Growth and development of Arabidopsis is affected by mutation of a member of the RNA recognition motif gene family that is required for RNA editing of more than 40% of the mitochondrial C targets in Arabidopsis.
Abstract
Plant RNA editosomes modify cytidines (C) to uridines (U) at specific sites in plastid and mitochondrial transcripts. Members of the RNA-editing factor interacting protein (RIP) family and Organelle RNA Recognition Motif-containing (ORRM) family are essential components of the Arabidopsis (Arabidopsis thaliana) editosome. ORRM2 and ORRM3 have been recently identified as minor mitochondrial editing factors whose silencing reduces editing efficiency at ∼6% of the mitochondrial C targets. Here we report the identification of ORRM4 (for organelle RRM protein 4) as a novel, major mitochondrial editing factor that controls ∼44% of the mitochondrial editing sites. C-to-U conversion is reduced, but not eliminated completely, at the affected sites. The orrm4 mutant exhibits slower growth and delayed flowering time. ORRM4 affects editing in a site-specific way, though orrm4 mutation affects editing of the entire transcript of certain genes. ORRM4 contains an RRM domain at the N terminus and a Gly-rich domain at the C terminus. The RRM domain provides the editing activity of ORRM4, whereas the Gly-rich domain is required for its interaction with ORRM3 and with itself. The presence of ORRM4 in the editosome is further supported by its interaction with RIP1 in a bimolecular fluorescence complementation assay. The identification of ORRM4 as a major mitochondrial editing factor further expands our knowledge of the composition of the RNA editosome and reveals that adequate mitochondrial editing is necessary for normal plant development.
C-to-U RNA editing in plants is confined to plastid and mitochondrial transcripts (Covello and Gray, 1989; Hiesel et al., 1989), with the exception of C-to-U editing events that have been observed at two sites within nuclear-encoded tRNAs of Arabidopsis (Arabidopsis thaliana; Zhou et al., 2014). A total of 43 Cs in plastids and more than 600 Cs in mitochondria have been reported to be editing sites in Arabidopsis (Chateigner-Boutin and Small, 2007; Bentolila et al., 2013; Ruwe et al., 2013). A large proportion of RNA editing events results in encoding of a different amino acid than the one predicted by the genome sequence, while in other cases, editing creates a start or stop codon (Wintz and Hanson, 1991; Kotera et al., 2005). The editing event usually generates a more conserved amino acid relative to the residue in homologous proteins from other organisms (Gualberto et al., 1989). RNA editing is regarded as a correction mechanism for T-to-C mutations in plastids and mitochondria that would otherwise impair the proper function of gene products, even leading to seedling lethality (Smith et al., 1997; Chateigner-Boutin et al., 2008; Gray, 2012).
RNA is processed by editosomes that are found between the 200 and 400 kD markers on size exclusion columns (Bentolila et al., 2012). Although the composition of the RNA editosome is not yet fully understood, cis- and trans-factors involved in the editing machinery have been identified. Cis-elements are sequences present in close proximity to the C targets (Chaudhuri and Maliga, 1996; Staudinger and Kempken, 2003; Hayes and Hanson, 2007). Members of the PLS subfamily of pentatricopeptide repeat (PPR) motif-containing proteins are trans-factors that specifically recognize cis-elements (Kotera et al., 2005; Zehrmann et al., 2009). The recognition code that specifies how PPR proteins interact with cis-elements was proposed recently based on in silico analysis and interactions were confirmed by RNA-protein binding assays (Barkan et al., 2012; Takenaka et al., 2013; Yin et al., 2013; Okuda et al., 2014). Many editing PPR proteins contain a C-terminal DYW domain (Lurin et al., 2004). The DYW domain exhibits sequence similarity to known cytidine deaminase motifs (Salone et al., 2007). Mutagenesis of conserved deaminase residues in DYW1, QED1, RARE1, OTP84, and CREF7 leads to loss of editing activity (Boussardon et al., 2014; Wagoner et al., 2015; Hayes et al., 2015), indicating the enzyme activity for catalyzing the C-to-U conversion may reside in the DYW domain.
In addition to the PPR proteins, members of the RIP/MORF family were also identified as RNA editing factors in plastids and mitochondria (Bentolila et al., 2012, 2013; Takenaka et al., 2012). RIP1 (for RNA-editing factor Interacting Protein 1) is a major editing factor that controls the editing extent of hundreds of plastid and mitochondrial C targets (Bentolila et al., 2012). Another family involved in RNA editing is called the ORRM (for Organelle RNA Recognition Motif-containing protein) family. The ORRM family is composed of a number of RRM (for RNA Recognition Motif)-containing proteins that are mostly predicted or proven to be organelle-targeted (Sun et al., 2013). The founder of this family, ORRM1, is a plastid editing factor discovered through a database search with the RIP1 protein sequence (Sun et al., 2013). Another plastid editing factor, OZ1 (for Organelle Zinc finger 1), was recently discovered by coimmunoprecipitation with ORRM1 (Sun et al., 2015).
By analyzing the ORRM family, we identified two proteins, ORRM2 and ORRM3, as minor mitochondrial editing factors (Shi et al., 2015). Unlike ORRM1, which contains a truncated RIP-RIP domain at its N terminus and an RRM domain at its C terminus, ORRM2 carries an RRM domain, while ORRM3 contains an N-terminal RRM domain and a C-terminal GR (for Gly-rich) domain. The fact that the RRM domain in ORRM1 and ORRM3 by itself is able to provide the editing activity of the corresponding protein in complementation tests testifies to the critical role of the RRM domain in plastid and mitochondrial editing (Sun et al., 2013; Shi et al., 2015).
Here we report the identification of another member of the ORRM family, encoded by At1g74230, as a major, novel mitochondrial editing factor. The protein, which we named ORRM4 (for Organelle RRM protein 4), contains an RRM domain at its N terminus and a GR domain at its C terminus. The absence of ORRM4 expression caused defective mitochondrial editing at ∼44% of all the mitochondrial sites assayed in this study. ORRM4 is therefore second only to RIP1, which controls the editing extent of 77% of all the mitochondrial sites (Bentolila et al., 2013), in its impact on mitochondrial RNA editing in Arabidopsis. ORRM4 expression is also required for normal plant development, as the orrm4 mutant exhibited slower growth and delayed flowering time. We were able to restore the editing defects in the orrm4 mutant by generating a stable transgenic line expressing the coding region of ORRM4. Our identification of ORRM4 further establishes the ORRM family as an important group in plastid and mitochondrial editing.
RESULTS
Transient Silencing of ORRM4 Causes Mitochondrial Editing Defects
To characterize the function of ORRM4 in RNA editing, we performed virus-induced gene silencing (VIGS) to transiently silence ORRM4 in Arabidopsis seedlings expressing GFP under a 35S promoter. GFP is used as a selective marker to track the silencing effectiveness. Two-week-old seedlings were inoculated with Agrobacterium transformed with a GFP-silencing construct, or with Agrobacterium carrying an ORRM4/GFP cosilencing construct. No morphological defects were observed upon the transient silencing of ORRM4. Editing extent in uninoculated plants, GFP-silenced plants, and ORRM4/GFP cosilenced plants was measured by a strand- and transcript-specific RNA-seq method (STS-PCRseq; Bentolila et al., 2013; Shi et al., 2015; Supplemental Dataset S1). The two biological replicates assayed for each sample were obtained from different plants grown and treated under the same conditions (Supplemental Fig. S1).
We performed quantitative reverse transcription PCR (qRT-PCR) to assay the reduction of ORRM4 expression level. The RNA expression level of ORRM4 is reduced to ∼16% compared to the uninoculated and the GFP-silenced controls (Fig. 1A). The specificity of the silencing experiment is validated by the fact that the expression level of ORRM4 is not significantly reduced in the ORRM2- and ORRM3-silenced plants (Fig. 1A). Out of the 564 mitochondrial editing sites analyzed, we found that the transient silencing of ORRM4 leads to a significant reduction of editing extents in both biological replicates at 51 mitochondrial sites (Δ ≥ 10%, P < 1.6e-6), which constitutes 9% of the total of the mitochondrial sites assayed (Fig. 1B; Supplemental Table S1). No significant plastid editing defects were observed in both silenced biological replicates. A previous study demonstrated that ORRM4 is localized to mitochondria in a GFP assay (Vermel et al., 2002).
Figure 1.
Transient silencing of ORRM4 causes mitochondrial editing defects. A, Relative RNA expression level of ORRM4 measured by qRT-PCR. B, Twenty-two editing sites that experienced a significant decrease of editing extent ≥ 10% upon the silencing of ORRM4. Editing extents were measured by STS-PCRseq. Two biological replicates were used in each silencing experiment. Not inoculated, Plants not inoculated with Agrobacterium; GFPsil, plants inoculated with Agrobacterium carrying a GFP-silencing construct; ORRM2sil, plants inoculated with Agrobacteria harboring an ORRM2/GFP cosilencing construct; ORRM3sil, plants inoculated with Agrobacteria harboring an ORRM3/GFP cosilencing construct; ORRM4sil, plants inoculated with Agrobacterium carrying an ORRM4/GFP cosilencing construct.
ORRM4 Mutation Recapitulates Mitochondrial Editing Defects Observed in ORRM4-Silenced Plants
In order to verify the involvement of ORRM4 in mitochondrial RNA editing, we obtained a T-DNA insertional line SALK_023061C in Columbia background from the ABRC stock center. The orrm4 mutant contains a T-DNA insertion in an intron separating the 5′UTR from the first exon of ORRM4 (Fig. 2A). It is a knockout mutant since the ORRM4 expression is decreased to an undetectable level measured by qRT-PCR (Fig. 2B). We performed STS-PCRseq to examine the editing extent of sites affected in the orrm4 mutant. The orrm4 mutation results in a significant reduction (Δ≥10%, P < 1.6e-6) of editing extent at 270 mitochondrial sites (Supplemental Table S3), 44% of the total mitochondrial sites assayed. A total of 96% of the sites identified by transient silencing of ORRM4 also showed defects in the orrm4 mutant (Supplemental Table S1). For example, at sites rpl5 C58, rpl5 C47, rps4 C88, and rpl5 C92, decreased editing extent was observed in both ORRM4-silenced plants and the orrm4 mutant compared to the controls (Fig. 2C). However, the effect on editing resulting from the orrm4 mutation is more drastic than that caused by ORRM4 silencing, as more sites are affected and editing extent is more intensely decreased in orrm4 mutant plants compared to the ORRM4-silenced plants (Fig. 2C; Supplemental Tables S1 and S3). This result can be explained by the absence of ORRM4 expression in the orrm4 mutant, while some ORRM4 expression occurs in the silenced tissue (Figs. 1A and 2B).
Figure 2.
ORRM4 gene model, RNA expression level, and editing extents in the orrm4 mutant. A, Gene structure of ORRM4. Triangle indicates the locus of the T-DNA insertion. Dashed box shows the gene-specific region selected for the VIGS experiment. B, Relative ORRM4 expression level measured by qRT-PCR. ORRM4 expression is reduced to an undetectable level in orrm4 homozygous mutants relative to wild-type plants (n = 3). ***, Significance of P < 0.001. C, ORRM4 mutant plants recapitulate the mitochondrial editing defects in the ORRM4-silenced plants assayed by STS-PCRseq. Not inoculated, Plants not inoculated with Agrobacterium (n = 2); GFP_sil, plants inoculated with Agrobacterium carrying a GFP-silencing construct (n = 2); wt, wild-type Arabidopsis (n = 2); ORRM4sil_1, one plant inoculated with Agrobacterium carrying an ORRM4/GFP cosilencing construct; orrm4, orrm4 homozygous mutant plants (n = 2).
Surprisingly, five plastid sites among the 38 sites assayed show a significant reduction of editing extent in both biological replicates of the orrm4 mutant plant (Supplemental Table S4). Given the mitochondrial localization of ORRM4 and the massive effect of the mutation on mitochondrial editing, it is likely that this effect is indirectly caused by altered mitochondrial function. Alternatively, the genetic background of the wild-type plant to which we are comparing the orrm4 mutant plants might have resulted in detection of different levels of plastid editing at the 5 sites. The insertional T-DNA orrm4 mutant plant is distributed as a homozygous plant; its genetic background is Columbia so that we used a generic Columbia wild-type as control. However, very slight differences might exist between the unavailable Columbia accession used to generate the orrm4 mutant and the one we used as control. Divergence between the mutant line’s wild-type sibling and our wild-type line could also explain the occurrence of the two mitochondrial sites that were significantly reduced in ORRM4-silenced plants but not in orrm4 mutant plants (Supplemental Table S1).
orrm4 Mutation or Silencing Affects Editing in a Site-Specific Way, Even When All the Sites on a Transcript Are Reduced in Their Editing Extent
Many sites affected in the orrm4 mutant occur together on a few transcripts (Fig. 3, A and B). An editing factor can possibly affect editing extent in either a site-specific or a transcript-specific manner, or both ways. To examine how ORRM4 affects editing, we represented the proportion of sites controlled by ORRM4 based on the transcripts where they are located and on the complex to which the transcripts belong (Fig. 3A).
Figure 3.
orrm4 mutation or silencing affects editing extent in a site-specific way. A, Proportion of sites affected by orrm4 mutation on each transcript. Each bar represents a transcript color-coded according to the complex to which it belongs. B, The decreased editing and the alteration of editing extents of sites on the rpl5 transcript measured by STS-PCRseq. C, The decreases in editing caused by ORRM4 silencing are highly correlated with the decreases caused by the orrm4 mutation on the rpl5 transcript. D, Editing at site nad4 C1101 is significantly reduced while editing at two sites next to it is not. E, Reduction of editing is observed at site ccmFc C160, but not at ccmFc C155. Not inoc, Plants not inoculated with Agrobacterium (n = 2); GFP_sil, plants inoculated with Agrobacterium carrying a GFP-silencing construct (n = 2); wt, wild-type Arabidopsis (n = 2); ORRM4sil_1, one plant inoculated with Agrobacterium carrying an ORRM4/GFP cosilencing construct; orrm4 mutants (n = 2), Δorrm4 = (% editing of wt − % editing of orrm4)/% editing of wt; ΔORRM4sil_1 = (% editing of Not inoc − % editing of ORRM4sil_1)/% editing of Not inoc.
A total of 262 sites showed decreased editing in the orrm4 mutant and are distributed on 31 mitochondrial gene transcripts or 43% of the sites assayed (orf114 was not included in this analysis; Fig. 3A). Three transcripts showed a reduction of editing extent at all the sites assayed, whereas the proportion of sites affected at 10 transcripts fell into a range from 60% to 90% (Fig. 3A). Eight transcripts showed a reduction of editing extent between 20% and 60% of sites, while 10 transcripts exhibited a reduction of editing extent on less than 20% of their sites (Fig. 3A). Among this latter group, three transcripts did not exhibit reduction of editing extent at any of their sites.
The proportion of reduced edited sites/transcript varies widely between and inside the mitochondrial complexes, with the transcripts encoding the ribosomal proteins being the most affected by the orrm4 mutation, followed by the cytochrome c biogenesis transcripts (Fig. 3A). Among this latter group, the ccmFc transcript is an outlier, as its proportion of reduced edited sites falls below 20% while the proportion of all the remaining ccm transcripts is above 60%. Likewise, two groups can be easily observed in the complex I transcripts: one group that is highly affected by orrm4 mutation is comprised of nad3, nad4L, and nad9, and a second group that is more weakly affected is comprised of the remaining nad transcripts. In summary, even though all Cs targets on some transcripts are affected by orrm4 mutation, other transcripts vary in the proportion of sites affected.
All the known editing sites on the rpl5 transcript were affected by mutation or silencing of ORRM4 (Fig. 3B, top; Supplemental Tables S1 and S3). However, the decrease in editing extent in rpl5 transcripts shows some site specificity, as it is not constant along the transcript and varies from 18% to 78% in the orrm4 mutant and from 14% to 63% in the ORRM4-silenced tissue (Fig. 3B, bottom). The alteration in editing extent of sites on rpl5 caused by orrm4 mutation correlates well with that caused by the silencing of ORRM4 (Fig. 3C). Likewise, all the known editing sites on the nad4L transcript are affected by mutation or silencing (Fig. 3A; Supplemental Fig. S2A; Supplemental Tables S1 and S3). As observed for rpl5, the reduction of editing extent along the nad4L transcript varies from 13% to 35% and from 8% to 15% in the orrm4 mutant and in the ORRM4-silenced tissue, respectively (Supplemental Fig. S2B). The reduction of editing extent on nad4L sites caused by the orrm4 mutation or its silencing are highly correlated (R2 = 0.89, Supplemental Fig. S2C). The reduction of editing extent at transcripts such as rpl5 and nad4L occurs on every editing site; however, the magnitude of the reduction is site specific as some sites experience a much more pronounced decrease than the others.
The location of sites affected by the knockdown or knockout of ORRM4 is dispersed in location on certain transcripts. For example, editing at site nad4 C1101 and site ccmFc C160 are significantly reduced, but the sites around them are not (Fig. 3, D and E; Supplemental Dataset S1; Supplemental Table S3).
orrm4 Mutation Affects the Abundance of Certain Transcripts Independently of Its Effect on Editing Extent
A possible reason for the strong effect of orrm4 mutation on all editing events located on the rpl5 transcript could be an alteration in the total abundance of rpl5 transcripts. We therefore examined the transcript abundance of two groups of transcripts showing either a strong (rps4 and rpl5) or a slight effect (nad4 and nad7) toward orrm4 mutation on their editing extent. Like rpl5, rps4 shows a reduction of editing extent for the majority of its sites in the orrm4 mutant, whereas less than 10% of the sites on nad4 and nad7 transcripts are affected by the orrm4 mutation (Fig. 3A). RNA blot hybridization indicates that the orrm4 mutation results in significantly decreased transcript abundance only for nad4 and rps4 when compared to the wild-type (P < 0.01), but not for nad7 or rpl5 transcripts (Fig. 4). The effect of orrm4 mutation on transcript abundance does not match with its effect on editing extent, as illustrated by the transcripts surveyed. Like rps4, nad4 exhibits a clear and significant reduction of its transcript abundance; but unlike rps4, nad4 shows very little effect on the editing extent of its sites (8% vs. 86%, respectively; Fig. 3A). Furthermore, both rps4 and rpl5 belong to the group with more sites controlled by ORRM4. The abundance of rpl5 transcripts is similar in the orrm4 mutants versus the wild-type, whereas rps4 shows a most pronounced reduction of transcript abundance in the orrm4 mutant (Fig. 4). Thus, our data are consistent with an effect of the orrm4 mutation on the abundance of transcripts, as detected by RNA blots, but this effect does not correlate with the effect on editing extent.
Figure 4.
orrm4 mutation can affect the transcript abundance. A, RNA blots hybridized with nad4, nad7, rps4, or rpl5 probes; wt, three wild-type plants; orrm4−/−, three homozygous mutant plants. The negative of the ethidium bromide staining gel in the bottom panel serves as a loading control. B, Quantification of the transcript abundance visualized in the RNA blots and normalized to the amount of RNA loaded demonstrates a significant reduction for nad4 and rps4 (**P < 0.01), but not for nad7 or rpl5.
Reduced Mitochondrial Editing Results in Slow Growth and Late Flowering
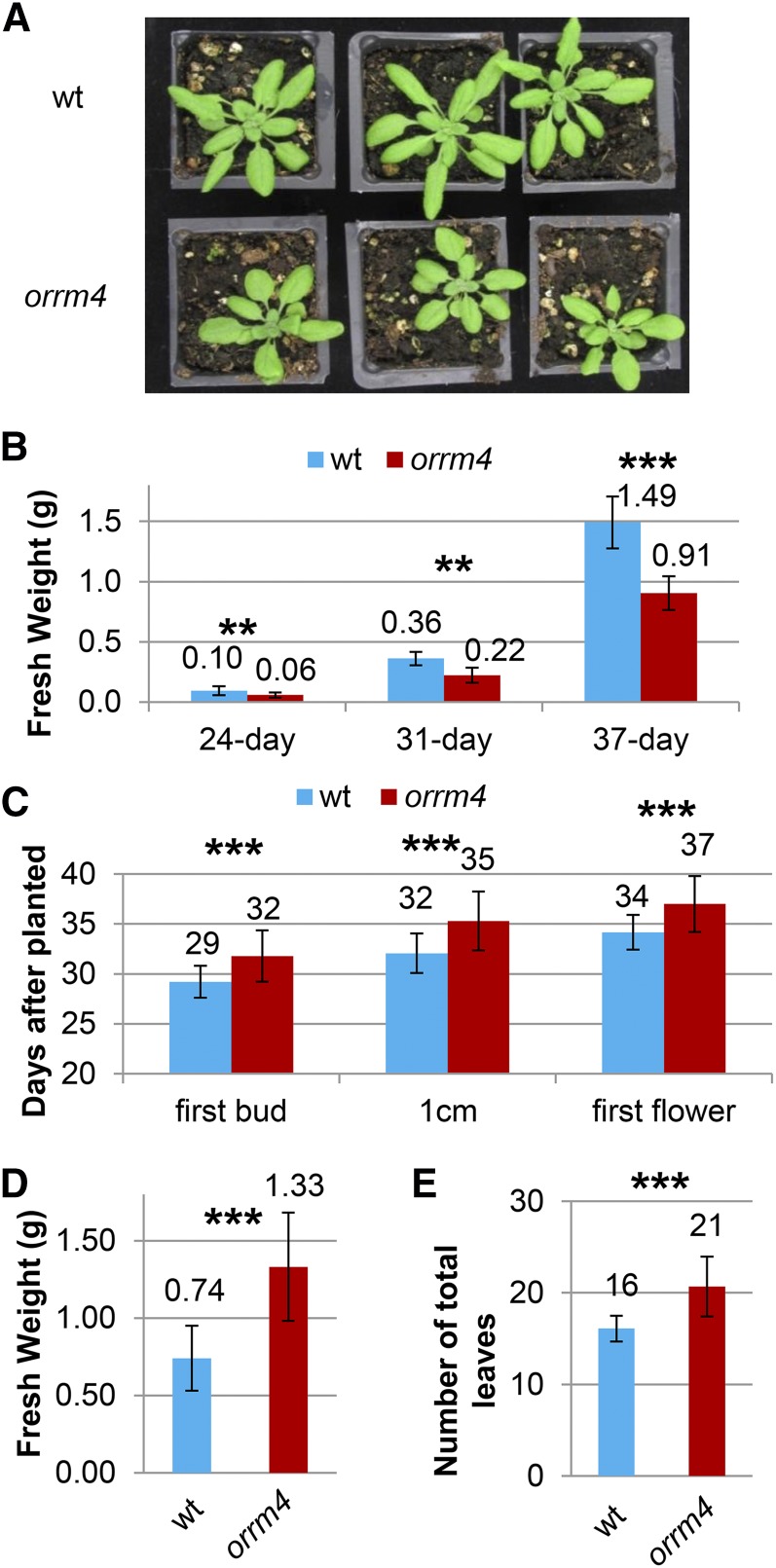
As shown in Figure 5A, the orrm4 homozygous mutant grows slower than the wild-type Arabidopsis ecotype Columbia. orrm4 mutants have lower fresh weight than the wild-type plants grown under long-day conditions (14 h of light/day) at the three time points that were measured (Fig. 5B). In order to examine the flowering phenotype of the mutant, we examined three flowering time-related traits. The results indicate that it took the mutants three more days on average for their first flower bud to become visible in the center of the rosette, for theirs inflorescence stems to reach 1 cm in height, and for their first flower to open in contrast to the wild-type (Fig. 5C). To determine whether the slow-growth phenotype is the cause of the late-flowering phenotype, we recorded the fresh weight and the number of total leaves of the mutant versus the wild type when the first flower opened. As shown in Figure 5, D and E, orrm4 mutant plants have higher fresh weight and number of total leaves than the controls when their first flower opened. The fact that the mutant requires a greater mass than the wild-type to flower indicates that the late-flowering phenotype is not solely a consequence of its slow-growth phenotype.
Figure 5.
orrm4 mutant plants show a slow-growth and late-flowering phenotype. A, Plant growth phenotype of orrm4 homozygous mutants (bottom) and wild-type plants (top) grown at 14 h of light per day for 29 d. B, Fresh weight of orrm4 homozygous mutants and wild-type plants grown at 14 h of light per day for 24 d, 31 d, and 37 d, respectively (n = 6). C, orrm4 homozygous mutants flower late compared to the wild-type plants. First bud, Days until visible flower buds in the center of the rosette; 1 cm, days until inflorescence stem reached 1 cm in height; first flower, days until first open flower (n = 30). D and E, The fresh weight and the number of total leaves of the orrm4 mutants versus the wild-type plants at the opening of the first flower (n = 30). ** P < 0.01, ***P < 0.001.
Stable Overexpression of ORRM4 in the orrm4 Mutant Background Complements the Mitochondrial Editing Defects
To verify that the mitochondrial editing defects in the orrm4 mutant are truly caused by the absence of ORRM4 expression, we transformed orrm4 homozygous mutants with a construct expressing ORRM4 driven by a 35S promoter. We performed genotyping of plants that survived in the Basta selection to verify their homozygosity for the orrm4 T-DNA insertion allele and the presence of the ORRM4 transgene. Afterward, we collected the seeds from one transgenic line carrying the transgene in the orrm4 mutant background and cultured them under long-day conditions for 4 weeks. Eleven segregating T1 plants were genotyped, and their editing extents were measured by poisoned primer extension (PPE) assays. Our genotyping data indicates that three plants did not carry any ORRM4 transgene, while eight plants carried one or two copies of the transgene. Two wild-type plants were grown under the same conditions, and their editing extents were also assayed. Compared to the homozygous mutants without the transgene, all plants expressing ORRM4 showed restoration of editing (Fig. 6). At site rpl5 C59, the editing extent increased in some transgenic lines to ∼50% or even to ∼65% relative to ∼40% in the mutant plants without the transgene (Fig. 6A). Likewise, editing extent at site rps4 C235 increases from ∼30% to ∼55% or ∼80% (Fig. 6B). Mitochondrial editing defects in the orrm4 mutant can be restored to a wild-type level or even higher.
Figure 6.
Stable expression of ORRM4 driven by a 35S promoter in orrm4 homozygous mutants restores the editing defects caused by the orrm4 mutation. A, Editing of rpl5 C59 assayed by PPE. B, Editing of rps4 C235 assayed by PPE. C, Editing of cox2 C138 assayed by PPE. orrm4−/−, Three plants from the T1 segregating population that do not contain the ORRM4 transgene; orrm4−/− w/ 35S::ORRM4, plants from the T1 segregating population that carry one or two copies of ORRM4 transgene driven by a 35S promoter in the orrm4 mutant background; wt, wild-type plants. Editing extents are shown as dark orange bars in complemented lines with excess editing of rpl5 C59 and rps4 C235 and decreased editing of cox2 C138. D, ORRM4 expression measured by qRT-PCR. The level of ORRM4 expression was normalized to the average detected in wild-type plants. The average expression in wild-type plants was arbitrarily chosen to be 1 (n = 3, log10 scale).
In order to test whether expression of ORRM4 increased editing not only in sites affected in the mutant, but also had any effect on other sites, we measured the editing extent of a site that was not affected in the orrm4 mutant by a PPE assay. As shown in Figure 6C, the editing extent of cox2 C138 in orrm4 mutants were unchanged from the wild-type level. However, three out of the eight transgenic plants carrying the ORRM4 transgene exhibited an unexpected decrease of editing from ∼85% to ∼70% (Fig. 6C). These three plants corresponded to the ones showing a higher extent of complementation at sites rpl5 C59 and rps4 C235 than the wild-type level (Fig. 6, A and B). The measurement of ORRM4 expression by qRT-PCR indicates that these three plants possess the highest level of ORRM4 expression (Fig. 6D), probably because they contain two copies of the transgene. Observation of the ORRM4 expression level clearly divides the transgenic lines into two groups: three plants presumably homozygous in ORRM4 transgene and five plants hemizygous for the ORRM4 transgene have an average ORRM4 expression that is ∼14 times and 5 times the wild-type level, respectively (Fig. 6). The ORRM4 expression level correlates well with the increase or decrease of editing extents as shown in Figure 6.
Because we found an effect of orrm4 mutation on the abundance of nad4 and rps4 transcripts, we measured the abundance of these transcripts in the complemented lines. As seen in Supplemental Figure S3, the presence of the transgene is able to significantly increase the abundance of nad4 and rps4 transcripts to close to or at wild-type levels (P < 0.01 and P < 0.001, respectively). The abundance of the nad7 transcript is increased, but not significantly, in the complemented plants, while the amount of rpl5 is invariant (Supplemental Fig. S3). This latter result further supports an absence of clear relationship between the effect of ORRM4 on transcript abundance and editing extent, as the restoration of rpl5 editing extent in complemented lines is not accompanied by a change in transcript abundance.
ORRM4 Interacts with RIP1
Both RIP1 and ORRM4 have a major impact on mitochondrial editing, as decreased expression of either result in editing defects at more than 250 mitochondrial sites. We consolidated the sites affected in either rip1 mutants or orrm4 mutants by subdividing them based on the transcripts where they are located. After ruling out the outliers, which are cox2, nad6, and genes encoding complex V subunits, we found that the proportion of sites per gene affected by rip1 mutation correlates well with those affected by the orrm4 mutation (Fig. 7A). Additionally, we compiled the data from this study with previous data (Bentolila et al., 2013) to determine which sites are affected by rip1, rip3, or orrm4 mutations jointly or independently of each other (Supplemental Dataset S2). A total of 253 sites, or 95% of the sites affected in the orrm4 mutant, are also affected in the rip1 mutant (Fig. 7B). As a comparison point, 88% of the sites affected in the rip3 mutant are also affected in the rip1 mutant (Supplemental Dataset S2; Fig. 7B). The large overlap of mitochondrial sites controlled by RIP1 and RIP3 is supported by a physical interaction between these two proteins, as demonstrated by yeast two-hybrid (Y2H), pulldown, and bimolecular fluorescence complementation (BiFC) assays (Zehrmann et al., 2015).
Figure 7.
The genetic interaction between ORRM4 and other mitochondrial editing factors. A, The correlation between percent of reduced sites per gene affected in the orrm4 mutant versus that in the rip1 mutant. B, Number of mitochondrial sites affected in rip1, rip3, or orrm4 mutants. C, Number of mitochondrial sites affected in ORRM2-silenced, ORRM3-silenced, or orrm4 mutant plants. Numbers in overlap region indicates the sites under the control of both/all. Number outside the circles shows the sites that are not affected by any of the three mutations.
To test whether the genetic interaction between RIP1 and ORRM4 is dependent on a physical interaction between their encoded products, we performed BiFC assays by transiently coexpressing a protein fused to the N-terminal half of GFP (GFPN) with another protein fused to the C-terminal half of GFP (GFPC) in Nicotiana benthamiana. No interaction would be expected between ORRM4 and MEF9, as editing at site nad7 C200 was impaired only in the mef9 mutant (Takenaka, 2010) and not in the orrm4 mutant (Supplemental Table S3). As predicted, no signal was observed when ORRM4-GFPN was coexpressed with MEF9-GFPC, or in the reciprocal pair MEF9-GFPN with ORRM4-GFPC (Fig. 8, A and B). In contrast, positive protein-protein interaction signals were observed when ORRM4-GFPN and RIP1-GFPC were coinoculated (Fig. 8C). Also, RIP1-GFPN interacts with RIP1-GFPC (Fig. 8D). In our Y2H assays, the predicted mature coding sequence (with the predicted transient peptides removed) was fused to the AD or BD domain. However, the interaction between ORRM4 and RIP1 is not informative in yeast, due to the auto-activation of both when fused to the BD domain.
Figure 8.
ORRM4 interacts with RIP1, ORRM3, and ORRM4 when transiently coexpressed in N. benthamiana leaves by BiFC assays. Each confocal image shows the merge of GFP signal (green) and chlorophyll autofluorescence (red). Scale bar (white line) represents 20 μm. White arrowheads point to the GFP signals indicating the interaction. A and B, ORRM4 does not interact with MEF9 serving as a negative control. C, ORRM4 interacts with RIP1. D, RIP1 forms homo-dimers. E, ORRM4 forms hetero-dimers with ORRM3. F, ORRM4 forms homo-dimers.
ORRM4 Interacts with ORRM3 and Itself
When we compared the editing sites affected by the silencing of ORRM3 (Shi et al., 2015) and those affected by the orrm4 mutation, we found that 15 sites are affected in both cases (Fig. 7C). These 15 sites constitute 47% of all the sites affected by the silencing of ORRM3. Likewise, about 54% of sites affected in ORRM2-silenced plants are also affected in the orrm4 mutant (Fig. 7C). Therefore, it is possible that ORRM4 interacts with ORRM2 and ORRM3. We performed Y2H and BiFC assays to test this hypothesis. Interaction between ORRM4 and ORRM3 was observed in both BiFC and Y2H assays (Figs. 8E and 9A). Our result also demonstrates that ORRM4 forms homo-dimers (Figs. 8F and 9B) but does not interact with ORRM2 in yeast (Fig. 9C). To examine the interaction between ORRM4 and PPR proteins, we selected MEF1 and MEF20, since decreased editing at site rps4 C956 was observed by either mef1 or orrm4 mutation while editing at site rps4 C226 is impaired in both mef20 and orrm4 mutants (Takenaka et al., 2010; Zehrmann et al., 2010). The results indicate that ORRM4 does not interact with MEF1 or MEF20 in Y2H (Fig. 9D).
Figure 9.
In the Y2H assay, ORRM4 interacts with ORRM3 and itself, and its C-terminal GR domain is required for the interaction. A, ORRM4 forms hetero-dimers with ORRM3. B, ORRM4 forms homo-dimers. C, ORRM4 does not interact with ORRM2 in yeast. D, ORRM4 does not interact with MEF1 or MEF20 in yeast. E, The C-terminal GR domain of ORRM4 is required for its interaction with itself, whereas its N-terminal RRM domain is not. F, The N-terminal RRM domain of ORRM3 interacts with the C-terminal GR domain of ORRM4. EM, Yeasts transformed with a vector carrying an empty GW cassette as a negative control.
ORRM4 contains an RRM domain at the N terminus and a GR domain at the C terminus (Fig. 10A). To further probe the role of different domains of ORRM4 in protein-protein interactions, we also performed Y2H assays between truncated proteins carrying either the N-terminal or C-terminal regions of the ORRMs. Our data indicate that the C-terminal GR domain of ORRM4 is indispensable for it to form a homo-dimer, while the N-terminal RRM domain is not (Fig. 9E). Likewise, the C-terminal GR of ORRM4, but not the N-terminal RRM domain, is required for its interaction with ORRM3 (Fig. 9F). The C-terminal GR domain of ORRM4 interacts with the N-terminal RRM domain of ORRM3 (Fig. 9F).
Figure 10.
The N-terminal RRM domain of ORRM4 is able to rescue RNA editing activity in the orrm4 mutant, whereas the C-terminal GR domain does not. A, The motif diagram of ORRM1, ORRM2, ORRM3, and ORRM4. B, Constructs used for transformation. nORRM4, Amino acids 1 to 112; cORRM4, amino acids 113 to 290; mtp, amino acids 1 to 29 of the nuclear-encoded coxIV from yeast. C, Editing of rps4 C77 and rpl5 C59 assayed by PPE. D, Editing of rps4 C77, rps4 C235, and rpl5 C59 assayed by PPE. wt, Wild-type plants (n = 2); orrm4 homozygous mutants (n = 2); orrm4 w/ 35S::nORRM4, orrm4 homozygous mutants transformed with the N-terminal RRM domain of ORRM4 under a 35S promoter; orrm4 w/ 35S::mtp-cORRM4, orrm4 homozygous mutants transformed with the C-terminal GR domain of ORRM4 under the control of a 35S promoter and an mtp. E, ORRM4 expression measured by qRT-PCR. The level of ORRM4 expression was normalized to the average detected in wild-type plants. The average expression in wild-type plants was arbitrarily chosen to be one (two plants assayed for wild type and orrm4; no. of technical replicates, three). The expression of the two constructs was measured with two different set of primers.
Stable Overexpression of the N-Terminal RRM Domain of ORRM4 Restores the Editing Defects in the orrm4 Mutant, While the C-Terminal GR Domain Only Has a Minor Effect on Editing
Given that the RRM domain of ORRM3 is required for its editing activity in mitochondria (Shi et al., 2015), we hypothesized that the RRM domain in ORRM4 might also rescue the editing defects caused by the orrm4 mutation. Thus, we transformed homozygous orrm4 mutants with two different transgenes. One construct expresses the N-terminal RRM domain of ORRM4 under the control of a 35S promoter (Fig. 10B). The other transgene expresses a mitochondrial transit peptide fused to the C-terminal GR domain of ORRM4 driven by a 35S promoter (Fig. 10B).
All the transgenic lines were morphologically normal. We selected three independent transgenic plants from the T0 generation of plants transformed with the N-terminal region of ORRM4, and six plants from the T0 generation of plants transformed with the mitochondrial transit peptide fused to the C-terminal ORRM4. Mutants expressing the N-terminal part of ORRM4 showed increased editing extent at the sites we monitored by PPE assays compared to the mutants without the transgene, whereas mutants expressing the C terminus of ORRM4 only exhibited a minor change of editing extent (Fig. 10C). For instance, editing extent at site rps4 C77 varies from 62% to 87% when the N-terminal portion of ORRM4 is expressed, indicating that editing can be recovered to a wild-type level. However, in mutants expressing the C terminus of ORRM4, the editing extent is altered only from 44% to 56% (Fig. 10C). The restoration of editing upon the expression of the N-terminal portion of ORRM4 was also observed at site rps4 C235 (Fig. 10C). In order to determine whether the increase of editing extent in the complemented lines is site-specific, we analyzed the editing extent of a site that was not affected in the orrm4 mutant. We found that editing at site cox2 C138 was not affected in any of the transgenic lines assayed; it was not decreased as has been observed previously for the three T1 plants transformed with the full length ORRM4 (Figs. 6C and 10D).
Examination of ORRM4 expression by qRT-PCR in the complemented lines shows a correlation between the editing restoration and the level of ORRM4 expression in the mutants expressing the N-terminal part of ORRM4 (nORRM4; Fig. 10E). The highest nORRM4-expressing transgenic plant shows also the highest level of editing restoration, while the lowest nORRM4-expressing transgenic plant shows the lowest level of editing restoration (Fig. 10, C and E). Given that the expression of nORRM4 in two of the transgenic plants reaches a comparable level as the one observed in transgenic T1 plants expressing the full length ORRM4 without a decrease in editing extent at site cox2 C138 (Figs. 6, C and D, and 10, D and E), we can conclude that the maintenance of wild-type levels of cox2 C138 editing might result from the absence of the C-terminal part of ORRM4.
The level of expression of the C-terminal GR domain of ORRM4 in transgenic lines with a range of 30× to 90× of the wild type demonstrates that the inability of this portion of ORRM4 to restore editing is not due to a faulty expression of the transgene (Fig. 10E). A level of nORRM4 expression that is 10 times higher than wild type is sufficient to restore editing extent in two of the three sites surveyed whereas a level of cORRM4 that is 30 times higher than wild type does not restore editing extent in the same two sites (Fig. 10, C and E). Thus, our data suggest that the editing ability of ORRM4 is mostly conferred by its RRM motif.
Stable Overexpression of ORRM3 Restores the Editing Defects in the orrm4 Mutant
Because ORRM3 and ORRM4 are both mitochondrial editing factors sharing high sequence similarity, particularly in the RRM domain (48% identical at the whole protein level, 70% identical in the RRM domain; Supplemental Fig. S4), we wondered whether we could compensate the editing default in the orrm4 mutant by overexpressing ORRM3. We transformed orrm4 homozygous mutants with a construct expressing ORRM3 driven by a 35S promoter (Fig. 11A). After selection and genotyping by PCR, transgenic plants along with two wild-type plants and two orrm4 homozygous mutant plants were grown and assayed for editing extent at three sites by PPE. Editing at site rps4 C77 and rpl5 C59 is controlled by both ORRM3 and ORRM4, whereas editing at site cox2 C138 is controlled by neither (Fig. 1A; Supplemental Tables S1 and S3; Shi et al., 2015). As seen in Figure 11B (left) at site rps4 C77, ORRM3 is able to restore editing in the orrm4 mutant up to the wild-type level in one T0 plant and even higher for another T0 plant. This latter plant also shows a transgressive editing extent at site rpl5 C59. Survey of ORRM3 expression by qRT-PCR indicates that the transgenic plant showing the highest editing extent also possesses the highest level of ORRM3 expression among the transgenic plants (110 times the wild-type level; Fig. 11C). As observed previously with certain T1 plants transformed with ORRM4 under the control of the 35S promoter (Fig. 6C), the level of editing extent is decreased at site cox2 C138 for the two plants exhibiting the highest level of ORRM3 expression (Fig. 11, B and C). This phenomenon occurs here at a much higher level of expression of the transgene ∼100 times the wild-type level for ORRM3 vs. ∼10 times the wild-type level for ORRM4 (Figs. 6D and 11C).
Figure 11.
Stable expression of ORRM3 or the N-terminal RRM domain of ORRM3 driven by a 35S promoter is able to rescue RNA editing activity in the orrm4 mutant. A, Constructs used for transformation. nORRM3, Amino acids 1 to 120; ORRM3, the coding region of ORRM3. B, Editing of rps4 C77, rpl5 C59, and cox2 C138 assayed by PPE. Left, Editing in mutant plants transformed with the full-length ORRM3; right, editing in mutant plants transformed with the N-terminal RRM domain of ORRM3; wt, wild-type plants (n = 2); orrm4 homozygous mutants (n = 2); orrm4 w/ 35S::ORRM3, orrm4 homozygous mutants transformed with the coding region of ORRM3 under a 35S promoter; orrm4 w/ 35S::nORRM3, orrm4 homozygous mutants transformed with the N-terminal RRM domain of ORRM3 under a 35S promoter. C, ORRM3 expression measured by qRT-PCR. The level of ORRM3 expression was normalized to the average detected in wild-type plants. The average expression in wild-type plants was arbitrarily chosen to be one (n = 3).
We also tested the ability of the N-terminal RRM domain of ORRM3 driven by a 35S promoter to complement the editing defects of the orrm4 mutant. orrm4 homozygous mutants were transformed with a construct expressing the N-terminal RRM domain of ORRM3 driven by a 35S promoter (Fig. 11A). As seen in the right panel of Figure 11B, one of the transgenic T0 plants is restored to wild-type level for the editing extent at site rpl5 C59. The same plant exhibits a pronounced increase in editing extent at the other site analyzed, rps4 C77. This result is somewhat similar to the experiment performed previously with N-terminal RRM domain of ORRM4 and further supports the RRM domain as being necessary for the editing ability of the protein.
DISCUSSION
In this study, we demonstrate that ORRM4 is a major mitochondrial editing factor that controls about 44% of mitochondrial sites in Arabidopsis. Impaired RNA editing was observed in both the orrm4 mutant and the ORRM4-silenced tissue. Stable overexpression of the ORRM4 coding region complements the editing defects in orrm4 homozygous mutants, further confirming ORRM4 as a mitochondrial editing factor.
ORRM4 resembles ORRM3 in many aspects. They have high similarity in gene structure and domain arrangement. As mitochondrial editing factors, both of them interact with RIP1 and form homo-dimers. Nevertheless, ORRM4 is a very different editing factor than ORRM3, as it has a more general impact on mitochondrial editing. The transient silencing of ORRM3 resulted in decreased editing extent at 32 mitochondrial sites, with one to five sites affected on each transcript (Shi et al., 2015). Even though the realm of influence of ORRM3 is underestimated, as ORRM3 expression is only reduced but not eliminated in the silenced tissue we analyzed, it is clear that the effect of ORRM3 on mitochondrial editing is site-specific and more limited than the effect of ORRM4 (Shi et al., 2015). orrm4 mutation has a more significant impact on mitochondrial editing; it affects the editing of 270 mitochondrial sites. Among all the mitochondrial editing factors known today, only RIP1 has been reported to affect more sites (Bentolila et al., 2012). However, unlike RIP1, whose defective expression resulted in an absence of detectable editing extent for ∼100 mitochondrial sites, residual editing extent was observed for all the sites under the control of ORRM4 in the mutant plant. Functional redundancy between mitochondrial ORRMs could prevent the elimination of editing in sites under the control of ORRM4 in the mutant plant. We have shown previously that this hypothesis was plausible for ORRM2 and ORRM3 (Shi et al., 2015). Silencing of ORRM2 in orrm3 mutant plants further impaired the editing extent of sites under the control of both ORRM2 and ORRM3.
The sites under the impact of ORRM4 are sometimes clustered together on particular transcripts, which was not observed when ORRM3 was silenced. However, evidence still supports ORRM4 as a site-specific, not merely transcript-specific, editing factor. First, even though sites affected by orrm4 mutation sometimes encompass the entire transcript, the reduction in editing extent at different sites is variable. Second, orrm4 mutation does not always affect all the editing sites on one transcript. In 90% of cases (28 out of 31 gene transcripts), reduced editing extent was only observed in a fraction of the editing sites, and the distribution of these sites is not contiguous. It is therefore unlikely that such specificity could be explained by general effects such as transcript stability, even though changes in transcript stability are expected to affect editing extent (Lu and Hanson, 1992). While orrm4 mutation affects the abundance of certain transcripts, our data clearly demonstrate an absence of relationship between the effect of orrm4 mutation on editing extent and transcript abundance. rpl5 transcript abundance is not reduced in the orrm4 mutant while its editing extent is decreased in all the editing sites. Restoration of rpl5 editing extent in orrm4 mutants expressing ORRM4 is not accompanied by a change in its abundance. Finally, we observed a direct interaction between ORRM4 and the mitochondrial editing factors RIP1and ORRM3, which further implicates ORRM4 as a component of the mitochondrial editosome. The interaction between ORRM4 and ORRM3 was shown to be mediated by the C-terminal GR domain of ORRM4. NMR studies of NtGR-RBP1, which contains an N-terminal RRM domain and a C-terminal GR domain like ORRM4, show that the GR domain is responsible for mediating self-association by transient interactions with its RRM domain (NtRRM; Khan et al., 2014).
In the complementation assay, we observed an unexpected decrease of editing at cox2 C138, a site unaffected in the orrm4 mutant, upon the stable expression of ORRM4. It only occurred in the transgenic plants that displayed a higher extent of complementation at sites controlled by ORRM4. The decreased editing in these plants is likely due to the overexpression of ORRM4. The overexpression of ORRM4 in these transgenic plants (∼10 times the wild-type level) might sequester other editing factors and thus result in a decrease of editing at sites for which the sequestered editing factors are needed. For example, sufficient RIP1 is obviously required for cox2 C138 editing, given that editing at site this site decreased from ∼86% to ∼8% in the rip1 mutant (Bentolila et al., 2012). Because ORRM4 interacts with RIP1, an excessive amount of ORRM4 proteins may withhold a large number of RIP1 proteins. Consequently, the amount of RIP1 proteins at the RIP1 C targets would be deficient, leading to editing defects at sites not controlled by ORRM4. We noticed that the unexpected drop of editing extent did not occur when we transformed orrm4 mutants with a construct carrying the N-terminal RRM domain driven by a 35S promoter. Given that we have shown that the interaction of ORRM4 with ORRM3 is mediated through the C-terminal GR domain of ORRM4, the absence of a reduction in editing in transgenic lines overexpressing the N-terminal RRM domain is expected, because other editing factors cannot be sequestered due to the absence of the C-terminal domain. This theory is also supported by the fact that a decrease of editing at cox2 C138 is observed in some orrm4 mutants expressing full-length ORRM3, but not in the plants expressing the N-terminal RRM domain of ORRM3.
Like ORRM3, ORRM4 carries an RRM domain at its N terminus and a GR domain at its C terminus. In this study, we demonstrated that the expression of the RRM domain of ORRM4, rather than the expression of its GR domain, is able to restore the editing defects in the orrm4 mutant. However, in the transgenic line complemented for editing extent at sites rps4 C77 and rps4 C235, editing extent at site rpl5 C59 did not reach the wild-type level. The level of expression of the transgene in this plant (∼10 times the wild-type level) is comparable to the one measured in transgenic line expressing the full length ORRM4 and complemented for rpl5 C59 editing extent. Thus, we cannot exclude a supportive role for the C-terminal GR domain of ORRM4 in contributing to the editing ability of the protein for certain sites. We showed that the N-terminal RRM domain of ORRM4 and ORRM3 were overexpressed in both complementation experiments. The excess amount of nORRM4 or nORRM3 may impair the specificity contributed by GR domain through interacting with other editing factors. In addition to our previous results showing that the RRM domain by itself is able to provide the editing activity of ORRM1 and ORRM3, we have now further demonstrated that the RRM domain plays a key role in both plastid and mitochondrial editing, whereas the GR domain may play a supportive role in editing.
Expression of ORRM3 in the orrm4 background supports the hypothesis that some specificity exists in the RRM domains of different editing factors. Approximately 100 times the expression of the wild-type level of ORRM3 was necessary to complement the editing defects in a transgenic plant expressing the full-length ORRM3, whereas ∼10 times the expression of the full-length ORRM4 was sufficient to restore editing in all the sites analyzed.
The RRM domain is a conserved ∼80-amino acid domain that binds to a wide variety of RNA molecules with a range of specificities and affinities (Kenan et al., 1991). It is one of the most abundant protein domains in eukaryotes and is involved in various processes of RNA metabolism, such as premRNA processing, mRNA splicing, mRNA stability, and RNA editing (Maris et al., 2005). The numerous biological functions of the RRM-containing proteins are likely due to the structural versatility of the RRM domain, as well as the presence of variable auxiliary domains. The extreme structural versatility of the RRM domain makes it capable of interacting with diverse partners, not only RNA molecules but also proteins, whereas the auxiliary domains, such as the zinc fingers, are also involved in different aspects of RNA metabolism (Maris et al., 2005). Taken together, the data presented in this paper and previous literature suggest that ORRM4 may specifically recognize editing targets; nevertheless, the specificity of ORRM4 for its RNA targets might be enhanced by its C-terminal GR domain, which binds to cognate editing factors.
In addition to the ORRMs, other RRM-containing proteins have also been identified as RNA-editing factors. For instance, the mammalian ACF (for apobec-1 complementation factor) contains three RRM domains (Mehta et al., 2000). It binds to the apo-B mRNA and docks APOBEC1 to deaminate its C target (Mehta et al., 2000). Another Arabidopsis RRM-containing protein, CP31A, belongs to a family of chloroplast ribonucleoproteins with an acidic residue-rich domain and two RRM domains (Hirose and Sugiura, 2001; Tillich et al., 2009; Kupsch et al., 2012). CP31A was first characterized as a plastid editing factor as the cp31a mutant showed editing defects at several plastid sites compared to the wild type under normal growth conditions (Tillich et al., 2009). Even though decreased abundance of some transcripts was observed, the RNA alterations did not correlate with the editing defects that were detected (Tillich et al., 2009). A subsequent study showed that CP31A is likely to play a more important role in mRNA stability than in editing, as the effect of its mutation on RNA editing is minor compared to the effect on rapid change of mRNA accumulation and RNA processing in response to cold stress (Kupsch et al., 2012). Whether CP31A interacts with known components of the RNA editosome is unknown, while interactions between ORRM4 and known editing factors were observed in this study. Additionally, the response of ORRM4 toward stress conditions is different from that of CP31A, as ORRM4 expression was not affected by cold, hydration, or salt stress (Kwak et al., 2005).
The GR domain is a feature that is present in the plant Gly-rich proteins (GRPs), consisting of sequence repeats that can be summarized by the formula (Gly)n-X (Sachetto-Martins et al., 2000). The presence of the Gly-rich repeats in GRPs is usually accompanied by some other conserved domains, such as RRM domains, zinc fingers, and cold-shock domains (Mangeon et al., 2010). Plant GRPs generally exhibit a developmentally regulated and tissue-specific expression pattern, and their expression is also modulated by several biotic and abiotic factors (Sachetto-Martins et al., 2000; Mangeon et al., 2010). Plant GRPs have diverse functions in variable aspects of cell metabolism. They were proposed to function in cell elongation, plant defense, plant stress responses, flowering, and development, etc. (Fusaro et al., 2007; Fu et al., 2007; Kim et al., 2007; Streitner et al., 2008; Mangeon et al., 2009), likely due to the variation of their domain arrangements. Despite of all the efforts in characterizing the functions of plant GRPs, little is known about the function of the GR domain in these GRPs. With regard to ORRM4, we found that the RRM domain of ORRM4 provides its editing activity, whereas the GR domain of ORRM4 is required for its interaction with other editing factors. This result reveals a possible supportive function of the GR domain in plant GRPs. In other words, when the GRP is part of a protein complex, the additional domains (such as the RRM domain) in plant GRPs play a more important role in the protein function while the GR domain is responsible for its interaction with other members of the protein complex.
This study provides new insights into the molecular function of GR-RBPs. ORRM4 was first identified as a mitochondrial RNA-binding protein named At-mRBP2a (for mitochondrial RNA-binding protein 2a), isolated by its affinity to ssDNA in purified potato mitochondria (Vermel et al., 2002). ORRM4 was also described as GR-RBP5 (for Gly-rich RNA-binding Protein 5), one of the eight GR-RBP family members in Arabidopsis (Lorković and Barta, 2002). All these GR-RBPs are classified as Class IVa GRPs (Mangeon et al., 2010) based on a N-terminal RRM domain and a C-terminal GR domain of variable length. Plant GR-RBPs have been implicated in the responses to various stress conditions, particularly to cold stress (Vermel et al., 2002; Kwak et al., 2005; Kim et al., 2007, 2008; Mangeon et al., 2010). Except for ORRM4 (GR-RBP5) and GR-RBP6, cold treatment greatly increased the expression of all the other GR-RBPs (GR-RBP-1 to -4, -7, and -8; Kwak et al., 2005). Though the molecular apparatus relevant to the function of these other GR-RBPs is still at large, several studies have revealed their involvement in RNA metabolism (Kwak et al., 2005; Schöning et al., 2008; Mangeon et al., 2010).
We observed a late-flowering phenotype in the orrm4 mutant compared to the wild type. Mutation or silencing of two other GR-RBPs, GR-RBP7 and GR-RBP8 (also named AtGRP7 and AtGRP8), also showed a late-flowering phenotype (Streitner et al., 2008). The late-flowering phenotype in the atgrp7 mutant is associated with an elevated transcript level of a MADS-box protein FLC (for Flowering Locus C), a key repressor of flowering (Streitner et al., 2008). Another study revealed that the silencing of CSDP2 (for Cold Shock Domain Protein 2, also named AtGRP2) resulted in an early flowering phenotype (Fusaro et al., 2007). GR-RBP7, GR-RBP8, and CSDP2 are all nucleo-cytoplasmic proteins (Fusaro et al., 2007; Fu et al., 2007); as a result of which, the mechanism underlying how these proteins regulate the flowering time is likely to be very different from the influence of ORRM4.
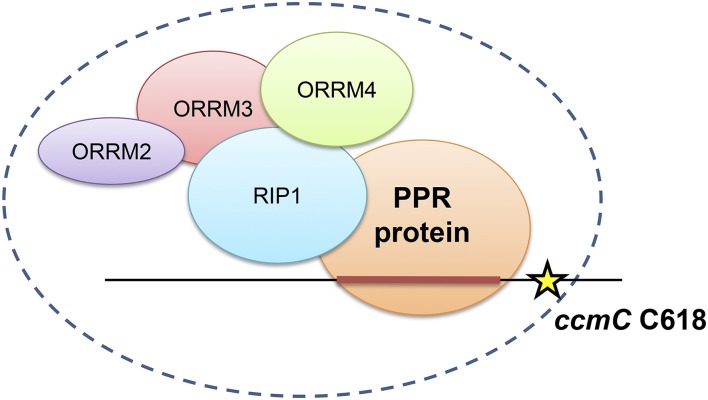
This study further supports the involvement of the ORRM family in organelle editosomes. Based on all of the interaction data we obtained in this study as well as previous studies, we propose an updated model for mitochondrial editosome. The mitochondrial editosome is a complex that contains various numbers of RIP/MORFs, ORRMs, and at least one PPR protein. It is heterogenous in its composition and has an unknown stoichiometry of subunits. Figure 12 is an example of a model for the editosome acting upon site ccmC C618 based on the interaction data obtained from Y2H and BiFC assays reported here and previously (Bentolila et al., 2012; Shi et al., 2015). In this model, RIP1 is a key component in the editosome as it acts like a bridge holding the PPR proteins and the ORRMs together, whereas ORRM proteins exert their functions through interacting with RIP1 and with each other. In this model, we have not represented the binding of ORRMs with the RNA even though it is likely to occur, given there is ample support from literature, including our own data from the study of the chloroplast editing ORRM1 (Sun et al., 2013). In addition, we represent this editosome model as a static complex where all the proteins are present all together, but the interaction between the different components might occur consecutively.
Figure 12.
Model of the mitochondrial editosome that operates on ccmC C618 based on protein-protein interaction data. Data are from Y2H and BiFC assays (this study; Bentolila et al., 2012; Shi et al., 2015). The stoichiometry of the components is not indicated, as it is unknown. The editable C at position 618 on the ccmC transcript is represented by a star. The cis-element upstream of the target C is shown as a red bar.
MATERIALS AND METHODS
Plant Material and Morphological Analysis
The Arabidopsis (Arabidopsis thaliana) T-DNA insertion line SALK_023061C in the ORRM4 gene was ordered from the Arabidopsis Biological Resource Center (https://abrc.osu.edu/). After 3 d of stratification, seeds from the mutant line were planted in soil and grown in a growth room (14 h of light/10 h of dark) at 26°C. Plants were genotyped by PCR with BioMix Red (Bioline) using primer pair SALK023061C_LP and SALK023061C_RP listed in Supplemental Table S5. The PCR product was sequenced at Cornell University Life Sciences Core Laboratories Center. Leaves were collected from 5-week-old plants for further analysis. The ORRM4 expression level was measured by qRT-PCR.
The fresh weight of SALK_023061C plants was measured 24, 31, and 37 d after planted. We recorded the number of days it took for visible flower buds to show in the center of the rosette, for inflorescence stem of the plant to reach 1 cm in height, and for its first flower to open. Information regarding to the fresh weight and the number of total leaves of the mutant versus the wild-type was recorded when the first flower bloomed.
VIGS
To perform VIGS, primers flanking a gene-specific region were selected from the CATMA database (Crowe et al., 2003). Fragments were amplified using Arabidopsis Columbia ecotype genomic DNA as a template with primer pairs ORRM4_VIGS_F and ORRM4_VIGS_R. The PCR product was first integrated into a PCR8/GW/TOPO vector (Invitrogen) and then transferred to a silencing vector PTRV2/GW/GFP upon a LR Clonase II (Invitrogen) recombination reaction. The silencing construct was subsequently transformed into Agrobacterium tumefaciens GV3101 by electroporation. Agrobacterium expressing the ORRM4/GFP silencing construct was used to infiltrate 2-week-old Arabidopsis seedlings expressing 35S-GFP as previously described (Bentolila et al., 2012). The green fluorescence of the inoculated plants was examined under UV light to track the silencing efficiency 2 weeks after infiltration. Leaves from 5-week-old GFP-silenced plants that exhibited dark red chlorophyll fluorescence, but no green fluorescence, were collected for further analysis. The silencing efficiency of ORRM4 was measured by qRT-PCR.
Use of STS-PCRseq Method to Assay Editing Extent
The STS-PCRseq technique was presented in detail in a previous paper (Bentolila et al., 2013). In brief, we amplified all transcripts encoding either plastid or mitochondrial genes with organelle-specific primers from the RNA extracted from the silenced plants, the mutant plants, and the controls. The RT-PCR products were mixed in equimolar ratio, sheared by sonication, and then used as a template to produce an Illumina TruSeq DNA Library. All the samples in this study were pooled in one sequencing lane of an Illumina HiSEquation 2500 instrument. The data we obtained was processed according to the guideline in Bentolila et al. (2013). Subsequent statistical analysis was previously described as in Bentolila et al. (2013) and in Shi et al. (2015). Briefly, we detected 618 mitochondrial editing sites and 38 plastid editing sites. Because of repetitive testing we adjusted the nominal significance test by a Bonferroni correction to P < 1.6e-6 or P < 2.6e-5 for the mitochondrial or plastid editing set, respectively. In the silencing experiments, we discarded 54 mitochondrial sites and 3 plastid sites from our analyses because they showed a significant reduction of editing extent in the GFP-silenced control when compared to the uninoculated plants. Finally, the only sites that were declared to be significantly reduced in their editing extent were those sites that exhibited a significant reduction in ORRM4-silenced plants when compared with both control plants, uninoculated and GFP-silenced.
Use of PPE Assay to Measure Editing Extents
RNA was extracted using PureLink RNA Mini Kit (Life Technologies), cleared from DNA contamination by using a turbo DNA-free kit (Ambion), followed by reverse transcription using the SuperScript III Reverse Transcriptase (Life Technologies). Primers used are listed in Supplemental Table S5 or provided in previous studies (Bentolila et al., 2008, 2012, 2013; Shi et al., 2015). Editing extent was examined by PPE assays as described previously (Peeters and Hanson, 2002; Robbins et al., 2009).
Generation of Transgenic Plants
The coding sequence of ORRM4 was reverse-transcribed with SuperScript III Reverse Transcriptase (Life Technologies) from RNA extracted from wild-type Arabidopsis Columbia using PureLink RNA Mini Kit (Life Technologies) and then cloned into a PCR8/GW/TOPO vector. Primers used were listed in Supplemental Table S5. The fragment was subsequently shuffled into a modified pBI121 vector using LR Clonase II. The sequence encoding the N-terminus of ORRM4 was amplified from the reverse-transcribed ORRM4 coding sequence with primer pair ORRM4_F and ORRM4_336R_TAG. The C-terminal ORRM4 with the mitochondrial transient peptide (amino acids 1–29 of the nuclear-encoded coxIV from yeast as described by Köhler et al. [1997]) was cloned using an overlap extension PCR method. The fragment amplified using primer pair ScCOX4_F and ScCOX4_R and the fragment amplified using ScCOX4R_ORRM4_337F and ORRM4_R were mixed and used as a template for a third PCR reaction with primer pair ScCOX4_F and ORRM4_R. The cloned N-terminal ORRM4 and the C-terminal ORRM4 with the transient peptide were each integrated into a PCR8/GW/TOPO vector and then transferred into the pBI121 vector by LR reaction. 35S-ORRM4, 35S-nORRM4, and 35S-ScCOX4-cORRM4 in the pBI121 vector were transformed into A. tumefaciens GV3101. Agrobacterium transformed with 35S-nORRM3 or 35S-ORRM3 was from a previous study (Shi et al., 2015). Floral dip transformation of homozygous orrm4 mutant plants was performed as described by Zhang et al. (2006). Plants were sprayed with Basta twice on soil for selection. The presence of the transgene and the homozygosity of the orrm4 mutant allele were verified by PCR reactions using primers listed in Supplemental Table S5. Leaves from 4- to 6-week-old transgenic plants were collected for further analysis. T1 generation of a 35S-ORRM4 transgenic line was planted on soil, and leaves from 4-week segregating T1 plants were collected.
Y2H Assay
The ORRM4 coding sequence with its predicted transit peptide removed (amino acids 1–30), was amplified from the reverse-transcribed ORRM4 cDNA clone with primer pair ORRM4_91F and ORRM4_R as listed in Supplemental Table S5. The coding sequence of N-terminal ORRM4 was cloned with primers ORRM4_91F and ORRM4_336R_TAG, whereas the coding sequence of C-terminal ORRM4 was cloned with primers ORRM4_337F and ORRM4_R using the ORRM4 cDNA clone as a template. The N-terminal ORRM3 (amino acids 1–36 removed) was amplified with primer pair ORRM3_109F and ORRM3_360R_TAG from the reverse-transcribed ORRM3 cDNA clone as in Shi et al. (2015). All these PCR products were integrated into PCR8/GW/TOPO vectors and then shuffled to pGADT7GW and pGBKT7GW vectors (Horák et al., 2008), respectively. The ORRM3 construct was from Shi et al. (2015).
Two mating types of yeast strain PJ69-4, a and α, were transformed with the constructs above followed by a protocol in Gietz et al. (1995). Double transformants were produced by mating single transformants of mating type a to mating type α, cultured in Leu- and Trp-dropout media (Clontech), and subsequently diluted with sterile water to 1 × 106, 1 × 105 cells /mL. A total of 10 µL of each dilution was spotted on Leu-, Trp-, adenine-, and His-dropout media plates. Yeasts transformed with empty vectors were used as negative controls to detect auto-activation. Data were collected from 2 d to 5 d after spotting.
Bimolecular Fluorescence Complementation Assay
The coding sequences of ORRM3, ORRM4, and RIP1 without the stop codon were amplified from the full-length cDNAs as described above and in previous studies (Bentolila et al., 2012; Shi et al., 2015), and cloned into pENTR/D-TOPO vectors (Invitrogen). These fragments were then shuffled into XNGW and XCGW vectors (Ohashi-Ito and Bergmann, 2006) by LR reactions. The coding sequence of MEF9 without the stop codon was reverse-transcribed from RNA extracted from wild-type Arabidopsis and subsequently cloned into a PCR8/GW/TOPO vector. The DNA was incubated with restriction endonucleases XhoI and XbaI for 4 h under 37°C and then transferred into the destination vectors XNGW and XCGW by LR reactions. Final vectors were validated by sequencing and transformed into A. tumefaciens GV3101. A range of 5 to 7 mL of each agrobacterial culture were incubated for 2 d at 28°C, and resuspended in 2-(N-morpholine)-ethanesulphonic acid buffer, pH 5.6, with 10 mm MgCl2 and 150 μm acetosyringone to an optical density of 1.0. Then we mixed the culture expressing GFPN to another culture expressing GFPC in equal volume to a final optical density of 0.5 each and incubated the mixtures in the dark at 28°C for about three hours. Afterward, these mixtures were used to infiltrate leaves from 4- to 6-week-old Nicotiana benthamiana plants grown at 10 h of light per day as described in Lin et al. (2014). Within 2 to 4 d after agroinfiltration, the leaf tissues were examined with a confocal microscope (Zeiss LSM 710) through a LD LCI Plan-Apochromat 25×/0.8 Imm Korr DIC M27 objective. The 488 nm line of an argon laser was used to excite GFP and chlorophyll. The images were collected and processed with ZEN 2012 microscope software (Carl Zeiss Microscopy).
Real-Time qRT-PCR Conditions and Analysis
The real-time qRT-PCR was performed as described in a previous study (Shi et al., 2015). Primers used in the reaction are listed in Supplemental Table S5. The expression levels of ORRM4 and the sequence encoding the N-terminal RRM domain of ORRM4 were measured with qRT-ORRM4-F and qRT-ORRM4-R. C-terminal GR domain of ORRM4 expression was measured with qRT-cORRM4-F and qRT-cORRM4-R. ORRM3 and N-terminal RRM domain of ORRM3 expression were measured with primer pairs qRT-ORRM3-F + qRT-ORRM3-R and qRT-nORRM3-F + qRT-nORRM3-R.
RNA Blots
RNA gel bot analysis was performed as described by Germain et al. (2011). Primers used to make the probes are listed in Bentolila et al. (2013).
Accession Numbers
Sequence data from this article can be found in the GenBank/EMBL data libraries under accession numbers BT002037.
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. High reproducibility of editing extent measured by STS-PCRseq between biological replicates.
Supplemental Figure S2. Variable effect of orrm4 mutation on nad4L editing extent.
Supplemental Figure S3. Effect of orrm4 mutation on transcript abundance can be reverted in complemented lines.
Supplemental Figure S4. Alignment of ORRM3 and ORRM4.
Supplemental Table S1. Sites showing a significant reduction of editing extent in ORRM4-silenced plants.
Supplemental Table S2. Plastid sites editing extent in ORRM4-silenced plants.
Supplemental Table S3. Mitochondrial sites showing a significant reduction of editing extent in orrm4 mutant plants.
Supplemental Table S4. Plastid sites showing a significant reduction of editing extent in orrm4 mutant plants.
Supplemental Table S5. Primers used in this study.
Supplementary Dataset S1. Number of reads at each editing site (gene-position) for each library (genotype).
Supplementary Dataset S2. Analysis of the influence of rip1, rip3, and orrm4 mutations on the editing extent of mitochondrial sites.
Supplementary Material
Acknowledgments
We thank Robert Bukowski for performing the primary analysis of the Illumina sequencing reads.
Glossary
- PPR
pentatricopeptide repeat
- GR
Gly-rich
- qRT-PCR
quantitative reverse transcription PCR
- VIGS
virus-induced gene silencing
- STS-PCRseq
strand- and transcript-specific RNA-seq method
- PPE
poisoned primer extension
- Y2H
yeast two-hybrid
- BiFC
bimolecular fluorescence complementation
- GRP
Gly-rich protein
Footnotes
This work was supported by the National Science Foundation Division of Molecular and Cellular Biosciences, Gene and Genome Systems (grant no. MCB–1330294 to M.R.H. and S.B.).
Articles can be viewed without a subscription.
References
- Barkan A, Rojas M, Fujii S, Yap A, Chong YS, Bond CS, Small I (2012) A combinatorial amino acid code for RNA recognition by pentatricopeptide repeat proteins. PLoS Genet 8: e1002910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentolila S, Elliott LE, Hanson MR (2008) Genetic architecture of mitochondrial editing in Arabidopsis thaliana. Genetics 178: 1693–1708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentolila S, Heller WP, Sun T, Babina AM, Friso G, van Wijk KJ, Hanson MR (2012) RIP1, a member of an Arabidopsis protein family, interacts with the protein RARE1 and broadly affects RNA editing. Proc Natl Acad Sci USA 109: E1453–E1461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentolila S, Oh J, Hanson MR, Bukowski R (2013) Comprehensive high-resolution analysis of the role of an Arabidopsis gene family in RNA editing. PLoS Genet 9: e1003584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boussardon C, Avon A, Kindgren P, Bond CS, Challenor M, Lurin C, Small I (2014) The cytidine deaminase signature HxE(x)n CxxC of DYW1 binds zinc and is necessary for RNA editing of ndhD-1. New Phytol 203: 1090–1095 10.1111/nph.12928 [DOI] [PubMed] [Google Scholar]
- Chateigner-Boutin A-L, Ramos-Vega M, Guevara-García A, Andrés C, de la Luz Gutiérrez-Nava M, Cantero A, Delannoy E, Jiménez LF, Lurin C, Small I, et al. (2008) CLB19, a pentatricopeptide repeat protein required for editing of rpoA and clpP chloroplast transcripts. Plant J 56: 590–602 [DOI] [PubMed] [Google Scholar]
- Chateigner-Boutin A-L, Small I (2007) A rapid high-throughput method for the detection and quantification of RNA editing based on high-resolution melting of amplicons. Nucleic Acids Res 35: e114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaudhuri S, Maliga P (1996) Sequences directing C to U editing of the plastid psbL mRNA are located within a 22 nucleotide segment spanning the editing site. EMBO J 15: 5958–5964 [PMC free article] [PubMed] [Google Scholar]
- Covello PS, Gray MW (1989) RNA editing in plant mitochondria. Nature 341: 662–666 [DOI] [PubMed] [Google Scholar]
- Crowe ML, Serizet C, Thareau V, Aubourg S, Rouzé P, Hilson P, Beynon J, Weisbeek P, van Hummelen P, Reymond P, et al. (2003) CATMA: a complete Arabidopsis GST database. Nucleic Acids Res 31: 156–158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fusaro AF, Bocca SN, Ramos RLB, Barrôco RM, Magioli C, Jorge VC, Coutinho TC, Rangel-Lima CM, De Rycke R, Inzé D, et al. (2007) AtGRP2, a cold-induced nucleo-cytoplasmic RNA-binding protein, has a role in flower and seed development. Planta 225: 1339–1351 [DOI] [PubMed] [Google Scholar]
- Fu ZQ, Guo M, Jeong BR, Tian F, Elthon TE, Cerny RL, Staiger D, Alfano JR (2007) A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 447: 284–288 [DOI] [PubMed] [Google Scholar]
- Germain A, Herlich S, Larom S, Kim SH, Schuster G, Stern DB (2011) Mutational analysis of Arabidopsis chloroplast polynucleotide phosphorylase reveals roles for both RNase PH core domains in polyadenylation, RNA 3′-end maturation and intron degradation. Plant J 67: 381–394 [DOI] [PubMed] [Google Scholar]
- Gietz RD, Schiestl RH, Willems AR, Woods RA (1995) Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11: 355–360 [DOI] [PubMed] [Google Scholar]
- Gray MW. (2012) Evolutionary origin of RNA editing. Biochemistry 51: 5235–5242 [DOI] [PubMed] [Google Scholar]
- Gualberto JM, Lamattina L, Bonnard G, Weil J-H, Grienenberger J-M (1989) RNA editing in wheat mitochondria results in the conservation of protein sequences. Nature 341: 660–662 [DOI] [PubMed] [Google Scholar]
- Hayes ML, Dang KN, Diaz MF, Mulligan RM (2015) A conserved glutamate residue in the C-terminal deaminase domain of pentatricopeptide repeat proteins is required for RNA editing activity. J Biol Chem 290: 10136–10142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes ML, Hanson MR (2007) Identification of a sequence motif critical for editing of a tobacco chloroplast transcript. RNA 13: 281–288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiesel R, Wissinger B, Schuster W, Brennicke A (1989) RNA editing in plant mitochondria. Science 246: 1632–1634 [DOI] [PubMed] [Google Scholar]
- Hirose T, Sugiura M (2001) Involvement of a site-specific trans-acting factor and a common RNA-binding protein in the editing of chloroplast mRNAs: development of a chloroplast in vitro RNA editing system. EMBO J 20: 1144–1152 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horák J, Grefen C, Berendzen KW, Hahn A, Stierhof Y-D, Stadelhofer B, Stahl M, Koncz C, Harter K (2008) The Arabidopsis thaliana response regulator ARR22 is a putative AHP phospho-histidine phosphatase expressed in the chalaza of developing seeds. BMC Plant Biol 8: 77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kenan DJ, Query CC, Keene JD (1991) RNA recognition: towards identifying determinants of specificity. Trends Biochem Sci 16: 214–220 [DOI] [PubMed] [Google Scholar]
- Khan F, Daniëls MA, Folkers GE, Boelens R, Saqlan Naqvi SM, van Ingen H (2014) Structural basis of nucleic acid binding by Nicotiana tabacum glycine-rich RNA-binding protein: implications for its RNA chaperone function. Nucleic Acids Res 42: 8705–8718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim JS, Jung HJ, Lee HJ, Kim KA, Goh C-H, Woo Y, Oh SH, Han YS, Kang H (2008) Glycine-rich RNA-binding protein 7 affects abiotic stress responses by regulating stomata opening and closing in Arabidopsis thaliana. Plant J 55: 455–466 [DOI] [PubMed] [Google Scholar]
- Kim JY, Park SJ, Jang B, Jung C-H, Ahn SJ, Goh C-H, Cho K, Han O, Kang H (2007) Functional characterization of a glycine-rich RNA-binding protein 2 in Arabidopsis thaliana under abiotic stress conditions. Plant J 50: 439–451 [DOI] [PubMed] [Google Scholar]
- Köhler RH, Zipfel WR, Webb WW, Hanson MR (1997) The green fluorescent protein as a marker to visualize plant mitochondria in vivo. Plant J 11: 613–621 [DOI] [PubMed] [Google Scholar]
- Kotera E, Tasaka M, Shikanai T (2005) A pentatricopeptide repeat protein is essential for RNA editing in chloroplasts. Nature 433: 326–330 [DOI] [PubMed] [Google Scholar]
- Kupsch C, Ruwe H, Gusewski S, Tillich M, Small I, Schmitz-Linneweber C (2012) Arabidopsis chloroplast RNA binding proteins CP31A and CP29A associate with large transcript pools and confer cold stress tolerance by influencing multiple chloroplast RNA processing steps. Plant Cell 24: 4266–4280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwak KJ, Kim YO, Kang H (2005) Characterization of transgenic Arabidopsis plants overexpressing GR-RBP4 under high salinity, dehydration, or cold stress. J Exp Bot 56: 3007–3016 [DOI] [PubMed] [Google Scholar]
- Lin MT, Occhialini A, Andralojc PJ, Devonshire J, Hines KM, Parry MAJ, Hanson MR (2014) β-Carboxysomal proteins assemble into highly organized structures in Nicotiana chloroplasts. Plant J 79: 1–12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lorković ZJ, Barta A (2002) Genome analysis: RNA recognition motif (RRM) and K homology (KH) domain RNA-binding proteins from the flowering plant Arabidopsis thaliana. Nucleic Acids Res 30: 623–635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu B, Hanson MR (1992) A single nuclear gene specifies the abundance and extent of RNA editing of a plant mitochondrial transcript. Nucleic Acids Res 20: 5699–5703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lurin C, Andrés C, Aubourg S, Bellaoui M, Bitton F, Bruyère C, Caboche M, Debast C, Gualberto J, Hoffmann B, et al. (2004) Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 16: 2089–2103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mangeon A, Junqueira RM, Sachetto-Martins G (2010) Functional diversity of the plant glycine-rich proteins superfamily. Plant Signal Behav 5: 99–104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mangeon A, Magioli C, Menezes-Salgueiro AD, Cardeal V, de Oliveira C, Galvão VC, Margis R, Engler G, Sachetto-Martins G (2009) AtGRP5, a vacuole-located glycine-rich protein involved in cell elongation. Planta 230: 253–265 [DOI] [PubMed] [Google Scholar]
- Maris C, Dominguez C, Allain FH-T (2005) The RNA recognition motif, a plastic RNA-binding platform to regulate post-transcriptional gene expression. FEBS J 272: 2118–2131 [DOI] [PubMed] [Google Scholar]
- Mehta A, Kinter MT, Sherman NE, Driscoll DM (2000) Molecular cloning of apobec-1 complementation factor, a novel RNA-binding protein involved in the editing of apolipoprotein B mRNA. Mol Cell Biol 20: 1846–1854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohashi-Ito K, Bergmann DC (2006) Arabidopsis FAMA controls the final proliferation/differentiation switch during stomatal development. Plant Cell 18: 2493–2505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okuda K, Shoki H, Arai M, Shikanai T, Small I, Nakamura T (2014) Quantitative analysis of motifs contributing to the interaction between PLS-subfamily members and their target RNA sequences in plastid RNA editing. Plant J 80: 870–882 [DOI] [PubMed] [Google Scholar]
- Peeters NM, Hanson MR (2002) Transcript abundance supercedes editing efficiency as a factor in developmental variation of chloroplast gene expression. RNA 8: 497–511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robbins JC, Heller WP, Hanson MR (2009) A comparative genomics approach identifies a PPR-DYW protein that is essential for C-to-U editing of the Arabidopsis chloroplast accD transcript. RNA 15: 1142–1153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruwe H, Castandet B, Schmitz-Linneweber C, Stern DB (2013) Arabidopsis chloroplast quantitative editotype. FEBS Lett 587: 1429–1433 [DOI] [PubMed] [Google Scholar]
- Sachetto-Martins G, Franco LO, de Oliveira DE (2000) Plant glycine-rich proteins: a family or just proteins with a common motif? Biochim Biophys Acta 1492: 1–14 [DOI] [PubMed] [Google Scholar]
- Salone V, Rüdinger M, Polsakiewicz M, Hoffmann B, Groth-Malonek M, Szurek B, Small I, Knoop V, Lurin C (2007) A hypothesis on the identification of the editing enzyme in plant organelles. FEBS Lett 581: 4132–4138 [DOI] [PubMed] [Google Scholar]
- Schöning JC, Streitner C, Meyer IM, Gao Y, Staiger D (2008) Reciprocal regulation of glycine-rich RNA-binding proteins via an interlocked feedback loop coupling alternative splicing to nonsense-mediated decay in Arabidopsis. Nucleic Acids Res 36: 6977–6987 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi X, Hanson MR, Bentolila S (2015) Two RNA recognition motif-containing proteins are plant mitochondrial editing factors. Nucleic Acids Res 43: 3814–3825 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith HC, Gott JM, Hanson MR (1997) A guide to RNA editing. RNA 3: 1105–1123 [PMC free article] [PubMed] [Google Scholar]
- Staudinger M, Kempken F (2003) Electroporation of isolated higher-plant mitochondria: transcripts of an introduced cox2 gene, but not an atp6 gene, are edited in organello. Mol Genet Genomics 269: 553–561 [DOI] [PubMed] [Google Scholar]
- Streitner C, Danisman S, Wehrle F, Schöning JC, Alfano JR, Staiger D (2008) The small glycine-rich RNA binding protein AtGRP7 promotes floral transition in Arabidopsis thaliana. Plant J 56: 239–250 [DOI] [PubMed] [Google Scholar]
- Sun T, Germain A, Giloteaux L, Hammani K, Barkan A, Hanson MR, Bentolila S (2013) An RNA recognition motif-containing protein is required for plastid RNA editing in Arabidopsis and maize. Proc Natl Acad Sci USA 110: E1169–E1178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun T, Shi X, Friso G, Van Wijk K, Bentolila S, Hanson MR (2015) A zinc finger motif-containing protein is essential for chloroplast RNA editing. PLoS Genet 11: e1005028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takenaka M. (2010) MEF9, an E-subclass pentatricopeptide repeat protein, is required for an RNA editing event in the nad7 transcript in mitochondria of Arabidopsis. Plant Physiol 152: 939–947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takenaka M, Verbitskiy D, Zehrmann A, Brennicke A (2010) Reverse genetic screening identifies five E-class PPR proteins involved in RNA editing in mitochondria of Arabidopsis thaliana. J Biol Chem 285: 27122–27129 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takenaka M, Zehrmann A, Brennicke A, Graichen K (2013) Improved computational target site prediction for pentatricopeptide repeat RNA editing factors. PLoS One 8: e65343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takenaka M, Zehrmann A, Verbitskiy D, Kugelmann M, Härtel B, Brennicke A (2012) Multiple organellar RNA editing factor (MORF) family proteins are required for RNA editing in mitochondria and plastids of plants. Proc Natl Acad Sci USA 109: 5104–5109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tillich M, Hardel SL, Kupsch C, Armbruster U, Delannoy E, Gualberto JM, Lehwark P, Leister D, Small ID, Schmitz-Linneweber C (2009) Chloroplast ribonucleoprotein CP31A is required for editing and stability of specific chloroplast mRNAs. Proc Natl Acad Sci USA 106: 6002–6007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vermel M, Guermann B, Delage L, Grienenberger J-M, Maréchal-Drouard L, Gualberto JM (2002) A family of RRM-type RNA-binding proteins specific to plant mitochondria. Proc Natl Acad Sci USA 99: 5866–5871 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagoner JA, Sun T, Lin L, Hanson MR (2015) Cytidine deaminase motifs within the DYW domain of two pentatricopeptide repeat-containing proteins are required for site-specific chloroplast RNA editing. J Biol Chem 290: 2957–2968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wintz H, Hanson MR (1991) A termination codon is created by RNA editing in the petunia mitochondrial atp9 gene transcript. Curr Genet 19: 61–64 [DOI] [PubMed] [Google Scholar]
- Yin P, Li Q, Yan C, Liu Y, Liu J, Yu F, Wang Z, Long J, He J, Wang H-W, et al. (2013) Structural basis for the modular recognition of single-stranded RNA by PPR proteins. Nature 504: 168–171 [DOI] [PubMed] [Google Scholar]
- Zehrmann A, Härtel B, Glass F, Bayer-Császár E, Obata T, Meyer E, Brennicke A, Takenaka M (2015) Selective homo- and heteromer interactions between the multiple organellar RNA editing factor (MORF) proteins in Arabidopsis thaliana. J Biol Chem 290: 6445–6456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zehrmann A, Verbitskiy D, Härtel B, Brennicke A, Takenaka M (2010) RNA editing competence of trans-factor MEF1 is modulated by ecotype-specific differences but requires the DYW domain. FEBS Lett 584: 4181–4186 [DOI] [PubMed] [Google Scholar]
- Zehrmann A, Verbitskiy D, van der Merwe JA, Brennicke A, Takenaka M (2009) A DYW domain-containing pentatricopeptide repeat protein is required for RNA editing at multiple sites in mitochondria of Arabidopsis thaliana. Plant Cell 21: 558–567 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X, Henriques R, Lin S-S, Niu Q-W, Chua N-H (2006) Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat Protoc 1: 641–646 [DOI] [PubMed] [Google Scholar]
- Zhou W, Karcher D, Bock R (2014) Identification of enzymes for adenosine-to-inosine editing and discovery of cytidine-to-uridine editing in nucleus-encoded transfer RNAs of Arabidopsis. Plant Physiol 166: 1985–1997 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.