Abstract
It should be emphasized that “129” is not simply a number but is also the designation of a mouse strain that has made a great contribution to modern biological science and technology. Embryonic stem cells derived from 129 mice were essential components of gene-targeting strategies in early research. More recently, 129 mice have provided superior donor genomes for cloning by nuclear transfer. Some factor or factors conferring genomic plasticity must exist in the 129 genome, but these remain unidentified.
Keywords: Embryonic stem cell, Genomic reprogramming, Mouse, Nuclear transfer
There are numerous strains of laboratory mice (Mus musculus) available for research in the fields of biology and basic medicine. Mice are small enough for easy handling, and can be propagated in a relatively short generation time (about 3–4 months). Historically, the laboratory mice were derived from “fancy” mice bred at the beginning of the last century [1, 2]. Since then, great efforts have been made to select them for unique phenotypic characteristics including coat colors and spontaneous diseases useful for biomedical research. Genetic background effects have been matters of interest, and inbreeding has been promoted to produce completely homozygous inbred strains, which are now broadly used worldwide as standard strains. The genetic diversity of laboratory mice is limited because they originate from a relatively small pool of fancy mice with genetic backgrounds composed predominantly of the Western European subspecies M. musculus domesticus and the Japanese subspecies M. musculus molossinus, specifically JF1/Ms [3]. Despite such a restricted genetic background, researchers have noted that there is a small group of mice showing characteristics that have diverged greatly from those of other strains. This is the “129” strain.
The 129 mouse strain: a superior source of embryonic stem cells
The 129 strain consists of many substrains and is thought to have been genetically contaminated to some extent in the course of research and exchanges between laboratories [4] (Fig. 1). These substrains can be divided into three major groups—129S, 129P and 129T—according to their origin [4]. As far as has been examined to date, all these substrains show unique and extensive genomic plasticity under specific circumstances. In early studies, 129 mice were characterized by a low but consistent incidence (1–8%) of spontaneous testicular germ cell tumors (TGCTs), which contributed to the derivation of embryonic carcinoma (EC) cells [5]. The methodologies established for EC cells during the 1960s and 1970s, including L.C. Stevens’ pioneering work [6], provided the important foundation for establishing embryonic stem (ES) cell lines. ES cells are pluripotent stem cells derived from the inner cell mass cells of blastocysts and are considered to be the non-tumor counterparts of EC cells. At that time, the 129 mouse was the only inbred strain that could give rise to stable ES cell lines [7, 8]. Interestingly, not only the homozygous but also the heterozygous set of the 129 genome can contribute to easier generation of ES cell lines, as evidenced by the generation of hybrid cell lines between 129 and “nonpermissive” strains such as the NOD, DDK, and Chinese Kunming mouse strains [9,10,11]. Currently, inbred strains other than 129—such as C57BL, NOD and MSM—can also be sources of ES cells thanks to the recent development of serum-free media containing inhibitors of MEK and GSK3 (2i) and LIF, which potentially can maintain the pluripotency of ES cells in the so-called “naïve” state [12]. Nevertheless, it is apparent that the contribution of the 129 strain to the advancements of biomedical science has been enormous, considering the thousands of gene-targeted mice generated with a 129 genetic background.
Fig. 1.
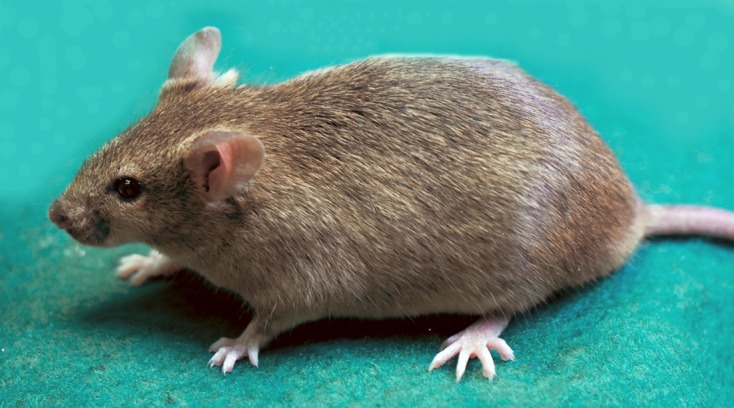

The 129 strain consists of many substrains due to its complex history [4]. The image shows a commonly used 129/Sv mouse, which has the full designation, 129S1/Sv-p<+> Tyr<+>, indicating its substrain group (“129S” indicates a Steel substrain group) and coat color (wild type). This strain is available from the RIKEN BioResource Center (RBRC00002).
The 129 genome: highly reprogrammable by nuclear transfer
The uniqueness of the 129 strain was further recognized in somatic cell nuclear transfer (SCNT) experiments in mice. Since the first report on the birth of mice cloned from cumulus cells in 1998 [13], F1 hybrid mice such as B6D2F1 (cross of a C57BL/6 female mouse and a DBA/2 male mouse) have been most frequently used for donors [14]. The so-called “hybrid vigor,” which is known to increase the survivability of mouse embryos in vitro, is particularly advantageous in the development of cloned embryos [15]. By contrast, cloning mice using SCNT from inbred strains is extremely difficult. The only exception of this is the 129 strain, as evidenced by the births of cloned offspring from cumulus cells [16, 17], immature Sertoli cells [17], and tail-tip cells [18] with the 129 genetic background. A global gene expression analysis we performed using blastocysts revealed the superior reprogrammability of the 129 genome as a donor for SCNT (Fig. 2). It is interesting to note that, as in the case of ES cells, the presence of the haploid component of the 129 genome in the donor genome has a beneficial effect. Specifically, we have been able to produce cloned mice successfully using new somatic cell types (neural stem cells, natural killer T cells, hematopoietic stem cells and primordial germ cells) with the F1 hybrid 129 genotype [19,20,21,22]. Hematopoietic cells, for example, are difficult to clone by nuclear transfer, but those from the 129 strain can be reprogrammed easily (Table 1). Similarly, hybrids between wild-derived mice (JF1) and 129 mice were cloned successfully for genomic imprinting studies, which analyzed the polymorphisms between these parental strains [17, 23,24,25]. Furthermore, one of the most outstanding experiments in mammalian cloning— the serial recloning of mice over 25 generations—was achieved with a donor genome partially comprised of the 129 genome [26]. In addition to its effect on promoting development, the 129 genome exerts corrective effects on abnormal phenotypes associated with SCNT. It is well known that cloned placentas show hyperplastic enlargement with abnormal histology [27, 28] (Fig. 3). This abnormality was not corrected even after significant improvements in cloning efficiency by correction of Xist gene expression [29] or the treatment of cloned embryos with trichostatin A [30]. However, the cloned placentas of the 129 strain are nearly normal in appearance and histology [16] (Fig. 3).
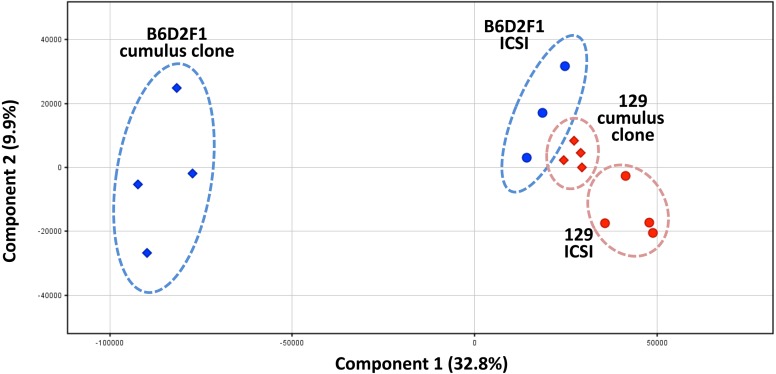
Fig. 2.
Principal component analysis for global gene expression profiles of blastocysts generated by ICSI (intracytoplasmic sperm injection) or SCNT (cumulus clone) with the B6D2F1 or 129 genotypes. Components 1 and 2 mean principal components with the first and second highest levels of variance, respectively. The percentages in parentheses indicate the contribution ratios of the components. Visualization of gene expression similarities clearly distinguished the B6D2F1 and 129 groups. SCNT-derived blastocysts from the B6D2F1 strain showed SCNT-specific expression profiles (enclosed with blue dotted lines), but those from 129 did not (enclosed with red dotted lines), indicating that the 129 genome was reprogrammed more correctly than the B6D2F1 genome by nuclear transfer.
Table 1. Development in vitro of embryos cloned from hematopoietic stem cells from B6D2F1 or 129 strains of mice.
| Donor strain | No. Embryos cultured | No. (%) 2-cells embryos | No. (%) 4-cells embryos | No. (%) blastocysts |
| B6D2F1 | 44 | 39 (89) | 8 (18)a | 4 (9)c |
| 129/Sv-ter | 58 | 48 (83) | 40 (69)b | 30 (52)d |
Different superscript letters within columns indicate significance at P < 0.0001 (Fisher’s exact test).
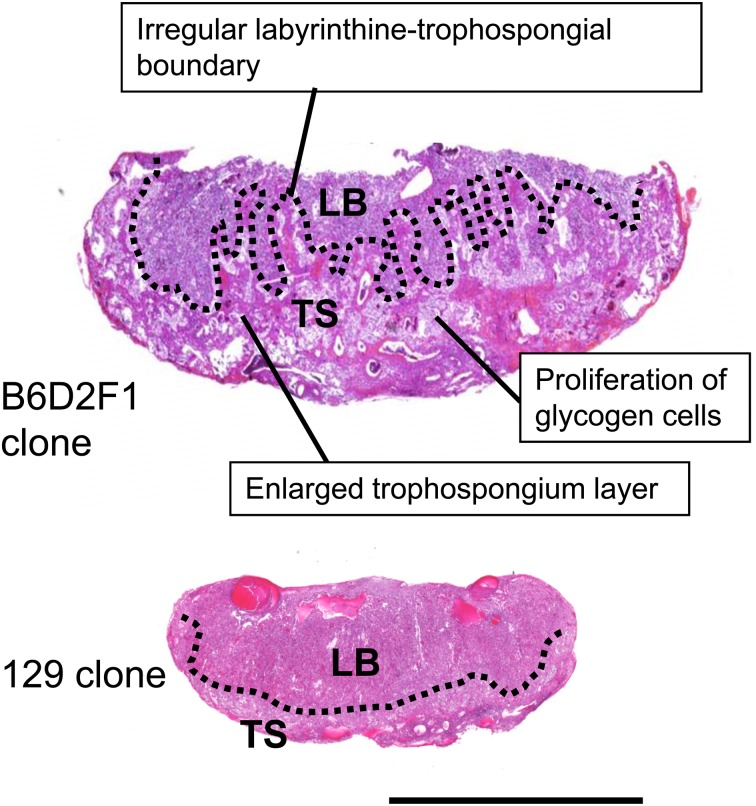
Fig. 3.
The histology of a typical term placenta produced by SCNT. The image shows two- to three-fold enlargement (hyperplasia) with characteristic histology (upper). However, the presence of the 129 genome in the donor nucleus ameliorated the SCNT-associated placental abnormalities (lower). LB, labyrinth layer. TS, trophospongium layer. Bar = 5 mm.
What is the nature of 129-associated genomic plasticity?
These 129-specific features identified with ES cells and cloned embryos cumulatively suggest the presence of some factor (or factors) that promotes genomic flexibility or plasticity of the 129 genome. Although the nature of this factor remains elusive, we suppose that it might cause lability of epigenetic regulation in a genome-wide manner. There are some findings that support this assumption. Transgenic mice with a 129 genetic background showed a moderate loss in DNA methylation of transgenes, although the methylation levels varied with the site of integration into the recipient genome [31]. The naïve status of both ES cells and embryonic germ (EG) cells was maintained by global DNA hypomethylation [32]. It is known that, like ES cells, EG cells can also be easily established from 129 mice [33]. Taken together, the plasticity factor in the 129 strain might endow the genome with a propensity towards DNA hypomethylation in a genome-wide manner.
In contrast to the situations of ES cells and EG cells, 129 is known to be nonpermissive for the derivation of germline stem (GS) cells, which were originally developed by Shinohara’s group [34]. GS cells are derived from neonatal spermatogonial stem cells and have an unlimited ability to proliferate in vitro while also maintaining full differentiation capacity in vivo [34,35,36]. It is possible that the inability of 129 spermatogonia to contribute to the generation of GS cells might also be explained by DNA methylation. Genome-wide analysis of DNA methylation levels revealed that GS cells and neonatal spermatogonia are the most hypermethylated cells among different types of germ and stem cells [37]. Therefore, the derivation of GS cells from spermatogonia requires maintenance of the hypermethylated status in vitro, which might be interfered with by the propensity of the 129 genome. Notably, transformation of GS cells into ES cell-like cells (multipotent GS cells; mGS cells [38]) was made possible by the induction of DNA demethylation [39, 40], indicating that GS cells are more methylated than pluripotent mGS cells. Thus, although still speculative, the epigenetic characters of GS cells and mGS cells might also be consistent with our assumption that the putative plasticity factor within the 129 genome might help the entire genome in acquiring a hypomethylated status.
How can the plasticity factor be identified?
So far, the search for genetic traits using the 129 strain has focused mainly on TGCTs. Among the major substrain groups, the 129T group (the ‘T’ comes from teratoma) is known to be susceptible to TGCTs. Genetic analysis of segregating populations generated from 129 mice and from mice that do not form TGCTs suggested that at least 3–5, or at maximum 15, genes control the susceptibility to TGCT [5]. It is possible that one or more genes among them may be associated with the genomic plasticity of 129 mice. Quantitative trait locus (QTL) analysis identified several genetic regions or specific genes, such as Ter, Steel and Trp53 [5]. However, these recessive genes are not strong candidates for the 129 plasticity factor because it exerts its effect as a dominant, not recessive, trait. Experiments using chromosome substitution (consomic) strains between 129 (susceptible) and MOLF (non-susceptible) strains identified that genes conferring strong TGCT susceptibility exist on chromosome (Chr) 18 and Chr 19 [41, 42]. Interestingly, a Chr 18 consomic strain, 129-Chr 18MOLF, showed not only a lower TGCT frequency but also a worse capacity for ES cell derivation than the 129 strain [42, 43]. This suggests that TGCT susceptibility and the capacity for ES cell derivation in 129 strains can be attributed to a common gene(s) on Chr 18.
It has been reported that ES cells from permissive strains including 129 activated the JAK-Stat3 pathway rather than the MAP kinase pathway downstream of LIF, while those from nonpermissive strains showed the opposite pattern [44]. However, in this study, C57BL/6 was also classified as a permissive strain, so any 129-specific characters associated with ES cells remained unclear. Further experiments that discriminate 129 from the C57BL/6 strains might give clues to understanding the plasticity factor in the 129 genome.
We sought to identify the plasticity factor present in the 129 genome by a forward genetics strategy using nine recombinant inbred (RI) strains between 129 and C57BL/6. These carry randomly distributed homozygous loci derived from either parental strain, so a set of RI strains can be used for identifying genomic regions or genes responsible for phenotypes of interest.
First, we performed SCNT experiments using cumulus cells from eleven strains (nine RI strains and two parental strains), and the resultant cloned blastocysts were subjected to global gene expression analysis by microarray. As mentioned above, we expected that the RI strains that carry the putative 129-derived plasticity factor would show a high level of genomic reprogrammability and more normal gene expression profiles. However, the gene expression profiles obtained were too diverse among the strains, so we failed to identify the RI strains that are close to the 129 strain in their gene expression profile. Next, we examined the birth rates of clones and the morphology of placentas at term in all RI strains. In total, we reconstructed 7454 embryos, and 6671 of them were transferred into recipient pseudopregnant females. Six of the nine RI strains gave rise to cloned offspring, and the remaining three strains did not. Based on the birth rates, the placental weights and the genomic data of the RI strains, we ultimately identified four candidate genomic regions that might be responsible for the plasticity of the 129 genome. These regions include several genes related to epigenetic regulation (unpublished). We are now searching for polymorphisms between the 129 and C57BL/6 strains in these candidate genes, while preparing another series of experiments on the RI strains based on derivation of ES cell lines.
Future prospects
Genomic reprogramming comprises the central part of induced pluripotent stem cell (iPSC) technology and SCNT cloning [45]. Therefore, if we could identify the genomic plasticity factor (or factors) in 129 mice, it would contribute greatly to the safe and efficient development of new technologies in regenerative medicine, farm animal production, and the pharmaceutical industries. At present, we do not know the real identity of this factor. However, it should be encoded in the 129 genome and could be transferred into other mammalian species, provided the mechanisms involved are not specific to the mouse. It is possible that novel agents that enhance genomic reprogramming could be synthesized if we could identify new mechanisms for epigenetic regulation from the 129 factor. They would indeed comprise a genomic treasure to be mined in the 129 genome, and humans would benefit from this in many aspects of science and medicine.
References
- 1.Wade CM, Kulbokas EJ, 3rd, Kirby AW, Zody MC, Mullikin JC, Lander ES, Lindblad-Toh K, Daly MJ. The mosaic structure of variation in the laboratory mouse genome. Nature 2002; 420: 574–578. [DOI] [PubMed] [Google Scholar]
- 2.Abe K, Noguchi H, Tagawa K, Yuzuriha M, Toyoda A, Kojima T, Ezawa K, Saitou N, Hattori M, Sakaki Y, Moriwaki K, Shiroishi T. Contribution of Asian mouse subspecies Mus musculus molossinus to genomic constitution of strain C57BL/6J, as defined by BAC-end sequence-SNP analysis. Genome Res 2004; 14: 2439–2447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Takada T, Ebata T, Noguchi H, Keane TM, Adams DJ, Narita T, Shin-I T, Fujisawa H, Toyoda A, Abe K, Obata Y, Sakaki Y, Moriwaki K, Fujiyama A, Kohara Y, Shiroishi T. The ancestor of extant Japanese fancy mice contributed to the mosaic genomes of classical inbred strains. Genome Res 2013; 23: 1329–1338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Simpson EM, Linder CC, Sargent EE, Davisson MT, Mobraaten LE, Sharp JJ. Genetic variation among 129 substrains and its importance for targeted mutagenesis in mice. Nat Genet 1997; 16: 19–27. [DOI] [PubMed] [Google Scholar]
- 5.Jiang LI, Nadeau JH. 129/Sv mice—a model system for studying germ cell biology and testicular cancer. Mamm Genome 2001; 12: 89–94. [DOI] [PubMed] [Google Scholar]
- 6.Stevens LC. A new inbred subline of mice (129-terSv) with a high incidence of spontaneous congenital testicular teratomas. J Natl Cancer Inst 1973; 50: 235–242. [DOI] [PubMed] [Google Scholar]
- 7.Kawase E, Suemori H, Takahashi N, Okazaki K, Hashimoto K, Nakatsuji N. Strain difference in establishment of mouse embryonic stem (ES) cell lines. Int J Dev Biol 1994; 38: 385–390. [PubMed] [Google Scholar]
- 8.Suzuki O, Matsuda J, Takano K, Yamamoto Y, Asano T, Naiki M, Kusanagi M. Effect of genetic background on establishment of mouse embryonic stem cells. Exp Anim 1999; 48: 213–216. [DOI] [PubMed] [Google Scholar]
- 9.Brook FA, Evans EP, Lord CJ, Lyons PA, Rainbow DB, Howlett SK, Wicker LS, Todd JA, Gardner RL. The derivation of highly germline-competent embryonic stem cells containing NOD-derived genome. Diabetes 2003; 52: 205–208. [DOI] [PubMed] [Google Scholar]
- 10.Kress C, Vandormael-Pournin S, Baldacci P, Cohen-Tannoudji M, Babinet C. Nonpermissiveness for mouse embryonic stem (ES) cell derivation circumvented by a single backcross to 129/Sv strain: establishment of ES cell lines bearing the Omd conditional lethal mutation. Mamm Genome 1998; 9: 998–1001. [DOI] [PubMed] [Google Scholar]
- 11.Yu S, Yan X, Liu H, Cai X, Cao S, Shen L, Zuo Z, Deng J, Ma X, Wang Y, Ren Z. Improved establishment of embryonic stem (ES) cell lines from the Chinese Kunming mice by hybridization with 129 mice. Int J Mol Sci 2014; 15: 3389–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B, Woodgett J, Cohen P, Smith A. The ground state of embryonic stem cell self-renewal. Nature 2008; 453: 519–523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wakayama T, Perry AC, Zuccotti M, Johnson KR, Yanagimachi R. Full-term development of mice from enucleated oocytes injected with cumulus cell nuclei. Nature 1998; 394: 369–374. [DOI] [PubMed] [Google Scholar]
- 14.Ogura A, Inoue K, Wakayama T. Recent advancements in cloning by somatic cell nuclear transfer. Philos Trans R Soc Lond B Biol Sci 2013; 368: 20110329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Eggan K, Akutsu H, Loring J, Jackson-Grusby L, Klemm M, Rideout WM, 3rd, Yanagimachi R, Jaenisch R. Hybrid vigor, fetal overgrowth, and viability of mice derived by nuclear cloning and tetraploid embryo complementation. Proc Natl Acad Sci USA 2001; 98: 6209–6214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wakayama T, Yanagimachi R. Mouse cloning with nucleus donor cells of different age and type. Mol Reprod Dev 2001; 58: 376–383. [DOI] [PubMed] [Google Scholar]
- 17.Inoue K, Ogonuki N, Mochida K, Yamamoto Y, Takano K, Kohda T, Ishino F, Ogura A. Effects of donor cell type and genotype on the efficiency of mouse somatic cell cloning. Biol Reprod 2003; 69: 1394–1400. [DOI] [PubMed] [Google Scholar]
- 18.Ogura A, Inoue K, Takano K, Wakayama T, Yanagimachi R. Birth of mice after nuclear transfer by electrofusion using tail tip cells. Mol Reprod Dev 2000; 57: 55–59. [DOI] [PubMed] [Google Scholar]
- 19.Inoue K, Noda S, Ogonuki N, Miki H, Inoue S, Katayama K, Mekada K, Miyoshi H, Ogura A. Differential developmental ability of embryos cloned from tissue-specific stem cells. Stem Cells 2007; 25: 1279–1285. [DOI] [PubMed] [Google Scholar]
- 20.Inoue K, Wakao H, Ogonuki N, Miki H, Seino K, Nambu-Wakao R, Noda S, Miyoshi H, Koseki H, Taniguchi M, Ogura A. Generation of cloned mice by direct nuclear transfer from natural killer T cells. Curr Biol 2005; 15: 1114–1118. [DOI] [PubMed] [Google Scholar]
- 21.Inoue K, Ogonuki N, Miki H, Hirose M, Noda S, Kim J-M, Aoki F, Miyoshi H, Ogura A. Inefficient reprogramming of the hematopoietic stem cell genome following nuclear transfer. J Cell Sci 2006; 119: 1985–1991. [DOI] [PubMed] [Google Scholar]
- 22.Miki H, Inoue K, Kohda T, Honda A, Ogonuki N, Yuzuriha M, Mise N, Matsui Y, Baba T, Abe K, Ishino F, Ogura A. Birth of mice produced by germ cell nuclear transfer. Genesis 2005; 41: 81–86. [DOI] [PubMed] [Google Scholar]
- 23.Lee J, Inoue K, Ono R, Ogonuki N, Kohda T, Kaneko-Ishino T, Ogura A, Ishino F. Erasing genomic imprinting memory in mouse clone embryos produced from day 11.5 primordial germ cells. Development 2002; 129: 1807–1817. [DOI] [PubMed] [Google Scholar]
- 24.Okae H, Matoba S, Nagashima T, Mizutani E, Inoue K, Ogonuki N, Chiba H, Funayama R, Tanaka S, Yaegashi N, Nakayama K, Sasaki H, Ogura A, Arima T. RNA sequencing-based identification of aberrant imprinting in cloned mice. Hum Mol Genet 2014; 23: 992–1001. [DOI] [PubMed] [Google Scholar]
- 25.Kamimura S, Hatanaka Y, Hirasawa R, Matsumoto K, Oikawa M, Lee J, Matoba S, Mizutani E, Ogonuki N, Inoue K, Kohda T, Ishino F, Ogura A. Establishment of paternal genomic imprinting in mouse prospermatogonia analyzed by nuclear transfer. Biol Reprod 2014; 91: 120. [DOI] [PubMed] [Google Scholar]
- 26.Wakayama S, Kohda T, Obokata H, Tokoro M, Li C, Terashita Y, Mizutani E, Nguyen VT, Kishigami S, Ishino F, Wakayama T. Successful serial recloning in the mouse over multiple generations. Cell Stem Cell 2013; 12: 293–297. [DOI] [PubMed] [Google Scholar]
- 27.Wakayama T, Yanagimachi R. Cloning of male mice from adult tail-tip cells. Nat Genet 1999; 22: 127–128. [DOI] [PubMed] [Google Scholar]
- 28.Wakisaka N, Inoue K, Ogonuki N, Miki H, Sekita Y, Hanaki K, Akatsuka A, Kaneko-Ishino T, Ishino F, Ogura A. Ultrastructure of placental hyperplasia in mice: comparison of placental phenotypes with three different etiologies. Placenta 2008; 29: 753–759. [DOI] [PubMed] [Google Scholar]
- 29.Inoue K, Kohda T, Sugimoto M, Sado T, Ogonuki N, Matoba S, Shiura H, Ikeda R, Mochida K, Fujii T, Sawai K, Otte AP, Tian XC, Yang X, Ishino F, Abe K, Ogura A. Impeding Xist expression from the active X chromosome improves mouse somatic cell nuclear transfer. Science 2010; 330: 496–499. [DOI] [PubMed] [Google Scholar]
- 30.Kishigami S, Mizutani E, Ohta H, Hikichi T, Thuan NV, Wakayama S, Bui H-T, Wakayama T. Significant improvement of mouse cloning technique by treatment with trichostatin A after somatic nuclear transfer. Biochem Biophys Res Commun 2006; 340: 183–189. [DOI] [PubMed] [Google Scholar]
- 31.Schumacher A, Koetsier PA, Hertz J, Doerfler W. Epigenetic and genotype-specific effects on the stability of de novo imposed methylation patterns in transgenic mice. J Biol Chem 2000; 275: 37915–37921. [DOI] [PubMed] [Google Scholar]
- 32.Leitch HG, McEwen KR, Turp A, Encheva V, Carroll T, Grabole N, Mansfield W, Nashun B, Knezovich JG, Smith A, Surani MA, Hajkova P. Naive pluripotency is associated with global DNA hypomethylation. Nat Struct Mol Biol 2013; 20: 311–316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Matsui Y, Zsebo K, Hogan BL. Derivation of pluripotential embryonic stem cells from murine primordial germ cells in culture. Cell 1992; 70: 841–847. [DOI] [PubMed] [Google Scholar]
- 34.Kanatsu-Shinohara M, Ogonuki N, Inoue K, Miki H, Ogura A, Toyokuni S, Shinohara T. Long-term proliferation in culture and germline transmission of mouse male germline stem cells. Biol Reprod 2003; 69: 612–616. [DOI] [PubMed] [Google Scholar]
- 35.Kanatsu-Shinohara M, Ogonuki N, Iwano T, Lee J, Kazuki Y, Inoue K, Miki H, Takehashi M, Toyokuni S, Shinkai Y, Oshimura M, Ishino F, Ogura A, Shinohara T. Genetic and epigenetic properties of mouse male germline stem cells during long-term culture. Development 2005; 132: 4155–4163. [DOI] [PubMed] [Google Scholar]
- 36.Ishii K, Kanatsu-Shinohara M, Shinohara T. Cell-cycle-dependent colonization of mouse spermatogonial stem cells after transplantation into seminiferous tubules. J Reprod Dev 2014; 60: 37–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ikeda R, Shiura H, Numata K, Sugimoto M, Kondo M, Mise N, Suzuki M, Greally JM, Abe K. Large, male germ cell-specific hypomethylated DNA domains with unique genomic and epigenomic features on the mouse X chromosome. DNA Res 2013; 20: 549–565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kanatsu-Shinohara M, Inoue K, Lee J, Yoshimoto M, Ogonuki N, Miki H, Baba S, Kato T, Kazuki Y, Toyokuni S, Toyoshima M, Niwa O, Oshimura M, Heike T, Nakahata T, Ishino F, Ogura A, Shinohara T. Generation of pluripotent stem cells from neonatal mouse testis. Cell 2004; 119: 1001–1012. [DOI] [PubMed] [Google Scholar]
- 39.Takashima S, Hirose M, Ogonuki N, Ebisuya M, Inoue K, Kanatsu-Shinohara M, Tanaka T, Nishida E, Ogura A, Shinohara T. Regulation of pluripotency in male germline stem cells by Dmrt1. Genes Dev 2013; 27: 1949–1958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tanaka T, Kanatsu-Shinohara M, Hirose M, Ogura A, Shinohara T. Pluripotent cell derivation from male germline cells by suppression of Dmrt1 and Trp53. J Reprod Dev 2015; 61: 473–484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Anderson PD, Nelson VR, Tesar PJ, Nadeau JH. Genetic factors on mouse chromosome 18 affecting susceptibility to testicular germ cell tumors and permissiveness to embryonic stem cell derivation. Cancer Res 2009; 69: 9112–9117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Matin A, Collin GB, Asada Y, Varnum D, Nadeau JH. Susceptibility to testicular germ-cell tumours in a 129.MOLF-Chr 19 chromosome substitution strain. Nat Genet 1999; 23: 237–240. [DOI] [PubMed] [Google Scholar]
- 43.Najm FJ, Chenoweth JG, Anderson PD, Nadeau JH, Redline RW, McKay RD, Tesar PJ. Isolation of epiblast stem cells from preimplantation mouse embryos. Cell Stem Cell 2011; 8: 318–325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ohtsuka S, Niwa H. The differential activation of intracellular signaling pathways confers the permissiveness of embryonic stem cell derivation from different mouse strains. Development 2015; 142: 431–437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Halley-Stott RP, Pasque V, Gurdon JB. Nuclear reprogramming. Development 2013; 140: 2468–2471. [DOI] [PubMed] [Google Scholar]