Abstract
The recent application of mass cytometry (CyTOF) to biology provides a ‘systems’ approach to monitor concurrent changes in multiple host cell factors at the single cell level. We used CyTOF to evaluate T cells infected with varicella zoster virus (VZV) infection, documenting virus-mediated phenotypic and functional changes caused by this T cell tropic human herpesvirus. Here we summarize our findings using two complementary panels of antibodies against surface and intracellular signaling proteins to elucidate the consequences of VZV-mediated perturbations on the surface and in signaling networks of infected T cells. CyTOF data was analyzed by several statistical, analytical and visualization tools including hierarchical clustering, orthogonal scaling, SPADE, viSNE, and SLIDE. Data from the mass cytometry studies demonstrated that VZV infection led to ‘remodeling’ of the surface architecture of T cells, promoting skin trafficking phenotypes and associated with concomitant activation of T-cell receptor and PI3-kinase pathways. This method offers a novel approach for understanding viral interactions with differentiated host cells important for pathogenesis.
1. Introduction
Varicella Zoster virus (VZV) is a human alphaherpesvirus that causes varicella (or chicken pox) as the primary infection, and zoster (or shingles), caused by reactivation from latency in sensory ganglion neurons. VZV has a 125 kbp DNA genome coding for 70+ proteins. Although VZV is neurotropic, like other alphaherpesviruses, its T cell tropism is a unique and essential part of the viral life cycle in the human host [1]. Based on clinical observations and our studies of VZV pathogenesis in the severe combined immunodeficiency mouse model, T cells in tonsils and regional lymphoid tissue become infected by spread of the virus from mucosal epithelial sites of inoculation and transport the virus to skin where lesion formation allows transmission to other susceptible hosts [2,3]. Studying VZV interactions with differentiated human T cells has been challenging because of the intrinsic heterogeneity of these cells. In addition, VZV inoculum titers that can be generated in tissue culture do not allow a fully synchronous infection of primary T cells in vitro. The recent application of single cell mass cytometry to biology offers a powerful new strategy to simultaneously measure the induction or increased expression of multiple proteins occurring within a diverse cell population [4]. We have used this approach to investigate VZV T cell tropism as a model system to determine the efficacy of single cell mass cytometry for understanding virus-host cell interactions that underlie a critical event in pathogenesis [5].
Single cell mass cytometry resembles flow cytometry in using antibodies specific for known proteins to profile their expression in individual cells. However, antibodies are tagged with lanthanide metal-isotopes which are then analyzed by time of flight mass spectrometry. Using elemental tags overcomes the technical problems of background autofluorescence and the need to compensate for spectral overlap commonly encountered in using flow cytometry. As a result, the mass cytometry approach allows the detection of many more proteins in each cell. Proof of concept for this method was obtained first in investigations of tumor cell and immune cell populations, including the evaluation of phenotypic and signaling changes in human T cells following stimulation with exogenous cytokines [4]. Therefore, we hypothesized that single cell mass cytometry could be used to reveal the consequences of viral infection on differentiated cells targeted during pathogenesis, including their surface characteristics and the signaling pathways that regulate these characteristics, and specifically, that this approach would generate new knowledge about how VZV infection affects the biology of human tonsil T cells [5].
Because single cell mass cytometry experiments generate an enormous number of data points, data visualization and interpretation requires the application of statistical methods appropriate for large-scale data analysis as well as the development of new analytical tools. These approaches include orthogonal scaling (Principal Component Analysis; PCA), agglomerative hierarchical clustering, and computational algorithms like Spanning Tree Progression Analysis of Density Normalization Events (SPADE) [6,7]. As described, we also developed Single Cell Linkage using Distance Estimation (SLIDE), a novel statistical algorithm, during the course of our studies of VZV-T cell interactions [5]. This method, based on principles of nearest neighbor analysis, revealed that the phenotypic profiles of VZV-infected T cells are a consequence of remodeling of differentiated T cells by the virus following entry and replication, instead of reflecting a process of preferential infection of T cells with particular phenotypes, as we had assumed to be the case from our prior studies of VZV pathogenesis [5,9]. Here we have also used our data set to demonstrate the applicability of a new visualization tool based on the t-Distributed Stochastic Neighbor Embedding algorithm (viSNE), which confirmed conclusions from our prior analyses [8].
Strikingly, remodeling due to VZV infection as demonstrated by single cell mass cytometry was associated with enhanced expression of cell surface proteins on memory T cells that are known to facilitate their trafficking to skin, or with the induction of these proteins on VZV-infected naive T cells [5]. Thus, the fate of the T cell was altered by VZV infection to favor targeting to skin where the virus is released for replication. Skin infection results in the formation of cutaneous lesions that contain high concentrations of infectious virus, which allows rapid spread within the host population. Here, we summarize our experience with the single cell mass cytometry method and the analytic tools that supported these conclusions as an example of the potential uses of this strategy to probe how other viral pathogens may engineer the take over of fully differentiated host cells important for the life cycle of the virus in its natural host.
2. Materials and Methods
2.1. T cell preparation and VZV infection
As previously described, human tonsils were obtained according to a protocol approved by the Stanford University Committee for Research involving human subjects [2,5]. T cells were purified from tonsil specimens collected at surgery from healthy donors undergoing routine tonsillectomy. A single cell suspension of homogenized tonsil tissue was prepared and subjected to nylon wool-column purification to separate T cells from macrophages and B cells. The eluate containing about 75–80% T cells was then further enriched to up to 99% purity using the EasySep Human T cell (STEMCELL Technologies, Vancouver, Canada). The purified T cells were infected with recombinant VZV, rOka-ORF10-GFP, expressing ORF10 as an ORF10-GFP fusion protein (VZV-GFP) cultivated in human embryonic lung fibroblasts (HELF) [5]. Approximately 20–25 million purified T cells were co-cultured with VZV-GFP-infected or mock-infected HELF for 24 hr, collected and incubated in tissue culture media for another 24 hr. The efficiency of T cell infection was monitored by fluorescence microscopy and confirmed by flow cytometry using antibodies to CD3 and CD4 T cell markers and GFP to detect virus infection.
2.2. Staining T cells for Mass Cytometry
2.2.1 Antibodies
The antibodies used and their elemental isotope tags, clone, and source are listed in Table 1. Antibodies to cell proteins were either obtained from DVS Sciences (Richmond Hill, Ontario, Canada) or prepared in the Nolan Lab, Stanford University [4]. VZV gE was chosen as the viral marker to identify infected cells from bystander cells; it is an abundant surface glycoprotein expressed at early times post infection [10]. For gE conjugation to lanthanide Dy164, 100 μg of purified monoclonal gE antibody (EMD Millipore Corp., Billerica, MA) was labeled using the MaxPAR antibody conjugation kit (DVS Sciences, Toronto, Canada).
Table 1.
Antibodies and Elemental Isotope Tags.
| Antigen Target (Human) | Clone Number | Supplier | Elemental Isotope | Final Concentration (μg/mL) |
|---|---|---|---|---|
| CD10 | HI10a | Biolegend | Gd156 | 3 |
| CD11c | 3.9 | Biolegend | Tb159 | 5 |
| CD3ε | UCHT1 | DVS | Er170 | 3 |
| CD3δ (Qdot 655) | S4.1 | Invitrogen | Cd110, 111, 112, 114 | 1:500 |
| CD33 | 67.6 | BD Biosciences | Yb173 | 1.5 |
| CD34 | 8G12 | BD Biosciences | Nd148 | 3 |
| CD38 | HIT2 | Biolegend | Er168 | 1 |
| CD4 | RPA-T4 | DVS | Nd145 | 1:100 |
| CD44 | G44-26 | BD Biosciences | Yb171 | 1 |
| CD45 | HI30 | Biolegend | In115 | 2 |
| CD45RA | HI100 | DVS | Eu153 | 1:100 |
| CD7 | M-T701 | BD Biosciences | Yb176 | 2 |
| CD8a | RPA-T8 | DVS | Nd146 | 1:100 |
| IgM | G20-127 | BD Biosciences | Lu175 | 2 |
| CD22 | HIB22 | Biolegend | Nd143 | 3 |
| CD49d | 9F10 | Biolegend | Nd144 | 2 |
| CD57 | HCD57 | Biolegend | Sm147 | 1 |
| CD45RO | UCHL1 | Biolegend | Sm149 | 4 |
| CD11a | HI111 | Biolegend | Nd150 | 2 |
| CD127 | HCD127 | Biolegend | Yb172 | 4 |
| CTLA4 (CD152) | L3D10 | Biolegend | Yb174 | 2 |
| CD69 | FN50 | DVS | Dy162 | 1:100 |
| CCR7 | 150503 | RnD Systems | Er166 | 2 |
| CD27 (DVS) | L128 | DVS | Er127 | 1:100 |
| CD25 (DVS) | 2A3 | DVS | Tm169 | 1:100 |
| CD279 (PD-1) | EH12.1 | BD Biosciences | Ho165 | 4 |
| CD28 | CD28.2 | BD Biosciences | Pr141 | 2 |
| IκBα | L35A5 | Cell Signaling Technology | Er166 | 2 |
| CREB (pS133) | 87G3 | Cell Signaling Technology | Yb176 | 1 |
| MAPKAPK 2 (pT334) | 27B7 | Cell Signaling Technology | Gd158 | 2 |
| S6 (pS235/pS236) | N7-548 | BD Biosciences | Yb172 | 2 |
| SLP76 (pY128) | J141-668.36.58 | BD Biosciences | Gd156 | 2 |
| ZAP70/Syk (pY319/pY352) | 17a | BD Biosciences | Gd160 | 2 |
| cPARP | F21-852 | BD Biosciences | La139 | 1 |
| 4EBP1 (T37/46) | 236B4 | Cell Signaling Technology | Nd143 | 3 |
| FAK (pY397) | Polyclonal | Cell Signaling Technology | Nd148 | 3 |
| AKT (S473) | D9E | Cell Signaling Technology | Tb159 | 4 |
| Erk1/2 (T202/Y204) | 197G2 | Cell Signaling Technology | Er167 | 3 |
| CD3z (Y142) | K25-407.69 | BD Biosciences | Lu175 | 3 |
2.2.2. Staining Procedure
For each experiment, approximately 20 million T cells were harvested 48 hpi. The T cells were treated immediately with 25 μM cisplatin (Enzo Life Sciences, Farmingdale, NY) for 1 min at room temperature and the reaction was quenched by adding RPMI + 10% FCS. Cisplatin is a platinum-containing drug that when added to cells in a one minute incubation step labels non-viable cells thus allowing identification of viable cells for downstream analysis [11]. The cells were then fixed in 1.6% paraformaldehyde and stored at −80°C in aliquots of 2–3 million cells. For staining, the aliquots were thawed on ice and treated with isotope-labeled antibodies. The antibodies to surface proteins (Table 1) were added in a total reaction volume of 100 μl; the cells were incubated on ice for 30 min, washed and permeabilized with cold methanol for 10 min at 4°C. After permeabilization, the cells were incubated with antibodies to the intracellular proteins (Table 1) for 30 min on ice. The washed cells were resuspended in 1:4000 191/193Ir DNA intercalator [12] (DVS Sciences, Richmond Hill, Ontario, Canada), incubated overnight at 4°C, and washed and resuspended in distilled water prior to analyzing on the CyTOF™ mass cytometer (DVS Sciences) [13]. Cytometer setting and data acquisition was done as described previously, with approximately 200,000 events acquired per sample [4]. To make all samples maximally comparable, all data was acquired using internal metal isotope bead standards and normalized to the standards before data analysis [14].
3. Data Analysis and Results
In order to determine how VZV infection altered expression of T cell surface and intracellular proteins, we used single cell mass cytometry analysis of VZV-GFP and mock-infected T cells to generate an equivalent of ~107 single cell measurements/experiment. These large-scale data sets were then analyzed using statistical and visualization tools as described below, with special emphasis on SLIDE, the novel algorithm that we developed for this purpose.
3.1. Preliminary Data Filtration
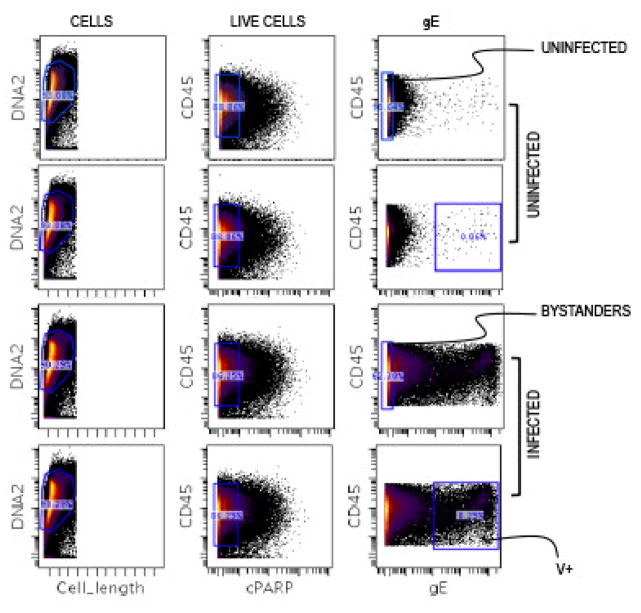
Normalized and background subtracted FCS files were imported from the mass cytometer into Cytobank (https://www.cytobank.org) [15]. Cells used in the single cell analyses were extracted following sequential gating for DNA content, control beads, viability and VZV gE expression (Fig. 1). Typically in all the experiments, ~10% of the T cells were infected by VZV (gE+). The T cells that were co-cultured with VZV-infected HELF were classified into infected (V+) and bystander (Bys) T cells, using a highly conservative gating strategy based on high levels of gE expression to identify V+ T cells. Uninfected (UI) T cells were prepared by co-culture with HELF.
Fig. 1. Gating Scheme.
Normalized data collected on the mass cytometer was transferred to www.cytobank.org. The T cells were consequently sequentially gated for DNA content to identify nucleated cells, and viability markers to extract uninfected (UI), bystander (Bys) and VZV-infected (V+) populations for downstream analysis. The UI cells chosen for comparative analysis were matched to be on the co-ordinates as the Bys cells from the VZV-infected sample (containing both Bys and V+ cells).
Two antibody panels were designed to determine the phenotypic (25 cell surface proteins; Panel 1) and functional characteristics (12 intracellular phosphoproteins; Panel 2) of V+ T cells compared to the Bys and UI T cells. The proteins recognized by these antibodies included well-defined cell surface markers that differentiate the major T cell subpopulations and identify functions such as tissue migratory phenotypes, and those that are components of established signaling pathways (Table 1). Based on conventional histogram analysis to detect overall expression/non-expression of each tested protein, we confirmed that immune cells expressing B cell and dendritic cell markers, e.g., IgM and CD11c, were not present, while those expressing CD22, CD57, CD34, CD10, CD33, and CTLA4 were not detected or were present at very low levels. The phosphorylation state of the 12 intracellular signaling proteins was assessed along with expression of 11 surface proteins and VZV gE. In each of these datasets, the UI and Bys population sizes were ~105 cells whereas the V+ population was ~104 T cells. Following population gating on Cytobank, the data files were either analyzed by SPADE or viSNE on Cytobank or were exported in the text format and used in downstream analysis by PCA and hierarchical clustering with the commercially available Partek Genomic Suite software 6.0, and for SLIDE.
3.2. Principal Component Analysis and Agglomerative Hierarchical Clustering
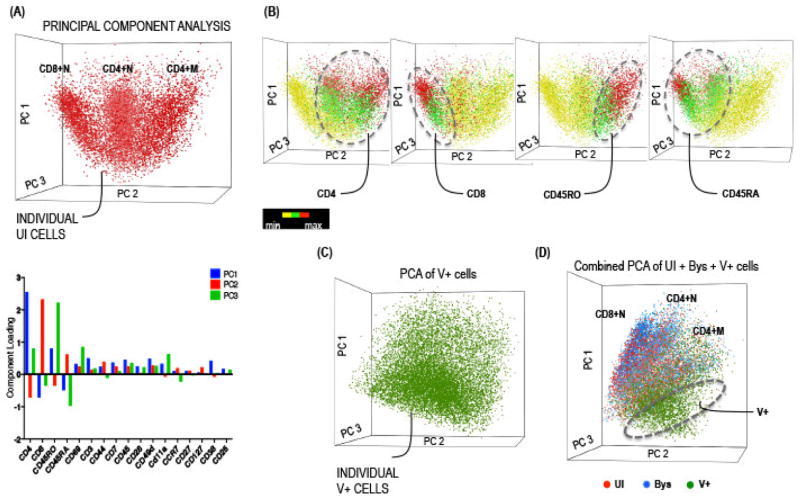
First, the phenotypic profile and hierarchy of the UI tonsil T cells (baseline control) was determined for comparison with the profiles exhibited by V+ and Bys T cells. We used the PCA-based dimension reduction technique on the multi-dimensional data cloud representing all of the surface markers to obtain three orthogonal directions that explained the maximum variability of the data [16]. The PCA scatter plot in which each cell in the reduced dimensional space is represented as a dot, demonstrated that the variability (~60%) captured by the first three components can be largely attributed to the differential expression of four core markers, including CD4, CD8, CD45RO (memory) and CD45RA (naive) T cell proteins (Fig. 2A). Thus, UI T cells were distributed into three predominant cell clouds corresponding to CD4+ memory (M), CD4+ naive (N), and CD8+N T cells; the distribution and expression of each of the four cluster determining factors, CD4, CD8, CD45RO and CD45RA, on UI T cells is shown in Fig. 2B. This analysis was an important first step because a comprehensive survey of tonsil T cell subpopulations had not been reported previously. When Bys and V+ T cells were analyzed similarly, the Bys T cell distribution was identical to UI T cells whereas that of V+ T cells was distinct (Fig. 2C–D). To determine the phenotypic distribution of the V+ compared to UI and Bys T cells, an equivalent number of cells from UI, Bys, and V+ groups was pooled (n= ~5000 cells per group per experiment) and scatter plots were generated by PCA. Because there were fewer V+ T cells, they were compared with an equivalent randomly selected number of UI and Bys T cells in multiple reiterative analyses. The data reproducibly demonstrated that V+ T cells formed a distinct cluster when compared with UI and Bys cells (Fig. 2D).
Fig. 2. Principal Component Analysis.
(A) 3D visualization of UI T cells on a PCA scatterplot obtained using the Partek Genomic software. Each dot on the plot represents a cell, the location of which is determined by the co-ordinates of its first three principal components. The PCA parameter loading components for the first three components, Principal component 1 (PC1; blue), PC2 (red), and PC3 (green), for UI T cells was plotted as a bar diagram shown below the scatterplot. The UI cells distributed into three major subpopulations corresponding to CD4+ memory, CD4+ naive, and CD8+ naive T cells. (B) The subpopulations were annotated based on the expression levels of each of the four core markers, CD4, CD8, CD45RO and CD45RA, ranging from minimum (yellow) to maximum (red). (C–D) PCA scatter plot of V+ T cells and of V+ T cells (green dots) pooled with an equivalent number of UI (red dots) and Bys (blue dots) T cells obtained as described above. The hashed sphere indicates the unique location of V+ cells compared to the overlapping UI and Bys cells. The V+ T cells were iteratively compared with randomly selected equal number of UI and Bys T cells and a representative iteration is shown here.
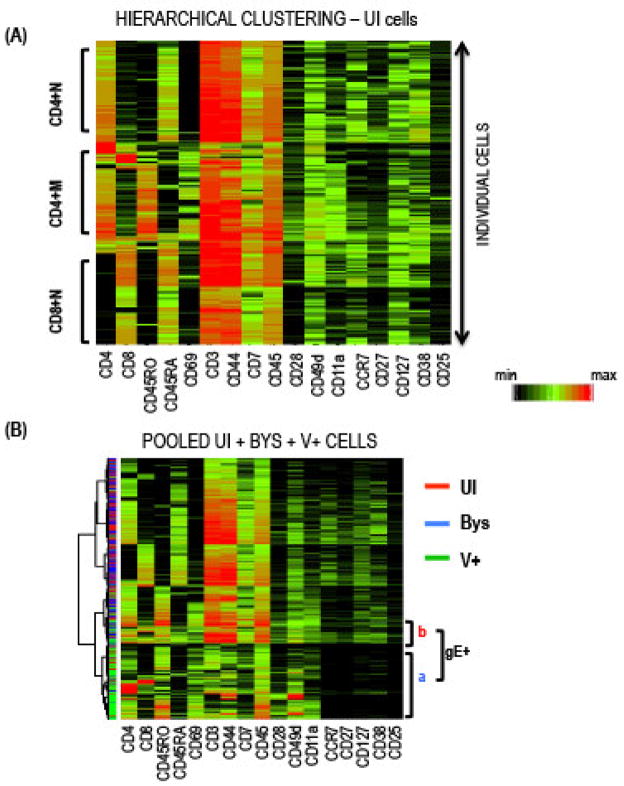
Agglomerative hierarchical clustering measures similarity between cells by linking them into the same or different groups based on assigned parameters. We used 17 T cell surface proteins for this analysis (Fig. 3). In the Partek software, the cells were clustered using the Euclidean dissimilarity measure based on all the phenotypic markers and Ward’s minimum variance method. The results were represented as a heatmap where each row and column represented a single T cell and the intensity of protein expression, respectively (Fig. 3A). Similar to PCA, the clustering analysis showed that (a) the UI T cells from tonsils could be categorized into three major subpopulations based on the expression of the four core proteins, and (b) the V+ T cells formed a distinct hierarchical group when analyzed together with an equal number of UI and Bys T cells (Fig. 3B), supporting the PCA analysis shown in Fig. 2D.
Fig. 3. Agglomerative Hierarchical Clustering.
(A) Heatmap displaying the hierarchical clustering of UI T cells; each row of the heatmap represents a cell and each column the indicated protein; the color scale indicates protein expression levels from minimum (black) to maximum for each protein. Dendrograms on the left denote the Euclidean distance between the clustered cells. (B) Hierarchical relationship of pooled UI, Bys, and V+ T cells based on surface proteins by PCA (left) and agglomerative clustering (right). A majority of the V+ cells clustered together on the heatmap distinct from the overlapping UI and Bys cells indicated by the red and blue lines, respectively, shown on the left of the heatmap.
3.3. Spanning Tree Progression Analysis of Density Normalization Events
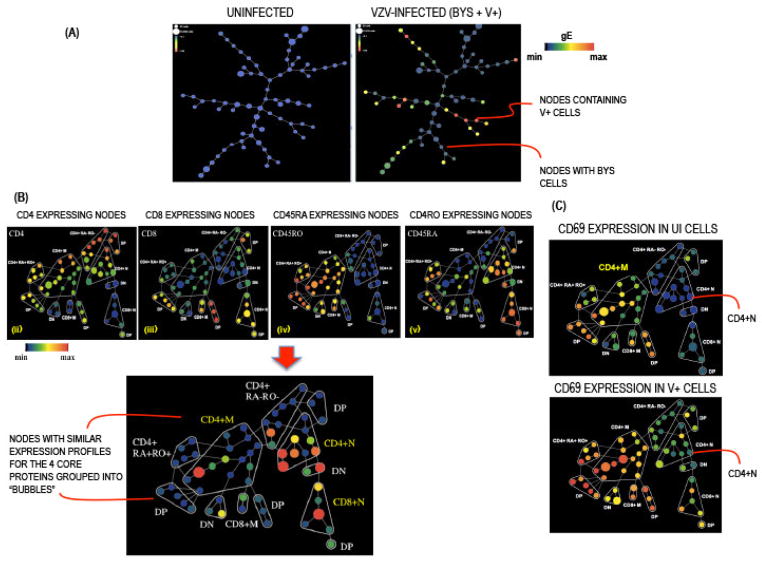
SPADE is a module of four analytical components including downsampling, agglomerative clustering, linking by minimum spanning, and upsampling that assembles cells with common protein expression profiles into hierarchies that reveal subgroup characteristics [7]. SPADE was developed for mass cytometry data analysis and is an important tool because it identifies rare populations of cells [4]. The SPADE analysis output consists of a tree with interconnected nodes; each node is a collection of cells with similar phenotypic profiles and is connected to nodes that are most closely related. SPADE trees were constructed on www.cytobank.org using default settings with a user defined target of 50 nodes and clustering channels, in this case the four core proteins, CD4, CD8, CD45RO and CD45RA. The SPADE program distributed the cells into nodes visualized as colored circles where color is proportional to the average intensity of expression of a given parameter and size represents the number of cells in that node. As shown in Fig. 4A, gE expression was only detected in the infected group of cells (showing both the Bys and V+ T cells) and its intensity of expression varied from high (red) to low (green) to none (blue; Bys T cells). For downstream SPADE analysis, we evaluated the Bys and the V+ T cells separately. The SPADE trees for UI, Bys, and V+ cells were annotated into subpopulations or “bubbles” based on expression of the four core proteins (Fig. 4B). The expression patterns of each of the surface proteins and intracellular signaling proteins were subsequently visualized on the annotated trees generated for each of the UI, Bys, and V+ groups [5].
Fig. 4. Spanning Tree Progression Analysis of Density Normalization Events.
(A) SPADE tree from a representative experiment in which the UI (left panel) and VZV-infected (unseparated Bys and V+ T cells; right panel) T cells were distributed into 50 nodes based on expression of the four core markers CD4, CD8, CD45RO and CD45RA; each node denotes a collection of cells with a similar profile placed in proximity to nodes with closely related profiles. The size of the nodes is directly proportional to the number cells in that node. The color of the nodes from blue (minimum) to red (maximum) denotes increasing median intensity of gE expression in UI and VZV-infected samples. (B) The nodes on the SPADE trees were annotated into subpopulations based on the expression of CD4, CD8, CD45RO and CD45RA as shown in respective panels; DP = double positive (CD4+CD8+), DN = double negative (CD4−CD8−). (C) SPADE trees showing the differential expression of CD69 on UI and V+ T cells across phenotypic hierarchies.
The SPADE trees revealed the differential expression patterns of each protein within the UI, Bys, and V+ T cells and also between the three groups. For example, in the UI group, CD69 expression was relatively higher in the CD4+M (memory) T cells compared to the CD4+N (naive) T cells. Between the UI and V+ groups, expression of CD69 was higher both in the CD4+M and CD4+N subpopulations in the V+ group compared to the corresponding UI subpopulations (Fig. 4C). A similar comparison made between the UI-Bys and Bys-V+ T cells revealed that the expression profiles of most of the proteins tested were modulated on the V+ T cells compared to the UI and Bys T cells, which displayed nearly identical profiles. These changes were bi-directional, including increased expression of CD69, CD28, CD49d, CD11a and programmed cell death-1 (PD-1; CD279) and reduced expression of CD3, CD7, CCR7, CD27, CD44, CD38 and CD127. CD45 and CD3D expressions were unchanged [5].
The increased expression of CD69, CD279 and CD28 together with decreased CCR7, CD27, and CD127 were indicative of activated T cells with tissue homing characteristics. These surface characteristics of V+ T cells were further modulated for skin migration by the increased expression of integrins, CD11a and CD49d, and reduced CD7 surface expression. Preferential migration of T cells with these surface features to the skin was demonstrated in cutaneous T cell lymphoma (CTCL). These observations were further supported by flow cytometry analysis that showed VZV-associated modulation of CXCR5 and CLA expression, which determine emigrant (from tonsils) and skin homing phenotypes. In summary, most infected cells displayed properties typical of activated effector memory T cells with tissue homing traits that are preferential for skin trafficking, and would be expected to lack immune function capacity.
3.4. Single Cell Linkage using Distance Estimation
Although PCA scatter plots and agglomerative clustering heatmaps showed that the V+ T cells express distinct phenotypic profiles compared to UI and Bys T cells as a consequence of modulation of the expression of several surface proteins (SPADE tree), the question remained whether the V+ T cells represented a subpopulation of T cells that were rare among the initial UI T cell population but were infected preferentially, or whether the V+ T cells had been remodeled as a consequence of VZV infection. In order to answer this question, we developed SLIDE, which is an analytical method that measures ‘distances’ between individual cells based on their expression of surface proteins. SLIDE was specifically designed to take into account the complexities of two-sample comparisons, each with a different sample size. Unlike unsupervised methods like SPADE which cluster cellular populations into varied sub-populations, SLIDE was designed to optimally test for homogeneity between two clearly demarcated sub-groups of cells.
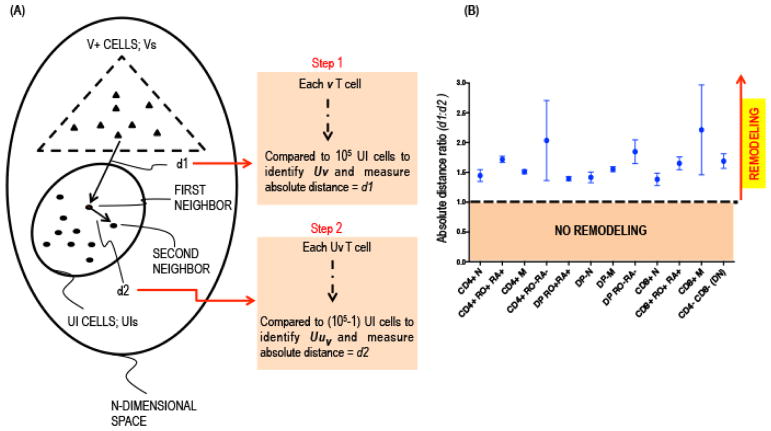
As an initial step, the T cells from the UI and V+ groups were clustered into 16 subpopulations based on the binary expression of the four core proteins, CD4, CD8, CD45RO, and CD45RA. We applied the SLIDE test to each subpopulation to determine statistically significant changes in the average prototype between the corresponding UI and V+ T cells. For each sub-population, we investigated the hypothesis of remodeling of the characteristics of cells in the V+ group (Vs) compared to the respective UI group (UIs). To study remodeling in a group of T cells that has a different sample size, the V+ group in this case, from the control UI group, we applied the nearest neighbor approach [17,18]. Each VZV-infected T cell (v in Vs) was projected into its corresponding UI group (UIs) and the nearest uninfected T cell (uv) with similar expression pattern for the 13 surface markers was extracted and the absolute distance, d1, measured (Fig. 5A), i.e.,
Fig. 5. Single Cell Linkage using Distance Estimation.
(A) Schematic representation of single cell linkage distance estimation (SLIDE) analysis. Each V+ T cell within any given subpopulation was matched with every UI T cell to identify its closest neighbor, UIv, based on combinatorial expression of 13 non-core surface markers and the absolute distance between the 2 cells were recorded as d1. The UIv T cell was subsequently matched to its nearest neighbor, Uuv, in the UI population and the distance between the two UI cells was measured as d2. (B) The d1:d2 ratio for each cell in a given subpopulation (x-axis) was calculated, averaged and plotted (y-axis) with confidence intervals incorporating 95% of V+ T cells.
for each v ∈ Vs we find:
Each of the nearest neighbor UI cell (uv) was further projected into the remaining UI data cloud (UIs − uv) and their distances (d2{v}) from UIs{v} was also evaluated, i.e.,
for each v ∈ Vs & uv we find:
If m and n, respectively, are the sizes of the UI and V+ sub-populations then for each V+ cell, the first step involves m comparisons and the second step involves m−1 comparisons (Fig. 5A). Typically for UI and V+ groups, m ~ 105 and n ~ 104 and so the number of comparisons computed was in the order of ~ 109.
If m is large, under the null hypothesis of no remodeling of the cell surface markers in the V+ T cells, the ratio of the distances r(v) = d1{v}/d2{v} for v ∈ Vs will have a nearly symmetric distribution around 1. Under the alternative hypothesis of remodeling in the V+ cell population, the V+ T cells would be distant from its UI neighbor and the ratio r(v) would have significant upward distortion from 1. We used non-parametric tests to compare the paired distances d1 and d2 for all viral cells in any given sub-population. From these distance estimations, the average d1:d2 ratio was calculated for each subpopulation and plotted on the y-axis (Fig. 5B). The average d1:d2 ratio (y-axis) was ≫1 in all subpopulations (x-axis), thereby providing statistical evidence that VZV infection directs remodeling of the surface architecture of the infected T cells. Once significant evidence for VZV remodeling of the average prototype was confirmed, co-ordinate wise tests with False Discovery Rate (FDR) based multiplicity corrections were conducted to detect the nature of changes in the remodeled virus-infected T cell clusters [5]. In the context of VZV infection and the data obtained by SPADE analysis, the bi-directional changes in expression of the surface proteins that promote skin trafficking was induced during infection, resulting in remodeled T cells that would more effectively transport VZV to the skin.
3.5. t-Distributed Stochastic Neighbor Embedding
An advanced analytical tool called viSNE was recently made available on Cytobank for analysis of data generated using CyTOF. viSNE allows visualization of single cell data based on the t-Distributed Stochastic Neighbor Embedding (t-SNE) algorithm [8]. The viSNE map is represented as a bi-axial scatter plot in which every dot is a single cell and its location on the plot is based on non-linear 2D-projection of the combinatorial expression of all dimensions, where a protein contributes each dimension. viSNE maps differ from PCA scatter plots where the position of the cells is determined by a linear 2D-projection in which it was empirically observed that the expressions of dominant dimensions or markers for example, CD4 and/or CD8 play a significant role. From viSNE maps, the phenotypic profile of single cells can be visualized with reference to all other cells. We applied the program to generate two separate viSNE maps for ~ 3×105 VZV-infected T cells (unseparated Bys and V+ T cells) based on (a) the expression of all 17 T cell surface markers (Figs. 6A–B) and (b) the expression of 13 non-core surface markers, excluding the four core proteins and gE (Figs. 6C–D). Each dot (cell) on the viSNE map is colored on a scale ranging from blue through green, indicating increasing expression of the indicated protein and the x- and y-axis represent the tSNE values in the non-linearly reduced 2D space. As seen in Figs. 6A and 6C, the V+ T cells (green dots) formed a cell cloud distinct from the Bys T cells (blue dots) that is consistent with Fig. 3B where the V+ T cells were pooled with an equal number of UI and Bys T cells and analyzed by PCA and clustering. Since viSNE also calculates distances between cells to determine their location on the map based on expression of all proteins, this analysis also underscores our finding that the V+ T cells occupy a unique hierarchical position due to the remodeling capacity of VZV infection as determined by SLIDE. Shown in Figs. 6B and 6D are examples of the expression levels of three surface proteins CD49d, CD3 and CD7 in V+ T cells relative to the Bys cells.
Fig. 6. t-Distributed Stochastic Neighbor Embedding algorithm.
viSNE maps showing the distribution of the VZV-infected T cells based on the expression (A) 17 surface markers including the core proteins and (B) 13 non-core surface markers. Each dot on the biaxial scatter plot represents a cell and the intensity of protein expression is determined by the color ranging from blue (minimum) to green (maximum). (C–D) visNE maps, generated using conditions A & B, showing the expression pattern of CD49d, CD7, and CD3 in Bys and V+ cells.
3.6. Activation Index of Signaling Proteins
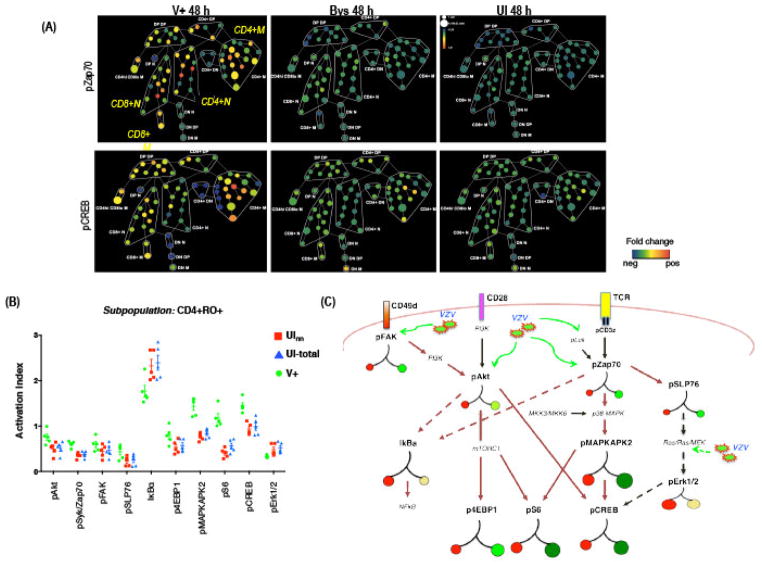
The SPADE analysis revealed differential effects of VZV infection on signaling proteins and pathways across different T cell subpopulations. As an example, the SPADE trees showing fold increase in expressions of phospho-Syk/Zap and pCREB in Bys, and V+ T cells relative to UI T cells (baseline) are given in Fig. 7A. Most nodes on the V+ T cell SPADE tree showed increased levels of pSy/Zap70 and pCREB while the Bys T cells showed no increase (fold over the UI) in expression of the two proteins. Since the overall increase in expression values for a given protein is a function of both higher signal intensity within a single cell and/or induced expression in higher proportion of total T cells, we derived an Activation Index (AI) parameter for each signaling protein [5]. AI is defined as the product of the mean intensity of expression (standardized) and the proportion of T cells expressing the protein within a specific subpopulation. Since the expression intensity is variable across experiments, we standardized the expression values in the V+ cells based on the corresponding UI group for each experiment. Thus, if X = {X1, X2, …, Xm} is the vector containing the standardized intensities of all the expressed values of the protein p in a sub-population S then the corresponding activation index was given by
Fig. 7. Activation Index of Signaling Proteins.
(A) SPADE tree showing the fold-change in expression levels of pZap70 and pCREB in V+ and Bys T cells relative to UI T cells by SPADE analysis. In this case the node colors indicate fold-change in expression from minimum (blue) to maximum (red). (B) AIs for each phosphoprotein in V+, UV, and total UI T cells in the CD4+RO+ subpopulation was plotted. The mean AI (±SD) (y-axis) for each protein in V+, UInn, and total UI T cells was calculated from five independent experiments. P values were determined by one-sided Student t test (p <0.05) for each protein compared between V+ and UV T cells and for V+ and total UI T cells. (C) Schematic representation of the T cell signaling pathways indicating the hierarchical locations of the different phosphoproteins that were analyzed in this study. VZV particles and green arrows indicate the points at which activation (solid lines) or inhibition (dashed lines) of the signaling pathway was detected in V+ T cells. The colored circles represent the activation index (AI) for each phospho-protein in V+ and UI T cells. Basal activation state of each protein in UI T cells is represented as red circles on the left branch and the size is proportionate to the AI. The right branches indicate the fold changes in AI for each protein in V+ T cells relative to UI T cells, green circles indicate an increase and yellow circles indicate a decrease in AI. Proteins indicated in small font were not tested but are important intermediary components of the T cell pathways.
where |S| denotes the population size and denotes the intensity of protein expression. AI can be also be expressed as the product of the proportion expressed (α̂) and the mean of only the expressed values ( ). The fold increase between the UI and V+ populations was given by where pv, pu are proportion expressed and are the mean of the expressed values in original scale in the V+ and UI populations, respectively.
Fig. 7B shows the activation index (AI), product of signal intensity and proportion of positive cells, of the signaling proteins in T cells from the CD4+M memory subpopulation combined over five independent experiments. The mean AI of each signaling protein in V+ T cells (green) was calculated and plotted against the mean AI of the UI nearest neighbor (Uv; red) and total UI T cells. The results indicated that VZV infection stimulates the pSyk/Zap70 and the pAkt pathways. Based on what has been established about the regulation of T cell surface proteins, this remodeling of the surface proteins can be explained as a consequence of perturbation of the T cell signaling pathways by VZV infection of T cells.
3.7. Signaling pathway Activation
To study the stochastic nature of the flow of activation signal through different phospho-proteins within a given pathway, viz., the pSyk/Zap70 and pAkt pathways, we discretized the continuous expression values of the proteins into two categories: (i) none/low or (ii) high. For each protein, V+ T cells that showed expression above the 70th percentile of the overall expression in the UI T cells, were considered to be highly expressing that protein. With this discretization strategy, we determined how the proportions of signal flow through different pathways were altered during VZV infection. In the continuous set-up, high positive correlations confirmed the hypothesis of increased flows in the V+ cells. The change in each signaling protein was quantified and visualized on a simplified pathway chart (Fig. 7C). Applying single cell resolution methods, we found that V+ T cells had an overall increase in expression intensity at each node along both the pSyk/Zap70 and pAkt pathways. Interestingly, despite increased pSyk/Zap70, no concomitant increase in pErk1/2 was detected which suggests that VZV may make proteins during its replication cycle that block the pSyk/Zap70-pErk1/2 pathway, thereby preventing cytokine synthesis and secretion.
4. Discussion
Recognition of the inherent heterogeneity of the biological systems within differentiated mammalian cells has underscored the importance of studying virus host interactions in single cells. Given the many viruses that are important human pathogens, evaluating the complex interplay of proteins, both host cell and virus derived, within infected host cells, is critical to achieving a better understanding of the basic biology that underlies the diseases they cause. This knowledge is relevant for the development of antiviral drugs and new strategies for designing attenuated viral vaccines. Our experience with studies of VZV T cell tropism is that single cell mass cytometry is a valuable new tool to probe these interactions.
To place the utility of single cell mass cytometry in context, most studies of the consequences of viral replication have been evaluated in cells or cell lines considered to resemble the target cell and infected in bulk culture. Studying specific viral and cell proteins and their interactions during infection of a uniform population of cultured cells provides important insights about effects identifiable by averaged measurements. However, under physiological conditions viral pathogens encounter differentiated target cells exhibiting inherently stochastic states and the process of altering conditions to support pathogenesis must occur within each cell. Differential viral replication kinetics and variability in levels of viral protein expression could further add to the differences in responses of each cell to the pathogen. Under such circumstances, the traditional methods providing averaged measurements from a bulk cell population could mask effects; for example, bi-directional changes when a protein is upregulated in some cells and downregulated in others would be recorded as no or only a modest effect and small but important variations in expression could be undetectable. Obtaining precise information about simultaneous multi-parametric changes within individual cells provides a systems approach that generates new knowledge and provides the basis for developing new hypotheses and disease models. Importantly, this approach makes it possible to re-assess assumptions about virus-host cell interactions, such as the prior assumption that VZV preferentially infects memory CD4 T cells to facilitate viral transport to skin.
Specifically, the data we obtained with this method demonstrated that VZV orchestrates a continuum of changes within heterogeneous host T cells, regardless of their basal state, which could not have been determined by traditional averaged measurements. As a further proof of concept underscoring the value of this approach, single cell mass cytometry using multiple markers was particularly important because there is no single cell factor that causes a T cell to exhibit skin homing behavior. A constellation of simultaneous changes, including both up-regulation and down-regulation of several T cell surface proteins, elicits a skin trafficking profile. Using single cell analysis of the surface protein profiles of each individual V+ T cell compared to each UI T cell by SLIDE demonstrated that VZV infection directs the changes observed on T cells, with consequences for cell function and that preferential infection of memory T cells is not the pathogenic mechanism. These experiments showing differential consequences of viral infection depending on baseline cell conditions offer a proof of concept likely to be generalizable to other virus-host cell interactions.
Using single cell mass cytometry for simultaneous measurements of the activation state of multiple phosphoproteins also made it possible to show that VZV infection modulates known T cell pathways that are typically triggered through the T cell receptor (TCR). These experiments demonstrated that this pathway can be altered under conditions of viral infection, separately from its initiation by a receptor binding event. Further, probing expression of cell pathways linked the remodeling of T cell surface characteristics to the concurrent VZV activation of cell signaling proteins. Again, similar insights about manipulation of cell processes can be expected when this method is applied to the investigation of other virus-host cell interactions
Our application of single cell mass cytometry to investigate how VZV perturbs T cell dynamics demonstrated that the method is useful when changes in individual cells are more heterogeneous and subtle. The experiments expand upon previous studies that validated the approach for revealing the hierarchy of immune cells by stimulation with exogenous cytokines that elicit more uniform and robust effects on particular cell proteins. In addition, analytic tools for interpreting mass cytometry data, such as PCA and SPADE, use dimension reduction and clustering methods that do not utilize the richness of the single cell data sets. In the scenario of the more varied and subtle consequences of viral infection, it was important to fully exploit the capacity to detect multi-parametric changes in each individual cell, which was achieved by developing SLIDE. Employing multiple statistical methods and choosing the optimal data analysis tool is an essential component of using this high throughput single cell mass cytometry technique. As indicated by SLIDE and viSNE, further advances in data analytics are likely to occur as applications of the method are expanded. Finally, posting the primary data sets allows later refinements of data interpretation and opportunities for new findings both about the virus-infected cells and more broadly, about the cell biology of the uninfected differentiated host cells, such as tonsil T cells, tested in parallel.
In summary, the significant principle about viral pathogenesis that emerged using single cell mass cytometry as an experimental tool is that infection with a virus such as VZV, which is highly adapted to its host, can dramatically modulate intracellular signaling and change the surface properties of differentiated target cells despite their basal heterogeneity. Since other viral pathogens are likely to have similar capacities, we predict that single cell mass spectrometry will be widely used by virologists to obtain novel insights about virus-induced changes in differentiated primary cells that are necessary for pathogenesis and that cannot be demonstrated by other approaches.
Highlights.
Deep protein profiling of virus infected cells by single cell mass cytometry
Single cell mass cytometry (CyTOF) as a novel technique to study virus-host pathogenesis taking into account the heterogeneity of differentiated host cells
CyTOF analysis of VZV-infected human tonsil T cells using 40+ parameters reveals VZV activates cell signaling pathways and remodels T cell surface proteins regardless of the basal state or T cell subpopulation.
Statistical tools to analyze virus mediated changes using mass cytometry data
Development of single cell linkage density estimation (SLIDE) to quantify T cell remodeling by VZV
Acknowledgments
We thank Sean Bendall, and Prof. Garry Nolan for providing the reagents and facilities for the CyTOF experiments. We thank Adrish Sen for the inputs provided during data analysis and development of SLIDE. This work was supported by NIH AI20459 and AI05346 grants.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Arvin AM, Gilden D. In: Fields Virology. Knipe D, Howley P, editors. Lippincot: Williams & Wilkins; 2013. [Google Scholar]
- 2.Ku CC, Zerboni L, Ito H, Graham BS, Wallace M, Arvin AM. J Exp Med. 2004;200:917–925. doi: 10.1084/jem.20040634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zerboni L, Sen N, Oliver S, Arvin AM. Nat Rev Microbiol. 2014;12:197–210. doi: 10.1038/nrmicro3215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bendall SC, Simonds EF, Qiu P, Amir ED, Krutzik PO, Finck R, Bruggner RV, Melamed R, Trejo A, Ornatsky OI, Balderas RS, Plevritis SK, Sachs K, Pe’er D, Tanner SD, Nolan GP. Science. 2011;332:687–696. doi: 10.1126/science.1198704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sen N, Mukherjee G, Sen A, Bendall SC, Sung P, Nolan GP, Arvin AM. Cell Rep. 2014;8:633–645. doi: 10.1016/j.celrep.2014.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Newell EW, Sigal N, Bendall SC, Nolan GP, Davis MM. Immunity. 2012;36:142–152. doi: 10.1016/j.immuni.2012.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Qiu P, Simonds EF, Bendall SC, Gibbs KD, Jr, Bruggner RV, Linderman MD, Sachs K, Nolan GP, Plevritis SK. Nature Biotechnol. 2011;29:886–891. doi: 10.1038/nbt.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Amir ED, Davis KL, Tadmor MD, Simonds EF, Levine JH, Bendall SC, Shenfeld DK, Krishnaswamy S, Nolan GP, Pe’er D. Nature Biotechnol. 2013;31:545–552. doi: 10.1038/nbt.2594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ku CC, Padilla JA, Grose C, Butcher EC, Arvin AM. J Virol. 2002;76:11425–11433. doi: 10.1128/JVI.76.22.11425-11433.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Reichelt M, Brady J, Arvin AM. J Virol. 2009;83:3904–18. doi: 10.1128/JVI.02137-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fienberg HG, Simonds EF, Fantl WJ, Nolan GP, Bodenmiller B. Cytometry A. 2012;81:467–475. doi: 10.1002/cyto.a.22067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ornatsky OI, Lou X, Nitz M, Schafer S, Sheldrick WS, Baranov VI, Bandura DR, Tanner SD. Anal Chem. 2008;80:2539–2547. doi: 10.1021/ac702128m. [DOI] [PubMed] [Google Scholar]
- 13.Bandura DR, Baranov VI, Ornatsky OI, Antonov A, Kinach R, Lou X, Pavlov S, Vorobiev S, Dick JE, Tanner SD. Anal Chem. 2009;81:6813–6822. doi: 10.1021/ac901049w. [DOI] [PubMed] [Google Scholar]
- 14.Finck R, Simonds EF, Jager A, Krishnaswamy S, Sachs K, Fantl W, Pe’er D, Nolan GP, Bendall SC. Cytometry Part A: J Int Soc Analytical Cytology. 2013;83:483–494. doi: 10.1002/cyto.a.22271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kotecha N, Krutzik PO, Irish JM. Current Protocols in Cytometry. 2010;Chapter 10(Unit10.17) doi: 10.1002/0471142956.cy1017s53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Genser B, Cooper PJ, Yazdanbakhsh M, Barreto ML, Rodrigues LC. BMC Immunol. 2007;8:27. doi: 10.1186/1471-2172-8-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Devroye L, Wagner TJ. Handbook of Statistics. 1982;2:193–197. [Google Scholar]
- 18.Hastie T, Tibshirani R. IEEE Transactions on Pattern Analysis and Machine Intelligence. 1996;18:607–616. [Google Scholar]