Abstract
The multifunctional Saccharomyces cerevisiae Pif1 DNA helicase affects the maintenance of telomeric, ribosomal, and mitochondrial DNAs, suppresses DNA damage at G-quadruplex motifs, influences the processing of Okazaki fragments, and promotes breakage induced replication. All of these functions require the ATPase/helicase activity of the protein. Owing to Pif1’s critical role in the maintenance of mitochondrial DNA, pif1Δ strains quickly generate respiratory deficient cells and hence grow very slowly. This slow growth makes it difficult to carry out genome-wide synthetic genetic analysis in this background. Here, we used a partial loss of function allele of PIF1, pif1-m2, which is mitochondrial proficient but has reduced abundance of nuclear Pif1. Although pif1-m2 is not a null allele, pif1-m2 cells exhibit defects in telomere maintenance, reduced suppression of damage at G-quadruplex motifs and defects in breakage induced replication. We performed a synthetic screen to identify nonessential genes with a synthetic sick or lethal relationship in cells with low abundance of nuclear Pif1. This study identified eleven genes that were synthetic lethal (APM1, ARG80, CDH1, GCR1, GTO3, PRK1, RAD10, SKT5, SOP4, UMP1, and YCK1) and three genes that were synthetic sick (DEF1, YIP4, and HOM3) with pif1-m2.
Keywords: Pif1, helicase, synthetic lethality, yeast, S. cerevisiae
Introduction
Pif1 family DNA helicases are found in all three kingdoms (reviewed in Bochman et al. 2010) . The best studied of these helicases is the founding member of the family, the Saccharomyces cerevisiae Pif1. There are two isoforms of Pif1 that depend on whether the first or second methionine is used to initiate translation of the mRNA (Schulz and Zakian 1994; Zhou et al. 2000). One isoform is targeted to the mitochondria (mt) where it is critical for the maintenance of mtDNA and for respiratory competence. The second isoform is localized to the nucleus and functions in multiple pathways that affect genome integrity. Pif1 is a negative regulator of telomere lengthening and de novo telomere addition by virtue of its ability to displace telomerase from DNA ends (Schulz and Zakian 1994; Boule et al. 2005; Phillips et al. 2015). It is required to generate long flap Okazaki fragments (Pike et al. 2009) and to promote breakage induced replication (Saini et al. 2013; Wilson et al. 2013). Pif1 promotes DNA replication through G-quadruplex (G4) motifs, which are sequences that form G4 structures in vitro, and suppresses DNA damage at G4 motifs (Ribeyre et al. 2009; Paeschke et al. 2011, 2013; Piazza et al. 2012). Additionally, Pif1 helps maintain the replication fork barrier (RFB) in the ribosomal DNA (rDNA) (Ivessa et al. 2002). Although Pif1 has weak unwinding activity on conventional 5′ tailed duplex DNA substrates, it robustly unwinds G4 structures and RNA/DNA hybrids in vitro (Boule and Zakian 2007; Ribeyre et al. 2009; Paeschke et al. 2011; Zhou et al. 2014).
Despite its multiple and diverse functions, PIF1 is not an essential gene. We anticipated that other genes might act in parallel with PIF1 to carry out its various cellular functions. S. cerevisiae encodes a second Pif1 family helicase, Rrm3, whose helicase domain is 40% identical to that of Pif1. However, the functions of Rrm3 and Pif1 are largely nonoverlapping, except at G4 motifs (Paeschke et al. 2013). Rrm3 does not appear to be a backup for Pif1 at many of its genomic targets (Ivessa et al. 2000, 2002; O’Rourke et al. 2005). We predicted that PIF1 might have synthetic interactions with genes involved in regulating telomere length, Okazaki fragment maturation, breakage induced replication, and G-quadruplex unwinding. Additionally, because Pif1 binds in vivo to the promoters of ∼130 genes (C. F. Chen, S. Pott, and V. A. Zakian, unpublished results), Pif1 might have as yet undescribed roles in transcriptional regulation, which could result in interactions with transcription factors. We anticipated that we might detect indirect synthetic lethal relationships owing to Pif1’s effect on gene expression and/or genome integrity. In addition, as pif1-m2 cells are more sensitive to proteasomal inhibition and have a higher basal level of autophagy, pif1-m2 cells may be more dependent on the proteasome for cellular maintenance and survival (J. L. Stundon and V. A. Zakian, unpublished results). Thus, pif1-m2 might have synthetic interactions with other genes with roles in autophagy and proteasomal function.
Rationale for screen
As we are particularly interested in the nuclear functions of Pif1, we sought to identify genes whose deletion affected the viability of or reduced the growth rate of pif1-m2 cells, which are deficient in the nuclear form of Pif1 (Schulz and Zakian 1994; Zhou et al. 2000). This strategy avoided the difficulty of using pif1Δ cells, which are very slow growing, behavior that might obscure synthetic interactions.
Materials and Methods
Screen design
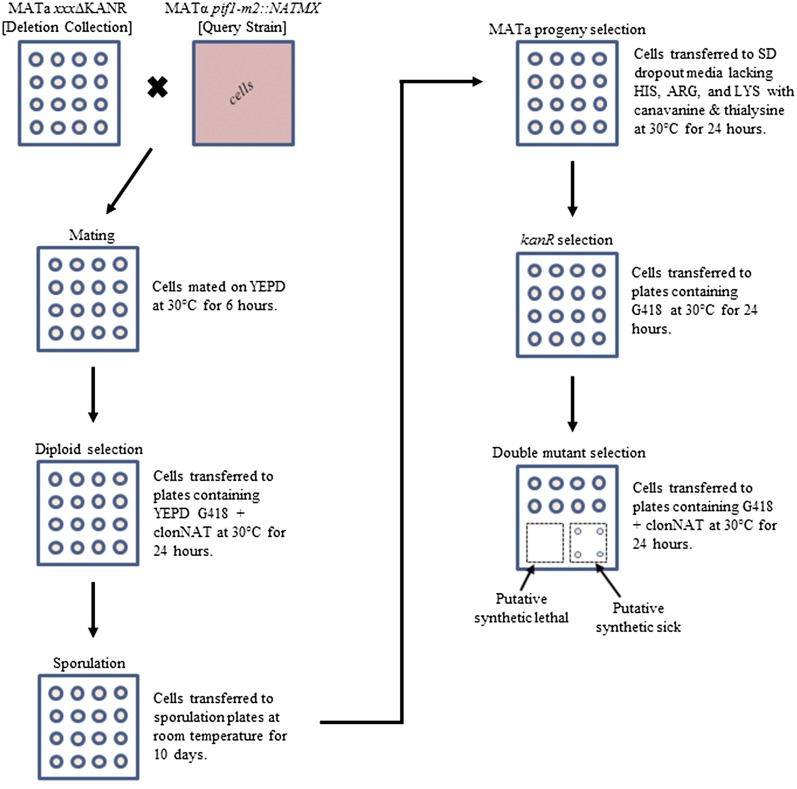
Strains and plasmids used in this study are listed in Table 1 and Table 2. The prototroph deletion collection, which contains 4783 strains with a single deletion of a nonessential gene, tagged with the kanMX antibiotic resistance marker, was used. The pif1-m2 query strain was created using the pvs31 plasmid, an integrating plasmid with a URA3 selectable marker (Schulz and Zakian 1994). The pvs31 plasmid was linearized with HindIII (NEB) and transformed into DBY11087 using lithium acetate transformation (Becker and Lundblad 2001). After introduction of the pif1-m2 mutation, the natMX resistance cassette was added proximal to the pif1-m2 gene (Goldstein and McCusker 1999). The pif1-m2 mutation was confirmed by polymerase chain reaction and sequencing and shown to segregate 2:2 with the natMX marker. Mating, sporulation and selection were performed as described (Tong and Boone 2006). Synthetic genetic analysis was performed as described (Tong and Boone 2006) as outlined in Figure 1. Briefly, the mutant and query strains were grown on yeast extract peptone dextrose media at 30°, and then mated, and diploids were selected using yeast extract peptone dextrose with G418 + clonNAT. The strains were transferred to sporulation media, then MATa haploids were selected using drop out media lacking HIS, ARG and LYS with canavanine and thialysine, followed by selection with drop out media with G418, and finally by selection of double mutant haploids on drop out media with G418 + clonNAT.
Table 1. Strains used in this study.
| Strain Name | Mutation, Strain Background, Previous Name/Previous Study if Applicable |
|---|---|
| Query-pif1m2::NATMX | pif1m2 mutation and NATMX cassette added to: DBY11087; S288C, MATα, his3∆ leu2∆ ura3∆ lyp1∆ met15∆cyh2∆ LYS2 can1::Pste2-S.P. his5 |
| Control-ho∆::NATMX | ho∆::natMX deletion added to: DBY11087; MATα, his3∆ leu2∆ ura3∆ lyp1∆ met15∆cyh2∆ LYS2 can1::Pste2-S.P. his5 |
| Prototrophic Deletion Mutation Array | DBY15001 W303 derived, MATa. Prototrophic deletion collection; created by Drs. Amy Caudy and David Hess. (Klosinska et al. 2011) |
Table 2. Plasmids used in this study.
| Plasmid Name | Description of Plasmid Use |
|---|---|
| pAG25 | Insertion of the NATMX cassette (Goldstein and McCusker 1999) |
| pvs31 | Insertion of the pif1m2 mutant via pop-in/pop-out (Schulz and Zakian 1994) |
Figure 1.
Schematic of steps for synthetic genetic analysis. All cells grown in quadruplicate as shown. Use of the hoΔNATMX control strain in parallel is not pictured. YEPD, yeast extract peptone dextrose.
Phenotypes
Each strain was mated in quadruplicate with the control (hoΔ::NATMX) and query (pif1-m2::NATMX) strains to generate double mutant diploids, which were then sporulated. Double mutant haploid clones (i.e., pif1-m2 geneXΔ) were derived from these spores. Strains that failed to form viable double mutant haploids when mated with the pif1-m2 strain but successfully formed viable double mutant haploids when mated with the control strain were considered putative synthetic lethal interactors. Strains that formed slow growing double mutant haploids when mated with the pif1-m2 strain (determined by visual inspection as being <50% of the size of either single mutant) were considered candidates for putative synthetic sick interactors. The use of the robotic pins, and the mixing steps utilized in the RoTOR robot (Singer, RoTOR-HDA), prevented the visualization of less severe synthetic growth differences. The synthetic sick mutants were not tested for mitochondrial proficiency.
Verification of mutants
Each putative synthetic relationship was re-examined by mating the appropriate strains by hand, selecting for diploids, which were sporulated and the resulting tetrads dissected. In some cases, random spore analysis was used as described (Lichten 2014).
Results
The genetic screen identified eleven genes that were synthetic lethal and three genes that were synthetic sick with pif1-m2. Surprisingly, this screen did not identify any of the over 100 genes that have been shown or proposed to encode a helicase as having a synthetic relationship with pif1-m2, including sgs1Δ and rad3Δ, which were shown previously to be synthetic sick with pif1Δ (Wagner et al. 2006; Moriel-Carretero and Aguilera 2010). The fact that pif1-m2 is not a null allele and retains residual nuclear function (Schulz and Zakian 1994) most likely explains why we did not observe synthetic relationships between pif1-m2 and other helicases and/or genes previously reported to have a synthetic phenotype with pif1Δ. Alternatively, synthetic phenotypes reported earlier may be due to the respiratory deficiencies, rather than the nuclear defects, of pif1Δ cells (Supporting Information, Table S1). It is also possible that the W303 based prototrophic deletion collection used here may contribute to the differences between this study and earlier analyses, as several earlier studies were completed in the BY4741 background (Pan et al. 2004, 2006), others used S288c (Zhang and Durocher 2010), and some used a combination of strain backgrounds (Osman et al. 2009; Moriel-Carretero and Aguilera 2010).
Synthetic lethal genes
Eleven genes were identified whose deletion from a pif1-m2 cell generated inviable cells. Here, we list those genes and provide some information on their functions and potential interactions with PIF1.
APM1 encodes a protein that is a subunit of the clathrin-associated protein complex. It is involved in the vesicular transport process (Nakayama et al. 1991; Stepp et al. 1995). apm1Δ cells have abnormal vacuolar transportation and abnormal Golgi protein sorting (Phelan et al. 2006).
ARG80 encodes a transcription factor that is involved in the regulation of arginine responsive genes (Dubois et al. 1987). arg80Δ cells have abnormal vacuolar morphology and decreased fitness (Michaillat and Mayer 2013). Given that Pif1 binds promoters of many genes (see Introduction), this synthetic phenotype may reflect a transcriptional problem in the double mutant.
CDH1 encodes a protein that activates the anaphase promoting complex/cyclosome (Visintin et al. 1997; Harper et al. 2002; Woodbury and Morgan 2007). Cdh1 is a cell-cycle regulated protein that directs the ubiquitination of cyclins and helps to orchestrate the mitotic exit from the cell cycle. cdh1Δ cells have increased telomere length and abnormal cell cycle progression (Visintin et al. 1997; Askree et al. 2004). Pif1 abundance is also cell-cycle regulated in a proteasome dependent manner, suggesting a potential relationship between Cdh1 and Pif1 (Mateyak and Zakian 2006). In addition, the essential telomerase subunit Est1 is cell cycle regulated (Taggart et al. 2002) in a proteasome and Cdh1-dependent manner (Osterhage et al. 2006; Ferguson et al. 2013). Moreover, many of the proteins that copurify with yeast telomerase, as determined by mass spectrometry analysis, affect ubiquitin and proteolysis (Lin et al. 2015). Thus, the lethality of pif1-m2 cdh1Δ cells may be due to impaired proteolysis that affects telomere length or other Pif1 functions.
GCR1 encodes a DNA binding protein that interacts with the transcriptional activator Gcr2 to promote transcriptional activation of genes involved in glycolysis (Clifton et al. 1978; Holland et al. 1987). As with ARG80, this interaction may be due to a transcriptional function of Pif1.
GTO3 encodes a glutathione transferase with a poorly defined function. It is putatively localized to the cytosol and gto3Δ cells have abnormal vacuolar morphology (Herrero 2005; Garcera et al. 2006).
PRK1 encodes a serine/threonine protein kinase that is involved in cytoskeletal organization and actin function (Byrne and Wolfe 2005; Zeng and Kinsella 2010). Endocytosis is reduced in prk1Δ cells (Henry et al. 2003).
RAD10 encodes a single-stranded DNA endonuclease with roles in both nucleotide excision repair and single-strand annealing-mediated recombination (Ivanov and Haber 1995; de Laat et al. 1999; Symington 2002). Pif1 inhibits telomerase-mediated double-strand break repair (Schulz and Zakian 1994). Rad10 promotes the creation of gross chromosomal rearrangements (GCR), which are increased in both pif1-m2 and pif1Δ cells (Myung et al. 2001; Hwang et al. 2005; Piazza et al. 2012; Paeschke et al. 2013). We speculate that pif1-m2 rad10Δ cells may be inviable due to combined defects in two different DNA repair pathways. Surprisingly, even though Rad1 and Rad10 act together in nucleotide excision repair and single strand annealing, this screen did not identify a synthetic relationship between pif1-m2 and RAD1. This result might indicate that RAD10 has a function that is distinct from RAD1, which is responsible for the synthetic relationship between RAD10 and pif1-m2. For example, a telomere-dedicated single strand annealing pathway that is RAD10- but not RAD1-dependent might act on the highly repetitive telomeric DNA.
SOP4 encodes an endoplasmic reticulum membrane protein that is involved in the export of Pma1 and Pma1-7, proteins that regulate cytoplasmic pH and help to maintain the plasma membrane potential from the endoplasmic reticulum (Luo and Chang 1997; Luo et al. 2002). sop4Δ cells have abnormal vacuolar morphology (Michaillat and Mayer 2013).
SKT5 encodes a protein that activates the chitin synthetase Chs3 that helps form spore walls (Iwamoto et al. 2005). skt5Δ cells have decreased vegetative growth rate and decreased viability (Kozubowski et al. 2003; Byrne and Wolfe 2005).
UMP1 encodes a protein that is a chaperone required for the maturation of the 20S proteasome (Ramos et al. 1998; Ishikawa et al. 2005). In ump1Δ cells, the proteasome is functionally impaired (Ramos et al. 1998) and data from our lab shows that pif1-m2 cells are more sensitive to proteasomal inhibition (J. L. Stundon and V. A. Zakian, unpublished results). ump1Δ cells with decreased nuclear Pif1 may be inviable due to strain on the proteasomal machinery. As with CDH1, the synthetic effects of pif1-m2 and ump1Δ may result from impaired Pif1 proteolysis.
YCK1 encodes a palmitoylated membrane-bound casein kinase that is involved in endocytic trafficking and glucose sensing (Robinson et al. 1992; Reddi and Culotta 2013). yck1Δ cells have abnormal vacuolar morphology (Michaillat and Mayer 2013).
Synthetic sick genes
Our screen also identified genes whose deletion in a pif1-m2 cell resulted in a slow growth phenotype where double mutants grew to <50% of the size of the single mutant strain. This analysis identified three such genes, DEF1, YIP4, and HOM3.
The identification of a synthetic relationship between PIF1 and DEF1 was particularly exciting, as both genes function in genome maintenance, telomere length, and maintenance of mtDNA. Def1 forms a complex with Rad26, a protein that functions in transcription-coupled repair (Woudstra et al. 2002; Somesh et al. 2005; Jordan et al. 2007). Def1 also plays a role in the maintenance of telomeres, as def1Δ telomeres are 200 bp shorter than the wild-type length of ∼300 bp (Chen et al. 2005). Like pif1Δ cells, def1Δ cells display increased mitophagy and abnormal vacuolar morphology (Michaillat and Mayer 2013; Bockler and Westermann 2014). We speculate that the reduced growth rate of the pif1-m2 def1Δ strain is due to their shared roles in DNA repair and telomere length.
YIP4 encodes a protein that interacts with Rab GTPases and is involved in vesicle-mediated transport (Samanta and Liang 2003; Inadome et al. 2007).
HOM3 encodes an aspartate kinase that is localized to the cytoplasm and catalyzes methionine and threonine biosynthesis (Mountain et al. 1991).
Discussion
The identified genes, whose deletion had synthetic effects with pif1-m2, support the known multifunctional nature of Pif1. In addition to its multiple described functions, our data suggest potential new roles for Pif1 in proteasome function, transcription coupled repair, endocytosis and vacuolar morphology, although some of these effects may be indirect if Pif1 has a transcriptional function. Future work will focus on elucidating the connections between PIF1 and the genes identified in this study, with a particular interest in examining the relationship between PIF1, CDH1, RAD10, and DEF1, as these genes all affect telomeres and/or DNA repair (Schulz and Zakian 1994; Woudstra et al. 2002; Askree et al. 2004; Chen et al. 2005). A second major focus will be on the role of Pif1 in proteasomal function as the synthetic relationships between pif1-m2 and CDH1 and UMP1 reported here, coupled with unpublished data from our lab, suggest a role for Pif1 in proteasomal function, which may be important for cells to tolerate stress. The synthetic effects of CDH1 and pif1-m2 and UMP1 might additionally help elucidate a role for PIF1 in autophagy. Future work will also examine the role of PIF1 on transcriptional regulation, which may help identify indirect interactions that are responsible for some of the synthetic genetic relationships reported here.
Supplementary Material
Acknowledgments
We thank David Botstein for providing strains. Research reported in this publication was supported by the National Institute on Aging of the National Institutes of Health (NIH) under Award Number F30AG042217 (to J. S.) and by NIH grant GM26938 (to V. A. Z.).
Footnotes
Supporting information is available online at www.g3journal.org/lookup/suppl/doi:10.1534/g3.115.021139/-/DC1
Communicating Editor: S. L. Jaspersen
Literature Cited
- Askree S. H., Yehuda T., Smolikov S., Gurevich R., Hawk J., et al. , 2004. A genome-wide screen for Saccharomyces cerevisiae deletion mutants that affect telomere length. Proc. Natl. Acad. Sci. USA 101: 8658–8663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker D. M., Lundblad V., 2001. Introduction of DNA into Yeast Cells in Current Protocols in Molecular Biology, John Wiley & Sons, Inc., Hoboken, New Jersey. [DOI] [PubMed] [Google Scholar]
- Bochman M. L., Sabouri N., Zakian V. A., 2010. Unwinding the functions of the Pif1 family helicases. DNA Repair (Amst.) 9: 237–249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bockler S., Westermann B., 2014. Mitochondrial ER contacts are crucial for mitophagy in yeast. Dev. Cell 28: 450–458. [DOI] [PubMed] [Google Scholar]
- Boule J. B., Zakian V. A., 2007. The yeast Pif1p DNA helicase preferentially unwinds RNA DNA substrates. Nucleic Acids Res. 35: 5809–5818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boule J. B., Vega L. R., Zakian V. A., 2005. The yeast Pif1p helicase removes telomerase from telomeric DNA. Nature 438: 57–61. [DOI] [PubMed] [Google Scholar]
- Byrne K. P., Wolfe K. H., 2005. The Yeast Gene Order Browser: combining curated homology and syntenic context reveals gene fate in polyploid species. Genome Res. 15: 1456–1461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y. B., Yang C. P., Li R. X., Zeng R., Zhou J. Q., 2005. Def1p is involved in telomere maintenance in budding yeast. J. Biol. Chem. 280: 24784–24791. [DOI] [PubMed] [Google Scholar]
- Clifton D., Weinstock S. B., Fraenkel D. G., 1978. Glycolysis mutants in Saccharomyces cerevisiae. Genetics 88: 1–11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Laat W. L., Jaspers N. G., Hoeijmakers J. H., 1999. Molecular mechanism of nucleotide excision repair. Genes Dev. 13: 768–785. [DOI] [PubMed] [Google Scholar]
- Dubois E., Bercy J., Messenguy F., 1987. Characterization of two genes, ARGRI and ARGRIII required for specific regulation of arginine metabolism in yeast. Mol. Gen. Genet. 207: 142–148. [DOI] [PubMed] [Google Scholar]
- Ferguson J. L., Chao W. C., Lee E., Friedman K. L., 2013. The anaphase promoting complex contributes to the degradation of the S. cerevisiae telomerase recruitment subunit Est1p. PLoS One 8: e55055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garcera A., Barreto L., Piedrafita L., Tamarit J., Herrero E., 2006. Saccharomyces cerevisiae cells have three Omega class glutathione S-transferases acting as 1-Cys thiol transferases. Biochem. J. 398: 187–196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldstein A. L., McCusker J. H., 1999. Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast 15: 1541–1553. [DOI] [PubMed] [Google Scholar]
- Harper J. W., Burton J. L., Solomon M. J., 2002. The anaphase-promoting complex: it’s not just for mitosis any more. Genes Dev. 16: 2179–2206. [DOI] [PubMed] [Google Scholar]
- Henry K. R., D’Hondt K., Chang J. S., Nix D. A., Cope M. J., et al. , 2003. The actin-regulating kinase Prk1p negatively regulates Scd5p, a suppressor of clathrin deficiency, in actin organization and endocytosis. Curr. Biol. 13: 1564–1569. [DOI] [PubMed] [Google Scholar]
- Herrero E., 2005. Evolutionary relationships between Saccharomyces cerevisiae and other fungal species as determined from genome comparisons. Rev. Iberoam. Micol. 22: 217–222. [DOI] [PubMed] [Google Scholar]
- Holland M. J., Yokoi T., Holland J. P., Myambo K., Innis M. A., 1987. The GCR1 gene encodes a positive transcriptional regulator of the enolase and glyceraldehyde-3-phosphate dehydrogenase gene families in Saccharomyces cerevisiae. Mol. Cell. Biol. 7: 813–820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang J. Y., Smith S., Myung K., 2005. The Rad1-Rad10 complex promotes the production of gross chromosomal rearrangements from spontaneous DNA damage in Saccharomyces cerevisiae. Genetics 169: 1927–1937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inadome H., Noda Y., Kamimura Y., Adachi H., Yoda K., 2007. Tvp38, Tvp23, Tvp18 and Tvp15: novel membrane proteins in the Tlg2-containing Golgi/endosome compartments of Saccharomyces cerevisiae. Exp. Cell Res. 313: 688–697. [DOI] [PubMed] [Google Scholar]
- Ishikawa T., Unno K., Nonaka G., Nakajima H., Kitamoto K., 2005. Isolation of Saccharomyces cerevisiae RNase T1 hypersensitive (rns) mutants and genetic analysis of the RNS1/DSL1 gene. J. Gen. Appl. Microbiol. 51: 73–82. [DOI] [PubMed] [Google Scholar]
- Ivanov E. L., Haber J. E., 1995. RAD1 and RAD10, but not other excision repair genes, are required for double-strand break-induced recombination in Saccharomyces cerevisiae. Mol. Cell. Biol. 15: 2245–2251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ivessa A. S., Zhou J. Q., Zakian V. A., 2000. The Saccharomyces Pif1p DNA helicase and the highly related Rrm3p have opposite effects on replication fork progression in ribosomal DNA. Cell 100: 479–489. [DOI] [PubMed] [Google Scholar]
- Ivessa A. S., Zhou J. Q., Schulz V. P., Monson E. K., Zakian V. A., 2002. Saccharomyces Rrm3p, a 5′ to 3′ DNA helicase that promotes replication fork progression through telomeric and subtelomeric DNA. Genes Dev. 16: 1383–1396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwamoto M. A., Fairclough S. R., Rudge S. A., Engebrecht J., 2005. Saccharomyces cerevisiae Sps1p regulates trafficking of enzymes required for spore wall synthesis. Eukaryot. Cell 4: 536–544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan P. W., Klein F., Leach D. R., 2007. Novel roles for selected genes in meiotic DNA processing. PLoS Genet. 3: e222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klosinska M. M., Crutchfield C. A., Bradley P. H., Rabinowitz J. D., Broach J. R., 2011. Yeast cells can access distinct quiescent states. Genes Dev. 25: 336–349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozubowski L., Panek H., Rosenthal A., Bloecher A., DeMarini D. J., et al. , 2003. A Bni4-Glc7 phosphatase complex that recruits chitin synthase to the site of bud emergence. Mol. Biol. Cell 14: 26–39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lichten M., 2014. Tetrad, random spore, and molecular analysis of meiotic segregation and recombination. Methods Mol. Biol. 1205: 13–28. [DOI] [PubMed] [Google Scholar]
- Lin K. W., McDonald K. R., Guise A. J., Chan A., Cristea I. M., et al. , 2015. Proteomics of yeast telomerase identified Cdc48-Npl4-Ufd1 and Ufd4 as regulators of Est1 and telomere length. Nat. Commun. 6: 8290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo W., Chang A., 1997. Novel genes involved in endosomal traffic in yeast revealed by suppression of a targeting-defective plasma membrane ATPase mutant. J. Cell Biol. 138: 731–746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luo W. J., Gong X. H., Chang A., 2002. An ER membrane protein, Sop4, facilitates ER export of the yeast plasma membrane [H+]ATPase, Pma1. Traffic 3: 730–739. [DOI] [PubMed] [Google Scholar]
- Mateyak M. K., Zakian V. A., 2006. Human PIF helicase is cell cycle regulated and associates with telomerase. Cell Cycle 5: 2796–2804. [DOI] [PubMed] [Google Scholar]
- Michaillat L., Mayer A., 2013. Identification of genes affecting vacuole membrane fragmentation in Saccharomyces cerevisiae. PLoS One 8: e54160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moriel-Carretero M., Aguilera A., 2010. A postincision-deficient TFIIH causes replication fork breakage and uncovers alternative Rad51- or Pol32-mediated restart mechanisms. Mol. Cell 37: 690–701. [DOI] [PubMed] [Google Scholar]
- Mountain H. A., Bystrom A. S., Larsen J. T., Korch C., 1991. Four major transcriptional responses in the methionine/threonine biosynthetic pathway of Saccharomyces cerevisiae. Yeast 7: 781–803. [DOI] [PubMed] [Google Scholar]
- Myung K., Chen C., Kolodner R. D., 2001. Multiple pathways cooperate in the suppression of genome instability in Saccharomyces cerevisiae. Nature 411: 1073–1076. [DOI] [PubMed] [Google Scholar]
- Nakayama Y., Goebl M., O’Brine Greco B., Lemmon S., Pingchang Chow E., et al. , 1991. The medium chains of the mammalian clathrin-associated proteins have a homolog in yeast. Eur. J. Biochem. 202: 569–574. [DOI] [PubMed] [Google Scholar]
- O’Rourke T. W., Doudican N. A., Zhang H., Eaton J. S., Doetsch P. W., et al. , 2005. Differential involvement of the related DNA helicases Pif1p and Rrm3p in mtDNA point mutagenesis and stability. Gene 354: 86–92. [DOI] [PubMed] [Google Scholar]
- Osman C., Haag M., Potting C., Rodenfels J., Dip P. V., et al. , 2009. The genetic interactome of prohibitins: coordinated control of cardiolipin and phosphatidylethanolamine by conserved regulators in mitochondria. J. Cell Biol. 184: 583–596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osterhage J. L., Talley J. M., Friedman K. L., 2006. Proteasome-dependent degradation of Est1p regulates the cell cycle-restricted assembly of telomerase in Saccharomyces cerevisiae. Nat. Struct. Mol. Biol. 13: 720–728. [DOI] [PubMed] [Google Scholar]
- Paeschke K., Capra J. A., Zakian V. A., 2011. DNA replication through G-quadruplex motifs is promoted by the Saccharomyces cerevisiae Pif1 DNA helicase. Cell 145: 678–691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paeschke K., Bochman M. L., Garcia P. D., Cejka P., Friedman K. L., et al. , 2013. Pif1 family helicases suppress genome instability at G-quadruplex motifs. Nature 497: 458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan X., Yuan D. S., Xiang D., Wang X., Sookhai-Mahadeo S., et al. , 2004. A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16: 487–496. [DOI] [PubMed] [Google Scholar]
- Pan X., Ye P., Yuan D. S., Wang X., Bader J. S., et al. , 2006. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124: 1069–1081. [DOI] [PubMed] [Google Scholar]
- Phelan J. P., Millson S. H., Parker P. J., Piper P. W., Cooke F. T., 2006. Fab1p and AP-1 are required for trafficking of endogenously ubiquitylated cargoes to the vacuole lumen in S. cerevisiae. J. Cell Sci. 119: 4225–4234. [DOI] [PubMed] [Google Scholar]
- Phillips J. A., Chan A., Paeschke K., Zakian V. A., 2015. The Pif1 Helicase, a Negative Regulator of Telomerase, Acts Preferentially at Long Telomeres. PLoS Genet. 11: e1005186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piazza A., Serero A., Boule J. B., Legoix-Ne P., Lopes J., et al. , 2012. Stimulation of Gross Chromosomal Rearrangements by the Human CEB1 and CEB25 Minisatellites in Saccharomyces cerevisiae Depends on G-Quadruplexes or Cdc13. PLoS Genet. 8: 17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pike J. E., Burgers P. M., Campbell J. L., Bambara R. A., 2009. Pif1 helicase lengthens some Okazaki fragment flaps necessitating Dna2 nuclease/helicase action in the two-nuclease processing pathway. J. Biol. Chem. 284: 25170–25180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramos P. C., Hockendorff J., Johnson E. S., Varshavsky A., Dohmen R. J., 1998. Ump1p is required for proper maturation of the 20S proteasome and becomes its substrate upon completion of the assembly. Cell 92: 489–499. [DOI] [PubMed] [Google Scholar]
- Reddi A. R., Culotta V. C., 2013. SOD1 integrates signals from oxygen and glucose to repress respiration. Cell 152: 224–235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ribeyre C., Lopes J., Boule J. B., Piazza A., Guedin A., et al. , 2009. The yeast Pif1 helicase prevents genomic instability caused by G-quadruplex-forming CEB1 sequences in vivo. PLoS Genet. 5: e1000475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson L. C., Hubbard E. J., Graves P. R., DePaoli-Roach A. A., Roach P. J., et al. , 1992. Yeast casein kinase I homologues: an essential gene pair. Proc. Natl. Acad. Sci. USA 89: 28–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saini N., Ramakrishnan S., Elango R., Ayyar S., Zhang Y., et al. , 2013. Migrating bubble during break-induced replication drives conservative DNA synthesis. Nature 502: 389–392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Samanta M. P., Liang S., 2003. Predicting protein functions from redundancies in large-scale protein interaction networks. Proc. Natl. Acad. Sci. USA 100: 12579–12583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schulz V. P., Zakian V. A., 1994. The saccharomyces PIF1 DNA helicase inhibits telomere elongation and de novo telomere formation. Cell 76: 145–155. [DOI] [PubMed] [Google Scholar]
- Somesh B. P., Reid J., Liu W. F., Sogaard T. M., Erdjument-Bromage H., et al. , 2005. Multiple mechanisms confining RNA polymerase II ubiquitylation to polymerases undergoing transcriptional arrest. Cell 121: 913–923. [DOI] [PubMed] [Google Scholar]
- Stepp J. D., Pellicena-Palle A., Hamilton S., Kirchhausen T., Lemmon S. K., 1995. A late Golgi sorting function for Saccharomyces cerevisiae Apm1p, but not for Apm2p, a second yeast clathrin AP medium chain-related protein. Mol. Biol. Cell 6: 41–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Symington L. S., 2002. Role of RAD52 epistasis group genes in homologous recombination and double-strand break repair. Microbiol. Mol. Biol. Rev. 66: 630–670 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taggart A. K., Teng S. C., Zakian V. A., 2002. Est1p as a cell cycle-regulated activator of telomere-bound telomerase. Science 297: 1023–1026. [DOI] [PubMed] [Google Scholar]
- Tong A. H., Boone C., 2006. Synthetic genetic array analysis in Saccharomyces cerevisiae. Methods Mol. Biol. 313: 171–192. [DOI] [PubMed] [Google Scholar]
- Visintin R., Prinz S., Amon A., 1997. CDC20 and CDH1: a family of substrate-specific activators of APC-dependent proteolysis. Science 278: 460–463. [DOI] [PubMed] [Google Scholar]
- Wagner M., Price G., Rothstein R., 2006. The absence of Top3 reveals an interaction between the Sgs1 and Pif1 DNA helicases in Saccharomyces cerevisiae. Genetics 174: 555–573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson M. A., Kwon Y., Xu Y., Chung W. H., Chi P., et al. , 2013. Pif1 helicase and Poldelta promote recombination-coupled DNA synthesis via bubble migration. Nature 502: 393–396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodbury E. L., Morgan D. O., 2007. Cdk and APC activities limit the spindle-stabilizing function of Fin1 to anaphase. Nat. Cell Biol. 9: 106–112. [DOI] [PubMed] [Google Scholar]
- Woudstra E. C., Gilbert C., Fellows J., Jansen L., Brouwer J., et al. , 2002. A Rad26-Def1 complex coordinates repair and RNA pol II proteolysis in response to DNA damage. Nature 415: 929–933. [DOI] [PubMed] [Google Scholar]
- Zeng X., Kinsella T. J., 2010. BNIP3 is essential for mediating 6-thioguanine- and 5-fluorouracil-induced autophagy following DNA mismatch repair processing. Cell Res. 20: 665–675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang W., Durocher D., 2010. De novo telomere formation is suppressed by the Mec1-dependent inhibition of Cdc13 accumulation at DNA breaks. Genes Dev. 24: 502–515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J., Monson E. K., Teng S. C., Schulz V. P., Zakian V. A., 2000. Pif1p helicase, a catalytic inhibitor of telomerase in yeast. Science 289: 771–774. [DOI] [PubMed] [Google Scholar]
- Zhou R., Zhang J., Bochman M. L., Zakian V. A., Ha T., 2014. Periodic DNA patrolling underlies diverse functions of Pif1 on R-loops and G-rich DNA. eLife 3: e02190. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.