Abstract
The right heart is functionally and structurally different from the left heart; however, potential differences in Akt signaling and the expression of metabolic genes between the right heart and left heart in different rodents are still unknown. Using Western blotting and real time quantification polymerase chain reaction, we measured the levels of total Akt, phosphorylated Akt and its downstream targets as well as metabolism genes including glucose transporter 1, glucose transporter 4 (GLUT4), peroxisome proliferator-activated receptor α, peroxisome proliferator-activated receptor γ, peroxisome proliferator-activated receptor δ, peroxisome proliferator-activated receptor gamma coactivator 1α (PGC-1α), and pyruvate dehydrogenase lipoamide kinase isozyme 4. We found that phosphorylated Akt and proline-rich Akt substrate 40 levels were significantly increased in the RV compared with the LV in rats but only had an increased trend in mice. Correspondingly, GLUT4 was significantly increased in the RV compared with the LV both in mice and rats. PGC-1α was significantly increased in the RV compared with the LV in mice but only had an increased trend in rats. Moreover, Akt signaling activity and metabolism genes’ expression in the IVS were similar to the RV in mice but to the LV in rats. There were some differences in the activity of Akt signaling and in the levels of metabolism genes among the right ventricle, interventricular septum and left ventricle. Also, the diversity of activity of Akt and metabolism genes between the right ventricle and left ventricle are different between rats and mice. In conclusion, the activity of Akt signaling and the levels of metabolism genes are different among the right ventricle, interventricular septum and left ventricle providing some potential clues for exploring the roles of Akt signaling and cardiac metabolisms in different parts of the heart. Additionally, the differences in Akt activity and metabolism genes’ levels between the right and left ventricles are different between mice and rats, to which we should pay attention when using different animal model in heart study.
Keywords: Right ventricle, left ventricle, Akt signaling, cardiac metabolism, rodent
Introduction
Limited information related to right heart suggests that the right ventricle (RV) is an important contributor and that further understanding of these issues is of pivotal importance [1]. There are important differences in developmental origin and postnatal function of left and right ventricles (LV and RV, respectively) [2,3]. The LV originates from the first heart field progenitor cells whereas the RV from the second heart field progenitor cells. Functionally, the tolerance of the RV for pressure-overload is weaker than that of LV. Conversely, the RV can tolerate more volume-overload than LV. Accordingly, the RV cannot be understood simply by extrapolating data and experience from the LV [2,3].
Protein kinase B (Akt) is a key molecule in energy metabolism and has been reported to be involved in cardiac hypertrophy, remodeling and cardiomyocyte proliferation in the LV [4,5]. A recent study showed that Akt signaling plays a very important role in the development of the RV [6]. However, it is still unknown whether there are differences in Akt activity between the RV and LV. The relative expression of metabolism genes between the RV and LV is also undefined.
Rodents are extensively used as human disease models, including for many types of heart disease [7]. Therefore, it is important to know the characteristics of distribution of Akt signaling pathway and metabolism genes’ expression in RV and LV between rat and mouse.
In the present study, we measured the activity of Akt signaling and the levels of metabolic genes in the RV and LV of mice and rats. We found the Akt activity and metabolism genes’ levels are different between the RV and LV. The differences themselves differed between mice and rats.
Materials and methods
Animal model
Male Sprague Dawley rats (200-250 g) fed a standard diet were anesthetized with sodium pentobarbital (50 mg/kg) i.p. The hearts were excised and the RV, interventricular septum (IVS) and LV were dissected on ice. Male mice (20-25 g) on a C57BL/6 genetic background were housed in groups with 12 h dark/light cycles and free access to food. After anesthetization (sodium pentobarbital 70-80 mg/kg), the RV, IVS and LV were dissected. All experiments were in accordance with the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health (NIH publication no. 85-23, revised in 1996) and the regulations on mouse welfare and ethics of Gansu Provincial Hospital. All procedures were approved by the Ethics Committee of Gansu Provincial Hospital (Lanzhou, China).
Western blotting analysis
Heart lysates of mice and rats were prepared in lysis buffer (20 mM Tris, 150 mM NaCl, 10% glycerol, 20 mM glycerophosphate, 1% NP40, 5 mM EDTA, 0.5 mM EGTA, 1 mM Na3VO4, 0.5 mM PMSF, 1 mM benzamidine, 1 mM DTT, 50 mM NaF, 4 µM leupeptin, pH=8.0). Samples were resolved on 10% SDS-PAGE gels and transferred to PVDF membranes (Millipore). Membranes were blocked with 5% non-fat milk in TBST (50 mM Tris, 150 mM NaCl, 0.5 mM Tween-20, pH=7.5) and then incubated with primary antibodies overnight. Antibodies used in this study were purchased from Cell Signaling Technology (CST), BioWorld: total Akt (CST #4691), phospho-Akt (Ser308) (CST #4060), phospho-Akt (Thr473) (CST #13038), phospho-GSK3β (Ser9) (CST #5558), LC3A/B (CST #12741), PRAS40 (CST #2691), phospho-PRAS40 (Thr246) (CST #13175), PTEN (CST #9188), phospho-PTEN (Ser380) (CST #9551), GAPDH (#AP0063), Anti-rabbit IgG, (HRP-linked Antibody) (CST #7074). Image J software (NIH) was used to perform densitometric analysis (http://rsb.info.nih.gov/ij/).
Quantitative real-time PCR for metabolism related genes and fetal genes
Total RNA was extracted from the LV, IVS and RV using TRIZOL reagent (Invitrogen), according to the manufacturer’s protocol. One microgram of total RNA from each specimen was reverse transcribed to cDNA using SuperScript Reverse Transcriptase and random hexamers as primers (Invitrogen). Quantitative real-time PCR (qRT-PCR) was performed with an ABI Stepone plus instrument (Applied Biosystems) using 1 ×I TaQ SYBR green Supermix Kit (Bio-Bad, Reinach, Switzerland) and 300 nmol/l for forward and reverse primers in a total volume of 20 μl. The mRNA level was based on the critical threshold (Ct) value. Primer sequences for quantitative real-time PCR were showed in Table 1. Gapdh was used as internal control.
Table 1.
Primer sequences of metabolism genes in rat and mouse
| Species | Primers | Forward | Reverse |
|---|---|---|---|
| Rat | GLUT1 | GCTGTGGCTGGCTTCTCTAA | CCGGAAGCGATCTCATCGAA |
| Rat | GLUT4 | GGCAATGCCAAATTGCTCCA | GGCACAGTTAAGGTCCCCTC |
| Rat | PPARα | GAGTAGCCTGGGCTGCTTTT | CTGATCACCAGCAGAGGTCC |
| Rat | PPARγ | TACCACGGTTGATTTCTC | TCTACTTTGATCGCACTTT |
| Rat | PPARδ | CATCCGTTCTCTACCCAGCC | AATTCTGAGCCCGGAGTTGG |
| Rat | PGC-1α | TGGAGTGACATAGAGTGTGCTG | TATGTTCGCGGGCTCATTGT |
| Rat | PDK4 | ACAATTCACGGAATGCCCCT | TACTTGGCGTAGAGACGGGA |
| Mouse | GLUT1 | GCTGTGCTTATGGGCTTCTC | CACATACATGGGCACAAAGC |
| Mouse | GLUT4 | ACTCTTGCCACACAGGCTCT | CCTTGCCCTGTCAGGTATGT |
| Mouse | PPARα | TGCAAACTTGGACTTGAACG | GATCAGCATCCCGTCTTTGT |
| Mouse | PPARγ | GAGTGTGACGACAAGATTTG | GGTGGGCCAGAATGGCATCT |
| Mouse | PPARδ | TGGAGCTCGATGACAGTGAC | GTACTGGCTGTCAGGGTGGT |
| Mouse | PGC-1α | CACGCAGCCCTATTCA | GTCGTACCTGGGCCTA |
| Mouse | PDK4 | CCGCTGTCCATGAAGCA | GCAGAAAAGCAAAGGACGTT |
Abbreviation: GLUT1, glucose transporter 1; GLUT4, glucose transporter 4; PPARα, peroxisome proliferator-activated receptor α; PPARγ, peroxisome proliferator-activated receptor γ; PPARδ, peroxisome proliferator-activated receptor δ; PGC-1α, peroxisome proliferator-activated receptor gamma coactivator 1α; PDK4, Pyruvate dehydrogenase lipoamide kinase isozyme.
Statistics
Results of calculations are presented as means ± SEM. Differences in means between two groups were evaluated with paired two-tailed Student t tests. All statistics were performed with GraphPad Prism 4.0 software (GraphPad, San Diego, CA, USA). p values of <0.05 were considered statistically significant.
Results
Akt activity is slightly stronger in the RV and IVS than in the LV in mice
Compared with the LV, phosphorylation of Akt was slightly higher at both sites 473 and 308 in the RV and IVS (Figure 1A-C). Total Akt had no significant differences between the LV, RV and IVS (Figure 1A, 1D). Correspondingly, total PTEN, an upstream inhibitor of Akt activation, was slightly lower in the RV and IVS than in the LV. Phosphorylated PTEN was a little increased in the IVS but not in the RV compared with the LV (Figure 1A, 1E, 1F). PRAS40, an indicator of Akt activity, had an increasing trend in phosphorylated form but not in total form in the RV and IVS compared with the LV (Figure 1A, 1H, 1I). Phosphorylated GSK3β, another downstream target of Akt, had a similar trend as phosphorylated PRAS40 (Figure 1A, 1G). Furthermore, LC3B also had an increased trend in the RV and IVS compared with the LV (Figure 1A, 1J). Taken together, there is an increasing trend in Akt signaling indicated by its phosphorylated form and downstreams in the RV and IVS compared with the LV.
Figure 1.

Akt signaling in LV, RV and IVS in mice. A. Representative Western blot. B-J. Quantifications of A (n=6). Note: Akt, protein kinase B; PTEN, phosphatase and tensin homolog; GSK3β, Glycogen synthase kinase 3β; PRAS40, proline-rich Akt substrate 40 kDa; LC3B, Autophagy marker Light Chain 3B; *P<0.05, versus LV.
Glucose transporter 4 (GLUT4) and peroxisome proliferator-activated receptor gamma coactivator 1α (PGC-1α) are more highly expressed in the RV and IVS than in the LV in mice
Compared with the LV, GLUT4 was significantly increased in the RV and IVS. The difference in PGC-1α levels between the RV and LV was a trend that was not statistically significant, but the difference between the IVS and LV was statistically significant. There were no significant differences in glucose transporter 1 (GLUT1), peroxisome proliferator-activated receptor α (PPARα), peroxisome proliferator-activated receptor γ (PPARγ), peroxisome proliferator-activated receptor δ (PPARδ), pyruvate dehydrogenase lipoamide kinase isozyme 4 (PDK4) between the LV, RV and IVS. However, compared with the LV, PPARα, PPARγ, PPARδ and PDK4 had an obvious increasing trend in the RV and IVS (Figure 2). Taken together, several metabolic genes including GLUT4 in the RV and IVS and PGC-1α in the IVS had significantly higher expression than in the LV.
Figure 2.

Metabolism genes in LV, RV and IVS in mice (n=6). Note: GLUT1, glucose transporter 1; GLUT4, glucose transporter 4; PPARα, peroxisome proliferator-activated receptor α; PPARγ, peroxisome proliferator-activated receptor γ; PPARδ, peroxisome proliferator-activated receptor δ; PGC-1α, peroxisome proliferator-activated receptor gamma coactivator 1α; PDK4, Pyruvate dehydrogenase lipoamide kinase isozyme 4; *P<0.05, **P<0.01, versus LV.
Akt activity is significantly stronger in the RV than in the LV and IVS in rats
Compared with the LV, phosphorylation of Akt was significantly stronger at site 308 but not at 473 in the RV and IVS (Figure 3A-C). Compared with LV, total Akt had no significant differences in the LV, RV and IVS but it was significantly increased in IVS than in RV (Figure 3A, 3D). Correspondingly, PRAS40, an indicator of Akt activity, was significantly increased in phosphorylation but not total level in the RV compared with the LV. Compared with RV, phosphorylated PRAS40 had a decreasing trend in the IVS (Figure 3A, 3H, 3I). Also, phosphorylated GSK3β, another downstream target of Akt, was significantly increased in the IVS but not in the RV compared with the LV (Figure 3A, 3G). However, both phosphorylated PTEN and total PTEN, upstream of Akt, had no significantly differences among the LV, RV and IVS (Figure 3A, 3E, 3F). As in mice, LC3B was a little but not significantly increased in the RV and IVS compared with the LV (Figure 3A, 3J). Taken together, Akt activity indicated by its phosphorylated form and downstream targets was significantly stronger in the RV than in the IVS and LV.
Figure 3.
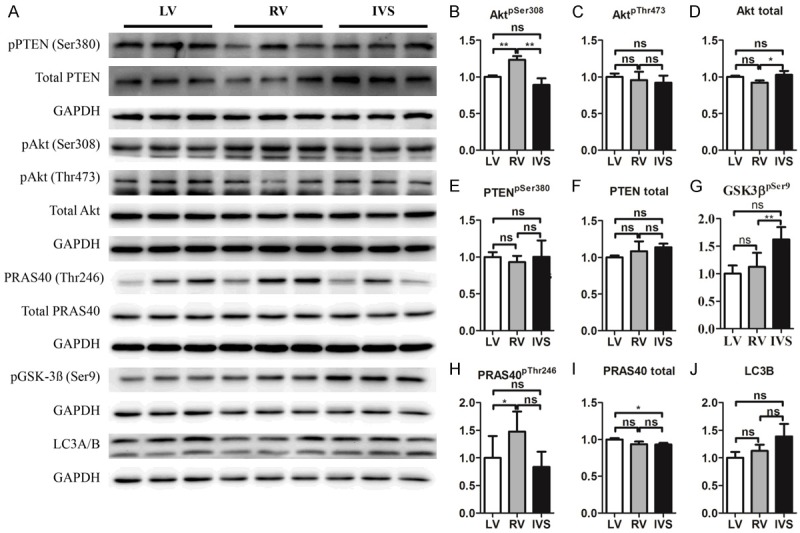
Akt signaling in LV, RV and IVS in rats. A. Representative Western blot. B-J. Quantifications of A (n=6). Note: Akt, protein kinase B; PTEN, phosphatase and tensin homolog; GSK3β, Glycogen synthase kinase 3β; PRAS40, proline-rich Akt substrate 40 kDa; LC3B, Autophagy marker Light Chain 3B; *P<0.05, **P<0.01, versus LV.
Higher expression of metabolism genes in the RV than in the LV and IVS in rats
Compared with the LV and IVS, GLUT4 was significantly increased in the RV. There was a strong trend towards increased GLUT1, PPARα, PPARγ, PPARδ, PGC-1α and PDK4 in the RV compared with the LV and IVS. In a word, there were some differences in the expression of metabolism genes among LV, RV and IVS (Figure 4).
Figure 4.
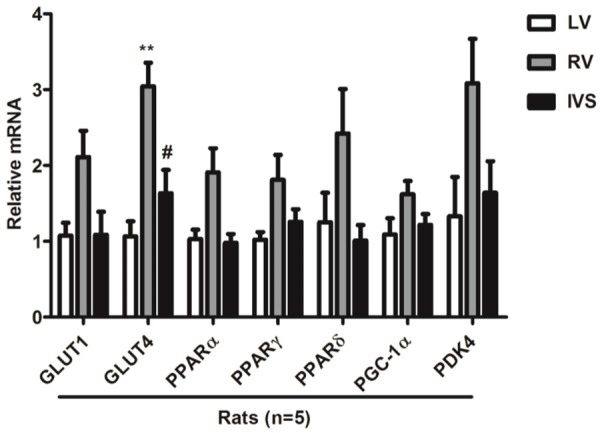
Metabolism genes in LV, RV and IVS in rats (n=5). Note: GLUT1, glucose transporter 1; GLUT4, glucose transporter 4; PPARα, peroxisome proliferator-activated receptor α; PPARγ, peroxisome proliferator-activated receptor γ; PPARδ, peroxisome proliferator-activated receptor δ; PGC-1α, peroxisome proliferator-activated receptor gamma coactivator 1α; PDK4, Pyruvate dehydrogenase lipoamide kinase isozyme 4; *P<0.05, **P<0.01, versus LV; #P<0.05, versus RV.
Discussion
The RV is very different in embryonic origin, anatomy and postnatal physiologic function from the LV. The IVS has the same embryonic origin as the RV, but is more functionally similar to the LV [2,3]. Mounting evidence has shown that Akt signaling plays a vital role in LV remodeling through effects on energy metabolism [5]. Recently, altered Akt signaling has been reported to affect RV development [6]. However, it still unclear that the differences in Akt signaling and the expression of metabolism genes between the LV, RV and IVS in mice and rats. Here we report that there are differences in Akt signaling and the expression of metabolism genes between the LV, RV and IVS in mice and rats.
Compared with the LV, Akt signaling has an increased trend but not significantly in the RV and IVS of mice. However, in rats, Akt signaling is significantly stronger in the RV but not in the IVS than the LV. Meanwhile, metabolism genes including GLUT4 and PGC-1α are consistently increased in the RV compared with the LV. Previous studies have shown that the oxygen demand of the RV is physiologically different from the LV [8] and that the energy metabolism of the RV is different from the LV under some pathological conditions. In view of the fact that Akt plays a central role in regulating heart metabolism by directly or indirectly controlling GLUT4 [9], GSK-3β [10], and mitochondrial function [11,12], it could be reasonable that increased Akt signaling is responsible for increased GLUT4 and PGC-1α in the RV compared with the LV. Collectively, the RV relies more on Akt signaling than LV under physiological status which may be related to its physiological features [3].
Although both mice and rats are rodents, there are really some differences in some signaling molecules including G protein-coupled receptor 78 (GPR78), inositol-requiring enzyme 1 (IRE1), sterol regulatory element-binding protein 1 (SREBP-1) and sodium-glucose cotransporter-2 (SGLT2) et al. between rats and mice [13-18]. In this study, we found that the IVS has Akt activity similar to the LV in rats whereas it is more similar to the RV in mice. Rats also differed from mice in the relative levels of several metabolism genes including GLUT4 and PGC-1α between the RV, IVS and LV. One of the possible reasons leading to these differences is genetic discrepancy between rats and mice [19,20]. These findings suggest that we should pay attention to the diversity of these two species which are frequently used as an experimental models in heart study [7].
In conclusion, the present study shows differences in Akt signaling activity and metabolism genes levels between the LV, RV and IVS, as well as between mice and rats. Our findings may be useful for future heart studies in rodent models.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81460072) grants to Yunshan Cao. Thanks to Colin Platt from Cardiovascular Division of Massachusetts General Hospital (MGH) and Harvard Medical School for providing language help and proof reading the article.
Disclosure of conflict of interest
None.
References
- 1.Mehra MR, Park MH, Landzberg MJ, Lala A, Waxman AB. Right heart failure: toward a common language. Pulm Circ. 2013;3:963–967. doi: 10.1086/674750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Haddad F, Doyle R, Murphy DJ, Hunt SA. Right ventricular function in cardiovascular disease, part II: pathophysiology, clinical importance, and management of right ventricular failure. Circulation. 2008;117:1717–1731. doi: 10.1161/CIRCULATIONAHA.107.653584. [DOI] [PubMed] [Google Scholar]
- 3.Haddad F, Hunt SA, Rosenthal DN, Murphy DJ. Right ventricular function in cardiovascular disease, part I: Anatomy, physiology, aging, and functional assessment of the right ventricle. Circulation. 2008;117:1436–1448. doi: 10.1161/CIRCULATIONAHA.107.653576. [DOI] [PubMed] [Google Scholar]
- 4.Altomare DA, Khaled AR. Homeostasis and the importance for a balance between AKT/mTOR activity and intracellular signaling. Curr Med Chem. 2012;19:3748–3762. doi: 10.2174/092986712801661130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sussman MA, Volkers M, Fischer K, Bailey B, Cottage CT, Din S, Gude N, Avitabile D, Alvarez R, Sundararaman B, Quijada P, Mason M, Konstandin MH, Malhowski A, Cheng Z, Khan M, McGregor M. Myocardial AKT: the omnipresent nexus. Physiol Rev. 2011;91:1023–1070. doi: 10.1152/physrev.00024.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Luo W, Zhao X, Jin H, Tao L, Zhu J, Wang H, Hemmings BA, Yang Z. Akt1 signaling coordinates BMP signaling and beta-catenin activity to regulate second heart field progenitor development. Development. 2015;142:732–742. doi: 10.1242/dev.119016. [DOI] [PubMed] [Google Scholar]
- 7.Zaragoza C, Gomez-Guerrero C, Martin-Ventura JL, Blanco-Colio L, Lavin B, Mallavia B, Tarin C, Mas S, Ortiz A, Egido J. Animal models of cardiovascular diseases. J Biomed Biotechnol. 2011;2011:497841. doi: 10.1155/2011/497841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zong P, Tune JD, Downey HF. Mechanisms of oxygen demand/supply balance in the right ventricle. Exp Biol Med (Maywood) 2005;230:507–519. doi: 10.1177/153537020523000801. [DOI] [PubMed] [Google Scholar]
- 9.Brownsey RW, Boone AN, Allard MF. Actions of insulin on the mammalian heart: metabolism, pathology and biochemical mechanisms. Cardiovasc Res. 1997;34:3–24. doi: 10.1016/s0008-6363(97)00051-5. [DOI] [PubMed] [Google Scholar]
- 10.Markou T, Cullingford TE, Giraldo A, Weiss SC, Alsafi A, Fuller SJ, Clerk A, Sugden PH. Glycogen synthase kinases 3alpha and 3beta in cardiac myocytes: regulation and consequences of their inhibition. Cell Signal. 2008;20:206–218. doi: 10.1016/j.cellsig.2007.10.004. [DOI] [PubMed] [Google Scholar]
- 11.Miyamoto S, Murphy AN, Brown JH. Akt mediated mitochondrial protection in the heart: metabolic and survival pathways to the rescue. J Bioenerg Biomembr. 2009;41:169–180. doi: 10.1007/s10863-009-9205-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sussman MA. Mitochondrial integrity: preservation through Akt/Pim-1 kinase signaling in the cardiomyocyte. Expert Rev Cardiovasc Ther. 2009;7:929–938. doi: 10.1586/erc.09.48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bai Y, Sanderson MJ. The contribution of Ca2+ signaling and Ca2+ sensitivity to the regulation of airway smooth muscle contraction is different in rats and mice. Am J Physiol Lung Cell Mol Physiol. 2009;296:L947–958. doi: 10.1152/ajplung.90288.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Francis C, Natarajan S, Lee MT, Khaladkar M, Buckley PT, Sul JY, Eberwine J, Kim J. Divergence of RNA localization between rat and mouse neurons reveals the potential for rapid brain evolution. BMC Genomics. 2014;15:883. doi: 10.1186/1471-2164-15-883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Swindell WR. Dietary restriction in rats and mice: a meta-analysis and review of the evidence for genotype-dependent effects on lifespan. Ageing Res Rev. 2012;11:254–270. doi: 10.1016/j.arr.2011.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shinohara M, Ji C, Kaplowitz N. Differences in betaine-homocysteine methyltransferase expression, endoplasmic reticulum stress response, and liver injury between alcohol-fed mice and rats. Hepatology. 2010;51:796–805. doi: 10.1002/hep.23391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Downs SM. Mouse versus rat: Profound differences in meiotic regulation at the level of the isolated oocyte. Mol Reprod Dev. 2011;78:778–794. doi: 10.1002/mrd.21377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Sabolic I, Vrhovac I, Eror DB, Gerasimova M, Rose M, Breljak D, Ljubojevic M, Brzica H, Sebastiani A, Thal SC, Sauvant C, Kipp H, Vallon V, Koepsell H. Expression of Na+-D-glucose cotransporter SGLT2 in rodents is kidney-specific and exhibits sex and species differences. Am J Physiol Cell Physiol. 2012;302:C1174–1188. doi: 10.1152/ajpcell.00450.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Jensen-Seaman MI, Furey TS, Payseur BA, Lu Y, Roskin KM, Chen CF, Thomas MA, Haussler D, Jacob HJ. Comparative recombination rates in the rat, mouse, and human genomes. Genome Res. 2004;14:528–538. doi: 10.1101/gr.1970304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Prasad A, Kumar SS, Dessimoz C, Bleuler S, Laule O, Hruz T, Gruissem W, Zimmermann P. Global regulatory architecture of human, mouse and rat tissue transcriptomes. BMC Genomics. 2013;14:716. doi: 10.1186/1471-2164-14-716. [DOI] [PMC free article] [PubMed] [Google Scholar]


