Abstract
The AC40 and AC19 subunits (encoded by RPC40 and RPC19) are shared by yeast RNA polymerases I and III and have a local sequence similarity to prokaryotic alpha subunits. Mutational analysis of the corresponding "alpha motif" indicated that its integrity is essential on AC40 subunit but is not essential on AC19 subunit. By applying the two-hybrid method, these two polypeptides were shown to associate in vivo. Extragenic suppression of rpc19 and rpc40 mutations confirmed that AC19 and AC40 subunits interact with each other in vivo and revealed an interaction with ABC10 beta subunit [encoded by RPB10; Woychick, N. A. & Young, R.A. (1990) J. Biol. Chem. 265, 17816-17819], one of the five polypeptides common to all three nuclear RNA polymerases. A correction of the RPB10 sequence showed that ABC10 beta subunit is a 70-amino acid polypeptide, as confirmed by peptide microsequencing. These results suggest that the assembly of RNA polymerase I and III requires the association of ABC10 beta subunit with an AC19/AC40 heterodimer.
Full text
PDF
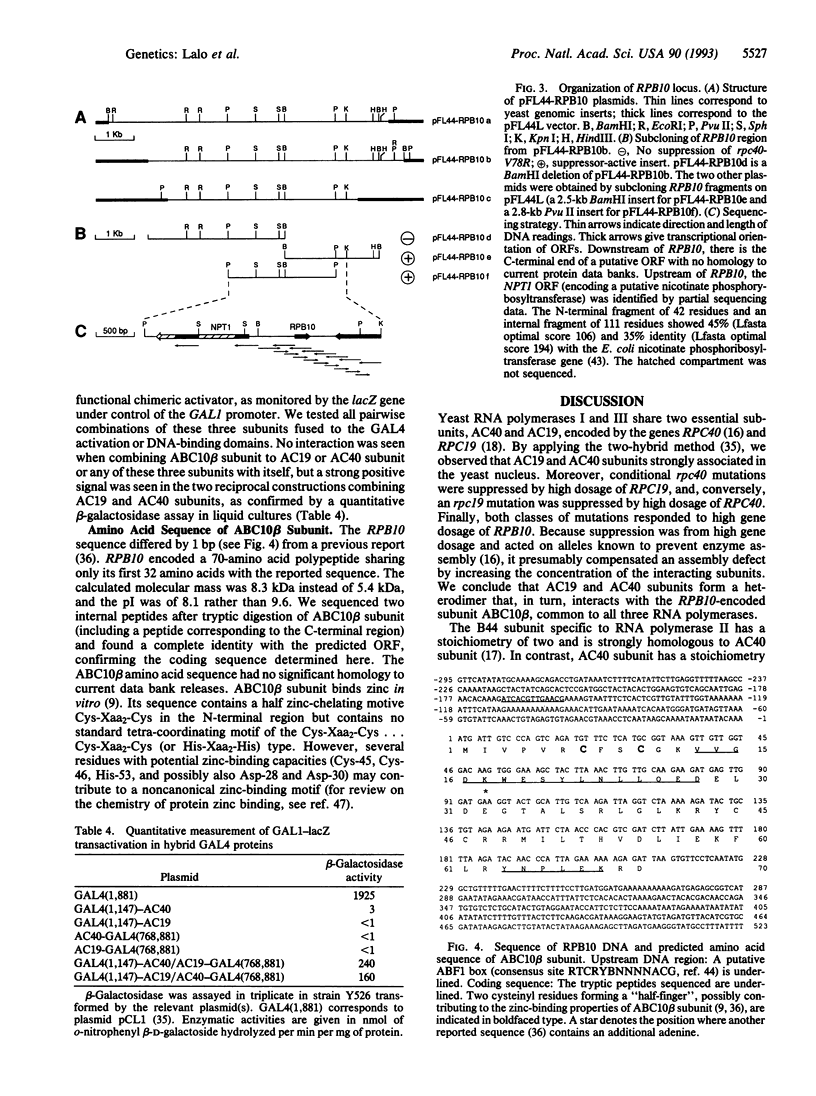

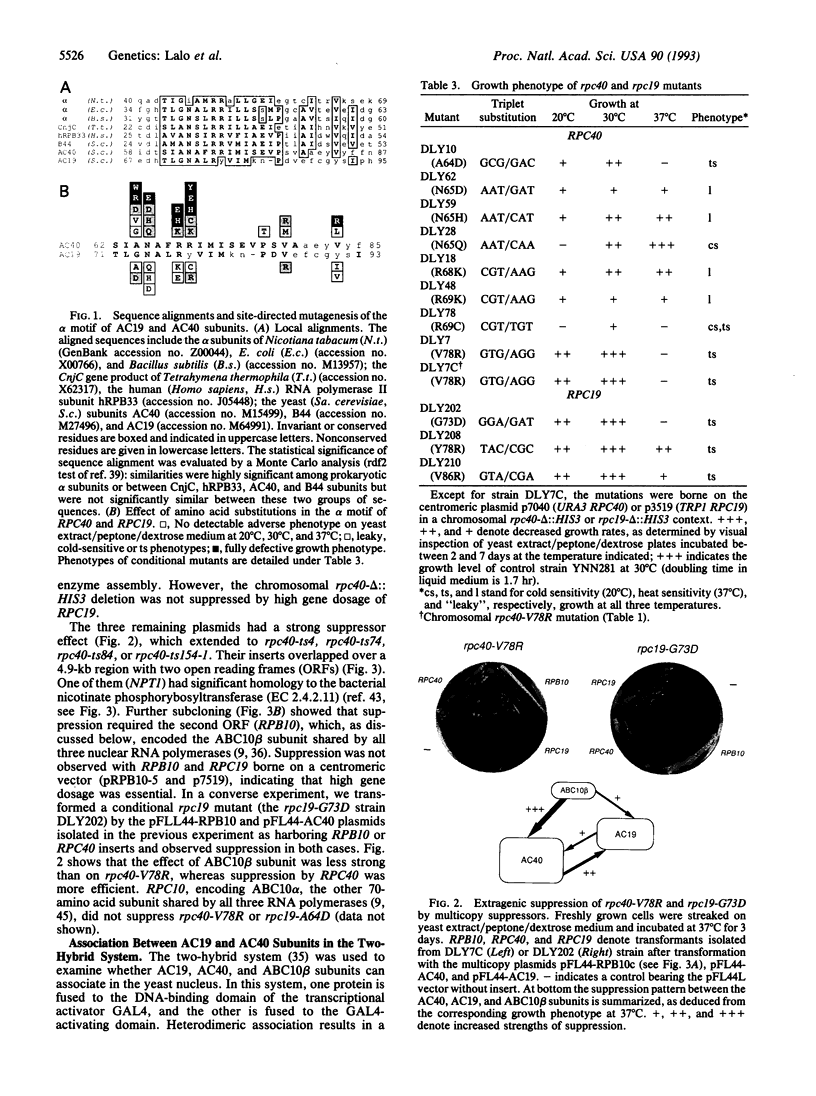


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allison L. A., Moyle M., Shales M., Ingles C. J. Extensive homology among the largest subunits of eukaryotic and prokaryotic RNA polymerases. Cell. 1985 Sep;42(2):599–610. doi: 10.1016/0092-8674(85)90117-5. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Burgess R. R. Separation and characterization of the subunits of ribonucleic acid polymerase. J Biol Chem. 1969 Nov 25;244(22):6168–6176. [PubMed] [Google Scholar]
- Carles C., Treich I., Bouet F., Riva M., Sentenac A. Two additional common subunits, ABC10 alpha and ABC10 beta, are shared by yeast RNA polymerases. J Biol Chem. 1991 Dec 15;266(35):24092–24096. [PubMed] [Google Scholar]
- Chiannilkulchai N., Stalder R., Riva M., Carles C., Werner M., Sentenac A. RPC82 encodes the highly conserved, third-largest subunit of RNA polymerase C (III) from Saccharomyces cerevisiae. Mol Cell Biol. 1992 Oct;12(10):4433–4440. doi: 10.1128/mcb.12.10.4433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chien C. T., Bartel P. L., Sternglanz R., Fields S. The two-hybrid system: a method to identify and clone genes for proteins that interact with a protein of interest. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9578–9582. doi: 10.1073/pnas.88.21.9578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corden J. L., Cadena D. L., Ahearn J. M., Jr, Dahmus M. E. A unique structure at the carboxyl terminus of the largest subunit of eukaryotic RNA polymerase II. Proc Natl Acad Sci U S A. 1985 Dec;82(23):7934–7938. doi: 10.1073/pnas.82.23.7934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Della Seta F., Treich I., Buhler J. M., Sentenac A. ABF1 binding sites in yeast RNA polymerase genes. J Biol Chem. 1990 Sep 5;265(25):15168–15175. [PubMed] [Google Scholar]
- Dequard-Chablat M., Riva M., Carles C., Sentenac A. RPC19, the gene for a subunit common to yeast RNA polymerases A (I) and C (III). J Biol Chem. 1991 Aug 15;266(23):15300–15307. [PubMed] [Google Scholar]
- Elledge S. J., Davis R. W. A family of versatile centromeric vectors designed for use in the sectoring-shuffle mutagenesis assay in Saccharomyces cerevisiae. Gene. 1988 Oct 30;70(2):303–312. doi: 10.1016/0378-1119(88)90202-8. [DOI] [PubMed] [Google Scholar]
- Faust D. M., Renkawitz-Pohl R., Falkenburg D., Gasch A., Bialojan S., Young R. A., Bautz E. K. Cloning and identification of the gene coding for the 140-kd subunit of Drosophila RNA polymerase II. EMBO J. 1986 Apr;5(4):741–746. doi: 10.1002/j.1460-2075.1986.tb04276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fields S., Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989 Jul 20;340(6230):245–246. doi: 10.1038/340245a0. [DOI] [PubMed] [Google Scholar]
- Fleig U. N., Pridmore R. D., Philippsen P. Construction of LYS2 cartridges for use in genetic manipulations of Saccharomyces cerevisiae. Gene. 1986;46(2-3):237–245. doi: 10.1016/0378-1119(86)90408-7. [DOI] [PubMed] [Google Scholar]
- Grachev M. A., Kolocheva T. I., Lukhtanov E. A., Mustaev A. A. Studies on the functional topography of Escherichia coli RNA polymerase. Highly selective affinity labelling by analogues of initiating substrates. Eur J Biochem. 1987 Feb 16;163(1):113–121. doi: 10.1111/j.1432-1033.1987.tb10743.x. [DOI] [PubMed] [Google Scholar]
- Hayward R. S., Igarashi K., Ishihama A. Functional specialization within the alpha-subunit of Escherichia coli RNA polymerase. J Mol Biol. 1991 Sep 5;221(1):23–29. doi: 10.1016/0022-2836(91)80197-3. [DOI] [PubMed] [Google Scholar]
- Hieter P., Mann C., Snyder M., Davis R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell. 1985 Feb;40(2):381–392. doi: 10.1016/0092-8674(85)90152-7. [DOI] [PubMed] [Google Scholar]
- Huet J., Riva M., Sentenac A., Fromageot P. Yeast RNA polymerase C and its subunits. Specific antibodies as structural and functional probes. J Biol Chem. 1985 Dec 5;260(28):15304–15310. [PubMed] [Google Scholar]
- Igarashi K., Fujita N., Ishihama A. Sequence analysis of two temperature-sensitive mutations in the alpha subunit gene (rpoA) of Escherichia coli RNA polymerase. Nucleic Acids Res. 1990 Oct 25;18(20):5945–5948. doi: 10.1093/nar/18.20.5945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishihama A., Fujita N., Glass R. E. Subunit assembly and metabolic stability of E. coli RNA polymerase. Proteins. 1987;2(1):42–53. doi: 10.1002/prot.340020106. [DOI] [PubMed] [Google Scholar]
- Kolodziej P. A., Young R. A. Mutations in the three largest subunits of yeast RNA polymerase II that affect enzyme assembly. Mol Cell Biol. 1991 Sep;11(9):4669–4678. doi: 10.1128/mcb.11.9.4669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kolodziej P., Young R. A. RNA polymerase II subunit RPB3 is an essential component of the mRNA transcription apparatus. Mol Cell Biol. 1989 Dec;9(12):5387–5394. doi: 10.1128/mcb.9.12.5387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Lacroute F. Regulation of pyrimidine biosynthesis in Saccharomyces cerevisiae. J Bacteriol. 1968 Mar;95(3):824–832. doi: 10.1128/jb.95.3.824-832.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liljelund P., Mariotte S., Buhler J. M., Sentenac A. Characterization and mutagenesis of the gene encoding the A49 subunit of RNA polymerase A in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1992 Oct 1;89(19):9302–9305. doi: 10.1073/pnas.89.19.9302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma J., Ptashne M. A new class of yeast transcriptional activators. Cell. 1987 Oct 9;51(1):113–119. doi: 10.1016/0092-8674(87)90015-8. [DOI] [PubMed] [Google Scholar]
- Mann C., Buhler J. M., Treich I., Sentenac A. RPC40, a unique gene for a subunit shared between yeast RNA polymerases A and C. Cell. 1987 Feb 27;48(4):627–637. doi: 10.1016/0092-8674(87)90241-8. [DOI] [PubMed] [Google Scholar]
- Mann C., Micouin J. Y., Chiannilkulchai N., Treich I., Buhler J. M., Sentenac A. RPC53 encodes a subunit of Saccharomyces cerevisiae RNA polymerase C (III) whose inactivation leads to a predominantly G1 arrest. Mol Cell Biol. 1992 Oct;12(10):4314–4326. doi: 10.1128/mcb.12.10.4314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marck C. 'DNA Strider': a 'C' program for the fast analysis of DNA and protein sequences on the Apple Macintosh family of computers. Nucleic Acids Res. 1988 Mar 11;16(5):1829–1836. doi: 10.1093/nar/16.5.1829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martindale D. W. A conjugation-specific gene (cnjC) from Tetrahymena encodes a protein homologous to yeast RNA polymerase subunits (RPB3, RPC40) and similar to a portion of the prokaryotic RNA polymerase alpha subunit (rpoA). Nucleic Acids Res. 1990 May 25;18(10):2953–2960. doi: 10.1093/nar/18.10.2953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mosrin C., Riva M., Beltrame M., Cassar E., Sentenac A., Thuriaux P. The RPC31 gene of Saccharomyces cerevisiae encodes a subunit of RNA polymerase C (III) with an acidic tail. Mol Cell Biol. 1990 Sep;10(9):4737–4743. doi: 10.1128/mcb.10.9.4737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearson W. R. Rapid and sensitive sequence comparison with FASTP and FASTA. Methods Enzymol. 1990;183:63–98. doi: 10.1016/0076-6879(90)83007-v. [DOI] [PubMed] [Google Scholar]
- Riva M., Carles C., Sentenac A., Grachev M. A., Mustaev A. A., Zaychikov E. F. Mapping the active site of yeast RNA polymerase B (II). J Biol Chem. 1990 Sep 25;265(27):16498–16503. [PubMed] [Google Scholar]
- Russo F. D., Silhavy T. J. Alpha: the Cinderella subunit of RNA polymerase. J Biol Chem. 1992 Jul 25;267(21):14515–14518. [PubMed] [Google Scholar]
- Sentenac A. Eukaryotic RNA polymerases. CRC Crit Rev Biochem. 1985;18(1):31–90. doi: 10.3109/10409238509082539. [DOI] [PubMed] [Google Scholar]
- Stettler S., Mariotte S., Riva M., Sentenac A., Thuriaux P. An essential and specific subunit of RNA polymerase III (C) is encoded by gene RPC34 in Saccharomyces cerevisiae. J Biol Chem. 1992 Oct 25;267(30):21390–21395. [PubMed] [Google Scholar]
- Stinchcomb D. T., Mann C., Davis R. W. Centromeric DNA from Saccharomyces cerevisiae. J Mol Biol. 1982 Jun 25;158(2):157–190. doi: 10.1016/0022-2836(82)90427-2. [DOI] [PubMed] [Google Scholar]
- Stotz A., Linder P. The ADE2 gene from Saccharomyces cerevisiae: sequence and new vectors. Gene. 1990 Oct 30;95(1):91–98. doi: 10.1016/0378-1119(90)90418-q. [DOI] [PubMed] [Google Scholar]
- Sweetser D., Nonet M., Young R. A. Prokaryotic and eukaryotic RNA polymerases have homologous core subunits. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1192–1196. doi: 10.1073/pnas.84.5.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treich I., Carles C., Riva M., Sentenac A. RPC10 encodes a new mini subunit shared by yeast nuclear RNA polymerases. Gene Expr. 1992;2(1):31–37. [PMC free article] [PubMed] [Google Scholar]
- Treich I., Carles C., Sentenac A., Riva M. Determination of lysine residues affinity labeled in the active site of yeast RNA polymerase II(B) by mutagenesis. Nucleic Acids Res. 1992 Sep 25;20(18):4721–4725. doi: 10.1093/nar/20.18.4721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vallee B. L., Galdes A. The metallobiochemistry of zinc enzymes. Adv Enzymol Relat Areas Mol Biol. 1984;56:283–430. doi: 10.1002/9780470123027.ch5. [DOI] [PubMed] [Google Scholar]
- Werner M., Hermann-Le Denmat S., Treich I., Sentenac A., Thuriaux P. Effect of mutations in a zinc-binding domain of yeast RNA polymerase C (III) on enzyme function and subunit association. Mol Cell Biol. 1992 Mar;12(3):1087–1095. doi: 10.1128/mcb.12.3.1087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson T. E., Fahrner T. J., Johnston M., Milbrandt J. Identification of the DNA binding site for NGFI-B by genetic selection in yeast. Science. 1991 May 31;252(5010):1296–1300. doi: 10.1126/science.1925541. [DOI] [PubMed] [Google Scholar]
- Woychik N. A., Lane W. S., Young R. A. Yeast RNA polymerase II subunit RPB9 is essential for growth at temperature extremes. J Biol Chem. 1991 Oct 5;266(28):19053–19055. [PubMed] [Google Scholar]
- Woychik N. A., Liao S. M., Kolodziej P. A., Young R. A. Subunits shared by eukaryotic nuclear RNA polymerases. Genes Dev. 1990 Mar;4(3):313–323. doi: 10.1101/gad.4.3.313. [DOI] [PubMed] [Google Scholar]
- Woychik N. A., Young R. A. RNA polymerase II subunit RPB10 is essential for yeast cell viability. J Biol Chem. 1990 Oct 15;265(29):17816–17819. [PubMed] [Google Scholar]
- Wubbolts M. G., Terpstra P., van Beilen J. B., Kingma J., Meesters H. A., Witholt B. Variation of cofactor levels in Escherichia coli. Sequence analysis and expression of the pncB gene encoding nicotinic acid phosphoribosyltransferase. J Biol Chem. 1990 Oct 15;265(29):17665–17672. [PubMed] [Google Scholar]
- Young R. A. RNA polymerase II. Annu Rev Biochem. 1991;60:689–715. doi: 10.1146/annurev.bi.60.070191.003353. [DOI] [PubMed] [Google Scholar]





