Abstract
Peroxiredoxin 6 (PRDX6) is one of six members of the PRDX family, which have peroxidase and antioxidant activity. PRDX6 is unique, containing only one conserved cysteine residue (C47) rather than the two found in other PRDXs. A yeast two-hybrid screen found PRDX6 to be a potential binding partner of the C-terminal tail of anion exchanger 1 (AE1), a Cl−/HCO3− exchanger basolaterally expressed in renal α-intercalated cells. PRDX6 immunostaining in human kidney was both cytoplasmic and peripheral and co-localized with AE1. Analysis of native protein showed it was largely monomeric, whereas expressed tagged protein was more dimeric. Two methionine oxidation sites were identified. In vitro and ex vivo pulldowns and immunoprecipitation assays confirmed interaction with AE1, but mutation of the conserved cysteine resulted in loss of interaction. Prdx6 knockout mice had a baseline acidosis with a major respiratory component and greater AE1 expression than wild type animals. After an oral acid challenge, PRDX6 expression increased in wild type mice, with preservation of AE1. However, AE1 expression was significantly decreased in knockout animals. Kidneys from acidified mice showed widespread proximal tubular vacuolation in wild type but not knockout animals. Knockdown of PRDX6 by siRNA in mammalian cells reduced both total and cell membrane AE1 levels. Thus, PRDX6-AE1 interaction contributes to maintenance of AE1 during cellular stress such as during metabolic acidosis.
Keywords: distal tubule, renal pathology
Introduction
Anion exchanger 1 (AE1), also known as Band 3, is a sodium-independent Cl−/HCO3− exchanger member of the SLC4 gene family expressed in erythrocytes (eAE1) and at the basolateral surface of intercalated cells in the kidney (kAE1). AE1 plays a crucial role in intracellular and systemic pH regulation, and maintenance of intracellular Cl− levels and cell volume.1 Compared with eAE1, human kAE1 lacks the N-terminal 65 amino acids.2 This truncation is known to preclude kAE1 protein binding to some known eAE1 partners (such as aldolase and ankyrin), but does not interfere with anion exchange.3, 4, 5, 6, 7 The C-terminus of AE1 (AE1(C)), containing residues 876-911, has proved interesting, with the elucidation firstly of AE1(C) mutations that result in distal renal tubular acidosis, including a truncating mutation (AE1(C)Δ11) and a point mutation (M909T) causing aberrant kAE1 membrane targeting, and secondly, C-terminal protein binding partners including GAPDH, the sodium pump’s β1 subunit, kinesin 3B, carbonic anhydrase and AP1.8, 9, 10, 11, 12, 13, 14, 15, 16, 17 To investigate the protein binding of AE1(C), a yeast two-hybrid screen was designed using both intact AE1(C)WT and AE1(C)Δ11 against a human kidney cDNA library.13 A number of candidate binding partners were identified, including peroxiredoxin 6 (PRDX6), which we investigated in this work.
The peroxiredoxins, a family of at least 6 members in mammals, have peroxidase activity, catalyzing H2O2 to H2O and acting as antioxidant proteins. While maintaining these functions, PRDX6 is also uniquely capable of phospholipase A2 activity, hydrolyzing the sn-2 acyl bond of phospholipids to release arachidonic acid and lysophospholipids.18 PRDX6 has only one conserved cysteine (C47) that is buried within the protein’s tertiary structure,19 whereas the other PRDX family members all contain two conserved cysteine residues and typically form homodimers.20 The crystal structure of PRDX6 reveals the exposed catalytic triad for phospholipase A2 activity (His26, Ser32, Asp140) and the buried conserved cysteine (C47), which is the catalytic site for peroxidase activity;18 other structural work indicates that heterodimerization with glutathione accompanies PRDX6’s activity.21 Northern and Western blotting have shown significant tissue expression of PRDX6 in brain, heart, kidney, lung and testis,22 and PRDX6 expression has recently been identified in all sections of the nephron.23 Here we report the interaction between AE1 and PRDX6 in the kidney, and demonstrate a role for renal PRDX6 in pH homeostasis.
Results
PRDX6 identified as a potential binding partner for AE1(C)
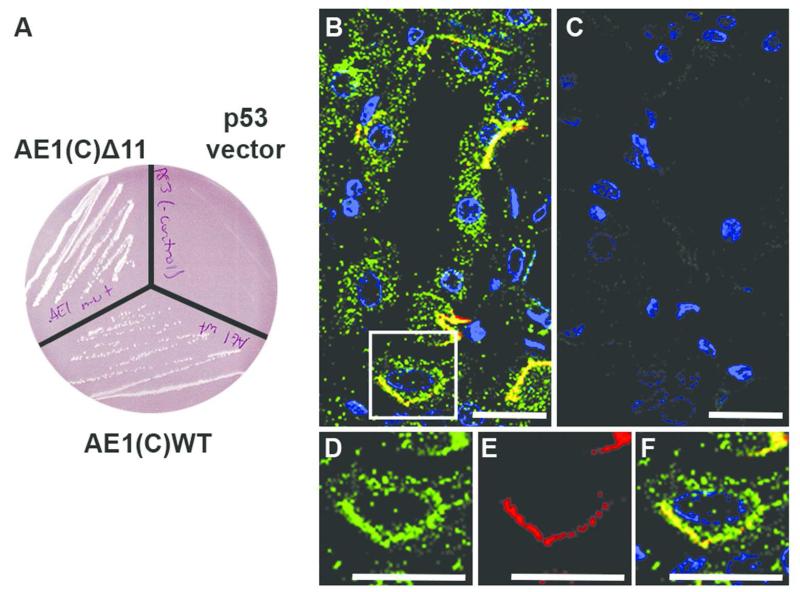
Initial evidence of an interaction between AE1 and PRDX6 from yeast two-hybrid screening was validated by obtaining colonies following specific mating of both the AE1(C)WT and AE1(C)Δ11 bait strains, but not p53 vector strain alone, with the PRDX6 strain (Figure 1A). This indicates that the interaction takes place upstream of the R901 truncation site within AE1’s 36-residue C-terminal cytoplasmic domain.
Figure 1. Confirmation of PRDX6 as binding candidate of kAE1and co-staining of intercalated cell.
(A) Yeast mating test of both AE1(C)WT and AE1(C)Δ11 with PRDX6 in yeast cells shows protein potential interaction indicated by the blue colonies on selection plates with no cell growth of the negative control (p53 empty vector). (B-F) Immunofluorescence in human renal collecting duct. PRDX6 (green) and AE1 (red) co-localize in a number of collecting duct cells (B) with absence of staining when primary antisera were omitted (C). An enlarged image of an intercalated cell shows widespread PRDX6 expression (D) and basolateral AE1 (Bric 170) expression (E). Merged panel (F) indicates co-localization of PRDX6 and AE1 at the basolateral surface. (Scale bars = 20 μm)
PRDX6 is present in human and mouse kidney
Localization of the two proteins in human kidney was examined using dual immunostaining. As expected, kAE1 was constrained to the basolateral surface of intercalated cells (Figure 1B-F, red), while PRDX6 (green) was observed in all cells and its distribution was more widespread in both cytoplasm and at the cell periphery. Clear co-localization of PRDX6 and kAE1 was observed (Figure 1F). Absence of staining when primary antibodies were omitted confirmed specificity (C). Further confirmation of the PRDX6 antibodies’ specificity was demonstrated on Prdx6+/+ and Prdx6−/− kidney sections and lysates (Supplementary Figure 1).
Analysis of native PRDX6 expression in human and mouse kidney under non-reducing conditions demonstrated largely monomeric expression as a doublet (~26 kDa) with a small amount of dimer (~52 kDa) (Figure 2A). Reducing conditions yielded an expected single monomeric band. Given the embedded location of the cysteine, any in vivo dimerization is most likely via non-covalent bonding.19
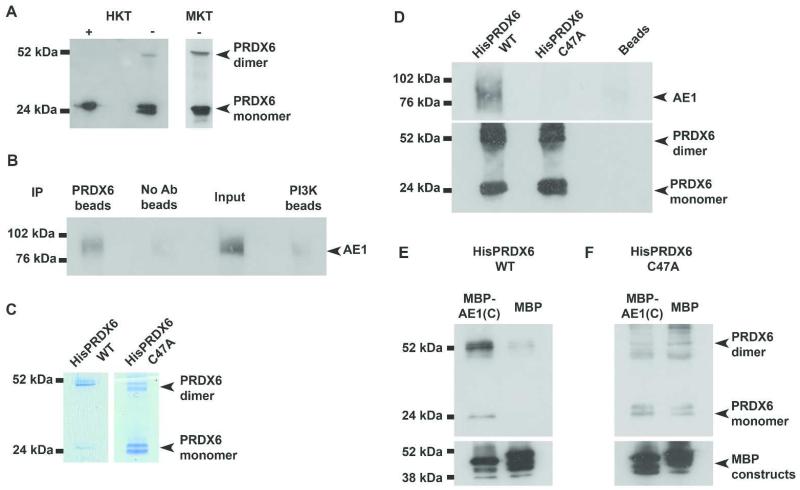
Figure 2. In vitro and ex vivo experimental evidence of PRDX6/AE1 interaction and loss of interaction with mutagenized PRDX6.
(A) PRDX6 from solubilized human kidney fraction (HKT) immunoblotted under reducing conditions (+) and non-reducing conditions (−), showing PRDX6 in monomer only and monomer and dimer forms respectively; PRDX6 from mouse kidney fraction (MKT) immunoblotted under non-reducing conditions shows monomer and dimer forms. (B) Co-immunoprecipitation by bead bound PRDX6 antibody of kAE1 from solubilized membrane protein fractions of human kidney. Negative controls (no antibody and PI3K antibody) did not immunoprecipitate kAE1, with the input (human kidney membrane) providing a control for the Western blot. (C) Coomassie stained gel of expressed HisPRDX6 (1.5 μg) and HisPRDX6-C47A (4.4 μg) showing monomer and dimer forms of the protein. (D) Immobilized HisPRDX6-WT and HisPRDX6-C47A proteins incubated with human kidney membrane showed HisPRDX6-WT could precipitate kAE1; HisPRDX6-C47A or beads alone could not. (E-F) Immobilized MBP-AE1(C) was able to directly pull down HisPRDX6-WT (E) indicating binding but this interaction was not replicated with HisPRDX6-C47A (F). MBP alone does not interact with either form of PRDX6.
PRDX6 co-immunoprecipitates with kAE1
In vivo confirmation of the kAE1-PRDX6 association was provided by co-immunoprecipitation (co-IP) of the two from human kidney membrane preparations. The completed IP was probed with Bric 170 (anti-AE1 antibody) and a band was observed in the bead-bound anti-PRDX6 lane (Figure 2B) but not in either of the two control lanes (beads alone and a bead-bound extraneous precipitating antibody (PI3K)). However, although these results supported an in vivo interaction between kAE1 and PRDX6, they could not differentiate between direct and indirect linkage between the two proteins.
Protein expression and HisPRDX6 mutagenesis
Expressed and purified N-terminal histidine-tagged PRDX6 (HisPRDX6) run on a coomassie gel under non-reducing conditions yielded approximately equal ratios of monomeric (~26 kDa) and dimeric (~52 kDa) states (Figure 2C), in contrast to the behaviour of the untagged protein (where monomer prevailed; Figure 2A). Mass spectrometry of all four bands revealed the purified tagged protein in four different oxidation states via the two methionine residues per molecule (Supplementary Figure 2); this likely also accounts for the doublets observed in Figure 2A.
The single conserved cysteine (C47) was next mutated to alanine (HisPRDX6-C47A) and here monomer prevailed over dimer (Figure 2C). In vitro dimerization is potentially influenced by both non-covalent bonding between two HisPRDX6 proteins and by the interaction between N-terminal His tags.24
Binding of PRDX6 to AE1 confirmed by pulldown
To further support the interaction between AE1 and PRDX6, a pulldown using purified bead-bound HisPRDX6 to precipitate kAE1 from a human kidney membrane protein fraction was also performed, showing that HisPRDX6 precipitates kAE1. When mutagenized HisPRDX6-C47A was used, a loss of binding to kAE1 was demonstrated (Figure 2D). Beads alone provided a negative control.
Direct protein-protein interaction
The direct interaction between purified HisPRDX6 and expressed and purified N-terminal maltose binding protein-tagged AE1(C) (MBP-AE1(C)) was assessed by pulldown using the two recombinant proteins under non-reducing conditions. Bead bound MBP-AE1(C) or MBP alone were incubated with HisPRDX6, and precipitated proteins were analyzed by Western blotting using anti-PRDX6. The same blot was re-probed for MBP as a loading control. As with the previous in vitro assay, HisPRDX6 was seen largely in its dimeric form (Figure 2E). Again mutagenized HisPRDX6-C47A showed a loss of binding (Figure 2F). Supplementary Figure 3 confirms the successful expression of AE1-MBP, although partial degradation or aggregation of MBP cannot be excluded in these preparations.
Thus both indirect and direct pulldown techniques demonstrated loss of kAE1 binding to the mutant HisPRDX6. Given the known inaccessible location of C47 within the protein,19 it is likely that non-covalent binding between kAE1 and the tertiary PRDX6 structure is disrupted by the C47A mutation.
PRDX6 and kAE1 expression variation in Prdx6+/+ and Prdx6−/− mice
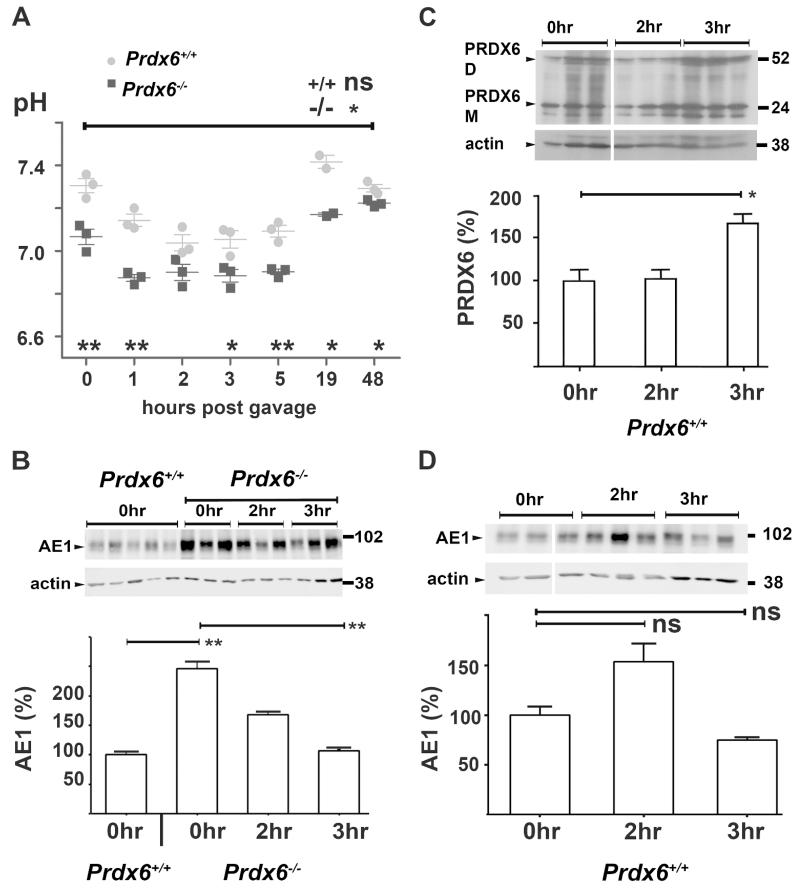
Prdx6−/− mice are reported as having no readily evident phenotype under ambient conditions25 but males have reduced fertility.26 Here we found that systemic pH in knockout mice was lower than that of wild type animals (+/+, 7.31 ± 0.03; −/− 7.07 ± 0.03, p<0.01; Figure 3A, t=0), with elevated pCO2 (Supplementary Table 1) indicating a respiratory component. This was accompanied by significantly more kAE1 protein at baseline in Prdx6−/− animals than wild-type animals (Figure 3B, t=0 hr), implying some, but not sufficient, attempt at renal compensation. Since kAE1 is critically involved in pH homeostasis, we next investigated the ability of these animals to handle oral acid loading with NH4Cl,27 examining kAE1 and/or PRDX6 expression. Both Prdx6+/+ and Prdx6−/− animals responded to acid loading with a drop in pH (Figure 3A); the nadir was reached more rapidly (by 1 hr) in Prdx6−/− than in Prdx6+/+ animals (2 hr post-gavage). At 3 hr post gavage, PRDX6 expression was significantly increased in Prdx6+/+ animals (Figure 3C), while a transient but non-significant rise in kAE1 expression had resolved (Figure 3D). Interestingly, kAE1 levels dropped significantly at 3 hr post gavage in the Prdx6−/− mice (Figure 3B).
Figure 3. PRDX6 and kAE1 expression studies in acidified Prdx6+/+ and Prdx6−/− mice.
(A) pH of terminally collected whole-blood from Prdx6+/+ (grey circles) and Prdx6−/− (black squares) animals at time points after acid gavage. (B) Baseline kAE1 expression (normalized to actin and relative to WT as 100% at t=0hr) is significantly greater in Prdx6−/− animals compared to Prdx6+/+ animals and at 3hrs post gavage, Prdx6−/− animals have significantly decreased expression. (C) PRDX6 expression (combined monomer and dimer normalized to actin relative to t=0hr) is maximally increased in Prdx6+/+ animals 3 hr post gavage. (D) kAE1 expression in Prdx6+/+ animals is not significantly changed. (all time points refer to hours post gavage, error bars represent ± standard error, * = p<0.05, ** = p<0.01)
Additionally, at 48 hours post-gavage, Prdx6+/+ animals had returned to their baseline pH; however, Prdx6−/− mice significantly overshot and failed to return to their baseline pH (Figure 3A). Taken together, these results suggest that PRDX6 plays a role in protecting kAE1 during metabolic acidification and works within the molecular pathways that maintain and restore pH.
Cellular changes in acidified Prdx6+/+ and Prdx6−/− mice
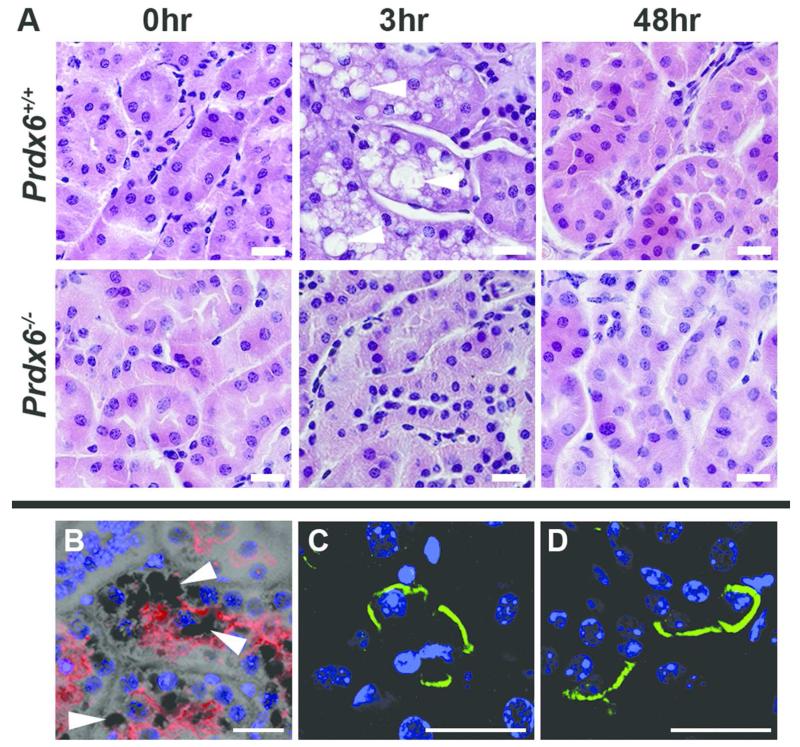
In support of this, when wild-type mice were acidified, widespread cellular vacuolation developed in their kidneys (Figure 4A, white arrows), which resolved when pH normalized, but in the knockout animals (n=3) at the same time points, vacuolation was minimal. Staining with megalin showed vacuolation to be in the proximal tubule and not in other nephron segments (Figure 4B) with ~30% of proximal tubules affected. Vacuolation did not conform to known organelles including lysosomes and autophagosomes (Supplementary Figure 4A-E). Apoptosis was not detected by TUNEL staining (data not shown) and there was no detectable activated Caspase-3 in either genotype by 3 hrs post gavage (Supplementary Figure 4F,G). Staining for AE1 in wild type and knockout mouse kidney revealed that it is maintained in a basolateral location in the absence of PRDX6 (Figure 4C and D respectively).
Figure 4. Cellular changes in acidified adult Prdx6+/+ and Prdx6−/− mouse kidney.
(A) H+E stained mouse renal cortex of Prdx6+/+ and Prdx6−/− at baseline, 3 hours and 48 hours after acid gavage showing tubular vacuolation in Prdx6+/+ animals (white arrowheads) which later resolved. This vacuolation was not evident in the cortex of Prdx6−/− animals after the same treatment. (B) Megalin staining (red) confirmed the proximal tubular location of the vacuoles (white arrowheads). AE1’s basolateral location (green) was similarly maintained in Prdx6+/+ (C) and Prdx6−/− (D) animals. (Scale bar = 20 μm)
siRNA knockdown of PRDX6 leads to loss of kAE1 in mammalian cells
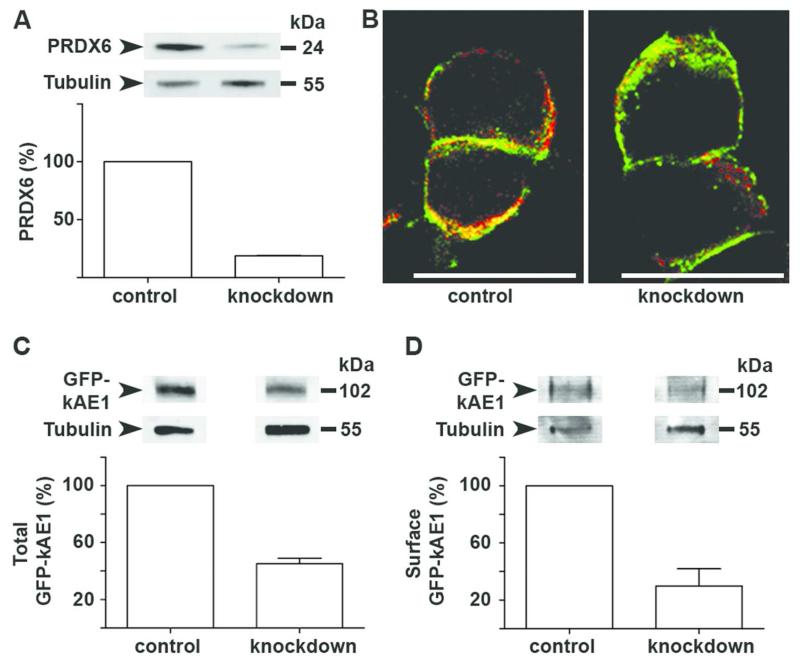
HEK-ΔpMEP-eGFP-kAE1 cells stably express GFP–tagged kAE1 evenly on the plasma membrane.14 Using a specific human siRNA for PRDX6, ~80% knockdown of expression was achieved whereas a control oligo had no effect (Figure 5A); the knockdown did not cause a difference in viability between the treatment groups. Immunolocalization following knockdown showed preservation of kAE1 appearances (Figure 5B) but total levels of kAE1 were lower in knockdown cells than control cells (Figure 5C). Surface biotinylation confirmed that surface kAE1 was depleted in approximately the same ratio as total kAE1 (Figure 5D). These data indicate that PRDX6 is required to maintain the amount of kAE1 present in the cell but does not affect its trafficking to the cell membrane.
Figure 5. PRDX6 knockdown in mammalian cells.
(A) Stably expressing HEK-ΔpMEP-eGFP-kAE1 cells transfected with siRNA against PRDX6 (knockdown) or negative control siRNA (control) over 48 hours show ~80% knockdown of PRDX6 protein. (B) Surface membrane localization of GFP-tagged kAE1 (green) was preserved in both control and knockdown cells and colocalization (yellow) with PRDX6 (red) was observed; scale bars = 20 μm. (C) Total GFP-kAE1and (D) cell surface GFP-kAE1 are reduced by similar amounts when PRDX6 is knocked down. Tubulin was used for normalization, with control oligo results set at 100%, and blots are representative of 3 experiments. (Error bars indicate SEM)
Discussion
This study includes the first data concerning PRDX6 in native human kidney. To date, most work regarding PRDX6 function has been in tissues other than the kidney and/or in rodents; rodent renal PRDX6 expression was recognized in 200122, 28 and there is evidence for PRDX6 up-regulation in renal cancer.29 While expression of all members of the PRDX superfamily has been identified within the distal nephron of rats,23, 28 our yeast two-hybrid screen only elicited PRDX6 as a potential binding partner for AE1. As well as ex vivo and in vitro experimental verification of an interaction between PRDX6 and the C-terminus of AE1, our work demonstrates a role for PRDX6 in maintenance of acid-base homeostasis, in which AE1 also plays an important part.
Unlike the other peroxiredoxins, PRDX6 does not form a homodimer through a disulfide bond of the single conserved cysteine; any in vitro dimers are rather either through hydrogen bonds or intermolecular interactions between residues in β-strands of the two monomers.19, 21 Our results from ex vivo kidney bear this out, with a minimal amount of dimerization observed. We wished to better understand the role of the C47 residue, a known catalytic site; thus it was mutagenized to alanine in a His-tagged PRDX6 construct. This did dimerize in vitro, whether or not the cysteine was present, further confirming previous assertions that dimerization is not via C47. The likeliest explanation for this difference is in vitro interaction between tags,24 however we cannot exclude the possibility that removing a cysteine residue may cause sufficient conformational change to affect intermolecular interactions. Importantly, complete loss of PRDX6-AE1 interaction was observed when the cysteine was absent, in both in vitro and ex vivo experiments. This implies that C47 is required for normal folding of a separate binding moiety in the PRDX6 structure. Very recently, PRDX6 structural differences based upon PLA2 versus peroxidase catalytic activity have also been shown, indicating flexibility of PRDX6 structure.30
For in vivo understanding, we pursued murine studies. Prdx6−/− mice have a resting acidosis that is largely respiratory, with correspondingly a greater expression of kAE1 compared to Prdx6+/+ animals. These results are in line firstly with PRDX6’s known expression in the lung, and secondly with the role of kAE1 to reabsorb bicarbonate into the blood to increase pH.
Exogenously induced metabolic acidosis in knockout animals led to a drop in kAE1 expression, but from a higher baseline than in WT animals. This suggests firstly that AE1 is also interacting with other stabilising protein(s) enabling it to contribute to compensation for chronic acidosis, and secondly that in the acute situation, PRDX6 stabilizes kAE1, so without PRDX6, kAE1 expression cannot be maintained under increasingly acidotic conditions. Systemic pH in fact overshot in Prdx6−/− compared to +/+ animals following acute acidification, again suggesting that normal compensatory mechanisms are not in place. We were not surprised that Prdx6−/− animals showed a baseline increase in AE1 abundance, since many other interacting and chaperone proteins are involved in the response to acidosis. It is known that on the basis of an initially normal acid-base status, AE1 increases at the cell surface in response to an induced acidosis via cellular trafficking and its total amount trends (not significantly) to an increase.31, 32 However, quantitative responses to the situation in our mice, where there was already an underlying respiratory acidotic component, do not appear in the literature for comparison. We speculate that the difference in kinetics between PRDX6 WT and KO animals is due to an abnormal cellular stress response. It is also likely that PRDX6 may interact with other proteins in the response pathway.
In support of PRDX6 playing a stabilizing role, siRNA knockdowns of PRDX6 in HEK cells stably expressing kAE1 demonstrated a total reduction of kAE1 expression but a similarly proportionate amount localizing to the cell surface as in control cells. Unfortunately, further correlations between in vitro and physiologic data could not be made because the buffered pH of cell culture media (7.5-8) is significantly different from the chronically acidic systemic pH in the mouse model and we could not mimic this in vitro.
Whether it is PRDX6’s phospholipase or peroxidase activity that is important for kAE1 maintenance in intercalated cells during an acid challenge is unfortunately unanswerable in the absence of an intercalated cell line in which to explore it, although either mechanism is plausible. In addition, purified native or recombinant enzyme in vitro generally lacks GSH peroxidase activity because of oxidation of its single conserved cysteine.21 Maximal PRDX6 enzymatic activity has been shown to be pH dependent with PLA2 activity optimal at pH 4 versus peroxidase activity at pH 7 – 7.4.33 Despite this difference, it has been suggested that it is PLA2 activity that is important for function.18 However, others indicate that PRDX6 peroxidase activity is crucial for gene expression via ROS stabilization.26, 34 kAE1 stabilization could either be due to PRDX6 stabilizing the cell membrane through PLA2 activity, which would explain kAE1 expression decreasing in acidified mice, or decreased kAE1 expression could be due to decreased gene expression resulting from increased ROS with decreased pH.
Histological changes were noted in the kidneys of acutely acidified Prdx6+/+ animals that were not found in Prdx6−/− mice. Neither strain showed cellular vacuolation at baseline; however, after NH4Cl treatment, Prdx6+/+ mice showed considerable amounts of proximal tubular cellular vacuolation, which did not develop in knockout animals. The vacuoles did not conform to any known organelles or lipid storage vacuoles. Cellular vacuolization is a known response to various forms of cellular stress35, 36 and addition of NH4Cl to chick fibroblast cells causes reversible vacuolation with no effect on functionality.36 These authors suggested a mechanism involving sequestration of base left from NH4Cl dissociation, to minimise alkalinization of the cytosol. Additionally, NH4Cl inhibits the maturation of lysosomes, which would normally sequester hydrogen ions.37 Thus, another possibility is that vacuoles directly sequester hydrogen ions. However, in this scenario, we would have expected vacuolation to be present at baseline in −/− animals but possibly not to worsen. A further hypothesis would be that vacuoles sequester oxidized glutathione in an effort to maintain the cell’s reducing capacity, a phenomenon which is seen in yeast,38 but which the lack of PRDX6 would render inefficient.
In summary these data, together with earlier studies of ours and others, suggest that while PRDX6 cannot be the sole AE1 stabilising protein (such that AE1 can contribute to compensation for chronic acidosis in Prdx6−/− animals), in the acute situation, PRDX6 plays a direct or indirect role in stabilising kAE1. Thus without PRDX6, kAE1 expression cannot be maintained under increasingly acidotic conditions and the recovery from acidosis is also compromised.
Materials and Methods
Yeast Two-Hybrid Assay
AE1(C)WT (residues 876-911) or AE1(C)Δ11 (residues 876-900) were used as bait protein to screen the Pre-Transformed Human Kidney Matchmaker cDNA Library (Clontech) as previously described.13 Confirmation of potential interaction with PRDX6 was assessed by mating back to the original bait of either AE1(C)WT, AE1(C)Δ11 or a control vector (p53). Confirmed clones were sequenced and identified through BLAST searches (blast.ncbi.nlm.nih.gov).
Histological and Immunofluorescence Staining
With ethical approval from the Cambridge Biomedical Research Centre Tissue Bank, 10% formalin fixed, paraffin wax-embedded normal human kidney discarded following tumour removal was obtained. After de-paraffinization and rehydration in an ethanol series, proteinase K antigen retrieval was undertaken (50 μg/ml) for 6 minutes at room temperature. Following washing and blocking, primary antibodies were applied overnight at 4°C (anti-PRDX6 (Abcam 59543) at 1:500; Bric 170 (IBGRL 9450) at 1:100). Fluorochrome conjugated secondary antibodies (1:200) were applied for 1hr at room temperature. After mounting with Vector Shield Mounting Medium (Vector Laboratories) slides were examined using a Zeiss LSM880 confocal microscope.
Mouse kidneys were fixed in 10% neutral buffer formalin, paraffin-embedded and sectioned before hematoxylin/eosin staining. Primary antibodies for immunolocalisation were against megalin (sc-16478) at 1:200, LC3 (NB100-2220 Novus) at 1:500, LAMP2 (ab13524) at 1:50, calreticulin (ab92516) at 1:250, perilipin (cs9349) at 1:250 and EEA1 (ab2900) at 1:500, PRDX6 (ab16947 or ab59543) at 1:500, AE1 (gift of C. Wagner) at 1:1000. TUNEL staining was carried out using a colorimetric TUNEL analysis kit (DeadEnd, Promega). The amount of vacuolation was assessed by counting the percentage of megalin positive tubules that displayed vacuolation in a 1000 μm2 area of cortex.
Co-Immunoprecipitation
Protein A beads were incubated with either PRDX6 antibody (Abcam 16947), PI3K antibody (Abcam 86714) as the control antibody or no antibody in solubilization buffer (20 mM Tris, pH 7.4, 150 mM NaCl, 1% NP40, 2 mM PMSF, EDTA-free Protease Inhibitor Cocktail (Roche), 10% glycerol) overnight at 4°C. Human kidney total protein fraction was solubilized with the solubilization buffer (30 mins, 4°C) before centrifugation. Recovered supernatant, after addition of 20% PEG, was incubated with the bead samples at 4°C for 4 h. The beads were washed with Buffer A (20 mM Tris, pH 7.4, 5 mM NaN3, 0.1% n-Nonyl-β-D-glucopyranoside (n-NDG)) and Buffer B (Buffer A + 500 mM NaCl). Bound proteins were eluted from beads by incubating in SDS sample buffer [0.175 M Tris/HCl (pH 6.8), 5.14% SDS, 18% glycerol, 0.3 M DTT, 0.006% bromophenol blue] for 5 min at 95°C. Samples were subjected to SDS/PAGE and Western blotting before probing with Bric 170.
Expression of Wild Type and mutagenized HisPRDX6
PRDX6 coding sequence inserted into a modified pRSET-A vector (N-terminal His tag, gift from J. Clarke) was used to express full-length N-terminal His-tagged PRDX6 (HisPRDX6) in BL21 cells according to standard methods. Expressed proteins were visualized following purification on Ni-NTA resin and Western Blotting of SDS-PAGE gels. The procedure was repeated with a construct created using the QuikChange Site-Directed Mutagenesis kit (Stratagene) to alter the 47th residue in the PRDX6 coding sequence from cysteine to alanine (HisPRDX6-C47A).
Mass Spectrometry
HisPRDX6 was separated on a 12% SDS-PAGE gel under non-reducing conditions and stained with InstantBlue (Expedeon). The bands were excised and individually trypsin digested. The peptides were analyzed with a MALDI-TOF-TOF mass spectrometer (ABI 4800 model, Applied Biosystems) and identified using the Mascot database.
Expression of MBP and MBP-AE1(C)
pMAL-c4X vector alone was used to express MBP; and AE1(C) (residues 876-911) coding sequence inserted into pMAL-c4X polylinker vector (N-terminal MBP tag) was used to express full-length N-terminal MBP-tagged AE1(C) (MBP-AE1(C)) in BL21 cells according to standard methods. Expressed proteins were visualized following purification on maltose resin and Western Blotting of SDS-PAGE gels (Supplementary Figure 3).
HisPRDX6 Pulldown
Equal moles of HisPRDX6 wild type (WT) or HisPRDX6-C47A were immobilized on Ni-NTA beads. Human kidney total fractions were solubilized with solubilization buffer (10 mM Tris, pH 7.4, 1 mM EDTA, 1 mM DTT, 1.5% n-NDG, 10% glycerol) for 30 mins at 4°C prior to centrifugation (Beckman Ultracentrifuge, 100,000 × g, 1 h, 4°C). The solubilized kidney fractions were added to the bead bound fusion proteins and incubated at 4°C overnight. The beads were washed with Buffer A (1× PBS, 0.1% Triton X-100) and Buffer B (Buffer A + 500 mM NaCl). Bound proteins were eluted from beads by incubating in SDS sample buffer for 5 min at 95°C and analyzed by SDS-PAGE and Western blot using Bric 170.
Direct MBP-AE1(C) and HisPRDX6 Pulldowns
Equal moles of MBP-AE1(C) or MBP alone were incubated with amylose resin for 3+ hours at 4°C. Equivalent amounts of HisPRDX6 ligand were added to beads with bound MBP-AE1(C) or MBP. NaCl (final concentration of 250 mM) was added to each tube. The combined constructs were incubated overnight at 4°C. The beads were washed (20 mM Tris-HCl, pH 7.4 buffer) containing 250-500 mM NaCl. Bound proteins were eluted from beads by incubating in SDS sample buffer for 5 min at 95°C and analyzed by Western blot using anti-PRDX6 antibody (Abcam 16947). Blots were reprobed with anti-MBP (NEB 8032S) to verify loading.
Acute acid challenge Prdx6+/+ and Prdx6−/− animals
Groups of adult Prdx6+/+ and Prdx6−/− mice were maintained on standard diets with ad lib access to food and water. Animals were gavage-fed either water or 0.28M NH4Cl (0.8 g NH4Cl/kg of body weight in a maximum volume of 10 mL/kg). After gavage, groups of mice were terminally anesthetized and arterial blood collected at various time points and blood analyzed on an i-STAT portable clinical analyzer (Abbott).
Protein Fractionation and Western Blotting
Mouse kidneys were homogenized in 500 μl of cold homogenization buffer (10 mM Tris pH 7.4, 150 mM NaCl, 1 mM EGTA, 1 mM EDTA, 2 mM PMSF, and protease inhibitors) followed by centrifugation and collection of the total protein fraction (supernatant). Lysate concentration was estimated using a BioRad protein binding assay (BioRad) and subjected to SDS/PAGE and Western blot. Primary antibodies used were PRDX6 (Abcam 16947), AE1 (gift of C. Wagner) or Caspase-3 (9665 Cell Signalling) with actin (Abcam 6276) as a loading control. PRDX6 analysis was carried out by combining the density of both the monomer and dimer forms relative to actin loading control.
Protein expression in mammalian cells, knockdown, biotinylation and staining
Stable HEK-ΔpMEP-GFP-kAE1 cells were cultured on tissue culture plates (for knockdown) or glass cover slips (for staining) in DMEM (Sigma) supplemented with 10% FBS, penicillin (100 U/ml)/streptomycin (100 μg/ml), and L-glutamine (2mM) at 37°C with a 5% CO2 atmosphere in a humidified incubator. GFP-kAE1 expression was induced with the addition of 100 mM ZnCl2 and 2 mM CdCl2. Endogenous PRDX6 expression was depleted with a siRNA oligonucleotide (117286; Ambion). The oligonucleotide, or a nonspecific siRNA (Silencer Negative Control No. 1 siRNA; 4611 Ambion) were transfected into the cells at a final concentration of 50 nM using Lipofectamine RNAiMAX reagents (Invitrogen). 48 hours after transfection the cells were evaluated by Western blot of total cell lysates or biotinylation of the plasma membrane using a Cell Surface Protein Isolation Kit (Pierce) followed by Western blot. Antibodies used were Bric170, PRDX6 (Abcam 16947) and α-tubulin (Sigma T9026) as a loading control. For immunofluorescence the cells were fixed in 4% paraformaldehyde and then permeabilized with PBS containing 0.1% Triton X-100. After blocking in 1% BSA the cells were incubated with PRDX6 (Abcam16947) for 1hr and after washing a fluorochrome conjugated secondary antibody was applied. The slides were mounted with Vectashield medium (Vector Laboratories).
Supplementary Material
Acknowledgements
Human kidney sections were prepared by Suzy Haward, Addenbrooke’s Human Research Tissue Bank, which is supported by the Cambridge Biomedical Research Centre. We thank Dr. Aron Fisher (Institute for Environmental Science, University of Pennsylvania) for the kind gift of Prdx6−/− mice and reagents, Carsten Wagner (Zurich) for antisera, Jane Clarke (University of Cambridge) for modified pRSET-A vector and Kamburapola Jayawardena for mass spectrometry (CIMR). This work was funded by the Wellcome Trust (award 088489/Z/09/Z to FEKF and Strategic award 100140/Z/12/Z to the Cambridge Institute for Medical Research), and the Jack Kent Cooke Foundation (scholarship to SLS).
Footnotes
Disclosure
No author has any disclosures.
Supplementary information is available at Kidney International’s website.
Part of this work was presented in Abstract form at the 2014 American Society of Nephrology meeting.
References
- 1.Alper SL. Molecular physiology and genetics of Na+-independent SLC4 anion exchangers. J Exp Biol. 2009;212:1672–1683. doi: 10.1242/jeb.029454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Brosius FC, 3rd, Alper SL, Garcia AM, et al. The major kidney band 3 gene transcript predicts an amino-terminal truncated band 3 polypeptide. J Biol Chem. 1989;264:7784–7787. [PubMed] [Google Scholar]
- 3.Wang CC, Moriyama R, Lombardo CR, et al. Partial characterization of the cytoplasmic domain of human kidney band 3. J Biol Chem. 1995;270:17892–17897. doi: 10.1074/jbc.270.30.17892. [DOI] [PubMed] [Google Scholar]
- 4.Ding Y, Casey JR, Kopito RR. The major kidney AE1 isoform does not bind ankyrin (Ank1) in vitro. An essential role for the 79 NH2-terminal amino acid residues of band 3. J Biol Chem. 1994;269:32201–32208. [PubMed] [Google Scholar]
- 5.Drenckhahn D, Merte C. Restriction of the human kidney band 3-like anion exchanger to specialized subdomains of the basolateral plasma membrane of intercalated cells. Eur J Cell Biol. 1987;45:107–115. [PubMed] [Google Scholar]
- 6.Janoshazi A, Ojcius DM, Kone B, et al. Relation between the anion exchange protein in kidney medullary collecting duct cells and red cell band 3. J Membr Biol. 1988;103:181–189. doi: 10.1007/BF01870948. [DOI] [PubMed] [Google Scholar]
- 7.Zhang D, Kiyatkin A, Bolin JT, et al. Crystallographic structure and functional interpretation of the cytoplasmic domain of erythrocyte membrane band 3. Blood. 2000;96:2925–2933. [PubMed] [Google Scholar]
- 8.Karet FE, Gainza FJ, Gyory AZ, et al. Mutations in the chloride-bicarbonate exchanger gene AE1 cause autosomal dominant but not autosomal recessive distal renal tubular acidosis. Proc Natl Acad Sci U S A. 1998;95:6337–6342. doi: 10.1073/pnas.95.11.6337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Quilty JA, Cordat E, Reithmeier RA. Impaired trafficking of human kidney anion exchanger (kAE1) caused by hetero-oligomer formation with a truncated mutant associated with distal renal tubular acidosis. Biochem J. 2002;368:895–903. doi: 10.1042/BJ20020574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Devonald MA, Smith AN, Poon JP, et al. Non-polarized targeting of AE1 causes autosomal dominant distal renal tubular acidosis. Nat Genet. 2003;33:125–127. doi: 10.1038/ng1082. [DOI] [PubMed] [Google Scholar]
- 11.Toye AM, Banting G, Tanner MJ. Regions of human kidney anion exchanger 1 (kAE1) required for basolateral targeting of kAE1 in polarised kidney cells: mis-targeting explains dominant renal tubular acidosis (dRTA) J Cell Sci. 2004;117:1399–1410. doi: 10.1242/jcs.00974. [DOI] [PubMed] [Google Scholar]
- 12.Fry AC, Su Y, Yiu V, et al. Mutation conferring apical-targeting motif on AE1 exchanger causes autosomal dominant distal RTA. J Am Soc Nephrol. 2012;23:1238–1249. doi: 10.1681/ASN.2012020112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Su Y, Blake-Palmer KG, Fry AC, et al. Glyceraldehyde 3-phosphate dehydrogenase is required for band 3 (anion exchanger 1) membrane residency in the mammalian kidney. Am J Physiol Renal Physiol. 2011;300:F157–166. doi: 10.1152/ajprenal.00228.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Su Y, Al-Lamki RS, Blake-Palmer KG, et al. Physical and functional links between anion exchanger-1 and sodium pump. J Am Soc Nephrol. 2015;26:400–409. doi: 10.1681/ASN.2013101063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Duangtum N, Junking M, Sawasdee N, et al. Human kidney anion exchanger 1 interacts with kinesin family member 3B (KIF3B) Biochem Biophys Res Commun. 2011;413:69–74. doi: 10.1016/j.bbrc.2011.08.050. [DOI] [PubMed] [Google Scholar]
- 16.Vince JW, Reithmeier RA. Carbonic anhydrase II binds to the carboxyl terminus of human band 3, the erythrocyte C1−/HCO3− exchanger. J Biol Chem. 1998;273:28430–28437. doi: 10.1074/jbc.273.43.28430. [DOI] [PubMed] [Google Scholar]
- 17.Sawasdee N, Junking M, Ngaojanlar P, et al. Human kidney anion exchanger 1 interacts with adaptor-related protein complex 1 mu1A (AP-1 mu1A) Biochem Biophys Res Commun. 2010;401:85–91. doi: 10.1016/j.bbrc.2010.09.015. [DOI] [PubMed] [Google Scholar]
- 18.Chen JW, Dodia C, Feinstein SI, et al. 1-Cys peroxiredoxin, a bifunctional enzyme with glutathione peroxidase and phospholipase A2 activities. J Biol Chem. 2000;275:28421–28427. doi: 10.1074/jbc.M005073200. [DOI] [PubMed] [Google Scholar]
- 19.Choi HJ, Kang SW, Yang CH, et al. Crystal structure of a novel human peroxidase enzyme at 2.0 A resolution. Nat Struct Biol. 1998;5:400–406. doi: 10.1038/nsb0598-400. [DOI] [PubMed] [Google Scholar]
- 20.Wood ZA, Schroder E, Robin Harris J, et al. Structure, mechanism and regulation of peroxiredoxins. Trends Biochem Sci. 2003;28:32–40. doi: 10.1016/s0968-0004(02)00003-8. [DOI] [PubMed] [Google Scholar]
- 21.Manevich Y, Feinstein SI, Fisher AB. Activation of the antioxidant enzyme 1-CYS peroxiredoxin requires glutathionylation mediated by heterodimerization with pi GST. Proc Natl Acad Sci U S A. 2004;101:3780–3785. doi: 10.1073/pnas.0400181101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Fujii T, Fujii J, Taniguchi N. Augmented expression of peroxiredoxin VI in rat lung and kidney after birth implies an antioxidative role. Eur J Biochem. 2001;268:218–225. doi: 10.1046/j.1432-1033.2001.01843.x. [DOI] [PubMed] [Google Scholar]
- 23.Lee JW, Chou CL, Knepper MA. Deep Sequencing in Microdissected Renal Tubules Identifies Nephron Segment-Specific Transcriptomes. J Am Soc Nephrol. 2015 doi: 10.1681/ASN.2014111067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wu J, Filutowicz M. Hexahistidine (His6)-tag dependent protein dimerization: a cautionary tale. Acta Biochim Pol. 1999;46:591–599. [PubMed] [Google Scholar]
- 25.Wang X, Phelan SA, Forsman-Semb K, et al. Mice with targeted mutation of peroxiredoxin 6 develop normally but are susceptible to oxidative stress. J Biol Chem. 2003;278:25179–25190. doi: 10.1074/jbc.M302706200. [DOI] [PubMed] [Google Scholar]
- 26.Ozkosem B, Feinstein SI, Fisher AB, et al. Advancing age increases sperm chromatin damage and impairs fertility in peroxiredoxin 6 null mice. Redox Biol. 2015;5:15–23. doi: 10.1016/j.redox.2015.02.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wrong O, Davies HE. The excretion of acid in renal disease. Q J Med. 1959;28:259–313. [PubMed] [Google Scholar]
- 28.Oberley TD, Verwiebe E, Zhong W, et al. Localization of the thioredoxin system in normal rat kidney. Free Radic Biol Med. 2001;30:412–424. doi: 10.1016/s0891-5849(00)00486-x. [DOI] [PubMed] [Google Scholar]
- 29.Sun CY, Zang YC, San YX, et al. Proteomic analysis of clear cell renal cell carcinoma. Identification of potential tumor markers. Saudi Med J. 2010;31:525–532. [PubMed] [Google Scholar]
- 30.Rivera-Santiago RF, Harper SL, Zhou S, et al. Solution Structure of the Reduced Form of Human Peroxiredoxin-6 Elucidated Using Zero-Length Chemical Cross-linking And Homology Modeling. Biochem J. 2015 doi: 10.1042/BJ20141463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sabolic I, Brown D, Gluck SL, et al. Regulation of AE1 anion exchanger and H+-ATPase in rat cortex by acute metabolic acidosis and alkalosis. Kidney Int. 1997;51:125–137. doi: 10.1038/ki.1997.16. [DOI] [PubMed] [Google Scholar]
- 32.Purkerson JM, Tsuruoka S, Suter DZ, et al. Adaptation to metabolic acidosis and its recovery are associated with changes in anion exchanger distribution and expression in the cortical collecting duct. Kidney Int. 2010;78:993–1005. doi: 10.1038/ki.2010.195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Manevich Y, Shuvaeva T, Dodia C, et al. Binding of peroxiredoxin 6 to substrate determines differential phospholipid hydroperoxide peroxidase and phospholipase A(2) activities. Arch Biochem Biophys. 2009;485:139–149. doi: 10.1016/j.abb.2009.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kubo E, Hasanova N, Tanaka Y, et al. Protein expression profiling of lens epithelial cells from Prdx6-depleted mice and their vulnerability to UV radiation exposure. Am J Physiol Cell Physiol. 2010;298:C342–354. doi: 10.1152/ajpcell.00336.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Merski JA, Meyers MC. Light- and electron-microscopic evaluation of renal tubular cell vacuolation induced by administration of nitrilotriacetate or sucrose. Food Chem Toxicol. 1985;23:923–930. doi: 10.1016/0278-6915(85)90109-7. [DOI] [PubMed] [Google Scholar]
- 36.Yang WC, Strasser FF, Pomerat CM. Mechanism of drug-induced vacuolization in tissue culture. Exp Cell Res. 1965;38:495–506. doi: 10.1016/0014-4827(65)90373-3. [DOI] [PubMed] [Google Scholar]
- 37.Imort M, Zuhlsdorf M, Feige U, et al. Biosynthesis and transport of lysosomal enzymes in human monocytes and macrophages. Effects of ammonium chloride, zymosan and tunicamycin. Biochem J. 1983;214:671–678. doi: 10.1042/bj2140671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Noctor G, Mhamdi A, Queval G, et al. Regulating the redox gatekeeper: vacuolar sequestration puts glutathione disulfide in its place. Plant Physiol. 2013;163:665–671. doi: 10.1104/pp.113.223545. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.