in mammals (e.g., Pezic et al. 2014), the existence of a factor with equivalent function seems very plausible outside Drosophilids.
How Silencio triggers heterochromatin formation is unclear. Our data suggest that it directly or indirectly recruits SetDB1 to target loci or licenses SetDB1's methylation activity. The specificity for this process almost certainly relies on Piwi–piRNA complexes. In fact, Silencio interacts with target-engaged Piwi. In this sense, Silencio could be considered as an adaptor protein that bridges Piwi to basic cellular activities involved in the establishment and maintenance of heterochromatin. Such a model is highly reminiscent of the biology of TE silencing in mammals, where, besides the gonadal piRNA pathway, a large family of KRAB-type zinc finger transcription factors recruits the adaptor protein KAP1/Trim28 to target TE sequences (Rowe et al. 2010). KAP1 acts as a binding platform for various chromatin effector proteins, including SetDB1 and HP1a. The concept of an adaptor protein is also evident for small RNA-guided post-transcriptional silencing, where GW182 bridges the sequence-specific AGO–miRNA complex to the cellular pathways that mediate mRNA turnover and translational repression (Behm-Ansmant et al. 2006).
The most remarkable aspect about Silencio is its ability to trigger heterochromatin formation if tethered to the nascent RNA. In fact, overcoming the transient interaction of the target RNA with chromatin does not seem to be a trivial task, as tethering of HP1a, which is a potent silencer if recruited directly to promoter regions, is unable to induce heterochromatin formation and silencing under these conditions. Whether Silencio links to activities that tether the target RNA to chromatin or whether this indicates that Silencio and HP1a signal to different downstream effectors with different silencing capacities or kinetics is unclear. In this respect, it is interesting to note that tethering of Ago1 as well as other RITS (RNA-induced transcriptional silencing) complex components to nascent transcripts in S. pombe is very inefficient in initiating heterochromatin as well (Buhler et al. 2006). A recent report indicates that impairing transcription elongation by loss of the Paf1 complex enables trans-silencing by RITS, presumably as the residence time of the nascent transcript at the target locus is increased (Kowalik et al. 2015).
The role of SetDB1 and Su(var)3-9 in piRNA-guided heterochromatin formation
Our data point to a major role of Eggless/SetDB1 in the establishment of H3K9me3-marked heterochromatin downstream from nuclear Piwi and Silencio. Consistent with this, loss of SetDB1, but not of Su(var)3-9, leads to the derepression of piRNA targeted TEs as well as a genome-wide loss of H3K9me3 at targeted TE insertions. In contrast to Su(var)3-9, which harbors an H3K9me3-binding chromodomain, SetDB1 seems to require other specificity determinants as a guide (e.g., piRNAs and KRAB zinc fingers). Whether SetDB1 is required for only the initiation or also the maintenance of the H3K9me3 domain is currently unclear and will require sophisticated experimental systems similar to those established in yeast (Audergon et al. 2015; Ragunathan et al. 2015). In terms of TE repression, Su(var)3-9 plays a less important role downstream from Piwi. Instead, it seems to primarily mediate the spreading of the H3K9me3 mark to genomic regions that are flanking piRNA targets. This is very consistent with a central role of Su(var)3-9 in position effect variegation (Tschiersch et al. 1994; Rea et al. 2000; Schotta et al. 2002). Also, phenotypically, the described division of labor between these two methyltransferases is consistent: eggless/SetDB1 mutant flies are subviable, and escapers exhibit severe oogenesis defects and are sterile (Clough et al. 2007). Instead, Su(var)3-9 mutant flies are viable and fertile.
The precise role of H3K9 methylation in piRNA-guided heterochromatin formation is not clear. It might serve as a binding platform for downstream effector complexes such as chromatin remodelers or histone deacetylase complexes, as has been shown previously (Czermin et al. 2001; Sugiyama et al. 2007). In mammals, it is probably also involved in guiding the de novo DNA methylation at CpG dinucleotides (Matsui et al. 2010). Alternative/additional roles of the H3K9me3 mark could be to aid higher-order chromatin structures (Sun et al. 2001) or control the subnuclear localization of silenced loci (Towbin et al. 2012) as well as prevent the ectopic recombination of TE loci that are identical in sequence at many genomic locations (Ellermeier et al. 2010).
Clearly, we are only beginning to reveal the molecular events that occur downstream from a Piwi–piRNA complex upon interacting with a complementary target transcript at chromatin. Understanding the mechanistic logic of piRNA-guided heterochromatin formation promises to provide a more general insight into the connections and interplay between heterochromatin formation and transcription.
Materials and methods
Genetics and transgenes
The RNA-tethering constructs (subcloned into pUASP/attB or act5C vectors) express the λN domain (Baron-Benhamou et al. 2004) followed by a single HA tag fused to the N terminus of the full-length ORFs of GW182, Piwi, Gtsf1, Maelstrom, CG9754, or HP1a. The empty λN-HA vector served as a control. The DNA-tethering constructs express LacI fused to GFP, Piwi, Gtsf1, Maelstrom, CG9754, or HP1a. For the boxB sensor construct, the α-tubulin-84D promoter drives GFP cDNA followed by five boxB sites in the 3′ UTR. In the DNA sensor construct, eight lacO sites precede a nanos promoter expressing a GFP-Piwi cDNA (including the first intron of piwi, which causes expression in somatic cells) (Hayashi et al. 2014). The UASP-λN and UASP-LacI transgenes were inserted into attP40, and the sensor constructs were inserted at attP2 (Markstein et al. 2008). Other transgenes were MTD-Gal4 and shRNAi lines (all inserted into attP2) targeting white, piwi, zuc, gtsf1, maelstrom, CG9754, eggless, and Su(var)3-9 (Ni et al. 2009; Olivieri et al. 2012; Sienski et al. 2012; Donertas et al. 2013; Mohn et al. 2014). Novel CG9754 alleles were isolated in a white1118 background isogenic for the second and third chromosomes using two independent guide RNAs (Supplemental Table S2) and were compared with Cas9-generated piwi alleles that have the same genetic background (Senti et al. 2015).
Antibodies
Mouse monoclonal antibodies (used for Western blotting and immunoprecipitations) against His-tagged full-length CG9754/Silencio were generated by the Max F. Perutz Laboratories Monoclonal Antibody Facility. Rabbit antisera (used for immunofluorescence) were raised against the MEAPMKLEVKVENYVECGINC peptide of CG9754. Rabbit anti-Piwi is described in Brennecke et al. (2007), rabbit anti-Rbp3 was raised against full-length Drosophila Rbp3, and rabbit anti-Eggless was raised as described in Clough et al. (2007). The commercially available antibodies used were MAb 16B12 anti-HA (Covance), MAb anti-LacI (Abcam), anti-H3K9me3 39161 (ActiveMotif), anti-RNA Pol II 8WG16 (Abcam), MAb C1A9 anti-HP1a (Developmental Studies Hybridoma Bank), and rabbit polyclonal PRB291C anti-HP1a (Covance). Rabbit anti-Su(var)3-9 was a kind gift of G. Reuter (Schotta et al. 2002), and rabbit anit-dG9a was a kind gift of M. Yamaguchi (Kato et al. 2008).
Cell culture
OSCs were cultured as described (Niki et al. 2006; Saito et al. 2009) and transfected with Cell Line Nucleofector kit V (Amaxa Biosystems, program T-029). The reporter constructs were cloned into pAcM (Saito et al. 2010) and cotransfected with siRNA targeting endogenous piwi transcript or Luciferase transcript. The luciferase assay was performed as described (Behm-Ansmant et al. 2006).
RT-qPCR
Primer sequences used for RT-qPCR are listed in Supplemental Table S1.
Transposon qPCR analysis
cDNA was prepared via random priming of 1 µg of total RNA isolated from ovaries of 2- to 3-d-old flies. Each experiment was performed in biological duplicates with technical triplicates. Relative RNA levels were normalized to rp49 levels. Fold enrichments were calculated in comparison with respective RNA levels obtained from white1118 flies.
Small RNA cloning
Small RNA cloning and sequencing was performed as described in Jayaprakash et al. (2011). For small RNA cloning after immunoprecipitation, the RNAs associated with Piwi or CG9754 were isolated from the respective immunoprecipitates (total nuclear small RNAs used as a control). Libraries were sequenced with a HiSeq 2500 (Illumina).
Sequencing of mRNAs (mRNA-seq)
Total RNA from OSCs was isolated using TRIzol. Twenty micrograms of RNA was subjected to DNaseI digestion (Qiagen) and purified using RNeasy kit (Qiagen). mRNAs were enriched using double purification with oligo-d(T) beads (Ambion) followed by RNA fragmentation and first and second strand cDNA synthesis. The libraries were constructed with the NEBNext Ultra DNA library preparation kit (Illumina), amplified with the KAPA HiFi polymerase (Kapa Biosystems), and sequenced with a HiSeq 2500 (Illumina).
Immunoprecipitation
OSCs were harvested by trypsinization, and nuclear extract was prepared with lysis buffer (50 mM Tris at pH 7.5, 0.5% Triton, 137.5 mM NaCl, 5 mM EDTA, 10% glycerol). The soluble nuclear fraction was incubated with α-Piwi (rabbit), α-CG9754 (mouse), or control IgG (mouse), each coupled to Protein G Dynabeads (Invitrogen) for 8 h at 4°C and washed (30 mM Hepes at pH 7.5, 300 mM NaCl, 2 mM MgCl2, 10% glycerol, 0.25% Triton, 0.5% NP40). Purified complexes were eluted with SDS sample buffer and analyzed by gel electrophoresis and Western blotting.
ChIP
ChIP was carried out according to Lee et al. (2006). In brief, 50 µL of dissected ovaries was fixed with 1% formaldehyde (PFA), and the prepared chromatin was sonicated. Two-thirds of the material was used for immunoprecipitation of RNA Pol II (8WG16), and one-third was used for immunoprecipitation of H3K9me3 (anti-H3K9me3; Active Motif, #39161) followed by decross-linking, purification of DNA, and qPCR analysis. ChIP analysis in OSCs was done similarly with the following modifications. Twenty million cells depleted for studied proteins were fixed with 1% PFA followed by chromatin extraction and sonication. For the RNA Pol II ChIP, we incubated 75% of the chromatin with α-Rbp3 and 25% with α-H3K9me3. After immunoprecipitation, purification, and decross-linking, DNA was extracted and used for qPCR measurements and/or library preparation for sequencing (NEBNext Ultra DNA library preparation kit, Illumina).
Computational analyses
For details regarding the computational analyses, see Supplemental Document S1.
Accession numbers
Illumina deep sequencing data sets were deposited at NCBI Gene Expression Omnibus (GSE74097).
Supplementary Material
Acknowledgments
We thank the Max F. Perutz Laboratories Monoclonal Antibody Facility for generating α-CG9754; M. Madalinski for peptide synthesis and antibody purifications; the Campus Science Support Facilities Next-Generation Sequencing unit for sequencing; J. Gokcezade for fly injections; P. Duchek for generation of CRISPR/Cas9 alleles; the Harvard Transgenic RNAi Project and Bloomington Stock Center for flies; the Developmental Studies Hybridoma Bank, G. Reuter, and M. Yamaguchi for antibodies; and E. Izaurralde for λN-GW182 and boxB-Luciferase plasmids. This work was supported by the Austrian Academy of Sciences, the European Community's 7th Framework Programme (FP7/2007-2013; ERC grant no. 260711EU) and the Austrian Science Fund (FWF; Y 510-B12, F4303-B09, and W12-7-B09). G.S. and J.B. are supported by Boehringer Ingelheim fellowships, and K.-A.S. was supported by a Marie Curie Intra-European Fellowship grant by the European Union (FP7-MC-IEF-253912).
Footnotes
Supplemental material is available for this article.
Article published online ahead of print. Article and publication date are online at http://www.genesdev.org/cgi/doi/10.1101/gad.271908.115.
Note added in proof
Hannon and colleagues (Yu et al. 2015) independently identified CG9754 as a central bridging factor between Piwi and the heterochromatin machinery and named this factor Panoramix.
References
- Audergon PN, Catania S, Kagansky A, Tong P, Shukla M, Pidoux AL, Allshire RC. 2015. Epigenetics. Restricted epigenetic inheritance of H3K9 methylation. Science 348: 132–135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aygun O, Mehta S, Grewal SI. 2013. HDAC-mediated suppression of histone turnover promotes epigenetic stability of heterochromatin. Nat Struct Mol Biol 20: 547–554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ayyanathan K, Lechner MS, Bell P, Maul GG, Schultz DC, Yamada Y, Tanaka K, Torigoe K, Rauscher FJ III. 2003. Regulated recruitment of HP1 to a euchromatic gene induces mitotically heritable, epigenetic gene silencing: a mammalian cell culture model of gene variegation. Genes Dev 17: 1855–1869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baron-Benhamou J, Gehring NH, Kulozik AE, Hentze MW. 2004. Using the λN peptide to tether proteins to RNAs. Methods Mol Biol 257: 135–154. [DOI] [PubMed] [Google Scholar]
- Bayne EH, White SA, Kagansky A, Bijos DA, Sanchez-Pulido L, Hoe KL, Kim DU, Park HO, Ponting CP, Rappsilber J, et al. 2010. Stc1: a critical link between RNAi and chromatin modification required for heterochromatin integrity. Cell 140: 666–677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behm-Ansmant I, Rehwinkel J, Doerks T, Stark A, Bork P, Izaurralde E. 2006. mRNA degradation by miRNAs and GW182 requires both CCR4:NOT deadenylase and DCP1:DCP2 decapping complexes. Genes Dev 20: 1885–1898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brennecke J, Aravin AA, Stark A, Dus M, Kellis M, Sachidanandam R, Hannon GJ. 2007. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128: 1089–1103. [DOI] [PubMed] [Google Scholar]
- Brower-Toland B, Riddle NC, Jiang H, Huisinga KL, Elgin SC. 2009. Multiple SET methyltransferases are required to maintain normal heterochromatin domains in the genome of Drosophila melanogaster. Genetics 181: 1303–1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buhler M, Verdel A, Moazed D. 2006. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125: 873–886. [DOI] [PubMed] [Google Scholar]
- Bulut-Karslioglu A, De La Rosa-Velazquez IA, Ramirez F, Barenboim M, Onishi-Seebacher M, Arand J, Galan C, Winter GE, Engist B, Gerle B, et al. 2014. Suv39h-dependent H3K9me3 marks intact retrotransposons and silences LINE elements in mouse embryonic stem cells. Mol Cell 55: 277–290. [DOI] [PubMed] [Google Scholar]
- Castel SE, Martienssen RA. 2013. RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet 14: 100–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clough E, Moon W, Wang S, Smith K, Hazelrigg T. 2007. Histone methylation is required for oogenesis in Drosophila. Development 134: 157–165. [DOI] [PubMed] [Google Scholar]
- Cox DN, Chao A, Lin H. 2000. piwi encodes a nucleoplasmic factor whose activity modulates the number and division rate of germline stem cells. Development 127: 503–514. [DOI] [PubMed] [Google Scholar]
- Czech B, Preall JB, McGinn J, Hannon GJ. 2013. A transcriptome-wide RNAi screen in the Drosophila ovary reveals factors of the germline piRNA pathway. Mol Cell 50: 749–761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Czermin B, Schotta G, Hulsmann BB, Brehm A, Becker PB, Reuter G, Imhof A. 2001. Physical and functional association of SU(VAR)3-9 and HDAC1 in Drosophila. EMBO Rep 2: 915–919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donertas D, Sienski G, Brennecke J. 2013. Drosophila Gtsf1 is an essential component of the Piwi-mediated transcriptional silencing complex. Genes Dev 27: 1693–1705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellermeier C, Higuchi EC, Phadnis N, Holm L, Geelhood JL, Thon G, Smith GR. 2010. RNAi and heterochromatin repress centromeric meiotic recombination. Proc Natl Acad Sci 107: 8701–8705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fritsch L, Robin P, Mathieu JR, Souidi M, Hinaux H, Rougeulle C, Harel-Bellan A, Ameyar-Zazoua M, Ait-Si-Ali S. 2010. A subset of the histone H3 lysine 9 methyltransferases Suv39h1, G9a, GLP, and SETDB1 participate in a multimeric complex. Mol Cell 37: 46–56. [DOI] [PubMed] [Google Scholar]
- Gokcezade J, Sienski G, Duchek P. 2014. Efficient CRISPR/Cas9 plasmids for rapid and versatile genome editing in Drosophila. G3 (Bethesda) 4: 2279–2282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall IM, Shankaranarayana GD, Noma K, Ayoub N, Cohen A, Grewal SI. 2002. Establishment and maintenance of a heterochromatin domain. Science 297: 2232–2237. [DOI] [PubMed] [Google Scholar]
- Handler D, Meixner K, Pizka M, Lauss K, Schmied C, Gruber FS, Brennecke J. 2013. The Genetic Makeup of the Drosophila piRNA Pathway. Mol Cell 50: 762–777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi R, Handler D, Ish-Horowicz D, Brennecke J. 2014. The exon junction complex is required for definition and excision of neighboring introns in Drosophila. Genes Dev 28: 1772–1785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang XA, Yin H, Sweeney S, Raha D, Snyder M, Lin H. 2013. A major epigenetic programming mechanism guided by piRNAs. Dev Cell 24: 502–516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishida M, Shimojo H, Hayashi A, Kawaguchi R, Ohtani Y, Uegaki K, Nishimura Y, Nakayama J. 2012. Intrinsic nucleic acid-binding activity of Chp1 chromodomain is required for heterochromatic gene silencing. Mol Cell 47: 228–241. [DOI] [PubMed] [Google Scholar]
- Jayaprakash AD, Jabado O, Brown BD, Sachidanandam R. 2011. Identification and remediation of biases in the activity of RNA ligases in small-RNA deep sequencing. Nucleic Acids Res 39: e141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337: 816–821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jonas S, Izaurralde E. 2015. Towards a molecular understanding of microRNA-mediated gene silencing. Nat Rev Genet 16: 421–433. [DOI] [PubMed] [Google Scholar]
- Kato Y, Kato M, Tachibana M, Shinkai Y, Yamaguchi M. 2008. Characterization of Drosophila G9a in vivo and identification of genetic interactants. Genes Cells 13: 703–722. [DOI] [PubMed] [Google Scholar]
- Kowalik KM, Shimada Y, Flury V, Stadler MB, Batki J, Buhler M. 2015. The Paf1 complex represses small-RNA-mediated epigenetic gene silencing. Nature 520: 248–252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachner M, O'Carroll D, Rea S, Mechtler K, Jenuwein T. 2001. Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410: 116–120. [DOI] [PubMed] [Google Scholar]
- Lee TI, Johnstone SE, Young RA. 2006. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat Protoc 1: 729–748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Le Thomas A, Rogers AK, Webster A, Marinov GK, Liao SE, Perkins EM, Hur JK, Aravin AA, Toth KF. 2013. Piwi induces piRNA-guided transcriptional silencing and establishment of a repressive chromatin state. Genes Dev 27: 390–399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Danzer JR, Alvarez P, Belmont AS, Wallrath LL. 2003. Effects of tethering HP1 to euchromatic regions of the Drosophila genome. Development 130: 1817–1824. [DOI] [PubMed] [Google Scholar]
- Markstein M, Pitsouli C, Villalta C, Celniker SE, Perrimon N. 2008. Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nat Genet 40: 476–483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsui T, Leung D, Miyashita H, Maksakova IA, Miyachi H, Kimura H, Tachibana M, Lorincz MC, Shinkai Y. 2010. Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 464: 927–931. [DOI] [PubMed] [Google Scholar]
- Mohn F, Sienski G, Handler D, Brennecke J. 2014. The rhino–deadlock–cutoff complex licenses noncanonical transcription of dual-strand piRNA clusters in Drosophila. Cell 157: 1364–1379. [DOI] [PubMed] [Google Scholar]
- Muerdter F, Guzzardo PM, Gillis J, Luo Y, Yu Y, Chen C, Fekete R, Hannon GJ. 2013. A genome-wide RNAi screen draws a genetic framework for transposon control and primary piRNA biogenesis in Drosophila. Mol Cell 50: 736–748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakayama J, Rice JC, Strahl BD, Allis CD, Grewal SI. 2001. Role of histone H3 lysine 9 methylation in epigenetic control of heterochromatin assembly. Science 292: 110–113. [DOI] [PubMed] [Google Scholar]
- Ni JQ, Liu LP, Binari R, Hardy R, Shim HS, Cavallaro A, Booker M, Pfeiffer BD, Markstein M, Wang H, et al. 2009. A Drosophila resource of transgenic RNAi lines for neurogenetics. Genetics 182: 1089–1100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niki Y, Yamaguchi T, Mahowald AP. 2006. Establishment of stable cell lines of Drosophila germ-line stem cells. Proc Natl Acad Sci 103: 16325–16330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noma K, Sugiyama T, Cam H, Verdel A, Zofall M, Jia S, Moazed D, Grewal SI. 2004. RITS acts in cis to promote RNA interference-mediated transcriptional and post-transcriptional silencing. Nat Genet 36: 1174–1180. [DOI] [PubMed] [Google Scholar]
- Ohtani H, Iwasaki YW, Shibuya A, Siomi H, Siomi MC, Saito K. 2013. DmGTSF1 is necessary for Piwi-piRISC-mediated transcriptional transposon silencing in the Drosophila ovary. Genes Dev 27: 1656–1661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olivieri D, Senti KA, Subramanian S, Sachidanandam R, Brennecke J. 2012. The cochaperone shutdown defines a group of biogenesis factors essential for all piRNA populations in Drosophila. Mol Cell 47: 954–969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pezic D, Manakov SA, Sachidanandam R, Aravin AA. 2014. piRNA pathway targets active LINE1 elements to establish the repressive H3K9me3 mark in germ cells. Genes Dev 28: 1410–1428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Post C, Clark JP, Sytnikova YA, Chirn GW, Lau NC. 2014. The capacity of target silencing by Drosophila PIWI and piRNAs. RNA 20: 1977–1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ragunathan K, Jih G, Moazed D. 2015. Epigenetics. Epigenetic inheritance uncoupled from sequence-specific recruitment. Science 348: 1258699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rangan P, Malone CD, Navarro C, Newbold SP, Hayes PS, Sachidanandam R, Hannon GJ, Lehmann R. 2011. piRNA production requires heterochromatin formation in Drosophila. Curr Biol 21: 1373–1379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rea S, Eisenhaber F, O'Carroll D, Strahl BD, Sun ZW, Schmid M, Opravil S, Mechtler K, Ponting CP, Allis CD, et al. 2000. Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature 406: 593–599. [DOI] [PubMed] [Google Scholar]
- Robine N, Lau NC, Balla S, Jin Z, Okamura K, Kuramochi-Miyagawa S, Blower MD, Lai EC. 2009. A broadly conserved pathway generates 3′UTR-directed primary piRNAs. Curr Biol 19: 2066–2076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rowe HM, Jakobsson J, Mesnard D, Rougemont J, Reynard S, Aktas T, Maillard PV, Layard-Liesching H, Verp S, Marquis J, et al. 2010. KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463: 237–240. [DOI] [PubMed] [Google Scholar]
- Rozhkov NV, Hammell M, Hannon GJ. 2013. Multiple roles for Piwi in silencing Drosophila transposons. Genes Dev 27: 400–412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saito K, Inagaki S, Mituyama T, Kawamura Y, Ono Y, Sakota E, Kotani H, Asai K, Siomi H, Siomi MC. 2009. A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila. Nature 461: 1296–1299. [DOI] [PubMed] [Google Scholar]
- Saito K, Ishizu H, Komai M, Kotani H, Kawamura Y, Nishida KM, Siomi H, Siomi MC. 2010. Roles for the Yb body components Armitage and Yb in primary piRNA biogenesis in Drosophila. Genes Dev 24: 2493–2498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarot E, Payen-Groschene G, Bucheton A, Pelisson A. 2004. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166: 1313–1321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schotta G, Ebert A, Krauss V, Fischer A, Hoffmann J, Rea S, Jenuwein T, Dorn R, Reuter G. 2002. Central role of Drosophila SU(VAR)3-9 in histone H3-K9 methylation and heterochromatic gene silencing. EMBO J 21: 1121–1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Senti KA, Jurczak D, Sachidanandam R, Brennecke J. 2015. piRNA-guided slicing of transposon transcripts enforces their transcriptional silencing via specifying the nuclear piRNA repertoire. Genes Dev 29: 1747–1762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sienski G, Donertas D, Brennecke J. 2012. Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151: 964–980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stabell M, Eskeland R, Bjorkmo M, Larsson J, Aalen RB, Imhof A, Lambertsson A. 2006. The Drosophila G9a gene encodes a multi-catalytic histone methyltransferase required for normal development. Nucleic Acids Res 34: 4609–4621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugiyama T, Cam HP, Sugiyama R, Noma K, Zofall M, Kobayashi R, Grewal SI. 2007. SHREC, an effector complex for heterochromatic transcriptional silencing. Cell 128: 491–504. [DOI] [PubMed] [Google Scholar]
- Sun FL, Cuaycong MH, Elgin SC. 2001. Long-range nucleosome ordering is associated with gene silencing in Drosophila melanogaster pericentric heterochromatin. Mol Cell Biol 21: 2867–2879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towbin BD, Gonzalez-Aguilera C, Sack R, Gaidatzis D, Kalck V, Meister P, Askjaer P, Gasser SM. 2012. Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell 150: 934–947. [DOI] [PubMed] [Google Scholar]
- Tschiersch B, Hofmann A, Krauss V, Dorn R, Korge G, Reuter G. 1994. The protein encoded by the Drosophila position-effect variegation suppressor gene Su(var)3-9 combines domains of antagonistic regulators of homeotic gene complexes. EMBO J 13: 3822–3831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verdel A, Jia S, Gerber S, Sugiyama T, Gygi S, Grewal SI, Moazed D. 2004. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303: 672–676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Volpe TA, Kidner C, Hall IM, Teng G, Grewal SI, Martienssen RA. 2002. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297: 1833–1837. [DOI] [PubMed] [Google Scholar]
- Wang SH, Elgin SC. 2011. Drosophila Piwi functions downstream of piRNA production mediating a chromatin-based transposon silencing mechanism in female germ line. Proc Natl Acad Sci 108: 21164–21169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamada T, Fischle W, Sugiyama T, Allis CD, Grewal SI. 2005. The nucleation and maintenance of heterochromatin by a histone deacetylase in fission yeast. Mol Cell 20: 173–185. [DOI] [PubMed] [Google Scholar]
- Yu Y, Gu J, Jin Y, Luo Y, Preall JB, Ma J, Czech B, Hannon GJ. 2015. Panoramix enforces piRNA-dependent cotranscriptional silencing. Science 350: 339–342. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.