Abstract
Glutathione S-transferases (GSTs) are enzymes which expressed in many tissues and play important roles in neutralization of toxic compounds, and protecting hosts against cancer. Among several GSTs, Glutathione S-transferases mu (GSTM) has been drawn attention upon the association with the genetic risk for many types of cancers. But whether the GSTM1 polymorphisms confer the susceptibility to colorectal cancer in Asians has not been well established. We searched the PubMed database with GSTM1, polymorphism and colorectal cancer, attempting to identify the eligible studies. In total, 33 case-control studies in Asian populations with 8502 colorectal cancer patients and 13699 controls were included in the current meta-analysis. The association between the polymorphism and susceptibility to colorectal cancer was evaluated by the odds ratio (OR) and 95% confidence intervals (CI). The pooled meta-analysis suggested that GSTM1 null variant was correlated to the colorectal cancer risk in Asians. There was a marginal heterogeneity among these eligible studies. Nevertheless, cumulative meta-analysis observed a trend of an obvious association between the GSTM1 null genotype and colorectal cancer risk in Asians. In summary, the meta-analysis suggested that GSTM1 null polymorphism confer the susceptibility to colorectal cancer in Asians, especially in Chinese populations.
Colorectal cancer is the second leading common cancer and a major cause of cancer-related deaths in the world-wide, which accounts for about 9.7% death rate of all cancers1,2. It has been reported that about 1.2 million new cases of colorectal cancer in the world in 20082. Although the incidence rate of colorectal cancer decreases in the western developed countries3,4, the rate still rises in the developing countries, especially in China. The etiology and development of colorectal cancer was not fully understood yet, but considering the previous studies, the colorectal cancer was proven to be complicated and multifactorial cancer5,6. Previous evidence suggested that the environmental risk factors and genetic factors both affected the pathogenesis of colorectal cancer7. It was reported that familial colorectal cancer accounts for around 5–15%8. Fearon ER and his colleagues6 found that many mutants/variants contribute to the pathogenesis of the sporadic and inherited forms colorectal cancer, including several GSTs variants.
GSTs are a super-family of phase II detoxification enzymes which play critical roles in the detoxification of exogenous and endogenous reactive species through conversion of toxic compounds to hydrophilic metabolites9,10. To date, there are 8 classes of GSTs including alpha (α), kappa (κ), mu (μ), omega (ω), pi (π), sigma (σ), theta (θ) and zeta (ζ) have been clarified11, and polymorphisms have been identified among several of them12. Glutathione S-transferase M1 (GSTM1) encodes the mu class of GSTs, which plays vital roles in protecting hosts against cancers13. The mu GST enzymes were demonstrated to be more effective at the process of detoxifying cytotoxic and genotoxic reactive species than other GSTs13. The null variant (GSTM1*0 allele) is the most common variant of the GSTM1, leading to the loss of enzyme activity, and the variant-carriers were proven to be associated with increased risk to cancers9. GSTM1 has been identified to be involved in the pathogenesis and development of certain cancers, inclusive of colorectal cancer14.
Many association studies and evidence were conducted and highlighted the GSTM1 null variant was correlated with the risk of cancers15,16. Moreover, the GSTM1 null genotype has been demonstrated to be linked to early onset for colorectal cancer17. However, the previous studies which aimed to investigate the association of GSTM1 polymorphism and the colorectal cancer susceptibility in Chinese populations were controversial, and so were the genetic results in Asians. Inclusive of 33 case-control studies, the present meta-analysis was performed to further explore the relationship between GSTM1 null variant and the susceptibility to colorectal cancer more comprehensively in Asians.
Methods
Literature search and inclusion criteria
We systematically searched the PubMed database (http://www.ncbi.nlm.nih.gov/pubmed/) to identify the eligible case-control studies using the following keywords: (“glutathione S-transferase” or “GST” or “glutathione S-transferase M1” or “GSTM1”), (“polymorphism” or “Single Nucleotide Polymorphism”) and (“colorectal cancer” or “CRC” or “colorectal carcinoma” or “colorectal adenoma”). There was no language limitation in the literature search. Titles and abstracts of these searching studies were primarily screened and full papers were further retrieved to confirm eligibility, the reference lists were also examined to find other relevant studies. These studies were reported from 1996 to 2014. These given studies included into our meta-analysis had to meet the following inclusion criteria: (1) case-control validation study, (2) the studies should estimate the association of the GSTM1 null variant with risk to colorectal cancer, (3) the studies should provide odds ratio (OR) with 95% CI or available data, (4) the studies should be conducted in Asian populations. Reviews and duplicate studies were excluded from the analysis. If the same case-control study were overlapped in multiple publications, only the most complete or most recent literature was included in the present study.
Data extraction
The available data originated in the eligible studies was independently extracted by two co-authors. The following information was collected: first author’s name, year of publication, country of the study conducted, ethnicity of participants, numbers of cases and controls and the genotype distributions of GSTM1 null variant. All subjects followed the principles of the Declaration of Helsinki.
Statistical analysis
The association between GSTM1 null genotype with colorectal cancer risk in Asians was calculated by pooled OR with 95% CI. The ORs were evaluated according to the extracted data. A 95% CI was used for standard for statistical significance and 95% CI without 1 for OR indicated that the genotype may increase or decrease the cancer genetic risk significantly. Firstly, the I2 statistic was estimated to quantify the heterogeneity among all eligible studies18, and an I2 < 50% suggested low heterogeneity. In general, the meta-analysis was performed using fixed-effect19 or random-effect20 models according to the effect estimates in the presence (I2 ≤ 50%) or absence (I2 > 50%) of significant heterogeneity. So, the random-effect model should be chosen once the obvious heterogeneity was observed. Interesting, the heterogeneity was also considered to be significant when P < 0.1018,21,22. But it was widely accepted that the cutoff for significance of heterogeneity is P < 0.05 in the meta-analysis23. Furthermore, Sensitivity analysis was conducted by omitting those studies in turns to estimate the overall pooled ORs in the present study. Additionally, the Begg’s funnel plot and the Egger’s regression plot were considered to be preferred method to assess the publication bias24,25. An asymmetric funnel plot suggested a relationship between effect and study size, indicating the possibility of either publication bias or a systematic difference between smaller and larger studies26,27. We further performed a cumulative meta-analysis to investigate a framework for updating a case-control-effect from all eligible studies and to assess how much the genetic effect changes as statistical power accumulates, and to find the trend in risk effect28. In the cumulative meta-analysis, studies were ordered by publication year, and the pooled ORs were calculated at the end of each study. All statistical analyses were assessed by using STATA software, version 12.0 (StataCorp LP, College Station, TX, USA). All P values were two-sided.
Results
Characteristics of the case-control studies
The flow chart of eligible studies was shown in Fig. 1. Firstly, 89 potential studies were screened after literature search followed with the search strategy, but only 33 eligible studies remained according to the inclusion criteria. All these association studies were published from 1996 to 2014, and 19 studies were in English, and the other 14 studies were published in Chinese, which shown in the Table 1. In total, there are 22201 subjects, comprising of 8502 colorectal cancer patients and 13699 matched controls were included in the current study from these 33 studies. The characteristics of those eligible studies in Asians were shown in Table 1.
Figure 1. Flow chart of eligible studies.
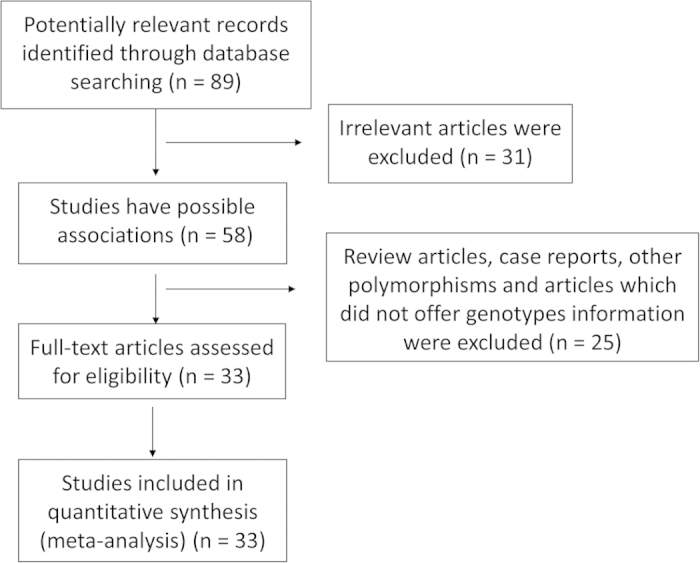
Table 1. Detailed information of the eligible studies included in the meta-analysis.
| Reference | Population | Ethnicity | No of cases |
case/GSTM1 |
control/GSTM1 |
|||
|---|---|---|---|---|---|---|---|---|
| null | present | No of controls | null | present | ||||
| Cong 201433 | Chinese men | Asian | 264 | 142 | 122 | 317 | 135 | 182 |
| Vogtmann 201434 | Chinese men | Asian | 335 | 201 | 134 | 638 | 379 | 259 |
| Hamachi 201335 | Japan men | Asian | 455 | 255 | 200 | 1052 | 546 | 506 |
| Huang X 201236 | Chinese | Asian | 100 | 42 | 58 | 130 | 59 | 71 |
| Koh 201137 | Chinese | Asian | 480 | 234 | 246 | 1167 | 526 | 641 |
| Wang 201138 | India | Asian | 302 | 100 | 202 | 291 | 76 | 215 |
| Nisa 201039 | Japan | Asian | 685 | 357 | 328 | 778 | 422 | 356 |
| Yang 201040 | Chinese | Asian | 322 | 189 | 133 | 1247 | 729 | 518 |
| Yeh 201041 | Taiwan | Asian | 722 | 401 | 321 | 733 | 410 | 323 |
| Piao 200942 | Korean | Asian | 1829 | 1004 | 825 | 1699 | 923 | 776 |
| Lin LM 200843 | Chinese | Asian | 120 | 69 | 51 | 204 | 90 | 114 |
| Huang LR 200744 | Chinese | Asian | 57 | 40 | 17 | 68 | 35 | 33 |
| Xia XP 200745 | Chinese | Asian | 112 | 67 | 45 | 140 | 63 | 77 |
| Yoshida 200746 | Japan | Asian | 66 | 36 | 30 | 121 | 62 | 59 |
| Fan CH 200647 | Chinese | Asian | 138 | 80 | 58 | 339 | 188 | 151 |
| Fu QH 200648 | Chinese | Asian | 315 | 229 | 86 | 438 | 321 | 117 |
| Luo JG 200649 | Chinese | Asian | 56 | 20 | 36 | 143 | 48 | 95 |
| Probst-Hensch 200650 | Chinese | Asian | 300 | 132 | 168 | 1168 | 525 | 643 |
| Chen K 200451 | Chinese | Asian | 126 | 69 | 56 | 343 | 188 | 151 |
| Huang P 200352 | Chinese | Asian | 82 | 46 | 36 | 82 | 28 | 54 |
| Yang J 200353 | Chinese | Asian | 58 | 40 | 18 | 65 | 29 | 35 |
| Seow 200254 | Chinese | Asian | 213 | 108 | 105 | 1190 | 537 | 653 |
| Wu 200255 | Taiwan | Asian | 356 | 173 | 183 | 278 | 136 | 142 |
| Zhu YQ 200256 | Chinese | Asian | 104 | 59 | 45 | 101 | 47 | 54 |
| Saadat 200157 | Iran | Asian | 46 | 25 | 21 | 131 | 53 | 78 |
| Zhang YC 200158 | Chinese | Asian | 52 | 22 | 30 | 52 | 25 | 27 |
| Inoue 200059 | Japan | Asian | 205 | 108 | 97 | 220 | 123 | 97 |
| Zhou JN 200060 | Chinese | Asian | 55 | 34 | 21 | 62 | 33 | 29 |
| Yoshioka 199961 | Japan | Asian | 106 | 56 | 50 | 100 | 42 | 58 |
| Gao JR 199862 | Chinese | Asian | 19 | 7 | 12 | 70 | 25 | 45 |
| Lee 199863 | Singapore | Asian | 300 | 128 | 172 | 183 | 89 | 94 |
| Guo 199664 | Chinese | Asian | 19 | 7 | 12 | 23 | 6 | 17 |
| Katoh 199665 | Japan | Asian | 103 | 56 | 47 | 126 | 55 | 71 |
Overall analysis
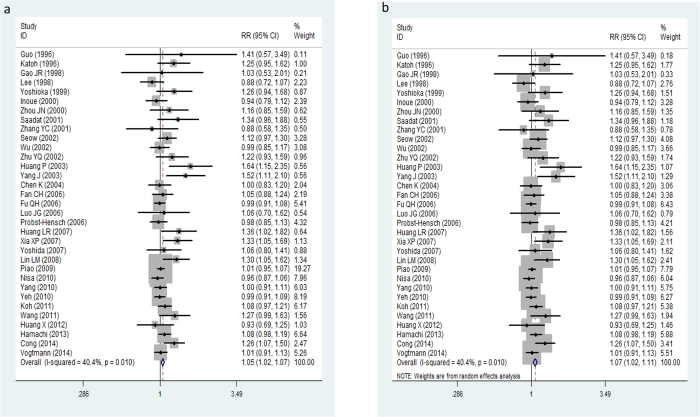
There was a marginal heterogeneity among these 33 validation studies in Asian cohorts (P = 0. 01, I2 = 40.4%) (Fig. 2). Overall, the pooled meta-analysis of eligible case-control studies suggested that GSTM1 null genotype was significantly associated with the risk to colorectal cancer in Asian populations (Z = 3.32, P = 0.001, OR = 1.05, 95% CI: 1.02–1.07) in a fixed-effect model (Fig. 2a). The association still remained (Z = 3.17, P = 0.002, OR = 1.07, 95% CI: 1.02–1.11) (Fig. 2b) under the random-effect model. Sensitivity analyses by omitting one study at a time did not materially alter the overall pooled ORs (supplementary Figure S1).
Figure 2.
Forest plot for the association between GSTM1 null polymorphism and colorectal cancer in Asians under fixed-effect model (a) and random-effect model (b).
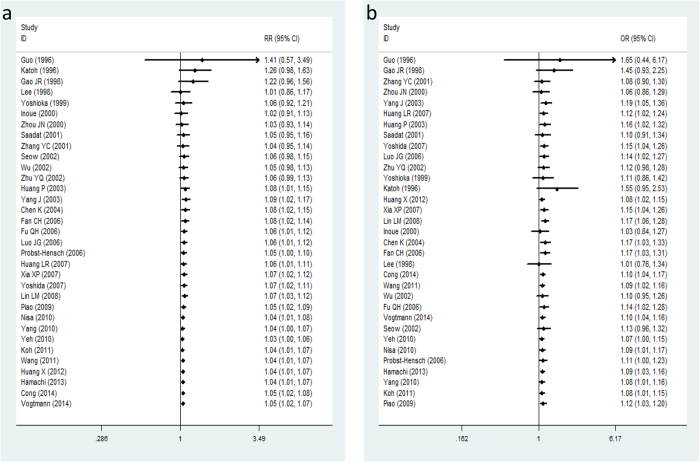
The cumulative meta-analysis further showed a trend of an obvious association between the GSTM1 null genotype and colorectal cancer risk in Asians as information accumulated by year (Fig. 3a), moreover, the cumulative analysis accumulated by the sample size also supported the result (Fig. 3b).
Figure 3.
Forest plot in the cumulative meta-analysis accumulated by publication year (a) and sample size (b).
Publication bias
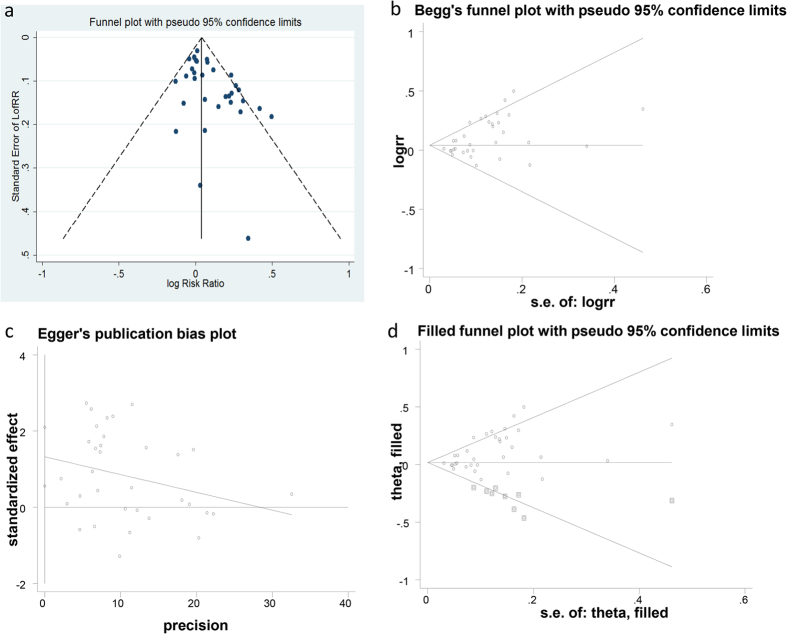
The funnel plot, Begg’s adjust rank correlation test, Egger’s regression test, trim and fill method are four corresponding methods applied to assess the publication bias in the current meta-analysis. Funnel plot, Begg’s funnel plot, Egger’s regression, plot trim and fill funnel plot were produced by these four methods respectively (Fig. 4a–d). There was no obvious asymmetry the shape of the funnel plot and the Begg’s funnel plot (Fig. 4a). However, publication bias was marginal when the Begg’s rank correlation method (z = 2.34, P = 0.019 < 0.05) was used, the publication bias was more significant in the Egger’s regression test (t = 3.51, P = 0.001 < 0.05) (supplementary Figure S3). Nine missing studies should be filled in the trim and fill method, furthermore, LogRR and its 95%CI altered significantly after the application of trim and fill method (supplementary Figure S4). The combined analyses indicated that there is publication bias in the present study.
Figure 4.
Publication bias test for the meta-analysis: Funnel plot (a), Begg’s funnel plot (b), Egger’s test (c) and Trim and fill test (d).
Sub-group analysis
There were 21 studies of Chinese, 6 in Japanese, 2 of Taiwan subjects, each single study in Korean, Singapore, India and Iran in the meta-analysis. Sub-analysis based on the different countries was conducted to verify the heterogeneity, it seemed that there may exist a marginal heterogeneity in Chinese populations (P = 0. 023, I2 = 42%)
Discussion
The etiology of colorectal cancer has not been well established, and previous evidence suggested that the pathogenesis of colorectal cancer was intricate and influenced by complicated interactions between genetic factors and environmental risk. Previous studies reported that the GSTM1 null genotype was in high prevalence in human population, about 40–60% in Europeans29 and about 50% in Asians30. It has been widely accepted that some susceptible genes correlated to colorectal cancer risk, besides smoking, diet and other environmental factors. Up to date, xenobiotic-metabolizing enzymes were primarily concerned on the pathogenesis of cancers31.
GSTM1 is one of the most main subtypes GSTs, which are more effective on protecting host from cancer than others13. GSTM1 null genotype is the most common polymorphism and has been proven to be associated with risk of several cancers15,16, including colorectal cancer17. There were many association studies on relationship between GSTM1 null polymorphism and colorectal cancer in various ethnicities32. Previous validation evidence and meta-analysis studies supported that the GSTM1 null genotype was significantly related to increased risk to colorectal cancer in Caucasian populations32, but the association in Asians has not been revealed yet. Large amount of validation studies have been conducted to detect the association in Asians, but produced inconsistent results. Therefore, we preformed the current meta-analysis to further explore whether the GSTM1 null polymorphism associated with the susceptibility to colorectal cancer.
In our meta-analysis, 33 eligible studies with 8502 colorectal cancer patients and 13699 healthy controls were pooled to calculate the association between GSTM1 polymorphism and colorectal cancer. Significant association between GSTM1 null variant and colorectal cancer in Asian populations was observed in the pooled meta-analysis under both fixed-effect model and random-effect model (Fig. 2a,b). Even though publication bias analyses suggested that obvious bias may exist in the included studies (Fig. 4a–d). The sensitivity analysis indicated the results of meta-analysis were credible and stable (supplementary Figure S1). Furthermore, the cumulative meta-analysis accumulated by publication year or the sample size further confirmed the obvious association of the GSTM1 null variant colorectal cancer in Asians (Fig. 3a,b). In summary, the meta-analysis verified that the GSTM1 null variant was linked to the genetic risk of colorectal cancer in Asians, which in accordance with the previous meta-analysis study26.
Compared to the previous meta-analyses, more validation studies were pooled in the present meta-analysis. The pooled meta-analysis, the cumulative meta-analyses by publication year and the sample size, and the sensitivity analysis correspondingly supported the GSTM1 null polymorphism contribute the genetic risk to colorectal cancer in Asians, especially in Chinese. Nevertheless, it should be noted that there are several limitations in the meta-analysis. First, the association between GSTM1 null variant and colorectal cancer was the only aspect we focused on in this meta-analysis, but the other potential susceptible factors, such as age, sex and smoking status were not considered in the current study, because great majority of these eligible studies did not provide the available information or data. Second, there exist a marginal heterogeneity in Asians (P = 0. 01, I2 = 40.4%). Third, evident bias also exists accordingly to the publication bias analyses. More case-control studies should be added to conduct the meta-analysis, then to further detect the potential gene-gene and gene-environment association.
In summary, the meta-analyses implied that the GSTM1 null variant was significantly associated with the susceptibility to colorectal cancer in Asians, which supporting the genetic factors play vital roles in the pathogenesis of colorectal cancer. Further validation studies should be included to solidify the current conclusions.
Additional Information
How to cite this article: Li, J. et al. GSTM1 polymorphism contribute to colorectal cancer in Asian populations: a prospective meta-analysis. Sci. Rep. 5, 12514; doi: 10.1038/srep12514 (2015).
Supplementary Material
Acknowledgments
We thank all authors of these eligible association studies. This study was supported by two projects: (1) Guangzhou scientific and technological project (No. 2012J4300091).(2) Guangzhou project of clinical and translational research center (early gastrointestinal cancers, No. 7415696196402).
Footnotes
Author Contributions Li Jing gathered and selected the eligible studies, then performed the meta-analysis, and prepared the manuscript. Xu Wen, Fang Liu and Huang Silin participated in the data collecting and selecting process. He Meirong conceived of the study, and participated in its design and coordination and helped to draft the manuscript. All authors reviewed and approved the final manuscript.
References
- Ferlay J. et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. International journal of cancer. Journal international du cancer 127, 2893–2917, 10.1002/ijc.25516 (2010). [DOI] [PubMed] [Google Scholar]
- Cunningham D. et al. Colorectal cancer. Lancet 375, 1030–1047, 10.1016/S0140-6736(10)60353-4 (2010). [DOI] [PubMed] [Google Scholar]
- Center M. M., Jemal A., Smith R. A. & Ward E. Worldwide variations in colorectal cancer. CA: a cancer journal for clinicians 59, 366–378, 10.3322/caac.20038 (2009). [DOI] [PubMed] [Google Scholar]
- Dai Z. et al. [Analysis and prediction of colorectal cancer incidence trend in China]. Zhonghua yu fang yi xue za zhi [Chinese journal of preventive medicine] 46, 598–603 (2012). [PubMed] [Google Scholar]
- Markowitz S. D. & Bertagnolli M. M. Molecular origins of cancer: Molecular basis of colorectal cancer. The New England journal of medicine 361, 2449–2460, 10.1056/NEJMra0804588 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fearon E. R. Molecular genetics of colorectal cancer. Annual review of pathology 6, 479–507, 10.1146/annurev-pathol-011110-130235 (2011). [DOI] [PubMed] [Google Scholar]
- Holley S. L. et al. Polymorphisms in the glutathione S-transferase mu cluster are associated with tumour progression and patient outcome in colorectal cancer. International journal of oncology 28, 231–236 (2006). [PubMed] [Google Scholar]
- Evans D. G. et al. Incidence of hereditary non-polyposis colorectal cancer in a population-based study of 1137 consecutive cases of colorectal cancer. The British journal of surgery 84, 1281–1285 (1997). [PubMed] [Google Scholar]
- Hayes J. D. & Strange R. C. Glutathione S-transferase polymorphisms and their biological consequences. Pharmacology 61, 154–166, 28396 (2000). [DOI] [PubMed] [Google Scholar]
- Strange R. C., Spiteri M. A., Ramachandran S. & Fryer A. A. Glutathione-S-transferase family of enzymes. Mutation research 482, 21–26 (2001). [DOI] [PubMed] [Google Scholar]
- Aghajany-Nasab M., Panjehpour M., Samiee S. M., Rahimi F. & Movahedian A. Glutathione S-transferase mu gene variants and colorectal cancer development--use of sequence-specific probes for an Iranian population. Asian Pacific journal of cancer prevention : APJCP 12, 1511–1515 (2011). [PubMed] [Google Scholar]
- Mannervik B. & Danielson U. H. Glutathione transferases--structure and catalytic activity. CRC critical reviews in biochemistry 23, 283–337 (1988). [DOI] [PubMed] [Google Scholar]
- Strange R. C. et al. The human glutathione S-transferases: a case-control study of the incidence of the GST1 0 phenotype in patients with adenocarcinoma. Carcinogenesis 12, 25–28 (1991). [DOI] [PubMed] [Google Scholar]
- Hengstler J. G. et al. Resistance factors in colon cancer tissue and the adjacent normal colon tissue: glutathione S-transferases alpha and pi, glutathione and aldehyde dehydrogenase. Cancer letters 128, 105–112 (1998). [DOI] [PubMed] [Google Scholar]
- Wang J. et al. Association of GSTT1 gene polymorphisms with the risk of prostate cancer: an updating meta-analysis. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine 34, 1431–1440, 10.1007/s13277-012-0640-8 (2013). [DOI] [PubMed] [Google Scholar]
- Wei Y. et al. Significant associations between GSTM1/GSTT1 polymorphisms and nasopharyngeal cancer risk. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine 34, 887–894, 10.1007/s13277-012-0623-9 (2013). [DOI] [PubMed] [Google Scholar]
- Rebbeck T. R. Molecular epidemiology of the human glutathione S-transferase genotypes GSTM1 and GSTT1 in cancer susceptibility. Cancer epidemiology, biomarkers & prevention : a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 6, 733–743 (1997). [PubMed] [Google Scholar]
- Higgins J. P., Thompson S. G., Deeks J. J. & Altman D. G. Measuring inconsistency in meta-analyses. Bmj 327, 557–560, 10.1136/bmj.327.7414.557 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mantel N. & Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. Journal of the National Cancer Institute 22, 719–748 (1959). [PubMed] [Google Scholar]
- DerSimonian R. & Laird N. Meta-analysis in clinical trials. Controlled clinical trials 7, 177–188 (1986). [DOI] [PubMed] [Google Scholar]
- Lau J., Ioannidis J. P. & Schmid C. H. Quantitative synthesis in systematic reviews. Annals of internal medicine 127, 820–826 (1997). [DOI] [PubMed] [Google Scholar]
- Dickersin K. & Berlin J. A. Meta-analysis: state-of-the-science. Epidemiologic reviews 14, 154–176 (1992). [DOI] [PubMed] [Google Scholar]
- Whitehead A. & Whitehead J. A general parametric approach to the meta-analysis of randomized clinical trials. Statistics in medicine 10, 1665–1677 (1991). [DOI] [PubMed] [Google Scholar]
- Begg C. B. & Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics 50, 1088–1101 (1994). [PubMed] [Google Scholar]
- Egger M., Davey Smith G., Schneider M. & Minder C. Bias in meta-analysis detected by a simple, graphical test. Bmj 315, 629–634 (1997). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai X., Yang L., Chen H. & Wang C. An updated meta-analysis of the association between GSTM1 polymorphism and colorectal cancer in Asians. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine 35, 949–953, 10.1007/s13277-013-1125-0 (2014). [DOI] [PubMed] [Google Scholar]
- Teng Z., Wang L., Zhang J., Cai S. & Liu Y. Glutathione S-transferase M1 polymorphism and colorectal cancer risk in Chinese population. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine 35, 2117–2121, 10.1007/s13277-013-1281-2 (2014). [DOI] [PubMed] [Google Scholar]
- Muellerleile P. & Mullen B. Sufficiency and stability of evidence for public health interventions using cumulative meta-analysis. American journal of public health 96, 515–522, 10.2105/AJPH.2003.036343 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garte S. et al. Metabolic gene polymorphism frequencies in control populations. Cancer epidemiology, biomarkers & prevention: a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology 10, 1239–1248 (2001). [PubMed] [Google Scholar]
- Benhamou S. et al. Meta- and pooled analyses of the effects of glutathione S-transferase M1 polymorphisms and smoking on lung cancer risk. Carcinogenesis 23, 1343–1350 (2002). [DOI] [PubMed] [Google Scholar]
- Norppa H. Genetic susceptibility, biomarker respones, and cancer. Mutation research 544, 339–348 (2003). [DOI] [PubMed] [Google Scholar]
- Economopoulos K. P. & Sergentanis T. N. GSTM1, GSTT1, GSTP1, GSTA1 and colorectal cancer risk: a comprehensive meta-analysis. European journal of cancer 46, 1617–1631, 10.1016/j.ejca.2010.02.009 (2010). [DOI] [PubMed] [Google Scholar]
- Cong N., Liu L., Xie Y., Shao W. & Song J. Association between Glutathione S-Transferase T1, M1, and P1 Genotypes and the Risk of Colorectal Cancer. Journal of Korean medical science 29, 1488–1492, 10.3346/jkms.2014.29.11.1488 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogtmann E. et al. Cruciferous vegetables, glutathione S-transferase polymorphisms, and the risk of colorectal cancer among Chinese men. Annals of epidemiology 24, 44–49, 10.1016/j.annepidem.2013.10.003 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamachi T. et al. CYP1A1, GSTM1, GSTT1 and NQO1 polymorphisms and colorectal adenomas in Japanese men. World journal of gastroenterology : WJG 19, 4023–4030, 10.3748/wjg.v19.i25.4023 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X., Tan Z. R. & Zhang Y. H. [GSTT1 gene polymorphisms and susceptibility to colorectal cancer in Guangxi Zhuang relationship]. J Guangxi Med Univ 29, 106–108 (2012). [Google Scholar]
- Koh W. P. et al. Glutathione S-transferase (GST) gene polymorphisms, cigarette smoking and colorectal cancer risk among Chinese in Singapore. Carcinogenesis 32, 1507–1511, 10.1093/carcin/bgr175 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J. et al. Genetic polymorphisms of glutathione S-transferase genes and susceptibility to colorectal cancer: a case-control study in an Indian population. Cancer epidemiology 35, 66–72, 10.1016/j.canep.2010.07.003 (2011). [DOI] [PubMed] [Google Scholar]
- Nisa H. et al. Cigarette smoking, genetic polymorphisms and colorectal cancer risk: the Fukuoka Colorectal Cancer Study. BMC cancer 10, 274, 10.1186/1471-2407-10-274 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang G. et al. Isothiocyanate exposure, glutathione S-transferase polymorphisms, and colorectal cancer risk. The American journal of clinical nutrition 91, 704–711, 10.3945/ajcn.2009.28683 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yeh C. C. et al. Protein carbonyl levels, glutathione S-transferase polymorphisms and risk of colorectal cancer. Carcinogenesis 31, 228–233, 10.1093/carcin/bgp286 (2010). [DOI] [PubMed] [Google Scholar]
- Piao J. M. et al. Glutathione-S-transferase (GSTM1, GSTT1) and the risk of gastrointestinal cancer in a Korean population. World journal of gastroenterology : WJG 15, 5716–5721 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin L. M. et al. [Glutathione S-transferases polymorphisms and sporadic colorectal cancer susceptibility in Zhejiang province]. Chin J Intern Med 47, 413–414 (2008). [Google Scholar]
- Huang L. R. [Genetic polymorphisms of GSTMl and GSTTl and colorecta tumor susceptibility]. Fujian Medical University (2007). [Google Scholar]
- Xia X. P. et al. [Glutathione S-transferase M1 genotype with ulcerative colitis and colorectal cancer susceptibility]. Chin J Intern Med 46, 583–585 (2007). [Google Scholar]
- Yoshida K. et al. Association of CYP1A1, CYP1A2, GSTM1 and NAT2 gene polymorphisms with colorectal cancer and smoking. Asian Pacific journal of cancer prevention : APJCP 8, 438–444 (2007). [PubMed] [Google Scholar]
- Fan C. H. et al. [Association between genetic polymorphisms of metabolic enzymes and susceptibility of colorectal cancer]. Zhonghua yu fang yi xue za zhi [Chinese journal of preventive medicine] 40, 13–17 (2006). [PubMed] [Google Scholar]
- Fu Q. H. et al. [Polymorphisms of GSTT1, GSTM1 and GSTP1 and susceptibility of colorectal cancer]. Pract J Cancer 21, 247–250 (2006). [Google Scholar]
- Luo J. G., He M. J. & Liu X. H. [Relationship between polymorphisms in glutathione-S-transferase M1 gene and susceptibility to colorectal cancer]. Anat Res 28, 52–54 (2006). [Google Scholar]
- Probst-Hensch N. M. et al. The effect of the cyclin D1 (CCND1) A870G polymorphism on colorectal cancer risk is modified by glutathione-S-transferase polymorphisms and isothiocyanate intake in the Singapore Chinese Health Study. Carcinogenesis 27, 2475–2482, 10.1093/carcin/bgl116 (2006). [DOI] [PubMed] [Google Scholar]
- Chen K. et al. [Associations between genetic polymorphisms of glutathione S-transferase M1 and T1, smoking and susceptibility to colorectal cancer: a case-control study]. Zhonghua zhong liu za zhi [Chinese journal of oncology] 26, 645–648 (2004). [PubMed] [Google Scholar]
- Huang P. et al. [GSTM1 and GSTT1 polymorphisms and colorectal cancer susceptibility in Chongqing people]. Acta Acad Med Militaris Tertiae 25, 1714–1717 (2003). [Google Scholar]
- Yang J., Peng R. X., Kong R. & Le J. [A study on CYP2E1 and GSTM1 gene polymorphisms and colon cancer susceptibility]. Chin Pharmacology 20, 35 (2003). [Google Scholar]
- Seow A. et al. Dietary isothiocyanates, glutathione S-transferase polymorphisms and colorectal cancer risk in the Singapore Chinese Health Study. Carcinogenesis 23, 2055–2061 (2002). [DOI] [PubMed] [Google Scholar]
- Wu M. S. et al. Genetic polymorphisms of cytochrome p450 2E1, glutathione S-transferase M1 and T1, and susceptibility to gastric carcinoma in Taiwan. International journal of colorectal disease 17, 338–343, 10.1007/s00384-001-0383-2 (2002). [DOI] [PubMed] [Google Scholar]
- Zhu Y. Q., Deng C. S., Zhang Y. C., Zhou X. & He X. L. [The relationship between GSTM1, GSTT1 gene polymorphisms and susceptibility to sporadic colorectal adenocarcinoma]. Zhonghua nei ke za zhi 41, 538–540 (2002). [PubMed] [Google Scholar]
- Saadat I. & Saadat M. Glutathione S-transferase M1 and T1 null genotypes and the risk of gastric and colorectal cancers. Cancer letters 169, 21–26 (2001). [DOI] [PubMed] [Google Scholar]
- Zhang Y. C., Deng C. S., Zhu Y. Q., Zhou X. & He X. L. [Relationship between GSTM1 null genotypes and genetic susceptibility to colonic cancers]. Med J Wuhan Univ 22, 131–133 (2001). [Google Scholar]
- Inoue H. et al. Cigarette smoking, CYP1A1 MspI and GSTM1 genotypes, and colorectal adenomas. Cancer research 60, 3749–3752 (2000). [PubMed] [Google Scholar]
- Zhou J. N. et al. [The relationship between polymorphism of GSTM1 and GSTT1 gene andgenetic susceptibility to colorectal cancer]. J Jiangsu Clin Med 4, 90–94 (2000). [Google Scholar]
- Yoshioka M. et al. Glutathione S-transferase (GST) M1, T1, P1, N-acetyltransferase (NAT) 1 and 2 genetic polymorphisms and susceptibility to colorectal cancer. Journal of UOEH 21, 133–147 (1999). [DOI] [PubMed] [Google Scholar]
- Gao J. R., Chen C. F. & Zhang Q. [Study on the relationship between GSTM1 genetic polymorphism and lung cancer, colon cancer susceptibility]. J Zhenjiang Med Coll 8, 446–447 (1998). [Google Scholar]
- Lee E. et al. Genetic polymorphism of conjugating enzymes and cancer risk: GSTM1, GSTT1, NAT1 and NAT2. The Journal of toxicological sciences 23 Suppl 2, 140–142 (1998). [PubMed] [Google Scholar]
- Guo J. Y., Wan D. S., Zeng R. P. & Zhang Q. The polymorphism of GSTM1, mutagen sensitivity in colon cancer and healthy control. Mutation research 372, 17–22, 10.1016/S0027-5107(96)00093-0 (1996). [DOI] [PubMed] [Google Scholar]
- Katoh T. et al. Glutathione S-transferase M1 (GSTM1) and T1 (GSTT1) genetic polymorphism and susceptibility to gastric and colorectal adenocarcinoma. Carcinogenesis 17, 1855–1859 (1996). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.