Abstract
Aim
We performed genome-wide and transcriptome-wide profiling to identify genes and single nucleotide polymorphisms (SNPs) associated with the response of triglycerides (TG) to exercise training.
Methods
Plasma TG levels were measured before and after a 20-week endurance training programme in 478 white participants from the HERITAGE Family Study. Illumina HumanCNV370-Quad v3.0 BeadChips were genotyped using the Illumina BeadStation 500GX platform. Affymetrix HG-U133+2 arrays were used to quantitate gene expression levels from baseline muscle biopsies of a subset of participants (N=52). Genome-wide association study (GWAS) analysis was performed using MERLIN, while transcriptomic predictor models were developed using the R-package GALGO.
Results
The GWAS results showed that eight SNPs were associated with TG training-response (ΔTG) at p<9.9×10−6, while another 31 SNPs showed p values <1×10−4. In multivariate regression models, the top 10 SNPs explained 32.0% of the variance in ΔTG, while conditional heritability analysis showed that four SNPs statistically accounted for all of the heritability of ΔTG. A molecular signature based on the baseline expression of 11 genes predicted 27% of ΔTG in HERITAGE, which was validated in an independent study. A composite SNP score based on the top four SNPs, each from the genomic and transcriptomic analyses, was the strongest predictor of ΔTG (R2=0.14, p=3.0×10−68).
Conclusions
Our results indicate that skeletal muscle transcript abundance at 11 genes and SNPs at a number of loci contribute to TG response to exercise training. Combining data from genomics and transcriptomics analyses identified a SNP-based gene signature that should be further tested in independent samples.
INTRODUCTION
Elevated triglycerides (TG) are strongly associated with increased risk of cardiovascular disease (CVD) in epidemiological studies.1–4 Furthermore, Mendelian randomisation studies of genetic variants affecting TG levels have suggested a causal role of TG on CVD and all-cause mortality.5,6 Physical activity is considered a major target for therapeutic lifestyle changes in the prevention and treatment of elevated TG.7,8 On average, exercise training induces reductions in TG levels ranging from 4 to 38 mg/dL.9 However, there is large inter-individual variation in the magnitude of changes in plasma TG levels derived from standardised exercise training. In fact, we recently reported that across six exercise intervention studies (N=1687), 10.3% of participants experienced a response of TG (increase of 0.42 mmol/L or greater) to exercise training that was qualified as being adverse.10
Innovative research strategies are needed to identify the molecular factors contributing to individual differences in response to regular exercise and to translate them into clinically useful applications, such as personalised exercise programmes.11 Genetic factors need to be taken into account in comprehensive personalised exercise medicine approaches, as their contributions are typically strong across populations studied to date and they are powerful determinants of the ability to benefit from regular exercise.11 In the HERITAGE Family Study (HERITAGE), the maximal heritability estimates for exercise-induced changes in TG were 32% in black participants and 29% in white participants.12 However, the genetic variants responsible for variation in TG response to regular exercise remain poorly understood. A limited number of candidate gene studies have provided evidence of the association of DNA sequence variants with TG response to lifestyle and exercise interventions, with nominal associations found for variants in the APOE, LIPC and PGS1 genes.13,14 However, these candidate genes explain only a small percentage of the variance in plasma TG response to exercise training. Thus, there is a need for innovative, unbiased approaches, such as integrated omics profiling, in the search for the genes and DNA sequence variants contributing to plasma TG response to regular exercise.
Integrated omics profiling (eg, combining data from genomics, epigenomics, transcriptomics, metabolomics and/or proteomics) allows for the profiling of the molecular factors that may impact the tolerance, effects and performance expected from regular exercise.11 We have previously shown that global RNA profiling of skeletal muscle combined with targeted genotyping increased the explanatory power of a gene signature for VO2max response to exercise training.15 A better understanding of the genetic or molecular factors associated with TG response to exercise could inform about the potential clinical utility of including this information in personalised exercise prescriptions. Therefore, the purpose of the present study was to perform both genome-wide and transcriptome-wide profiling in order to identify a single nucleotide polymorphism (SNP)-based gene signature that predicts the responsiveness of TG levels to exercise training.
METHODS
A detailed description of the study design, methodology and analyses can be found in the online only supplementary material.
HERITAGE Family Study
The participants, study design and exercise training protocol of the HERITAGE Family Study have been described elsewhere.16 This study sample includes data from 478 white participants from 99 nuclear families who completed the 20 week exercise programme and have baseline and post-training plasma lipid measurements. Participants were sedentary at baseline, normotensive or mildly hypertensive (<160/100 mm Hg) without medications for hypertension, diabetes or dyslipidemia.16 The study protocol had been approved by the Institutional Review Boards at each of the five participating centres of the HERITAGE Family Study consortium. Written informed consent was obtained from each participant.
Exercise training programme
Each participant in HERITAGE exercised three times per week for 20 weeks on cycle ergometers controlled by direct heart rate (HR) monitoring. Details of the exercise programme can be found elsewhere.16 Briefly, participants exercised at the HR associated with 55% of baseline maximal oxygen uptake (VO2max) for 30 min per session for the first 2 weeks. The duration and intensity were gradually increased every 2 weeks, until reaching 50 min and 75% of the HR associated with baseline VO2max. This level was maintained for the final 6 weeks of training. The protocol was standardised across all clinical centres and supervised to ensure that the equipment was working properly and that the participants were compliant with the protocol.
Determination of plasma lipids
Blood samples for plasma lipid assays were obtained from an antecubital vein into Vacutainer tubes containing EDTA in the morning after a 12-h fast with participants in a semirecumbent position. The blood samples were collected twice at baseline (on separate days), and again at 24 h and 72 h after the last exercise session. TG levels were determined in plasma by enzymatic methods using a Technicon RA-500 Analyzer (Bayer Corporation Inc, Tarrytown, New York, USA). The reproducibility of TG measurements in HERITAGE has been previously examined, with a coefficient of variation of 21.8, intraclass correlation of 0.79 and technical error of 0.21 (mmol/L).17 The response to exercise training (Δ) was computed as the difference between the average post-exercise training TG measures and the average baseline TG measures.
GWAS SNP genotyping
Genomic DNA was prepared from immortalised lymphoblastoid cell lines by commercial DNA extraction kits (Gentra Systems, Inc, Minneapolis, Minnesota, USA). Genome-wide association study (GWAS) SNPs were genotyped using Illumina HumanCNV370-Quad v3.0 BeadChips on the Illumina BeadStation 500GX platform. The choice of the CNV370 BeadChips was dictated by cost at the time (2008), as the chips met our need of genotyping at the genome-wide level in a cost-effective way. Overall, 277 133 SNPs with minor allele frequency (MAF) ≥0.10 passed all quality control (QC) measures and were tested in the GWAS analysis. More details of the geno-typing procedures and QC measures can be found in the online only supplementary material.
GWAS statistical analyses
For the individual SNP GWAS analyses, normalised trait residuals were used as phenotypes.18 The dependent variable (ΔTG) was adjusted for age (up to a cubic polynomial), baseline body mass index (BMI) and baseline TG using a stepwise regression procedure. Associations between genotyped GWAS SNPs and ΔTG were analysed using the total association model of MERLIN.19 The Bonferroni adjusted p value threshold of 1.8×10−7 represents genome-wide statistical significance (N=277 133 SNPs tested).
Regression models
Multivariable regression procedures were used to evaluate the overall contribution of the most significant GWAS SNPs on ΔTG. All GWAS SNPs with p≤1.0×10−4 were included. First, a regression model with backward elimination was used to filter out redundant SNPs (mainly due to strong pairwise linkage dis-equilibrium). The threshold for keeping SNPs in the model was p=0.05. The SNPs retained in the backward elimination model were then analysed in a forward stepwise regression model.
Conditional heritability analyses
Whether the most significant SNPs from the regression model contribute to the heritability of ΔTG was tested using conditional heritability analysis in MERLIN. If an SNP contributes to the genetic variance (heritability) of a trait, the heritability estimate should decrease when the marker is included as a covariate in the model. Starting with the SNP showing the greatest partial R2 in the final regression model, the SNPs were added one at a time in the heritability model until the estimate reached zero (heritability fully accounted for by the covariates (SNPs)) or no more SNPs remained.
Pathway analysis of GWAS associations
We performed pathway analysis of the GWAS SNP associations via gene set enrichment analysis (GSEA) using Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways as gene sets.20,21
We employed two different approaches to calculate gene-level statistics for GSEA: (1) the second most significantly associated SNP was used to summarise all SNPs for each gene22 and (2) Stouffer's method23 that combines p values from multiple individual statistical tests and calculates the gene set statistic using the inverse normal cumulative distribution function.
Affymetrix microarray analysis
Muscle biopsies of the vastus lateralis muscle were performed at baseline using the percutaneous needle biopsy technique in 52 HERITAGE white participants from the Québec Clinical Center. Total RNA was isolated from frozen muscle biopsies preserved in Tissue-Tek using Trizol and messenger RNA (mRNA) amplified with Ambion MessageAmp Premier following the manufacturer's instructions.
Affymetrix HG-U133+2 arrays were used to quantitate global gene expression levels. Raw microarray data have been deposited with Gene Expression Omnibus under accession number: GSE47874.23a After QC and removal of outliers (N=2 with Cook's distance >0.5),24 baseline microarray data were available for 49 participants.
Baseline RNA gene signature
We used GALGO,25 an R package for multivariate variable selection based on a genetic algorithm (GA) methodology, to derive a multivariate regression model of ΔTG in HERITAGE using baseline mRNA expression levels. In brief, this computational search procedure tries to find the best subset of genes for maximising the fitness of the regression model (defined by R2). During the GA search procedure, ¾ of the samples (N=37) were used for training, whereas the remaining 12 samples were completely omitted and used for validation. In order to better estimate the model accuracy (owing to the number of samples), the training data set was split in 100 different training and test sets and 4000 simulations were run. Owing to variability in the model performance across the splits (likely caused by interindividual differences), we chose to focus on the subset of models (N=512) that performed well in most splits (ie, low overall deviation and high accuracy; see online supplementary figure S1).
Replication in an independent cohort
We tested our predictive model in an independent training study, for which Affymetrix Gene 1.1 ST microarray data are publicly available (GSE53598), consisting of 18 overweight to obese normoglycaemic middle-aged men who underwent 12 weeks of mixed exercise training (2 days/week aerobic, 1 day/week resistance training).26,27
Association of top SNPs from RNA signature genes
The association of SNPs, in or near (±20 kb) the top RNA predictor genes, with ΔTG was tested in all HERITAGE white participants using imputed SNPs. Approximately 2.54 million HapMap SNPs (release 22, build 36, CEU population) were imputed using the HERITAGE QC'ed genotyped GWAS data and MACH software,28 as previously described.29 After removing monomorphic SNPs (N=32 904) and zeroing out Mendelian errors, an additional 114 394 SNPs were removed that had R2 <0.3 (indicating poor quality), missing rate over 5%, MAF <1%, and/or a Hardy-Weinberg equilibrium p value <10−6. More details of the imputation methods and QC can be found in the online only supplementary material. The association between imputed SNPs from the selected genes and ΔTG was tested using the same total association models described above for the GWAS analyses.
SNP summary score
An SNP summary score for ΔTG was constructed by combining the GWAS SNPs that accounted for all of the heritability in the conditional heritability analyses (N=4) with the top SNPs from the RNA-based classifier genes (N=4). Each SNP was recoded to reflect the number of favourable responses to regular exercise alleles (eg, favourable defined as a decrease in TG) in the following manner: 0 for no copy of the favourable response allele, 1 for one copy and 2 for two copies of the favourable response allele. The sum of the recoded SNPs was used as the SNP summary score. The association between the SNP summary score and TG response to exercise training in HERITAGE white participants (N=476) was analysed using a general linear model (Proc GLM in SAS V.9.4) adjusting for age, sex, baseline BMI and baseline TG.
RESULTS
The basic characteristics, including mean lipid values at baseline and in response to exercise training, of HERITAGE white participants with valid GWAS or gene expression data are shown in table 1. On average, high-density lipoprotein cholesterol and lipoprotein lipase (LPL) activity increased with regular exercise, while TG and hepatic lipase activity decreased. More detailed descriptions of the lipid trait responses to regular exercise in HERITAGE have been previously published.30–32
Table 1.
Descriptive data, including baseline and response to exercise training values for lipid, lipoprotein and lipase phenotypes, for HERITAGE white participants with valid GWAS (left) and gene expression (right) data
| Participants with GWAS data (N=478) |
Subsample with gene expression data (N=49) |
||||
|---|---|---|---|---|---|
| Variable | Mean (SD) | 95% CI | Range | Mean (SD) | 95% CI |
| Age, years | 35.9 (14.5) | 17.0 to 65.2 | 32.9 (14.3) | ||
| BMI, kg/m2 | |||||
| Baseline | 25.9 (5.0) | 17.0 to 47.5 | 25.9 (4.1) | ||
| Response to exercise training | –0.09 (0.7) | –0.15 to –0.02 | –3.5 to 2.9 | –0.004 (0.8) | –0.22 to 0.21 |
| Triglycerides, mmol/L | |||||
| Baseline | 1.4 (0.8) | 0.4 to 6.3 | 1.4 (0.8) | ||
| Response to exercise training | –0.02 (0.4) | –0.06 to 0.02 | –1.8 to 2.3 | –0.2 (0.5) | –0.28 to –0.03 |
| HDL-C, mmol/L | |||||
| Baseline | 1.0 (0.3) | 0.5 to 2.0 | 1.1 (0.2) | ||
| Response to exercise training | 0.04 (0.1) | 0.03 to 0.05 | –0.3 to 0.6 | 0.07 (0.1) | 0.04 to 0.11 |
| LDL-C, mmol/L | |||||
| Baseline | 3.0 (0.8) | 0.9 to 6.0 | 2.8 (0.8) | ||
| Response to exercise training | –0.004 (0.4) | –0.04 to 0.03 | –1.2 to 1.5 | 0.01 (0.4) | –0.09 to 0.12 |
| PH-LPL, nmol/min/mL | |||||
| Baseline | 56.7 (30.1) | 2.8 to 205.5 | 57.2 (26.7) | ||
| Response to exercise training | 7.6 (28.1) | 5.0 to 10.2 | –98.3 to 96.2 | 9.4 (26.0) | 2.1 to 16.7 |
| PH-HL, nmol/min/mL | |||||
| Baseline | 209.2 (70.7) | 53.9 to 379.7 | 216.6 (77.8) | ||
| Response to exercise training | –11.5 (48.4) | –16.1 to –7.0 | –203.2 to 143.6 | –9.0 (51.9) | –23.5 to 5.5 |
BMI, body mass index, GWAS, genome-wide association study; HDL-C, high-density lipoprotein cholesterol; LDL-C, low-density lipoprotein cholesterol; PH-LPL, post-heparin lipoprotein lipase; PH-HL, post-heparin hepatic lipase.
GWAS associations for TG response to exercise training
Online supplementary figure S2 displays a Manhattan plot of the GWAS results for the response of TG to exercise training across 22 autosomes. In the individual SNP analyses, 39 SNPs showed associations of p<1×10−4 (table 2). None of the previously identified TG GWAS loci33 that we tested showed evidence of an association with TG response to exercise training (see online supplementary table S1).
Table 2.
List of GWAS SNPs associated with TG response to exercise training at p<1.5×10–4 in HERITAGE white participants
| SNP | Chromosome | Position* | Allele | Freq | β † | H2 | p Value | Gene‡ |
|---|---|---|---|---|---|---|---|---|
| rs2396190 | 2 | 225 718 082 | A | 0.77 | –0.45 | 5.67 | 3.30×10–7 | DOCK10 (90 kb) |
| rs222158 | 21 | 26 794 032 | A | 0.67 | –0.38 | 4.95 | 1.76×10–6 | CYYR1 |
| rs3906453 | 18 | 49 512 336 | A | 0.76 | 0.39 | 4.43 | 2.80×10–6 | DCC (200 kb) |
| rs3862435 | 15 | 88 883 536 | G | 0.91 | –0.60 | 4.77 | 6.67×10–6 | CRTC3 |
| rs3862436 | 15 | 88 883 873 | G | 0.91 | –0.60 | 4.76 | 6.67×10–6 | CRTC3 |
| rs3861882 | 9 | 131 505 125 | A | 0.72 | –0.38 | 4.75 | 6.85×10–6 | PRRX2 |
| rs2646822 | 1 | 215 602 483 | G | 0.77 | 0.42 | 4.92 | 9.84×10–6 | GPATCH2 (69 kb) |
| rs2646817 | 1 | 215 591 874 | C | 0.77 | 0.42 | 4.91 | 9.84×10–6 | GPATCH2 (80 kb) |
| rs12153753 | 5 | 134 692 476 | G | 0.85 | –0.46 | 4.12 | 1.83×10–5 | C5orf66 |
| rs7712997 | 5 | 134 690 996 | C | 0.89 | –0.51 | 4.05 | 1.95×10–5 | C5orf66 |
| rs9357234 | 6 | 37 138 195 | C | 0.69 | 0.35 | 4.09 | 1.97×10–5 | FGD2 (33 kb) |
| rs726553 | 2 | 225 724 738 | A | 0.63 | –0.33 | 4.14 | 2.29×10–5 | DOCK10 (100 kb) |
| rs1452404 | 4 | 109 599 177 | A | 0.90 | 0.52 | 3.96 | 2.59×10–5 | LEF1 (290 kb) |
| rs13093483 | 3 | 68 376 923 | A | 0.88 | –0.48 | 4.00 | 2.97×10–5 | FAM19A1 |
| rs738958 | 22 | 33 001 012 | G | 0.74 | 0.36 | 4.09 | 3.12×10–5 | LARGE (350 kb) |
| rs2722171 | 12 | 102 973 617 | C | 0.80 | –0.41 | 4.32 | 3.33×10–5 | GLT8D2 |
| rs7850237 | 9 | 89 643 439 | A | 0.86 | –0.46 | 4.13 | 4.18×10–5 | SPATA31C1 (75 kb) |
| rs713765 | 22 | 33 005 903 | A | 0.78 | 0.38 | 3.96 | 4.59×10–5 | LARGE (350 kb) |
| rs4742057 | 9 | 4 943 916 | G | 0.58 | 0.30 | 3.57 | 5.09×10–5 | JAK2 (32 kb) |
| rs795602 | 4 | 140 801 382 | A | 0.52 | 0.31 | 3.82 | 5.29×10–5 | MGST2 (5 kb) |
| rs11666431 | 19 | 2 904 087 | G | 0.58 | 0.32 | 3.92 | 5.29×10–5 | ZNF77 (9 kb) |
| rs4045101 | 19 | 24 238 309 | A | 0.63 | –0.31 | 3.70 | 5.64×10–5 | ZNF254 (140 kb) |
| rs7645395 | 3 | 64 326 912 | A | 0.79 | –0.36 | 3.45 | 5.71×10–5 | PRICKLE2 & ADAMTS9 (15 kb) |
| rs1906058 | 16 | 6 084 649 | C | 0.53 | 0.30 | 3.69 | 5.74×10–5 | RBFOX1 |
| rs7253132 | 19 | 24 170 037 | A | 0.60 | –0.31 | 3.73 | 5.91×10–5 | ZNF254 (65 kb) |
| rs9469986 | 6 | 11 857 166 | G | 0.80 | 0.39 | 3.94 | 5.95×10–5 | ADTRP |
| rs7185541 | 16 | 47 222 182 | A | 0.64 | –0.32 | 3.82 | 6.20×10–5 | LOC105371240 & N4BP1 (21 kb) |
| rs1914037 | 2 | 189 553 760 | A | 0.75 | 0.35 | 3.71 | 6.57×10–5 | COL3A1 |
| rs28862711 | 19 | 32 470 668 | A | 0.61 | –0.31 | 3.58 | 6.78×10–5 | Only a pseudogene within 1 Mb |
| rs10420243 | 19 | 32 640 702 | A | 0.68 | –0.31 | 3.32 | 6.79×10–5 | Only a pseudogene within 1 Mb |
| rs2190798 | 19 | 33 141 651 | G | 0.75 | –0.36 | 3.93 | 7.37×10–5 | LOC102724694 |
| rs12659606 | 5 | 123 591 568 | A | 0.89 | –0.48 | 3.61 | 7.55×10–5 | ZNF608 (400 kb) |
| rs748731 | 3 | 133 175 696 | A | 0.82 | –0.38 | 3.52 | 7.68×10–5 | CPNE4 |
| rs3736487 | 2 | 189 564 188 | G | 0.76 | 0.35 | 3.56 | 7.71×10–5 | COL3A1 |
| rs1901163 | 5 | 165 909 270 | G | 0.82 | –0.40 | 3.62 | 7.81×10–5 | TENM2 (409 kb) |
| rs6584162 | 10 | 99 631 761 | A | 0.67 | 0.31 | 3.36 | 8.98×10–5 | CRTAC1 |
| rs10520872 | 5 | 21 709 737 | A | 0.86 | –0.44 | 3.66 | 9.13×10–5 | LOC105374685 & CDH12 (77 kb) |
| rs2158244 | 7 | 111 610 670 | G | 0.69 | 0.32 | 3.49 | 9.26×10–5 | DOCK4 |
| rs894417 | 9 | 131 497 934 | A | 0.90 | –0.49 | 3.31 | 9.40×10–5 | PRRX2 |
| rs7197966 | 16 | 6 086 332 | G | 0.59 | –0.30 | 3.58 | 9.51×10–5 | RBFOX1 |
| rs2593324 | 3 | 22 094 225 | G | 0.62 | –0.30 | 3.42 | 9.8×10–5 | ZNF385D |
| rs1889879 | 6 | 69 720 601 | A | 0.61 | –0.29 | 3.18 | 9.91×10–5 | BAI3 |
H2 is the proportion of total phenotypic variance explained by the SNP.
Positions are relative to Human Genome National Center for Biotechnology Information (NCBI) Build 36.3.
A positive β means that the modelled allele is associated with an increase in TG in response to exercise training, while a negative β reflects the converse.
The gene located nearest to the SNP. Distance to the gene in kilo bases (1000 bp) is shown in parentheses. If no distance is shown, the SNP is located within the gene locus.
Freq, allele frequency; GWAS, genome-wide association study; SNP, single nucleotide polymorphisms; TG, triglycerides.
The strongest evidence of association with ΔTG was detected at rs2396190 (p=3.3×10−7) located 90 kb from DOCK10 (2q36.2), followed by rs222158 (p=1.8×10−6) located in CYYR1 (21q21.2) (see online supplementary figure S3). In the final forward regression model, the top 10 SNPs explained 32.0% of the variance in ΔTG (table 3). CYYR1 rs222158 was the strongest predictor of ΔTG in the model, explaining 5.5% of the total variance, followed by GLT8D2 rs2722171, which explained 4.1% of the variance. As shown in the last column of table 3, the top four SNPs were shown to be sufficient to account for the genetic component of TG response to exercise training in white participants HERITAGE families. These four SNPs were retained for the ΔTG SNP summary score.
Table 3.
Results of the GWAS-based multivariate regression model with forward selection for TG response to exercise training in HERITAGE white participants (N=478)
| SNP | Chromosome | Position* | Gene† | MAF | Regression model |
Remaining Heritability‡ | ||
|---|---|---|---|---|---|---|---|---|
| Partial R2 | Model R2 | p Value | ||||||
| rs222158 | 21 | 26 794 032 | CYYR1 | 0.33 | 0.055 | 0.055 | 2.32×10–7 | 9.48% |
| rs2722171 | 12 | 102 973 617 | GLT8D2 | 0.20 | 0.041 | 0.097 | 4.70×10–6 | 6.4% |
| rs1906058 | 16 | 6 084 649 | RBFOX1 | 0.47 | 0.039 | 0.135 | 6.19×10–6 | 2.6% |
| rs2593324 | 3 | 22 094 225 | ZNF385D | 0.38 | 0.037 | 0.172 | 6.98×10–6 | 0% |
| rs12659606 | 5 | 123 591 568 | ZNF608 (400 kb) | 0.11 | 0.032 | 0.204 | 1.82×10–5 | NA |
| rs2190798 | 19 | 33 141 651 | LOC102724694 | 0.25 | 0.028 | 0.231 | 5.30×10–5 | NA |
| rs726553 | 2 | 225 724 738 | DOCK10 (100 kb) | 0.37 | 0.025 | 0.256 | 8.40×10–5 | NA |
| rs7850237 | 9 | 89 643 439 | SPATA31C1 (75 kb) | 0.14 | 0.027 | 0.283 | 3.66×10–5 | NA |
| rs9357234 | 6 | 37 138 195 | FGD2 (33 kb) | 0.31 | 0.020 | 0.303 | 0.0003 | NA |
| rs2646822 | 1 | 215 602 483 | GPATCH2 (69 kb) | 0.23 | 0.017 | 0.320 | 0.0007 | NA |
| rs3736487 | 2 | 189 564 188 | COL3A1 | 0.24 | 0.012 | 0.332 | 0.0038 | NA |
| rs13093483 | 3 | 68 376 923 | FAM19A1 | 0.12 | 0.012 | 0.344 | 0.0038 | NA |
| rs1889879 | 6 | 69 720 601 | BAI3 | 0.39 | 0.011 | 0.356 | 0.0046 | NA |
| rs10520872 | 5 | 21 709 737 | LOC105374685 | 0.14 | 0.010 | 0.366 | 0.0066 | NA |
| rs1452404 | 4 | 109 599 17 7 | LEF1 (290 kb) | 0.10 | 0.011 | 0.377 | 0.0053 | NA |
| rs11666431 | 19 | 2 904 087 | ZNF77 (9 kb) | 0.42 | 0.009 | 0.386 | 0.0103 | NA |
| rs3861882 | 9 | 131 505 125 | PRRX2 | 0.28 | 0.008 | 0.394 | 0.0143 | NA |
| rs9469986 | 6 | 11 857 166 | ADTRP | 0.20 | 0.008 | 0.401 | 0.0165 | NA |
| rs2158244 | 7 | 111 610 670 | DOCK4 | 0.31 | 0.007 | 0.408 | 0.0218 | NA |
| rs4742057 | 9 | 4 943 916 | JAK2 (32 kb) | 0.42 | 0.006 | 0.414 | 0.0333 | NA |
Positions are relative to Human Genome National Center for Biotechnology Information (NCBI) Build 36.3.
The gene located nearest to the SNP. Distance to the gene in kilo bases (1000 bp) is shown in parentheses. If no distance is shown, the SNP is located within the gene locus.
Remaining heritability estimate when a given SNP (plus preceding SNPs) is included as covariate(s) in the MERLIN heritability model.
GWAS, genome-wide association study; MAF, minor allele frequency; NA, not applicable; SNP, single nucleotide polymorphisms; TG, triglycerides.
Pathway analysis of GWAS associations
The results of the pathway analysis using the second best p value and Stouffer's methods can be found in online supplementary tables S2–S4. Briefly, the glycosphingolipid biosynthesis gene set was enriched using both approaches. Glycosphingolipid biosynthesis-related gene sets were the second, fifth and sixth most enriched positive gene sets using the second best p value method (false discovery rate (FDR): 0.07–0.32), while it was the third ranked gene set using Stouffer's method (FDR=0.30). The most enriched positive gene set using the second best p value method was the heparan sulfate glycosaminoglycan bio-synthesis gene set (FDR=0.097), while cell adhesion molecules were the most enriched negative gene set (FDR=0.12).
RNA expression-based gene signature of TG response to exercise training
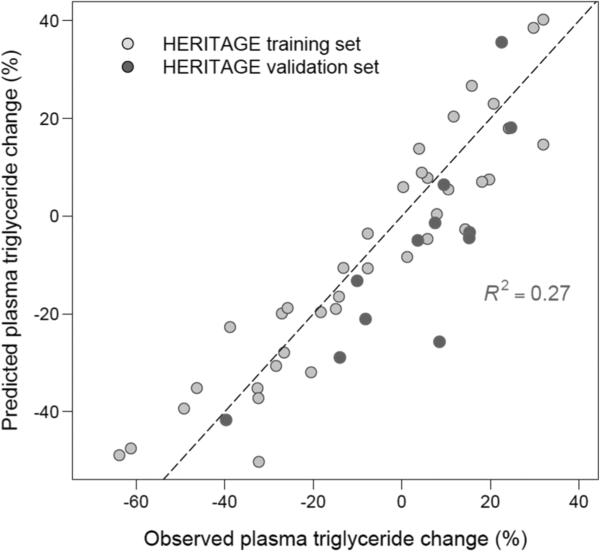
By means of forward stepwise regression, ranking Affymetrix probesets by their selection frequency (high to low) in the different GA-derived predictor models, an 11-gene linear regression model was developed (table 4). This model was able to explain 80% of the variance in the training set (F value=13.2, p<0.0001). We then evaluated the model performance on the remaining 12 HERITAGE samples that were omitted from the GA search procedure. As shown in figure 1, the predictor model was able to explain 27% of the variance. Importantly, the test set spanned a broad range (−40 to +20%) in terms of the training-induced changes in TG.
Table 4.
Results of the RNA-based multivariate regression model with forward selection for TG response to exercise training in HERITAGE white participants (N=37)
| Variable | β | SE | t Value | p Value |
|---|---|---|---|---|
| Sex | 0.21 | 0.17 | 1.1 | 0.27 |
| DYX1C1 | 0.59 | 0.11 | 5.6 | 9.9×10–6 |
| ZNF30 | –0.01 | 0.12 | –0.09 | 0.93 |
| BTG2 | 0.21 | 0.12 | 1.7 | 0.10 |
| MACROD1 | 0.10 | 0.20 | 0.5 | 0.62 |
| UBE2L3 | 0.16 | 0.15 | 1.1 | 0.29 |
| C21orf88 | –0.14 | 0.13 | –1.1 | 0.28 |
| EEF2K | 0.21 | 0.10 | 2.1 | 0.05 |
| NCBP2 | 0.22 | 0.16 | 1.5 | 0.15 |
| FASTK | 0.39 | 0.15 | 2.5 | 0.02 |
| C2orf69 | –0.03 | 0.12 | –0.3 | 0.80 |
| NSA2 | –0.28 | 0.10 | –2.8 | 0.009 |
Model R2=0.80, p=8.6×10–8.
Figure 1.
Performance of the RNA-based regression model derived from the training set (N=37, grey dots) in the test set (N=12, red dots) for the prediction of exercise training-induced changes in triglycerides in HERITAGE.
In order to examine the general applicability of the molecular signature, we took advantage of a previously published Affymetrix gene expression data set from an independent exercise-training cohort.27 Since the gene-chip technology significantly differed between cohorts (ie, 3′-based vs the newer whole transcript-based methodologies), we performed all possible subsets regression. We found that the model containing six genes (BTG2, C2orf69, C21orf88, DYX1C1, NSA2, UBE2L3) performed the best, as this model (F value=3.7, p=0.03) explained 48% of the variance in TG changes, while also having the lowest Bayesian Information Criterion (BIC) score (see online supplementary figure S4). Further, a resampling procedure in which 10 000 random multivariate models were developed confirmed the significance of the six gene model in terms of R2 performance (data not shown).
The association of SNPs (N=498), in or near (±20 kb) the 11 predictor genes, with ΔTG was tested in all HERITAGE white participants (N=481) using imputed SNPs (data not shown). Only SNPs from four genes (NSA2, FASTK, MACROD1 and EEF2 K) showed nominal (p<0.05) associations with ΔTG (see online supplementary tables S5–S8). The top associated SNP (p≤0.01) from each of the four genes (rs1043968, rs3793336, rs594461, rs11646610) was used in the SNP summary score.
Pathway analysis of mRNA predictor models
We evaluated the enrichment of pathways among genes in the subset of 512 predictive models using Ingenuity Pathways Analysis. We found that pathways related to mitochondrial dysfunction and oxidative phosphorylation were enriched (p<0.0003) (see online supplementary figure S5).
Association of SNP score and ΔTG
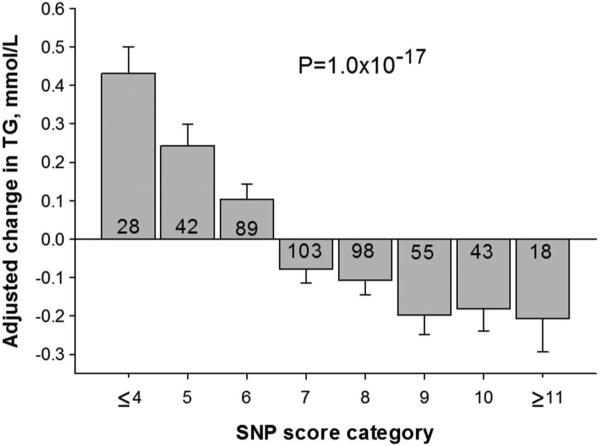
The SNP score was created by combining the four top SNPs from the GWAS analysis (rs222158, rs2722171, rs1906058, rs2593324) and the four top SNPs from the targeted SNP analysis of the RNA predictor genes (rs1043968, rs3793336, rs594461, rs11646610). The SNP score values ranged from 2 to 12 in HERITAGE white participants. The adjusted mean decrease in TG in participants with 11 or 12 favourable alleles (N=18) was −18.2 mg/dL (−0.21 mmol/L), while those with 4 or less favourable alleles (N=284) experienced an adjusted mean increase of 38.1 mg/dL (0.43 mmol/L) after exercise training (figure 2).
Figure 2.
Adjusted mean response to exercise training triglycerides (ΔTG) across eight single nucleotide polymorphisms (SNP) summary score categories in HERITAGE white participants. Values were adjusted for age, sex, baseline body mass index and baseline triglyceride level. Number of participants within each SNP score category is indicated inside each histogram bar.
After backwards elimination of over 50 baseline variables related to blood pressure, body composition, fitness, lipids and lipoproteins, and insulin and glucose metabolism, a forward regression model for ΔTG showed that the SNP score was the strongest predictor variable (p=3.0×10−68), explaining 14.4% of the variance in ΔTG (approximately 8% from 4 GWAS SNPs, approximately 6% from 4 RNA predictor SNPs), while baseline TG explained 7.4% of the variance in responsiveness (table 5).
Table 5.
Final forward regression model, after backwards elimination of over 50 baseline variables, for predictors of exercise-induced changes in TG in HERITAGE white participants
| ΔTG predictors | Partial R2 | Model R2 | p Value |
|---|---|---|---|
| SNP summary score | 0.1435 | 0.1435 | 3.0×10–68 |
| Baseline TG (mmol/L) | 0.0742 | 0.2177 | 5.1×10–10 |
| Baseline large LDL-P (nmol/L) | 0.0167 | 0.2344 | 0.002 |
| Baseline fasting insulin (pmol/L) | 0.0078 | 0.2422 | 0.037 |
| Baseline AIRg (pmol/l×40 min) | 0.009 | 0.2512 | 0.025 |
AIRg, acute insulin response to glucose, defined as the integrated area under the insulin curve between 0 and 40 min of the intravenous glucose tolerance test. In addition to age and sex, backwards elimination models included baseline variables related to blood pressure, body composition, fitness, lipids and lipoproteins, and insulin and glucose metabolism; LDL-P, low-density lipoprotein particle; SNP, single nucleotide polymorphisms; ΔTG, response to exercise training triglycerides.
DISCUSSION
Our results indicate that multiple genes and sequence variants contribute to the genetic and transcriptomic variation in the response of TG to exercise training. A novel finding of this study is that a small number of SNPs (N=4) accounted for all of the genetic variance of TG response to exercise training, as quantified in white participants from HERITAGE. Furthermore, we found that a molecular signature based on the baseline expression levels of 11 genes predicted 27% of TG exercise response in HERITAGE, which was validated in an independent study. Lastly, our SNP summary score results provide an example of how parsimonious panels of associated SNPs from multiple omics platforms could potentially be used as a priori predictors of a trait response to regular exercise, so that more effective means of treatment and prevention can be identified. If validated in independent studies, this information could potentially be used to individually tailor exercise training programmes targeted at managing TG levels.
The pathway analysis of our GWAS results revealed that the genetic effect of TG response to regular exercise may be exerted through pathways related to heparan sulfate glycosaminoglycan and glycosphingolipid biosynthesis and cell-adhesion molecules. Heparan sulfate has several biological functions, including cell adhesion and notably cell surface binding of LPL. Several studies have shown that heparan sulfate glycosaminoglycan modified proteoglycans act in the hydrolysis of TG-rich lipoproteins.34 In HERITAGE, LPL activity significantly increased with exercise training, concomitant with decreases in TG levels (r=−0.21, p<0.001). Glycosphingolipids mediate and modulate intercellular coordination in multicellular organisms.35 They cluster in lipid rafts, which are enriched in cholesterol and sphingolipids such as sphingomyelin. Lipid rafts are involved in many cellular processes, including membrane sorting and trafficking, cell polarisation and signal transduction processes.36 As such, the specialised structure of lipid rafts may play a critical role in the trafficking of lipids between lipoproteins and cells. However, it is unknown how exercise affects lipid rafts and glycosphingolipids.
The pathway analysis of the transcriptomic predictor models revealed that mitochondrial dysfunction and oxidative phosphorylation pathways were enriched in relation to TG response to exercise training. Defects in mitochondrial oxidation and phosphorylation are associated with insulin resistance, type 2 diabetes and ectopic TG accumulation (eg, intramyocellular and intrahepatic lipids), among others.37–39 Exercise is known to improve insulin sensitivity and risk of diabetes40,41 and has also been shown to increase intramyocellular lipid (IMCL) content, which leads to a higher proportion of IMCL in direct contact with mitochondria and improved lipid flux.42,43
In summary, our bioinformatics analysis suggests that different subsets of pathways are enriched when variants from genomic and transcriptomic data are interrogated for their association with TG response to exercise. Interestingly, we did not find any associations between the existing TG GWAS loci33 and the response of TG to regular exercise. Thus, our results suggest that the genes most important in modifying changes in TG in response to regular exercise are most likely different from the loci contributing to variation in population TG levels. It is not clear how many of the markers and genes identified here are functionally related to the response to regular exercise of TG, but, if confirmed in independent studies, they could inform on the potential mechanisms involved in TG changes with regular exercise. Irrespective of the biological plausibility of the identi-fied variants, identifying predictors of favourable or unfavourable TG response to regular exercise could help optimise therapeutic strategies.
This study is based on a relatively small sample size compared to commonly accepted standards of GWAS. Thus, it is not surprising that we did not find SNPs reaching genome-wide significance. However, it is important to appreciate that human experimental studies are by definition characterised by much smaller sample sizes than epidemiological and observational studies, but are also less likely to be negatively impacted by any number of uncontrolled confounders. HERITAGE remains the largest fully controlled exercise intervention study thus far. The family structure and well defined and twice measured phenotypes help to minimise the influence of confounding factors. Moreover, since one component of the environment (ie, exercise) has been rigorously controlled in HERITAGE, we had predicted larger effect sizes than is commonly seen in observational studies. Thus, we concluded that it would be useful to undertake hypothesis free and unbiased GWAS explorations for the response of TG to regular exercise, as it could generate hypotheses and candidate genes deserving further research. Furthermore, our global microarray data from a subset of HERITAGE participants complements our GWAS results and provides a separate unbiased exploration of the genes potentially involved in TG exercise response.
We acknowledge that there is still a possibility of false discovery and that the parsimonious SNP score associated with ΔTG in HERITAGE is likely to overfit our data, as the score was tested on the same data set used to create it, possibly resulting in biased conclusions about the strength of the findings. To somewhat address this limitation, in a proof-of-concept analysis we reran the GWAS in half the sample (49 randomly selected families) and tested the top resulting SNPs in the other half of the sample (remaining 49 families). Of the top SNPs (p<1×10−4, N=30), only the association with CYYR1 rs222158 (p=2.9×10−5) was replicated, as it explained 3.2% of the variance in ΔTG (p=0.005) in the second half of the sample. Interestingly, CYYR1 rs222158 was also the top predictor SNP in the regression models in the whole cohort. Thus, although the splitting of the sample in half decreases the sample size, the separate discovery and replication data sets provide additional support for the results found in the larger cohort. There is an obvious need to replicate our results in other samples and studies.
This study suggests that regular exercise may not influence TG levels at the same rate for all individuals and that the efficacy of regular exercise could be related in part to multiple genetic variants. Identifying the genes underlying the variability in lipid traits due to regular exercise would significantly contribute to the biology of adaptation to exercise and the development of an exercise component of personalised preventive and therapeutic medicine. The ability to identify individuals whose TG levels are likely to positively respond to lifestyle interventions could help optimise prevention and treatment strategies designed to decrease the risk of hypertriglyceridaemia.
Supplementary Material
What are the findings?
▶ A genome-wide association study identified four SNPs that accounted for the genetic variance of triglyceride (TG) response to exercise training in HERITAGE white participants.
▶ Analysis of baseline RNA expression levels identified a subset of 11 genes that significantly predicted TG exercise response in HERITAGE white participants, which was validated in an independent cohort.
▶ Integrating data from genomic and transcriptomic analyses produced a strong predictor of TG response to regular exercise.
How might it impact on clinical practice in the future?
▶ Provides potential genetic markers related to the ability to improve plasma triglyceride levels with endurance exercise training.
▶ Further research is needed to replicate and further test these findings in independent studies and other populations and exercise programmes.
▶ If validated in independent studies, this information could potentially be used to individually tailor exercise training programmes targeted at managing triglyceride levels.
Acknowledgements
The authors would like to thank Dr Arthur Leon, Dr James Skinner and the late Dr Jack Wilmore for their contributions to the prior phases of the HERITAGE Family Study. They also thank Jessica Watkins for her expert contributions to GWAS genotyping and DNA bank maintenance.
Funding This work was supported by the AHA Greater Southeast Affiliate Postdoctoral Fellowship number 10POST3670006 (MS), the NIH/NIGMS Center of Biomedical Research Excellence (COBRE) program award P20 GM103528 (MS), and the NIH/NIGMS Louisiana Clinical and Translational Science Center 1 U54 GM104940 (MS). The HERITAGE Family Study is supported by the National Heart, Lung and Blood Institute grant HL-45670 (CB). CB is partially supported by the JWBarton, Sr. Endowed Chair in Genetics and Nutrition.
Footnotes
▶ Additional material is published online only. To view please visit the journal online (http://dx.doi.org/10.1136/bjsports-2015-095179).
To cite: Sarzynski MA, Davidsen PK, Sung YJ, et al. Br J Sports Med 2015;49: 1524–1531.
Contributors MKCH, PS, DCR and CB participated in the design of their respective exercise training studies. MAS, TKR, DCR and CB contributed to the GWAS analysis. PKD and FF conducted the transcriptomic and pathway analyses. MAS and PKD drafted the manuscript, while all authors contributed to its critical revision for important intellectual content.
Correction notice This paper has been amended since it was published Online First. A sentence to the methods section has been added. After the sentence “Affymetrix HG-U133+2 arrays were used to quantitate global gene expression levels” the following sentence and reference 23a have been added: “Raw microarray data have been deposited with Gene Expression Omnibus under accession number: GSE47874.”
Competing interests MAS is an advisor for Genetic Direction, Dallas, Texas, USA.
Ethics approval Institutional Review Boards (IRB).
Provenance and peer review Not commissioned; externally peer reviewed.
REFERENCES
- 1.Miller M, Stone NJ, Ballantyne C, et al. Triglycerides and cardiovascular disease: a scientific statement from the American Heart Association. Circulation. 2011;123:2292–333. doi: 10.1161/CIR.0b013e3182160726. [DOI] [PubMed] [Google Scholar]
- 2.Nordestgaard BG, Varbo A. Triglycerides and cardiovascular disease. Lancet. 2014;384:626–35. doi: 10.1016/S0140-6736(14)61177-6. [DOI] [PubMed] [Google Scholar]
- 3.Hokanson JE, Austin MA. Plasma triglyceride level is a risk factor for cardiovascular disease independent of high-density lipoprotein cholesterol level: a meta-analysis of population-based prospective studies. J Cardiovasc Risk. 1996;3:213–19. [PubMed] [Google Scholar]
- 4.Sarwar N, Danesh J, Eiriksdottir G, et al. Triglycerides and the risk of coronary heart disease: 10,158 incident cases among 262,525 participants in 29 Western prospective studies. Circulation. 2007;115:450–8. doi: 10.1161/CIRCULATIONAHA.106.637793. [DOI] [PubMed] [Google Scholar]
- 5.Jorgensen AB, Frikke-Schmidt R, West AS, et al. Genetically elevated non-fasting triglycerides and calculated remnant cholesterol as causal risk factors for myocardial infarction. Eur Heart J. 2013;34:1826–33. doi: 10.1093/eurheartj/ehs431. [DOI] [PubMed] [Google Scholar]
- 6.Thomsen M, Varbo A, Tybjaerg-Hansen A, et al. Low nonfasting triglycerides and reduced all-cause mortality: a mendelian randomization study. Clin Chem. 2014;60:737–46. doi: 10.1373/clinchem.2013.219881. [DOI] [PubMed] [Google Scholar]
- 7.National Cholesterol Education Program Third Report of the National Cholesterol Education Program (NCEP) Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults (Adult Treatment Panel III) final report. Circulation. 2002;106:3143–421. [PubMed] [Google Scholar]
- 8.Berglund L, Brunzell JD, Goldberg AC, et al. Evaluation and treatment of hypertriglyceridemia: an Endocrine Society clinical practice guideline. J Clin Endocrinol Metab. 2012;97:2969–89. doi: 10.1210/jc.2011-3213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Durstine JL, Grandjean PW, Davis PG, et al. Blood lipid and lipoprotein adaptations to exercise: a quantitative analysis. Sports Med. 2001;31:1033–62. doi: 10.2165/00007256-200131150-00002. [DOI] [PubMed] [Google Scholar]
- 10.Bouchard C, Blair SN, Church TS, et al. Adverse metabolic response to regular exercise: is it a rare or common occurrence? PLoS ONE. 2012;7:e37887. doi: 10.1371/journal.pone.0037887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bouchard C, Antunes-Correa LM, Ashley EA, et al. Personalized preventive medicine: genetics and the response to regular exercise in preventive interventions. Prog Cardiovasc Dis. 2015;57:337–46. doi: 10.1016/j.pcad.2014.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rice T, Despres JP, Perusse L, et al. Familial aggregation of blood lipid response to exercise training in the health, risk factors, exercise training, and genetics (HERITAGE) Family Study. Circulation. 2002;105:1904–8. doi: 10.1161/01.cir.0000014969.85364.9f. [DOI] [PubMed] [Google Scholar]
- 13.Bray MS, Hagberg JM, Perusse L, et al. The human gene map for performance and health-related fitness phenotypes: the 2006–2007 update. Med Sci Sports Exerc. 2009;41:35–73. doi: 10.1249/mss.0b013e3181844179. [DOI] [PubMed] [Google Scholar]
- 14.Huggins GS, Papandonatos GD, Erar B, et al. Do genetic modifiers of high-density lipoprotein cholesterol and triglyceride levels also modify their response to a lifestyle intervention in the setting of obesity and type-2 diabetes mellitus?: the Action for Health in Diabetes (Look AHEAD) study. Circ Cardiovasc Genet. 2013;6:391–9. doi: 10.1161/CIRCGENETICS.113.000042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Timmons JA, Knudsen S, Rankinen T, et al. Using molecular classification to predict gains in maximal aerobic capacity following endurance exercise training in humans. J Appl Physiol. 2010;108:1487–96. doi: 10.1152/japplphysiol.01295.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bouchard C, Leon AS, Rao DC, et al. The HERITAGE family study. Aims, design, and measurement protocol. Med Sci Sports Exerc. 1995;27:721–9. [PubMed] [Google Scholar]
- 17.Despres JP, Gagnon J, Bergeron J, et al. Plasma post-heparin lipase activities in the HERITAGE Family Study: the reproducibility, gender differences, and associations with lipoprotein levels. HEalth, RIsk factors, exercise Training and GEnetics. Clin Biochem. 1999;32:157–65. doi: 10.1016/s0009-9120(98)00106-4. [DOI] [PubMed] [Google Scholar]
- 18.Bouchard C, Rankinen T, Chagnon YC, et al. Genomic scan for maximal oxygen uptake and its response to training in the HERITAGE Family Study. J Appl Physiol. 2000;88:551–9. doi: 10.1152/jappl.2000.88.2.551. [DOI] [PubMed] [Google Scholar]
- 19.Abecasis GR, Cherny SS, Cookson WO, et al. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet. 2002;30:97–101. doi: 10.1038/ng786. [DOI] [PubMed] [Google Scholar]
- 20.Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mootha VK, Lindgren CM, Eriksson KF, et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–73. doi: 10.1038/ng1180. [DOI] [PubMed] [Google Scholar]
- 22.Ghosh S, Vivar JC, Sarzynski MA, et al. Integrative pathway analysis of a genome-wide association study of (V)O(2max) response to exercise training. J Appl Physiol (1985) 2013;115:1343–59. doi: 10.1152/japplphysiol.01487.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Stouffer SA, Suchman EA, DeVinney LC, et al. The American soldier, Vol 1: adjustment during Army Life. Princeton University Press; Princeton: 1949. [Google Scholar]
- 23a.Phillips BE, Williams JP, Gustafsson T, et al. Molecular networks of human muscle adaptation to exercise and age. PLoS Genet. 2013;9:e1003389. doi: 10.1371/journal.pgen.1003389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Cook RD. Detection of influential observations in linear regression. Technometrics. 1977;19:15–18. [Google Scholar]
- 25.Toth PP, Barter PJ, Rosenson RS, et al. High-density lipoproteins: a consensus statement from the National Lipid Association. J Clin Lipidol. 2013;7:484–525. doi: 10.1016/j.jacl.2013.08.001. [DOI] [PubMed] [Google Scholar]
- 26.Catoire M, Mensink M, Kalkhoven E, et al. Identification of human exercise-induced myokines using secretome analysis. Physiol Genomics. 2014;46:256–67. doi: 10.1152/physiolgenomics.00174.2013. [DOI] [PubMed] [Google Scholar]
- 27.Meex RC, Schrauwen-Hinderling VB, Moonen-Kornips E, et al. Restoration of muscle mitochondrial function and metabolic flexibility in type 2 diabetes by exercise training is paralleled by increased myocellular fat storage and improved insulin sensitivity. Diabetes. 2010;59:572–9. doi: 10.2337/db09-1322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Y, Willer CJ, Ding J, et al. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol. 2010;34:816–34. doi: 10.1002/gepi.20533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sung YJ, Wang L, Rankinen T, et al. Performance of genotype imputations using data from the 1000 Genomes Project. Hum Hered. 2012;73:18–25. doi: 10.1159/000334084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Leon AS, Rice T, Mandel S, et al. Blood lipid response to 20 weeks of supervised exercise in a large biracial population: the HERITAGE Family Study. Metabolism. 2000;49:513–20. doi: 10.1016/s0026-0495(00)80018-9. [DOI] [PubMed] [Google Scholar]
- 31.Leon AS, Gaskill SE, Rice T, et al. Variability in the response of HDL cholesterol to exercise training in the HERITAGE Family Study. Int J Sports Med. 2002;23:1–9. doi: 10.1055/s-2002-19270. [DOI] [PubMed] [Google Scholar]
- 32.Bergeron J, Couillard C, Despres JP, et al. Race differences in the response of postheparin plasma lipoprotein lipase and hepatic lipase activities to endurance exercise training in men: results from the HERITAGE Family Study. Atherosclerosis. 2001;159:399–406. doi: 10.1016/s0021-9150(01)00515-9. [DOI] [PubMed] [Google Scholar]
- 33.Willer CJ, Schmidt EM, Sengupta S, et al. Discovery and refinement of loci associated with lipid levels. Nat Genet. 2013;45:1274–83. doi: 10.1038/ng.2797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhang L, editor. Glycosaminoglycans in Development, Health, and Disease. Academic Press; San Diego: 2010. [DOI] [PubMed] [Google Scholar]
- 35.Schnaar RL, Suzuki A, Stanley P. Glycosphingolipids. In: Cummings RD, Esko JD, Freeze HH, et al., editors. Essentials of Gkycobiology. 2nd edn. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY: 2009. pp. 129–143. [PubMed] [Google Scholar]
- 36.Simons K, Ehehalt R. Cholesterol, lipid rafts, and disease. J Clin Invest. 2002;110:597–603. doi: 10.1172/JCI16390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Snel M, Jonker JT, Schoones J, et al. Ectopic fat and insulin resistance: pathophysiology and effect of diet and lifestyle interventions. Int J Endocrinol. 2012;2012:983814. doi: 10.1155/2012/983814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Roden M. Muscle triglycerides and mitochondrial function: possible mechanisms for the development of type 2 diabetes. Int J Obes (Lond) 2005;29(Suppl 2):S111–15. doi: 10.1038/sj.ijo.0803102. [DOI] [PubMed] [Google Scholar]
- 39.Vankoningsloo S, Piens M, Lecocq C, et al. Mitochondrial dysfunction induces triglyceride accumulation in 3T3-L1 cells: role of fatty acid beta-oxidation and glucose. J Lipid Res. 2005;46:1133–49. doi: 10.1194/jlr.M400464-JLR200. [DOI] [PubMed] [Google Scholar]
- 40.Borghouts LB, Keizer HA. Exercise and insulin sensitivity: a review. Int J Sports Med. 2000;21:1–12. doi: 10.1055/s-2000-8847. [DOI] [PubMed] [Google Scholar]
- 41.Knowler WC, Barrett-Connor E, Fowler SE, et al. Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med. 2002;346:393–403. doi: 10.1056/NEJMoa012512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tarnopolsky MA, Rennie CD, Robertshaw HA, et al. Influence of endurance exercise training and sex on intramyocellular lipid and mitochondrial ultrastructure, substrate use, and mitochondrial enzyme activity. Am J Physiol Regul Integr Comp Physiol. 2007;292:R1271–8. doi: 10.1152/ajpregu.00472.2006. [DOI] [PubMed] [Google Scholar]
- 43.Corcoran MP, Lamon-Fava S, Fielding RA. Skeletal muscle lipid deposition and insulin resistance: effect of dietary fatty acids and exercise. Am J Clin Nutr. 2007;85:662–77. doi: 10.1093/ajcn/85.3.662. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.