Abstract
In the enteric nervous system (ENS), glia outnumber neurons by 4-fold and form an extensive network throughout the gastrointestinal tract. Growing evidence for the essential role of enteric glia in bowel function makes it imperative to understand better their molecular marker expression and how they relate to glia in the rest of the nervous system. We analyzed expression of markers of astrocytes and oligodendrocytes in the ENS and found, unexpectedly, that proteolipid protein 1 (PLP1) is specifically expressed by glia in adult mouse intestine. PLP1 and S100β are the markers most widely expressed by enteric glia, while glial fibrillary acidic protein (GFAP) expression is more restricted. Marker expression in addition to cellular location and morphology distinguishes a specific subpopulation of intramuscular enteric glia, suggesting that a combinatorial code of molecular markers can be used to identify distinct subtypes. To assess the similarity between enteric and extra-enteric glia, we performed RNA sequencing analysis (RNA-Seq) on PLP1-expressing cells in the mouse intestine and compared their gene expression pattern to that of other types of glia. This analysis shows that enteric glia are transcriptionally unique and distinct from other cell types in the nervous system. Enteric glia express many genes characteristic of the myelinating glia, Schwann cells and oligodendrocytes, although there is no evidence of myelination in the murine ENS.
Keywords: Enteric Nervous System, Neuroglia, RNA-Seq, Gastrointestinal Tract
Introduction
Glial cells play critical roles in the development, function and plasticity of the nervous system. The diverse roles of glia are matched by a diversity of glial subtypes with distinct locations, functions, morphologies and molecular markers. The ENS is a major autonomic division that can independently regulate gastrointestinal functions including motility, secretion, mucosal maintenance, and immunity (Furness 2006). It is organized into two interconnected networks of ganglia, the myenteric and submucosal plexuses, where enteric neuronal perikarya are located. Enteric glia outnumber neurons by several fold, and are more widely distributed throughout the intestine (Gulbransen and Sharkey 2012). Enteric glia are known to be essential for normal gastrointestinal function (Aube et al. 2006; Bush et al. 1998; Savidge et al. 2007); yet, the degree of their similarity to glia in the rest of the nervous system remains unclear.
Enteric glia were initially considered to be the Schwann cells of the gut because they too are neural crest derivatives. As electron microscopy studies began to reveal differences between Schwann cells in peripheral nerves and glia in the bowel, this classification began to change. Gabella described the ultrastructure of glia in the myenteric plexus as a small, central perikaryon from which numerous processes extended, and was the first to suggest that this stellate morphology as well as their relationship to neurons was more similar to that of astrocytes than to Schwann cells (Gabella 1971). Jessen and Mirsky later showed that the astrocytic marker, GFAP, was expressed within enteric ganglia (Jessen and Mirsky 1980), fueling the idea, which has long prevailed, that glia in the ENS are analogous to CNS astrocytes (Gershon and Rothman 1991; Jessen and Mirsky 2005). To date, GFAP and S100β remain the most commonly used markers to identify enteric glia. More recent work has added the transcription factor, Sox10, as another marker of glia in the mouse intestine (Hoff et al. 2008; Laranjeira et al. 2011; Young et al. 2003).
Although molecular markers have often been used interchangeably, growing evidence of morphologic differences among enteric glia suggests underlying diversity. Hanani and Reichenbach (1994) first identified distinct morphological types of glia within the guinea pig myenteric plexus and proposed a classification scheme based on their findings. Gulbransen and Sharkey (2012) later suggested expanding this morphologic classification, or alternatively, grouping enteric glia based on their location along the serosa-to-lumen axis. Morphological analysis of Sox10-expressing cells using the mosaic analysis with double markers (MADM) system has since confirmed the existence of at least 4 distinct glial morphologies within the muscular layers of the mouse intestine (Boesmans et al. 2014). This work also showed that GFAP, Sox10, and S100β are differentially expressed by cells in the myenteric plexus (Boesmans et al. 2014). Differences in dye filling, calcium transients and receptor expression, even among glia in a single myenteric ganglion, further hint that enteric glia are functionally diverse (Boesmans et al. 2014; Maudlej and Hanani 1992; Nasser et al. 2006b). With a growing number of studies implicating enteric glia in digestive and neurologic disorders (Clairembault et al. 2014; Neunlist et al. 2013), it is becoming essential to understand better the normal distribution and diversity of glial cells in the ENS.
To date, much of the work investigating enteric glial diversity has relied on preparations of longitudinal muscle with adherent myenteric plexus (LMMP), in which the mucosa, submucosa, and circular muscle are discarded (Boesmans et al. 2014; Hanani and Reichenbach 1994). LMMP preparations thus exclude many enteric glia, including mucosal glia, which populate one of the most unique microenvironments in the nervous system. To ascertain more fully enteric glial distribution, diversity, and similarity to CNS and PNS glial subtypes, we analyzed expression of glial markers throughout the length and breadth of the mouse intestine and identified a new molecular marker of enteric glia, PLP1. We then isolated PLP1-expressing cells from the small and large intestine, performed transcriptional profiling and compared enteric glial gene expression to that of extra-intestinal glia. Our data show that enteric glia are a distinct, heterogeneous, and unique class of mammalian glial cells. This work provides new insights into enteric glia and identifies novel molecular tools to manipulate these cells in vivo, facilitating further analysis of glial function in the bowel.
Methods
Mouse lines
PLP1-eGFP hemizygous mice (Mallon et al., 2002), PLPCreERT mice (Doerflinger et al., 2003) and Rosa26-lox-stop-lox-TdTomato (LSLTdTomato) mice have been previously described (Madisen et al., 2010). Male and female mice were used in all experiments, and FVBN mice were utilized for wildtypes. For morphological studies, mice hemizygous for PLPCreERT and heterozygous for LSLTdTomato were administered 0.1μg tamoxifen by oral gavage and analyzed 7 days later. Mice were handled and housed in accordance with the IACUC guidelines of Boston Children’s Hospital and Columbia University Medical Center.
Immunocytochemistry
For immunostaining cryosections, intestinal segments were fixed in 4% PFA/PBS at 4°C for 2h, equilibrated in 30% sucrose/PBS overnight at 4°C, and embedded in OCT. 14μm sections were obtained, incubated for 1h in blocking solution (PBS + 0.1% Triton + 1% heat-inactivated goat serum [HINGS]) and then overnight at 4°C in primary antibody diluted in blocking solution. Slides were washed 3 times with PBS, incubated for 1h at room temperature (RT) in secondary antibody diluted 1:500, washed 3 times with PBS, and then mounted in Vectashield containing DAPI (Vector Labs H-1200). For whole mount immunostaining, intestinal segments were dissected, trimmed into 1.5cm cylinders, opened along the mesenteric border, pinned flat onto Sylgard plates, and fixed in 4% PFA/PBS for 75 minutes at 4°C. Samples were rinsed twice in PBS, and processed in a modified version of the protocol described by Li et al. (2011). Samples were unpinned, washed 6 × 20min with PBST (PBS + 0.5% Triton), incubated in primary antibody diluted in blocking solution (PBST + 20% DMSO + 5% HINGS) at RT for 48–72 hours, washed 6 × 20min with PBST, incubated in secondary antibody at RT for 24 hours, washed 6 × 20min with PBST, taken through a graded methanol series, equilibrated with 1:1 methanol:BABB solution [BABB = 1 part benzyl alcohol to 2 parts benzyl benzoate] for 2 hours, and then incubated overnight in BABB for optical clearing. PLP1-eGFP signal in Figures 1, 2 and 4 represents endogenous GFP fluorescence. Primary antibodies used: Rat anti-PLP1 1:500 (gift from W. Macklin), Rabbit anti-S100β 1:500 (DAKO Z0311), Rabbit anti-GFAP 1:1000 (Sigma G9269), Chicken anti-GFAP 1:1000 (Millipore AB5541), Goat anti-Sox10 1:50 (Santa Cruz sc-17342), Chicken anti-GFP 1:1000 (Aves GFP-1020), Rabbit anti-GFP 1:1000 (Invitrogen A11122), and Rabbit anti-DsRed 1:500 (Clontech 632496). Secondary antibodies used were Alexa-Fluor 594, 568, or 488 conjugates generated in either Goat or Donkey (Invitrogen).
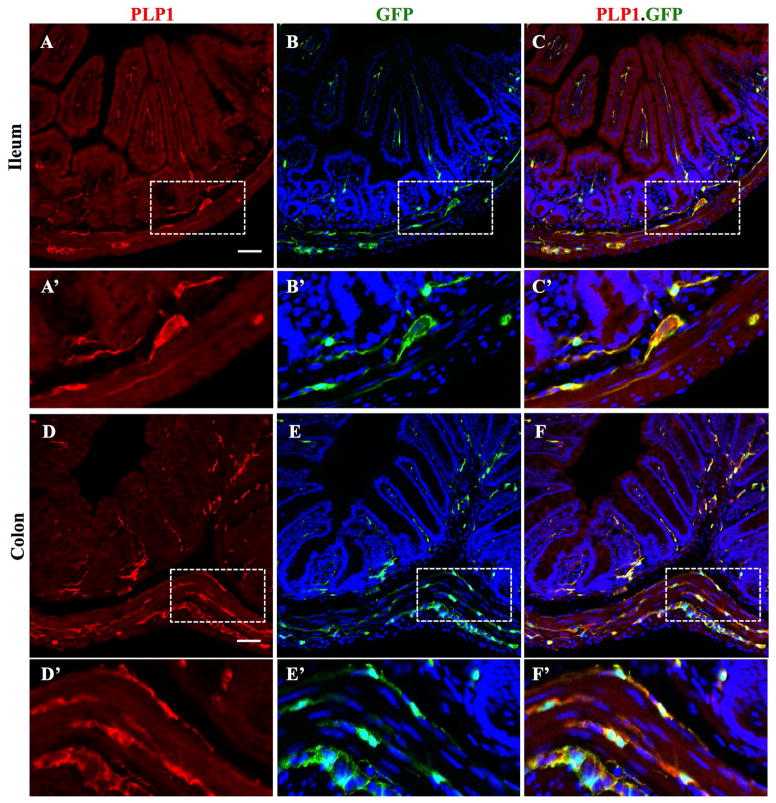
Figure 1. PLP1 is expressed in the mouse intestine and colocalizes with eGFP expressed under the control of the PLP1 promoter.
(A, D) Immunostaining for PLP1 protein in intestinal cross-sections from PLP1-eGFP transgenic mouse at P28.
(B, E) eGFP fluorescence with DAPI nuclear stain in blue.
(C, F) Merged images. Boxed areas are displayed at higher magnification below respective images.
Scale bar represents 50μm (panels A–F) or 20μm (panels A′–F′).
Figure 2. PLP1 and S100β are co-expressed in the adult mouse intestine.
(A, D, G) eGFP fluorescence in PLP1-eGFP transgenic mouse intestine at P56, with DAPI nuclear stain in blue.
(B, E, H) S100β immunostaining
(C, F, I) Merged images. Boxed areas displayed at higher magnification below respective images show that most PLP1-expressing glia co-express S100β, but there are rare GFP+ cells that are S100β− (*). Scale bar represents 50μm (panels A–I) or 14μm (panels G′–I′).
Figure 4. GFAP is expressed in a subset of PLP1-expressing cells.
(A, D, G) eGFP fluorescence in the PLP1-eGFP transgenic mouse at P56, with DAPI nuclear stain in blue.
(B, E, H) GFAP immunostaining
(C, F, I) Merged images. Boxed areas displayed at higher magnification below respective images show that many extraganglionic PLP1-expressing glia in the circular muscle exhibit low (*) or no detectable (arrowhead) GFAP immunoreactivity. Scale bar represents 50μm (panels A–I) or 14μm (panels G′–I′).
Imaging and quantification
Single planar images of cryosections and whole mounts were obtained on either LSM710 or Nikon A1R confocal microscopes for all figures except supplemental figures 1 & 2 (voxel size 0.31–0.50μm × 0.31–0.50μm × 1μm, resolution 2–3 pixels/μm). Images in supplemental figures 1 & 2 represent maximum intensity projections of 16μm stacks (voxel size 0.31μm × 0.31μm × 2μm, resolution 3.2 pixels/μm). Decomposition into individual channels was done using ImageJ software (Schneider et al. 2012). Quantification of Plp1+, S100β+, and GFAP+ cells in cryosections was performed using CellProfiler (Carpenter et al. 2006) followed by manual analysis. Images of 10 fields per intestinal segment (duodenum, ileum, and colon) were obtained from each of 3 PLP1-eGFP mice immunostained for S100β or GFAP (minimum of 1200 cells analyzed per each marker in each intestinal segment). All thresholds were calculated using the Otsu Global method with two-class thresholding, using intensity to distinguish objects with the following criteria: DAPI+ nuclei (threshold correction factor 4, diameter 6–40 pixel units), S100β+ cells (threshold correction factor 10, diameter 14–100 pixel units), Plp1+ cells (threshold correction factor 7, diameter 14–100 pixel units), and GFAP+ cells (threshold correction factor 14, diameter 14–100 pixel units). For each image, the myenteric ganglia, the muscular layer (longitudinal and circular muscle), and non-muscular layers were then manually outlined using nuclear staining for reference, and the CellProfiler-identified cells were further screened to eliminate those outside the tissue area (i.e. in intestinal lumen) or without an associated cell nucleus. The number of cells that were Plp1+/GFAP+, Plp1+/S100β+, only Plp1+, only S100β+ and only GFAP+ in each layer were then quantified, and the percentages of cells in each category were calculated per animal. Percentages for S100β and GFAP were then compared for each intestinal segment by unpaired t-test. Sox10-PLP1 colocalization was quantified using the ImageJ Cell Counter macro to analyze 300μm × 300μm regions of interest in 6 images per intestinal segment (n= 4 animals).
Isolation of enteric glia and RNA Sequencing
4cm segments of ileum and colon were dissected from PLP1-eGFP mice between P20 and P27, cut into 1mm fragments, and then incubated in digestion medium (1X HEPES, Collagenase A 1mg/mL, Dispase II 2.9mg/ml) for 1h at 37°C. After trituration, samples were spun down at 600g for 8min, and incubated in 0.25% Trypsin-EDTA at 37°C for 5min. FBS was added, cells were spun down at 600g for 8min, resuspended in HBSS, triturated with a series of flame polished pipets, and then filtered through a 70μm cell strainer. Cells were spun down again and resuspended in HBSS with propidium iodide (1μg/ml). FACS was performed on a BD Aria Flow Sorter and cells were collected in Trizol. Gates used for collection are illustrated on representative FACS plots in Supplemental Figure 2. Cells were homogenized in Trizol, lysates from 3 animals were pooled per each of 4 samples (Ileum GFP+ and −, Colon GFP+ and −), and then total RNA was purified using the Qiagen RNEasy Micro kit. Bioanalyzer PicoChip measurement of RNA quality showed RIN > 8.0. Total RNA (10ng) was then reverse transcribed for library construction, using the Total RNA-Seq IntegenX-Low Input kit. One microgram of DNA from each sample was sheared to generate the library and then sequenced by Illumina HiSeq 2000 to obtain 100bp paired-end reads.
Mapping and Analysis of RNA Sequencing Data
Approximately 80–100 million 100bp reads were obtained for each of the 4 samples, and mapped to the mouse genome using Bowtie and Tophat (Trapnell et al., 2012). The mitochondrial ribosome and nuclear ribosome accounted for 52–79% of all reads. Unmapped reads represented 4–9%, and were composed primarily of long polyA and polyT stretches that likely correspond to 3′ mRNA. Of the remaining reads, ~60% uniquely mapped within annotated genes. Assuming an mRNA abundance of 5 × 105 total mRNAs per cell (Galau et al., 1977), this sequencing depth corresponds to an average of 10–20 mapped fragments for an mRNA expressed at 1 copy per cell. To prevent bias due to mitochondrial overrepresentation, we followed a two-stage mapping strategy. Reads were first mapped to a “contamination library” containing the mitochondrial chromosome and nuclear ribosome sequences using Bowtie2 (Langmead and Salzberg, 2012). If either read in a pair mapped to the contamination library, both reads were excluded from further analysis. This “trimmed” library (containing 18–40 million reads, 20–50% of the total) was then aligned to the M. musculus genome (UCSC version mm10) using TopHat2 (Trapnell et al., 2012). Reads were aligned to annotated transcripts before mapping to the remainder of the genome (option -G). Mapped reads were next assembled into transcripts with CuffLinks (Trapnell et al., 2012) using the annotated mouse transcriptome to guide assembly. Transcript abundance estimates were corrected using the upper-quartile normalization, fragment bias correction, and multi-read correction options in CuffLinks. Finally, differential expression was tested with Cuffdiff (Trapnell et al. 2012), treating GFP+ and GFP− samples from ileum and colon as biological replicates. Loci with <10 alignments were excluded from hypothesis testing.
Quantitative PCR (qPCR)
For qPCR validation, RNA was obtained from PLP1-eGFP animals as described above (each set generated from FACS-sorted cells from 1 animal for n=3 biological replicates), reverse transcribed with iScript (BioRad), and then used as template for qPCR with SYBR Green Select (Applied Biosystems) on a 7500 Fast Real-Time PCR System (Applied Biosystems). The standard 7500 Fast Protocol followed by the thermal denaturing step was used. All reactions were performed in duplicate with GAPDH run in parallel. Data were analyzed with 7500 Fast System SDS software version 2.0.6 (Applied Biosystems) and the thresholds of each gene were standardized between experiments. All primers utilized had efficiency ≥ 97% on positive control cDNA and resulting PCR products were confirmed by sequencing. The effect of tissues (colon vs. ileum) was not significant for the 5 genes tested (p = 0.7, two way ANOVA excluding samples where no signal was detected), and therefore results were pooled for comparison of GFP+ to GFP−. P values for differences in expression between GFP+ and GFP− samples were estimated with the Mann-Whitney U test, and corrected for multiple hypotheses using Holm’s method.
Global Comparisons of Gene Expression
To compare gene expression between enteric glia and other cell types (Supplemental Table 1), every gene was first ranked according to its differential expression in each cell type. For cell types in Cahoy, et al. (2008) and Zhang, et al. (2014), differential expression was calculated according to the methods used in these papers. For Schwann cells, we used a large meta-analysis of mouse microarrays (Zheng-Bradley et al., 2010) that contained data from purified Schwann cells (Buchstaller et al., 2004) and data from many other studies, to serve as negative controls. The expression of each gene in the Buchstaller P0 Schwann cell dataset was compared to the same gene’s expression in every other sample, using a one-sided Mann-Whitney U test, and genes were then ranked according to their p value. Cell type enriched genes were also predicted using the “gene-driven” cell type differential expression algorithm, CellMapper (Nelms, et al. in review). CellMapper searches for genes that have a similar expression profile to an established set of cell type-specific markers, ranking every gene based on its tendency to be expressed at a similar level to the marker genes in a given dataset. For CellMapper searches, microarray data was gathered from two large meta-analyses of human gene expression (Engreitz et al., 2010; Lukk et al., 2010). These meta-analyses were divided into “intestine” and “non-intestine” samples based on the presence or absence of the terms “colon”, “intestin*”, “gut”, “enteric”, “mucosa”, and “bowel” within the sample titles or descriptions using the R package GEOmetadb (Zhu et al., 2008). Enteric glial genes were predicted by searching the “intestine” sample subset for genes with similar expression to PLP1. Schwann cell genes were predicted by searching the “non-intestine” sample subset for genes with similar expression to MPZ and P0. Finally, genes for CNS cell types were predicted by searching the Allen Brain Atlas human microarray dataset (Hawrylycz et al., 2012) for genes with similar expression to L1CAM (Neurons), ALDH1L1 (Astrocytes), or MOG (Oligodendrocytes). After ranking genes based on their differential expression in each cell type, the rank lists were compared. A similarity score (Yang et al., 2006) was calculated using the R package OrderedList (Lottaz et al., 2006). This score assesses how often the same genes appear at the top of two ordered genes lists, assigning greater weight to genes ranked at the top of both lists than genes in the middle. A free parameter, alpha, controls how quickly the weights fall off with gene rank; alpha was set to 0.01 for all calculations (results were similar for alpha between 0.005 – 0.1). To assess the variance explained by cell type combination, each similarity score was assigned to a group depending on the set of cell types being compared. Then the within-group variance and between group variance were calculated (η2), and the statistical significance of the variance explained was estimated using a permutation test in which the cell type assigned to each individual rank list was scrambled and then the variance explained was recalculated. Similarity of enteric glia to CNS cell types was also examined by looking for enrichment of cell type markers from Cahoy et al. (2008), using the top 50 ranked genes for astrocytes, neurons, and oligodendrocytes from Cahoy et al. (2008) as “established markers” for these cell types. Then the position of these markers within the ranked lists for enteric glia were plotted graphically, or an unweighted (p = 0) GSEA enrichment statistic was calculated (Subramanian et al. 2005).
Results
PLP1 is widely expressed by enteric glia
While surveying expression of glial markers in the GI tract, we unexpectedly discovered that PLP1 is widely expressed in the adult mouse small and large intestine (Figure 1A, D). To determine the identity of PLP1-expressing cells, we examined the co-localization of PLP1 with established markers of enteric glia and other intestinal cell types in tissues from a PLP1-eGFP transgenic mouse line (Mallon et al. 2002). In this line, PLP1-expressing cells in the CNS are marked by expression of green fluorescent protein (GFP) (Mallon et al. 2002). We found that GFP expression mirrored PLP1 immunoreactivity in the intestine of PLP1-eGFP mice (Figure 1B, E), and co-localized with S100β throughout the bowel (Figure 2). PLP1 did not co-localize with the neuronal marker, ANNA-1, the smooth muscle marker, alpha smooth muscle actin, or the pericyte marker NG2 (data not shown), suggesting that enteric PLP1 expression is limited to glia.
To assess the extent of overlap in expression between PLP1 and known enteric glial markers, we immunostained S100β and GFAP in serial sections of proximal small intestine (duodenum), distal small intestine (ileum), and large intestine (colon) from adult PLP1-eGFP animals. S100β and PLP1 were largely co-expressed throughout the intestine (Figure 2). S100β was expressed by 84.5 ± 2.7% of GFP+ cells in the duodenum, 83 ± 5.2% in the ileum, and 76 ± 2.4% in the colon (mean ± S.E.M.; Figure 3). In contrast, GFAP co-expression with PLP1 was much more restricted (35–54%; Figure 3 & 4). GFP+/GFAP− cells could readily be identified throughout the intestine, and were most numerous in the colon (Figure 4). In the colon, where the largest difference was observed, 65% of PLP1-expressing cells did not express GFAP, highlighting the observation that GFAP is not a marker for all enteric glia (Boesmans et al. 2014; Jessen and Mirsky 1983).
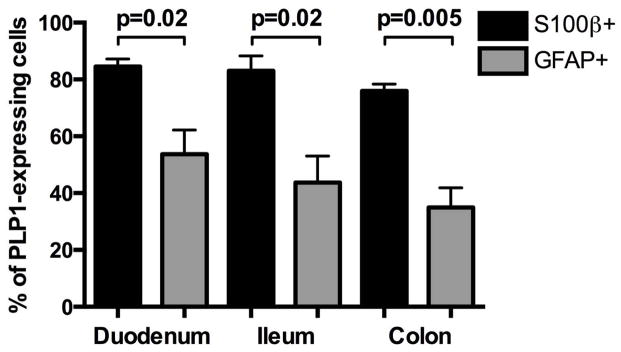
Figure 3. S100β is expressed by the majority of PLP1-expressing cells while GFAP expression is more limited.
Serial intestinal cross-sections from PLP1-eGFP mice at P56 were immunostained for either S100β or GFAP. The number of cells expressing eGFP and/or either S100β or GFAP was then counted from 10 cross-sections per intestinal segment per animal (n=3 mice; mean ± s.e.m; p values obtained by unpaired t-test).
The ENS is organized into two sets of ganglia: myenteric ganglia, located in between the longitudinal and circular muscle layers, and a smaller set of submucosal ganglia located closer to the luminal epithelium. Enteric neuronal cell bodies are restricted to these ganglia, while enteric glia are not. Given the widespread distribution of glia within the laminar organization of the bowel, one proposed classification of enteric glia is based on their location along the radial (serosa-to-lumen) axis (Gulbransen and Sharkey 2012). To determine whether marker expression varies based on glial position along this axis, we quantified expression of PLP1 and GFAP based on segregation of glia into 2 locations: myenteric (glia within myenteric ganglia and extraganglionic glia dispersed in the muscular layers), or extra-myenteric (submucosal and mucosal glia). We elected to group submucosal and mucosal glia because these two subtypes could not be reliably distinguished in our automated quantitation approach. Patterns of marker expression were similar in myenteric glia from proximal to distal intestine (Figure 5). Extra-myenteric glia, in contrast, exhibited a larger fraction of PLP1+/GFAP− cells as well as greater differences in marker expression from one intestinal segment to another, consistent with the markedly different luminal and epithelial environments in the various parts of the intestine (Figure 5). Mucosal glia within the lamina propria of villi, for example, were typically PLP1+/GFAP+ in the duodenum but variably expressed GFAP in the ileum (Figure 4, Supplemental Figure 1). Many colonic mucosal glia were also GFAP− (Supplemental Figure 2). To determine if intraganglionic glia exhibit different marker expression from extraganglionic glia, we focused on myenteric glia, where we could easily assess the localization of cells. Intraganglionic myenteric glia exhibited similar patterns of marker expression throughout the intestine, with the majority expressing both PLP1 and GFAP (Figure 5). In contrast, extraganglionic intramuscular glia were overwhelmingly GFAP negative in all 3 intestinal locations examined (Figure 5). These observations suggest that glial expression of molecular markers varies along both the proximal-distal and radial axes of the intestine, and that a specific subset of enteric glia located in the muscular layers outside of ganglia expresses PLP1 but not GFAP.
Figure 5. PLP1 and GFAP expression vary based on glial position along the proximal-distal and radial axes of the digestive tract.
Analysis of PLP1 and GFAP expression in PLP-eGFP mice at P56 shows that “myenteric” glia (those within myenteric ganglia and extraganglionic glia distributed within the muscular layers) exhibit similar patterns of marker expression along the length of the intestine with the majority expressing GFAP; in contrast, greater than 50% of “extra-myenteric” glia (mucosal and submucosal glia) express PLP1 but not GFAP. Segregating myenteric glia into intraganglionic and extraganglionic compartments reveals that the majority of extraganglionic intramuscular glia also express PLP1 but not GFAP. All cells expressing one or both markers were counted from 10 cross-sections per intestinal segment from each of 3 PLP1-eGFP mice at P56 (mean ± s.e.m). Kruskal-Wallis test was used to compare the distribution of medians across the 6 groups for each intestinal segment for all enteric glia (p=0.025, 0.022, and 0.0097 for duodenum, ileum and colon, respectively), and for myenteric glia either within or outside ganglia (p=0.01, 0.04, and 0.0096 for duodenum, ileum and colon, respectively). “Total glia” refers to sum of all cells expressing PLP1 only, GFAP only, or both markers.
Since GFAP is an intermediate filament protein, we considered the possibility that PLP1+/GFAP− glia outside of myenteric ganglia on cross-sections might reflect undetected co-localization of molecular markers with different subcellular distributions. We thus used whole mount immunostaining to compare PLP1, GFAP, S100β, and Sox10 expression in myenteric intraganglionic and intramuscular glia of the small and large intestine. S100β expression overlapped with PLP1, consistent with quantitation of sections (Figure 6A–C). Similarly, 99.5% of Sox10+ myenteric and intramuscular glia in the duodenum, ileum and colon expressed PLP1 (p<0.0001), supporting the idea that PLP1, S100β and Sox10 are co-expressed by most enteric glia. Within myenteric ganglia, most of the PLP1+/S100β+ cells were also GFAP+ (Figure 6D–F). There was variation in levels of GFAP expression within myenteric ganglia, consistent with previous reports (Boesmans et al. 2014; Jessen and Mirsky 1983). Interestingly, whole mount imaging confirmed that the majority of PLP1+ glia interspersed in the muscular layers outside of myenteric ganglia did not express GFAP (Figure 6D–F). These GFAP-negative extraganglionic glia therefore represent a distinct subset of enteric glia that can be clearly distinguished by molecular marker expression (PLP1+, S100β+, Sox10+, GFAP−) and location (extraganglionic, intramuscular).
Figure 6. Extraganglionic intramuscular glia with Type IV morphology express PLP1 and S100β, but not GFAP.
(A–C) S100β immunostaining and PLP1-eGFP fluorescence in mouse colon at P56 shows high degree of S100β and PLP1 colocalization in intramuscular and intraganglionic myenteric glia.
(D–F) GFAP immunostaining and PLP1-eGFP fluorescence in mouse colon at P56 shows that PLP1 and GFAP expression overlap within myenteric ganglia, but extraganglionic intramuscular glia express PLP1 but not GFAP.
(G–I) Immunostaining for GFAP in colonic segments from PLPCreERT:: LSLTdTomato mice treated with low dose tamoxifen to induce sparse labeling of PLP1-expressing cells. TdTomato expression is detected in enteric glia with Type I (arrowheads), Type II (+), Type III (arrow), and Type IV (*) morphology. See Supplemental Figure 3 for higher magnification images of morphological types.
Scale bar = 50um. All images acquired at a single z plane at the level of the myenteric plexus from whole mount immunostained colonic segments.
Previous work exploring the morphological heterogeneity of enteric glia suggests that there are at least 4 types: star-shaped “protoplasmic gliocytes” within myenteric ganglia (Type I), elongated “fibrous gliocytes” within fiber tracts (Type II), subepithelial glia with long processes (Type III) and elongated intramuscular gliocytes (Type IV) (Hanani & Reichenbach 1994; Gulbransen and Sharkey, 2012). Recently, Boesmans et al., (2014) showed that extraganglionic, intramuscular glia with 4 primary processes could also be distinguished and proposed that they be grouped within Type III. To investigate whether the PLP1 promoter is active in all these morphological subtypes, we crossed PLPCreERT transgenic mice with Rosa26-lox-stop-lox-TdTomato (LSLTdTomato) animals to generate mice in which enteric glia can be induced to express the fluorescent reporter TdTomato. By inducing reporter expression with low doses of tamoxifen, we sparsely labeled enteric glia for morphological analysis and found that that TdTomato+ cells exhibited all 4 reported morphologies (Figure 6G–I; Supplemental Figure 3). TdTomato-expressing glia types I–III within and adjacent to the myenteric plexus exhibited varying degrees of GFAP co-localization. In contrast, none of the examined TdTomato+ Type IV intramuscular glia expressed GFAP (Figure 6G–I). Taken together, our data show that PLP1 is among the most widely expressed markers of enteric glia and that genetic tools employing the PLP1 promoter to isolate enteric glia can be used to gain a better understanding of these cells.
Transcriptional Profiling of Enteric Glia
Enteric glia are neural crest-derived; however, the nature of their relationship with enteric neurons as well as their location in an environment filled with an abundance of cell types, such as fibrocytes, macrophages, smooth muscle, lymphocytes, enterocytes, and enteroendocrine cells makes them unique among glia in the mammalian PNS and CNS (Bohorquez et al. 2014; Liu et al. 2013). To gain insight into the functions of glia in the gastrointestinal tract, we determined the transcriptional profile of PLP1+ cells by RNA-Seq. We focused on acutely isolated GFP+ glial cells from ileum and colon of PLP1-eGFP mice between the ages of P21–P28. Since the gastrointestinal epithelium undergoes major changes around the time of weaning (Henning 1981), we reasoned that enteric glia at this age might upregulate genes that are important for their functions in the mature ENS. Ileum and colon were chosen because they are physically proximate segments of the GI tract with different functions, microbiota, and epithelial surfaces. A protocol was developed to make single cell suspensions from segments of mouse intestine and optimized to achieve at least 75% cell survival by trypan blue exclusion. GFP+ enteric glial cells were then isolated by Fluorescence Activated Cell Sorting (FACS; Supplemental Figure 4), and represented 0.8% of live cells in the ileum and 2% of live cells in the colon. RT-PCR revealed that S100β expression was limited to GFP+ cells, suggesting that enteric glial cells were successfully depleted from the GFP− sample (Figure 7A). In contrast, villin1, a gene uniquely expressed by epithelial cells in the intestine (el Marjou et al. 2004), was detected in both samples (Figure 7A). Since this suggested that epithelial cells were not completely eliminated from the GFP+ preparation, we analyzed gene expression in both GFP+ and GFP− cells in order to identify transcripts enriched in GFP+ enteric glia. Total RNA obtained from GFP+ and GFP− cells from ileum and colon of PLP1-eGFP animals was sequenced, and differential expression of transcripts was assessed. GFP+ samples from colon and ileum were treated as biological replicates and compared to GFP− samples, which represents a conservative strategy that should highlight genes selectively expressed in enteric glia across the intestine.
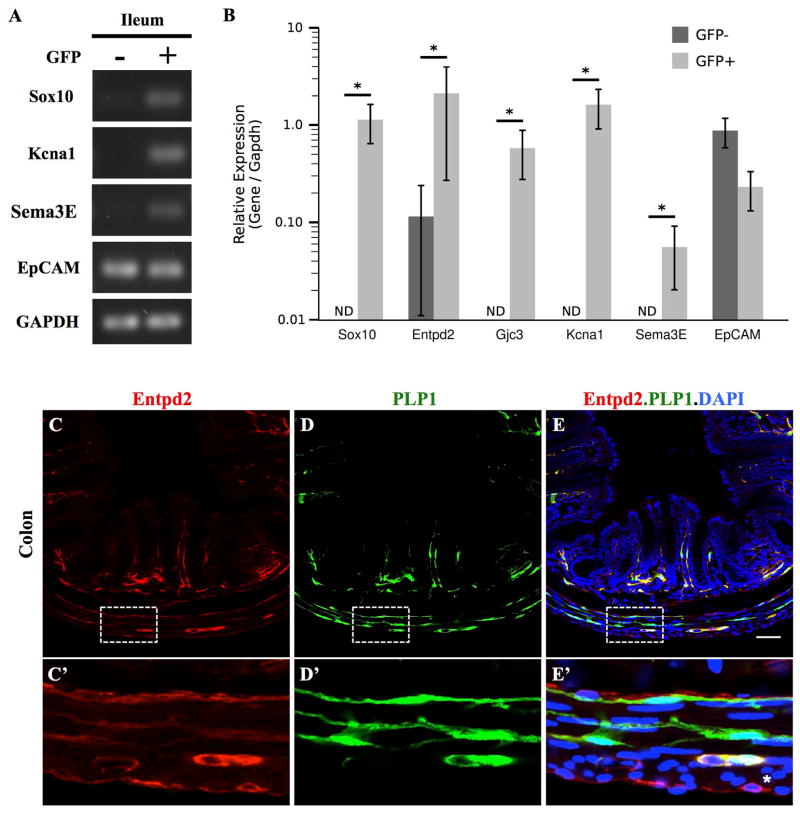
Figure 7. RNA from PLP1+ cells isolated from postnatal intestine is enriched for enteric glial marker genes.
(A) RT-PCR of RNA extracted from FACS-sorted cells utilized for RNA Seq analysis shows enrichment for S100β mRNA but not Villin1 in the GFP+ fraction.
(B) Relative difference in expression of cell-type specific genes in RNA-Seq data from GFP+ versus GFP− samples. To avoid numerical instability (dividing by 0) in cases where a gene was very weakly expressed, a pseudocount of 0.01 was added to all measured FPKM prior to calculating fold enrichment. This will, in general, decrease the estimated fold change, providing a more conservative value.
To validate our RNA-Seq results, we checked the measured expression of established marker genes for several intestinal cell types. The enteric glial genes S100β, Gfap, Plp1, and Sox10 were strongly enriched in GFP+ samples, while markers of epithelial cells, smooth muscle, and endothelia were all depleted (Figure 7B). Despite the suspected presence of some epithelial cells in our GFP+ samples, our comparative analysis strategy successfully differentiated glial genes from epithelial genes. A subset of neuronal genes that are not expressed by enteric glia were somewhat enriched in the GFP+ sample (Figure 7B), suggesting that this sample contained some neurons. Regardless, glial genes were enriched to levels at least 20-fold higher than that of neuronal genes in the GFP+ sample; therefore, the presence of some neurons is unlikely to confound further analysis, which focuses on the most differentially expressed genes.
We identified 292 differentially expressed genes between GFP+ and GFP− samples at a false discovery rate (FDR) cutoff of 0.1. Table 1 lists the top 25 genes enriched in PLP1+ cells in the intestine, which includes several genes previously reported to be important for enteric glial development, such as Sox10 and Foxd3 (Laranjeira et al. 2011; Mundell et al. 2012). To validate our RNA-Seq results further, we selected 5 genes for additional analysis. Enteric glia are known to express 2 of these genes, Sox10 and Entpd2 (Braun et al. 2004; Lavoie et al. 2011), while 3 genes had not previously been tested (Gjc3, Sema3E and Kcna1). Quantitative RT-PCR (qPCR) on 3 biological replicates of GFP+ and GFP− cells showed that 4 of the 5 genes were undetectable in GFP− cells and enriched in GFP+ cells, validating both our isolation technique and RNA-Seq results (Figure 8A–B). The only one of these 5 genes detectable in GFP− cells, Entpd2, had 27-fold greater expression in GFP+ cells (Figure 8B). Consistent with this, Entpd2 immunoreactivity was predominantly found in glial cells in the mouse intestine, but was also observed in a small population of cells adjacent to crypts and along the serosa that were not glia (Figure 8C–E). Notably, Entpd2 was expressed by many but not all PLP1+ glia. Similar to GFAP, Entpd2 was not detected in many extraganglionic intramuscular glia (Figure 8C–E). In summary, these findings suggest that RNA-Seq profiling of enteric PLP1+ cells identified genes expressed by all enteric glia as well as some expressed only by subsets, and support the use of this dataset as a validated resource of gene expression in enteric glia.
Table 1.
Top 25 genes enriched in PLP1+ enteric glia.
| Gene | GFP− (FPKM) | GFP+ (FPKM) | Log 2 Fold Change | FDR |
|---|---|---|---|---|
| Kcna1 | 0.60 | 116.25 | 7.59 | 0.0072 |
| Gjc3 | 0.09 | 22.56 | 8.01 | 0.0118 |
| Gpr37l1 | 0.16 | 40.98 | 8.02 | 0.0118 |
| Plp1 | 2.46 | 479.20 | 7.61 | 0.0118 |
| Cdh19 | 0.34 | 42.12 | 6.97 | 0.0118 |
| Grik3 | 0.06 | 27.04 | 8.79 | 0.0148 |
| Kcna6 | 0.20 | 34.02 | 7.38 | 0.0159 |
| Fign | 0.14 | 15.56 | 6.83 | 0.0159 |
| Tacr3 | 0.05 | 10.98 | 7.79 | 0.0200 |
| Nell2 | 0.17 | 19.39 | 6.87 | 0.0200 |
| Col20a1 | 0.31 | 74.31 | 7.91 | 0.0200 |
| Sostdc1 | 1.60 | 198.28 | 6.96 | 0.0200 |
| Slc35f1 | 0.49 | 39.29 | 6.34 | 0.0207 |
| Foxd3 | 0.06 | 22.09 | 8.51 | 0.0207 |
| Sema3e | 0.20 | 33.07 | 7.36 | 0.0216 |
| Kcna2 | 0.73 | 55.70 | 6.24 | 0.0284 |
| Col28a1 | 0.16 | 13.11 | 6.39 | 0.0284 |
| Sox10 | 0.02 | 42.14 | 11.00 | 0.0362 |
| Ptprz1 | 0.93 | 32.43 | 5.12 | 0.0474 |
| Wdr86 | 0.07 | 30.86 | 8.69 | 0.0476 |
| Zfp451 | 1.91 | 57.93 | 4.92 | 0.0498 |
| Megf10 | 0.05 | 4.58 | 6.64 | 0.0522 |
| Lrrc4c | 0.03 | 5.95 | 7.83 | 0.0553 |
| Gfra2 | 2.01 | 56.40 | 4.81 | 0.0643 |
| Cadm3 | 1.47 | 41.12 | 4.81 | 0.0665 |
Fragments per kilobase of transcript per million mapped reads (FPKM) for each gene in GFP− and GFP+ RNA-Seq samples from PLP1-eGFP mouse intestine. FDR = False discovery rate.
Figure 8. RNA-Seq identifies genes enriched in enteric glia.
(A–B) Quantitative RT-PCR validation of genes enriched in enteric glia according to RNA-Seq. (A) Representative agarose gel electrophoresis of RT-PCR (B) Bar plots of qPCR data show median +/− 1.48 times the median absolute deviation of each gene’s expression level relative to Gapdh (n = 6); * = p < 0.05, Mann-Whitney U test).
(C) Entpd2 immunostaining of colon from PLP1-eGFP mouse at P21. Boxed areas displayed at higher magnification immediately below respective images show that intraganglionic PLP1-expressing glia in the muscular layer exhibit strong Entpd2 immunoreactivity, while extraganglionic glia are dim (* indicates a non-glial cell expressing Entpd2). Scale bar represents 50μm (panels C–E) or 11μm (panels C′–E′).
Enteric glia are transcriptionally distinct from other types of glia
Enteric glia share the same developmental origins as Schwann cells, but have traditionally been considered to be analogous to astrocytes based on their morphology and expression of GFAP. To explore the transcriptional similarity between enteric glia and other types of glia, we compared our data to previous studies of gene expression in murine astrocytes, neurons, oligodendrocytes, microglia and Schwann cells (Supplemental Table 1). In a second complementary approach, we used an in silico cell type-expression deconvolution algorithm, called CellMapper (Nelms et al., in review), to predict genes expressed in enteric glia and other neural cell types using human microarray data. The CellMapper algorithm employs a “gene-driven” strategy, in which established cell-type specific markers can be used to query large databases to identify genes that share a similar expression profile. In total, these two approaches provide independent gene expression measurements for each cell type, allowing us to determine which cell type-similarities are robust across methods.
There are several technical challenges when comparing gene expression between studies: different cell isolation protocols, RNA purification methods, and expression technologies, all leading to “lab effects” that can substantially alter an overall expression profile (Zilliox and Irizarry 2007). To overcome these uncertainties, we designed a strategy to distinguish true biological differences between cell types while mitigating technical variability. First, we compared genes upregulated in a given cell type (relative expression) rather than raw expression levels. Second, we assessed similarity of gene expression between each experiment by calculating a “similarity score” (Yang et al. 2006) (Figure 9A). This score emphasizes genes that are most differentially expressed in an experiment over genes that do not change substantially and contribute to noise. To test this approach, we calculated similarity scores between every pair of studies and estimated how much of the variability in these scores could be explained by the combination of cell types being compared (e.g. Neuron-Neuron, Neuron-Enteric Glia). Cell type combination explained ~80% of the variance in similarity scores (η2 = 83%, p < 10−4 by permutation test), indicating that the similarity score strategy can highlight true biological differences between cell types. Similarity scores between every combination of samples are visualized as a heatmap in Figure 9A. This plot shows the consistency across methods aimed at measuring expression for the same cell type. For example, at a global level, gene expression in astrocytes is highly similar whether measured by RNA-Seq, microarray or CellMapper. Previously known similarities and differences between cell types are also evident. For instance, oligodendrocytes, astrocytes and neurons have very little gene expression similarity with each other, and consequently similarity scores between these cell types are very low. Schwann cells show strong similarity with oligodendrocytes, the other myelinating cell type, but only weak similarity to astrocytes and no similarity to neurons. Enteric glia display the greatest similarity to Schwann cells, and then to oligodendrocytes. They also exhibit some similarity to astrocytes, but it is limited and no greater than the similarity between Schwann cells and astrocytes. Enteric glia share no significant similarity with the mesoderm-derived microglia, pericytes or endothelia. Of note, enteric glia exhibit significant similarity scores with neurons, unlike any of the other types of glia. It is unclear if this similarity is due to neuronal contamination of the enteric glial data set, or shared expression of some biological pathways. The consistency of this observation across the RNA-Seq and CellMapper data supports the possibility of common pathways.
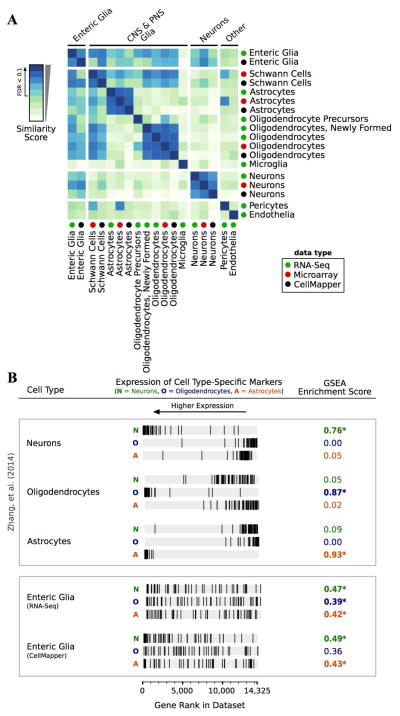
Figure 9. Global comparisons of enteric glial transcriptional profiles to that of other PNS and CNS cell types.
(A) Heatmap illustrating global similarity scores between gene expression in glial and non-glial cell types. See Supplemental Table 1 for source data for each cell type. False discovery rate (FDR) < 0.1 annotated on color scale.
(B) Gene set enrichment analysis (GSEA) showing the rank of markers from neurons (N), oligodendrocytes (O), and astrocytes (A) within the transcriptomes of CNS cell types (top box) versus enteric glia (bottom box).
The global comparisons of transcriptional similarity suggest that enteric glia express many of the same genes as several different types of glia. To investigate this suggestion further, we examined our RNA-Seq dataset for expression of a defined set of established markers of astrocytes, oligodendrocytes, and neurons. We selected the 50 top-ranked marker genes for each of these cell types from a previous microarray study (Cahoy et al. 2008), and asked whether these genes were enriched in enteric glia by gene set enrichment analysis (GSEA) (Subramanian et al. 2005). To validate this approach, we first assessed the rank of these markers within RNA-Seq data sets for each of the CNS cell types (Zhang et al. 2014). As illustrated in Figure 9B, neuronal genes were enriched only in neurons, astrocytic genes in astrocytes, and oligodendrocyte genes in oligodendrocytes, confirming that the marker genes chosen were consistent across studies and represented a reproducible set of cell type-specific markers within the CNS.
When we carried out GSEA using these markers in the enteric glia data set, the results were striking. Markers for all 3 CNS cell types were enriched, with some markers being strongly expressed in enteric glia (ranking among the top enriched enteric glial genes), while other markers were undetected (Figure 9B). This result was the same whether we analyzed the enteric glial expression profile predicted by CellMapper or the RNA-Seq data, showing that this finding was consistent across methods (Figure 9B). Certain subsets of markers of each CNS cell type were strongly enriched in our GFP+ sample. For instance, a subset of genes important in myelinating glia including Sox10, Plp1, Mbp, and Mpz, were all strongly enriched while others, such as Mog, Mobp, and Mag were not detected (Supplemental Figure 5). Astrocytic genes such as Gfap, Entpd2, and Dio2 were all highly enriched in enteric glia while other widely used markers of astrocytes, such as Aldh1l1 and the glutamate transporter Slc1a3 (GLAST), were not expressed (Supplemental Figure 5). Taken together, the GSEA and global analyses of transcriptional similarity show that enteric glia are a unique class of glia, without direct analogy to any other type of glial cell examined.
Discussion
Molecular markers of enteric glia
Enteric glia have previously been related to other types of glia, such as Schwann cells and astrocytes, based on the expression of single molecular markers or cellular morphology. We analyzed the distribution and co-localization of the most commonly used glial markers along both the longitudinal and radial axes of the murine intestinal tract, to investigate the diversity and distribution of glial cells in the ENS. We found that most enteric glia co-express S100β, PLP1 and Sox10, and that these 3 genes represent the most widely expressed markers for glia in the ENS. In contrast, only subsets of enteric glia in each intestinal segment express GFAP. In the colon, for example, GFAP immunoreactivity was absent from many extraganglionic glia in the muscle layers, and present at variable levels of intensity even within myenteric ganglia. Our observations, together with work by others that focused on GFAP expression in myenteric plexus (Boesmans et al. 2014; Jessen and Mirsky 1983), show that GFAP expression is absent from many enteric glia, and therefore does not serve as a universal marker for these cells.
In astrocytes, high GFAP levels are a marker of “reactivity” and cellular activation (Silver and Miller 2004). GFAP expression in enteric glia may be similarly dynamic. Upon lineage tracing GFAP-expressing cells within myenteric ganglia, Boesmans et. al (2014) found that 23% of intraganglionic myenteric glia that expressed GFAP at one time point did not express it 7 days later. This observation suggests that GFAP expression may be dynamic within myenteric ganglia, and is consistent with the variable intensity of GFAP immunoreactivity we detected. Our data show that a specific subset of glia in the mature ENS, extraganglionic intramuscular glia with Type IV morphology, is consistently GFAP-negative. Future lineage tracing studies will be needed to determine whether these and other GFAP-negative enteric glia, such as those in the mucosa and submucosa, arise from GFAP-expressing cells, and if so, when they downregulate GFAP expression.
Precise characterization of molecular marker expression and definition of subsets is essential to interpreting the growing literature on enteric glia. For example, a number of groups have analyzed enteric glial function by targeted ablation in vivo. These studies have implicated enteric glia in a variety of epithelial processes in addition to neuronal support (Aube et al. 2006; Bush et al. 1998; Savidge et al. 2007). To date, virtually all of these studies targeted GFAP-expressing cells. In contrast, one study using a non-selective gliotoxin to ablate enteric glia showed changes in small intestinal motility without epithelial dysfunction (Nasser et al. 2006a). Our data show that large subsets of enteric glia, such as extraganglionic, intramuscular enteric glia, do not express GFAP. The conflicting findings in previous ablation studies may reflect the underlying heterogeneity of enteric glia, and suggests that the functional roles of some enteric glial subsets likely remain undiscovered.
“Myelin gene” expression in non-myelinating glia
One observation in our data is that PLP1 is widely expressed by enteric glia. Taking advantage of this expression, we utilized PLP1-eGFP transgenic mice to isolate PLP1-expressing cells from the murine intestine and carry out RNA-seq. The resulting transcriptional profile revealed that enteric glia express, not only PLP1, but also many other genes that are important for formation of compact and non-compact myelin, including myelin basic protein (MBP), myelin protein zero (MPZ), 2′,3′-cyclic nucleotide 3′ phosphodiesterase (CNP) and Sox10. Over the past 50 years, numerous ultrastructural studies have closely examined the relationship between neurons and glia in the rodent ENS and found no evidence of myelination in the small or large intestine (Cook and Burnstock 1976; Gabella 1971; Gabella 1972; Wilson et al. 1981). It remains unclear, therefore, why enteric glia express these particular genes. The expression of myelin genes in mature, non-myelinating glia is not unprecedented in the nervous system. For example, the PLP1 promoter is also active in perisynaptic Schwann cells and a subset of supporting cells in the inner ear, neither of which form myelin (Duregotti et al. 2015; Gomez-Casati et al. 2010; Morris et al. 2006). The role that these molecules play outside the context of myelin, however, remains unclear.
We found that not only is the PLP1 promoter transcriptionally active in enteric glia, but that PLP1 protein is widely detectable in these cells (Figure 1). PLP1 is a profoundly hydrophobic integral membrane protein and, together with myelin basic protein (MBP), accounts for more than 50% of the protein in CNS myelin (Deber and Reynolds 1991). Similarly, myelin protein zero (MPZ; P0) and MBP together constitute up to 65% of the protein in PNS myelin (Garbay et al. 2000). Why are classic myelin proteins expressed in non-myelinating glia in the intestine? One possibility is that myelin proteins may play more than a structural role within glia. Studies of cultured oligodendrocytes show that PLP1 can alter expression of other myelin genes and inhibit differentiation of oligodendrocyte precursor cells (Karim et al. 2007; Miyamoto et al. 2012). Similarly, MBP is increasingly recognized as a multifunctional molecule that with a potential role as a scaffolding protein (Harauz and Boggs 2013). Although enteric glia do not form myelin, proteins such as PLP1 and MBP may be important for organizing signaling microdomains that bring specific molecules together. Proteomic studies assessing the interaction partners of these classic myelin proteins within enteric glia could provide new insights into the signaling and other capabilities of these proteins.
Enteric glia are heterogeneous and distinct from other glia
To explore the similarities between enteric glia and glia in other parts of the nervous system, we utilized two different approaches to compare genes enriched in enteric glia with those enriched in astrocytes, oligodendrocytes, Schwann cells, microglia, and neurons. Remarkably, these global comparisons of transcriptional profiles, as well as our studies of individual molecular marker expression, converged on the observation that enteric glia express genes and features of other glial types, but do not exhibit a transcriptional signature identical to any of them. This hybrid pattern of similarity likely reflects the unique nature of enteric glia. Our comparative analysis of transcriptional profiles was based on publicly available data sets, most of which were obtained by purifying cell types from early postnatal animals (Supplemental Table 1). The Schwann cell data, for example, was obtained at P0 when peripheral nerves are populated by many Schwann cell precursors that subsequently give rise to both myelinating and non-myelinating Schwann cells (Buchstaller et al. 2004). Future studies using data obtained from mature glial types in adult animals, when common developmental programs are extinguished, may be even more revealing. In summary, our comparative analysis shows that enteric glia are transcriptionally most similar to Schwann cells and oligodendrocytes, although they also express a subset of genes in common with astrocytes. Contrary to traditional ideas, enteric glia and Schwann cells exhibit similar degrees of transcriptional similarity to astrocytes, although the genes expressed in common may not necessarily be the same.
Another explanation for the hybrid pattern of transcriptional similarity observed is that it reflects the underlying diversity of enteric glial subtypes. It is possible that each enteric glial subtype bears a strong resemblance to a particular class of glia elsewhere in the nervous system, leading to a hybrid gene expression pattern when all enteric glia are assessed as a whole. We discovered that Type IV intramuscular glia as well as a subset of glia in the mucosa and submucosa do not express GFAP, but do express PLP1 and S100β. Further investigation of transcripts enriched in the RNA-Seq screen might reveal additional genes that are differentially expressed by subsets of enteric glia. For instance, our preliminary analysis of Entpd2 suggests that, like GFAP, it is not expressed by many extraganglionic intramuscular glia. Entpd2 and GFAP are both enriched in mature astrocytes and not detected in myelinating glia (Cahoy et al. 2008; Wink et al. 2006; Zhang et al. 2014); consequently, enteric glia that express these two proteins may share functions in common with astroglia.
Taken together, our studies of individual glial marker expression and RNA-Seq profiling show that the ENS is populated by a heterogeneous group of glia that are transcriptionally distinct from glia in other parts of the nervous system. Future studies, such as single cell transcriptional profiling, will be needed to assess more exhaustively the diversity of enteric glial subtypes and their relationship to other types of CNS and PNS glia. Enteric glia have increasingly been implicated in the pathophysiology of a wide range of neurologic and digestive disorders (Clairembault et al. 2014; Garcia et al. 2014). The finding that the PLP1 promoter is active in virtually all enteric glia opens the door to a new array of genetic tools for manipulating enteric glial function in vivo. These new tools will help to explore the roles of enteric glia in both the physiology and pathophysiology of the bowel.
Supplementary Material
Supplemental Figure 1. PLP1 is more commonly expressed by mucosal glia in the small intestine than GFAP. Maximum intensity projections of confocal stacks from PLP1-eGFP adult mouse duodenum (A–C) and ileum (D–I) immunostained for GFAP (A–F) or S100β (G–I). DAPI nuclear stain is represented in blue. Scale bar = 50μm.
Supplemental Figure 2. Many mucosal glia in the colon marked by PLP1 expression, do not express GFAP. Maximum intensity projections of confocal stacks from PLP1-eGFP adult mouse colon immunostained for GFAP. DAPI nuclear stain is represented in blue. Scale bar = 50μm.
Supplemental Figure 3. All 4 morphological types of enteric glia express PLP1.
Confocal single planar images of TdTomato fluorescence in colonic segments from PLPCreERT:: LSL-TdTomato mice treated with low dose tamoxifen.
Supplemental Figure 4. Isolation of PLP1+ enteric glia from postnatal mouse intestine.
FACS plots of propidium iodide (PI) fluorescence vs. GFP fluorescence generated from ileums of control (A) and PLP1-eGFP (B) littermate mice, with cross bars marking gates used to sort and collect GFP+ and GFP− cells.
Supplemental Figure 5. RNA from PLP1+ cells in the intestine is enriched for subsets of oligodendrocyte and astrocyte marker genes.
Relative difference in expression of cell-type specific genes in RNA-Seq data from GFP+ versus GFP− samples. To avoid numerical instability (dividing by 0), a pseudocount of 0.01 was added to all measured FPKM prior to calculating fold enrichment. ND = not detected.
Source data utilized for comparative analysis of glia Datasets used for comparative analysis strategy to compare the transcriptional profile of enteric glia to that of other cell types. CM = Cell Mapper.
Main Points.
Proteolipid protein 1 (PLP1) is expressed by enteric glia throughout the intestine
Transcriptional profiling of PLP1+ enteric glia reveals that they are most similar to myelinating glia, not astrocytes
Acknowledgments
This study was supported by the NIDDK (M.R., F32DK098903), NASPGHAN Young Investigator Award (M.R.), NINDS (G.C., R01 NS35884), Development Disability Research Center (DDRC) National Institutes of Health Grant (G.C., P30-HD18655), NSF Graduate Research Fellowship (B.D.N.), pilot grant from the Milton Fund (G.C.), and research support from the Divisions of Pediatric Gastroenterology, Hepatology and Nutrition at Boston Children’s Hospital and Columbia University Medical Center. We thank W. Macklin for PLP1 antibody; N. Francis & E. Lamousé-Smith for assistance with FACS analysis; A. Chalazonitis, and L. Waldron for critical reading of the manuscript, and the DDRC Imaging and FACS core facilities at Boston Children’s Hospital.
Footnotes
The authors declare no competing financial interests.
References
- Aube AC, Cabarrocas J, Bauer J, Philippe D, Aubert P, Doulay F, Liblau R, Galmiche JP, Neunlist M. Changes in enteric neurone phenotype and intestinal functions in a transgenic mouse model of enteric glia disruption. Gut. 2006;55:630–7. doi: 10.1136/gut.2005.067595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boesmans W, Lasrado R, Vanden Berghe P, Pachnis V. Heterogeneity and phenotypic plasticity of glial cells in the mammalian enteric nervous system. Glia. 2015;63:229–41. doi: 10.1002/glia.22746. [DOI] [PubMed] [Google Scholar]
- Bohorquez DV, Samsa LA, Roholt A, Medicetty S, Chandra R, Liddle RA. An enteroendocrine cell-enteric glia connection revealed by 3D electron microscopy. PLoS One. 2014;9:e89881. doi: 10.1371/journal.pone.0089881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braun N, Sevigny J, Robson SC, Hammer K, Hanani M, Zimmermann H. Association of the ecto-ATPase NTPDase2 with glial cells of the peripheral nervous system. Glia. 2004;45:124–32. doi: 10.1002/glia.10309. [DOI] [PubMed] [Google Scholar]
- Buchstaller J, Sommer L, Bodmer M, Hoffmann R, Suter U, Mantei N. Efficient isolation and gene expression profiling of small numbers of neural crest stem cells and developing Schwann cells. J Neurosci. 2004;24:2357–65. doi: 10.1523/JNEUROSCI.4083-03.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bush TG, Savidge TC, Freeman TC, Cox HJ, Campbell EA, Mucke L, Johnson MH, Sofroniew MV. Fulminant jejuno-ileitis following ablation of enteric glia in adult transgenic mice. Cell. 1998;93:189–201. doi: 10.1016/s0092-8674(00)81571-8. [DOI] [PubMed] [Google Scholar]
- Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA, et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci. 2008;28:264–78. doi: 10.1523/JNEUROSCI.4178-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carpenter AE, Jones TR, Lamprecht MR, Clarke C, Kang IH, Friman O, Guertin DA, Chang JH, Lindquist RA, Moffat J, et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 2006;7:R100. doi: 10.1186/gb-2006-7-10-r100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clairembault T, Kamphuis W, Leclair-Visonneau L, Rolli-Derkinderen M, Coron E, Neunlist M, Hol EM, Derkinderen P. Enteric GFAP expression and phosphorylation in Parkinson’s disease. J Neurochem. 2014;130:805–15. doi: 10.1111/jnc.12742. [DOI] [PubMed] [Google Scholar]
- Cook RD, Burnstock G. The ultrastructure of Auerbach’s plexus in the guinea-pig. II. Non-neuronal elements. J Neurocytol. 1976;5:195–206. doi: 10.1007/BF01181656. [DOI] [PubMed] [Google Scholar]
- Deber CM, Reynolds SJ. Central nervous system myelin: structure, function, and pathology. Clin Biochem. 1991;24:113–34. doi: 10.1016/0009-9120(91)90421-A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duregotti E, Negro S, Scorzeto M, Zornetta I, Dickinson BC, Chang CJ, Montecucco C, Rigoni M. Mitochondrial alarmins released by degenerating motor axon terminals activate perisynaptic Schwann cells. Proc Natl Acad Sci U S A. 2015;112:E497–505. doi: 10.1073/pnas.1417108112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- el Marjou F, Janssen KP, Chang BH, Li M, Hindie V, Chan L, Louvard D, Chambon P, Metzger D, Robine S. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis. 2004;39:186–93. doi: 10.1002/gene.20042. [DOI] [PubMed] [Google Scholar]
- Engreitz JM, Daigle BJ, Jr, Marshall JJ, Altman RB. Independent component analysis: mining microarray data for fundamental human gene expression modules. J Biomed Inform. 2010;43:932–44. doi: 10.1016/j.jbi.2010.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furness JB. The enteric nervous system. xiii. Malden, Mass: Blackwell Pub; 2006. p. 274. [Google Scholar]
- Gabella G. Glial cells in the myenteric plexus. Z Naturforsch B. 1971;26:244–5. doi: 10.1515/znb-1971-0313. [DOI] [PubMed] [Google Scholar]
- Gabella G. Fine structure of the myenteric plexus in the guinea-pig ileum. J Anat. 1972;111:69–97. [PMC free article] [PubMed] [Google Scholar]
- Garbay B, Heape AM, Sargueil F, Cassagne C. Myelin synthesis in the peripheral nervous system. Prog Neurobiol. 2000;61:267–304. doi: 10.1016/s0301-0082(99)00049-0. [DOI] [PubMed] [Google Scholar]
- Garcia SB, Stopper H, Kannen V. The contribution of neuronal-glial-endothelial-epithelial interactions to colon carcinogenesis. Cell Mol Life Sci. 2014;71:3191–7. doi: 10.1007/s00018-014-1642-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gershon MD, Rothman TP. Enteric glia. Glia. 1991;4:195–204. doi: 10.1002/glia.440040211. [DOI] [PubMed] [Google Scholar]
- Gomez-Casati ME, Murtie J, Taylor B, Corfas G. Cell-specific inducible gene recombination in postnatal inner ear supporting cells and glia. J Assoc Res Otolaryngol. 2010;11:19–26. doi: 10.1007/s10162-009-0191-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gulbransen BD, Sharkey KA. Novel functional roles for enteric glia in the gastrointestinal tract. Nat Rev Gastroenterol Hepatol. 2012;9:625–32. doi: 10.1038/nrgastro.2012.138. [DOI] [PubMed] [Google Scholar]
- Hanani M, Reichenbach A. Morphology of horseradish peroxidase (HRP)-injected glial cells in the myenteric plexus of the guinea-pig. Cell Tissue Res. 1994;278:153–60. doi: 10.1007/BF00305787. [DOI] [PubMed] [Google Scholar]
- Harauz G, Boggs JM. Myelin management by the 18.5-kDa and 21.5-kDa classic myelin basic protein isoforms. J Neurochem. 2013;125:334–61. doi: 10.1111/jnc.12195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawrylycz MJ, Lein ES, Guillozet-Bongaarts AL, Shen EH, Ng L, Miller JA, van de Lagemaat LN, Smith KA, Ebbert A, Riley ZL, et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature. 2012;489:391–9. doi: 10.1038/nature11405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henning SJ. Postnatal development: coordination of feeding, digestion, and metabolism. Am J Physiol. 1981;241:G199–214. doi: 10.1152/ajpgi.1981.241.3.G199. [DOI] [PubMed] [Google Scholar]
- Hoff S, Zeller F, von Weyhern CW, Wegner M, Schemann M, Michel K, Ruhl A. Quantitative assessment of glial cells in the human and guinea pig enteric nervous system with an anti-Sox8/9/10 antibody. J Comp Neurol. 2008;509:356–71. doi: 10.1002/cne.21769. [DOI] [PubMed] [Google Scholar]
- Jessen KR, Mirsky R. Glial cells in the enteric nervous system contain glial fibrillary acidic protein. Nature. 1980;286:736–7. doi: 10.1038/286736a0. [DOI] [PubMed] [Google Scholar]
- Jessen KR, Mirsky R. Astrocyte-like glia in the peripheral nervous system: an immunohistochemical study of enteric glia. J Neurosci. 1983;3:2206–18. doi: 10.1523/JNEUROSCI.03-11-02206.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jessen KR, Mirsky R. The origin and development of glial cells in peripheral nerves. Nat Rev Neurosci. 2005;6:671–82. doi: 10.1038/nrn1746. [DOI] [PubMed] [Google Scholar]
- Karim SA, Barrie JA, McCulloch MC, Montague P, Edgar JM, Kirkham D, Anderson TJ, Nave KA, Griffiths IR, McLaughlin M. PLP overexpression perturbs myelin protein composition and myelination in a mouse model of Pelizaeus-Merzbacher disease. Glia. 2007;55:341–51. doi: 10.1002/glia.20465. [DOI] [PubMed] [Google Scholar]
- Laranjeira C, Sandgren K, Kessaris N, Richardson W, Potocnik A, Vanden Berghe P, Pachnis V. Glial cells in the mouse enteric nervous system can undergo neurogenesis in response to injury. J Clin Invest. 2011;121:3412–24. doi: 10.1172/JCI58200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lavoie EG, Gulbransen BD, Martin-Satue M, Aliagas E, Sharkey KA, Sevigny J. Ectonucleotidases in the digestive system: focus on NTPDase3 localization. Am J Physiol Gastrointest Liver Physiol. 2011;300:G608–20. doi: 10.1152/ajpgi.00207.2010. [DOI] [PubMed] [Google Scholar]
- Liu YA, Chung YC, Pan ST, Shen MY, Hou YC, Peng SJ, Pasricha PJ, Tang SC. 3-D imaging, illustration, and quantitation of enteric glial network in transparent human colon mucosa. Neurogastroenterol Motil. 2013;25:e324–38. doi: 10.1111/nmo.12115. [DOI] [PubMed] [Google Scholar]
- Lukk M, Kapushesky M, Nikkila J, Parkinson H, Goncalves A, Huber W, Ukkonen E, Brazma A. A global map of human gene expression. Nat Biotechnol. 2010;28:322–4. doi: 10.1038/nbt0410-322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallon BS, Shick HE, Kidd GJ, Macklin WB. Proteolipid promoter activity distinguishes two populations of NG2-positive cells throughout neonatal cortical development. J Neurosci. 2002;22:876–85. doi: 10.1523/JNEUROSCI.22-03-00876.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maudlej N, Hanani M. Modulation of dye coupling among glial cells in the myenteric and submucosal plexuses of the guinea pig. Brain Res. 1992;578:94–8. doi: 10.1016/0006-8993(92)90234-z. [DOI] [PubMed] [Google Scholar]
- Miyamoto Y, Torii T, Tanoue A, Yamauchi J. Pelizaeus-Merzbacher disease-associated proteolipid protein 1 inhibits oligodendrocyte precursor cell differentiation via extracellular-signal regulated kinase signaling. Biochem Biophys Res Commun. 2012;424:262–8. doi: 10.1016/j.bbrc.2012.06.101. [DOI] [PubMed] [Google Scholar]
- Morris JK, Maklad A, Hansen LA, Feng F, Sorensen C, Lee KF, Macklin WB, Fritzsch B. A disorganized innervation of the inner ear persists in the absence of ErbB2. Brain Res. 2006;1091:186–99. doi: 10.1016/j.brainres.2006.02.090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mundell NA, Plank JL, LeGrone AW, Frist AY, Zhu L, Shin MK, Southard-Smith EM, Labosky PA. Enteric nervous system specific deletion of Foxd3 disrupts glial cell differentiation and activates compensatory enteric progenitors. Dev Biol. 2012;363:373–87. doi: 10.1016/j.ydbio.2012.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasser Y, Fernandez E, Keenan CM, Ho W, Oland LD, Tibbles LA, Schemann M, MacNaughton WK, Ruhl A, Sharkey KA. Role of enteric glia in intestinal physiology: effects of the gliotoxin fluorocitrate on motor and secretory function. Am J Physiol Gastrointest Liver Physiol. 2006a;291:G912–27. doi: 10.1152/ajpgi.00067.2006. [DOI] [PubMed] [Google Scholar]
- Nasser Y, Ho W, Sharkey KA. Distribution of adrenergic receptors in the enteric nervous system of the guinea pig, mouse, and rat. J Comp Neurol. 2006b;495:529–53. doi: 10.1002/cne.20898. [DOI] [PubMed] [Google Scholar]
- Neunlist M, Van Landeghem L, Mahe MM, Derkinderen P, des Varannes SB, Rolli-Derkinderen M. The digestive neuronal-glial-epithelial unit: a new actor in gut health and disease. Nat Rev Gastroenterol Hepatol. 2013;10:90–100. doi: 10.1038/nrgastro.2012.221. [DOI] [PubMed] [Google Scholar]
- Savidge TC, Newman P, Pothoulakis C, Ruhl A, Neunlist M, Bourreille A, Hurst R, Sofroniew MV. Enteric glia regulate intestinal barrier function and inflammation via release of S-nitrosoglutathione. Gastroenterology. 2007;132:1344–58. doi: 10.1053/j.gastro.2007.01.051. [DOI] [PubMed] [Google Scholar]
- Schneider CA, Rasband WS, Eliceiri KW. NIH Image to ImageJ: 25 years of image analysis. Nat Methods. 2012;9:671–5. doi: 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silver J, Miller JH. Regeneration beyond the glial scar. Nat Rev Neurosci. 2004;5:146–56. doi: 10.1038/nrn1326. [DOI] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012;7:562–78. doi: 10.1038/nprot.2012.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson AJ, Furness JB, Costa M. The fine structure of the submucous plexus of the guinea-pig ileum. I. The ganglia, neurons, Schwann cells and neuropil. J Neurocytol. 1981;10:759–84. doi: 10.1007/BF01262652. [DOI] [PubMed] [Google Scholar]
- Wink MR, Braganhol E, Tamajusuku AS, Lenz G, Zerbini LF, Libermann TA, Sevigny J, Battastini AM, Robson SC. Nucleoside triphosphate diphosphohydrolase-2 (NTPDase2/CD39L1) is the dominant ectonucleotidase expressed by rat astrocytes. Neuroscience. 2006;138:421–32. doi: 10.1016/j.neuroscience.2005.11.039. [DOI] [PubMed] [Google Scholar]
- Yang X, Bentink S, Scheid S, Spang R. Similarities of ordered gene lists. J Bioinform Comput Biol. 2006;4:693–708. doi: 10.1142/s0219720006002120. [DOI] [PubMed] [Google Scholar]
- Young HM, Bergner AJ, Muller T. Acquisition of neuronal and glial markers by neural crest-derived cells in the mouse intestine. J Comp Neurol. 2003;456:1–11. doi: 10.1002/cne.10448. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Chen K, Sloan SA, Bennett ML, Scholze AR, O’Keeffe S, Phatnani HP, Guarnieri P, Caneda C, Ruderisch N, et al. An RNA-Sequencing Transcriptome and Splicing Database of Glia, Neurons, and Vascular Cells of the Cerebral Cortex. J Neurosci. 2014;34:11929–47. doi: 10.1523/JNEUROSCI.1860-14.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zilliox MJ, Irizarry RA. A gene expression bar code for microarray data. Nat Methods. 2007;4:911–3. doi: 10.1038/nmeth1102. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental Figure 1. PLP1 is more commonly expressed by mucosal glia in the small intestine than GFAP. Maximum intensity projections of confocal stacks from PLP1-eGFP adult mouse duodenum (A–C) and ileum (D–I) immunostained for GFAP (A–F) or S100β (G–I). DAPI nuclear stain is represented in blue. Scale bar = 50μm.
Supplemental Figure 2. Many mucosal glia in the colon marked by PLP1 expression, do not express GFAP. Maximum intensity projections of confocal stacks from PLP1-eGFP adult mouse colon immunostained for GFAP. DAPI nuclear stain is represented in blue. Scale bar = 50μm.
Supplemental Figure 3. All 4 morphological types of enteric glia express PLP1.
Confocal single planar images of TdTomato fluorescence in colonic segments from PLPCreERT:: LSL-TdTomato mice treated with low dose tamoxifen.
Supplemental Figure 4. Isolation of PLP1+ enteric glia from postnatal mouse intestine.
FACS plots of propidium iodide (PI) fluorescence vs. GFP fluorescence generated from ileums of control (A) and PLP1-eGFP (B) littermate mice, with cross bars marking gates used to sort and collect GFP+ and GFP− cells.
Supplemental Figure 5. RNA from PLP1+ cells in the intestine is enriched for subsets of oligodendrocyte and astrocyte marker genes.
Relative difference in expression of cell-type specific genes in RNA-Seq data from GFP+ versus GFP− samples. To avoid numerical instability (dividing by 0), a pseudocount of 0.01 was added to all measured FPKM prior to calculating fold enrichment. ND = not detected.
Source data utilized for comparative analysis of glia Datasets used for comparative analysis strategy to compare the transcriptional profile of enteric glia to that of other cell types. CM = Cell Mapper.