Abstract
Background
Pseudomonas aeruginosa establishes life-long chronic airway infections in cystic fibrosis (CF) patients. As the disease progresses, P. aeruginosa pathoadaptive variants are distinguished from the initially acquired strain. However, the genetic basis and the biology of host-bacteria interactions leading to a persistent lifestyle of P. aeruginosa are not understood. As a model system to study long term and persistent CF infections, the P. aeruginosa RP73, isolated 16.9 years after the onset of airways colonization from a CF patient, was investigated. Comparisons with strains RP1, isolated at the onset of the colonization, and clonal RP45, isolated 7 years before RP73 were carried out to better characterize genomic evolution of P. aeruginosa in the context of CF pathogenicity.
Results
Virulence assessments in disease animal model, genome sequencing and comparative genomics analysis were performed for clinical RP73, RP45, RP1 and prototype strains. In murine model, RP73 showed lower lethality and a remarkable capability of long-term persistence in chronic airways infection when compared to other strains. Pathological analysis of murine lungs confirmed advanced chronic pulmonary disease, inflammation and mucus secretory cells hyperplasia. Genomic analysis predicted twelve genomic islands in the RP73 genome, some of which distinguished RP73 from other prototype strains and corresponded to regions of genome plasticity. Further, comparative genomic analyses with sequential RP isolates showed signatures of pathoadaptive mutations in virulence factors potentially linked to the development of chronic infections in CF.
Conclusions
The genome plasticity of P. aeruginosa particularly in the RP73 strain strongly indicated that these alterations may form the genetic basis defining host-bacteria interactions leading to a persistent lifestyle in human lungs.
Electronic supplementary material
The online version of this article (doi:10.1186/s12864-015-2276-8) contains supplementary material, which is available to authorized users.
Keywords: Cystic fibrosis, P. aeruginosa, Genome, Adaptation, Chronic infection, Mouse model
Background
The opportunistic pathogen Pseudomonas aeruginosa has broad capabilities to thrive in diverse ecological niches and to establish serious human infections [1]. Poor clinical outcome of P. aeruginosa-associated infection was described in immunocompromised patients and those in intensive care units, connected to mechanical ventilation or other invasive devices. P. aeruginosa is also the leading cause of chronic lung infections and death in patients with cystic fibrosis (CF), as well as a frequent cause of exacerbations in individuals with advanced chronic obstructive pulmonary disease (COPD) [2].
The genetic basis of P. aeruginosa leading to acute or chronic infection is not yet understood [3]. Genome sequencing projects are underway with the aim of providing new data to dissect the molecular basis of P. aeruginosa infections. Seventeen completely sequenced and assembled genomes are currently available and draft genomes exist for 561 additional genomes. The genome size of P. aeruginosa is larger than those of most sequenced bacteria and varies between 5.2 and 7 Mbp, with ~5500 ORFs [4]. A significant number (8,4 %) of P. aeruginosa genes are predicted to be involved in regulation, which at the time of publication was the largest fraction of regulators among sequenced bacterial genomes. Irrespective of their origin, P. aeruginosa isolates share a remarkable amount of similarity in their genome content and in virulence traits (core genome). The extent of divergence between strains is determined by extra-chromosomal elements like plasmids or blocks of DNA inserted into the chromosome at various loci [5]. These genetic features are likely to be acquired by horizontal gene transfer from different sources including other species or genera and can be present in subgroups of the P. aeruginosa population but may also be unique to single strains, accounting for most of intra- and inter-clonal P. aeruginosa genome diversity. These strain-specific segments of the genome are not scattered randomly through the core genome; rather, they tend to be clustered in certain loci, referred to as regions of genome plasticity (RGPs) [6]. The genetic sequences occupying many RGPs are often referred to as genomic islands (GIs) and islets. Therefore, the P. aeruginosa chromosome presents a picture of a mosaic, consisting of a conserved core component, interrupted in each strain by the inserted parts of the accessory genome. Genetic elements within the accessory genome may encode properties that contribute to niche-specific adaptation of the particular strains that harbor them.
Furthermore, mutations of single nucleotides also confer specific P. aeruginosa phenotypes that are advantageous under certain conditions [7–10]. Long-term colonization of the CF host is maintained by P. aeruginosa pathoadaptive lineages, which are clonal with the initially acquired strain and carry phenotypic variants. Pathoadaptive mutations are frequent in virulence genes, essential for acute infection but no longer compatible with the novel lifestyle of the P. aeruginosa in CF airways. However, little is known about the genetic basis and the biology of host-bacteria interactions leading to a persistent lifestyle of P. aeruginosa.
To define the genetic basis of P. aeruginosa persistent lifestyle, longitudinal isolates from CF patient were selected. In particular, P. aeruginosa RP73 was isolated after long-term chronic infection and compared with the preceding RP1 and clonal RP45, as well as prototype PAO1 and PA14 strains. When murine model of chronic lung infection was used, RP73 showed a marked persistent lifestyle. Thus, genome sequencing and comparative genomics analysis were carried out. Our results show the links between genomic properties and pathogenic potential of RP73 that may define the basis of long-term chronic infection by P. aeruginosa. The significance of these results is discussed in the context of understanding disease pathogenesis.
Results and discussion
Chronic colonization of a CF patient’s airways with the P. aeruginosa RP isolates
CF was suspected in the exocrine-insufficient patient index case RP (CFTR genotype: F508del/R1162X) by a positive meconium test at birth and was confirmed by pathological sweat tests at the age of 4 months. The RP patient’s airways became colonized with P. aeruginosa by the age of 7 years (Fig. 1). The CF clinic in Hannover has collected sequential isolates from this patient since the onset of colonization for up to 28 years [11]. The patient was chronically carrying P. aeruginosa isolates of clone type OC2E, for the first eleven years. During this time period strains of the clone type OC4A were sporadically isolated, but thereafter OC4A has become the dominant clone type until today. The RP patient received one to four annual 2-week courses of intravenous (iv) antipseudomonal chemotherapy since onset of colonization and was administered aerosolized colistin on a daily basis during the last 17 years. The last clone type OC2E strain was isolated from the patient’s sputum four months after the start of colistin inhalation. The patient’s clinical status remained stable during the 28 years of chronic airway infection. Lung function parameters fluctuated between 70 and 90 % predicted for forced vital capacity (FVC) and 60–80 % forced expiratory volume (FEV1) during the last 20 years with no tendency to irreversible decline.
Fig. 1.

P. aeruginosa sequential isolates from patient RP. Two clone types (OC2E and OC4A) of P. aeruginosa strains were isolated from patient RP who is heterozygous for F508del and R1162X mutations in the CFTR gene. OC2E was isolated at the onset of chronic colonization for the first eleven years. Thereafter OC4A became the dominant clone. Strain RP1 belongs to the clone type OC2E and was the first P .aeruginosa strains isolated. Strains RP45 and RP7 belong to the clone type OC4A and were isolated after 10 and 16.9 years respectively after the onset of chronic colonization of the patient’s airways with P. aeruginosa (Additional file 1 and Cramer et al. [12]). Lung function parameters at the time of P. aeruginosa isolation are indicated
In this study three P. aeruginosa isolates from RP patient were selected for genetic and biological characterization. RP1 was the first P. aeruginosa isolate and belongs to clone type OC2E, while RP45 and RP73, isolated after 10 and 16.9 years from the onset of colonization, belong to clone type OC4A (Additional file 1 and Cramer et al. [12]). Thus, RP73 P. aeruginosa isolate was able to establish long-term infection replacing the initially RP1 acquired isolate and likely adapting within CF airways respect to RP45.
Pathogenicity of P. aeruginosa RP isolates in murine model of airways infection
To translate data from CF patients into disease models, P. aeruginosa clinical isolates RP1, RP45 and RP73 were tested in the agar beads mouse model of chronic airways infection in comparison to prototype PAO1 and PA14 strains [13, 14]. Bacteria embedded in the immobilizing agents appear to grow in the microaerobic/anaerobic conditions in form of microcolonies, similarly to the growth in the mucus of patients with CF [15, 16]. RP1 isolate, as well as prototype PA14 strain, caused death in all mice (100 %) within the first three days of P. aeruginosa infection (Fig. 2 and Additional file 2). Lower incidence of mortality (50 %) was recorded after infection with RP45 strain, while RP73 were not lethal (0 %). Thus, despite their clonality, RP45 and RP73 were significantly different in the risk of death. As previously reported, prototype PAO1 showed 24 % of acute mortality [17]. Next, the capacity to establish chronic infection in the surviving mice was assessed at 14 days. Nearly all mice had chronic airways colonization by RP45 (80 %) and RP73 (90 %), demonstrating the persistent lifestyle of this lineage among surviving mice. PAO1 strain showed less capacity to establish chronic infection (24 %). The ability of the clinical isolate RP73 to achieve long-term chronic infection associated with no risk of mortality in mice was superior to all other P. aeruginosa clinical strains tested in previous studies [17].
Fig. 2.

Virulence of P. aeruginosa RP isolates in comparison with prototype strains in murine model. C57Bl/6NCrlBR mice were infected with 1-2*106 CFU/lung P. aeruginosa RP1, RP45, RP73, PAO1 and PA14 strains embedded in agar beads. Mortality induced by bacteremia (black) and survival (light gray) was evaluated on challenged mice. Clearance (white) and capacity to establish chronic infection (dark gray) were determined on surviving mice after 14 days. The data show the percentage of mice infected with single P. aeruginosa strains analyzed in two to three independent experiments. Statistical significance by Chi-square test is indicated: *P < 0.05, **P < 0.01, ***P < 0.001
To assess clinical trait of chronic infection, lung histopathology was performed after 14 days from P. aeruginosa challenge with the persistent RP73 isolate. Chronic pulmonary disease, including inflammation and mucus secretory cells, was detected. The bronchi were filled by a massive neutrophil inflammation, whereas the parenchyma was infiltrated by macrophages, lymphocytes and some neutrophils (Fig. 3a). Agar beads were observed in bronchial lumina (Fig. 3b). Mucous secretory cells hyperplasia (Fig. 3c) was found. These features resembled lesions found in CF patients with advanced chronic pulmonary disease [18].
Fig. 3.

Histological lesions after chronic P. aeruginosa infection in mice. Mice were infected with 2x106 cfu/lung of P. aeruginosa RP73 strain embedded in agar beads and lung harvested after 14 days from challenge. Histopathological analysis of lungs chronically infected by RP73 strain are characterised by acute and chronic lesions; the pulmonary parenchyma is infiltrated by macrophages, lymphocytes and some neutrophils (a). Agar beads (arrow) containing bacteria macrocolonies (*) are localised in the bronchia and surrounded by a massive neutrophils inflammation (b). Alcian blue staining shows mucus secretory cells hyperplasia (circle) (c)
Genome sequences of RP isolates and comparative genomics analyses
To link the persistent lifestyle with a genetic basis, we sequenced the genome of RP73 [19], in addition to those of preceding RP1 and clonal RP45 isolates, and performed comparative genomic analysis. The fully assembled RP73 genome consists of a single circular chromosome of 6,342,034 base pairs (Additional file 3 for genome description). Twelve genomic islands were predicted in this genome (Table 1); three of them distinguished RP73 from other prototype strains and corresponded to regions of genome plasticity (Fig. 4) [5]. They include known genomic islands PAGI-9, which is similar to rearrangement hot spots (Rhs) [20], and plasmid pKLC102, which carries the pil gene cluster and chvB glucan synthetase [21]. Nucleotide blast search on NCBI limited to P. aeruginosa showed that the former can be found in multiple clinical isolates, while the latter is identical to RP73 only in strain 8380, isolated from the human gut. However, plasmid pKLC102 is often partially present [22]. A SMC4389 CRISPR repeat sequence also differentiates RP73 from most prototype strains [6]. In fact, blast search for this sequence resulted in a single hit from soil strain Azotobacter chroococcum NCIMB 8003. The RP73 genome also contains full length LESGI-4, which was identified in the Liverpool epidemic strain (LES) [23]. Genomic islands predicted in RP73 were investigated in the draft genomes of RP1 and RP45. While RP45 carries all 12, RP1 lacks full-length plasmid pKLC102 and an ABC transporter protein. A circular map comparing the 3 sequenced RP genomes clearly shows the genomic similarity between RP45 and RP73 on one hand, and between RP1 and strain PA14, which showed similar results in the murine infection model, on the other (Fig. 4).
Table 1.
Predicted genomic islands in the genome of P. aeruginosa RP73 and comparison with other RP isolates and prototype strains
| RP73 genomic island annotation | Other strains from patient RP | Prototype strains | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Predicted Island | Start position | End position | Size (bp) | Annotation | Region of genome plasticity | RP45 | RP1 | PA14 | PAO1 | LESB58 | PA7 |
| 1–3 | 1023578 | 1126198 | 102620 | Pseudomonas aeruginosa strain C plasmid pKLC102 | 41 | P | * | * | - | - | * |
| 4 | 1372342 | 1385723 | 13381 | Major facilitator transporter, 2-isopropylmalate synthase, putative membrane-bound lytic murein transglycolase A, hypothetical proteins | na | P | P | P | P | P | - |
| 5 | 1408824 | 1421034 | 12210 | Pseudomonas aeruginosa strain SMC4389 CRISPR repeat sequence | 12 | P | P | - | - | - | - |
| 6 | 2576892 | 2587628 | 10736 | Non-ribosomal peptide synthetases, hypothetical proteins | na | P | P | P | P | P | - |
| 7 | 3092401 | 3101220 | 8819 | Type II secretion system proteins, ABC transporter permease, hypothetical proteins | na | P | P | P | P | P | - |
| 8 | 3764047 | 3768307 | 4260 | Hypothetical proteins | na | P | P | P | P | P | - |
| 9 | 3818768 | 3828628 | 9860 | Non-ribosomal peptide synthetase, FAD-dependent monooxygenase, short chain dehydrogenase, cytochrome P450, 3-oxoacyl-(acyl carrier protein) synthase III, acyl carrier protein, major facilitator transporter | na | P | P | P | P | P | - |
| 10 | 4356197 | 4363888 | 7691 | Pseudomonas aeruginosa PAGI-9 genomic island sequence | 89 | P | P | - | - | - | - |
| 11 | 5866390 | 5877731 | 11341 | Putative short-chain dehydrogenase, ABC transporter ATP-binding protein, hypothetical proteins | na | P | * | * | * | * | * |
| 12 | 6002572 | 6007043 | 4471 | Hypothetical proteins | na | P | P | - | P | P | - |
Genomic islands were predicted using Island Viewer [52] and described based on annotation with xBase [53] using P. aeruginosa PAO1 as a reference genome
Colocalization with regions of genome plasticity previously described by Klockgether et al. [5], P present, *: partially present (20–90 % coverage), -: absent
Fig. 4.
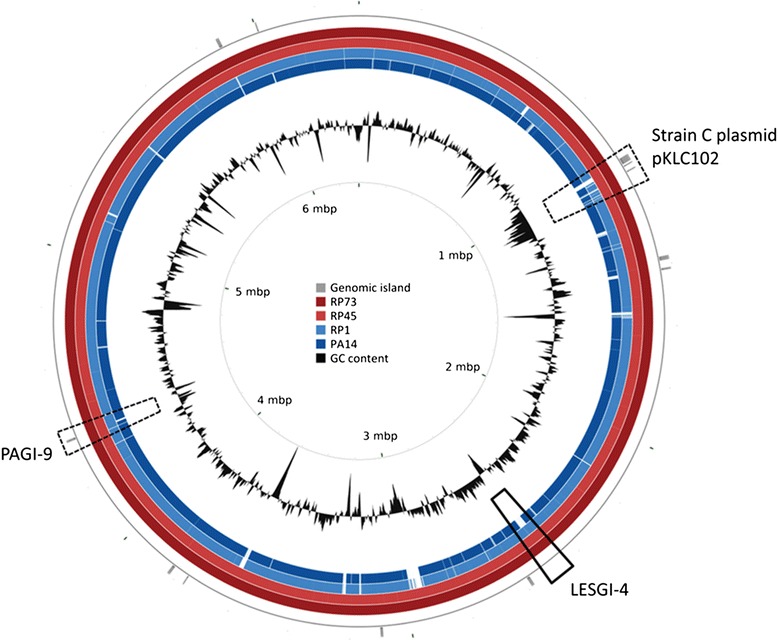
Circular map of P. aeruginosa RP isolates and prototype strain PA14. Circular map constructed with the CGView Comparison Tool [56]. Starting from the outside: genomic islands predicted with IslandViewer (see Table 1 for details) [52], RP73, RP45, RP1, PA14 and GC content. Colored regions are shared with RP73 according to blast search. Dotted lines: known genomic islands (GIs) that distinguish RP73 because they are incomplete or absent in the 12 complete P. aeruginosa genomes available at pseudomonas.com. RP isolates also carry LESGI-4, identified in the Liverpool epidemic strain
In order to study the evolution of chronic infection and determine which genetic determinants are involved in this process, further comparisons were made among RP isolates. First, analyses were carried out to study the relationship between these three strains. A core genome phylogeny was performed using 53 sequences from a previous study [24] representing an extensive sampling of P. aeruginosa’s diversity. In the resulting tree (Fig. 5), RP73 and RP45 cluster together while RP1 is found in a different and independent branch. A multi-locus sequence typing (MLST) analysis was also performed. RP73 and RP45 shared the same MLST profile while RP1 showed a different one (Additional file 4). All these analyses suggest that RP73 and RP45 are close to each other from a genetic point of view, while RP1 is more distantly related.
Fig. 5.

Core genome phylogeny for RP isolates and strains representative of P. aeruginosa diversity. The figure represents a partial view of the tree to show the relationships between RP1, RP45 and RP73. The position of RP1 is indicated in blue, while the position of RP45 and RP73 is indicated in red. PA14 is distantly related to all these strains
In order to explain the molecular bases of the development of chronic infection in a CF context, SNPs were determined between PA14 and RP1 compared to RP45 and RP73. A genomic comparison between PAO1 and RP1 was already performed by Hilker et al. [22]. A total of 54,621 SNPs was found by taking PA14 as reference strain and searching for positions at which at least one among RP1, RP45 and RP73 strains showed a different nucleotide compared to PA14. About one fifth of these SNPs (10,819) were non-synonymous substitutions (Additional file 5) that are likely to have an effect on protein function and structure. Several of these SNPs were located in virulence genes representing good candidates to explain the diversity in patterns of mortality and chronicity we observed.
Virulence factors of RP isolates and their putative role in pathogenicity
The genome of RP isolates contains most of the virulence factors, described for other P. aeruginosa genomes and identified in the Virulence Factor (VF) database (http://www.mgc.ac.cn/VFs/) [25], with few exceptions. Several VFs of RP73 isolate shows signatures of pathoadaptive mutations within the genome when compared to the preceding clonal RP45 isolate as reported in Additional files 5 and 6. Phenotypic characterization of RP isolates is reported (Table 2) and their putative role in pathogenicity discussed.
Table 2.
Phenotypic characterization of P. aeruginosa RP isolates and prototype strains
| Strain | Mucoidy | Twitching* (ø cm) | Swarming* (ø cm) | Protease (ø cm) | Siderophore (ø cm) | LasR henotype§ | Pyocyanin # | Biofilm° | Antibiotic resistance |
|---|---|---|---|---|---|---|---|---|---|
| RP1 | - | 2.3 | 3.4 | 2.1 | 1.2 | - | 0.094 ± 0.016 | 0.332 ± 0.141 | - |
| RP45 | - | - | - | - | - | + | 0.051 ± 0.015 | 0.493 ± 0.146 | GEN |
| RP73 | - | - | - | - | 1.3 | + | 0.05 ± 0.013 | 1.289 ± 0.596 | AMK; CAZ; GEN; IMP; MER |
| PAO1 | - | 1.2 | 2.6 | 1.8 | 2.0 | - | 0.127 ± 0.015 | 1.719 ± 0.217 | - |
| PA14 | - | 1.2 | 5 | 2.1 | 2.0 | - | 0.09 ± 0 | 3.553 ± 0.457 | - |
*Indicates twitching and swarming motility zone diameter, as measured by subsurface stab assay
§Isolates with iridescent and metallic sheen of the colony surface, that is typical for a lasR mutant, are indicated (+)
#Indicates mean value ± SD at 26 h. Values ≤ 0.05 indicate no production of pyocyanin
°Indicates mean value ± SD at 24 h
AMK amikacin, CAZ ceftazidime, GEN gentamicin, IMP imipenem, MER meropenem
Motility, adherence and cell interaction
Pili, flagella and outer membrane proteins promote motility, attach to epithelial or endothelial cells, activate or inactivate host cellular pathways and immune responses [26]. These P. aeruginosa VFs play a key role in acute infection and are present in RP1 isolate. Variations in the twitching and swimming motility are common in P. aeruginosa isolates from CF patients and described to be hallmarks of bacterial adaptation to the airways [7, 27]. Both RP73 and its clonal RP45 isolate did not encode pilV, pilW, pilY2 and fimT and carried a premature stop mutation in pilO (Table 3); while pilA and pilC were deleted in all isolates from RP patient. The RP73 and RP45 phenotypes are consistent with the absence of twitching and swimming motility (Table 2). Lack of motility was associated with decreased virulence in models of acute infection [28, 29] and increased risk of chronic infection [17]. Our results obtained in RP73 and RP45-infected mice (Fig. 2) are consistent with the observation that unlike strains RP1, PA14 and PAO1, favor long-term persistence.
Table 3.
Comparative pathogenomics of mutations present in the major virulence factors of RP isolates respect to the prototype strain PAO1 and evaluation in others P. aeruginosa sequenced strains
| Virulence factors | ORFs | PAO11 | PA141 | LESB581 | PA71 | RP12 | RP452 | Mutation in RP45 | RP732 | Mutation in RP73 |
|---|---|---|---|---|---|---|---|---|---|---|
| Type IV Pili biosynthesis | pilA | PA4525 | PA14_58730 | PLES_49071 | PSPA7_5161 | deleted3 | deleted3 | deleted | ||
| pilC | PA4527 | PA14_58760 | PLES_49101 | PSPA7_5163 | deleted3 | deleted3 | deleted | |||
| pilO | PA5042 | PA14_66640 | PLES_54321 | PSPA7_5779 | present | present | stop codon after 138 aa | present | stop codon after 138 aa | |
| pilV | PA4551 | deleted | PLES_49341 | PSPA7_5191 | present | deleted4 | deleted | |||
| pilW | PA4552 | PA14_60290 | PLES_49351 | PSPA7_5192 | present | deleted4 | deleted | |||
| pilY2 | PA4555 | deleted | PLES_49381 | PSPA7_5195 | present | deleted4 | deleted | |||
| fimT | PA4549 | deleted | PLES_49321 | PSPA7_5189 | present | deleted3 | deleted | |||
| Alginate regulation | algP/algR3 | PA5253 | PA14_69370 | PLES_56471 | PSPA7_5998 | present | present | present | ||
| Pyochelin | pchD | PA4228 | PA14_09240 | PLES_06991 | PSPA7_0872 | present | present | stop codon after 276 aa | present | stop codon after 276 aa |
| Pyoverdin | pvdD | PA2399 | PA14_33650 | PLES_28971 | deleted | present5 | present | present | stop codon after 839 aa | |
| Phospholipase D | pldA | PA3487 | PA14_18970 | deleted | deleted | deleted4 | deleted4 | deleted | ||
| N-(3-oxo-dodecanoyl)-L-homoserine lactone QS system | lasR | PA1430 | PA14_45960 | PLES_39841 | PSPA7_3898 | present | present | stop codon after 220 aa | present | stop codon after 220 aa |
| Type III Secretion System | pscQ | PA1694 | PA14_42610 | PLES_36321 | deleted | present | present | deleted | ||
| pscP | PA1695 | PA14_42600 | deleted | deleted | present | present5 | deleted | |||
| pscO | PA1696 | PA14_42580 | PLES_36311 | deleted | present | present | deleted | |||
| pscN | PA1697 | PA14_42570 | PLES_36301 | deleted | present | present | deleted | |||
| poPN | PA1698 | PA14_42550 | PLES_36291 | deleted | present | present | deleted | |||
| pcr1 | PA1699 | PA14_42540 | PLES_36281 | deleted | present | present | deleted | |||
| pcr2 | PA1700 | PA14_42530 | PLES_36271 | deleted | present | present | deleted | |||
| pcr3 | PA1701 | PA14_42520 | PLES_36261 | deleted | present | present | deleted | |||
| pcr4 | PA1702 | PA14_42510 | PLES_36251 | deleted | present | present | deleted | |||
| pcrD | PA1703 | PA14_42500 | PLES_36241 | deleted | present | present | deleted | |||
| pcrR | PA1704 | PA14_42490 | PLES_36231 | deleted | present | present | deleted | |||
| pcrG | PA1705 | PA14_42480 | PLES_36221 | deleted | present | present | deleted | |||
| pcrV | PA1706 | PA14_42470 | PLES_36211 | deleted | present | present | deleted | |||
| pcrH | PA1707 | PA14_42460 | deleted | deleted | present | present | deleted | |||
| poPB | PA1708 | PA14_42450 | PLES_36201 | deleted | present | present | deleted | |||
| poPD | PA1709 | PA14_42440 | PLES_36191 | deleted | present | present | deleted | |||
| exsC | PA1710 | PA14_42430 | PLES_36181 | deleted | present | present | deleted | |||
| exsE | PA1711 | PA14_42410 | deleted | deleted | present | present | deleted | |||
| exsB | PA1712 | PA14_42400 | PLES_36171 | deleted | present | present | deleted | |||
| exsA | PA1713 | PA14_42390 | PLES_36161 | deleted | present | present | deleted | |||
| exsD | PA1714 | PA14_42380 | PLES_36151 | deleted | present | present | deleted | |||
| pscB | PA1715 | PA14_42360 | PLES_36141 | deleted | present | present | deleted | |||
| pscC | PA1716 | PA14_42350 | PLES_36131 | deleted | present | present | deleted | |||
| pscD | PA1717 | PA14_42340 | PLES_36121 | deleted | present5 | present | deleted | |||
| pscE | PA1718 | PA14_42320 | PLES_36111 | deleted | present | present | deleted | |||
| pscF | PA1719 | PA14_42310 | PLES_36101 | deleted | present | present | deleted | |||
| pscG | PA1720 | PA14_42300 | PLES_36091 | deleted | present | present | deleted | |||
| pscH | PA1721 | PA14_42290 | PLES_36081 | deleted | present | present | deleted | |||
| pscI | PA1722 | PA14_42280 | PLES_36071 | deleted | present | present | deleted | |||
| pscJ | PA1723 | PA14_42270 | PLES_36061 | deleted | present | present | deleted | |||
| pscK | PA1724 | PA14_42260 | PLES_36051 | deleted | present | present | deleted | |||
| pscL | PA1725 | PA14_42250 | PLES_36041 | deleted | present | present | deleted | |||
| LPS | lpxO2 | PA0936 | PA14_52150 | PLES_43801 | PSPA7_4774 | present | present | present | stop codon after 145 aa | |
| Efflux pumps | mexA | PA0425 | PA14_05530 | PLES_04231 | PSPA7_0525 | present | present | +29 aa at N-ter | present | +29 aa at N-ter |
| mexT | PA2492 | PA14_32410 | pseudogene | PSPA7_2746 | present | present | N-ter not conserved | present | N-ter not conserved | |
| Antibiotic resistance | arr | PA2818 | deleted | deleted | PSPA7_2339 | deleted3 | deleted3 | deleted |
1Sources for PAO1, PA14, LESB58 and PA7 information are the Virulence Factor Database
(http://www.mgc.ac.cn/VFs/) and Pseudomonas Database (www.pseudomonas.com)
2Identified by local blast (>90 % query coverage, >90 % sequence similarity) using the PAO1 gene sequences as query
3Presence of at least one flanking region (500 bp) confirmed by blast search on contigs; gene not located at contig edge
4Flanking regions not found, but absence of the gene is consistent with a deletion in isolate RP73
5Gene is present, but separated onto multiple contigs
Lipopolysaccharides (LPSs) are potent immune stimulants through their interactions with Toll-like receptor 4 (TLR4). P. aeruginosa strains isolated from CF patients evolved the capacity to reduce host immuno-detection by modulating LPS structure [30]. Biochemical and biological characterization of RP73 LPS showed to possess an under-acylated lipid A leading to a lower pro-inflammatory capacity in a murine model of intranasal instillation when compared to the LPS from the prototype strain PAO1 [31]. This structure is distinguished by the absence of hexa- or hepta-acylated lipid A species that are typical phenotypic changes that can occur on the LPS molecule of a P. aeruginosa chronic strain. Furthermore, RP73 carries an R-type LPS without O-antigen in which the lipid A is covalently linked to the core oligosaccharide region. Absence of LPS O-antigen in RP73 suggests an adaptation of this strain to persistent lifestyle.
Among genes responsible for lipid A modification (lpxO1, lpxO2, phoP, phoQ, pagL and oprH) [30], our genetic characterization showed that RP73 isolate carries a stop mutation and lacks of the C-terminus of the protein in lpxO2 compared to the preceding clonal RP45, as well as RP1 isolate and prototype PAO1 and PA14 strains (Table 3). Both RP45 and RP73 isolates carry a non-synonimous SNP in the sequence of pagL compared with RP1 and PA14 (Additional files 5 and 6). These data suggests that an adaptive process has occurred to the LPS structure of the RP lineage in the period of time between the isolation of RP45 and RP73.
Secretion systems and toxins
In P. aeruginosa genome, the genes (psc, pcr, exs and pop) encoding the type III secretion system (T3SS) are clustered together. The psc and prc genes primarily encode components of the bacterial secretion apparatus whereas the exs gene products are involved in regulation of TTSS. Two pop genes encode proteins (PopB and PopD) essential for the translocation of the effectors into host cells. Remarkably, RP73 isolate lost the entire 32-gene cluster encoding the T3SS which was present in the preceding clonal RP45, as well as RP1 and prototype PAO1 and PA14 strains (Table 3) [25]. However, the exoS, exoT, and exoY genes encoding for the “T3SS translocated effectors” are still present both in RP73 and RP45 genome with two non-synonymous SNPs recorded in the exoT when compared with RP1 and PA14 (Table 3 and Additional files 5 and 6). T3SS is an important virulence determinant of P. aeruginosa which may act at the site of infection and contribute to subversion of the host immune response. In contrast to acute infection, small proportion of isolates infecting CF patients secrete T3SS proteins and this proportion decreases with duration of infection [32]. We speculate that the differences in the risk of mortality associated to RP73 and RP45 isolates may be linked to absence or presence of the T3SS that changed during the progression of CF chronic infection.
Genes encoding for toxA, hcnA, hcnB and hcnC are present. However, RP73 has two non-synonymous SNPs compared with its clonal isolate RP45, as well as RP1 and PA14, in hcnC gene. If we consider the exotoxin A, we observed several non-synonymous SNPs (Table 3 and Additional file 5).
Finally, pldA coding for the periplasmic phospholipase D (Table 3), one of effectors of the type VI secretion system [33], is absent in RP73 genome. However pldA is reported to be not conserved among P. aeruginosa genomes [34].
Iron uptake and pigment
The ability to produce siderophores, like pyochelin and pyoverdine, has been linked to the bacterial pathogenic potential. Phenotypic assay showed that RP isolates secreted lower (RP73 and RP1) or no (RP45) levels of siderophores when compared to PAO1 and PA14 (Table 2). When we look at the genome sequence, both RP73 and RP45 carries stop mutations both in pchD, while PvdD protein is interrupted in RP73 only. PchD and PvdD are both involved in the synthesis of the two principal siderophores (Table 3). PvdD production was shown to be required for airways bacterial colonization in rat, lethal virulence in burned and immunosuppressed mouse models [35, 36]. Pyoverdine was detected in the sputa of CF patients [37], while in a larger study, one-third of sputa positive for P. aeruginosa contained no detectable pyoverdine [38]. These data suggests that pyoverdine-mediated iron uptake may not always be essential for chronic infection and other mechanisms are active in CF [39].
Phenotypic tests showed that RP73 and RP45 are able to secrete less pyocyanin when compared to RP1, PAO1 and PA14 strains (Table 2). No changes in genes involved in pyocyanin biosynthesis were found, however in RP73 and RP45 were found non-synonymous SNPs in several genes belonging to the two phenazine biosynthesis operons (phzA1B1C1D1E1F1G1 and phzA2B2C2D2E2F2G2) if compared with the non-clonal isolate RP1 and the prototype strain PA14 (Additional file 5). Pyocyanin is required for full virulence in animal models and has been detected in patients’ airway secretions, promoting virulence by interfering with several cellular functions in host cells including electron transport, cellular respiration, energy metabolism, gene expression, and innate immune mechanisms [40].
Mucoidy
RP isolates are phenotypically non-mucoid (Table 2) with absence of mutations in mucABD locus. Other key regulators of alginate pathway are present (Table 3). The phenotypic switch to mucoidy in P. aeruginosa infections is a well-established paradigm in CF. Infection with the mucoid phenotype, which produces large amounts of the exopolysaccharide alginate, has been associated with a more rapid decline in pulmonary function than infection with non-mucoid bacteria [41]. However, some but not all P. aeruginosa isolates became mucoid in the CF lung suggesting that a mucoid phenotype did not always confer a selective advantage to bacterial cells in persistence [42]. In mouse model, CF clonal strains, displaying a mucoid and a non-mucoid phenotype, showed a similar capacity of persistence [17]. Our data obtained in mouse model with RP73 and RP45 isolates support the notion that non-mucoid P. aeruginosa strains are able of long-term persistence.
Quorum sensing
In P. aeruginosa, many virulence determinants and secondary metabolites are regulated in a cell population density-dependent manner via cell-to-cell communication or “quorum sensing” (QS) [43]. P. aeruginosa possesses two N-acylhomoserine lactone (AHL)-dependent QS systems. These are termed the las and rhl systems, consisting of the LuxRI homologues, LasRI and RhlRI, respectively. RP isolates have no mutations in lasI, rhlR and rhlI genes. However, inactivation of the transcriptional regulator LasR, carrying a stop mutation in the gene sequence (Table 3), was found both in RP73 and RP45, while the RP1 has no changes. The distinctive lasR mutant phenotype was confirmed by colony morphology that includes surface iridescent sheen and colony flattening exclusively in RP73 and RP45 (Table 2) [44].
Mutations in lasR lead to several phenotypic changes of potential clinical significance, including a growth advantage in amino acid abundant CF airway secretions. LasR regulates the production of virulence factors (elastase, protease, alkaline protease and exotoxin A) affecting the immune response and antibiotic resistance [45]. Most importantly, lasR mutations are often associated with the progression of CF lung disease and may serve as a marker of early CF adaptive change of prognostic significance [46].
Antibiotic resistance
RP73 showed remarkable resistance to most of the antibiotic classes while the preceding RP45 and RP1 isolates were not, indicating an increased treatment-refractory during the course of the chronic infection in this CF patient (Table 2). A strong link between antibiotic resistance and hypermutation was observed in patients with CF [47]. However, RP73 strain does not have mutations in mutS, mutL and uvrD, described previously as responsible for the hypermutable phenotype [48]. Regarding efflux pumps, the RP73 strain did not show mutation in mexEF-oprN, mexCD-oprJ and mexXY. Respectively one and six non-synonymous SNPs are present in the sequence of mexC and mexD of RP73 when compared with its clonal isolate RP45. No insertions or deletions in ampC, ampR, mexR, mexZ and oprD were detected. An insertion at the N-terminal of MexA and a non-synonymous SNP was found in RP73 and RP45 (Table 3 and Additional file 5). MexA belongs to the efflux pump complex MexAB-OprM, which is anchored to the inner membrane via N-terminal fatty acids. Adaptive mutations in mexA have been reported in CF isolates [7].
Additional modifications were detected at the N-terminal of mexT, which is not conserved in RP73 and RP45 (Table 3). An additional non-synonymous aa change at position 128 was found in RP73 when compared to RP45 (Additional file 5). MexT plays a pleiotropic role in modulating P. aeruginosa virulence such as TTSS, pyocyanin production and early surface attachment [49]. Similarly to MexA, also MexT is an hallmark of P. aeruginosa adaptation in CF patients [7]. Among the additional 58 PAO1 coding sequence annotated as “antibiotic resistance and susceptibility”, only arr, a putative aminoglycoside response regulator, is absent in all strains from RP patient.
Conclusions
Taken together, our study combined clinical data, whole-genome analysis and animal models to link the persistent lifestyle of P. aeruginosa in CF lungs with the bacterial genetic basis. Starting from a clinical case of CF, P. aeruginosa RP73 was isolated after long-term chronic infection and compared with the preceding RP1 and clonal RP45, as well as prototype PAO1 and PA14 strains. When tested in the animal model, P. aeruginosa RP73 isolate, but not other strains, mimics most of the traits of airways infection and inflammation observed in CF patients. These results suggested that key features of RP73 isolate may contribute to its pathogenesis. The genome sequence of RP73 and comparative genomics analysis with other P. aeruginosa genomes, pointed clearly to signatures of pathoadaptive mutations within the genome. This in turn correlated with the major impact on the in vitro phenotypes and in vivo maintenance observed and described here. Our findings support and better define the hypothesis that genes encoding major virulence factors are deleted and/or contain beneficial mutations when P. aeruginosa establishes long-term chronic infection. The results presented in this study provide important information with respect to the P. aeruginosa mosaic genome structure and chronic infections found in CF patients.
Methods
Ethics statement
Animal studies adhered strictly to the Italian Ministry of Health guidelines for the use and care of experimental animals. The use of the clinical data is in line with study no. 3739 which has been approved by the Ethics Commission of Hannover Medical School. The patient and her parents provided informed consent prior to sampling of strains and storage of clinical data.
Bacterial strains and CF patient
CF clinical P. aeruginosa RP1, RP45 and RP73 isolates were chosen from the collection of the CF clinic Medizinische Hochschule Hannover, Germany. Genotypic analysis by multimarker array and phenotypic data of P. aeruginosa strains were partly published [50]. P. aeruginosa was cultured in Pseudomonas isolation agar (PIA) or Trypticase Soy Broth (TSB) at 37 °C. CF patient gave informed consent before the sample collection. Approval for storing of biological materials was obtained by the Hannover Medical School, Germany.
Phenotypic characterization
Swimming and twitching capacities, protease, siderophore and pyocyanin secretion, hemolytic activity, and LasR mutant phenotypic analysis were assayed as described in the online data supplement.
Genome sequencing
The genome of RP73 was previously published [19]. Genomic DNA from strains RP1 and RP45 was isolated from overnight cultures using the DNeasy Blood and Tissue Kit (QIAGEN). Genomic DNA (500 ng) was mechanically fragmented for 40 s using a Covaris M220 (Covaris, Woburn MA, USA) with default settings. Fragmented DNA was transferred to PCR tubes and library synthesis was performed with the Kapa Hyperprep kit (Kapa biosystems, Wilmington MA, USA) according to manufacturer’s instructions. TruSeq HT adapters (Illumina, SanDiego, CA, USA) were used to barcode the samples and each library was sequenced in 1/48 of an Illumina MiSeq 300 bp paired-end run at the Plateforme d’Analyses Génomiques of the Institut de Biologie Intégrative et des Systèmes (Laval University, Quebec, Canada). Sequencing data for each genome was assembled with the A5 pipeline [51]. Whole genome shotgun projects has been deposited at DDBJ/EMBL/GenBank under accessions LNBU00000000 (RP1) and LNDM00000000 (RP45).
Genomic analyses
Blast (NCBI) was used to compare the genome of RP73 to prototype strains and to the complete Pseudomonas aeruginosa content in Genbank. Genomic islands were predicted with Island Viewer [52] and annotated with xBase [53]. The core genome phylogeny was determined using the Harvest suite [51]. The data set of sequences we used to generate the core phylogeny includes 53 strains representative of P. aeruginosa diversity. MLST profiles were determined combining the results obtained from the pubmlst database (http://pubmlst.org) and the SRST2 software package [54]. SNPs between PA14 and RP1, RP45 and RP73 were detected with the Samtools software package [55] (samtools mpileup options: -C 50, SNPs with quality score of less than 30 were discarded).
Mouse model of P. aeruginosa acute and chronic lung infection
For chronic infection, C57BL/6NCrlBR male mice (20–22 g, Charles River) were infected with 1-2×106 CFUs of P. aeruginosa strains, embedded in agar beads [15, 17]. Fourteen days post-challenge lungs were recovered, homogenized and plated for CFUs counting. In additional group of mice, the lungs were excised for histopathology. Additional details are reported in the Online data supplement. Student’s t-test and the χ2 test considering p < 0.05 as the limit of statistical significance was performed.
Availability of supporting data
The data sets supporting the results of this article are included within the article and its additional files.
Acknowledgments
This study was supported by grants to A.B. from the Cystic Fibrosis Foundation (US) (BRAGON11I0) and Ministero della Salute (project GR/2009/1579812). R.C.L. is funded by Cystic Fibrosis Canada and by CIHR. J.J. holds a Cystic Fibrosis Canada postdoctoral research fellowship. We thank members of the IBIS Plateforme d’analyses génomiques and the IBIS bioinformatics group for genome assembly. Part of this work was carried out in ALEMBIC, an advanced microscopy laboratory established by the San Raffaele Scientific Institute and the Vita-Salute San Raffaele University (Milan).
Abbreviations
- AHL
N-acylhomoserine lactone
- CDSs
coding sequences
- CF
cystic fibrosis
- CFUs
colony forming units
- COPD
chronic obstructive pulmonary disease
- FEV
forced expiratory volume
- FVC
forced vital capacity
- GIs
genomic islands
- LES
liverpool epidemic strain
- LPSs
lipopolysaccharides
- MDR
multidrug-resistant
- ORFs
open reading frames
- PIA
Pseudomonas isolation agar
- PFGE
pulsed field gel electrophoresis
- TLR4
toll-like receptor 4
- TSB
trypticase soy broth
- T3SS
type III secretion system
- VF
virulence factor
Additional files
PFGE of P. aeruginosa RP isolates. (PNG 112 kb)
Virulence of P. aeruginosa RP isolates, and prototype strains in a murine model of chronic airways infection. (DOC 29 kb)
MLST typing of RP isolates [ 55 ]. (DOC 28 kb)
Non-synonymous SNPs between prototype strain PA14 and strains RP73, RP45 and RP1. (XLS 2195 kb)
Non-synonymous SNPs between prototype strain PA14 and strains RP73, RP45 and RP1 located in known virulence factors. (XLS 31 kb)
Footnotes
Irene Bianconi and Julie Jeukens contributed equally to this work.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
IB, JJ, LF, BAF, MF, BB and IKI performed research; IB, AM, BT, RL, and AB designed research; BT, AM, RL and AB contributed new reagents/analytic tools; IB, JJ, LF, BAF, MF, BB, AM and AB analyzed data; IB, JJ, LF, BAF, BT, RCL and AB wrote the paper. All authors read and approved the final manuscript.
Contributor Information
Irene Bianconi, Email: bianconi.irene@hsr.it.
Julie Jeukens, Email: julie.jeukens.1@ulaval.ca.
Luca Freschi, Email: luca.freschi@bio.ulaval.ca.
Beatriz Alcalá-Franco, Email: alcalafranco.beatriz@hsr.it.
Marcella Facchini, Email: facchini.marcella@hsr.it.
Brian Boyle, Email: Brian.Boyle@ibis.ulaval.ca.
Antonio Molinaro, Email: molinaro@unina.it.
Irena Kukavica-Ibrulj, Email: kukavica@fmed.ulaval.ca.
Burkhard Tümmler, Email: Tuemmler.Burkhard@mh-hannover.de.
Roger C. Levesque, Email: rclevesq@ibis.ulaval.ca
Alessandra Bragonzi, Phone: +39(02)26434189, Email: bragonzi.alessandra@hsr.it.
References
- 1.Lyczak JB, Cannon CL, Pier GB. Establishment of Pseudomonas aeruginosa infection: lessons from a versatile opportunist. Microbes Infect. 2000;2:1051–60. doi: 10.1016/S1286-4579(00)01259-4. [DOI] [PubMed] [Google Scholar]
- 2.Gellatly S, Hancock RE. Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog Dis. 2013;67(3):159–73. doi: 10.1111/2049-632X.12033. [DOI] [PubMed] [Google Scholar]
- 3.Yahr T, Greenberg EP. The genetic basis for the commitment to chronic versus acute infection in Pseudomonas aeruginosa. Mol Cell. 2004;16(4):497–8. doi: 10.1016/j.molcel.2004.11.009. [DOI] [PubMed] [Google Scholar]
- 4.Stover CK, Pham XQ, Erwin AL, Mizoguchi SD, Warrener P, Hickey MJ, et al. Complete genome sequence of Pseudomonas aeruginosa PA01, an opportunistic pathogen. Nature. 2000;406(6799):959–64. doi: 10.1038/35023079. [DOI] [PubMed] [Google Scholar]
- 5.Klockgether J, Cramer N, Wiehlmann L, Davenport CF, Tümmler B. Pseudomonas aeruginosa genomic structure and diversity. Front Microbiol. 2011;2:150. doi: 10.3389/fmicb.2011.00150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mathee K, Narasimhan G, Valdes C, Qiu X, Matewish JM, Koehrsen M, et al. Dynamics of Pseudomonas aeruginosa genome evolution. Proc Natl Acad Sci U S A. 2008;105(8):3100–5. doi: 10.1073/pnas.0711982105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Smith EE, Buckley DG, Wu Z, Saenphimmachak C, Hoffman LR, D’Argenio DA, et al. Genetic adaptation by Pseudomonas aeruginosa to the airways of cystic fibrosis patients. Proc Natl Acad Sci U S A. 2006;30(103 22):8487–92. doi: 10.1073/pnas.0602138103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Nguyen D, Singh PK. Evolving stealth: genetic adaptation of Pseudomonas aeruginosa during cystic fibrosis infections. Proc Natl Acad Sci U S A. 2006;30(103(22)):8305–6. doi: 10.1073/pnas.0602526103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yang L, Jelsbak L, Marvig RL, Damkiær S, Workman CT, Rau MH, et al. Evolutionary dynamics of bacteria in a human host environment. Proc Natl Acad Sci U S A. 2011;108(18):7481–6. doi: 10.1073/pnas.1018249108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Yang L, Jelsbak L, Molin S. Microbial ecology and adaptation in cystic fibrosis airways. Environ Microbiol. 2011;13(7):1682–9. doi: 10.1111/j.1462-2920.2011.02459.x. [DOI] [PubMed] [Google Scholar]
- 11.Cramer N, Wiehlmann L, Ciofu O, Tamm S, Høiby N, Tümmler B. Molecular epidemiology of chronic Pseudomonas aeruginosa airway infections in cystic fibrosis. PLoS ONE. 2012;7(11):e50731. doi: 10.1371/journal.pone.0050731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cramer N, Klockgether J, Wrasman K, Schmidt M, Davenport CF, Tümmler B. Microevolution of the major common Pseudomonas aeruginosa clones C and PA14 in cystic fibrosis lungs. Environ Microbiol. 2011;13(7):1690–704. doi: 10.1111/j.1462-2920.2011.02483.x. [DOI] [PubMed] [Google Scholar]
- 13.Bragonzi A. Murine models of acute and chronic lung infection with cystic fibrosis pathogens. IJMM. 2010;300(8):584–93. doi: 10.1016/j.ijmm.2010.08.012. [DOI] [PubMed] [Google Scholar]
- 14.Facchini M, De Fino I, Riva C, Bragonzi A. Long-term chronic Pseudomonas aeruginosa airway infection in mice. J Vis Exp. 2014;17:85. doi: 10.3791/51019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bragonzi A, Worlitzsch D, Pier GB, Timpert P, Ulrich M, Hentzer M, et al. Nonmucoid Pseudomonas aeruginosa expresses alginate in the lungs of patients with cystic fibrosis and in a mouse model. J Infect Dis. 2005;192(3):410–9. doi: 10.1086/431516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Worlitzsch D, Tarran R, Ulrich M, Schwab U, Cekici A, Meyer KC, et al. Effects of reduced mucus oxygen concentration in airway Pseudomonas infections of cystic fibrosis patients. J Clin Invest. 2002;109(3):317–25. doi: 10.1172/JCI0213870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bragonzi A, Paroni M, Nonis A, Cramer N, Montanari S, Rejman J, et al. Pseudomonas aeruginosa microevolution during cystic fibrosis lung infection establishes clones with adapted virulence. AJRCCM. 2009;180(2):138–45. doi: 10.1164/rccm.200812-1943OC. [DOI] [PubMed] [Google Scholar]
- 18.Bedrossian C, Greenberg SD, Singer DB, Hansen JJ, Rosenberg HS. The lung in cystic fibrosis. A quantitative study including prevalence of pathologic findings among different age groups. Hum Pathol. 1976;7(2):195–204. doi: 10.1016/S0046-8177(76)80023-8. [DOI] [PubMed] [Google Scholar]
- 19.Jeukens J, Boyle B, Bianconi I, Kukavica-Ibrulj I, Tümmler B, Bragonzi A, et al. Complete Genome Sequence of Persistent Cystic Fibrosis Isolate Pseudomonas aeruginosa Strain RP73. Genome Announc. 2013;1(4):e00568. doi: 10.1128/genomeA.00568-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Battle S, Rello J, Hauser AR. Genomic islands of Pseudomonas aeruginosa. FEMS Microbiol Lett. 2009;290:70–8. doi: 10.1111/j.1574-6968.2008.01406.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Klockgether J, Reva O, Larbig K, Tümmler B. Sequence analysis of the mobile genome island pKLC102 of Pseudomonas aeruginosa C. J Bacteriol. 2004;186(2):518–34. doi: 10.1128/JB.186.2.518-534.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hilker R, Munder A, Klockgether J, Losada PM, Chouvarine P, Cramer N, et al. Interclonal gradient of virulence in the Pseudomonas aeruginosa pangenome from disease and environment. Environ Microbiol. 2015;17(1):29–46. doi: 10.1111/1462-2920.12606. [DOI] [PubMed] [Google Scholar]
- 23.Winstanley C, Langille MG, Fothergill JL, Kukavica-Ibrulj I, Paradis-Bleau C, Sanschagrin F, et al. Newly introduced genomic prophage islands are critical determinants of in vivo competitiveness in the Liverpool Epidemic Strain of Pseudomonas aeruginosa. Genome Res. 2009;19(1):12–23. doi: 10.1101/gr.086082.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Stewart L, Ford A, Sangal V, Jeukens J, Boyle B, Kukavica-Ibrulj I, et al. Draft genomes of 12 host-adapted and environmental isolates of Pseudomonas aeruginosa and their positions in the core genome phylogeny. Pathog Dis. 2014;71(1):20–5. doi: 10.1111/2049-632X.12107. [DOI] [PubMed] [Google Scholar]
- 25.Yang J, Chen L, Sun L, Yu J, Jin Q. VFDB 2008 release: an enhanced web-based resource for comparative pathogenomics. Nucleic Acids Res. 2008;36:539–42. doi: 10.1093/nar/gkm951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Filloux A. Protein secretion systems in pseudomonas aeruginosa: an essay on diversity, evolution, and function. Front Microbiol. 2011;2:155. doi: 10.3389/fmicb.2011.00155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bianconi I, Milani A, Paroni M, Cigana C, Levesque RC, Bertoni G, et al. Positive signature-tagged mutagenesis in Pseudomonas aeruginosa: tracking patho-adaptive mutations promoting long-term airways chronic infection. PlosPathogen. 2011;7(2):e1001270. doi: 10.1371/journal.ppat.1001270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Tang HB, DiMango E, Bryan R, Gambello M, Iglewski BH, Goldberg JB, et al. Contribution of specific Pseudomonas aeruginosa virulence factors to pathogenesis of pneumonia in a neonatal mouse model of infection. Infect Immun. 1996;64(1):37–43. doi: 10.1128/iai.64.1.37-43.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sato H, Okinaga K, Saito H. Role of pili in the pathogenesis of Pseudomonas aeruginosa burn infection. Microbiol Immunol. 1988;32(2):131–9. doi: 10.1111/j.1348-0421.1988.tb01372.x. [DOI] [PubMed] [Google Scholar]
- 30.Cigana C, Curcurù L, Leone MR, Ieranò T, Lorè NI, Bianconi I, et al. Pseudomonas aeruginosa exploits lipid A and muropeptides modification as a strategy to lower innate immunity during cystic fibrosis lung infection. PLoS ONE. 2009;4(12):e8439. doi: 10.1371/journal.pone.0008439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Di Lorenzo F, Silipo A, Bianconi I, Lore’ NI, Scamporrino A, Sturiale L, et al. Persistent cystic fibrosis isolate Pseudomonas aeruginosa strain RP73 exhibits an under-acylated LPS structure responsible of its low inflammatory activity. Mol Immunol. 2014;S0161-5890(14):00084. doi: 10.1016/j.molimm.2014.04.004. [DOI] [PubMed] [Google Scholar]
- 32.Jain M, Ramirez D, Seshadri R, Cullina JF, Powers CA, Schulert GS, et al. Type III secretion phenotypes of Pseudomonas aeruginosa strains change during infection of individuals with cystic fibrosis. J Clin Microbiol. 2004;42(11):5229–37. doi: 10.1128/JCM.42.11.5229-5237.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Jiang F, Waterfield NR, Yang J, Yang G, Jin Q. A Pseudomonas aeruginosa type VI secretion phospholipase D effector targets both prokaryotic and eukaryotic cells. Cell Host Microbe. 2014;15(5):600–10. doi: 10.1016/j.chom.2014.04.010. [DOI] [PubMed] [Google Scholar]
- 34.Wilderman P, Vasil AI, Johnson Z, Vasil ML. Genetic and biochemical analyses of a eukaryotic-like phospholipase D of Pseudomonas aeruginosa suggest horizontal acquisition and a role for persistence in a chronic pulmonary infection model. Mol Microbiol. 2001;39(2):291–303. doi: 10.1046/j.1365-2958.2001.02282.x. [DOI] [PubMed] [Google Scholar]
- 35.Poole K, Dean C, Heinrichs D, Neshat S, Krebs K, Young L, et al. Siderophore-mediated iron transport in Pseudomonas aeruginosa. In: Nakazawa T, et al., editors. Molecular biology of Pseudomonas. Washington, DC: American Society for Microbiology; 1996. pp. 371–83. [Google Scholar]
- 36.Takase H, Nitanai H, Hoshino K, Otani T. Impact of siderophore production on Pseudomonas aeruginosa infections in immunosuppressed mice. Infect Immun. 2000;68(4):1834–9. doi: 10.1128/IAI.68.4.1834-1839.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Haas B, Kraut J, Marks J, Zanker SC, Castignetti D. Siderophore presence in sputa of cystic fibrosis patients. Infect Immun. 1991;59(11):3997–4000. doi: 10.1128/iai.59.11.3997-4000.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Martin L, Reid DW, Sharples KJ, Lamont IL. Pseudomonas siderophores in the sputum of patients with cystic fibrosis. Biometals. 2011;24(6):1059–67. doi: 10.1007/s10534-011-9464-z. [DOI] [PubMed] [Google Scholar]
- 39.Konings A, Martin LW, Sharples KJ, Roddam LF, Latham R, Reid DW, et al. Pseudomonas aeruginosa uses multiple pathways to acquire iron during chronic infection in cystic fibrosis lungs. Infect Immun. 2013;81(8):2697–704. doi: 10.1128/IAI.00418-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Rada B, Leto TL. Pyocyanin effects on respiratory epithelium: relevance in Pseudomonas aeruginosa airway infections. Trends Microbiol. 2013;21(2):73–81. doi: 10.1016/j.tim.2012.10.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Li Z, Kosorok MR, Farrell PM, Laxova A, West SE, Green CG, et al. Longitudinal development of mucoid Pseudomonas aeruginosa infection and lung disease progression in children with cystic fibrosis. Jama. 2005;293(5):581–8. doi: 10.1001/jama.293.5.581. [DOI] [PubMed] [Google Scholar]
- 42.Bragonzi A, Wiehlmann L, Klockgether J, Cramer N, Worlitzsch D, Döring G, et al. Sequence diversity of the mucABD locus in Pseudomonas aeruginosa isolates from patients with cystic fibrosis. Microbiology. 2006;152(Pt 11):3261–9. doi: 10.1099/mic.0.29175-0. [DOI] [PubMed] [Google Scholar]
- 43.Diggle S, Cornelis P, Williams P, Cámara M. 4-quinolone signalling in Pseudomonas aeruginosa: old molecules, new perspectives. Int J Med Microbiol. 2006;296(2-3):83–91. doi: 10.1016/j.ijmm.2006.01.038. [DOI] [PubMed] [Google Scholar]
- 44.D’Argenio DA, Wu M, Hoffman LR, Kulasekara HD, Déziel E, Smith EE, et al. Growth phenotypes of Pseudomonas aeruginosa lasR mutants adapted to the airways of cystic fibrosis patients. Mol Microbiol. 2007;64(2):512–33. doi: 10.1111/j.1365-2958.2007.05678.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Pesci E, Iglewski BH. The chain of command in Pseudomonas quorum sensing. Trends Microbiol. 1997;5(4):132–4. doi: 10.1016/S0966-842X(97)01008-1. [DOI] [PubMed] [Google Scholar]
- 46.Hoffman L, Kulasekara HD, Emerson J, Houston LS, Burns JL, Ramsey BW, et al. Pseudomonas aeruginosa lasR mutants are associated with cystic fibrosis lung disease progression. J Cyst Fibros. 2009;8(1):66–70. doi: 10.1016/j.jcf.2008.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Oliver A, Cantón R, Campo P, Baquero F, Blázquez J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science. 2000;288(5469):1251–4. doi: 10.1126/science.288.5469.1251. [DOI] [PubMed] [Google Scholar]
- 48.Montanari S, Oliver A, Salerno P, Mena A, Bertoni G, Tümmler B, et al. Biological cost of hypermutation in Pseudomonas aeruginosa strains from patients with cystic fibrosis. Microbiology. 2007;153:1445–54. doi: 10.1099/mic.0.2006/003400-0. [DOI] [PubMed] [Google Scholar]
- 49.Tian Z, Mac Aogáin M, O’Connor HF, Fargier E, Mooij MJ, Adams C, et al. MexT modulates virulence determinants in Pseudomonas aeruginosa independent of the MexEF-OprN efflux pump. Microb Pathog. 2009;47(4):237–41. doi: 10.1016/j.micpath.2009.08.003. [DOI] [PubMed] [Google Scholar]
- 50.Wiehlmann L, Wagner G, Cramer N, Siebert B, Gudowius P, Morales G, et al. Population structure of Pseudomonas aeruginosa. PNAS. 2007;104(19):8101–6. doi: 10.1073/pnas.0609213104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Tritt A, Eisen JA, Facciotti MT, Darling AE. An integrated pipeline for de novo assembly of microbial genomes. PLoS ONE. 2012;7(9):e42304. doi: 10.1371/journal.pone.0042304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Langille M, Brinkman FS. IslandViewer: an integrated interface for computational identification and visualization of genomic islands. Bioinformatics. 2009;25(5):664–5. doi: 10.1093/bioinformatics/btp030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chaudhuri R, Loman NJ, Snyder LA, Bailey CM, Stekel DJ, Pallen MJ. xBASE2: a comprehensive resource for comparative bacterial genomics. Comp Immunol Microbiol Infect Dis. 2008;31(1):47–62. doi: 10.1016/j.cimid.2007.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Inouye M, Dashnow H, Raven LA, Schultz MB, Pope BJ, Tomita T, et al. SRST2: Rapid genomic surveillance for public health and hospital microbiology labs. Genome Med. 2014;6(11):90. doi: 10.1186/s13073-014-0090-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25(16):2078–9. doi: 10.1093/bioinformatics/btp352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Grant J, Arantes AS, Stothard P. Comparing thousands of circular genomes using the CGView Comparison Tool. BMC Genomics. 2012;13:202. doi: 10.1186/1471-2164-13-202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Roy P, Tetu SG, Larouche A, Elbourne L, Tremblay S, Ren Q, et al. Complete genome sequence of the multiresistant taxonomic outlier Pseudomonas aeruginosa PA7. PLoS ONE. 2010;5(1):e8842. doi: 10.1371/journal.pone.0008842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Wu D, Ye J, Ou HY, Wei X, Huang X, He YW, et al. Genomic analysis and temperature-dependent transcriptome profiles of the rhizosphere originating strain Pseudomonas aeruginosa M18. BMC Genomics. 2011;12:438. doi: 10.1186/1471-2164-12-438. [DOI] [PMC free article] [PubMed] [Google Scholar]


