Abstract
While next-generation sequencing (NGS) has become the primary technology for discovering gene fusions, we are still faced with the challenge of ensuring that causative mutations are not missed while minimizing false positives. Currently, there are many computational tools that predict structural variations (SV) and gene fusions using whole genome (WGS) and transcriptome sequencing (RNA-seq) data separately. However, as both WGS and RNA-seq have their limitations when used independently, we hypothesize that the orthogonal validation from integrating both data could generate a sensitive and specific approach for detecting high-confidence gene fusion predictions. Fortunately, decreasing NGS costs have resulted in a growing quantity of patients with both data available. Therefore, we developed a gene fusion discovery tool, INTEGRATE, that leverages both RNA-seq and WGS data to reconstruct gene fusion junctions and genomic breakpoints by split-read mapping. To evaluate INTEGRATE, we compared it with eight additional gene fusion discovery tools using the well-characterized breast cell line HCC1395 and peripheral blood lymphocytes derived from the same patient (HCC1395BL). The predictions subsequently underwent a targeted validation leading to the discovery of 131 novel fusions in addition to the seven previously reported fusions. Overall, INTEGRATE only missed six out of the 138 validated fusions and had the highest accuracy of the nine tools evaluated. Additionally, we applied INTEGRATE to 62 breast cancer patients from The Cancer Genome Atlas (TCGA) and found multiple recurrent gene fusions including a subset involving estrogen receptor. Taken together, INTEGRATE is a highly sensitive and accurate tool that is freely available for academic use.
Chromosomal rearrangements represent the most prevalent category of somatic aberrations in cancer genomes, often leading to the juxtaposition of two genes, creating gene fusions. Gene fusions have served as exquisitely specific diagnostic markers, prognostic indicators, and therapeutic targets (Druker et al. 2006). The unparalleled depth of next-generation sequencing (NGS) has revealed novel gene fusions in numerous solid tumors as exemplified by fusions involving the E26 transformation-specific (ETS) transcription factor family members in prostate cancer (Tomlins et al. 2007), MAST and NOTCH kinases in breast cancer (Robinson et al. 2011), and ALK, ROS1, and RET fusions in lung cancer (Takeuchi et al. 2012).
To date, many groups have used whole-genome sequencing (WGS) to identify structural variations (SV), a subset of which may produce gene fusions. Despite some successes, existing bioinformatics tools such as BreakDancer (Chen et al. 2009), VariationHunter (Hormozdiari et al. 2010), CREST (Wang et al. 2011), and PRISM (Jiang et al. 2012) are hindered by intra-tumor heterogeneity, alignment to repetitive genomic sequences, technical artifacts (i.e., library preparation), poor coverage, and a large number of false-positive calls owing to sequencing errors. The failure to predict some SVs using WGS data would therefore result in the corresponding gene fusion product being missed. Additionally, it is unclear whether SVs predicted to produce a gene fusion are expressed in the absence of RNA-seq expression data. Therefore, many groups have focused on using RNA-seq for gene fusion discovery as it enriches for expressed events that are more likely to be functional.
Currently, many RNA-seq gene fusion discovery algorithms utilize spanning reads (one read partially aligns to both genes corresponding to the fusion junction) or encompassing reads (each read of a pair aligns to a different gene, thereby surrounding the fusion junction) such as TopHat-Fusion (Kim and Salzberg 2011), deFuse (McPherson et al. 2011a), ChimeraScan (Iyer et al. 2011), BreakFusion (Chen et al. 2012), FusionCatcher (Nicorici et al. 2014), pyPRADA (Torres-Garcia et al. 2014), and TRUP (Fernandez-Cuesta et al. 2015). However, despite the successful application of these algorithms to discover gene fusions, a recent comparison of eight gene fusion discovery tools revealed a lot of variability between callers. Most tools report a very high number of false-positive chimeras (Carrara et al. 2013), highlighting the ongoing struggle to balance sensitivity and specificity of fusion detection. Some of the various factors contributing to false positives and false negatives include artifacts and mapping errors in RNA-seq data, reliance on a comprehensive transcriptome reference, repetitive regions, and low-expressing gene fusions that may appear as background signal. Furthermore, while a large portion of unmapped reads may represent sequencing errors, artifacts, or the limitations of split-read mapping tools, it is possible that a subset of these unmapped reads could represent critical reads spanning the gene fusion junction. Therefore, more efficient split-read mapping methods are necessary to find these critical reads among the potential noise.
While WGS or RNA-seq have their limitations when analyzed independently, the orthogonal validation from integrating RNA-seq and WGS could generate a sensitive and specific approach for detecting high-confidence gene fusion predictions. Since WGS and RNA-seq data are generated separately, presumably they do not share the same artifacts and noise and therefore will result in fewer false positives. This in turn will facilitate the prioritization of gene fusion predictions as biologically relevant gene fusions that are often masked by false positives. Furthermore, weak sequence evidence by both WGS and RNA-seq could help detect gene fusions expressed at low levels that may have appeared as background noise when analyzing only WGS or RNA-seq data. In addition to improving gene fusion discovery, integrating WGS and transcriptome data can provide evidence about the gene fusion biology. For instance, recent studies have shown the importance of read-through chimeras, which involve two adjacent genes in the same coding orientation, in multiple cancers (Maher et al. 2009; Zhang et al. 2012; Varley et al. 2014). When analyzing RNA-seq data alone, a chimera transcript is typically classified as a read-through based on the proximity and orientation of the genes. However, it is not possible to rule out that the event is the by-product of a focal deletion. Therefore, the presence or absence of a genomic event by integrating WGS and RNA-seq could improve the classification of RNA chimeras.
In this study, we describe a new method, INTEGRATE, for detecting expressed gene fusions by leveraging the advantages of both WGS and RNA-seq generated from the same individual. Existing methods that use both whole-genome and RNA-seq data include Comrad (McPherson et al. 2011b), nFuse (McPherson et al. 2012), and BreakTrans (Chen et al. 2013). Comrad (McPherson et al. 2011b) is a dedicated gene fusion calling program that simultaneously uses both encompassing RNA-seq and WGS paired reads. It uses an integer linear programming algorithm to assign repetitive reads that minimizes differences of WGS data sets, RNA-seq data sets, and the reference genome. nFuse (McPherson et al. 2012) is a computational tool intended to identify complex genomic rearrangements from whole-genome data with the help of transcriptome sequencing data. More recently, BreakTrans (Chen et al. 2013) was developed to intersect predicted gene fusions that correspond with SV nominations by analyzing the output of independent gene fusion and SV prediction tools. Here, we developed INTEGRATE, which simultaneously uses both RNA-seq and WGS encompassing and spanning reads to focus on the discovery of expressed gene fusions. To prioritize expressed gene fusions caused by SVs, INTEGRATE first utilizes mapped and unmapped RNA-seq reads followed by analysis of WGS reads from tumor, and if available, a normal sample. To minimize run time and memory requirements without sacrificing accuracy, INTEGRATE uses discordant RNA-seq reads to construct a gene fusion graph connecting genes involved in a putative fusion event. This enables all of the unaligned RNA-seq reads that could serve as spanning junction reads to undergo split-read mapping against only the Burrows-Wheeler Transform (BWT) for the relevant gene pair in the fusion graph instead of against the whole genome or whole transcriptome (see Methods for details). The gene fusion graph also avoids nominating false positives since INTEGRATE realigns encompassing and spanning reads to only the BWTs for the relevant gene pair in the graph, thereby decreasing spurious mappings that may occur when aligning to the whole genome or whole transcriptome. Here, we will show that INTEGRATE is an efficient gene fusion discovery tool that has both high sensitivity and accuracy. INTEGRATE can be downloaded at https://sourceforge.net/projects/integrate-fusion/.
Results
Overview of INTEGRATE
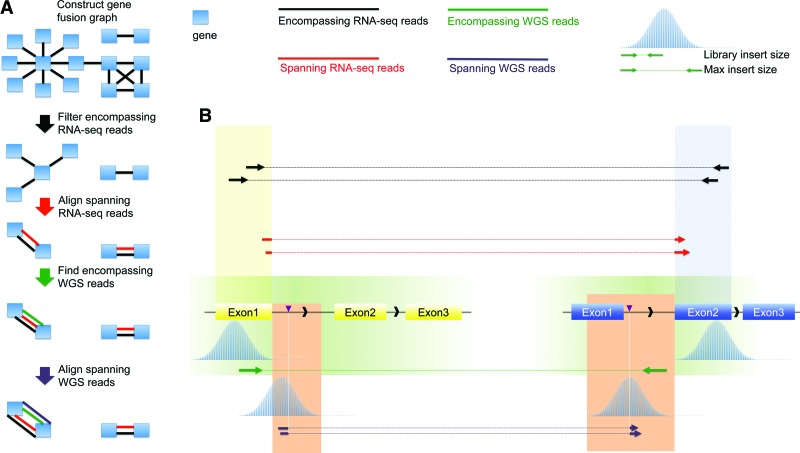
INTEGRATE is designed to discover gene fusions using RNA-seq and WGS paired-end sequencing reads properly aligned to the reference genome in BAM format. Unlike many gene fusion tools which are programmed to use a specific reads mapping tool, INTEGRATE is implemented with the flexibility to use reads aligned by different tools, including GSNAP (Wu and Nacu 2010), TopHat2 (Kim et al. 2013), and STAR (Dobin et al. 2013). Since we are most interested in expressed gene fusions, INTEGRATE first utilizes mapped and unmapped RNA-seq reads, followed by analysis of WGS reads from tumor and, if available, a normal sample. INTEGRATE uses two types of reads, encompassing and spanning reads. Encompassing reads are pairs of reads with each in the pair aligned to a different gene, thereby surrounding the fusion junctions or genomic breakpoints, and spanning reads are reads partially aligned to both genes corresponding to a fusion junction or both flanking regions of an SV. As shown in Figure 1, INTEGRATE is comprised of the following steps: (1) Construct gene fusion graph using discordant, or encompassing, RNA-seq reads; (2) remove edges corresponding to discordant reads that have a concordant suboptimal mapping or have low weights due to excessive multimapping; (3) map previously unaligned RNA-seq reads between gene nodes as split-reads to reconstruct fusion junctions (Supplemental Fig. 1); (4) retrieve encompassing WGS reads corresponding to focal regions surrounding fusion junctions; and (5) map spanning WGS reads to focal regions with the guidance of encompassing WGS reads to reconstruct genomic breakpoints. Once completed, INTEGRATE outputs the gene fusion candidates with the exact fusion junctions sorted according to the quantity of supporting WGS and RNA-seq reads.
Figure 1.
Overview of INTEGRATE. (A) INTEGRATE establishes a gene fusion graph using encompassing RNA-seq reads (black lines) to connect nodes or genes (blue rectangles). Edges of the fusion graph are removed following various filtering steps before undergoing a targeted split-read alignment involving the remaining edges. Encompassing and spanning split-read realignment and mapping are performed on BWTs of gene nodes (Supplemental Fig. 1). Encompassing WGS reads are retrieved from regions determined by spanning RNA-seq reads. Spanning WGS reads are aligned to the regions indicated by encompassing WGS reads (steps indicated by green and purple arrows; also see B). (B) When encompassing (black) and spanning (red) RNA-seq reads have been mapped to the genes involved in a gene fusion, the encompassing WGS reads (green) are expected from focal encompassing WGS regions (green area) bounded by maximum insert size upstream of or downstream from the fusion junctions of the transcripts. The spanning WGS reads (purple) are expected to align within focal WGS regions (orange area) bounded by fusion junction and maximum insert size downstream from the encompassing WGS reads.
To prioritize gene fusion candidates, INTEGRATE reports fusions in tiers corresponding to the level of sequencing support and potential biology (Supplemental Fig. 2). Tiers 1, 2, and 3 all involve gene fusions with canonical exonic boundaries. Tier 1 candidates have the highest confidence as they have both encompassing and spanning RNA-seq and WGS reads supporting a gene fusion. Tier 2 gene fusion candidates also have both WGS and RNA-seq read support; however, they only have encompassing WGS read support and lack spanning WGS reads. Tier 3 lacks any WGS read support but has both encompassing and spanning RNA reads. However, Tier 3 includes both non-read-through gene fusions (Tier 3-nr) and read-throughs (Tier 3-r).
Application to HCC1395 breast cancer cells
To evaluate INTEGRATE, we used HCC1395 breast cancer cells because both SVs and gene fusions have been previously characterized in this cell line (Stephens et al. 2009; Robinson et al. 2011; Kalyana-Sundaram et al. 2012), multiple gene fusions have been experimentally validated using qRT-PCR (Lambros et al. 2011), and because of the availability of a matched B lymphoblast cell line (HCC1395BL) to be used as our normal comparator. We sequenced ∼321 million tumor RNA-seq paired-end reads, ∼339 million tumor RNA-seq paired-end reads, ∼1884 million tumor WGS paired-end reads (∼63× coverage), and ∼1031 million normal WGS paired-end reads (∼34× coverage). INTEGRATE was run using the RNA-seq reads aligned by GSNAP (Wu and Nacu 2010), TopHat2 (Kim et al. 2013), and STAR (Dobin et al. 2013). WGS reads were aligned using BWA (Li and Durbin 2009). Interestingly, we found that different alignment tools affected the final list of gene fusion predictions. Using alignments from GSNAP, TopHat2, and STAR and running INTEGRATE with default parameters (i.e., two encompassing RNA-seq reads), we discovered 110, 68, and 68 gene fusion candidates, respectively.
To compare the performance of INTEGRATE to other available algorithms, we reanalyzed the HCC1395 data with three WGS and RNA-seq callers (Comrad [McPherson et al. 2011b], nFuse [McPherson et al. 2012], and BreakTrans [Chen et al. 2013]) and five commonly used and recently published RNA-seq gene fusion tools (TopHat-Fusion [Kim and Salzberg 2011], ChimeraScan [Iyer et al. 2011], FusionCatcher [Nicorici et al. 2014], pyPRADA [Torres-Garcia et al. 2014], and TRUP [Fernandez-Cuesta et al. 2015]). BreakTrans was provided with fusion and SV candidates called by BreakDancer (Chen et al. 2009). For all methods, common false-positive gene fusion predictions were filtered (Methods) to produce high-confidence predictions for each program. This resulted in a range of four to 110 gene fusion candidates across the programs. An aggregate of the top gene fusion candidates nominated by each program resulted in 240 gene fusion candidates (Supplemental Fig. 3; Supplemental Table 1). After applying our filtering steps, of the eight additional programs, nFuse nominated the most gene fusion candidates (n = 103). ChimeraScan, FusionCatcher, TopHat-Fusion, and pyPRADA had a moderate number of gene fusion candidates with 54, 36, 12, and 17, respectively. TRUP, Comrad, and BreakTrans nominated a limited number of gene fusion candidates with 6, 4, and 4, respectively.
Through our comprehensive analysis, we found that two previously reported gene fusions, KCNQ5-RIMS1 and BCAR3-ABCA4, are not called by any of the nine methods. Subsequent manual inspection did not identify any supporting reads for the two missed gene fusions (KCNQ5-RIMS1 and BCAR3-ABCA4) in our data set. This is not surprising for KCNQ5 and RIMS1, as the DNA coverage shows an obvious aberration but the expression levels of KCNQ5 and RIMS1 are very low (Supplemental Fig. 4). In contrast, the fusion between BCAR3 and ABCA4 was previously identified by RNA-seq but was never detected by WGS or experimentally validated. Therefore, its absence in our data could suggest that it may not be expressed in our data or it is a false positive. INTEGRATE is the only program to detect all seven previously discovered gene fusions, whereas the other programs detected between 2 and 5 of these gene fusions.
Validation of HCC1395 gene fusions
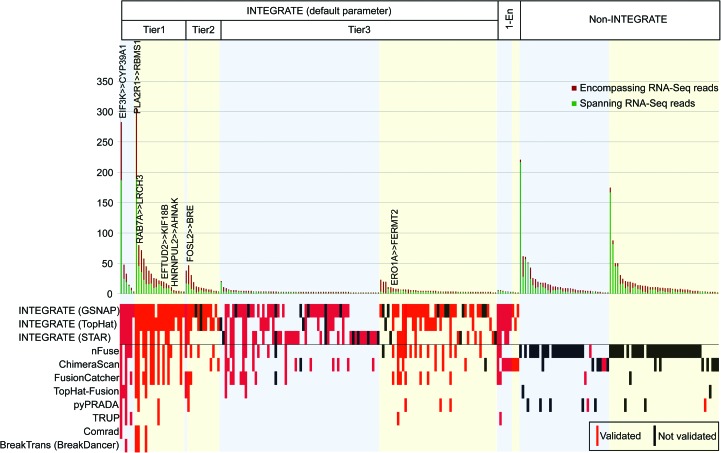
To evaluate the accuracy of the predictions from all these methods, we used cDNA-Capture (Methods; Cabanski et al. 2014) which combines RNA-seq with an enrichment step using custom probes targeting 240 gene fusion candidates (Supplemental Table 1). The 240 gene fusion candidates are an aggregate of the top gene fusion candidates nominated by each program. As shown in Figure 2, we experimentally validated 138 gene fusions (see Supplemental Table 2 for validated fusion junctions), of which 123 are called by INTEGRATE using default parameters (two encompassing RNA-seq reads), nine can be detected by INTEGRATE using one encompassing RNA-seq read, and only six were not detected by INTEGRATE.
Figure 2.
Gene fusion validation. Targeted cDNA-capture validation was attempted for 240 gene fusion candidates called by INTEGRATE and eight additional gene fusion detection methods, resulting in the validation of 138 gene fusions. Gene fusion candidates nominated by INTEGRATE using default parameters and a threshold of one encompassing RNA-seq read (1-En) are shown on the left, whereas candidates nominated by eight additional programs, and not INTEGRATE (“non-INTEGRATE”), are shown on the right. See Supplemental Figure 2 for tiers of INTEGRATE. In each category, gene fusion candidates are further divided into rearrangement classes: inter-chromosomal (blue shade) and intra-chromosomal (yellow shade), and sorted in descending order of total RNA-seq read support (dark red bar—encompassing RNA-seq reads; green bar—spanning RNA-seq reads). Previously reported gene fusions are written at the top of the bars. In the lower panel, each row corresponds to the gene fusion candidates nominated by each program. INTEGRATE, Comrad, BreakTrans, and nFuse use both RNA-seq and WGS data. INTEGRATE is shown using the RNA-seq alignments from GSNAP, TopHat2, and STAR, separately. ChimeraScan, Tophat-Fusion, FusionCatcher, pyPRADA, and TRUP use only RNA-seq data. Red boxes indicate nominated gene fusions that were experimentally validated, black boxes indicate nominated gene fusions that did not have validation read support, and the lack of a red or black box indicates an algorithm did not nominate the gene fusion candidate.
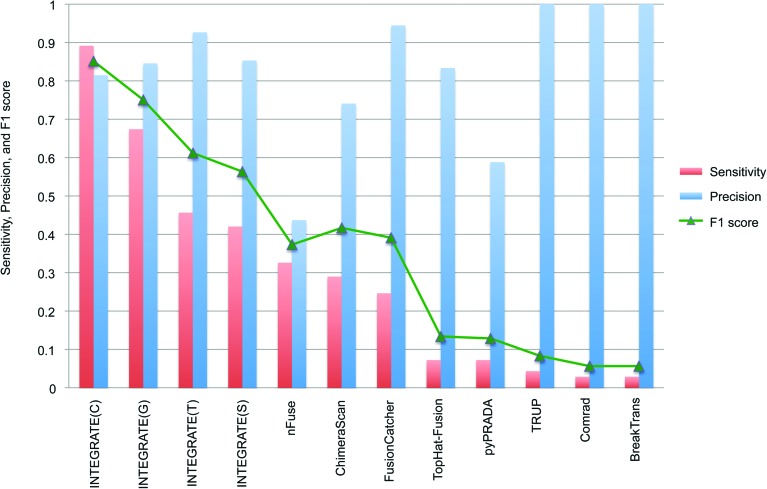
Next, we used the 138 validated gene fusions discovered using nine methods as the gold standard for comparing the sensitivity and precision of each method as shown in Figure 3. The combined set of INTEGRATE gene fusion predictions using all three alignment tools has the highest sensitivity (89%) while maintaining a high precision (81%). While the default parameters for INTEGRATE require two encompassing RNA-seq reads, a user could modify this threshold to one encompassing RNA-seq read, which had a sensitivity of 95.6% (132/138). Interestingly, while INTEGRATE only missed six validated fusions (out of 138), none of the other programs successfully detected all of the six remaining candidates (Fig. 2), indicating that a user would have to use multiple programs to detect all of the gene fusions. We also found variation based on the alignment tool used. The sensitivity of running INTEGRATE with default parameters using a single alignment tool is 67% for GSNAP, 46% for TopHat2, and 42% for STAR, while maintaining high precision of 85%, 93%, and 85%, respectively. Even when INTEGRATE uses STAR, which had the worst performance of the alignment tools, it still outperformed the next best program, nFuse, which discovered 45 gene fusions but missed 93 gene fusions resulting in a 33% sensitivity and 44% precision. ChimeraScan has a slightly lower sensitivity (29%) than nFuse but has a higher precision of 74%. FusionCatcher has an even lower sensitivity (25%) but a higher precision (95%). The remaining five methods (TopHat-Fusion, pyPRADA, TRUP, Comrad, and BreakTrans) have sensitivities lower than 10%. TRUP, Comrad, and BreakTrans all have a precision of 100%; however, they miss more than 132 validated gene fusions. Overall, the accuracy (F1 score) of INTEGRATE, based on the combination of sensitivity and precision, is the highest of all nine tools (Fig. 3).
Figure 3.
Comparison of INTEGRATE and eight additional fusion calling methods. Sensitivity and precision (red bar and blue bar, respectively) of each method were calculated using a gold standard, which is the experimentally validated gene fusion called by nine methods, sorted in decreasing order of sensitivity. INTEGRATE applied with default parameters, using aligned reads generated by GSNAP, TopHat2, and STAR is indicated by G, T, and S, respectively. The combination of the three alignment tools is indicated by C. The accuracy, or F1 score, based on the combination of sensitivity and precision is shown with a green triangle.
Overall, INTEGRATE is a highly sensitive method, resulting in the discovery of an additional 125 gene fusions that eluded earlier studies. Interestingly, all of the previously discovered gene fusions have more than five supporting RNA-seq reads (Fig. 2; Supplemental Table 1) with a maximum of 306 supporting RNA-seq reads. In contrast, most of the newly discovered fusions have low expression levels, as exemplified by 85 out of the 138 fusions having ≤5 RNA-seq reads and 44 of these 85 gene fusions having only one spanning read. Additionally, as shown in Figure 2, the eight additional programs typically detect the more highly expressed gene fusions, whereas the gene fusions with lower expression levels are typically missed by more methods. For example, the most highly expressed of the seven previously reported gene fusions, EIF3K-CYP39A1 (283 reads), PLA2R1-RBMS1 (306 reads), and RAB7A-LRCH3 (80 reads), are detected by seven, six, and seven methods, respectively. EFTUD2-KIF18B (20 reads), FOSL2-BRE (47 reads), and ERO1A-FERMT2 (nine reads) were found by three, three, and two methods, respectively. However, the lowly expressed gene fusion HNRNPUL2-AHNAK, having only six supporting reads, is only detected by INTEGRATE. Interestingly, while INTEGRATE shows increased sensitivity, as exemplified by its ability to detect lowly expressed gene fusions compared to other methods, it also discovered highly expressed gene fusions that were missed by other methods. For example, INTEGRATE was the only method that detected the Tier 1 gene fusion MAVS-PANK2 (Supplemental Fig. 5) that has 38 supporting RNA-seq reads.
We also compared the RNA-seq expression levels with the read support from cDNA-Capture, revealing a positive correlation (0.95) (Supplemental Fig. 6). This shows that gene fusion validation, similar to RNA-seq, is also heavily dependent on the expression of the gene fusion transcript. Therefore, it is unclear whether low-expressing gene fusions predicted by INTEGRATE that were not validated are actually false positives or whether additional validation sequencing would eventually confirm their presence as found by RNA-seq data. In contrast, 60 of the 71 gene fusion candidates that did not validate and were nominated by other methods, but not by INTEGRATE, had ≥5 RNA-seq reads. Therefore, the higher quantity of supporting RNA-seq reads coupled with lack of read support from cDNA-Capture further suggests that these nominations are false positives.
In addition to discovering Tier 1 and 2 gene fusions accurately by combining RNA-seq and WGS data (38 out 40 [95%] are validated) (see Supplemental Table 3 for genomic breakpoints of Tier 1 gene fusions), INTEGRATE also reliably detected candidates supported only by RNA-seq reads. Eighty-five out of 111 Tier 3-nr gene fusion candidates were validated, resulting in a precision of ∼77%. Fifty-one out of the 85 (60%) validated Tier 3-nr gene fusions were missed by all the other methods.
Overall, we found that INTEGRATE only missed six candidates; however, none of these predictions were found to have a high level of read support (all have <5 reads). Additionally, the six gene fusions missed by INTEGRATE were not nominated by a single program but were nominated by one of four programs (nFuse, ChimeraScan, FusionCatcher, and pyPRADA). Conversely, the other four programs (TopHat-Fusion, TRUP, Comrad, and BreakTrans) did not identify any validated gene fusions that were missed by INTEGRATE.
Application to TCGA breast cancer patient cohort
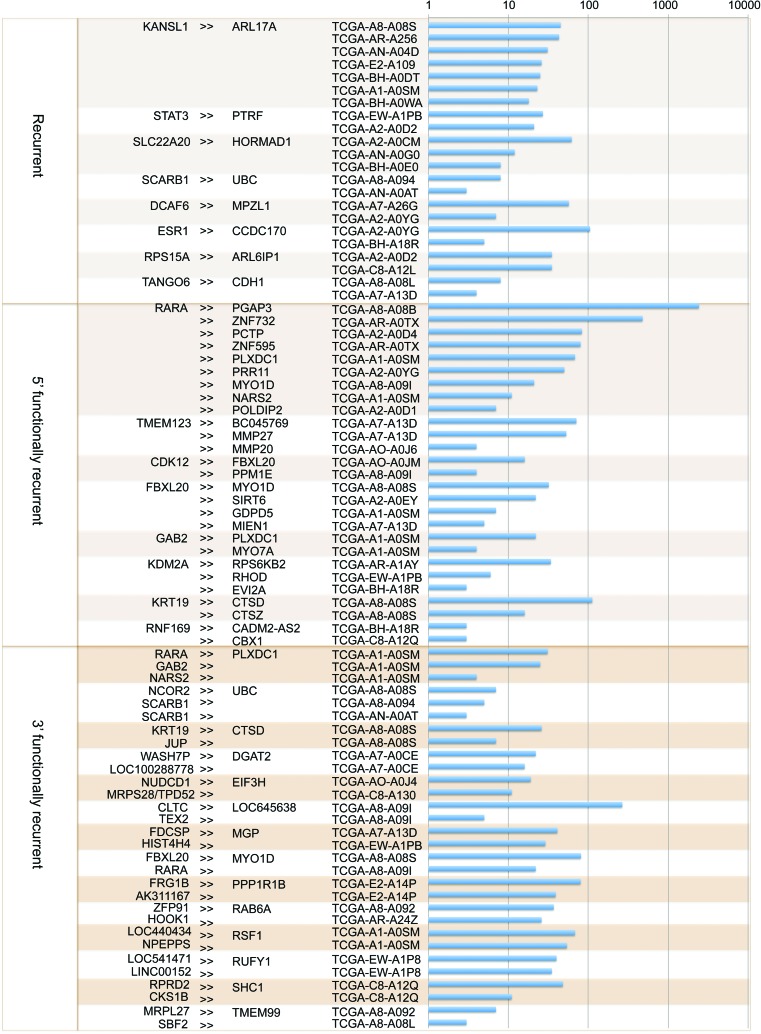
Next, we applied INTEGRATE to a cohort of 62 breast cancer patients (Supplemental Table 4), generated by The Cancer Genome Atlas (TCGA) Research Network (http://cancergenome.nih.gov/), that had both whole-genome and RNA-seq data available (The Cancer Genome Atlas Network 2012). INTEGRATE discovered 347 gene fusions involving both WGS and RNA-seq read support and 132 non-read-through gene fusions with only RNA-seq reads (Supplemental Table 4). This revealed eight recurrent gene fusions, six of which (DCAF6-MPZL1, ESR1-CCDC170, KANSL1-ARL17A, RPS15A-ARL6IP1, STAT3-PTRF, and TANGO6-CDH1) were reconstructed by INTEGRATE with the fusion junctions and genomic breakpoints (Fig. 4). SLC22A20-HORMAD1 and SCARB1-UBC were called by only supporting RNA-seq reads (Fig. 4). The most frequent gene fusion, KANSL1-ARL17A/ARL17B, has been recently reported (Wen et al. 2012). Additionally, the estrogen receptor gene fusion ESR1-CCDC170 (Fig. 5A,B) was recently reported in breast cancer (Sakarya et al. 2012).
Figure 4.
Recurrent and functionally recurrent gene fusions in a TCGA 62 breast cancer patient cohort. Gene fusions are listed in the order of recurrent, 5′ functionally recurrent, and 3′ functionally recurrent. The first column shows the 5′ genes and the second column shows the 3′ genes. The third column is the TCGA names of the samples. Bar chart in the fourth column shows the log scale value of the quantity of supporting RNA-seq reads for each gene fusion.
Figure 5.
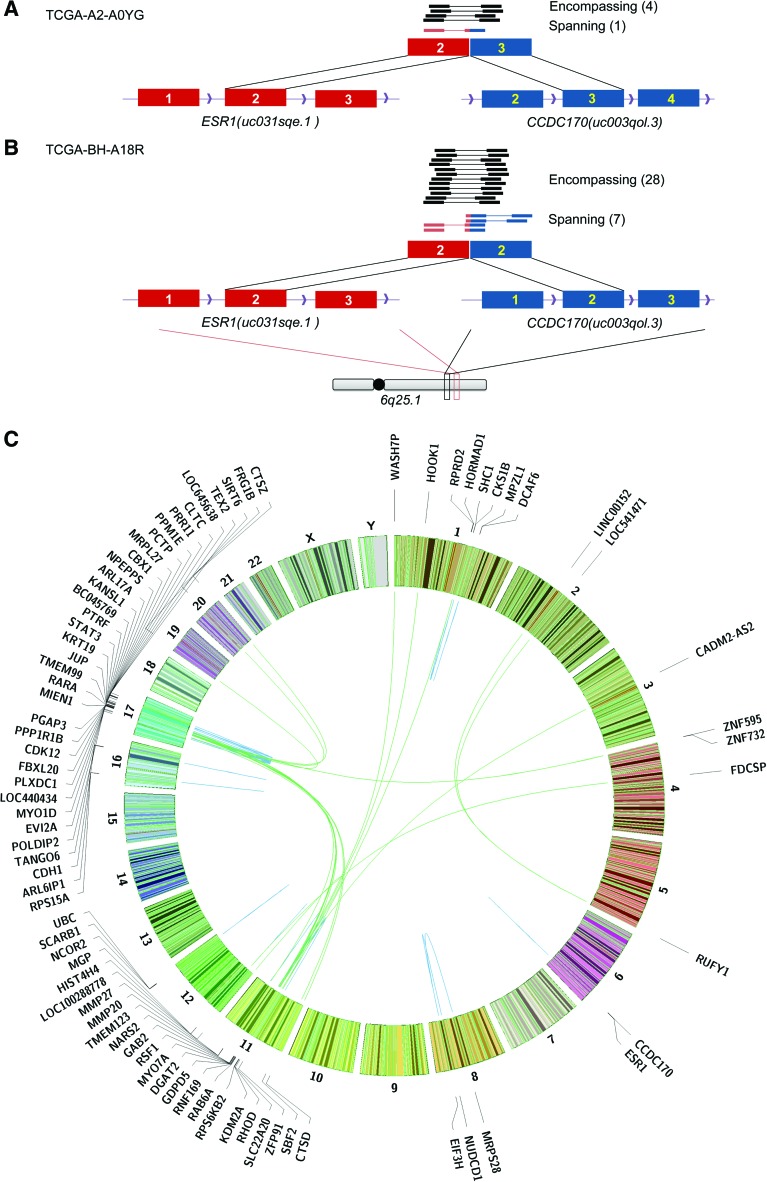
Hotspots of gene fusions in 62 TCGA breast cancer patients. (A) ESR1-CCDC170 fusion in TCGA-A2-A0YG. (B) ESR1-CCDC170 fusion in TCGA-BH-A18R. ESR1 (red) and CCDC170 (blue) are on the forward strand in region 6q25.1, and the 5′ gene ESR1 is downstream from the 3′ gene CCDC170. The two fusions share the same 5′ exon at ESR1 (Exon 2 of transcript uc031sqe.1), but 3′ exons of CCDC170 are different (Exons 2 and 3 of transcript uc003qol.3). (C) Circos plot of recurrent and functionally recurrent gene fusions detected by INTEGRATE. The green lines indicate inter-chromosomal gene fusions, and the blue lines indicate intra-chromosomal gene fusions. The names of the genes involved in each fusion are plotted on the outside of the circle. The gene fusions associate with several hotspots on Chromosomes 1, 11, and 17.
In addition to recurrent gene fusions, we hypothesized that there may be a selective pressure to alter a gene in order to achieve a similar molecular consequence, as exemplified by ETS family members (Papas et al. 1989), BRAF (Stratton et al. 2004), MAST/NOTCH (Robinson et al. 2011), which we refer to as being functionally recurrent. Therefore, we sought to identify genes that are involved in multiple fusions with the breakpoint occurring with the same 5′ or 3′ exon. We found eight conserved 5′ genes and 14 conserved 3′ genes representing 51 non-read-through gene fusions across 25 patients (Fig. 4). Many of these genes have been previously reported as gene fusion partners in breast cancer, e.g., RARA (Edgren et al. 2011), CDK12 (Asmann et al. 2011; Natrajan et al. 2014), FBXL20 (Mardis et al. 2009; Robinson et al. 2011; Kalyana-Sundaram et al. 2012), GAB2 (Stephens et al. 2009), PLXDC1 (Robinson et al. 2011), CTSD (Asmann et al. 2012), EIF3H (Edgren et al. 2011; Kalyana-Sundaram et al. 2012), MGP (Asmann et al. 2012; Kalyana-Sundaram et al. 2012), PPP1R1B (Robinson et al. 2011), and RAB6A (McPherson et al. 2011a).
Closer examination of the genomic locations for the recurrent and functionally recurrent gene fusions reveals multiple hotspots of gene fusions (Fig. 5C) on Chromosomes 1, 11, and 17. In total, 44 out of the 65 recurrent and functionally recurrent gene fusions harbor partners that reside in these three commonly altered regions (Supplemental Table 5). There are three recurrent and functionally recurrent gene fusions involving a gene in 1q21.3. Seventeen gene fusions involve a gene residing in 11q13.1–11q14.1. Thirty-three gene fusions involved a gene that resides in the consecutive region of bands 17q11.2, 17q12, and 17q21–23. It is plausible that many of these gene fusions may be passenger aberrations corresponding to recurrent amplicons. However, a subset may represent potentially relevant gene fusions in breast cancer as exemplified by ESR1 translocations (Veeraraghavan et al. 2014). Additionally, while the majority of gene fusions reside in commonly altered regions, there is still a subset of recurrent and functionally recurrent gene fusions that are not the by-product of a copy number event and therefore may warrant further exploration.
Read-through transcription
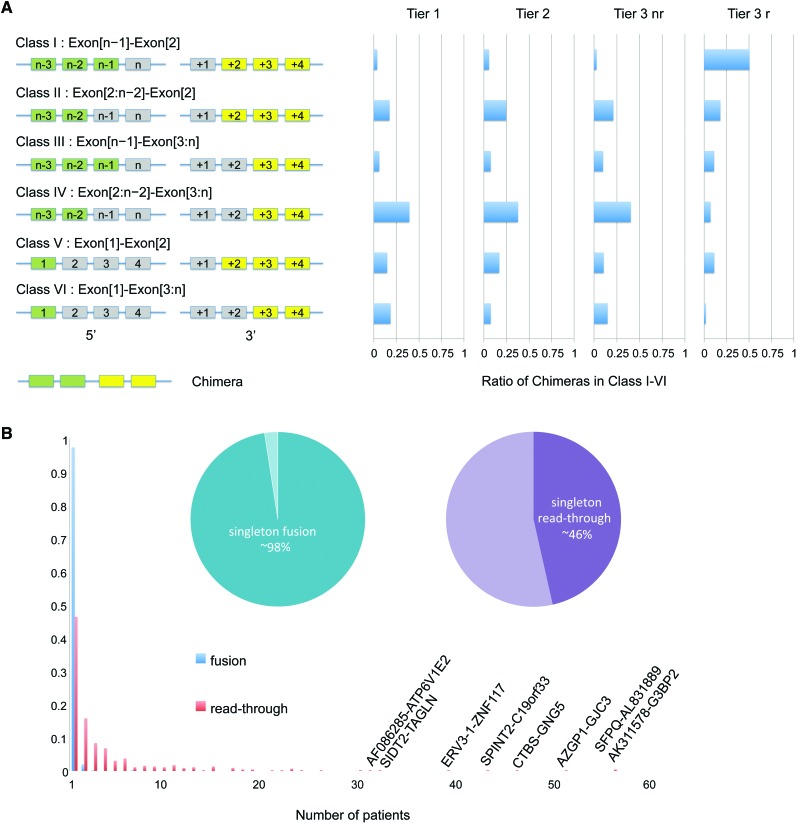
INTEGRATE classifies a read-through as a chimera involving two adjacent genes on the same strand, with the 5′ gene being upstream, but lacking any WGS read support. For example, INTEGRATE identified 288 read-throughs in HCC1395 cells and 453 read-throughs in a cohort of 62 TCGA patients (Supplemental Tables 6, 7). We examined the difference between gene fusions associated with genomic rearrangements (Tiers 1, 2, and 3-nr) and read-throughs (Tier 3-r) by classifying the patterns of exons involved in the fusion junctions. This includes any combination of a 5′ gene involving (1) the first exon (Exon[1]), (2) the second to the last exon (Exon[n−1], where n is the number of exons), and (3) all other exons (Exon[2:n−2]) with a 3′ gene involving either the second exon (Exon[2]) or any downstream exons (Exon[3:n]). This revealed six classes: (I) Exon[n−1]-Exon[2]; (II) Exon[2:n−2]-Exon[2]; (III) Exon[n−1]-Exon[3:n]; (IV) Exon[2:n−2]-Exon[3:n]; (V) Exon[1]-Exon[2]; and (VI) Exon[1]-Exon[3:n]. As shown in Figure 6A, we observed different patterns between gene fusions associated with genomic rearrangements and read-throughs. The majority of Tier 1, 2, and 3-nr gene fusions (38%–40%) belong to class IV (Exon[2:n−2]-Exon[3:n]), which involves random exons in the middle of the 5′ and 3′ fusion partners, in contrast to 7.2% of read-throughs belonging to class IV. Fifty-one percent of read-throughs belong to class I (Exon[n−1]-Exon[2]), which involves the second to the last exon of a 5′ transcript and the second exon of a 3′ transcript. In contrast, only 4%, 6%, and 3% of Tier 1, 2, and 3-nr fall into class I, respectively. Overall, the exon usage distribution of read-throughs is significantly different from the Tiers 1, 2, and 3-nr (Pearson's χ2 test, each tier has a P < 2.2 × 10−16), whereas Tier 2 and Tier 3-nr are not significantly different from Tier 1 (P-values 0.32 and 0.17, respectively).
Figure 6.
Different patterns between gene fusions and read-throughs. (A) Exons involved in gene fusions and read-throughs follow different patterns. A gene fusion (or read-through) transcript can be categorized into six classes involving the first, second to last, or any other exon of the 5′ gene with either the second or downstream exon of the 3′ gene. (B) Recurrence of gene fusions and read-throughs across 62 breast cancer patients. The horizontal axis is number of patients, and the vertical axis is fraction of events. Blue bars represent gene fusions and red bars represent read-throughs. The pie chart shows the percentage of singleton gene fusions (left) and read-throughs (right).
We were next interested in assessing the recurrence of read-throughs relative to gene fusions derived from genomic events. As shown in Figure 6B, within the 62 patient cohort, 98% of the non-read-throughs predicted by INTEGRATE occur in a single patient, whereas only 46% of read-throughs are singletons (Pearson's χ2 test has a P < 2.2 × 10−16). In addition to 54% of read-throughs occurring in multiple patients, we observed eight read-throughs occurring in more than 30 patients.
Discussion
INTEGRATE is a gene fusion discovery tool that leverages both RNA-seq and whole-genome data. By integrating orthogonal data sets, we demonstrate that INTEGRATE is highly sensitive. This can be exemplified by our discovery of 125 novel gene fusions in HCC1395 in addition to the previously discovered gene fusions reported by three earlier studies (Stephens et al. 2009; Robinson et al. 2011; Kalyana-Sundaram et al. 2012). Many of these gene fusions were expressed at very low levels. For instance, INTEGRATE identified 45 gene fusions with only one fusion junction spanning the RNA-seq read. Due to their low read support, these gene fusions are difficult to detect and may even be filtered by other programs depending on a minimum read support threshold to nominate a fusion. Furthermore, the sensitivity of INTEGRATE revealed low-expressing gene fusions that may have been expressed below a level of detection with the current validation sequencing depth. Unlike DNA-based validation, where we would expect more uniform coverage, the ability to detect and validate expressed gene fusions is highly dependent on expression. Our validation data could underrepresent the accuracy of INTEGRATE, and additional sequencing may confirm the presence of the remaining candidates.
One major reason (other than orthogonal data sets) for the improved sensitivity and accuracy of INTEGRATE is that we use a gene fusion graph that enables us to conduct alignment steps targeting specific gene nodes. In contrast to mapping the true positive spanning RNA-seq reads against the whole genome or whole transcriptome, reducing the search space to only a few relevant genes that may be involved in a gene fusion event increased sensitivity by (1) allowing for the alignment to shorter flanking regions of the spanning RNA-seq reads, (2) tolerating more mismatches and gaps in the alignments, and (3) avoiding false positive alignments that would have occurred outside of the relevant gene nodes caused by repeats and sequencing errors. Furthermore, INTEGRATE uses multiple realignment steps against a small number of relevant gene nodes to reduce false-positive encompassing and spanning reads to achieve a more accurate rate in discovering gene fusions. Interestingly, while INTEGRATE was designed with the intent of using both WGS and RNA-seq data, we found that, in the absence of WGS data, INTEGRATE can still be applied to find high-quality gene fusion candidates using only RNA-seq data. This is likely due to the thorough realignment steps to filter inaccurate alignments that may introduce false-positive gene fusion predictions.
INTEGRATE is also designed to be highly efficient due to multiple design aspects. First, the fusion gene graph that connects the gene nodes guarantees only the relevant RNA-seq reads, i.e., discordant encompassing reads mapped to two genes and spanning reads with their anchors in the graph, undergo thorough alignment and realignment steps. Second, alignments and realignments in gene nodes are performed on the BWTs of the gene nodes with time complexity linear of read length. Third, integration of WGS data is guided by the RNA-seq gene fusion candidates. Only relevant encompassing and spanning WGS reads residing within focal regions near the candidate fusion junctions are considered for detecting SVs.
Currently, only a few existing computational tools focus on using combined data with the intent of balancing sensitivity and specificity. To discover gene fusion candidates, Comrad was designed to simultaneously analyze encompassing, but not spanning, WGS and RNA-seq reads. However, spanning reads offer significant evidence for nominating candidates that can help balance specificity and accuracy. This also represents a significant limitation for being able to reconstruct the genomic breakpoint or the fusion junction accurately. nFuse has a higher sensitivity (33%) than all the other programs evaluated other than INTEGRATE; however, it also had the most nominations of any program (Fig. 2; Supplemental Fig. 3). This is not surprising as nFuse was intended to identify complex genomic rearrangements. Lastly, as BreakTrans integrates SV and gene fusion predictions called by separate tools, its overall performance relies on the sensitivity and accuracy of each individual tool run separately. This will therefore miss low-expressing gene fusions or low-frequency genomic events that are detectable by using orthogonal applications but are not easily detected by either RNA-seq or WGS alone. Additionally, gene fusions with strong transcriptome read support but no genomic evidence potentially due to lack of coverage, highly repetitive sequence, or representing an RNA chimera (i.e., read-through, trans-splicing events) will be missed by requiring both RNA-seq and WGS evidence. Taken together, our comparative analyses demonstrate the clear advances that INTEGRATE has achieved in gene fusion discovery from NGS data.
Unlike many gene fusion prediction tools that ignore read-through or trans-splicing events, INTEGRATE is able to provide valuable insight into RNA chimeras. For instance, RNA callers categorize read-throughs purely based on their close proximity and orientation but have no definitive way to discriminate between a genomic or transcriptomic event in the absence of genomic data. However, INTEGRATE can identify events that may be classified as read-throughs based on genomic location but in fact are due to a genomic deletion. This in turn could have important implications in the underlying biology. For instance, we have shown that read-throughs have different patterns of exon usage and prevalence across patients compared to genomic-based events. First, the patterns of exon utilization support that read-throughs are due to splicing of a 3′ exon in the 5′ partner to a 5′ exon of the 3′ gene partner. In contrast, genomic-based gene fusions appear to occur more randomly. Second, read-throughs appear to be recurrent whereas genomic events are typically patient-specific. Of the few recurrent genomic events, they typically occur in a small subset of patients, whereas read-throughs were observed in up to 30+ patients. Taken together, the ability to distinguish a genomic event from a common read-through event could reveal genomic mutations that could serve as valuable biomarkers.
INTEGRATE has also been implemented to improve the interpretation of gene fusion discovery. First, we have established a tier structure that incorporates the level of data support from RNA-seq and WGS data. This in turn provides more confidence beyond the total number of reads that support a particular candidate. For instance, a gene fusion with encompassing and spanning WGS and RNA-seq reads would be considered a reliable candidate compared to a candidate with only RNA-seq encompassing reads. Second, INTEGRATE is able to provide candidate gene fusion junctions at single-base resolution and the exact genomic breakpoints if spanning WGS reads were detected. Sequences and locations of the involved exons of the fusion junctions are provided to facilitate subsequent functional analysis and experimental validation. This also facilitates downstream analysis, such as finding complex gene fusions involving more than two genes, as exemplified by the seven instances observed in the cohort of 62 TCGA breast cancers (Supplemental Fig. 7). Furthermore, INTEGRATE is capable of detecting multiple alternative splicing isoforms for a fusion gene. For instance, of the seven previously discovered gene fusions in HCC1395, three of these gene fusions were captured by INTEGRATE with multiple isoforms (four isoforms for EIF3K-CYP39A1, three isoforms for RAB7A-LRCH3, and two isoforms for HNRNPUL2-AHNAK). This is important to ensure that an isoform producing a potentially in-frame novel protein is not overlooked.
Prior to this study, only a small number of gene fusions had been experimentally validated in the HCC1395. However, following our comprehensive analysis and validation, we have confirmed a large quantity of gene fusions. It is likely due to a number of factors. First, this is the most comprehensive analysis of this cell line conducted to date. Second, INTEGRATE was able to detect gene fusions missed by multiple programs suggesting that INTEGRATE is more sensitive and earlier analyses underrepresented the total number of gene fusions in a given sample. Third, a large portion of the HCC1395 cell line genome harbors copy number variation (Supplemental Fig. 8). As breast cancer gene fusions have been associated with copy number variation (Kalyana-Sundaram et al. 2012), it is not surprising that 106 (77%) of the 138 validated gene fusions have one or both genes residing in an amplicon. In total, 143 (59%) out of 276 genes involved in the 138 validated gene fusions reside in the amplicons, which is significantly higher (Pearson's χ2 test, P = 2.35 × 10−6) than the percentage of expressed genes residing in amplicons (FPKM ≥ 0.1; 6871 [38%] out of 18,235).
The application of INTEGRATE to a breast cancer cohort enabled the identification of novel gene fusions, a subset of which were recurrent. Interestingly, among the recurrent fusion candidates, ESR1 translocations (Figs. 4, 5) have been reported to be involved in hormone therapy resistance, exemplifying the potential biological significance of these candidates (Li et al. 2013). ESR1-CCDC170 was recently reported to markedly increase cell motility and anchorage-independent growth, reduced endocrine sensitivity, and enhanced xenograft tumor formation (Veeraraghavan et al. 2014). The mechanistic studies suggest that CCDC170 engages the GAB1 signalosome to potentiate growth factor signaling and enhance cell motility (Veeraraghavan et al. 2014). Additionally, STAT3-PTRF has been reported to be present in uterus/cervix cancer (Ojesina et al. 2014). Furthermore, many of the functionally recurrent gene fusions found by INTEGRATE involve genes related to previously reported fusions in breast cancer. Overall, given the high accuracy of INTEGRATE in detecting gene fusions, the locations of the nominated fusions are in hotspots of breast cancer, and many of the gene fusion partners have been previously implicated in breast cancer, it is possible that some of the remaining INTEGRATE recurrent and functionally recurrent gene fusion nominations may also be relevant to breast cancer progression.
Overall, based on our comparison of nine tools, INTEGRATE provides a significant advance, balancing sensitivity and specificity for improved gene fusion discovery. Various factors such as artifacts, low data coverage, mapping errors, repetitive regions, and low expression levels hinder the ability of using only WGS SV prediction or RNA-seq fusion detection strategies. Therefore, INTEGRATE takes advantage of both strategies due to the increasing availability of both whole genome and transcriptome sequencing data from the same patient to provide a highly accurate method for gene fusion discovery to unveil novel causative mutations.
Methods
INTEGRATE fusion calling using both RNA-seq and WGS data
Supplemental Figure 9 provides a detailed overview of INTEGRATE. The first step of INTEGRATE is to systematically store all of the encompassing reads aligned by a RNA-seq reads mapping tool in a graph where the nodes correspond to genes and the edges connect two genes involved in a putative fusion. At this point, the graph can be very dense, and INTEGRATE uses a series of filtering steps to remove false-positive gene fusion candidates according to the concordant suboptimal alignments and repetitiveness of the paired-end reads in the graph (Supplemental Methods). Next, INTEGRATE leverages the fusion gene graph that has been built to conduct a targeted split-read alignment to map either spanning reads or suboptimal concordant reads systematically in a single step instead of using two independent procedures (Supplemental Methods). If provided with WGS data sets, INTEGRATE attempts to identify SVs supporting the fusion candidates by alignment in focal regions (Supplemental Fig. 10; Supplemental Methods).
Due to the large quantity of encompassing reads that are realigned, coupled with unmapped reads that are evaluated as spanning reads, we implemented a fast split-read mapping algorithm. A BWT is created for each gene node (including exons and introns) (Supplemental Fig. 1) so that the prefix trie of each gene node can be simulated. A dynamic programming algorithm is designed to perform local alignment between a split-read and the prefix trie using a breadth first search that extends on the prefix trie according to the number of differences (mismatches and indels) of the best matches (highest scores in the dynamic programming). Pseudo code of the algorithm is given in Supplemental Figure 11, and details of the algorithm are explained in Supplemental Methods. The fast split-read mapping algorithm for aligning and realigning RNA-seq reads and the fast split-read mapping algorithm for aligning WGS reads in focal regions enable INTEGRATE to perform efficiently in run time (Supplemental Fig. 12). INTEGRATE also has a moderate space usage (Supplemental Fig. 12).
Validation of gene fusions in HCC1395 cell line
To enrich for the highest quality gene fusion predictions in our tool comparison, gene fusions with canonical exonic boundaries called by INTEGRATE and eight additional fusion calling methods (Comrad, nFuse, BreakTrans, TopHat-Fusion, ChimeraScan, FusionCatcher, pyPRADA, and TRUP) are considered as gene fusion candidates after additional filtering to remove false-positive candidates (i.e., gene fusions involving transcript variants such as HLA-A>>HLA-C, read-throughs, genes with overlapping isoforms) (Supplemental Fig. 3). For each gene fusion candidate, two sequencing probes were designed near the fusion junction. One probe was designed in the 5′ gene and the second probe was designed in the 3′ gene. For gene fusions with multiple fusion junctions corresponding to the alternative splicing of each gene fusion partner, we chose probes corresponding to the isoform with the greatest read support. All probes for the 240 gene fusion candidates are summarized in Supplemental Table 8 with their corresponding genes. Following cDNA hybridization, as previously described (Cabanski et al. 2014), ∼10 million 2 × 250 paired-end reads were generated using MiSeq. These reads were aligned using BWA to the predicted gene fusion junctions. The number of targeted validation reads spanning fusion junctions, with a conservative threshold of requiring the smaller flanking region to be longer than 30 nt, are summarized in Supplemental Table 9.
Application of INTEGRATE on TCGA human patient cohort
We downloaded whole-genome and RNA-seq data of 62 TCGA breast cancer patients. On average, INTEGRATE uses 8 h to process data for one patient (minimum was 1 h and maximum was 78 h). Average memory is 30 GB with a minimum of 24 GB and a maximum of 49 GB. When a gene fusion was predicted as a genomic event (intra- or inter-chromosomal) and a read-through event in multiple patients, then only if the percentage of patients with an intra- or inter-chromosomal gene fusion was >80% was it reported as a genomic fusion. For the less recurrent chimeras that could be explained as read-throughs or genomic events (due to false-positive encompassing WGS reads, medium-size deletions, or lack of WGS reads), three steps are performed before classifying them as genomic or read-through events. First, the two genes are >1 Mb apart. Second, for chimeras with genes closer than 1 Mb, the size of the deletions must be longer than 5000 nt. Third, if the fusion junction involves the second to last exon of the 5′ gene fusing into the second exon of the 3′ gene, characteristic of splicing that occurs in read-through transcripts, then we consider them as read-throughs.
Data access
The sequence data from this study have been submitted to the NCBI BioProject database (http://www.ncbi.nlm.nih.gov/bioproject/) under accession number PRJNA201238. The INTEGRATE software can be downloaded from SourceForge (https://sourceforge.net/projects/integrate-fusion/), and the source code is available in the Supplemental Material.
Supplementary Material
Acknowledgments
We thank T. Abbott, J. Walker, and D. Larson for their help in improving the code quality. We also thank C. Cabanski, N. Rockweiler, and A. Ramu for their feedback on the website. This work was supported by a National Institute of Health (NIH) Pathway to Independence Award (grant R00 CA149182; Principal Investigator: C.A.M.), R21 CA185983-01 (Principal Investigator: C.A.M.), Prostate Cancer Research Foundation Young Investigator Award (Principal Investigator: C.A.M.), and by the National Human Genome Research Institute (grant U54 HG003079; Principal Investigator: R.K.W.).
Footnotes
[Supplemental material is available for this article.]
Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.186114.114.
References
- Asmann YW, Hossain A, Necela BM, Middha S, Kalari KR, Sun Z, Chai HS, Williamson DW, Radisky D, Schroth GP, et al. 2011. A novel bioinformatics pipeline for identification and characterization of fusion transcripts in breast cancer and normal cell lines. Nucleic Acids Res 39: e100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asmann YW, Necela BM, Kalari KR, Hossain A, Baker TR, Carr JM, Davis C, Getz JE, Hostetter G, Li X, et al. 2012. Detection of redundant fusion transcripts as biomarkers or disease-specific therapeutic targets in breast cancer. Cancer Res 72: 1921–1928. [DOI] [PubMed] [Google Scholar]
- Cabanski CR, Magrini V, Griffith M, Griffith OL, McGrath S, Zhang J, Walker J, Ly A, Demeter R, Fulton RS, et al. 2014. cDNA hybrid capture improves transcriptome analysis on low-input and archived samples. J Mol Diagn 16: 440–451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Cancer Genome Atlas Network. 2012. Comprehensive molecular portraits of human breast tumours. Nature 490: 61–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carrara M, Beccuti M, Lazzarato F, Cavallo F, Cordero F, Donatelli S, Calogero RA. 2013. State-of-the-art fusion-finder algorithms sensitivity and specificity. BioMed Res Int 2013: 340620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, McGrath SD, Wendl MC, Zhang Q, Locke DP, et al. 2009. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods 6: 677–681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, Wallis JW, Kandoth C, Kalicki-Veizer JM, Mungall KL, Mungall AJ, Jones SJ, Marra MA, Ley TJ, Mardis ER, et al. 2012. BreakFusion: targeted assembly-based identification of gene fusions in whole transcriptome paired-end sequencing data. Bioinformatics 28: 1923–1924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen K, Navin NE, Wang Y, Schmidt HK, Wallis JW, Niu B, Fan X, Zhao H, McLellan MD, Hoadley KA, et al. 2013. BreakTrans: uncovering the genomic architecture of gene fusions. Genome Biol 14: R87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. 2013. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29: 15–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Druker BJ, Guilhot F, O'Brien SG, Gathmann I, Kantarjian H, Gattermann N, Deininger MW, Silver RT, Goldman JM, Stone RM, et al. 2006. Five-year follow-up of patients receiving imatinib for chronic myeloid leukemia. N Engl J Med 355: 2408–2417. [DOI] [PubMed] [Google Scholar]
- Edgren H, Murumagi A, Kangaspeska S, Nicorici D, Hongisto V, Kleivi K, Rye IH, Nyberg S, Wolf M, Borresen-Dale AL, et al. 2011. Identification of fusion genes in breast cancer by paired-end RNA-sequencing. Genome Biol 12: R6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernandez-Cuesta L, Sun R, Menon R, George J, Lorenz S, Meza-Zepeda LA, Peifer M, Plenker D, Heuckmann JM, Leenders F, et al. 2015. Identification of novel fusion genes in lung cancer using breakpoint assembly of transcriptome sequencing data. Genome Biol 16: 7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hormozdiari F, Hajirasouliha I, Dao P, Hach F, Yorukoglu D, Alkan C, Eichler EE, Sahinalp SC. 2010. Next-generation VariationHunter: combinatorial algorithms for transposon insertion discovery. Bioinformatics 26: i350–i357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iyer MK, Chinnaiyan AM, Maher CA. 2011. ChimeraScan: a tool for identifying chimeric transcription in sequencing data. Bioinformatics 27: 2903–2904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang Y, Wang Y, Brudno M. 2012. PRISM: pair-read informed split-read mapping for base-pair level detection of insertion, deletion and structural variants. Bioinformatics 28: 2576–2583. [DOI] [PubMed] [Google Scholar]
- Kalyana-Sundaram S, Shankar S, Deroo S, Iyer MK, Palanisamy N, Chinnaiyan AM, Kumar-Sinha C. 2012. Gene fusions associated with recurrent amplicons represent a class of passenger aberrations in breast cancer. Neoplasia 14: 702–708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Salzberg SL. 2011. TopHat-Fusion: an algorithm for discovery of novel fusion transcripts. Genome Biol 12: R72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. 2013. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14: R36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambros MBK, Wilkerson PM, Natrajan R, Patani N, Pawar V, Vatcheva R, Mansour M, Laschet M, Oelze B, Orr N, et al. 2011. High-throughput detection of fusion genes in cancer using the Sequenom MassARRAY platform. Lab Invest 91: 1491–1501. [DOI] [PubMed] [Google Scholar]
- Li H, Durbin R. 2009. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25: 1754–1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S, Shen D, Shao J, Crowder R, Liu W, Prat A, He X, Liu S, Hoog J, Lu C, et al. 2013. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep 4: 1116–1130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maher CA, Kumar-Sinha C, Cao X, Kalyana-Sundaram S, Han B, Jing X, Sam L, Barrette T, Palanisamy N, Chinnaiyan AM. 2009. Transcriptome sequencing to detect gene fusions in cancer. Nature 458: 97–101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mardis ER, Ding L, Dooling DJ, Larson DE, McLellan MD, Chen K, Koboldt DC, Fulton RS, Delehaunty KD, McGrath SD, et al. 2009. Recurring mutations found by sequencing an acute myeloid leukemia genome. New Engl J Med 361: 1058–1066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McPherson A, Hormozdiari F, Zayed A, Giuliany R, Ha G, Sun MG, Griffith M, Heravi Moussavi A, Senz J, Melnyk N, et al. 2011a. deFuse: an algorithm for gene fusion discovery in tumor RNA-Seq data. PLoS Comput Biol 7: e1001138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McPherson A, Wu C, Hajirasouliha I, Hormozdiari F, Hach F, Lapuk A, Volik S, Shah S, Collins C, Sahinalp SC. 2011b. Comrad: detection of expressed rearrangements by integrated analysis of RNA-Seq and low coverage genome sequence data. Bioinformatics 27: 1481–1488. [DOI] [PubMed] [Google Scholar]
- McPherson A, Wu C, Wyatt AW, Shah S, Collins C, Sahinalp SC. 2012. nFuse: discovery of complex genomic rearrangements in cancer using high-throughput sequencing. Genome Res 22: 2250–2261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Natrajan R, Wilkerson PM, Marchio C, Piscuoglio S, Ng CK, Wai P, Lambros MB, Samartzis EP, Dedes KJ, Frankum J, et al. 2014. Characterization of the genomic features and expressed fusion genes in micropapillary carcinomas of the breast. J Pathol 232: 553–565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicorici D, Satalan M, Edgren H, Kangaspeska S, Murumagi A, Kallioniemi O, Virtanen S, Kilkku O. 2014. FusionCatcher - a tool for finding somatic fusion genes in paired-end RNA-sequencing data. bioRxiv doi: http://dx.doi.org/10.1101/011650. [Google Scholar]
- Ojesina AI, Lichtenstein L, Freeman SS, Pedamallu CS, Imaz-Rosshandler I, Pugh TJ, Cherniack AD, Ambrogio L, Cibulskis K, Bertelsen B, et al. 2014. Landscape of genomic alterations in cervical carcinomas. Nature 506: 371–375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papas TS, Fisher RJ, Bhat N, Fujiwara S, Watson DK, Lautenberger J, Seth A, Chen ZQ, Burdett L, Pribyl L, et al. 1989. The ets family of genes: molecular biology and functional implications. Curr Top Microbiol Immunol 149: 143–147. [DOI] [PubMed] [Google Scholar]
- Robinson DR, Kalyana-Sundaram S, Wu YM, Shankar S, Cao X, Ateeq B, Asangani IA, Iyer M, Maher CA, Grasso CS, et al. 2011. Functionally recurrent rearrangements of the MAST kinase and Notch gene families in breast cancer. Nat Med 17: 1646–1651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakarya O, Breu H, Radovich M, Chen Y, Wang YN, Barbacioru C, Utiramerur S, Whitley PP, Brockman JP, Vatta P, et al. 2012. RNA-Seq mapping and detection of gene fusions with a suffix array algorithm. PLoS Comput Biol 8: e1002464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stephens PJ, McBride DJ, Lin ML, Varela I, Pleasance ED, Simpson JT, Stebbings LA, Leroy C, Edkins S, Mudie LJ, et al. 2009. Complex landscapes of somatic rearrangement in human breast cancer genomes. Nature 462: 1005–1010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stratton MR, Wooster RW, Futreal PA. 2004. The BRAF gene is frequently mutated in malignant melanoma. J Drugs Dermatol 3: 573–575. [PubMed] [Google Scholar]
- Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S, Hatano S, Asaka R, Hamanaka W, Ninomiya H, Uehara H, et al. 2012. RET, ROS1 and ALK fusions in lung cancer. Nat Med 18: 378–381. [DOI] [PubMed] [Google Scholar]
- Tomlins SA, Laxman B, Dhanasekaran SM, Helgeson BE, Cao X, Morris DS, Menon A, Jing X, Cao Q, Han B, et al. 2007. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448: 595–599. [DOI] [PubMed] [Google Scholar]
- Torres-Garcia W, Zheng S, Sivachenko A, Vegesna R, Wang Q, Yao R, Berger MF, Weinstein JN, Getz G, Verhaak RG. 2014. PRADA: pipeline for RNA sequencing data analysis. Bioinformatics 30: 2224–2226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varley KE, Gertz J, Roberts BS, Davis NS, Bowling KM, Kirby MK, Nesmith AS, Oliver PG, Grizzle WE, Forero A, et al. 2014. Recurrent read-through fusion transcripts in breast cancer. Breast Cancer Res Treat 146: 287–297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veeraraghavan J, Tan Y, Cao XX, Kim JA, Wang X, Chamness GC, Maiti SN, Cooper LJ, Edwards DP, Contreras A, et al. 2014. Recurrent ESR1–CCDC170 rearrangements in an aggressive subset of oestrogen receptor-positive breast cancers. Nat Commun 5: 4577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Mullighan CG, Easton J, Roberts S, Heatley SL, Ma J, Rusch MC, Chen K, Harris CC, Ding L, et al. 2011. CREST maps somatic structural variation in cancer genomes with base-pair resolution. Nat Methods 8: 652–654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wen H, Li Y, Malek SN, Kim YC, Xu J, Chen P, Xiao F, Huang X, Zhou X, Xuan Z, et al. 2012. New fusion transcripts identified in normal karyotype acute myeloid leukemia. PLoS One 7: e51203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu TD, Nacu S. 2010. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26: 873–881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Y, Gong M, Yuan H, Park HG, Frierson HF, Li H. 2012. Chimeric transcript generated by cis-splicing of adjacent genes regulates prostate cancer cell proliferation. Cancer Discov 2: 598–607. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.