Abstract
Vasopressin controls osmotic water transport in the renal collecting duct through regulation of aquaporin-2 (AQP2). We carried out bioinformatic analysis of quantitative proteomic data from the accompanying article to investigate the mechanisms involved. The experiments used stable isotope labeling by amino acids in cell culture in cultured mpkCCD cells to quantify each protein species in each of five differential-centrifugation (DC) fractions with or without the vasopressin analog 1-desamino-8-d-arginine-vasopressin (dDAVP). The mass spectrometry data and parallel Western blot experiments confirmed that dDAVP addition is associated with an increase in AQP2 abundance in the 17,000-g pellet and a corresponding decrease in the 200,000-g pellet. Remarkably, all subunits of the cytoplasmic ribosome also increased in the 17,000-g pellet in response to dDAVP (P < 10−34), with a concomitant decrease in the 200,000-g pellet. Eukaryotic translation initiation complex 3 (eIF3) subunits underwent parallel changes (P < 10−6). These findings are consistent with translocation of assembled ribosomes and eIF3 complexes into the rough endoplasmic reticulum in response to dDAVP. Conversely, there was a systematic decrease in small GTPase abundances in the 17,000-g fraction. In contrast, most proteins, including protein kinases, showed no systematic redistribution among DC fractions. Of the 521 protein kinases coded by the mouse genome, 246 were identified, but many fewer were found to colocalize with AQP2 among DC fractions. Bayes' rule was used to integrate the new colocalization data with prior data to identify protein kinases most likely to phosphorylate aquaporin-2 at Ser256 (Camk2b > Camk2d > Prkaca) and Ser261 (Mapk1 = Mapk3 > Mapk14).
Keywords: aquaporin-2, mass spectrometry, ribosome, translation, small GTPase, protein kinase, phosphorylation
in the accompanying article (38), we carried out comprehensive proteomic profiling of a vasopressin-sensitive mouse collecting duct cell line, mpkCCD-clone 11. The mpkCCD collecting duct cell line has been an important tool in the study of vasopressin-mediated regulation of the water channel aquaporin-2 (AQP2) (10, 20). The new proteomic data provide a “parts list” for these cells, which can be used as a tool in experimental design. In the accompanying article, for each protein identified, we also provided “virtual Western blots,” which map all identified peptides to positions in the SDS-polyacrylamide gels used to fractionate the proteins. The mapping provides comprehensive molecular weight information for all identified proteins, which we analyzed with respect to mechanisms involved in production of larger- or smaller-than-predicted proteins. An example is the water channel AQP2, which was found to have a previously undocumented “small” isoform of molecular mass <20 kDa.
The purpose of this study is to use the comprehensive data set from the accompanying article (38) to ask critical questions about the nature of vasopressin-stimulated signaling in mpkCCD cells. Signaling in all cell types is mediated by a limited number of processes that can be quantified by protein mass spectrometry, chiefly, 1) changes in the cellular abundances of individual proteins, 2) changes in posttranslational modifications of individual proteins, 3) changes in the subcellular distribution (translocation) of individual proteins, and 4) changes in binding interactions of proteins with other proteins, lipids, or nucleic acids. In our previous studies we used quantitative protein mass spectrometry to carry out proteome-wide profiling of vasopressin's effects on two of these processes in mpkCCD cells: 1) changes in individual protein abundances (18, 30) and 2) changes in posttranslational modifications, specifically phosphorylation events (4, 27). Among the phosphorylation sites regulated by vasopressin are four sites in the COOH-terminal tail (Ser256, Ser261, Ser264, and Ser269) that are believed to play roles in regulated trafficking of AQP2 (8, 9, 12–14, 22). In addition, we have carried out nuclear proteomics to identify proteins that move into or out of the nucleus in response to vasopressin (32). Here we use data from the accompanying article to characterize the five fractions profiled, the 1,000-g pellet (1K), the 4,000-g pellet (4K), the 17,000-g pellet (17K), the 200,000-g pellet (200Kp), and the 200,000-g supernatant (200Ks), with regard to the cellular components they contain. We then use the quantitative data to ask whether specific groups of proteins appear to undergo translocation among the five fractions profiled. We used stable isotope labeling by amino acids in cell culture (SILAC) (26) for quantitative comparison of abundances of individual proteins in vasopressin- vs. vehicle-treated cells. Analysis of the data identified several protein complexes that appear to translocate from one fraction to another in response to vasopressin.
We also use the data set from the accompanying article (38) to address another question relevant to vasopressin signaling: What protein kinases among the >500 kinases coded by the genome (7) are most likely to play roles in vasopressin-induced phosphorylation changes that were identified in prior studies (4, 27)? This analysis is based on the simple concept that, for a given kinase to phosphorylate a given substrate, the kinase and substrate must be present in the same location in the cell. Thus, by co-clustering the distributions of kinases and substrates among the five differential-centrifugation (DC) fractions studied, we obtain information about the likelihood that a given kinase and a given substrate interact in the cell. These data are used to rank all kinases, based on application of Bayes' rule, with regard to whether they are likely to phosphorylate the vasopressin-regulated water channel AQP2.1
METHODS
Western blotting.
Western blotting was carried out as described elsewhere (4) using DC fractions obtained as described in the accompanying article (38). Briefly, after solubilization in Laemmli buffer, mpkCCD protein samples (20 μg per lane) were resolved by SDS-PAGE using precast polyacrylamide gels (Any-kD TGX Gel, Bio-Rad, Hercules, CA) at 200 V for 45 min and transferred electrophoretically onto nitrocellulose membranes at 70 V for 60 min. The membranes were then blocked with Odyssey blocking buffer (LI-COR, Lincoln, NE), rinsed, and probed with primary antibody (rabbit anti-mouse AQP2; catalog no. NB110-74682, Novus) overnight at 4°C. The blots were washed and then incubated for 1 h with a fluorescently labeled goat anti-rabbit secondary antibody (IRDye 680RD, LI-COR). Fluorescence was imaged and quantified using the LI-COR Odyssey imaging system. Equal loading between vehicle- and vasopressin-treated samples from the same fraction was confirmed by staining an identically loaded gel run in parallel with Coomassie blue dye (Imperial protein stain, catalog no. 24615, Thermo Scientific) and imaging with the LI-COR imaging system.
Mass spectrometry.
The data analyzed in this study were from the quantitative protein mass spectrometry experiments described in the accompanying article (38). Briefly, for quantification, we used the SILAC protocol, in which cells were labeled with heavy amino acids ([13C615N4]arginine and [13C6]lysine) or light amino acids ([12C614N4]arginine and [12C6]lysine), allowing the mass spectrometer to read out the relative amounts of the heavy and light versions of the same peptide (26). The cells labeled with the heavy amino acids were treated with 0.1 nM 1-desamino-8-d-arginine-vasopressin (dDAVP) for 30 min, and the cells labeled with the light amino acids were treated with vehicle. dDAVP- and vehicle-treated samples were combined 1:1, subjected to DC, and analyzed by liquid chromatography-tandem mass spectrometry, yielding dDAVP-to-control abundance ratios for all proteins in all DC fractions. For further analysis, we used the median of dDAVP-to-control abundance ratios that mapped to the same protein. For bioinformatic analysis, we used the data set reported at https://helixweb.nih.gov/ESBL/Database/mpkFractions/proteomic_fractions_linear.html.
Gene/protein enrichment analysis: Gene Ontology Cellular Component terms in each DC fraction.
Gene Ontology (GO) Cellular Component terms for each protein were assigned using Automated Bioinformatics Extractor (ABE; http://helixweb.nih.gov/ESBL/ABE/). Enrichment for an individual GO term in a given DC fraction was calculated as the ratio of the percentage of proteins with that term in the DC fraction to the percentage in the entire data set.
Identification of proteins that change reciprocally in two or more fractions.
The change in abundance of each protein in each fraction Dij (where i indicates the protein and j the fraction) was calculated from the SILAC data. These values can be recapitulated from the data shown at https://helixweb.nih.gov/ESBL/Database/mpkFractions/proteomic_fractions_linear.html from the abundance of each protein in each fraction under the control condition (aij) and the median dDAVP-to-control abundance ratio (rij) as:
Thus, increases in a protein in a fraction are represented by positive values and decreases by negative values. (Since the values are derived from the area under the curve of ion currents from the MS1 vs. time plot, the units are nanocoulombs, although the values can best be thought of as being proportional to molecular abundance.) To identify proteins that may show reciprocal changes in two or more fractions, for further analysis, we selected the proteins that satisfied the following condition
where |ΣDij| represents the absolute value of the sum, while Σ|Dij| represents the sum of the absolute values. This selection criterion yielded 1,490 of 8,608 proteins in the full data set as “possibly translocating proteins.” To identify protein complexes that may translocate to or from one of the fractions, the Dij values were sorted for each fraction j, and the top-25th percentile (most-positive values) and bottom-25th percentile (most-negative values) were separately inserted into the Database for Annotation, Visualization, and Integrated Discovery (DAVID, National Institute of Allergy and Infectious Diseases, Bethesda, MD) functional annotation tool (15) to identify enriched GO Cellular Component terms and PANTHER molecular function terms using Fisher's exact test (threshold P = 10−2). The background for this analysis was all proteins identified in the study (8,608 RefSeq identifiers). Because a number of multiprotein complexes were identified in this analysis, negative controls were created and tested by downloading all members of selected multiprotein complexes from Comprehensive Resource of Mammalian Protein Complexes (Corum, http://mips.helmholtz-muenchen.de/genre/proj/corum/) and mapping abundance changes, i.e., Dij values, to these proteins.
Co-clustering of protein kinases and potential substrates.
Relative protein abundances of basophilic kinases (AGC and CAMK family members) and of proline-directed kinases (CMGC family members) in each fraction were extracted from the proteomic data presented in the accompanying article (38) (https://helixweb.nih.gov/ESBL/Database/mpkFractions/) normalized by the calculated molecular weight of each protein. Phosphorylation sites in mouse mpkCCD cells that change with vasopressin were downloaded from Rinschen et al. (27). Relative abundance values for kinases and potential target proteins were co-clustered by hierarchical clustering using R (Heatmap2 package). Overlap between kinases and possible substrates was quantified by forming the dot product across all fractions j, namely, Σ(akjasj) over all j, where akj is the abundance for a given kinase k in fraction j and asj is the abundance value for a given substrate s in fraction j. A higher value for dot product implies a greater colocalization.
Large-scale data integration using Bayes' rule.
We used colocalization information from the present study integrated with previously published data to identify protein kinases most likely to phosphorylate AQP2 at Ser256 and Ser261. We employed Bayes' rule to integrate the new data with prior information as described by Bradford et al. (6). We define Ai as phosphorylation of AQP2 at the indicated site by kinase i. Then, for each kinase i
where Pi(Ai) is the prior probability of Ai (before integration of the new data), and Pi(Ai|new data) is the probability of Ai updated by the new data. Pi(Bi) is the probability of Bi, and ΣPi(Bi) is the sum of probabilities of Bi over all i. At each stage of the calculation, Bi represents a new element of information about kinase i (see specific data elements in the following paragraph).
We began with a list of all 521 protein kinases in the mouse genome extracted from the UniProt/Swiss-Prot Protein Knowledgebase (http://www.uniprot.org/docs/pkinfam), giving each an equal prior probability P(A) of 1/521. We then sequentially calculated revised probabilities using the following data sets: 1) transcriptomic data from Affymetrix microarrays in mpkCCD cells (39) (Bi = kinase i mRNA expressed), 2) proteomic data from mpkCCD cells (18) (Bi = kinase i protein expressed), 3) data describing target specificities of individual protein kinase classes (7) (Bi = kinase i recognizes the amino acid sequence surrounding the phosphorylated amino acid), and 4) evidence for regulation by vasopressin (6, 31) (Bi = kinase i known to be regulated by vasopressin). In the final step, Bi = kinase i colocalizes with AQP2, allowing the colocalization data (dot products) to be used to estimate the final Bayes' probabilities for each kinase i.
RESULTS AND DISCUSSION
Distribution of subcellular marker proteins among DC fractions.
Before analysis of the proteomic data to identify protein groups that may translocate between fractions in response to vasopressin, it is pertinent to ask what structures are present in each fraction. Classically, the individual DC fractions that we analyzed in this study have been associated with particular elements of the cell (21, 25, 34). For example, the following rough associations are commonly made [summarized by Sachs et al. (29)]: the 1K fraction (unbroken cells and intact nuclei), the 4K fraction (mitochondria), the 17K fraction (endoplasmic reticulum, Golgi, endosomes, and plasma membrane), the 200Kp fraction (small intracellular vesicles and multiprotein complexes in cytoplasm, such as ribosomes and proteasomes), and the 200Ks fraction (cytosol). These expectations can be compared with an analysis of GO Cellular Component terms enriched in the various fractions (Fig. 1). In Fig. 1, the individual bars show the frequency of gene symbols associated with a given GO Cellular Component term normalized by the frequency of that term among all fractions taken together. Overall, the distribution of these terms among fractions is roughly compatible with the view summarized above, although there is a considerable overlap among fractions. Mitochondria appear to be relatively excluded from the 17K, 200Kp, and 200Ks fractions; endosomes appear to be relatively excluded from the 1K fraction; the endoplasmic reticulum appears to be relatively excluded from the 200Kp and 200Ks fractions; and the ribosome appears to be relatively excluded from the 200Ks fraction. Data for all proteins are made available as a publically accessible webpage (https://helixweb.nih.gov/ESBL/Database/mpkFractions/). Raw files, search results, and all spectra have been uploaded to PRIDE (http://www.ebi.ac.uk/pride/archive/; accession no. PXD002434).
Fig. 1.
Proteins associated with different subcellular components show distinct distributions among differential-centrifugation (DC) fractions from mouse mpkCCD cells. “Enrichment” is calculated as percentage of proteins in a given subcellular fraction that has a particular Gene Ontology (GO) Cellular Component term divided by the corresponding percentage that has that GO Cellular Component term in all proteins identified in all fractions. Bars represent the following DC fractions: 1,000-g pellet (1K), 4,000-g pellet (4K), 17,000-g pellet (17K), 200,000-g pellet (200Kp), and 200,000-g supernatant (200Ks). The term “ribosome” includes only the cytoplasmic ribosome, and not the mitochondrial ribosome. The analysis used all data from mass spectrometry-based proteomics of mouse mpkCCD cells (see https://helixweb.nih.gov/ESBL/Database/mpkFractions/).
The considerable overlap among fractions is further supported when typical markers for subcellular structures and organelles are mapped to the fractions (Fig. 2). Although the 200Ks fraction is relatively homogeneous (chiefly cytosolic proteins), the 200Kp fraction is heterogeneous and contains various multiprotein complexes, including ribosomes and proteasomes, as well as small intracellular membrane vesicles. The 17K fraction is also heterogeneous and contains not only proteins present predominantly in the plasma membrane, but also proteins characteristic of endoplasmic reticulum, Golgi apparatus, lysosomes, and endosomes. The 4K and 1K fractions are even more heterogeneous and contain mitochondrial and nuclear markers, as well as markers from various membrane structures. Interestingly, distribution of basolateral plasma membrane proteins (Atp1a1 and Cdh16) is different from distribution of apical plasma membrane proteins (Stx4a and Aqp2) and junctional proteins (Cldn4 and Crb3). Specifically, basolateral markers were predominant in the 1K fraction, while the apical and junctional markers were predominant in the 17K fraction (Fig. 2). We propose that the presence of basolateral plasma membrane markers in the 1K fraction is due to incomplete dissociation from basement membrane fragments, which are likely to be relatively dense. This observation could be useful in predicting the location of other plasma membrane proteins.
Fig. 2.
Distribution of subcellular marker proteins among DC fractions from mouse mpkCCD cells. Values are calculated as percentage of total among all fractions for each protein. Data columns represent DC fractions: 1K, 4K, 17K, 200Kp, and 200Ks. Yellow shading highlights values >10%. The analysis used data from mass spectrometry-based proteomic analysis of mouse mpkCCD cells (see https://helixweb.nih.gov/ESBL/Database/mpkFractions/).
Vasopressin-mediated redistribution of protein complexes.
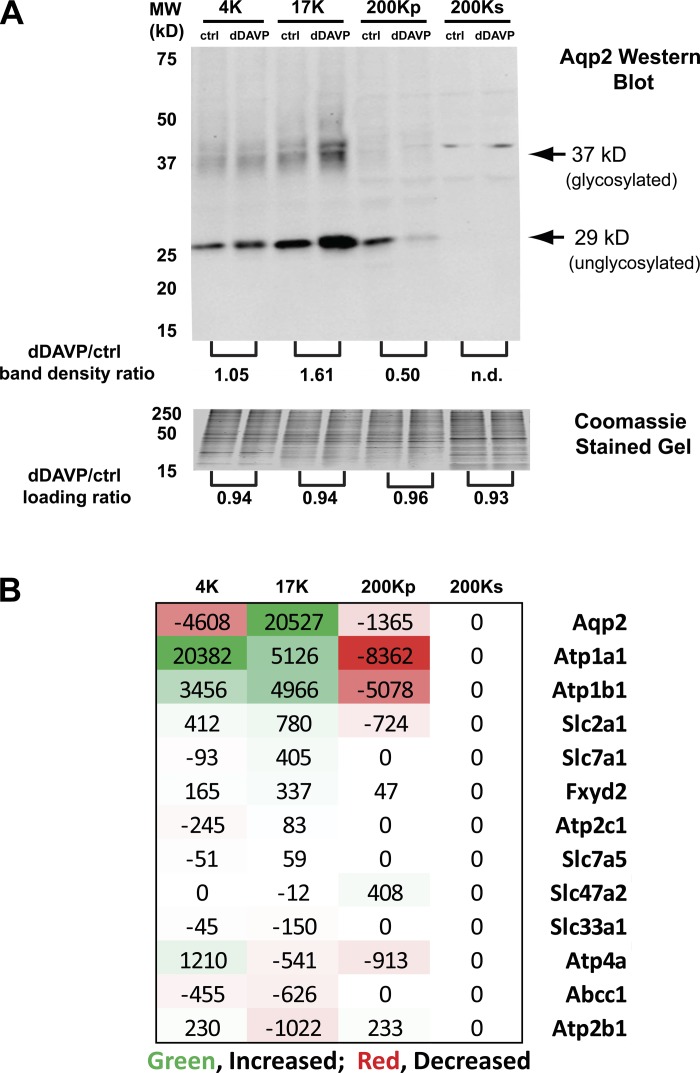
Previous studies using DC protocols similar to that used in this study have demonstrated in native collecting ducts that vasopressin triggers a shift in the subcellular distribution of AQP2, with a decrease in the 200Kp fraction and a concomitant increase in the 17K fraction (21). These studies are consistent with results involving electron microscopy (24, 37) and light microscopy (28, 37) that have shown that vasopressin increases the water permeability of collecting duct cells by triggering the translocation of AQP2-containing intracellular vesicles to fuse with the apical plasma membrane. To ensure that the mpkCCD cells used in this study respond to vasopressin in the same way, we carried out Western blotting of AQP2 in the fractions (Fig. 3A), which showed the expected shift (decrease in the 200Kp fraction and increase in the 17K fraction). We used SILAC quantification in the current experiments to identify and quantify changes in AQP2 and other proteins in the five DC fractions following addition of dDAVP [see accompanying article (38) for details]. As illustrated in Fig. 3B, the mass spectrometry experiments also showed a selective shift in AQP2 from the 200Kp to the 17K fraction, similar to the shift shown by Western blotting. Although a similar shift was seen for the two Na-K-ATPase subunits (Atp1a1 and Atp1b1), no redistribution was demonstrable for other integral membrane proteins. These data show therefore that quantitative mass spectrometry can be successfully used to detect shifts in particular proteins between subcellular fractions. Then we asked the following question: “Do other proteins undergo translocation?”
Fig. 3.
Immunoblotting and quantitative mass spectrometry confirm the previously demonstrated effect of vasopressin to cause redistribution of aquaporin-2 (AQP2) from the 200Kp fraction to the 17K fraction. A: immunoblot of AQP2 in DC fractions from mouse mpkCCD cells after treatment of the cells with the vasopressin analog 1-desamino-8-d-arginine-vasopressin (dDAVP) or vehicle (ctrl) for 30 min. Band density ratios (dDAVP/ctrl) are shown at bottom of blot. Bottom: Coomassie-stained loading gel (compressed to 40% in the vertical direction) demonstrating equal loading between vehicle (ctrl)- and dDAVP-treated samples. Mean pixel density measured for entire lane length. B: protein abundance changes from quantitative liquid chromatography-tandem mass spectroscopy in DC fractions from mouse mpkCCD cells after treatment of cells with dDAVP or vehicle (ctrl) for 30 min. Changes are in arbitrary units from total areas under the reconstructed chromatograms for light and heavy versions of each peptide using stable isotope labeling by amino acids in cell culture (SILAC). AQP2 and the 2 subunits of the Na-K-ATPase (Atp1a1 and Atp1b1) showed similar patterns, while other integral membrane proteins showed varying patterns.
Our strategy was to use the data from SILAC quantification to identify coordinated changes in specific groups of related proteins, especially components of large multiprotein complexes. Figure 4 shows specific groups of proteins (classified according to GO Cellular Component terms or PANTHER molecular function terms) whose members were increased or decreased in specific fractions out of proportion to the general population of proteins. We used the Benjamini-Hochberg test to account for multiple testing, reporting fractional false discovery rate (P). Standing out in Fig. 4 are two groups defined by the GO term “ribosome” [increased in the 17K fraction (P = 2.4 × 10−35) and decreased in the 200Kp fraction (P = 1.7 × 10−31)] and by the PANTHER term “small GTPase” [decreased in the 17K fraction (P = 5.4 × 10−22), decreased in the 200Kp fraction (P = 2.5 × 10−20), and increased in the 200Ks fraction (P = 9.8 × 10−8)]. (For all calculations, see Supplemental Data Set 1 in Supplemental Material for this article, available online at the Journal website.) The reasons for these extremely low P values can be seen when the changes in individual proteins that make up these two populations are viewed using heat-map visualizations (Figs. 5 and 6).
Fig. 4.
Protein populations that show reciprocal abundance changes in DC fractions in response to dDAVP. Proteins from https://helixweb.nih.gov/ESBL/Database/mpkFractions/ were aggregated into specific subpopulations based on GO Cellular Component terms or PANTHER molecular function terms. Change in abundance was calculated for each protein in each subpopulation using SILAC quantification data. Proportion of positive values (“increased by dDAVP”) or negative values (“decreased by dDAVP”) was compared with proportion in full data set using the Benjamini-Hochberg algorithm. P values represent fractional false discovery rates. (See Supplemental Table S1 for full calculations.) Only representative values are shown; most subpopulations tested showed no change in proportion increased or decreased, i.e., P > 10−2. See Figs. 5 and 6 for protein lists for some of these terms.
Fig. 5.
Changes in individual protein abundances in DC fractions in response to dDAVP, focusing on protein complexes involved with regulation of protein production or degradation. Proteins making up specific complexes were identified at the Corum website (http://mips.helmholtz-muenchen.de/genre/proj/corum/) or by GO terms.
Fig. 6.
Changes in individual small GTPase protein abundances in DC fractions in response to dDAVP. Note that virtually all small GTPase manifested negative abundance changes in the 17,000-g fraction. These divided into 2 groups, depending on whether the reciprocal increase in small GTPase protein abundances was in the 200,000-g supernatant (i.e., cytosol) or the 1,000-g pellet. Official gene symbols highlighted in yellow were previously identified by mass spectrometry as elements of AQP2-containing intracellular vesicles in rat inner medullary collecting duct cells (2).
Figure 5 shows all the proteins with the GO classifier ribosome (Fig. 5, left) compared with the components of several other multiprotein complexes involved in protein production and degradation in cells (Fig. 5, right). Note that virtually all proteins with the ribosome classifier (n = 65) show the same pattern, with a decrease in the 200Kp fraction and an increase in the 17K fraction. (Note: components of the mitochondrial ribosome were not included in this group and do not show this shift.) The most likely explanation for the shift in all ribosomal subunits is that fully assembled small (40S) and large (60S) ribosomal subunits are being assimilated into the rough endoplasmic reticulum in response to vasopressin. Such an explanation is consistent with the well-established view that integral membrane proteins are cotranslationally inserted into the endoplasmic reticulum, resulting in association of formerly free ribosomes with the rough endoplasmic reticulum (1, 16). This response would be expected if vasopressin increases the translation of integral membrane proteins. Indeed, in mpkCCD cells, we have shown that vasopressin increases the translation rate of AQP2 (an extremely abundant integral membrane protein) by ∼11-fold (30). A study carrying out simultaneous quantitative proteomics and transcriptomics in mpkCCD cells showed that AQP2 is, by far, the largest responder to long-term vasopressin treatment, in terms of protein abundance and transcript abundance (18). Thus the shift of ribosomes into the rough endoplasmic reticulum may simply be a consequence of increased transcription of the Aqp2 gene in response to vasopressin.
Among the other complexes shown in Fig. 5, proteins that make up the eukaryotic translation initiation factor complex 3 (eIF3) were seen to undergo a shift similar to that for ribosomal proteins. The eIF3 complex interacts with the 40S subunit of the ribosome to facilitate loading of the initiator methionine-tRNA and, subsequently, to assist in recruiting mRNAs to the ribosome (11). If the eIF3 complex is bound to ribosomes when they are recruited to the rough endoplasmic reticulum, the two complexes will presumably translocate as a unit, potentially explaining the similarity in their heat-map patterns. There is also evidence that eIF3 plays a role in STOP codon recognition (3), consistent with the idea that it remains bound to the ribosome throughout translational elongation. As seen in Fig. 5, the components of several multiprotein complexes do not show systematic changes, thus providing a negative control for the apparent translocation of ribosomes and the eIF3 complex. The protein components of two complexes involved in protein degradation, however, seem to undergo a different shift, from the 200Ks fraction to the 200Kp fraction. These were proteins with the GO Cellular Component terms “signalosome” and “proteasome.”
The COP9 signalosome is a multiprotein complex that mediates deneddylation (removal of Nedd8) of a scaffold component (one or more cullin isoforms) of ring-finger E3 ubiquitin-ligase complexes (17). A shift in COP9 signalosome subunits into the 200Kp fraction may be indicative of assembly of the complex or creation of a combined ubiquitin E3 ligase-signalosome complex in response to vasopressin. Either would tend to reduce the activity of the involved ubiquitin E3 ligases and stabilize proteins by reducing the rate of ubiquitylation. The proteasome is a multisubunit protease complex that is responsible for the degradation of cytosolic proteins. Appearance in the 200,000-g pellet in response to vasopressin may be indicative of assembly of additional proteasomal complexes. A number of proteins undergo changes in half-life in response to vasopressin (30), and it is possible that the changes in stability are related to alterations in ubiquitylation via signalosome changes or in proteasomal activity.
Figure 6 shows the dDAVP-induced changes in the relative abundances of proteins associated with the PANTHER molecular function term “small GTPase.” Remarkably, all identified small GTPase proteins show decreases in the 17,000-g pellet. They can be divided into two groups based on whether there is a corresponding increase in the 200,000-g supernatant (Fig. 6, left) or the 1,000-g pellet (Fig. 6, right). Small GTPases are molecular switches that provide points of regulation. Many of them, including Rab proteins, Arf proteins, and dynamins, are involved in regulation of membrane trafficking (33) and, therefore, have relevance to the mechanism by which vasopressin regulates the trafficking of AQP2 in the collecting duct. For example, cAMP-induced redistribution of AQP2 to the plasma membrane depends on inhibition of Rho family proteins (35). In previous studies, immunoisolation of AQP2-containing vesicles from rat inner medullary collecting duct cells followed by mass spectrometry identified a large number of small GTP-binding proteins in the Rab and Arf families with hypothetical roles in AQP2 trafficking (2). The parallel movement of multiple small GTPase proteins out of the 17,000-g pellet implies a general mechanism of regulation such as activation of one or more GTPase-activating proteins, since small GTPase proteins, unlike the ribosomal subunits, probably do not form a single complex that would explain tandem movement.
Another protein group that undergoes a coordinated change in two fractions is defined by the GO Cellular Component term “clathrin coat” (Fig. 4). This group consists of 10 proteins, including clathrin heavy chain (gene symbol Cltc) and several members of the AP-1 and AP-2 clathrin adaptor complexes (Ap1b1, Ap1g1, Ap1g2, Ap1m1, Ap1m2, Ap2a1, Ap2a2, Ap2b1, and Ap2m1). Each of these 10 proteins undergoes an increase in the 17,000-g pellet and a concomitant decrease in the 200,000-g supernatant in response to dDAVP. These clathrin adaptor complexes are responsible for recruitment of clathrin to the trans-Golgi complex and recycling endosomes (AP-1) and the plasma membrane (AP-2) as a prelude to vesicle budding (5). Consequently, the demonstrated shift is consistent with organization of clathrin coats in trans-Golgi, recycling endosomes, and plasma membranes in response to vasopressin. The clathrin adaptor complexes are also responsible for recognition of cargo proteins through specific amino acid sequences in their COOH-terminal tails. Interestingly, members of two other clathrin adaptor complexes, AP-3 and AP-4, did not show coordinated shifts.
Distribution of protein kinases and their substrates among fractions.
An important question in the physiology of the renal collecting duct is “What are the signaling pathways activated by vasopressin?” Previously, we carried out comprehensive phosphoproteomic studies to identify phosphorylation sites in mpkCCD proteins that change in response to vasopressin (4, 27). The protein kinases and/or phosphatases responsible for these phosphorylation changes have not been identified. In the present study we identified 246 protein kinases among all fractions that, in theory, could be involved in vasopressin signaling (see Supplemental Data Set 2). This number is 47% of the 521 kinases encoded by the mouse genome. The protein kinases responsible for vasopressin-dependent phosphorylation in mpkCCD cells can be predicted through integration of multiple data types reflecting kinase specificities, kinase expression levels in mpkCCD cells, and the regulatory characteristics of individual kinases (6). One factor that has not been taken into account is the subcellular localization of individual kinases, vis-à-vis the localization of their substrates. In theory, if a kinase and a potential substrate are not found in the same subcellular fraction(s), a functional interaction is less likely than if they are found in the same fractions. Here, we used the data set developed in this study to carry out this comparison (Figs. 7–10).
Fig. 7.
Co-clustering of subcellular distributions of basophilic kinases (black) and potential substrates (red) reveals colocalized kinase-substrate pairs. Basophilic kinases listed are members of the AGC or CAMK protein kinase families found in this study to be expressed in mouse mpkCCD cells. Substrates listed are proteins with vasopressin-regulated phosphorylation sites from Rinschen et al. (27) that are present in motifs compatible with phosphorylation by a basophilic kinase. Clustering was by hierarchical clustering using R (Heatmap2 package) represented by the tree at left.
Fig. 10.
Extent of subcellular colocalization of AQP2 protein and proline-directed kinases expressed in mouse mpkCCD cells based on dot product of distribution vectors. Distribution vectors are formed from relative abundance values in each of the 5 fractions obtained from Fig. 9. Colocalized proline-directed kinases are candidates to phosphorylate Ser261 of AQP2 in mouse mpkCCD cells.
Figure 7 shows a heat-map presentation of the subcellular distributions of the 62 expressed basophilic kinases clustered together with all regulated basophilic targets from Rinschen et al. (27). The protein kinases clustered into four distinct groups that define their likely domains of actions. The fourth cluster (Fig. 7, bottom) includes AQP2 as a substrate and 12 basophilic protein kinases. Figure 8 ranks the overlap between AQP2 localization and all basophilic kinases based on the dot products of their distributions among fractions (see Supplemental Data Set 3). The top-ranked kinases were calcium/calmodulin-dependent protein kinase IIδ (Camk2d) and AMP kinase-α1 (Prkaa1), followed by microtubule affinity-regulating kinase 2 (Mark2), CDC42-binding protein kinase-β (CDC42bpb), three more calmodulin kinase II isoforms, protein kinase A catalytic-α (Prkaca), and protein kinase Cι (Prkci). These kinases, therefore, may be considered to be candidates for roles in phosphorylation of AQP2 at Ser256, Ser264, and Ser269, which have basophilic signatures.
Fig. 8.
Extent of subcellular colocalization of AQP2 protein and basophilic kinases expressed in mouse mpkCCD cells based on dot product of distribution vectors provides a ranking of kinases. Distribution vectors are formed from relative abundance values in each of the 5 fractions obtained from Fig. 7. Only the top-40 basophilic kinases are shown. All others manifested no substantial colocalization. Colocalized basophilic kinases are candidates to phosphorylate Ser256, Ser264, and Ser269 of AQP2.
Figure 9 shows a heat-map representation of the distribution of expressed proline-directed kinases (CMGC family kinases) clustered together with all regulated proline-directed targets from Rinschen et al. (27). All these phosphorylation sites have a proline in position +1 relative to the phosphorylated amino acid. Ser261 of AQP2 is one of these targets and, like most of the other proline-directed sites, undergoes decreased phosphorylation in response to vasopressin (14). In Fig. 9, AQP2 clustered with four proline-directed kinases. Figure 10 ranks the overlap between AQP2 localization and all proline-directed kinases on the basis of the dot product of their distributions among fractions (see Supplemental Data Set 4). The top-ranked kinases based on dot product were cyclin-dependent kinase 1 (Cdk1), ERK1 (Mapk3), and ERK2 (Mapk1). Prior studies have provided evidence for a role for ERK1/ERK2 (13) and/or Cdk1 (27, 36) in the phosphorylation of AQP2 at Ser261. However, many additional proline-directed kinases (MAP kinases and cyclin-dependent kinases) were also present in the 17,000- and 4,000-g membrane fractions, where most of the AQP2 is found (Fig. 9, top). This includes several MAP kinases previously proposed to phosphorylate AQP2, JNK isoforms (Mapk8, Mapk9, and Mapk10) (27), and p38 isoforms (Mapk11, Mapk12, Mapk13, and Mapk14) (23, 27). Thus the protein kinase(s) responsible for phosphorylation of AQP2 at Ser261 appears to be an unsettled question.
Fig. 9.
Co-clustering of subcellular distributions of proline-directed kinases (black) and potential substrates (red) reveals colocalized kinase-substrate pairs. Proline-directed kinases listed are members of the CMGC protein kinase family that were found in this study to be expressed in mouse mpkCCD cells. Substrates listed are proteins in mouse mpkCCD cells with vasopressin-regulated phosphorylation sites from Rinschen et al. (27) that are present in motifs compatible with phosphorylation by a proline-directed kinase, i.e., with a proline in position +1 relative to the phosphorylated amino acid. Clustering was by hierarchical clustering using R (Heatmap2 package) represented by the tree at left.
Using Bayes' rule to rank protein kinases that may phosphorylate AQP2 in mpkCCD cells.
Previously, using Bayes' rule, we integrated data from a variety of sources to identify the protein kinases in the rat renal inner medullary collecting duct (IMCD) that are most likely to phosphorylate AQP2 at Ser256 (6). This phosphorylation site is a “keystone” site that must be phosphorylated prior to vasopressin-stimulated phosphorylation of downstream sites at Ser264 and Ser269 (12). The objective is to use data sets specific to mpkCCD cells to ascertain whether the top-ranked kinases match those of the rat IMCD. These data sets include 1) transcriptomic data, giving relative abundances of all expressed mRNAs (39), 2) proteomic data, providing lists of expressed proteins (18; present study), 3) substrate specificity data for kinase families (7), and 4) evidence from the literature for regulation of individual kinases by vasopressin (6, 31). To this, we add the data from Distribution of protein kinases and their substrates among fractions, describing the degree by which each individual kinase overlaps the subcellular distribution of AQP2. We assume that if the overlap is greater between substrate and kinase, the probability of a functional interaction is greater. Figure 11 shows the ranking results for all 521 protein kinases in the mouse genome with regard to likelihood of phosphorylating Ser256 of AQP2 (see Supplemental Data Set 5). The analysis resulted in the separation of a relatively small number of kinases from the entire population. Table 1 lists the 10 top-ranked kinases before and after the addition of colocalization data from the present study. The list for mpkCCD cells overlaps extensively with the list derived from investigations in rat native IMCD cells published by Bradford et al. (6). Both contain products of calcium/calmodulin-dependent protein kinase IIβ and IIδ genes, protein kinase Aα and Aβ genes, protein kinase Cδ and Cι genes, microtubule affinity-regulating kinase 2, and CDC42-binding protein kinase-β. The coincidence of these kinases from the Bayes' rule-based analyses of independent data sets from mouse mpkCCD cells and from native rat IMCD cells creates a particularly strong argument in favor of their possible roles in regulation of AQP2 in the collecting duct. Thus the systems approach described sets up future reductionist experiments aimed at resolving the roles of each of these eight protein kinases in vasopressin signaling in the collecting duct, a much easier task than if all 521 protein kinases needed to be screened.
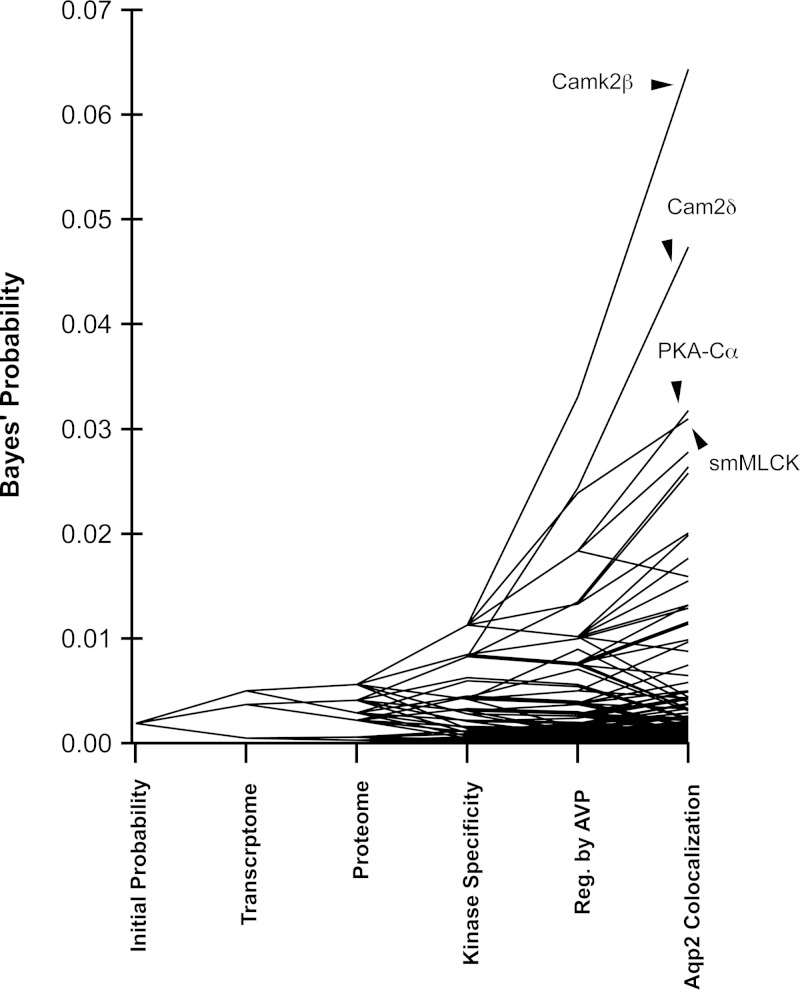
Fig. 11.
Result of large-scale data integration regarding the following question: “What protein kinase is most likely to phosphorylate AQP2 at Ser256 in mouse mpkCCD cells?” Starting with all 521 protein kinases in the mouse genome set to “equal probabilities,” Bayes' theorem was used to incorporate the following information (in order): transcript abundances in mpkCCD (“transcriptome”), presence or absence in proteomic studies of mpkCCD cells (“proteome”), degree of match between kinase family target specificity and AQP2 sequence surrounding Ser256 (“kinase specificity”), evidence for regulation by vasopressin (“Reg by AVP”), and colocalization with AQP2 using the dot products from Fig. 8 (“AQP2 colocalization”). See text for references. The top-ranking protein kinases are indicated. See Supplemental Data Set 5 for detail of calculations.
Table 1.
Ranking of mpkCCD kinases with regard to likelihood of a role in phosphorylation of Ser256 of aquaporin-2, based on Bayesian integration of multiple data sets
| Mouse mpkCCD Cells |
||||||||
|---|---|---|---|---|---|---|---|---|
| From prior data* |
After colocalization data† |
Rat IMCD‡ |
||||||
| Rank | Kinase | Bayes' probability | Rank | Kinase | Bayes' probability | Rank | Kinase | Bayes' probability |
| 1 | Camk2b | 0.033 | 1 | Camk2b | 0.064 | 1 | Camk2d | 0.045 |
| 2 | Camk2d | 0.024 | 2 | Camk2d | 0.047 | 2 | Akt1 | 0.025 |
| 2 | Mylk | 0.024 | 3 | Prkaca | 0.032 | 2 | Prkaca | 0.025 |
| 4 | Prkaca | 0.018 | 4 | Mylk | 0.031 | 2 | Prkacb | 0.025 |
| 4 | Prkacb | 0.018 | 5 | Prkacb | 0.028 | 5 | Cdc42bpb | 0.018 |
| 4 | Akt1 | 0.018 | 6 | Camk2 g | 0.026 | 5 | Prkcd | 0.018 |
| 7 | Camk2 g | 0.014 | 6 | Cdc42bpb | 0.026 | 7 | Cask | 0.014 |
| 8 | Cdc42bpb | 0.013 | 8 | Prkcd | 0.02 | 7 | Dapk3 | 0.014 |
| 8 | Prkcd | 0.013 | 8 | Mark2 | 0.02 | 7 | Mark2 | 0.014 |
| 10 | Mark2 | 0.01 | 10 | Prkci | 0.018 | 7 | Mark3 | 0.014 |
| 10 | Nuak2 | 0.01 | 7 | Nuak2 | 0.014 | |||
| 10 | Pkn2 | 0.01 | 7 | Pdpk1 | 0.014 | |||
| 10 | Prkci | 0.01 | 7 | Pkn1 | 0.014 | |||
| 10 | Rock2 | 0.01 | 7 | Pkn2 | 0.014 | |||
| 10 | Rps6ka1 | 0.01 | 7 | Prkci | 0.014 | |||
| 10 | Trio | 0.01 | 7 | Rock2 | 0.014 | |||
| 10 | Mark3 | 0.01 | 7 | Rps6ka1 | 0.014 | |||
| 10 | Pdpk1 | 0.01 | ||||||
| 10 | Mknk2 | 0.01 | ||||||
| 10 | Stk38 | 0.01 | ||||||
| 10 | Rock1 | 0.01 | ||||||
| 10 | Stk11 | 0.01 | ||||||
| 10 | Cdc42bpg | 0.01 | ||||||
| 10 | Chek1 | 0.01 | ||||||
| 10 | Melk | 0.01 | ||||||
| 10 | Rps6ka4 | 0.01 | ||||||
See Supplemental Data Set 5 in Supplemental Material for this article (available online at the Journal website) for details of calculations. IMCD, inner medullary collecting duct.
From experiments in mpkCCD cells as described in text.
From the present study.
From Bradford et al. (6).
Figure 12 and Table 2 show the results for a similar Bayesian analysis of what protein kinases might phosphorylate AQP2 at Ser261 (see Supplemental Data Set 6). Again, the analysis narrowed the list to relatively few proline-directed kinases (Fig. 12). The addition of colocalization data from the present study sharpened the separation of top-ranked kinases. ERK1 (Mapk3), ERK2 (Mapk1), and p38α (Mapk14) ranked highest (Fig. 12), concordant with prior evidence (13, 23, 27). These were followed by Cdk2, Cdk4, and Mapk13.
Fig. 12.
Result of large-scale data integration regarding the following question: “What protein kinase is most likely to phosphorylate AQP2 at Ser261 in mouse mpkCCD cells?” Starting with all 521 protein kinases in the mouse genome set to “equal probabilities,” Bayes' theorem was used to incorporate the following information (in order): transcript abundances in mpkCCD (“transcriptome”), presence or absence in proteomic studies of mpkCCD cells (“proteome”), degree of match between kinase family target specificity and AQP2 sequence surrounding Ser261 (“kinase specificity”), evidence for regulation by vasopressin (“Reg by AVP”), and colocalization with AQP2 using the dot products from Fig. 10 (“AQP2 colocalization”). See text for references. Top-ranking protein kinases are indicated. See Supplemental Data Set 6 for details of calculations.
Table 2.
Ranking of mpkCCD kinases with regard to likelihood of a role in phosphorylation of Ser261 of aquaporin-2, based on Bayesian integration of multiple data sets
| Mouse mpkCCD Cells* |
Mouse mpkCCD Cells (after colocalization data)† |
||||
|---|---|---|---|---|---|
| Rank | Kinase | Bayes' probability | Rank | Kinase | Bayes' probability |
| 1 | Mapk1 | 0.0237 | 1 | Mapk1 | 0.0393 |
| 1 | Mapk3 | 0.0237 | 1 | Mapk3 | 0.0393 |
| 1 | Mapk14 | 0.0237 | 3 | Mapk14 | 0.0349 |
| 4 | Gsk3a | 0.0174 | 4 | Cdk2 | 0.0218 |
| 5 | Cdk2 | 0.0131 | 4 | Cdk4 | 0.0218 |
| 5 | Cdk4 | 0.0131 | 4 | Mapk13 | 0.0218 |
| 5 | Cdk6 | 0.0131 | 7 | Srpk1 | 0.0194 |
| 5 | Cdk8 | 0.0131 | 7 | Nlk | 0.0194 |
| 5 | Cdk9 | 0.0131 | 7 | Gsk3b | 0.0194 |
| 5 | CdkL1 | 0.0131 | 7 | Cdk9 | 0.0194 |
| 5 | Clk1 | 0.0131 | 7 | Cdk6 | 0.0194 |
| 5 | Clk2 | 0.0131 | 7 | Mapk12 | 0.0194 |
| 5 | Dyrk1a | 0.0131 | |||
| 5 | Gsk3b | 0.0131 | |||
| 5 | Mapk12 | 0.0131 | |||
| 5 | Mapk13 | 0.0131 | |||
| 5 | Mapk6 | 0.0131 | |||
| 5 | Nlk | 0.0131 | |||
| 5 | Prpf4b | 0.0131 | |||
| 5 | Srpk1 | 0.0131 | |||
See Supplemental Data Set 6 for details of calculations.
From experiments in mpkCCD cells as described in text.
From the present study.
Conclusion
Systems biology provides a set of tools (e.g., protein mass spectrometry, deep sequencing of DNA, and computational methods) that allows all components of a biological system to be studied together in parallel to discover integrative function (19). A prerequisite is the identification of all components of the system, which must be achieved before the components are linked into network models. The comprehensive proteomics data presented in the accompanying article, curated from 35 gigabytes of spectral data, bring us close to knowing the entire expressed proteome in mpkCCD cells and is expected to be a useful resource for study of the regulation of water and solute transport in the mammalian collecting duct. In the present study we exploited the data via bioinformatics analysis to address two questions pertinent to vasopressin signaling in the collecting duct: 1) “Does vasopressin trigger translocation of specific subpopulations of proteins between subcellular fractions in the mpkCCD cell?” and 2) “What protein kinases colocalize with the vasopressin-regulated water channel AQP2 in the renal collecting duct and, consequently, are candidates for mediating AQP2 phosphorylation?” Our answers to these questions, provide grist for further studies using systems biology approaches and classical reductionist approaches to understand how vasopressin regulates the function of collecting duct cells.
GRANTS
This study was carried out in the National Heart, Lung, and Blood Institute Intramural Program (Projects HL-006129 and HL-001285, M. A. Knepper). The authors are affiliated with the National, Heart, Lung, and Blood Institute Division of Intramural Research and receive funding support through the intramural budget (Projects HL-006129 “Computational Tools for Proteomics” and HL-001285 “Solute and Water Transport in Renal Epithelia”).
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the authors.
AUTHOR CONTRIBUTIONS
C.-R.Y., V.R., and M.A.K. developed the concept and designed the research; C.-R.Y. and M.E. performed the experiments; C.-R.Y., V.R., M.E., P.C.S., and M.A.K. analyzed the data; C.-R.Y., V.R., M.E., P.C.S., and M.A.K. interpreted the results of the experiments; C.-R.Y., V.R., M.E., and M.A.K. prepared the figures; C.-R.Y., V.R., and M.A.K. drafted the manuscript; C.-R.Y., V.R., M.E., P.C.S., and M.A.K. edited and revised the manuscript; C.-R.Y., V.R., M.E., P.C.S., and M.A.K. approved the final version of the manuscript.
Supplementary Material
Footnotes
This article is the topic of an Editorial Focus by M. M. Rinschen (26a).
REFERENCES
- 1.Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular Biology of the Cell. New York: Garland Science, 2002. [Google Scholar]
- 2.Barile M, Pisitkun T, Yu MJ, Chou CL, Verbalis MJ, Shen RF, Knepper MA. Large scale protein identification in intracellular aquaporin-2 vesicles from renal inner medullary collecting duct. Mol Cell Proteomics 4: 1095–1106, 2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Beznoskova P, Cuchalova L, Wagner S, Shoemaker CJ, Gunisova S, von der HT, Valasek LS. Translation initiation factors eIF3 and HCR1 control translation termination and stop codon read-through in yeast cells. PLos Genet 9: e1003962, 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bolger SJ, Hurtado PA, Hoffert JD, Saeed F, Pisitkun T, Knepper MA. Quantitative phosphoproteomics in nuclei of vasopressin-sensitive renal collecting duct cells. Am J Physiol Cell Physiol 303: C1006–C1020, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bonifacino JS. Adaptor proteins involved in polarized sorting. J Cell Biol 204: 7–17, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bradford D, Raghuram V, Wilson JL, Chou CL, Hoffert JD, Knepper MA, Pisitkun T. Use of LC-MS/MS and Bayes' theorem to identify protein kinases that phosphorylate aquaporin-2 at Ser256. Am J Physiol Cell Physiol 307: C123–C139, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Caenepeel S, Charydczak G, Sudarsanam S, Hunter T, Manning G. The mouse kinome: discovery and comparative genomics of all mouse protein kinases. Proc Natl Acad Sci USA 101: 11707–11712, 2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Christensen BM, Zelenina M, Aperia A, Nielsen S. Localization and regulation of PKA-phosphorylated AQP2 in response to V2-receptor agonist/antagonist treatment. Am J Physiol Renal Physiol 278: F29–F42, 2000. [DOI] [PubMed] [Google Scholar]
- 9.Fenton RA, Moeller HB, Hoffert JD, Yu MJ, Nielsen S, Knepper MA. Acute regulation of aquaporin-2 phosphorylation at Ser-264 by vasopressin. Proc Natl Acad Sci USA 105: 3134–3139, 2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hasler U, Vinciguerra M, Vandewalle A, Martin PY, Feraille E. Dual effects of hypertonicity on aquaporin-2 expression in cultured renal collecting duct principal cells. J Am Soc Nephrol 16: 1571–1582, 2005. [DOI] [PubMed] [Google Scholar]
- 11.Hinnebusch AG. The scanning mechanism of eukaryotic translation initiation. Annu Rev Biochem 83: 779–812, 2014. [DOI] [PubMed] [Google Scholar]
- 12.Hoffert JD, Fenton RA, Moeller HB, Simons B, Tchapyjnikov D, McDill BW, Yu MJ, Pisitkun T, Chen F, Knepper MA. Vasopressin-stimulated increase in phosphorylation at Ser269 potentiates plasma membrane retention of aquaporin-2. J Biol Chem 283: 24617–24627, 2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hoffert JD, Nielsen J, Yu MJ, Pisitkun T, Schleicher SM, Nielsen S, Knepper MA. Dynamics of aquaporin-2 serine-261 phosphorylation in response to short-term vasopressin treatment in collecting duct. Am J Physiol Renal Physiol 292: F691–F700, 2007. [DOI] [PubMed] [Google Scholar]
- 14.Hoffert JD, Pisitkun T, Wang G, Shen RF, Knepper MA. Quantitative phosphoproteomics of vasopressin-sensitive renal cells: regulation of aquaporin-2 phosphorylation at two sites. Proc Natl Acad Sci USA 103: 7159–7164, 2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4: 44–57, 2009. [DOI] [PubMed] [Google Scholar]
- 16.Jan CH, Williams CC, Weissman JS. Principles of ER cotranslational translocation revealed by proximity-specific ribosome profiling. Science 346: 1257521, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kato JY, Yoneda-Kato N. Mammalian COP9 signalosome. Genes Cells 14: 1209–1225, 2009. [DOI] [PubMed] [Google Scholar]
- 18.Khositseth S, Pisitkun T, Slentz DH, Wang G, Hoffert JD, Knepper MA, Yu MJ. Quantitative protein and mRNA profiling shows selective post-transcriptional control of protein expression by vasopressin in kidney cells. Mol Cell Proteomics 10: M110, 2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Knepper MA. Systems biology in physiology: the vasopressin signaling network in kidney. Am J Physiol Cell Physiol 303: C1115–C1124, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Li Y, Shaw S, Kamsteeg EJ, Vandewalle A, Deen PM. Development of lithium-induced nephrogenic diabetes insipidus is dissociated from adenylyl cyclase activity. J Am Soc Nephrol 17: 1063–1072, 2006. [DOI] [PubMed] [Google Scholar]
- 21.Marples D, Knepper MA, Christensen EI, Nielsen S. Redistribution of aquaporin-2 water channels induced by vasopressin in rat kidney inner medullary collecting duct. Am J Physiol Cell Physiol 269: C655–C664, 1995. [DOI] [PubMed] [Google Scholar]
- 22.Moeller HB, Knepper MA, Fenton RA. Serine 269 phosphorylated aquaporin-2 is targeted to the apical membrane of collecting duct principal cells. Kidney Int 75: 295–303, 2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nedvetsky PI, Tabor V, Tamma G, Beulshausen S, Skroblin P, Kirschner A, Mutig K, Boltzen M, Petrucci O, Vossenkamper A, Wiesner B, Bachmann S, Rosenthal W, Klussmann E. Reciprocal regulation of aquaporin-2 abundance and degradation by protein kinase A and p38-MAP kinase. J Am Soc Nephrol 21: 1645–1656, 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Nielsen S, Chou CL, Marples D, Christensen EI, Kishore BK, Knepper MA. Vasopressin increases water permeability of kidney collecting duct by inducing translocation of aquaporin-CD water channels to plasma membrane. Proc Natl Acad Sci USA 92: 1013–1017, 1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Novikoff AB. Electron microscopy: cytology of cell fractions. Science 124: 969–972, 1956. [DOI] [PubMed] [Google Scholar]
- 26.Ong SE, Blagoev B, Kratchmarova I, Kristensen DB, Steen H, Pandey A, Mann M. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteomics 1: 376–386, 2002. [DOI] [PubMed] [Google Scholar]
- 26a.Rinschen MM. Water transport running deep. Focus on “Deep proteomic profiling of vasopressin-sensitive collecting duct cells.” Am J Physiol Cell Physiol (September 30, 2015). doi: 10.1152/ajpcell.00280.2015. [DOI] [PubMed] [Google Scholar]
- 27.Rinschen MM, Yu MJ, Wang G, Boja ES, Hoffert JD, Pisitkun T, Knepper MA. Quantitative phosphoproteomic analysis reveals vasopressin V2-receptor-dependent signaling pathways in renal collecting duct cells. Proc Natl Acad Sci USA 107: 3887, 2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sabolic I, Katsura T, Verbabatz JM, Brown D. The AQP2 water channel: effect of vasopressin treatment, microtubule disruption, and distribution in neonatal rats. J Membr Biol 143: 165–177, 1995. [DOI] [PubMed] [Google Scholar]
- 29.Sachs AN, Pisitkun T, Hoffert JD, Yu MJ, Knepper MA. LC-MS/MS analysis of differential centrifugation fractions from native inner medullary collecting duct of rat. Am J Physiol Renal Physiol 295: F1799–F1806, 2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sandoval PC, Slentz DH, Pisitkun T, Saeed F, Hoffert JD, Knepper MA. Proteome-wide measurement of protein half-lives and translation rates in vasopressin-sensitive collecting duct cells. J Am Soc Nephrol 24: 1793–1805, 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sanghi A, Zaringhalam M, Corcoran CC, Saeed F, Hoffert JD, Sandoval P, Pisitkun T, Knepper MA. A knowledge base of vasopressin actions in the kidney. Am J Physiol Renal Physiol 307: F747–F755, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Schenk LK, Bolger SJ, Luginbuhl K, Gonzales PA, Rinschen MM, Yu MJ, Hoffert JD, Pisitkun T, Knepper MA. Quantitative proteomics identifies vasopressin-responsive nuclear proteins in collecting duct cells. J Am Soc Nephrol 23: 1008–1018, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Segev N. GTPases in intracellular trafficking: an overview. Semin Cell Dev Biol 22: 1–2, 2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Siekevitz P, Palade GE. A cytochemical study on the pancreas of the guinea pig. I. Isolation and enzymatic activities of cell fractions. J Biophys Biochem Cytol 4: 203–218, 1958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tamma G, Klussmann E, Procino G, Svelto M, Rosenthal W, Valenti G. cAMP-induced AQP2 translocation is associated with RhoA inhibition through RhoA phosphorylation and interaction with RhoGDI. J Cell Sci 116: 1519–1525, 2003. [DOI] [PubMed] [Google Scholar]
- 36.Tamma G, Lasorsa D, Trimpert C, Ranieri M, Di MA, Mola MG, Mastrofrancesco L, Devuyst O, Svelto M, Deen PM, Valenti G. A protein kinase A-independent pathway controlling aquaporin 2 trafficking as a possible cause for the syndrome of inappropriate antidiuresis associated with polycystic kidney disease 1 haploinsufficiency. J Am Soc Nephrol 25: 2241–2253, 2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yamamoto N, Sasaki S, Fushimi K, Ishibashi K, Yaiota E, Kawasaki K, Marumo F, Kihara I. Vasopressin increases AQP-CD water channel in the apical membrane of collecting duct cells without affecting AQP3 distribution in Brattleboro rat. Am J Physiol Cell Physiol 268: C1546–C1551, 1995. [DOI] [PubMed] [Google Scholar]
- 38.Yang CR, Tongyoo P, Emamian M, Sandoval PC, Raghuram V, Knepper MA. Deep proteomic profiling of vasopressin-sensitive collecting duct cells. I. Virtual Western blots and molecular weight distributions. Am J Physiol Cell Physiol (August 26, 2015). doi: 10.1152/ajpcell.00213.2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yu MJ, Miller RL, Uawithya P, Rinschen MM, Khositseth S, Braucht DW, Chou CL, Pisitkun T, Nelson RD, Knepper MA. Systems-level analysis of cell-specific AQP2 gene expression in renal collecting duct. Proc Natl Acad Sci USA 106: 2441–2446, 2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.