Abstract
Human noroviruses are major causative agents of food and waterborne outbreaks of nonbacterial acute gastroenteritis. In this study, we report the epidemiological features of three outbreak cases of norovirus in Korea, and we describe the clinical symptoms and distribution of the causative genotypes. The incidence rates of the three outbreaks were 16.24% (326/2,007), 4.1% (27/656), and 16.8% (36/214), respectively. The patients in these three outbreaks were affected by acute gastroenteritis. These schools were provided unheated food from the same manufacturing company. Two genotypes (GII.3 and GII.4) of the norovirus were detected in these cases. Among them, major causative strains of GII.4 (Hu-jeju-47-2007KR-like) were identified in patients, food handlers, and groundwater from the manufacturing company of the unheated food. In the GII.4 (Hu-jeju-47-2007KR-like) strain of the norovirus, the nucleotide sequences were identical and identified as the GII.4 Sydney variant. Our data suggests that the combined epidemiological and laboratory results were closely related, and the causative pathogen was the GII.4 Sydney variant strain from contaminated groundwater.
Keywords: GII.4 Sydney variant, norovirus, outbreak
Norovirus is a major cause of epidemic acute nonbacterial gastroenteritis in humans worldwide. Norovirus infection is common in all age groups and is characterized by a low infection dose and efficient transmission with typical fecal-oral routes, as well as airborne spread and environmental contamination 1, 2, 3. Human norovirus GII.4 is prominent in all genogroups and genotypes, and GII.4 variants were divided into 13 subcluster types along with epidemic year and genetic characterization. In particular, the GII.4 Sydney strain, named as 2012 variant, is relatively new and predicted as the next prominent strain 4, 5, 6. There are several reports on GII.4-associated gastroenteritis, but there is a lack of environmental and molecular–epidemiological data. In Korea, three outbreaks associated with norovirus GII.4 Sydney variant occurred in a middle and high school setting in different cities on November 21st–30th, 2011. In this study, we describe the investigation of three outbreaks caused by the GII.4 Sydney variant, which was traced to contaminated groundwater from a supply manufacturing company.
In November 2011, three outbreaks of acute gastroenteritis occurred in three different regions. An epidemiological study was performed with a retrospective cohort study and cases were defined as patients who presented with diarrhea, nausea, abdominal pain, vomiting, and fever. Attack rates were 16.27% (326/2,007) in Outbreak A, 4.1% (27/656) in Outbreak B, and 16.8% (36/214) in Outbreak C. The main symptoms were diarrhea, nausea, abdominal pain, vomiting, and fever (Table 1). Patients from Outbreak A had mainly diarrheal symptoms (100%), patients from Outbreak B had abdominal pains (92.6%), and patients from Outbreak C had diarrhea (72.2%) and vomiting (72.2%). According to epidemiological findings, three outbreaks occurred in different regions, but the same food manufactures supplied the three regions with unheated food around the same time. This food manufacturer had used groundwater for the preparation of unheated food.
Table 1.
Comparison of epidemiological and laboratory characteristics.
| Features of outbreaks | Outbreak A | Outbreak B | Outbreak C |
|---|---|---|---|
| Epidemiological features | |||
| Attack rate (%) | 326/2007 (16.24) | 27/656 (4.1) | 36/214 (16.8) |
| Symptomatic patients | 326 | 27 | 36 |
| Date of onset | Nov 21, 2012 | Nov 30, 2012 | Nov 30, 2012 |
| Scale(s) | Large | Medium | Medium |
| Type of setting | School | School | School |
| Clinical symptoms (%) | |||
| Diarrhea | 326/326 (100) | 23/27 (85.2) | 26/36 (72.2) |
| Nausea | 223/326 (68.4) | 24/27 (88.9) | 24/36 (66.7) |
| Abdominal pain | 205/326 (62.9) | 25/27 (92.6) | 17/36 (47.2) |
| Vomiting | 156/326 (47.9) | 14/27 (51.9) | 26/36 (72.2) |
| Fever | 190/326 (58.3) | 14/27 (51.9) | 17/36 (47.2) |
| Laboratory test | |||
| No. of examinations of patients | 99 | 10 | 22 |
| Positive samples | 46 | 2 | 12 |
| Genogroup I | — | — | — |
| Genogroup II | 46 | 2 | 12 |
| No. of examinations of food-handlers | 28 | 14 | 5 |
| Positive samples | 2 | 5 | 2 |
| Genogroup I | — | — | — |
| Genogroup II | 2 | 5 | 2 |
| Environmental samples | 13 | ||
| Causative foods (unheated foods) | 9 | ||
| Genogroup I | — | ||
| Genogroup II | 9 | ||
| Supplemental water | 3 | ||
| Genogroup I | — | ||
| Genogroup II | 3 | ||
| Groundwater | 1 | ||
| Genogroup I | — | ||
| Genogroup II | 1 | ||
A laboratory test was performed on 131 fecal specimens of cases, 47 from food handlers, and 13 environmental samples. For virus detection, viral RNA was extracted using a commercialized RNA preparation kit (GM-AUTOPREP Kit, Green Mate Biotech Crop, Seoul, Korea). The norovirus was screened using commercialized real-time reverse transcription polymerase chain reaction (RT-PCR) (Bioneer, Daejeon, Korea) according to the manufacturer’s manual. From 131 fecal specimens from patients, 60 patients (45.8%) were determined as norovirus-positive with real-time RT-PCR. In each case, 46.5% (46/99) were norovirus-positive in Outbreak A, 20% (2/10) in Outbreak B, and 54.5% (12/22) in Outbreak C. Nine food handlers (19.1%) were determined as norovirus-positive from the 47 food handlers tested. All environmental samples tested were proven as norovirus-positive using real-time RT-PCR. Environmental samples included causative unheated food, supplemental water to prepare food in the manufacturer's company, and groundwater used as supplemental water (Table 1).
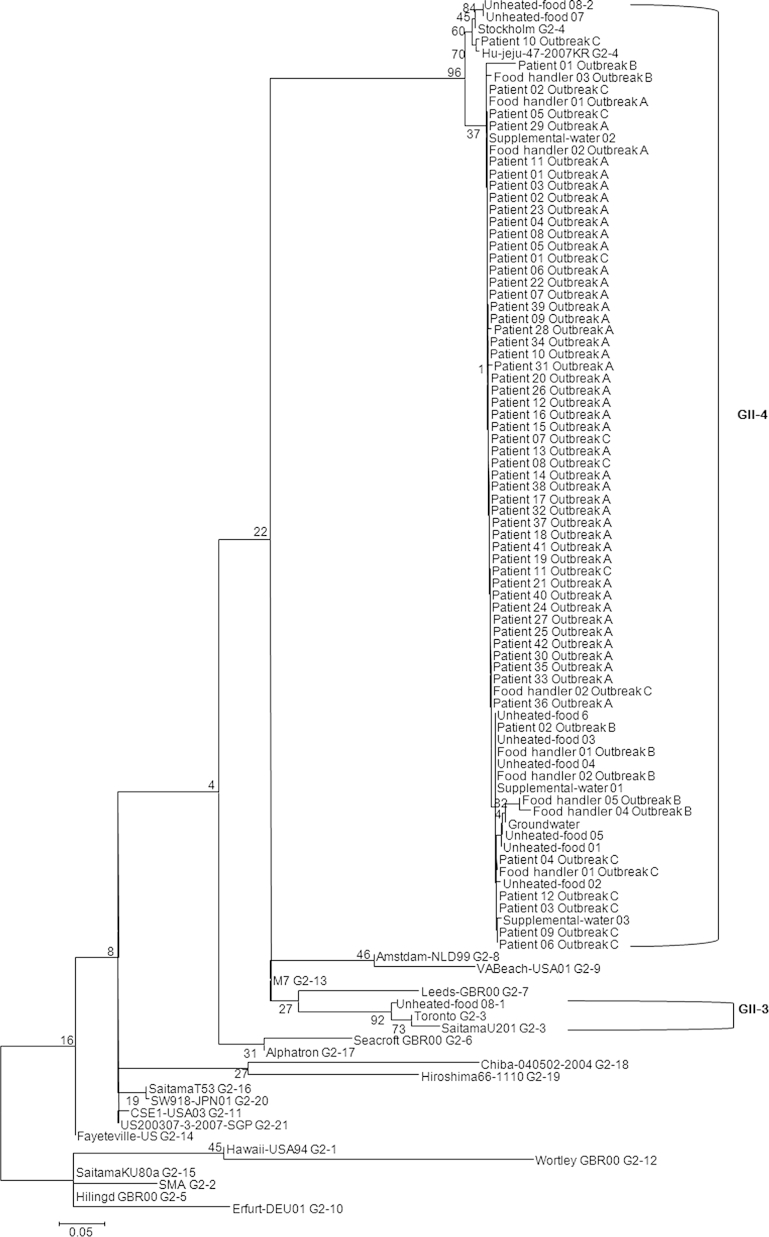
Amplification of norovirus genes were performed using one-step RT-PCR with norovirus specific primers listed in Table 2. Subsequently, seminested PCR was performed to partially amplify the norovirus capsid gene. Environmental samples were tested according to the methods of the Ministry of Food and Drug Safety and stool specimens were tested according to the methods of the Korea National Institute of Health. Amplified PCR products were purified using Millipore plate MSNU030 (Millipore SAS, Molsheim, France) and were sequenced with the Big Dye terminator version 3.1 sequencing kit and a 3730xl automated sequencer (Applied Bio systems, Foster City, CA, USA). Nucleic acid sequences were aligned with information of genotype sequence using MegAlign package (Window version 3.12e, DNASTAR, Madison, WI) in the DNASTAR program, and the phylogenetic tree was generated using the neighbor-joining method 7, 8. Most genotypes were determined as GII.4 (Hu-jeju-like) strains detected in patients, food handler fecal samples, and environmental samples. However, GII.3 was only detected in one sample collected from unheated food (Figure 1). To compare sequence identity, each group showed a similarity of 96.8–99.5% (Outbreaks A and B), 94.7–100% (Outbreaks A and C), 96.3–100% (Outbreak A and environmental samples), 89.9.3–99.5% (Outbreaks B and C), 95.2–100% (Outbreak B and environmental samples), and 94.7–100% (Outbreak C and environmental samples; data not shown).
Table 2.
Oligo nucleotide sequences of primers used in this study.
| Genogroup | Name of primer | Sequence (5′→3′) | Application |
|---|---|---|---|
| GI | NoGI-F1 | ATGGCCATGTTCCGITGGATG | Stool specimen |
| NoGI-F2 | CGGGCCCGAATTYGTAAATGATG | ||
| NoGI-R | CCAACCCARCCATTRTACATYTG | ||
| GI-F1M | CTGCCCGAATTYGTAAATGATGAT | Stool & environmental samples | |
| GI-F2 | ATGATGATGGCGTCTAAGGACGC | ||
| NoGI-R | CCAACCCARCCATTRTACATYTG | ||
| GII | NoGII-F1 | CCCTCGAGGGCGATCGCAATCT | Stool specimen |
| NoGII-F2 | CACAATTGTGAATGAAGATGGCGTCGA | ||
| NoGII-R | CCRCCIGCATRICCRTTRTACAT | ||
| GII-F1M | GGGAGGGCGATCGCAATCT | Stool & environmental samples | |
| GII-F3M | TTGTGAATGAAGATGGCGTCGART | ||
| NoGII-R | CCRCCIGCATRICCRTTRTACAT |
Figure 1.
Phylogenetic analysis of the norovirus detected from patients, food handlers, and environmental samples (unheated foods, supplemental water, and groundwater). The phylogenetic tree was constructed with the neighbor-joining method with norovirus partial capsid region (312-314bp). The numbers in the branches indicate the bootstrap values.
GII.4 variant analysis was performed using NoroNet alignment tools (http://noronet.nl) 4, 5, 6. Prominent strains were 2012 Sydney variants in these three outbreaks, although 2006b and 2010 New Orleans were also detected. Analysis of strains from Outbreaks A and B detected the GII-4 Sydney strain but a 2006b variant was detected in Outbreak C. In the environmental samples, the 2010 New Orleans strain was detected in two unheated food samples. However, the 2012 Sydney strain occupied all environmental samples (Table 3).
Table 3.
Distribution of norovirus GII-4 variant analyzed in this study.
| Name of variant | Outbreak A | Outbreak B | Outbreak C | Environmental samples |
|---|---|---|---|---|
| 2006b | — | — | 1 (7.1) | — |
| 2010 | — | — | — | 2 (16.7) |
| 2012 | 44 (100) | 7 (100) | 13 (92.9) | 10 (83.3) |
Data are presented as n (%).
Three outbreaks occurred with the same norovirus GII.4 Sydney variant in different cities at the same time and showed typical symptoms of norovirus-related acute gastroenteritis. These outbreaks were associated with unheated food which was commonly provided from the same manufacturer. However, there were several environmental factors that led to infection, such as food handlers, unheated food, supplemental water, and groundwater. GII.4 strains were already well-known to be prominent in humans and the current prominent Sydney variant has been well-reported in outbreak cases 4, 5, 6, 9, 10, 11. However, there is a lack of reports on 2012 Sydney variant-associated cases. Our data were fully analyzed based on clinical, environmental, and molecular analysis. In conclusion, this study revealed three outbreaks of acute gastroenteritis caused by norovirus GII that came from contaminated groundwater, and the major causative pathogen was the GII-4 Sydney variant.
Conflicts of interest
The authors have nothing to declare.
Acknowledgments
This study was supported by the National Norovirus Surveillance System (4851-304-210) and conducted by the Korea National Institute Health and Seoul and Gyungbuk Institute of Health and Environment Research.
Footnotes
This is an open-access article distributed under the terms of the Creative Commons Attribution-NonCommercial-No Derivative Works License (http://creativecommons.org/licenses/by-nc-nd/4.0) which permits non-commercial use, distribution, and reproduction in any medium, provided the original author and source are credited.
References
- 1.Rockx B., De Wit M., Vennema H. Natural history of human calicivirus infection: a prospective cohort study. Clin Infect Dis. 2002 Aug;35(3):246–253. doi: 10.1086/341408. [DOI] [PubMed] [Google Scholar]
- 2.Maunula L., Miettinen I.T., von Bonsdorff C.H. Norovirus outbreaks from drinking water. Emerg Infect Dis. 2005 Nov;11(11):1716–1721. doi: 10.3201/eid1111.050487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hewitt J., Bell D., Simmons G.C. Gastroenteritis outbreak caused by waterborne norovirus at a New Zealand ski resort. Appl Environ Microbiol. 2007 Dec;73(24):7853–7857. doi: 10.1128/AEM.00718-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Belliot G., Kamel A.H., Estienney M. Evidence of emergence of new GGII.4 norovirus variants from gastroenteritis outbreak survey in France during the 2007-to-2008 and 2008-to-2009 winter seasons. J Clin Microbiol. 2010 Mar;48(3):994–998. doi: 10.1128/JCM.01888-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Vega E., Barclay L., Gregoricus N. Novel surveillance network for norovirus gastroenteritis outbreaks, United States. Emerg Infect Dis. 2011 Aug;17(8):1389–1395. doi: 10.3201/eid1708.101837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.van Beek J., Ambert-Balay K., Botteldoorn N. Indications for worldwide increased norovirus activity associated with emergence of a new variant of genotype II.4, late 2012. Euro Surveill. 2013 Jan;18(1):8–9. [PubMed] [Google Scholar]
- 7.Felsenstein J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 1985 Jul;39(4):783–791. doi: 10.1111/j.1558-5646.1985.tb00420.x. [DOI] [PubMed] [Google Scholar]
- 8.Saitou N., Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987 Jul;4(4):406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- 9.Fonager J., Hindbæk L.S., Fischer T.K. Rapid emergence and antigenic diversification of the norovirus 2012 Sydney variant in Denmark, October to December, 2012. Euro Surveill. 2013 Feb;28;18(9):1–4. [PubMed] [Google Scholar]
- 10.Polkowska A., Ronnqvist M., Lepisto O. Outbreak of gastroenteritis caused by norovirus GII.4 Sydney variant after a wedding reception at a resort/activity centre, Finland, August 2012. Epidemiol Infect. 2014 Sep;142(9):1877–1883. doi: 10.1017/S0950268813002847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wong T.H., Dearlove B.L., Hedge J. Whole genome sequencing and de novo assembly identifies Sydney-like variant noroviruses and recombinants during the winter 2012/2013 outbreak in England. Virol J. 2013 Nov;10:335. doi: 10.1186/1743-422X-10-335. [DOI] [PMC free article] [PubMed] [Google Scholar]