Abstract
Triggering receptor expressed on myeloid cells-1 (TREM-1) is critical for inflammatory signal amplification. Humans have two forms of TREM-1: a membrane receptor (mbTREM-1), associated with the adaptor DAP12, and a soluble receptor detected at times of infection. The membrane receptor isoform acts synergistically with the TLR pathway to promote cytokine secretion and neutrophil migration while the soluble receptor functions as a counter regulatory molecule. In multiple models of sepsis, exogenous administration of soluble forms of TREM-1 attenuates inflammation and markedly improves survival. Despite intense interest in soluble TREM-1 both as a clinical predictor of survival and as a therapeutic tool, the origin of native soluble TREM-1 remains controversial. Utilizing human neutrophils, we identified a 15 kDa TREM-1 isoform in primary (azurophilic) and secondary (specific) granules. Mass spectrometric analysis, ELISA, and immunoblot confirm that the 15 kD protein is a novel splice variant of TREM-1 (TREM-1sv). Neutrophil stimulation with P. aeruginosa, LPS, or PAM(3)Cys4 resulted in degranulation and release of TREM-1sv. The addition of exogenous TREM-1sv inhibited TREM-1 receptor mediated proinflammatory cytokine production. Thus these data reveal that TREM-1 isoforms simultaneously activate and inhibit inflammation via the canonical membrane TREM-1 molecule and this newly discovered granular isoform, TREM-1sv.
Introduction
Neutrophils, the most abundant circulating leukocyte, play a pivotal role in the response to pathogen invasion (1, 2). A broad array of antimicrobial agents is stored within distinct intracellular granules to eliminate pathogens. Moreover these granules contain chemokines, cytokines, and immunoreceptors that shape the subsequent inflammatory response to many danger signals(3). Neutrophil granules are formed during cell maturation in the bone marrow and each subset contains a characteristic protein content that define the subset. Mobilization of these subsets is achieved with different stimuli and thus their contents are deployed in a regulated fashion facilitating neutrophil function as the cell moves from the blood, through the extracellular matrix, across epithelial barriers and into a variety of tissues in response to danger signals(4, 5).
To detect pathogens, neutrophils express several large genetically-encoded receptor families. These families include the toll-like receptor (TLR) family, C-type lectin receptors, and the Triggering Receptor Expressed on Myeloid (TREM) family (6-8). Our data and others have shown that TREM-1, expressed in monocytes and neutrophils, is critical for inflammatory signal amplification (9-12). More recently, studies with TREM-1-deficient mice demonstrate that this molecule is also required for neutrophil migration into the lung airspace and that abrogation of TREM-1-mediated inflammation is protective in a murine liver cancer model (13, 14). As such, TREM-1 has been implicated in a variety of diseases, including sepsis, ventilator-associated pneumonia, cancer development, cardiovascular disease, and autoimmunity (15-20).
TREM-1 is present in humans in two forms: as a membrane receptor (mbTREM-1), associated with the adaptor DAP12, and as a soluble receptor detected in plasma, gastric secretions, bronchioalveolar lavage (BAL) fluid, and urine at times of infection and inflammation (11, 15, 21, 22). The membrane form of this receptor synergizes with TLRs to amplify pro-inflammatory cytokine production in both neutrophils and monocytes (7, 9, 23). In contrast, soluble forms of TREM-1 act as counter regulatory molecules, attenuating inflammation and improving outcomes in animal models of sepsis, acute kidney injury, and inflammatory bowel disease (24-26).
Despite intense interest in soluble TREM-1 both as a predictor of outcomes in pneumonia and sepsis and as a therapeutic tool, the origin of soluble TREM-1 remains controversial. In vitro studies have suggested that the extracellular domain of this receptor is proteolytically processed by matrix metalloproteinases, resulting in soluble TREM-1 release (27). Other data report that de novo protein synthesis following stimulation is required for soluble TREM-1 release (28). Finally, a splice variant mRNA transcript (TREM-1sv) lacking the sequence that encoded the transmembrane domain was reported in adult and fetal tissues as well as peripheral blood-derived CD14 (+) monocytes, though protein expression was not assessed (29). Thus, whether soluble TREM-1 in clinical samples is generated by proteolytic cleavage of membrane expressed TREM-1 and/or if soluble TREM-1 is generated from a splice variant transcript is unclear in the literature. Moreover, whether a soluble form of TREM-1 is stored for immediate release at the time neutrophil activation is unknown.
To address these questions, we isolated human neutrophil granules and characterized their TREM-1 isoforms. This analysis resulted in the surprising identification of a 15kD form of TREM-1 in primary and secondary granules. Mass spectroscopy, immunoblot and ELISA together confirmed that the 15 kDa protein was a splice variant form of TREM-1 (TREM-1sv). RNA profiling identified the presence of this TREM-1sv transcript in human neutrophils. Finally, we demonstrated that TREM-1sv is released upon degranulation from human neutrophils following physiologically relevant stimulation, suggesting that this unique splice variant form of TREM-1 serves as a source of soluble TREM-1 during inflammation.
Methods
Isolation of human neutrophils and cell fractionation
Institutional Review Board at the University of Iowa reviewed and approved human studies involving healthy donors (200911748) and exempted the studies involving blood from critically ill patients. Patients were screened for presence or absence of sepsis in the intensive care units at the University of Iowa (30). Venous blood was drawn into tubes coated with heparin and neutrophils were isolated within 3 hours of collection(31). Neutrophil subcellular fractions were obtained by discontinuous 2-layer Percoll density gradient centrifugation with a slight modification (32). Cells were incubated with 1 mM Diisopropylfluorophosphate (Sigma-Aldrich #D0879), an irreversible serine protease inhibitor, for 20 minutes on ice prior to cell disruption by nitrogen cavitation. Amounts of each subcellular component recovered are expressed as Cell Equivalent (CE) of starting neutrophil counts. Whole cell lysates were obtained by suspending cells in a buffer consisting of 25mM Tris pH 7.4, 150mM Sodium Chloride, 5% Glycerol, 1% NP-40 with Complete Mini (Roche) protease inhibitor cocktail and PhosStop (Roche) phosphatase inhibitor cocktail. Cells were disrupted with a probe sonicator, centrifuged at 16000×g for 10 minutes, and supernatants were stored −80°C until use.
RNA isolation and cDNA synthesis
RNA was isolated from the neutrophils using the PureLink RNA Mini Kit (Ambion) with an added DNase I treatment. cDNA was obtained by reverse transcription of 0.5 μg RNA using the iScript cDNA Synthesis Kit (Bio-Rad).
Amplification of DNA using Polymerase Chain Reaction
TREM-1-specific forward (GCAGCCAGAAAGCTTGGCAGATAA) and reverse (ATCCACCAGCCAGGAGAATGACAA) primers (IDT, Inc.) PCR primers were designed to detect both membrane TREM-1 and TREM-1sv transcripts. Standard PCR, gel electrophoresis, and sequencing techniques were performed to detect amplification products.
Expression of TREM-1sv in HEK 293 cells
TREM-1sv was cloned from human neutrophils cDNA into pDNA3. Constructs were transfected into HEK 293 cells stably transfected with DAP12, the adaptor protein required for TREM-1 surface expression (ATCC) using Lipofectamine 2000 (Invitrogen) per the manufacturer's instructions. Analysis by ELISA as described below was performed at 48 hrs. after transfection.
Production of a polyclonal antibody against TREM-1sv
Analysis of the predicted protein sequences revealed one peptide unique to the splice variant (RCSTLSFSWLVDS)(Supplemental Figure 1). This peptide was synthesized (New England Peptide, Gardner, MA) and used to immunize rabbits. Antibody was affinity purified and assessed in an ELISA assay for its specificity for TREM-1sv as described below.
ELISA
Total TREM-1 in cell culture supernatants and whole cell lysates was quantified using a commercial ELISA kit (R&D Systems). TREM-1sv levels in the tissue culture samples and neutrophil supernatants were determined in a similar manner. The same polyclonal antibody used above (R&D Systems) which was generated against the extracellular immunoglobulin-like domain shared by mbTREM-1 and TREM-1sv was used as the capture antibody to pull down all TREM-1 species present in samples. E. coli expressed recombinant human TREM-1sv (aa 21-150, UniProt identifier Q9NP99-2) (R&D Systems) was used at concentration range of 10 pg/ml to 40 ng/ml to generate the standard calibration curve. Biotinylated rabbit anti-human TREM-1sv polyclonal antibody was subsequently used for detection (1ug/ml).
Western Blot
SDS-PAGE and subsequent Immunoblot was performed following standard methods. Briefly samples were mixed with sample loading buffer including β-mercaptoethanol and heated at 65°C for 15 minutes. Following electrophoresis on a Mini-Protean TGX 4-20% gradient gel, bands were blotted onto 0.45 micron nitrocellulose membrane and stained with Ponceau-S for total protein. Membrane was blocked for 4 hours at room temperature in a buffer with 3% BSA and incubated overnight at 4°C with either 0.8 μg/ml goat anti-human TREM-1 polyclonal antibody (R&D Systems #AF1278) or 2.0 ug/ml rabbit anti-human TREM-1sv polyclonal antibody. Secondary antibodies used were donkey anti-goat IgG-HRP and donkey anti-rabbit IgG-HRP respectively (both from Santa Cruz Biotechnologies) at a dilution of 1:15000. Blots were developed with Supersignal West Pico Chemiluminescent Substrate (Thermo Scientific) and visualized on film.
Neutrophil Stimulation Experiments
Neutrophils were freshly isolated from venous blood sampling of healthy donors. Following isolation, 1×107 neutrophils were resuspended in Hanks buffered saline solution supplemented with calcium and magnesium (Cellgro) and 1% human serum albumin and 0.1% glucose. Stimulants were added as follows: Heat killed P. aeruginosa were added at a multiplicity of infection 50:1, PAM3Cys4 (0.1 ug/ml), or LPS (0.1ug/ml) with 10% human serum. Following incubation for 30 minutes or 4 hours at 37°C in 5% CO2 incubator supernatants were collected by centrifugation.
Mass Spectrometry
37 μg of neutrophil and subcellular fraction cell lysates and 2 μg of expressed TREM-1sv protein standard were separated on precast gels under reducing and denaturing conditions. The gels were stained with Coomassie (G-250 Thermo Scientific). Relevant bands were excised for in-gel tryptic digestion following the procedure of Shevchenko et al. (33). The quality of the digest supernatant was determined prior to lyophilization by MALDI/TOF analysis on an Autoflex III TOF/TOF (Bruker). Peptides from digested samples were analyzed by nano-LC MS/MS using a Dionex 3000 UHP nano RSLC series HPLC system (Thermo-Electron) and linear ion-trap mass spectrometer (Thermo LTQ/XL, Thermo-Electron). MS/MS spectra were acquired in a data-dependent acquisition mode that automatically selected and fragmented the six most intense peaks from each MS spectrum. The same bands from a different gel were also analyzed on Triple TOF 5600 Plus MS (ABSciex) as detailed in supplemental figures.
LC-MS Data analysis
Peptide “.RAW” datasets were refined to a centroid list using Distiller (ver. 2.4, MatrixScience) and matched to protein sequences in the SwissProt and TrEMBl databases of July 23, 2012 with the Spectrum Mill Proteomics Workbench (Rev A.03.02.060, Agilent Technologies) accepting carbamidomethyl cysteine as a fixed modification and methionine oxidation as the single variable modification (34). Peptide and MS/MS mass tolerances were set at 1.8 and 0.4 Th, respectively. Taxonomy files were restricted to Homo Sapiens with common contaminants included. A Target:Decoy search strategy was used with mass tolerances set to 1.8 Th for peptide precursor ions and 0.4 Th for fragment ion data (35). ). Search requirements were set to a maximum local peptide FDR cut-off score of 0.5%, at least four y- or b-ion pairs and at least two unique peptides. The resulting peptide alignments were manually examined and accepted only if sequence identical peptides from the TREM-1sv standard protein revealed a consistent fragmentation pattern. Triple TOF MS data was analyzed using PeakView (ABSciex) and Mascot (Matrix Sciences).
Human Cytokine Methods
200,000 freshly isolated neutrophils from 3 healthy volunteers were incubated with 0.1ug/ml 21C7 or 0.1ug/ml 21C7 pre-incubated with 2ug/ml Trem-1sv or 0.1ug/ml mouse IgG1 for either 4 hours or 18 hours at 37°C/5% CO2. Each treatment was performed in triplicate in 3 separate wells of a 96-well plate. Supernatants of samples, centrifuged at 500xg for 10 minutes, were stored at −20°C until analysis.
Secreted IL-1β in supernatants were quantified using a customized multiplex bead assay (MILLIPLEX® MAP Human Cytokine Magnetic Bead Panel, EMD Millipore catalog # HCYTOMAG-60K-3plex) per the manufacturer-supplied protocol. A Bio-Plex Pro II magnetic bead plate washer (Bio-Rad) was used to wash magnetic beads. Fluorescence intensity was measured on a Bio-Plex 200 system using Bio-Plex Manager v6.1 software. Data collection parameters were set to 50 beads per region, 60-second timeout, bead map of 100 regions, sample size of 100 μl, and gate setting of 5,000–25,000. All samples were measured in triplicate.
Results
Detection of TREM-1sv in human primary neutrophils
To investigate the subcellular location of TREM-1 isoforms, we fractionated primary human neutrophils using nitrogen cavitation and discontinuous Percoll gradient (32). This procedure separates neutrophils into the following fractions: 1) gamma fraction enriched for plasma membranes and secretory vesicles, 2) alpha fraction which is enriched in primary or azurophilic granules, 3) beta fraction enriched for secondary (secretory) granules. As positive controls for the Western blots, we used recombinantly expressed tagged TREM-1 (Abcam), which migrates at ~48 kDa, and recombinant TREM-1sv protein lacking the transmembrane region, which migrates at 15kDa (R&D Systems). The predicted molecular weight of non-glycosylated native human monomeric membrane TREM-1 (mbTREM-1) is 23.8 kDa. Using an antibody that recognizes the conserved extracellular immunoglobulin (Ig)-like domain of TREM-1, four different TREM-1 species were detected in the fractions by Western blot (Figure 1). A ~55kDa band, ~42 kDa band, a ~24kDa band, and a ~15kDa band were detected in unfractionated neutrophils. The ~55kDa molecular weight species was also detected in the alpha and beta fractions. We speculate that this band is a multimeric form of TREM-1. The gamma fraction (membrane/secretory vesicles) was enriched for the 42kDa band, consistent with previously described aberrant migration of TREM-1 that has been attributed to glycosylation and/or dimerization of the molecule at the cell surface (36). A ~24 kDa isoform, consistent with full length monomeric TREM-1 was detected in only in the alpha fraction. A ~15kDa TREM-1 isoform was identified in the both alpha and beta fractions. This ~15 kDa species is consistent with the size the processed form of a putative spice variant of TREM-1, lacking the transmembrane domain.
Figure 1.

Western Blot of human neutrophil fractions using antibody against the TREM-1 Ig-like domain. Positive controls are tagged form of membrane TREM-1 (mb)(Abcam) migrating at 48 kDa and recombinant splice variant TREM-1 (sv) (RnD Systems) migrating at 15 kDa. Neutrophil samples include unfractionated neutrophils (WCL)(45×106 CE) where four TREM-1 species are noted (55kDa, 42kDa, and 24kDa and 15kDa), gamma (γ) fraction (86×106 CE)(42 kD TREM-1 species), beta fraction (β)(111×106 CE)(55kDa and 15kDa species) and alpha fraction (α)(97×106 CE)(55kDa, 24kDa and 15kDa TREM-1 species) respectively. Cell Equivalents (CE) Molecular size ladder (L)(Precision Plus Western Standards,BioRad). Representative blot shown, n=3 independent donors.
Detection of a splice variant form of TREM-1 mRNA in human neutrophils
The presence of a 15 kDa TREM-1 isoform in neutrophil primary and secondary granules lead us to hypothesize that this protein was the product of an alternatively spliced TREM-1 transcript. To investigate whether human neutrophils express a splice variant mRNA for TREM-1, we examined the RNA profiles from neutrophils isolated from both septic patients and healthy controls. Using PCR primers that amplify a portion of TREM-1 coding region (noted by arrows in Figure 2A), we analyzed the TREM-1 transcripts. Gel electrophoresis of PCR products revealed the presence of two products, one 299 bp and another 492 bp (Figure 2B) which were excised and sequenced. The PCR products encoded two distinct transcripts of human TREM-1, one sequence which included a predicted transmembrane region (“membrane TREM-1,” mbTREM-1) and a second product in which the cytoplasmic domain of TREM-1 was absent. We termed this latter product “splice variant TREM-1” (TREM-1sv). We next cloned and sequenced the full cDNAs for TREM-1sv and mbTREM-1 from human neutrophils. These cDNA sequences matched GenBank database sequences NM_001242590.1 and NM_018643.3, respectively (http://www.ncbi.nlm.nih.gov/genbank/). The presence of an mRNA lacking the transmembrane region led us to hypothesize that some or all of the soluble form of TREM-1 found in patient samples may be due to translation of this novel mRNA and not due to cleavage or shedding of mbTREM-1 as previously suggested (27).
Figure 2.
Membrane TREM-1 (mbTREM-1) and splice variant TREM-1 (TREM-1sv) transcripts isolated from primary human neutrophils. 1A) mRNA schematic of TREM-1 transcripts: exon 1 (signal sequence, 113 bp, dots), exon 2 (Ig-like domain, 357 bp, black), exon 3 (stalk, 193 bp, white) present in membrane isoform, but not splice variant, exon 4 (terminal region in membrane isoform, 106 bp, stripe, is truncated to 47 bp in splice variant due to a frame shift creating a novel stop codon. 1B) PCR products using forward primer GCAGCCAGAAAGCTTGGCAGATAA and reverse primer ATCCACCAGCCAGGAGAATGACAA (arrows). Primers span the splice region, resulting in a 492 bp product for mbTREM-1 and 299 bp product for TREM-1sv. β actin was used as load control.
Identification of TREM-1sv
To investigate whether the 15 kDa band in neutrophil granules was TREM-1sv, the band was excised and subjected to analysis by mass spectrometry (Figure 3, Supplemental Figure 2). Three peptides (85IILEDYHDHGLLR97, 76NSHPVQVGR84, 54AWQIIRDGEMPK65) were identified. IILEDYHDHGLLR (Figure 3, Panel 1) and NSHPVQVGR (Figure 3, Panel 2) were identified on the linear ion trap with Spectrum Mill protein scores of 14.74 and 5.83 with corresponding % spectral product ions (SPI) of 69.5 and 100, respectively. AWQIIRDGEMPK (Supplemental Figure 2) was identified using a triple time-of-flight (TOF) system with a Mascot protein score of 31 and at 0.05 significance. MS/MS spectra were compared to those obtained from in-gel digests of TREM-1sv standard in order to assign its identity.
Figure 3.
TREM-1 identification by mass spectroscopy. To confirm interpretation of spectra from 15 kDa excised band of neutrophil alpha fraction, SDS-Page band of recombinant splice variant TREM-1 was excised and analyzed by LCMSMS under identical conditions. Panel 1 and 2 show 2 different peptides identified in the 15kDa excised band. The predicted trypsin digest peptides with major b- and y- series ions are shown (A) of each panel where b- series are the sequential amino acid residues fragmented from the N-terminus and y- series are fragments starting from the C-terminus. The fragmentation pattern for the neutrophil derived band (B) is shown in comparison to corresponding peptide derived from recombinant TREM-1 (C). Spectra labeled with respective mass to charge (m/z) values in various charge states are shown with assigned b- or y- ions. Sequence interpretation was deemed to be correct because the major b- and y- series ions match.
Generation of a polyclonal antibody against TREM-1sv
Next we compared the primary sequence of mbTREM-1 and TREM-1sv proteins and identified a single peptide (RCSTLSFSWLVDS, Supplemental Figure 1) unique to TREM-1sv. A polyclonal antibody was raised against the peptide sequence (anti TREM-1sv)(see methods).
Recombinant TREM-1sv is translated into a soluble secreted protein
mbTREM-1 and TREM-1sv were cloned from human neutrophil cDNA. HEK 293 cells were transfected with expression vectors containing these constructs. mbTREM-1 was transfected into HEK 293 stably expressing DAP-12, which is required for TREM-1 surface expression. Whole cell lysates were analyzed by ELISA (R&D Systems). The capture and detection anti-TREM-1 antibodies in this kit are directed against the extracellular Ig-like domain shared by mbTREM-1 and TREM-1sv, and thus both TREM-1 isoforms can be detected by this ELISA assay. As expected, no TREM-1 was detected by ELISA in untransfected HEK 293 cells. TREM-1 was detected in lysates from cells transfected with either mbTREM-1 and TREM-1sv constructs (Figure 4A). Western blot analysis of the same cell lysates utilizing a polyclonal anti TREM-1 antibody revealed a ~27 kDa band present in cells transfected with mbTREM-1 but no reactivity with either untransfected cells or cells transfected with TREM-1sv. We hypothesized that TREM-1sv, lacking the transmembrane region and DAP-12 interacting lysine, was primarily secreted and thus we examined cell supernatants by ELISA and Western blot. For supernatant ELISA, we utilized a polyclonal anti TREM-1 capture antibody as above, followed by with detection a polyclonal antibody against the TREM-1sv peptide. TREM-1sv was observed in the supernatant of cells transfected with TREM-1sv (Figure 4C). No reactivity was seen with supernatant from untransfected cells, or cells transfected with mbTREM-1 construct (Figure 4C). Western blot analysis of supernatant using the anti TREM-1sv antibody demonstrated a 15kDa band present only the TREM-1sv transfected cells. These data show that the custom anti-TREM-1sv antibody specifically detects TREM-1sv and suggest that TREM-1sv is secreted by the transfected cells.
Figure 4.
Detection of mbTREM-1 and TREM-1sv proteins in transfected HEK 293 cells. A. ELISA of whole cell lysates using antibody which recognizes both TREM-1sv and mbTREM-1 isoforms. B. Western Blot of whole cell lysates using antibody which recognizes both TREM-1sv and mbTREM-1 isoforms. C. ELISA of supernatant from transfected HEK 293 cells using antibody generated against TREM-1sv specific peptide. D. Western blot of supernatant using TREM-1sv antibody.
TREM-1sv is released by human neutrophils following stimulation
Neutrophils and monocytes release a soluble form of TREM-1 into the supernatant following TLR ligation(9, 23). To investigate whether TREM-1sv is released under these conditions, freshly isolated human neutrophils from 7 individual donors were stimulated for 4 hours with LPS, P. aeruginosa, or PAM(3)Cys4 (Figure 5A). TREM-1sv was detected in neutrophil supernatant following exposure to P. aeruginosa and PAM(3)Cys4 by ELISA using the TREM-1sv antibody (P < 0.05). Western blot analysis of neutrophil supernatant following P. aeruginosa stimulation demonstrated the presence of a 15 kD band, consistent with TREM-1sv (Figure 5B). The release of primary granules under these conditions was confirmed by Western Blot analysis of supernatants for myeloperoxidase (MPO) (data not shown). Immunoblot, MS, and ELISA data, taken together, demonstrate that preformed TREM-1sv is released by mature human neutrophils under physiologic conditions.
Figure 5.

Release of TREM-1sv following in vitro stimulation of human neutrophils. A. TREM-1sv ELISA of neutrophil supernatant from 7 individual donors stimulated with LPS (10ng/ml), heat killed P. aeruginosa, or PAM(3)Cys4 (TLR1/2 agonist). ELISA data were not normally distributed and therefore are expressed in natural log for analysis. Symbols represent individual donors. * indicates p value< 0.05. B. Western blot analysis with anti TREM-1 Ig-like domain antibody of supernatant from unstimulated neutrophils (−) and neutrophils stimulated with P. aeruginosa (+). Recombinant TREM-1sv (SV) used as positive control. Molecular size ladder (L).
Function of TREM-1sv
To assess the functional role of TREM-1sv, we utilized the well characterized TREM-1 agonist antibody (21C7) as a surrogate TREM-1 ligand. Binding of this antibody mimics TREM-1 ligand binding resulting in proinflammatory cytokines secretion in primary cells and TREM-1 reporter cells lines(9, 37). Therefore to test whether TREM-1sv could act as a decoy receptor, inhibiting membrane TREM-1 signaling, we utilized 21C7 (gift from Marco Colonna) in a competitive bind assay. Freshly isolated human neutrophils were treated with 21C7 in the absence and presence of recombinant TREM-1sv (Figure 6). Indeed we found that the addition of exogenous soluble TREM-1sv abrogated 21C7 binding to membrane TREM-1 on the surface of human neutrophils as assessed with flow cytometry (Supplemental Figure 3). We next assessed whether soluble TREM-1sv could inhibit TREM-1 mediated cytokine secretion in human neutrophils (n=3 donors). Neutrophils were treated with total mouse IgG1, media only, 21C7, or 21C7 in the presence of TREM-1sv and cytokine release was measured using MILLIPLEX® MAP Human Cytokine Magnetic Beads per the manufacturer-supplied protocol. TREM-1 agonist Mab 21C7 significantly increased IL1β release in all 3 donors. However, the concomitant addition of 21C7 and exogenous TREM-1sv resulted in a significant decrease in proinflammatory cytokine secretion (Figure 6). While these data suggest that TREM-1sv competes with membrane TREM-1 for ligand binding, it is still possible that TREM-1sv could have other direct cellular effects. Nevertheless, these data do demonstrate in vitro that TREM-1sv inhibits membrane TREM-1 mediated proinflammatory cytokine secretion and suggests that TREM-1sv may function as a counter regulatory molecule in vivo.
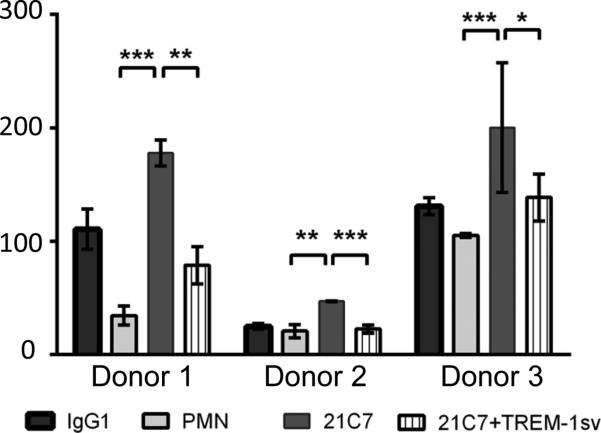
Figure 6.
TREM-1sv inhibits TREM-1 mediated IL1β release following in vitro stimulation of human neutrophils. IL1β production from human neutrophils treated with control IgG1, media, 21C7 or 21C7 plus TREM-1sv for 18 hours. 3 individual donors * p < 0.05, ** p<0.001, *** p <0.0001.
Discussion
Selective granule mobilization allows the neutrophil to rapidly alter proteins displayed on the cell surface and also quickly release needed soluble molecules into the local environment. The hierarchical release of these intracellular stores suggests that these organelles are specifically mobilized at different times during the neutrophil immune response(3). For example, secretory vesicles, the most readily released neutrophil organelles, contain large amounts of β2 integrin Mac-1(CD11b/CD18)(38). These vesicles are released at the time of neutrophil adherence and transendothelial migration, immediately increasing the surface expression of Mac-1, a molecule necessary for migration (38, 39). In contrast, the release of primary granules, which contain MPO among other destructive proteases, requires more vigorous neutrophil stimulation. Interestingly, our data demonstrate that TREM-1sv is present in these primary granules, as well as secondary granules.
The presence of a soluble TREM-1 isoform in neutrophil granules suggest that membrane TREM-1 signaling is simultaneously modulated by release of TREM-1sv at the time of neutrophil activation. Soluble forms of immune receptors often serve as counter regulatory molecules to modulate the inflammatory response (11-15). These soluble receptors temper the interaction between ligands and membrane-bound receptors to fine tune the immune response locally and restore homeostasis systemically following a pathologic insult. While the physiologic role of soluble TREM-1 in human disease is not clear, animal models suggest that the presence of this molecule in serum is anti-inflammatory. For example, administration of soluble forms of the TREM-1 molecule improves survival in rodent models of pneumonia and sepsis (16-18). This improved survival is accompanied by a decrease in systemic cytokine and chemokine production, suggesting that soluble TREM-1 functions in vivo as an anti-inflammatory molecule. We have previously shown that TREM-1 is required for neutrophil transepithelial migration into the lung during pneumonia (13). Based on this new data, we would speculate that, upon neutrophil activation, the membrane form of TREM-1 facilitates neutrophil migration across the epithelial barrier and synergizes with TLR signaling to amplify local inflammation. Concomitantly, granular exocytosis results in local release of the TREM-1sv. This TREM-1sv can potentially modulate the immune response in two ways. Locally, TREM-1sv release could inhibit additional mbTREM-1-mediated neutrophil migration and cytokine production and systemically it could prevent disseminated inflammation by serving as a decoy receptor for the as yet unknown TREM-1 ligand(s). Thus, this preformed reservoir of TREM-1sv circulating in the blood has important therapeutic implications. Moreover, these data suggest that selective neutrophil degranulation to release TREM-1sv could be an effective therapy to restore homeostasis during immune dysregulation.
Studies examining the granular contents of human neutrophils have been challenging for a number of reasons. By design, neutrophil granules contain a wide variety of protease, lipases, and reactive oxygen species which challenge our ability to recover quality protein for analysis. Moreover, these terminally differentiated cells must be freshly isolated from the blood as they do not survive in culture and no adequate cell lines exist. In recent years, several groups reported proteomic analysis of human neutrophil granular contents (40-43). To our knowledge, this is the first time that TREM-1, a well-known and highly expressed neutrophil protein, has been definitively identified using mass spectrometry.
The content of neutrophil subcellular compartments are a critical source of secretory and plasma membrane molecules that can be readily mobilized to shape local and systemic immune responses. Herein, we identify a novel TREM-1 isoform, TREM-1sv, present in human neutrophils. Moreover we demonstrate that TREM-1sv attenuates TREM-1 mediated proinflammatory cytokine secretion. These data set the stage for future studies to examine regulation of TREM-1sv release and ultimately whether TREM-1sv could be used for therapeutic purposes to modulate neutrophil mediated inflammation and transepithelial migration. A detailed understanding of the regulation of TREM-1sv production and release is necessary if we hope to utilize soluble forms of the TREM-1 molecule in the immune modulation of sepsis.
Supplementary Material
Acknowledgements
We thank Bill Nauseef (University of Iowa) and Marco Colonna (Washington University in St. Louis) for helpful discussions. For technical support we thank Kevin Leidal (University of Iowa) for help with neutrophil subcellular fractionation and Yalan Li (University of Iowa), Lake Paul (Purdue), and Victoria Hedrick (Purdue) for MS analysis of neutrophil subcellular granule lysates.
Footnotes
Authorship: JKT,SB,KK conceived of the study; JKT,SB,KK,MRP,MP,MV designed and performed experiments; JKT,SB,KK,MRP,KD,MV analyzed data; SB,MP performed analysis of mass spectrometry data and JKT,SB,KK,MV,MRP,KD manuscript preparation.
The authors have no relevant conflicts of interest to disclose.
References
- 1.Nauseef WM, Borregaard N. Neutrophils at work. Nat Immunol. 2014;15(7):602–611. doi: 10.1038/ni.2921. [DOI] [PubMed] [Google Scholar]
- 2.Mantovani A, Cassatella MA, Costantini C, Jaillon S. Neutrophils in the activation and regulation of innate and adaptive immunity. Nature Reviews Immunology. 2011;11(8):519–31. doi: 10.1038/nri3024. [DOI] [PubMed] [Google Scholar]
- 3.Borregaard N, Cowland JB. Granules of the human neutrophilic polymorphonuclear leukocyte. Blood. 1997;89(10):3503–21. [PubMed] [Google Scholar]
- 4.Borregaard N. Neutrophils, from marrow to microbes. Immunity. 2010;33(5):657–70. doi: 10.1016/j.immuni.2010.11.011. [DOI] [PubMed] [Google Scholar]
- 5.Kolaczkowska E, Kubes P. Neutrophil recruitment and function in health and inflammation. Nat Rev Immunol. 2013;13(3):159–175. doi: 10.1038/nri3399. [DOI] [PubMed] [Google Scholar]
- 6.Beutler B. Inferences, questions and possibilities in Toll-like receptor signaling. Nature. 2004;430(6996):257–63. doi: 10.1038/nature02761. [DOI] [PubMed] [Google Scholar]
- 7.Klesney-Tait J, Turnbull IR, Colonna M. The TREM receptor family and signal integration. Nature Immunology. 2006;7(12):1266–1273. doi: 10.1038/ni1411. [DOI] [PubMed] [Google Scholar]
- 8.Kawai T, Akira S. Toll-like Receptors and Their Crosstalk with Other Innate Receptors in Infection and Immunity. Immunity. 2011;34(5):637–650. doi: 10.1016/j.immuni.2011.05.006. [DOI] [PubMed] [Google Scholar]
- 9.Bouchon A, Dietrich J, Colonna M. Cutting edge: inflammatory responses can be triggered by TREM-1, a novel receptor expressed on neutrophils and monocytes. J. Immunol. 2000;164:4991–4995. doi: 10.4049/jimmunol.164.10.4991. [DOI] [PubMed] [Google Scholar]
- 10.Bouchon A, Facchetti F, Weigand MA, Colonna M. TREM-1 amplifies inflammation and is a crucial mediator of septic shock. Nature. 2001;410:1103–1107. doi: 10.1038/35074114. [DOI] [PubMed] [Google Scholar]
- 11.Gibot S. Soluble triggering receptor expressed on myeloid cells and the diagnosis of pneumonia. New England Journal of Medicine. 2004;350:451–458. doi: 10.1056/NEJMoa031544. [DOI] [PubMed] [Google Scholar]
- 12.Knapp S, Gibot S, de Vos A, Versteeg HH, Colonna M, van der Poll T. Cutting edge: expression patterns of surface and soluble triggering receptor expressed on myeloid cells-1 in human endotoxemia. J Immunol. 2004;173(12):7131–4. doi: 10.4049/jimmunol.173.12.7131. [DOI] [PubMed] [Google Scholar]
- 13.Klesney-Tait J, Keck K, Li X, Gilfillan S, Otero K, Baruah S, et al. Transepithelial migration of neutrophils into the lung requires TREM-1. The Journal of clinical investigation. 2013;123(1):138–149. doi: 10.1172/JCI64181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wu J, Li J, Salcedo R, Mivechi NF, Trinchieri G, Horuzsko A. The Proinflammatory Myeloid Cell Receptor TREM-1 Controls Kupffer Cell Activation and Development of Hepatocellular Carcinoma. Cancer Research. 2012;72(16):3977–3986. doi: 10.1158/0008-5472.CAN-12-0938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gibot S, Kolopp-Sarda MN, Bene MC, Cravoisy A, Levy B, Faure GC, et al. Plasma level of a triggering receptor expressed on myeloid cells-1: its diagnostic accuracy in patients with suspected sepsis. Ann Intern Med. 2004;141(1):9–15. doi: 10.7326/0003-4819-141-1-200407060-00009. [DOI] [PubMed] [Google Scholar]
- 16.Gibot S, Cravoisy A, Levy B, Bene MC, Faure G, Bollaert PE. Soluble triggering receptor expressed on myeloid cells and the diagnosis of pneumonia. N Engl J Med. 2004;350(5):451–8. doi: 10.1056/NEJMoa031544. [DOI] [PubMed] [Google Scholar]
- 17.Liao R, Sun TW, Yi Y, Wu H, Li YW, Wang JX, et al. Expression of TREM-1 in hepatic stellate cells and prognostic value in hepatitis B-related hepatocellular carcinoma. Cancer Sci. 2012;103(6):984–92. doi: 10.1111/j.1349-7006.2012.02273.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ford JW, McVicar DW. TREM and TREM-like receptors in inflammation and disease. Curr Opin Immunol. 2009;21(1):38–46. doi: 10.1016/j.coi.2009.01.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Dopheide JF, Doppler C, Scheer M, Obst V, Radmacher MC, Radsak MP, et al. Critical limb ischaemia is characterised by an increased production of whole blood reactive oxygen species and expression of TREM-1 on neutrophils. Atherosclerosis. 2013;229(2):396–403. doi: 10.1016/j.atherosclerosis.2013.05.029. [DOI] [PubMed] [Google Scholar]
- 20.Collins CE, La DT, Yang HT, Massin F, Gibot S, Faure G, et al. Elevated synovial expression of triggering receptor expressed on myeloid cells 1 in patients with septic arthritis or rheumatoid arthritis. Ann Rheum Dis. 2009;68(11):1768–74. doi: 10.1136/ard.2008.089557. [DOI] [PubMed] [Google Scholar]
- 21.Su LX, Feng L, Zhang J, Xiao YJ, Jia YH, Yan P, et al. Diagnostic value of urine sTREM-1 for sepsis and relevant acute kidney injuries: a prospective study. Crit Care. 2011;15(5):R250. doi: 10.1186/cc10508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Koussoulas V. Soluble triggering receptor expressed on myeloid cells (sTREM-1): a new mediator involved in the pathogenesis of peptic ulcer disease. Eur. J. Gastroenterol. Hepatol. 2006;18:375–379. doi: 10.1097/00042737-200604000-00010. [DOI] [PubMed] [Google Scholar]
- 23.Bleharski JR. A role for triggering receptor expressed on myeloid cells-1 in host defense during the early-induced and adaptive phases of the immune response. Journal of immunology. 2003;170:3812–3818. doi: 10.4049/jimmunol.170.7.3812. [DOI] [PubMed] [Google Scholar]
- 24.Gibot S. A soluble form of the triggering receptor expressed on myeloid cells-1 modulates the inflammatory response in murine sepsis. J. Exp. Med. 2004;200:1419–1426. doi: 10.1084/jem.20040708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Campanholle G, Mittelsteadt K, Nakagawa S, Kobayashi A, Lin SL, Gharib SA, et al. TLR-2/TLR-4 TREM-1 signaling pathway is dispensable in inflammatory myeloid cells during sterile kidney injury. PLoS One. 2013;8(7):e68640. doi: 10.1371/journal.pone.0068640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Schenk M, Bouchon A, Seibold F, Mueller C. TREM-1--expressing intestinal macrophages crucially amplify chronic inflammation in experimental colitis and inflammatory bowel diseases. J Clin Invest. 2007;117(10):3097–106. doi: 10.1172/JCI30602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gómez-Piña V, Soares-Schanoski A, Rodríguez-Rojas A, del Fresno C, García F, Vallejo-Cremades MT, et al. Metalloproteinases Shed TREM-1 Ectodomain from Lipopolysaccharide-Stimulated Human Monocytes. The Journal of Immunology. 2007;179(6):4065–4073. doi: 10.4049/jimmunol.179.6.4065. [DOI] [PubMed] [Google Scholar]
- 28.Mahdy AM. Production of soluble triggering receptor expressed on myeloid cells by lipopolysacccharide-stimulated human neutrophils involves de novo protein synthesis. Clin. Vaccine Immunol. 2006;13:492–495. doi: 10.1128/CVI.13.4.492-495.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Gingras MC, Lapillonne H, Margolin JF. TREM-1, MDL-1, and DAP12 expression is associated with a mature stage of myeloid development. Mol. Immunol. 2002;38:817–824. doi: 10.1016/s0161-5890(02)00004-4. [DOI] [PubMed] [Google Scholar]
- 30.Levy M, Fink MP, Marshall JC, Abraham E, Angus D, Cook D, et al. SCCM/ESICM/ACCP/ATS/SIS International Sepsis Definitions Conference Critical Care Medicine. 2001;2003(31):1250–1256. doi: 10.1097/01.CCM.0000050454.01978.3B. [DOI] [PubMed] [Google Scholar]
- 31.Clark RA, Nauseef WA. Isolation and Functional Analysis of Neutrophils. Current Protocols in Immunology. 2009;(Supplement 6):7.23. doi: 10.1002/0471142735.im0723s19. [DOI] [PubMed] [Google Scholar]
- 32.Borregaard N, Heiple JM, Simons ER, Clark RA. Subcellular localization of the b-cytochrome component of the human neutrophil microbicidal oxidase: translocation during activation. J Cell Biol. 1983;97(1):52–61. doi: 10.1083/jcb.97.1.52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shevchenko A, Tomas H, Havlis J, Olsen JV, Mann M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc. 2006;1(6):2856–60. doi: 10.1038/nprot.2006.468. [DOI] [PubMed] [Google Scholar]
- 34.Perkins DN, Pappin DJ, Creasy DM, Cottrell JS. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis. 1999;20(18):3551–67. doi: 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 35.Elias JE, Haas W, Faherty BK, Gygi SP. Comparative evaluation of mass spectrometry platforms used in large-scale proteomics investigations. Nat Methods. 2005;2(9):667–75. doi: 10.1038/nmeth785. [DOI] [PubMed] [Google Scholar]
- 36.Radaev S, Kattah M, Rostro B, Colonna M, Sun PD. Crystal structure of the human myeloid cell activating receptor TREM-1. Structure. 2003;11:1527–1535. doi: 10.1016/j.str.2003.11.001. [DOI] [PubMed] [Google Scholar]
- 37.Read CB, Kuijper JL, Hjorth SA, Heipel MD, Tang X, Fleetwood AJ, et al. Cutting Edge: Identification of Neutrophil PGLYRP1 as a Ligand for TREM-1. The Journal of Immunology. 2015;194(4):1417–1421. doi: 10.4049/jimmunol.1402303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sengelov H, Follin P, Kjeldsen L, Lollike K, Dahlgren C, Borregaard N. Mobilization of granules and secretory vesicles during in vivo exudation of human neutrophils. J Immunol. 1995;154(8):4157–65. [PubMed] [Google Scholar]
- 39.von Andrian UH, Chambers JD, McEvoy LM, Bargatze RF, Arfors KE, Butcher EC. Two-step model of leukocyte endothelial cell interaction in inflammation: distinct roles for LECAM-1 and the leukocyte beta 2 integrins in vivo. Proceedings of the National Academy of Sciences. 1991;88(17):7538–7542. doi: 10.1073/pnas.88.17.7538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Lominadze G, Powell DW, Luerman GC, Link AJ, Ward RA, McLeish KR. Proteomic analysis of human neutrophil granules. Mol Cell Proteomics. 2005;4(10):1503–21. doi: 10.1074/mcp.M500143-MCP200. [DOI] [PubMed] [Google Scholar]
- 41.Luerman GC, Powell DW, Uriarte SM, Cummins TD, Merchant ML, Ward RA, et al. Identification of phosphoproteins associated with human neutrophil granules following chemotactic peptide stimulation. Mol Cell Proteomics. 2011;10(3):M110 001552. doi: 10.1074/mcp.M110.001552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Uriarte SM, Powell DW, Luerman GC, Merchant ML, Cummins TD, Jog NR, et al. Comparison of proteins expressed on secretory vesicle membranes and plasma membranes of human neutrophils. J Immunol. 2008;180(8):5575–81. doi: 10.4049/jimmunol.180.8.5575. [DOI] [PubMed] [Google Scholar]
- 43.Rorvig S, Ostergaard O, Heegaard NH, Borregaard N. Proteome profiling of human neutrophil granule subsets, secretory vesicles, and cell membrane: correlation with transcriptome profiling of neutrophil precursors. J Leukoc Biol. 2013;94(4):711–21. doi: 10.1189/jlb.1212619. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.