Significance
Much research has focused on the molecular basis for protein thermodynamic stability; by comparison, kinetic stability is much less understood. Thermodynamics define the equilibrium fraction of unfolded protein while kinetics define the rate of unfolding; the latter can be of great practical importance for ensuring a protein remains folded under biological conditions. Using extensive experimental and modeling analyses we show that ThreeFoil, a small glycan binding protein without disulfides, exhibits outstanding kinetic stability against chemical denaturation and proteolytic degradation. We demonstrate that high kinetic stability is successfully modeled in terms of extensive long-range intramolecular interactions. These results show that topological complexity is a key determinant of kinetic stability which should help in designing proteins to withstand harsh conditions.
Keywords: SDS/protease resistance, protein folding, coarse-grained simulations, protein topology, contact order
Abstract
The design of stable, functional proteins is difficult. Improved design requires a deeper knowledge of the molecular basis for design outcomes and properties. We previously used a bioinformatics and energy function method to design a symmetric superfold protein composed of repeating structural elements with multivalent carbohydrate-binding function, called ThreeFoil. This and similar methods have produced a notably high yield of stable proteins. Using a battery of experimental and computational analyses we show that despite its small size and lack of disulfide bonds, ThreeFoil has remarkably high kinetic stability and its folding is specifically chaperoned by carbohydrate binding. It is also extremely stable against thermal and chemical denaturation and proteolytic degradation. We demonstrate that the kinetic stability can be predicted and modeled using absolute contact order (ACO) and long-range order (LRO), as well as coarse-grained simulations; the stability arises from a topology that includes many long-range contacts which create a large and highly cooperative energy barrier for unfolding and folding. Extensive data from proteomic screens and other experiments reveal that a high ACO/LRO is a general feature of proteins with strong resistances to denaturation and degradation. These results provide tractable approaches for predicting resistance and designing proteins with sufficient topological complexity and long-range interactions to accommodate destabilizing functional features as well as withstand chemical and proteolytic challenge.
The design of proteins with a desired stable fold and function is a much sought after goal. Although impressive recent successes have been reported in designing both natural and novel protein functions and/or structures (1–6), design remains difficult, often requiring multiple rounds of iterative improvements (7–10). In depth biophysical characterization of protein design outcomes and an understanding of their molecular basis have been limited, and these are critical for improving future designs. Combining designed function with structure is particularly difficult, in part because functional sites tend to be sources of thermodynamic instability (11, 12) and folding frustration (13–15). We investigate how an approach that considers both structure and function from the outset may be used to overcome such obstacles. Furthermore, we demonstrate how kinetic and related stabilities against denaturation can be rationally designed.
A promising emerging paradigm for protein design is the repetition of modular structural elements (1, 2, 5–7, 14, 16–20). This approach can simplify the design process and build on aspects of the evolution of natural repetition in proteins, as well as incorporate the inherent multivalent binding functionality of such structures (1, 21). Internal structural symmetry, resulting from the repetition of smaller elements of structure, is very common in natural proteins, with ∼20% of all protein folds (22) and the majority of the most populated globular protein folds (superfolds) (21) containing internal structural symmetry. Recent design successes, for helical proteins (5, 6), repeat proteins (18, 20, 23) and symmetric superfolds (1, 2, 7, 16, 17, 19, 24–26) recommend the simplification of the design process by using repetitive/symmetric folds as a particularly effective strategy.
The β-trefoil superfold is an interesting test case for design by repetition as bioinformatics analysis has revealed multiple and recent instances of the evolution of distinct proteins with this symmetric fold (1). The fold consists of three repeats, each containing four β-strands, and is adopted by numerous superfamilies with highly diverse binding functions (27). Our design of a completely symmetric β-trefoil, ThreeFoil (Fig. 1), used a hypothetical multivalent carbohydrate binding template and mutated 40 of the 141 residues (1). The mutations were based on a combination of consensus design using a limited set of close homologs (to preserve function), and energy scoring using Rosetta (28). The design was successful on the first attempt, producing a soluble, well folded, and functional monomer with very high resistance to structural fluctuations as indicated by high resistance to thermal denaturation and limited amide H/D exchange (1).
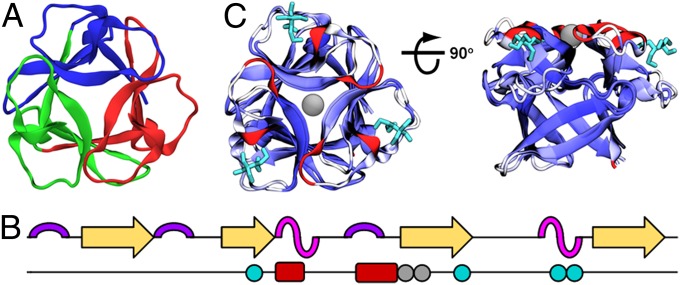
Fig. 1.
Design of ThreeFoil. (A) ThreeFoil (PDB: 3PG0) illustrating its three identical peptide subdomains (red, green, blue). (B) ThreeFoil’s secondary structure: turn (purple), β-strand/bridge (yellow), and 3/10-helix (magenta) and ligand binding residues indicated by colored circles and insertions shown in red. (C) Comparison of ThreeFoil with the independently designed Symfoil (PDB: 3O4D, 15% sequence identity), shown along (Left) and across (Right) the axis of symmetry. Backbones are colored by RMSD between the two structures (blue to white, 0–5 Å), with insertions in the loops of ThreeFoil relative to Symfoil colored red. ThreeFoil’s bound sodium shown in gray, and bis-Tris, which binds in the conserved carbohydrate binding sites, shown in cyan.
Here, we use a battery of biophysical and computational methods to perform an in depth analysis of Threefoil, which shows that it has remarkably slow unfolding and folding kinetics compared with natural and designed proteins due to an unusually high transition state energy barrier. Such kinetic stability against unfolding has been studied little to date. Furthermore, Threefoil is extremely resistant to chemical denaturation and proteolytic degradation. Analyses using Absolute Contact Order (ACO) (29) and Long-Range Order (LRO) (30) as well as Gō model folding simulations (31–33) show that ThreeFoil’s resistance can be explained by the high cooperativity of its folded structure, which includes many long-range interactions. Simulations also show that nonnative interactions or folding frustration arising from protein symmetry (34) do not create long-lived traps during folding or account for the high barrier. They also explain how ligand binding can chaperone folding, which can be an added advantage of designing the fold and function together. Notably, additional analyses using whole proteome screening and other experiments show that proteins with similar resistances as ThreeFoil generally have high ACO/LRO values. Thus, the design method used for ThreeFoil and the strategy of designing folds with many long-range contacts may be useful for designing functional proteins with high resistance to denaturation and degradation, as may be needed for challenging biotechnology applications.
Results
ThreeFoil Has Extremely Slow Kinetics and Substantial Thermodynamic Stability.
To better understand the basis for ThreeFoil’s very high apparent melting temperature (>90 °C) and slow amide exchange (1), we measured its folding kinetics and thermodynamic stability using chemical denaturation. ThreeFoil is extremely resistant to chemical denaturation, remaining folded in high concentrations of urea, with unfolding only observable after very long incubation times in high concentrations of the stronger denaturants guanidinium chloride and guanidinium thiocyanate (GuSCN) (Fig. 2 and SI Appendix, Fig. S1). ThreeFoil’s half-life for unfolding in the absence of denaturant is ∼8 y, although its folding half-life is on the order of 1 h (SI Appendix, Table S1). A comparison against natural and designed proteins of varying structural classes and lengths illustrates how unusually slow these kinetics are (Fig. 3). Despite ThreeFoil’s slow kinetics, unfolding is highly reversible and the rate constants measured by multiple optical probes vary linearly with denaturant concentration, indicating a two-state transition between folded and unfolded states (Fig. 2A and SI Appendix, Fig. S1). The very slow kinetics are indicative of a high free energy (un)folding transition state (Fig. 2B). Similarly, a high transition state barrier underlies the extremely slow unfolding of α-lytic protease (35); however, unlike this prototypical kinetically stable protein, which is thermodynamically unstable, ThreeFoil possesses substantial thermodynamic stability of ∼6 kcal/mol (SI Appendix, Table S1).
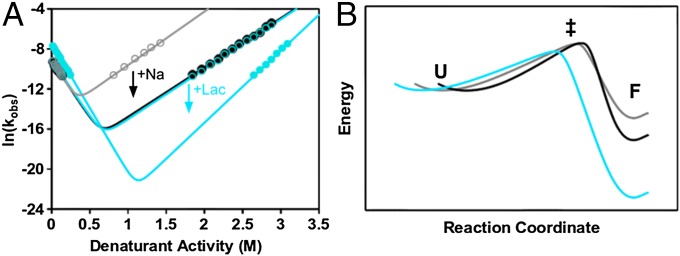
Fig. 2.
Folding and unfolding kinetics of ThreeFoil are modulated by ligand binding. (A) Chevron plots of observed folding and unfolding rate constants (in s−1) in GuSCN were determined by fluorescence. Measurements were without sodium (gray open circles), with sodium (300 mM, black filled circles) or sodium and 50 mM of either lactose (cyan filled circles) or sucrose (cyan open circles). (B) Energy diagram corresponding to the kinetic measurements (coloring as in A). The energy axis is given by -RTln(kobs) and the reaction coordinate follows the change in solvent accessible surface area as measured by mf and mu. The folded (F), transition (‡), and unfolded (U) states are indicated. Unfolded state energies and folded state reaction coordinates are set equal to facilitate comparisons.
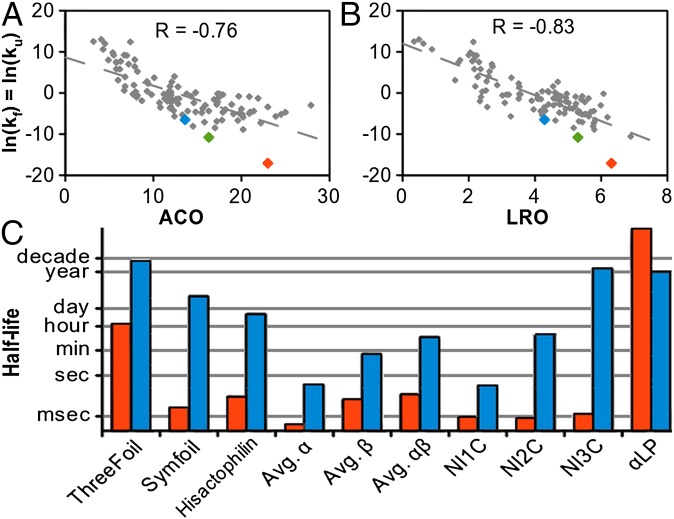
Fig. 3.
ThreeFoil folding/unfolding kinetics are extremely slow compared with other proteins. Rate constants (gray diamonds) at the transition midpoint (ln(kf) = ln(ku)) for a large dataset of proteins (SI Appendix, Table S2) (41), are correlated with ACO (A) and LRO (B). β-trefoil proteins: ThreeFoil (orange), Symfoil (green), and Hisactophilin (blue) are highlighted. (C) The half-lives for folding (orange) and unfolding (blue) are shown for β-trefoils and the averages for the large dataset in each major structural class (α, β, αβ). The prototypical kinetically stable protein α-lytic protease is shown for comparison (35). Ankyrin proteins with 1–3 consensus designed internal repeats (NI1C to NI3C) illustrate the effect of increasing interface area and cooperativity (23).
Various studies have found evidence that repetition in proteins can slow kinetics by creating folding frustration (34, 36). To further examine the role of sequence repetition on kinetics, we compare ThreeFoil with another fully symmetric β-trefoil, Symfoil, which was obtained using iterative rounds of rational design and sequence selection (7). Symfoil has <15% sequence identity to ThreeFoil and, although it has a higher thermodynamic stability of ∼11 kcal/mol, it both folds and unfolds much faster (one million- and 400-fold, respectively, SI Appendix, Table S1 and Fig. 3). Thus, symmetry does not a priori result in kinetic stability. Despite having a nearly identical core secondary structure to Symfoil, ThreeFoil has additional length and interactions for a set of loops involved in its carbohydrate binding function (Fig. 1 B and C). By contrast, heparin binding function, including binding residues in a loop of Symfoil’s template protein, acidic fibroblast growth factor, were eliminated during the many iterations of the design process (7). As ThreeFoil’s longer loops surround and create its ligand binding sites, we investigated the structure of these sites during folding.
Formation of Ligand Binding Sites During Folding.
Measurements of the kinetics of (un)folding in the presence of ligands can be used to monitor the formation of ligand binding sites (37). ThreeFoil has a single binding site for sodium, which is coordinated by three sets of residues distant in sequence, and three carbohydrate binding sites, which have binding residues close in sequence (Fig. 1 B and C) (1). The carbohydrate sites bind lactose and are highly specific for glycans with terminal galactose in a β-1,4 linkage (SI Appendix, Figs. S2A and S3). Sodium decreases the unfolding rate but has no effect on the folding rate, thus it binds only to the folded state and not to the transition state (Fig. 2B). In contrast, lactose not only decreases the unfolding rate but also increases the folding rate, indicating partial formation of the lactose binding sites in the transition state ensemble. The kinetic effects of lactose are specific and not general solvent properties, as no kinetic changes are observed for sucrose (Fig. 2 and SI Appendix, Table S1).
Interestingly, the addition of lactose also increases the denaturant-dependence of stability, m, which reports on the extent of solvent accessible surface burial for a structural transition. The m increases from a value that is 68% of that expected for a protein of this size to 85% (SI Appendix, Table S1). An increase in m may arise from increased burial of hydrophobic residues in the folded protein and/or decreased residual structure in the unfolded protein. Multiple lines of evidence support the latter explanation. Circular dichroism (CD) and NMR (SI Appendix, Figs. S2 and S4, respectively) experiments for folded ThreeFoil show no significant change in native structure upon lactose binding. Also, anilinonapthalenesulfonic acid (ANS), a dye that binds clusters of exposed hydrophobic groups, shows no binding to folded or denatured Threefoil (SI Appendix, Fig. S5). There is no apparent change in CD upon adding lactose to denatured ThreeFoil (SI Appendix, Fig. S2B); however, the CD spectra of denatured ThreeFoil show evidence for nonrandom structure. Similarly, quantitative CD analysis for OneFoil, a peptide consisting of just one of the constituent repeats of ThreeFoil, shows it has approximately half of the β-structure observed in folded ThreeFoil (SI Appendix, Fig. S2C). However, OneFoil shows no evidence for stable structure formation by NMR (1). These experiments strongly suggest the presence of fluctuating residual structure in the ensemble of denatured conformers for ThreeFoil. Together with folding simulations (described below), the results indicate that lactose binding enhances folding not only by binding to the transition state, but also by binding weakly to some conformations in the denatured ensemble and so decreasing nonnative residual structure.
Other studies have also shown that different types of ligands (e.g., metals, heme, nucleotides) can bind to partially folded proteins (denatured, intermediate, and transition states) and so promote folding (38–40). Thus, ligands may not only stabilize the native state, but also promote and chaperone protein folding by binding to other states and thereby smooth the folding energy landscape. In this way, ligand binding can increase the foldability of the protein when structure and function are designed concurrently rather than separately.
Modeling Reveals the Molecular Mechanisms for ThreeFoil’s Ligand Binding and Slow Kinetics.
The ligand binding loops make extensive contacts with distant residues in the primary sequence and so increase the ACO and LRO for ThreeFoil, which are notably high (Fig. 3 A and B). ACO/LRO are measures of topological complexity based on the sequence separation of contacting residues in the folded protein. We have shown recently that the rates of protein folding and unfolding both decrease with increasing ACO/LRO (41). LRO provides a more linear and stronger correlation and is normalized for increasing protein size, which also slows (un)folding (41, 42). As ACO/LRO provide just a simple measure of protein structural complexity, we used Gō models to further define the molecular origins of ThreeFoil’s high barrier.
Gō models encode the structure of the folded protein in their energy functions (31–33) and can be used to understand at higher resolution the effects of protein topological complexity on folding. Molecular dynamics (MD) simulations of such models for diverse proteins can capture trends in barrier heights as well as mechanistic details of folding (32). Here, a coarse-grained Gō model shows that ThreeFoil has a particularly high free energy barrier (Fig. 4A). Also, in the structure of the transition state ensemble (Fig. 4 B and C) residues around the carbohydrate binding site of the second repeat are almost completely folded. Therefore, lactose may bind to and lower the energy of the transition state ensemble and so increase the folding kinetics, whereas unfolding kinetics are slowed owing to even stronger lactose binding to the folded state (Fig. 2). In contrast, the residues in the sodium-binding site are quite unstructured early in the transition state (Fig. 4 B and C) showing that sodium only binds the folded state and therefore slows unfolding with no effect on folding. Thus, the Gō model simulations rationalize ThreeFoil’s slow experimental kinetics and also provide a molecular explanation for the kinetic effects of ligands. In addition, although a simple calculation of ACO/LRO indicates that ThreeFoil should (un)fold slower than Symfoil and Hisactophilin (see below) (Fig. 3 A and B), the more detailed Gō model simulations better capture the variations in these rates (Fig. 4G and SI Appendix, Fig. S6).
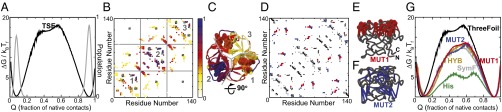
Fig. 4.
Structure-based simulations reveal the molecular origins of ThreeFoil’s large kinetic barrier. (A) The folding free energy of ThreeFoil in units of kBTf (left y axis) is plotted at the transition midpoint (Tf) as a function of the fraction of native contacts (Q) in black. The population distribution is plotted in gray (right y axis). The protein populates only the unfolded state (Q ∼ 0.1) and the folded state (Q ∼ 0.9). The two curves were calculated from simulations of a ThreeFoil model using all contacts shown in D. (B) Contact map of the transition state ensemble (TSE, Q ∼ 0.35 in A), colored based on degree of structure, with 1 indicating native levels of structure and 0 indicating no structure. Contacts between lactose binding site residues (cyan) and sodium binding residues (gray) are shown. The numbered squares contain intratrefoil contacts (see C). (C) Average level of structure derived from B (same coloring) illustrated for ThreeFoil partitioned into its three repeats by gray lines. The residues shown as spheres are part of the 3 symmetric lactose-binding sites, whereas those shown as sticks are part of the single sodium-binding site (Fig. 1). The rotation indicated gives the view in E and F. (D) Contact map of ThreeFoil, with contacts deleted in MUT1 and MUT2 shown in red and blue, respectively. All deleted contacts are long-range (far from diagonal). (E) Long-range contacts (red sticks) of the β2–β3 loop residues (red spheres at Cα positions) deleted in MUT1. (F) For MUT2 the same number of contacts were deleted such that MUT1 and MUT2 have a very similar ACO. However, these contacts (blue sticks) are spread over the entire protein. (G) Folding free energies of ThreeFoil (black, same as in A), Symfoil (SymF; gray), a hybrid protein with the ThreeFoil contact map projected on the Symfoil backbone (HYB; gold), the two ThreeFoil mutants (MUT1; red, MUT2; blue), and Hisactophilin (His; green) are plotted in units of their respective folding temperatures (kBTf, y axis) at their respective transition midpoints as a function of the fraction of their respective native contacts (x axis). The SymF, HYB, MUT1 and MUT2 free energy profiles have very similar barrier heights, in between those of the highly kinetically stable ThreeFoil and the much less stable His.
The largest differences between Threefoil and Symfoil are in the β2–β3 loops, at the edge of ThreeFoil’s carbohydrate binding sites (Fig. 1 B and C). Consequently, the differences in the contact maps of the two also occur mostly in the contacts of the β2–β3 loops, with Symfoil having shorter loops and fewer contacts in this region. To understand whether the high barrier for ThreeFoil is caused by differences in the length, conformation and packing of the β2–β3 loops or by differences in packing for the rest of the protein a hybrid construct (HYB) was created which has the Symfoil backbone with the ThreeFoil contact map; this construct has almost the same barrier as Symfoil (SymF) (Fig. 4G). Thus, the differences responsible for the higher barrier are the β2–β3 loops. To further define how the conformation and packing of the β2–β3 loops increases the barrier, a mutant of ThreeFoil (MUT1) was created with all long-range interactions of the β2–β3 loops deleted (Fig. 4 D and E). The mutation lowered the barrier height of MUT1 leaving it similar to that of HYB and SymF (Fig. 4G). These results show that the packing of the β2–β3 loops of a given repeat with parts of the other repeats create long-range contacts that markedly increase the barrier height. To test the effect of other long-range contacts (which reduce the overall ACO by an equivalent amount), a control mutant (MUT2) was created where the same number of other contacts with similar sequence separations were deleted (Fig. 4 D and F). The free energy barrier of MUT2 is similar to that of both MUT1 and HYB. Thus, the kinetic stability of a protein can be reduced by either a large loss in packing density localized in the structure (as in MUT1) or by an additive effect from many losses across the structure (as in MUT2). To further confirm the correlation between ACO/LRO, barrier heights, and kinetic stability, we also simulated Hisactophilin, a natural β-trefoil with a low barrier (SI Appendix, Table S1). As expected, the low ACO/LRO Hisactophilin has a much lower folding free energy barrier (Fig. 4G; green profile) than that of either ThreeFoil or SymFoil and has the lowest kinetic stability (Fig. 3). Distinct functional features for Hisactophilin contribute to its low ACO/LRO and barrier (43).
In principle, the internal symmetry of ThreeFoil might also contribute to slow folding by creating misfolded intermediates arising from internal subdomain swapping, analogous to domain swapping observed or inferred for proteins containing longer stretches of repeated sequence (34, 36). Such trapping was tested as a cause of ThreeFoil’s slow kinetics using simulations performed with the addition of symmetric nonnative contacts between the repeats. The results indicate that, close to the transition midpoint, nonnative interactions arising from symmetry do not create significant trapping (SI Appendix, Fig. S7). Thus, compared with the longer proteins the shorter repeat length of ThreeFoil aided by local structure formation, likely limits slowing of folding due to nonnative interrepeat interactions.
MD simulations were also used to investigate the effect of nonnative residual structure (in the unfolded ensemble) on ThreeFoil folding. We performed simulations where the local structure of all three repeats was biased to both the native ThreeFoil structure (as above) and to the most common OneFoil structure obtained in Rosetta ab initio simulations. The tertiary contacts between the repeats were calculated only from the native ThreeFoil structure. The ThreeFoil tertiary contacts appear to suppress the intra-OneFoil nonnative interactions and these nonnative interactions do not create significant trapping (SI Appendix, Fig. S8). Lastly, simulations of just OneFoil (including both native and nonnative structural biases) confirm that the presence of ligand, modeled as a strengthening of intrabinding-residue contacts, greatly suppresses the formation of nonnative structure (SI Appendix, Fig. S8 A and B). Overall, the simulations explain the effects of ligands during folding while also revealing that the extreme kinetic stability of ThreeFoil arises from its native topology and is unlikely to be caused by nonnative traps on the folding free energy landscape.
High Chemical and Protease Resistances of ThreeFoil and Other High ACO/LRO Proteins.
Extremely slow unfolding has been associated with the capacity to maintain native form and function under harsh conditions (44), such as high concentrations of protease (35, 45, 46) and detergent (46, 47). Protease resistance of a classic extremely kinetically stable protein, α-lytic protease, has been proposed to originate from its large and highly cooperative unfolding energy barrier resulting in a rigid native conformation with limited local openings and consequently limited proteolytically susceptible regions (35). Also, challenge by a high concentration of SDS has been used extensively for direct evaluation of protein kinetic stability based on the ability of SDS to induce denaturation by trapping hydrophobic residues exposed during even transient unfolding events (47, 48). Given its high barrier to unfolding, we tested ThreeFoil for resistance to protease and SDS.
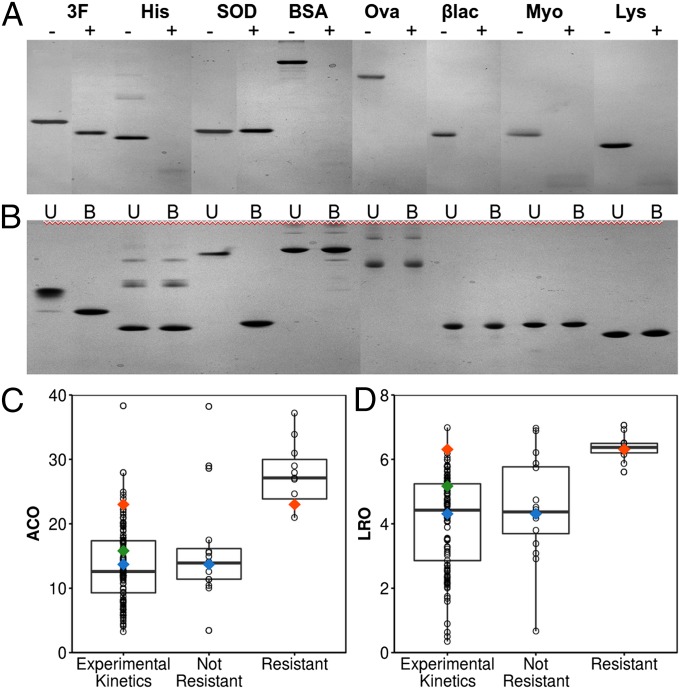
In the manner of Manning and Colón (46) in their profiling of protein kinetic stability, we incubated ThreeFoil with the aggressive and nonspecific protease, proteinase K. ThreeFoil demonstrated strong resistance, remaining intact for the full 4-d challenge by a high concentration of protease (Fig. 5A). A highly protease-resistant control protein, the dimeric human Cu,Zn superoxide dismutase (SOD) also remained intact, whereas histactophilin, which has greater thermodynamic stability but much faster unfolding kinetics than ThreeFoil (SI Appendix, Table S1), was completely degraded within an hour as were other commonly studied proteins (Fig. 5A). The results for the SDS challenge follow the same pattern with only ThreeFoil and SOD being resistant (Fig. 5B), although SOD depends on an intact disulfide bond for SDS resistance whereas ThreeFoil does not (SI Appendix, Fig. S9F).
Fig. 5.
ThreeFoil is highly resistant to protease and detergent. (A) Incubation with Proteinase K of: ThreeFoil (3F), Hisactophilin (His), human Cu,Zn superoxide dismutase (SOD), BSA, ovalbumin (Ova), β-lactoglobulin (βlac), myoglobin (Myo), and lysozyme (Lys). Protein before (−) and after incubation with protease (+) are shown. Retention of the protein band after incubation shows resistance to digestion. ThreeFoil and SOD are shown after 4 d (still nondegraded), whereas others are shown after 1 h (fully degraded). The molecular weight decrease for ThreeFoil after incubation is due to the loss of its unstructured his-tag (untagged ThreeFoil has a MW of 15.3 kDa and runs higher than intact Hisactophilin with a MW of 13.3 kDa, see also SI Appendix, Fig. S9A). Individual gels shown in SI Appendix, Fig. S9 B–E. (B) The same proteins tested for resistance to SDS. Where the unboiled (U) and boiled (B) samples are indistinguishable, no SDS resistance is observed, whereas a higher running unboiled sample indicates SDS is unable to penetrate and bind without thermal unfolding of the protein. Comparison of topological complexity as measured by ACO (C) and LRO (D) for proteins that have been kinetically characterized experimentally (SI Appendix, Table S2) and those with experimentally demonstrated resistance or nonresistance to protease and SDS (SI Appendix, Table S3). Resistant proteins generally have higher topological complexity. β-trefoil proteins are colored as in Fig. 3. Data shown as box-and-whisker plots, with a horizontal line at the median, box enclosing middle 50% of the data, whiskers drawn to 1.5*IQR (interquartile range).
Given the correlations between high topological complexity and slow unfolding (Fig. 3 A and B) (41) and between slow unfolding and SDS/protease resistance (44, 46), we asked whether these resistances could be predicted from topological complexity. We conducted experiments and surveyed the literature to identify proteins with experimentally demonstrated resistance, or lack thereof, to SDS or protease. The identified proteins include new (SI Appendix, Fig. S10) and previously reported (45, 47) results from whole proteome screening to identify kinetically stable proteins, as well as new (Fig. 5 A and B and SI Appendix, Fig. S9) and previously reported analyses of individual proteins (SI Appendix, Table S3). The results (Fig. 5 C and D) clearly show that resistant proteins have notably high ACO/LRO values, similar to ThreeFoil, whereas the nonresistant proteins tend to have much lower values. The few nonresistant proteins with a high ACO/LRO indicate that high topological complexity is necessary but not always sufficient for resistance. The preceding observations suggest the rough measure provided by ACO/LRO does not capture other requirements such as highly cooperative unfolding, needed to eliminate weak points in the structure, which provide opportunities for attack by chemical denaturants and proteases (35, 44, 46). Thus, a high ACO/LRO indicates potentially high resistance to degradation/denaturation but a more detailed simulation, as performed for ThreeFoil (Fig. 4), is needed for a more accurate prediction and understanding of molecular determinants for resistance.
Finally, we note that the distribution of ACO/LRO values for a large dataset of proteins with previously characterized kinetics, similar to the nonresistant cases, is markedly lower than for the resistant proteins (Fig. 5 C and D). Thus, although kinetically stable resistant proteins exist, they have received relatively little attention and using their folds or incorporating analogous long-range contacts provides attractive new avenues for designing resistance.
Discussion
.An in depth analysis of the folding characteristics of designed proteins, as we have performed for the threefold symmetric ThreeFoil, is rarely reported, yet is critical for ultimately understanding design outcomes and improving their reliability. We demonstrate a high level of design success for ThreeFoil as evidenced by its: (i) reversible, cooperative, two-state (un)folding; and (ii) well folded and functional native structure which has high solubility and monodispersity, well diffracting crystals, and great resistance against H/D exchange (1), denaturation by chaotropes and detergent, and degradation by protease.
Although the rational design of proteins with desired structure and function remains a great challenge and often require multiple cycles of design and/or selection to improve them, successes in designing both structures and/or functions, including ones not observed in nature, have been increasing (3, 4, 6, 8, 9, 18, 49, 50). These results demonstrate the increasing understanding of fundamental principles and utility of computational protein design. Recently, there have been multiple reports of success for common folds based on repeated structural elements, including relatively high success rates and stabilities for various helix-containing elongated repeat proteins (18, 51) and toroidal or globular superfolds (1–3, 7, 16, 17, 19). The great diversity of sequences observed for these symmetric protein structures may reflect an inherent capacity for stability, foldability and functionality that is especially amenable to both evolution and design (22).
Design strategies similar to that used to make ThreeFoil, which use repetition of structural elements designed using existing sequence information and structural modeling with the Rosetta energy function (28), have proven particularly fruitful, with several studies yielding well-folded proteins with high melting points on the first attempt (1, 16–18). Furthermore, we have shown that ThreeFoil possesses stability, cooperativity and multivalent binding function. These features may be “inherited” through the use of existing sequence information, generating a more naturally funneled energy landscape. Other proteins designed in a similar way, and not yet characterized in detail, may also capture favorable natural features (16–18). Also, our results show how ligand binding can further smooth the landscape by decreasing the formation of nonnative structure and so promote folding and design success.
Although evolution has provided a great range of sequences and structures that may be leveraged, it has also set limitations, which need not constrain rational protein design. As an example, natural proteins for which kinetics have been measured typically unfold on a timescale of seconds-hours (41); ready unfolding may be needed to facilitate protein transport, regulation or turnover. However, other natural proteins that must withstand harsh extracellular or thermophilic conditions tend to have high kinetic stability (35, 44); hence, fast unfolding is not an inherent constraint on proteins. Artificial proteins can be freed from various biological constraints allowing for uncommon properties such as extreme kinetic stability using suitable natural structures or novel ones with the requisite features.
It is important to note that although the energy landscape of a protein defines both its thermodynamic and kinetic stabilities, the two properties are distinct. Thermodynamic stability depends on the energy difference between folded and unfolded states, whereas kinetic stability depends on the energy barrier between the folded and transition states (Fig. 2). High kinetic stability is a particularly attractive feature for rational design, as it is linked to other benefits such as resistances against protein denaturation and degradation by detergents (by decreasing exposure of the hydrophobic core), proteases (by limiting accessibility of cut sites), and temperature (by producing a high energy transition state barrier that is unlikely to be crossed by thermal fluctuations) (44, 46). Such characteristics are highly desirable for industrial or biomedical applications that require a protein to remain folded and functional for a long time, even in challenging environments. Although it is known that kinetic stability and its associated resistances are the result of slow global unfolding and limited local opening (35, 44–46), little has been reported on how to rationally incorporate this into designed proteins. Our in depth experimental and modeling analyses of ThreeFoil provide valuable insight into the molecular basis for these characteristics. Specifically, the origin of ThreeFoil’s very slow global and limited local unfolding is a high and steep energy barrier which is a consequence of a folded topology that includes a large number and proportion of long-range and extensively distributed contacts. Thus, there are no weak points in the structure and it undergoes very cooperative folding to a native state that is highly resistant to local openings.
In summary, a simple calculation of ACO/LRO indicates whether a design has the capacity to be kinetically stable, whereas Gō model simulations give a more accurate prediction and can be used to determine the impact of specific contacts. This paves the way for rational design of resistance to harsh conditions. The mechanistic understanding of the structural determinants of resistance and the ability to design it, as well as the simplified and efficient design process of using structural repetition within the context of a symmetric and functional superfold, provide valuable avenues for improving future protein designs.
Materials and Methods
Protein Expression and Purification.
ThreeFoil was expressed in E. coli and inclusion bodies solubilized in urea before being purified on a Ni-NTA column and refolded by dialysis, as described (1). Details for removal of sodium are given in the SI Appendix, SI Materials and Methods.
Kinetic Measurements.
Measurements were performed at 27 °C and monitored by fluorescence (excitation 274 nm, emission 317 nm) using a SpectraMax M5 plate reader (Molecular Devices). Protein was diluted into varying concentrations of GuSCN by manual mixing and measured for up to 4 d. Additional details are given in the SI Appendix, SI Materials and Methods.
SDS Resistance.
Protein in H2O was diluted into SDS and Tris so that final samples contained 0.5 mg/mL protein and 1% SDS in 125 mM Tris (pH 6.8). Samples were then either boiled or incubated at room temperature for 10 min before analysis by SDS/PAGE using 15% (wt/vol) Acrylamide gels with 0.1% SDS in Tris/glycine running buffer (pH 8.3), and either without (Fig. 5B) or with (SI Appendix, Fig. S9 F and G) 7% (vol/vol) β-mercaptoethanol to reduce disulfides.
Protease Resistance.
Samples contained 0.5 mg/mL of protein in 25 mM Tris and 1 µM EDTA (pH 8.3). A time 0, control was taken before adding proteinase K (final concentration 0.02 mg/mL), and further samples taken after 1 h, 1 d, and 4 d of incubation at 25 °C. The reaction was stopped by mixing samples 1:1 with buffer [2.5 µM phenylmethylsulfonyl fluoride, 125 mM Tris, 4% SDS (wt/vol), 20% (vol/vol) glycerol, 15% (vol/vol) β-mercaptoethanol, at pH 6.8] and boiling for 10 min. SDS/PAGE was performed using the same gel conditions as for SDS Resistance.
Coarse-Grained Gō Models.
A common form of the Gō model (31, 32) was used to perform MD simulations. The inputs to this model are the coordinates of the Cα atoms of the protein and the contact map. Details of the model, contact map calculations, simulation conditions and analyses are given in SI Appendix, SI Materials and Methods. The simulations were performed using a previously developed enhanced sampling technique (52) based on the multicanonical method.
Supplementary Material
Acknowledgments
We thank Prof. Jayant B. Udgaonkar for monellin and SH3; Dr. Ranabir Das for ubiquitin; and Core H of the Consortium for Functional Glycomics, funded by the National Institute of General Medical Sciences (GM62116), for glycan array analysis. This work was supported by National Sciences and Engineering Research Council of Canada (to E.M.M.), Government of India-DAE and DST-Ramanujan Fellowship (SR/S2/RJN-63/2009, 5 years, wef 15/04/2010; to S.G.), and National Science Foundation Grant 1158375 (to W.C.).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1510748112/-/DCSupplemental.
References
- 1.Broom A, et al. Modular evolution and the origins of symmetry: Reconstruction of a three-fold symmetric globular protein. Structure. 2012;20(1):161–171. doi: 10.1016/j.str.2011.10.021. [DOI] [PubMed] [Google Scholar]
- 2.Longo LM, Kumru OS, Middaugh CR, Blaber M. Evolution and design of protein structure by folding nucleus symmetric expansion. Structure. 2014;22(10):1377–1384. doi: 10.1016/j.str.2014.08.008. [DOI] [PubMed] [Google Scholar]
- 3.Koga N, et al. Principles for designing ideal protein structures. Nature. 2012;491(7423):222–227. doi: 10.1038/nature11600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rajagopalan S, et al. Design of activated serine-containing catalytic triads with atomic-level accuracy. Nat Chem Biol. 2014;10(5):386–391. doi: 10.1038/nchembio.1498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Thomson AR, et al. Computational design of water-soluble α-helical barrels. Science. 2014;346(6208):485–488. doi: 10.1126/science.1257452. [DOI] [PubMed] [Google Scholar]
- 6.Huang P-S, et al. High thermodynamic stability of parametrically designed helical bundles. Science. 2014;346(6208):481–485. doi: 10.1126/science.1257481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lee J, Blaber SI, Dubey VK, Blaber M. A polypeptide “building block” for the β-trefoil fold identified by “top-down symmetric deconstruction”. J Mol Biol. 2011;407(5):744–763. doi: 10.1016/j.jmb.2011.02.002. [DOI] [PubMed] [Google Scholar]
- 8.Privett HK, et al. Iterative approach to computational enzyme design. Proc Natl Acad Sci USA. 2012;109(10):3790–3795. doi: 10.1073/pnas.1118082108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li Z, Yang Y, Zhan J, Dai L, Zhou Y. Energy functions in de novo protein design: Current challenges and future prospects. Annu Rev Biophys. 2013;42:315–335. doi: 10.1146/annurev-biophys-083012-130315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Korendovych IV, DeGrado WF. Catalytic efficiency of designed catalytic proteins. Curr Opin Struct Biol. 2014;27:113–121. doi: 10.1016/j.sbi.2014.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Meiering EM, Serrano L, Fersht AR. Effect of active site residues in barnase on activity and stability. J Mol Biol. 1992;225(3):585–589. doi: 10.1016/0022-2836(92)90387-y. [DOI] [PubMed] [Google Scholar]
- 12.Shoichet BK, Baase WA, Kuroki R, Matthews BW. A relationship between protein stability and protein function. Proc Natl Acad Sci USA. 1995;92(2):452–456. doi: 10.1073/pnas.92.2.452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Capraro DT, Roy M, Onuchic JN, Gosavi S, Jennings PA. β-Bulge triggers route-switching on the functional landscape of interleukin-1β. Proc Natl Acad Sci USA. 2012;109(5):1490–1493. doi: 10.1073/pnas.1114430109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ferreiro DU, Komives EA, Wolynes PG. Frustration in biomolecules. Q Rev Biophys. 2014;47(4):285–363. doi: 10.1017/S0033583514000092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gershenson A, Gierasch LM, Pastore A, Radford SE. Energy landscapes of functional proteins are inherently risky. Nat Chem Biol. 2014;10(11):884–891. doi: 10.1038/nchembio.1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fortenberry C, et al. Exploring symmetry as an avenue to the computational design of large protein domains. J Am Chem Soc. 2011;133(45):18026–18029. doi: 10.1021/ja2051217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Voet ARD, et al. Computational design of a self-assembling symmetrical β-propeller protein. Proc Natl Acad Sci USA. 2014;111(42):15102–15107. doi: 10.1073/pnas.1412768111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Parmeggiani F, et al. A general computational approach for repeat protein design. J Mol Biol. 2015;427(2):563–575. doi: 10.1016/j.jmb.2014.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Höcker B. Design of proteins from smaller fragments-learning from evolution. Curr Opin Struct Biol. 2014;27:56–62. doi: 10.1016/j.sbi.2014.04.007. [DOI] [PubMed] [Google Scholar]
- 20.Javadi Y, Itzhaki LS. Tandem-repeat proteins: Regularity plus modularity equals design-ability. Curr Opin Struct Biol. 2013;23(4):622–631. doi: 10.1016/j.sbi.2013.06.011. [DOI] [PubMed] [Google Scholar]
- 21.Orengo CA, Jones DT, Thornton JM. Protein superfamilies and domain superfolds. Nature. 1994;372(6507):631–634. doi: 10.1038/372631a0. [DOI] [PubMed] [Google Scholar]
- 22.Balaji S. Internal symmetry in protein structures: Prevalence, functional relevance and evolution. Curr Opin Struct Biol. 2015;32:156–166. doi: 10.1016/j.sbi.2015.05.004. [DOI] [PubMed] [Google Scholar]
- 23.Wetzel SK, Settanni G, Kenig M, Binz HK, Plückthun A. Folding and unfolding mechanism of highly stable full-consensus ankyrin repeat proteins. J Mol Biol. 2008;376(1):241–257. doi: 10.1016/j.jmb.2007.11.046. [DOI] [PubMed] [Google Scholar]
- 24.Yadid I, Tawfik DS. Functional β-propeller lectins by tandem duplications of repetitive units. Protein Eng Des Sel. 2011;24(1-2):185–195. doi: 10.1093/protein/gzq053. [DOI] [PubMed] [Google Scholar]
- 25.Carstensen L, et al. Conservation of the folding mechanism between designed primordial (βα)8-barrel proteins and their modern descendant. J Am Chem Soc. 2012;134(30):12786–12791. doi: 10.1021/ja304951v. [DOI] [PubMed] [Google Scholar]
- 26.Höcker B, Claren J, Sterner R. Mimicking enzyme evolution by generating new (betaalpha)8-barrels from (betaalpha)4-half-barrels. Proc Natl Acad Sci USA. 2004;101(47):16448–16453. doi: 10.1073/pnas.0405832101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ponting CP, Russell RB. Identification of distant homologues of fibroblast growth factors suggests a common ancestor for all beta-trefoil proteins. J Mol Biol. 2000;302(5):1041–1047. doi: 10.1006/jmbi.2000.4087. [DOI] [PubMed] [Google Scholar]
- 28.Leaver-Fay A, et al. ROSETTA3: An object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 2011;487:545–574. doi: 10.1016/B978-0-12-381270-4.00019-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ivankov DN, et al. Contact order revisited: Influence of protein size on the folding rate. Protein Sci. 2003;12:2057–2062. doi: 10.1110/ps.0302503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gromiha MM, Selvaraj S. Comparison between long-range interactions and contact order in determining the folding rate of two-state proteins: Application of long-range order to folding rate prediction. J Mol Biol. 2001;310(1):27–32. doi: 10.1006/jmbi.2001.4775. [DOI] [PubMed] [Google Scholar]
- 31.Nymeyer H, García AE, Onuchic JN. Folding funnels and frustration in off-lattice minimalist protein landscapes. Proc Natl Acad Sci USA. 1998;95(11):5921–5928. doi: 10.1073/pnas.95.11.5921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chavez LL, Onuchic JN, Clementi C. Quantifying the roughness on the free energy landscape: Entropic bottlenecks and protein folding rates. J Am Chem Soc. 2004;126(27):8426–8432. doi: 10.1021/ja049510+. [DOI] [PubMed] [Google Scholar]
- 33.Hyeon C, Thirumalai D. Capturing the essence of folding and functions of biomolecules using coarse-grained models. Nat Commun. 2011;2:487. doi: 10.1038/ncomms1481. [DOI] [PubMed] [Google Scholar]
- 34.Borgia MB, et al. Single-molecule fluorescence reveals sequence-specific misfolding in multidomain proteins. Nature. 2011;474(7353):662–665. doi: 10.1038/nature10099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jaswal SS, Sohl JL, Davis JH, Agard DA. Energetic landscape of alpha-lytic protease optimizes longevity through kinetic stability. Nature. 2002;415(6869):343–346. doi: 10.1038/415343a. [DOI] [PubMed] [Google Scholar]
- 36.Javadi Y, Main ERG. Exploring the folding energy landscape of a series of designed consensus tetratricopeptide repeat proteins. Proc Natl Acad Sci USA. 2009;106(41):17383–17388. doi: 10.1073/pnas.0907455106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Sancho J, Meiering EM, Fersht AR. Mapping transition states of protein unfolding by protein engineering of ligand-binding sites. J Mol Biol. 1991;221(3):1007–1014. doi: 10.1016/0022-2836(91)80188-z. [DOI] [PubMed] [Google Scholar]
- 38.Wittung-Stafshede P. Role of cofactors in protein folding. Acc Chem Res. 2002;35(4):201–208. doi: 10.1021/ar010106e. [DOI] [PubMed] [Google Scholar]
- 39.Stigler J, Rief M. Calcium-dependent folding of single calmodulin molecules. Proc Natl Acad Sci USA. 2012;109(44):17814–17819. doi: 10.1073/pnas.1201801109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Liu P-F, Park C. Selective stabilization of a partially unfolded protein by a metabolite. J Mol Biol. 2012;422(3):403–413. doi: 10.1016/j.jmb.2012.05.044. [DOI] [PubMed] [Google Scholar]
- 41.Broom A, Gosavi S, Meiering EM. Protein unfolding rates correlate as strongly as folding rates with native structure. Protein Sci. 2015;24:580–587. doi: 10.1002/pro.2606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Thirumalai D. From minimal models to real proteins - time scales for protein folding kinetics. J Phys I. 1995;5:1457–1467. [Google Scholar]
- 43.Gosavi S. Understanding the folding-function tradeoff in proteins. PLoS One. 2013;8(4):e61222. doi: 10.1371/journal.pone.0061222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sanchez-Ruiz JM. Protein kinetic stability. Biophys Chem. 2010;148(1-3):1–15. doi: 10.1016/j.bpc.2010.02.004. [DOI] [PubMed] [Google Scholar]
- 45.Park C, Zhou S, Gilmore J, Marqusee S. Energetics-based protein profiling on a proteomic scale: Identification of proteins resistant to proteolysis. J Mol Biol. 2007;368(5):1426–1437. doi: 10.1016/j.jmb.2007.02.091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Manning M, Colón W. Structural basis of protein kinetic stability: Resistance to sodium dodecyl sulfate suggests a central role for rigidity and a bias toward beta-sheet structure. Biochemistry. 2004;43(35):11248–11254. doi: 10.1021/bi0491898. [DOI] [PubMed] [Google Scholar]
- 47.Xia K, et al. Identifying the subproteome of kinetically stable proteins via diagonal 2D SDS/PAGE. Proc Natl Acad Sci USA. 2007;104(44):17329–17334. doi: 10.1073/pnas.0705417104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Xia K, et al. Quantifying the kinetic stability of hyperstable proteins via time-dependent SDS trapping. Biochemistry. 2012;51(1):100–107. doi: 10.1021/bi201362z. [DOI] [PubMed] [Google Scholar]
- 49.Watters AL, et al. The highly cooperative folding of small naturally occurring proteins is likely the result of natural selection. Cell. 2007;128(3):613–624. doi: 10.1016/j.cell.2006.12.042. [DOI] [PubMed] [Google Scholar]
- 50.Best RB, Hummer G, Eaton WA. Native contacts determine protein folding mechanisms in atomistic simulations. Proc Natl Acad Sci USA. 2013;110(44):17874–17879. doi: 10.1073/pnas.1311599110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Boersma YL, Plückthun A. DARPins and other repeat protein scaffolds: Advances in engineering and applications. Curr Opin Biotechnol. 2011;22(6):849–857. doi: 10.1016/j.copbio.2011.06.004. [DOI] [PubMed] [Google Scholar]
- 52.Gosavi S, Chavez LL, Jennings PA, Onuchic JN. Topological frustration and the folding of interleukin-1 beta. J Mol Biol. 2006;357(3):986–996. doi: 10.1016/j.jmb.2005.11.074. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.