Abstract
Distinctive patterns of chromatin modification control gene expression and define cellular identity during development and cell differentiation. Polycomb repressive complex 2 (PRC2), the sole mammalian enzymatic complex capable of establishing gene-repressive high-degree methylation of histone H3 at lysine 27 (H3K27), plays crucial roles in regulation of normal and malignant hematopoiesis. Recently, increasing evidence has indicated that recurrent gain-of-function mutation and overexpression of EZH2, the catalytic subunit of PRC2, drive and promote malignant transformation such as B-cell lymphomagenesis, providing a rationale for PRC2 inhibition as a novel anticancer strategy. Here, we summarize the recently developed strategies for inhibition of PRC2, which include a series of highly specific, highly potent, small-molecule inhibitors of EZH2 and EZH1, an EZH2-related methyltransferase. PRC2 establishes functional crosstalk with numerous epigenetic machineries during dynamic regulation of gene transcription. Perturbation of such functional crosstalk caused by genetic events observed in various hematologic cancers, such as inactivation of SNF5 and somatic mutation of UTX, confers PRC2 dependence, thus rendering an increased sensitivity to PRC2 inhibition. We discuss our current understanding of EZH2 somatic mutations frequently found in B-cell lymphomas and recurrent mutations in various other epigenetic regulators as novel molecular predictors and determinants of PRC2 sensitivity. As recent advances have indicated a critical developmental or tumor-suppressive role for PRC2 and EZH2 in various tissue types, we discuss concerns over potentially toxic or even adverse effects associated with EZH2/1 inhibition in certain biological contexts or on cancer genetic background. Collectively, inhibition of PRC2 catalytic activity has emerged as a promising therapeutic intervention for the precise treatment of a range of genetically defined hematologic malignancies and can be potentially applied to a broader spectrum of human cancers that bear similar genetic and epigenetic characteristics. Published by Elsevier Inc. on behalf of ISEH - International Society for Experimental Hematology.
Chromatin modification provides a fundamental means for regulating DNA accessibility in a wide range of DNA-templated processes such as gene transcription and DNA damage repair [1]. At the very least, the regulatory mechanisms for defining distinctive chromatin states include DNA methylation [2], posttranslational modification of histones [3], ATP-dependent chromatin remodeling [4], and utility of histone variants [5]. Deregulation of these chromatin regulatory mechanisms is directly involved in oncogenesis, including hematopoietic malignancies [3,4,6–9].
Polycomb repressive complex 2 (PRC2) probably represents the only enzymatic machinery capable of catalyzing the high-level methylation of histone H3 at lysine 27 (H3K27me3/2) in mammalian cells [10,11]. H3K27 trimethylation (H3K27me3) strongly associates with transcriptional repression [10,11]. Extensive studies have indicated that PRC2 and, by extension, H3K27me3 play critical roles in cell fate determination during development including hematopoietic lineage specification [12–15]. Biochemically, PRC2 uses either enhancer of zeste homologue 2 (EZH2) or a related homolog (EZH1) as the catalytic subunit, and other core components such as EED and SUZ12 are requisite for forming a stable and enzymatically active methyltransferase complex [10,11]. The DNA- and histone-binding properties harbored by PRC2 core components [10,11] and several PRC2-associated proteins [16–20] or noncoding RNAs [21] have been found to regulate PRC2’s target specificity and mediate its chromatin association and/or spreading on chromatin. Hyperactivation of PRC2 caused by mutation or overexpression of EZH2 has been identified as a frequent genetic event among B-cell lymphomas [22–24]. In the next sections, we focus on our current understanding of the lymphoma-associated EZH2 mutations and recent exciting advances in developing EZH2/1-specific inhibitors as a novel molecularly targeted therapy. Because of the high degree of crosstalk between PRC2 and other chromatin regulatory machineries, we also cover recent findings of novel molecular determinants yielding sensitivity to PRC2 inhibition and discuss the potential applications for small-molecule PRC2 inhibitors as therapies in several genetically defined hematologic cancers.
Lymphoma-associated gain-of-function mutation of EZH2
During germinal-center (GC) B-cell development, EZH2 exhibits a dynamic expression pattern: it is massively upregulated when B cells undergo rapid proliferation and immunoglobulin affinity maturation on immune activation, and decreased after completion of these processes [14,25], suggesting a key role for EZH2 in GC development. EZH2 is expressed in a wide range of B-cell neoplasms including Burkitt’s lymphoma, mantle cell lymphomas (MCLs), follicular lymphoma (FL), and diffuse large B-cell lymphomas (DLBCLs) [26,27]. It is upregulated in MCLs as compared with the normal tissue counterparts from which the lymphoma derived [28]. EZH2 deregulation indeed associates with B-cell lymphomagenesis [26], with its expression levels positively correlated with aggressiveness and unfavorable prognosis [29,30]. The importance of PRC2 in lymphomagenesis is further strengthened by the recent identification of recurrent, heterozygous missense mutations of EZH2 in B-cell lymphomas of GC origin such as DLBCLs and FLs. These mutations specifically target the catalytic SET domain of EZH2. The most prevalent is a point mutation of the Tyr641 residue found mutated to either Asn, Phe, Cys, Ser, or His (Fig. 1A, the numeration of EZH2 amino acids based on a short isoform [NCBI Accession No. Q15910.2]) in ~10%–20% of DLBCLs and FLs [22,31,32]; two other rare EZH2 mutations, A677G and A687V (Fig. 1A), were reported in about ~1%–3% of B-cell lymphoma cases [23,31–33].
Figure 1.

Gain-of-function EZH2 mutations affect substrate specificity of the PRC2 complex. (A) Depiction of EZH2 and EZH1 domain structure with the site of gain-of-function mutations (either the hotspot Y641 mutation, A667G, or A687V) in the catalytic SET domain of EZH2 highlighted with a yellow asterisk. (B) Wild-type EZH2 is more efficient at catalyzing the turnover of H3K27me0 and H3K27me1 than H3K27me2 (shown in inset, black line). Y641N (blue bars) and A677G (green bars) exhibit the opposite trend. A687V (purple bars) is equipotent at catalyzing the turnover of H3K27me1 and H3K27me2, relative to WT EZH2. WT = wild-type. (Color version of figure is available online.)
Biochemically, lymphoma-associated EZH2 mutations alter substrate specificity of the PRC2 complex [31,34,35]. EZH2 can induce mono-, di-, and trimethylation of H3K27 (i.e., H3K27me1/2/3), with H3K27me3/2 most strongly associated with gene silencing. Kinetic studies in vitro indicate that PRC2 complexes assembled by the wild-type EZH2 (i.e., PRC2-EZH2WT) have the greatest catalytic efficiency for converting nonmethylated H3K27 (H3K27me0) to monomethylated H3K27 (H3K27me1) and exhibit diminished efficiency for subsequent (H3K27me1 to H3K27me2 and H3K27me2 to H3K27me3) reactions [31,34,35] (Fig. 1B, inset). In contrast, PRC2 complexes bearing a lymphoma-associated EZH2 Tyr641 hotspot mutation such as Y641N (i.e., PRC2-EZH2Y641N) display very limited ability to methylate nonmethylated H3K27, but once H3K27 is monomethylated, they can catalyze the turnover of H3K27me1 to H3K27me2 and, then, much more rapidly catalyze the H3K27me2-to-H3K27me3 reaction [31,34,35] (Fig. 1B, blue bars). The enzymatic differences between PRC2-EZH2WT and PRC2-EZH2Y641N require cooperation to produce the abnormally high level of H3K27me3 seen in the lymphoma cells [31,34,35]; a similar phenomenon was observed for other Tyr641 mutations such as Y641F [31,34,35]. Two other rare EZH2 SET domain mutations, A677G and A687V (Fig. 1A), also affect the substrate specificity of PRC2; however, they have kinetic properties that differ from those of the Y641 mutants [31,32,36,37]. For example, the A677G mutation leads to a form of EZH2 that is capable of efficiently methylating all the H3K27 substrates (Fig. 1B, green bars). Contrarily, the A687V mutation yields a PRC2 complex that is equipotent at methylating H3K27me1 and H3K27me2, but like PRC2-EZH2Y641N, is incapable of methylating H3K27me0 [31,32,36,37] (Fig. 1B, purple bars). Collectively, lymphoma-associated mutations of EZH2 are gain-of-function mutations and induce PRC2 hyperactivity through distinct molecular mechanisms, leading to a globally elevated H3K27me3 phenotype seen in patient-derived lymphoma cells [26]. The reduced ability to methylate H3K27me0 by EZH2 bearing the hotspot mutation provides an explanation for the exclusively heterozygous mutation pattern observed in lymphoma patients [22,31,32]. A potentially additional aspect of this “cooperative” model is existence of EZH1 in vivo, to which we believe the model can be extended. In the future, generation of a murine EZH2 Y641-mutated knockin model that faithfully recapitulates the human disease would be very useful for proving the “cooperative” model genetically.
Homology modeling [31,34] and the recently solved apo structure of the EZH2 SET domain [38,39] have provided mechanistic insights into the aforementioned trends in substrate specificity that was seen with the EZH2 gain-of-function mutants. Y641 is believed to be important for both recognizing and limiting the H3K27 methylation states. Specifically, the ε-amino lysine nitrogen of the H3K27 substrate is within proximity of the phenolic oxygen of Y641 to create a hydrogen bond (Fig. 2A, dashed line, 2.9 Å) [31,34]. This hydrogen bond is maintained through successive steps of mono- and dimethylation and is believed to be part of the reason EZH2WT prefers to stop methylation with an H3K27me2 product [31]. The functional relevance of this hydrogen bond is manifold [31]: first, it is important for engaging H3K27me0 as substrate, because all Y641 mutants possess little catalytic activity with unmethylated substrates (Fig. 1B, H3K27me0); second, Y641 helps create a constrained “pocket” that reduces the ability of H3K27me2 to rotate and accept a third methyl group from the methyl donor (Fig. 2A). A mutation that results in a smaller amino acid at the Y641 position, such as EZH2Y641N (Fig. 2B), changes both the hydrogen bonding ability and local geometry of the substrate-binding pocket. EZH2Y641N is not as proficient at hydrogen bonding with H3K27, because the mutated N641 form is no longer within the acceptable range to form a hydrogen bond [31] (Fig. 2B, dashed line, 5.6 Å). Furthermore, as the pocket dimensions change, the dimethylated lysine of the H3K27me2 substrate is able to rotate more freely, which accelerates the transfer of an additional methyl group. Similar effects were observed for other EZH2 Y641 mutations (Y641 to F, C, S, or H), but the most drastic effects were seen with EZH2Y641N [31].
Figure 2.
Mechanistic insights into substrate selectivity of EZH2 mutants. (A) Wild-type EZH2 uses Y641 to control the methylation state of H3K27 by maintaining a hydrogen bond (dotted line, 2.9 Å) with the dimethylated substrate (green). SAH (purple), the methylation product of cofactor SAM, and the SAM-competitive EZH2 inhibitor UNC1999 (light blue) are also included in the model to illustrate their overlapped binding sites. (B) Mutation to Y641N enlarges the pocket and reduces the propensity for a hydrogen bond between the asparagine and H3K27me2 (dotted line, 5.6 Å), promoting trimethylation of H3K27. (C) Mutation of A677 to the smaller glycine residue (A677G) leads to a slight conformational change, enlarging the pocket and weakening the hydrogen bond between Y641 and H3K27me2 (dotted line, 4.2 Å), which enhances the ability of EZH2A677G to trimethylate H3K27. (D) Mutation of A687 to valine (A687V) leads to a similar conformational change, which leads to a slightly larger pocket, and reduces the strength of the hydrogen bond between Y641 and H3K27me2 (dotted line, 3.2 Å). SAM = S-adenosylmethionine.
Structural modeling also provided an explanation for the altered substrate preference associated with the EZH2 A677G and A687V mutations. A677 and A687 are two residues that surround the substrate recognition groove and appear to play no direct role in substrate recognition (Fig. 2C, D); however, their mutation leads to a change in geometry of the substrate binding groove, which alters the kinetics of the corresponding PRC2 complex. This change in geometry has two effects on substrate recognition [31,37]: first, both mutations result in a slightly larger pocket, which allows for more rotation of the lysine residue; second, the phenolic oxygen of Y641 is shifted further away from H3K27, resulting in a less stable hydrogen bond, which reduces the ability of EZH2 to recognize the methylation state (Fig. 2C and D, 3.2 Å and 4.2 Å). Because of the smaller effect on pocket dimensions seen in the A677G and A687V mutants, they are less effective in generating H3K27me3 than Y641N mutants (Fig. 1B, H3K27me2 bars) [31], but EZH2A677G is as effective as EZH2WT at methylating H3K27me0 (Fig. 1B, green bars). A recent electron microscopy analysis of PRC2 complexes has indicated that the EZH2 SET domain can form intracomplex interactions with other PRC2 components [40]. Therefore, further investigations aimed at solving the structural details of PRC2 complexes will allow a better understanding of PRC2/EZH2 enzymology and its lymphoma-associated mutations.
Development of PRC2 inhibitors
As EZH2 overexpression is frequently found in various human cancers [3], EZH2 and PRC2 are proposed as attractive drug targets for cancer therapy, which has inspired several pharmaceutical companies to endeavor on high-throughput screening campaigns for inhibitors of EZH2. Independent efforts by Epizyme, GlaxoSmithKline (GSK), and Novartis have revealed a structurally similar, cofactor-competitive, small molecule, which potently suppresses EZH2-catalyzed H3K27me3 [41–47] (Table 1). These independent hits were then optimized via traditional medicinal chemistry to enhance the drug-likeness of these compounds.
Table 1.
Structures of the disclosed small-molecule inhibitors of EZH2, as well as their activity against EZH2, either wild-type (WT) or mutant, and EZH1a
| Compound | Structure | In vitro activity, IC50 (nmol/L)b
|
In vivo activity? | Orally bioavailable? | Fold selectivity for EZH2 | ||||
|---|---|---|---|---|---|---|---|---|---|
| WT | Y641N | A677G | A687V | EZH1 | |||||
| EPZ005687 |

|
54 | 41 | 10 | ND | 2713 | No | No | 50 |
| GSK126 |

|
10 | 29 | 24 | 441 | 680 | Yes | No | 150 |
| GSK343 |

|
4 | ND | ND | ND | 240 | No | No | 60 |
| El1 |

|
9 | ND | ND | ND | 1340 | No | No | 140 |
| UNC1999 |

|
10 | 46 | ND | ND | 45 | Yes | Yes | 5 |
| EPZ6438 |

|
11 | 38 | 2 | 2 | 392 | Yes | Yes | 35 |
| Constellation Compound 3 |

|
21 | 197 | ND | ND | 213 | No | No | 10 |
ND = not determined.
Compounds appear in the order they were disclosed in the literature. All compounds except Constellation compound 3 bear a dimethylpyridone motif that is requisite for activity. Additionally, all compounds remain active against EZH2 gain-of-function mutants tested and are fairly selective for EZH2 over EZH1, with UNC1999 displaying the most EZH1 inhibition. GSK126 exhibited the first activity in vivo, and UNC1999 was the first EZH2/1 inhibitor to exhibit oral bioavailability.
IC50 values were calculated on the basis of the Cheng–Prusoff equation for competitive inhibitors.
The first two published EZH2 inhibitors are EPZ005687 [42] and GSK126 [41] (Table 1), which potently inhibit wild-type and lymphoma-associated mutants of EZH2 and are highly selective for EZH2 over a range of unrelated methyltransferases. Although EPZ005687 from Epizyme was not suitable for in vivo studies, it was an essential tool compound for target validation and overall study of EZH2 biology. GSK126 from GSK exhibited in vivo potency via intraperitoneal administration [41]. These compounds share very similar pharmacophoric features and are fairly selective for EZH2 versus EZH1, the only EZH2-related enzyme (50- to 150-fold selectivity [Table 1]), indicating the high specificity of these compounds. Soon after the disclosure of GSK126, GSK343 was published with similar potency against EZH2, and differs from GSK126 in that it contains an indazole core and several different substitutions such as the piperazine-substituted pyridine (Table 1). El1 (Table 1) is a compound that was optimized from a hit of a high-throughput screening campaign at Novartis [43] and has structural features and selectivity similar to those of EPZ005687 and GSK126, but did not have any in vivo activity. UNC1999 (Table 1) represents the first orally bioavailable inhibitor of EZH2 and is the most panactive EZH2/1 inhibitor to date [44]. UNC1999 is similar in structure to GSK343, differing only in the substitution of the pyridine and the capping group of the piperazine. However, this small modification has a large effect on the pharmacokinetic properties of the compound and EZH2/1 selectivity. EZH1 compensates the function of EZH2 [48,49] and emerges as a regulator of hematopoietic neoplasms [50,51]. This EZH2/1 panactivity poses a potential advantage for UNC1999 in cancer cells that rely on both EZH enzymes such as MLL-rearranged acute leukemia. Epizyme also released an orally bioavailable derivative of EPZ005687—EPZ-6438 [45] (Table 1). Unlike previously published compounds that contain a bicyclic core, EPZ-6438 has a phenyl core. The only non–pyridone-containing inhibitor of EZH2 is Constellation Pharmaceuticals compound 3 (Table 1) [46]. This compound is not suitable for in vivo studies and has about tenfold selectivity for EZH2 versus EZH1. All of these inhibitors exhibit high potency and high selectivity toward wild-type EZH2 and its lymphoma-associated mutant forms [41–47] (Table 1). In addition, enzymology assays revealed that all of the EZH2/1 inhibitors block EZH2 enzymatic activity through a cofactor S-adenosylmethionine (SAM)-competitive mechanism (Fig. 2A; inhibitor in purple overlapped with SAM shown in cyan), rather than disruption of PRC2 complex formation or alteration of PRC2 protein stability.
Targeting EZH2 hyperactivity for B-cell lymphoma therapy
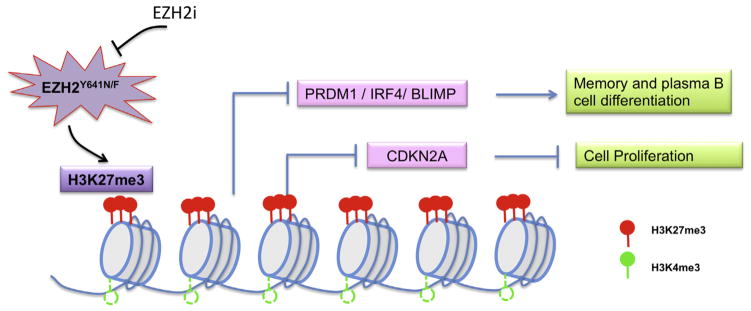
Recent studies have indicated that acquisition of EZH2 gain-of-function mutations indeed perturbs normal B-cell development and promotes lymphomagenesis in animal models [14,15,53,54]. Mechanistically, EZH2 gain-of-function mutations cause increased H3K27me3 occupancy and transcriptional repression of critical genes associated with B-cell differentiation, such as BLIMP1, IRF4, and PRDM1, as well as the cell cycle regulator genes CDKN2A and CDKN1A/1B [14,15,53,54] (Fig. 3). Thus, lymphoma-associated EZH2 mutations are supposed to reinforce B cells at an immature and proliferative state and represent ideal molecular targets. Indeed, early success has been achieved using highly selective EZH2 inhibitors for treatment of GC-cell lymphomas bearing EZH2 gain-of-function mutations [15,41–47]. Studies with GSK126 [41], EPZ005687, and EPZ-6438 [42,55] indicate their preferential effectiveness in suppressing growth of EZH2-mutated lymphomas in comparison to those with wild-type EZH2. However, B-cell studies performed in DLBCLs [15], MCLs, and Burkitt’s lymphoma [47,56] indicate that overexpression of EZH2 may confer a similar PRC2 addiction, arguing a general sensitivity to EZH2 inhibitors regardless of EZH2 mutations. Besides cell-based studies, early success with EZH2-selective inhibitors in vivo was also achieved in treatment of xenografted DLBCL models in which GSK126 was well tolerated [41]. It remains to be evaluated by several ongoing clinical studies whether the selective PRC2 inhibitors can provide clinical benefits to lymphoma patients. It is, however, noteworthy that the first two patients reported as responders in a recent Epizyme clinical trial are actually EZH2 wild type, with a mediastinal B-cell lymphoma patient reportedly bearing no germinal center features [57]. These unexpected findings raise questions regarding what genetic markers in these responder cases render sensitivity to EZH2 inhibitors clinically or which of the molecular response predictors identified in preclinical mouse and cell line studies will actually translate to the clinic.
Figure 3.
EZH2 gain-of-function mutations in driving B-cell lymphomagenesis. H3K27me3 and H3K4me3 often coexist at multiple genomic loci in germinal-center B cells, rendering these genes in a repressive but poised status. EZH2 gain-of-function mutants reinforce H3K27me3 occupancy and repression of the “bivalent genes” that are crucial for antiproliferation (such as CDKN2A) or terminal differentiation (such as PRDM1, IRF4, and BLIMP), promoting hyperplasia or cancerous transformation of germinal-center B cells.
Collectively, PRC2 inhibition represents a promising way to treat B-cell lymphomas, especially those DLBCLs, FLs, and MCLs that exhibit PRC2 dependency because of EZH2 somatic mutation or overexpression.
Crosstalk between PRC2 and other chromatin regulatory pathways
Evidence from different cellular and organismal systems has revealed complex crosstalk between PRC2 and other chromatin machineries. PRC2 activity is influenced by a myriad of cooperative and antagonistic interactions, allowing dynamic regulation of PRC2 target genes and the landscape of H3K27me3. It can be anticipated that certain genetic events during cancer development may affect these crosstalk and genetic interactions, thus conferring a “synthetic” or collateral sensitivity to PRC2 or EZH2 blockade.
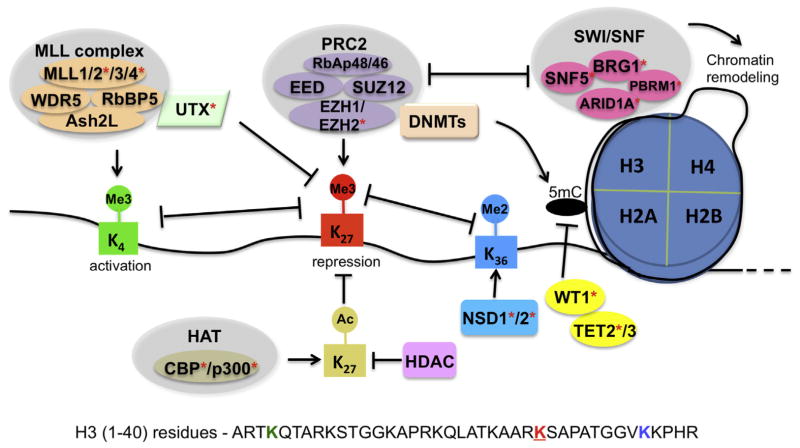
Several mechanisms were found to reverse or interfere with PRC2 enzymatic activity (Fig. 4). While PRC2 establishes or “writes” H3K27me3 on chromatin, the KDM6 family of H3K27 demethylases such as UTX removes or “erases” the mark by converting H3K27me3 to the low-or unmethylated H3K27 [58]. Furthermore, catalysis of H3K27me3 is influenced by the surrounding chromatin environment, such as the existence of other histone modifications [59,60] and nucleosomal density [61]. Methylation of histone H3 Lys 4 (H3K4me) and methylation of histone H3 Lys 36 (H3K36me) are two prominent histone marks that demarcate the promoter and gene body regions, respectively, of actively transcribed genes [3,62]. Pre-existence of H3K4me3 or H3K36me2/3 on nucleosomes directly inhibits enzymatic activities of PRC2 [59,60]. In addition, because a single lysine cannot be simultaneously modified by methylation and acetylation at the ε-amino group, acetylation of H3K27 (H3K27ac) by histone acetyltransferases such as p300 and CBP also strongly suppresses addition of H3K27me3 by PRC2 [63,64]. Thus, the existence of active histone marks (H3K4me3, H3K36me3/2, H3K27ac) and associated enzymatic machineries directly inhibits PRC2-mediated H3K27me3 establishment (Fig. 4).
Figure 4.
Crosstalk between PRC2 and other epigenetic factors. PRC2 is the sole methyltransferase complex capable of catalyzing H3K27me3 to induce and enforce gene repression. Various epigenetic machineries have crosstalk with PRC2 complex, either cooperatively or antagonistically. The KDM6 family of lysine demethylases such as UTX removes H3K27me3. Methylations of H3K4 and H3K36, as well as acetylation of H3K27, are prominent histone marks associated with transcriptional activation, which are established by the MLL complexes, the NSD family proteins, and the CBP/p300 acetyltransferases, respectively; preinstallation of H3K4me3, H3K36me2/3, and/or H3K27ac inhibits the activity of PRC2 and interferes with H3K27me3 installation. UTX physically interacts with the MLL complex linking H3K4 methylation with H3K27 demethylation. SWI/SNF is an ATP-dependent chromatin remodeling complex that can also antagonize PRC2. PRC2 was found to interact with other epigenetic factors such as histone deacetylases (HDACs) and DNA methyltransferases (DNMTs) to further reinforce a repressive state of polycomb target genes. WT1, however, serves as a platform for recruiting DNA demethylases TET2 and TET3, thus facilitating conversion of 5-methylcytosine (5 mC) to 5-hydroxymethylcytosine (5hmC) and attenuating the DNA methylation-mediated repression. Genetic alterations that affect such a myriad of epigenetic factors (red stars) are identified as the recurrent event in various hematopoietic malignancies, causing global or focal perturbation of PRC2 activity and H3K27me3 and thereby rendering cancer cells sensitive to PRC2 inhibition.
Furthermore, genetic studies in Drosophila have revealed that the SWI/SNF family of chromatin remodeling complexes antagonizes PRC2 (Fig. 4): mutation of SWI/SNF suppressed developmental defects caused by mutation of PRC2 homologues in flies [65]. Although the exact underlying mechanism remains unclear, SWI/SNF remodeling complexes modify DNA accessibility and nucleosomal density [66], a factor known to influence PRC2 enzymatic activity [61]. It has been hypothesized that interplay between SWI/SNF and PRC2 complexes defines a dynamic balance of the chromatin state [67].
Factors also exist that cooperate with PRC2 and promote its function (Fig. 4). For instance, PRC2 physically interacts with the histone deacetylases HDAC1-3 [68], which remove the acetyl group from H3K27ac and thus make the lysine available for methylation by PRC2. PRC2 also associates with de novo DNA methyltransferases [69], and it was reported that H3K27me3 and PRC2 targets are positively correlated to DNA methylation [11,69–71].
Taken together, mounting data have indicated that PRC2 activity and functional outcomes are regulated by additional chromatin contexts. Epigenetic factors that influence PRC2 activity or catalytic outcomes such as UTX [72,73] and MMSET/NSD2 [74,75] were recurrently found mutated in a range of hematopoietic malignancies. We therefore anticipate that these genetic lesions may potentially affect PRC2 and confer tumor cell sensitivity to PRC2 inhibition. In the next sections, we summarize recent studies that have indicated PRC2 inhibition as a promising therapeutic option in several genetically defined hematologic malignancies. These studies have expanded the potential applications of PRC2 inhibitors and will improve the personalized medicine and treatment of hematologic cancers in future.
Genetic lesions that confer PRC2 dependence and sensitivity to PRC2 inhibition
UTX mutation
UTX (also known as KDM6A) is a ubiquitously expressed H3K27 demethylase that antagonizes PRC2 in gene regulation and development [76–78]. Localized on the X chromosome, UTX is one of a limited number of genes known to escape X inactivation in females [79]. Somatic mutations of UTX were frequently identified in a range of tumors including hematologic cancers such as multiple myeloma (MM), acute lymphoblastic leukemia (ALL), and myeloid leukemia [72,73,80–83] (Fig. 5A). These UTX mutations are often loss-of-function gene deletions, truncations and frameshifts, or missense mutations found centered in a hotspot region within the demethylase enzymatic domain [72,80,81]. UTX mutations coexist with the hyperactivation mutation of NOTCH1, one of the most common lesions associated with T-cell ALLs (T-ALLs) [80]. Deletion of UTX significantly accelerated tumor progression in a NOTCH1 mutation-induced T-ALL model [80], whereas restoration of UTX expression in UTX-mutated T-ALL cell lines induced apoptosis and suppressed tumor growth [81]. These observations support a bona fide tumor-suppressive role for UTX. Although UTX deficiency did not change the global levels of H3K27me3 or H3K4me3, it caused a genomewide redistribution of H3K27me3 [72,80,81]. In the NOTCH1 mutation-induced T-ALL model, loss of Utx aberrantly repressed several putative tumor suppressors such as Pcgf2 and Lzts2, which is concurrent with gain in H3K27me3 at their promoters [80] (Fig. 5A). Conversely, gene derepression after UTX restoration in UTX-null tumor lines was associated with a decrease in H3K27me3 occupancy at target genes [72,80], demonstrating direct involvement of UTX-mediated demethylation in dynamic control of critical gene expression and malignant transformation. Considering the aberrant H3K27me3 accumulation and accompanying gene repression at critical location upon UTX mutation, it is reasonable to postulate that UTX-mutated leukemia and lymphoma would be more sensitive to PRC2 inhibition. Indeed, van der Meulen et al. have recently reported that UTX-mutated T-ALL lines are more sensitive to DZNep, a nonspecific methyltransferase inhibitor that causes EZH2 degradation, than UTX wild-type T-ALL lines, despite the similar reduction in H3K27me3 seen in tested lines after compound treatment [80]. Similarly, Ezponda et al. recently reported that GSK343 (Table 1) is more effective in treating UTX-mutant MM lines in comparison to the UTX–wild-type lines [84]. These findings provide rationale for treating UTX-mutated hematopoietic tumors with the more specific PRC2 inhibitors.
Figure 5.
Genetic mutations found in hematopoietic malignancies confer on cancer cells sensitivity to PRC2 or EZH2 inhibition. (A) Although they do not affect the global H3K27me3 level, UTX inactivating mutations cause focal H3K27me3 enrichment in genes involved in antiproliferation and cell adhesion to promote cancerous transformation. (B) Biallelic inactivation of a core component of the SWI/SNF chromatin remodeling complex, SNF5 (also known as SMARCB1), enhances PRC2-mediated suppression of antiproliferation and lineage differentiation signature genes; other subunits of the SWI/SNF complex also function as tumor suppressors and their mutations in malignancies may confer a similar PRC2 dependence. Similarly, PRC2 is also required for acute leukemia caused by MLL rearrangements such as MLL-AF9. In addition to leukemia stem cell programs, MLL-AF9 oncoproteins directly promote expression of EZH2. PRC2 complexes assembled by both EZH2 and EZH1 repress myeloid differentiation and antiproliferation programs to promote acute leukemogenicity. (C) Because of the abnormal chromosomal translocation or gain-of-function somatic mutation (E1099K) found in multiple myeloma patients, hyperactivation of MMSET/NSD2 causes a global increase in H3K36me2 and decrease in H3K27me3. However, H3K27me3 is maintained and even enriched at certain critical regions protected by CTCF, an insulator factor. Focal enrichment of EZH2 leads to an enhanced suppression of lymphoid signatures and miR126, a negative regulator of c-MYC, thus enhancing tumorigenesis. (D) Loss-of-function mutation of WT1 induces a DNA hypermethylation phenotype in patients with acute myeloid leukemia (AML). Methylated genes found specific to WT1-mutated AMLs largely overlap with PRC2 gene targets such as myeloid maturation signatures, indicating cooperation between DNMTs and PRC2. WT1-mutated AMLs exhibit sensitivity to PRC2 inhibition. * = mutation of the labeled gene found in hematologic cancers.
Loss-of-function mutation of the SWI/SNF complex
The mammalian SWI/SNF complex is an ATP-dependent chromatin remodeling complex that is critical for regulating cell differentiation and proliferation [85]. Loss-of-function mutations that “hit” various components of the SWI/SNF complex have been reported in nearly 20% of human cancers, including hematologic cancers, indicating its tumor suppressive role [86,87] (Figs. 4 [stars] and 5B). SNF5 is the first subunit of SWI/SNF functionally linked to PRC2 deregulation during tumorigenesis [88]. Biallelic inactivation of SNF5 exists in a majority of human malignant rhabdoid tumors (MRTs) [89,90] and sporadic cases of T-cell malignancies [91]. Conditional inactivation of Snf5 in mouse peripheral T cells causes completely penetrant CD8+ T-cell lymphomas [88]. It has been found that Snf5 mutation drives tumor progression by disrupting several key pathways related to tumor suppression, differentiation, and cell cycle progression [66]. The mechanisms that underlie functional antagonisms between SNF5 and PRC2 are manifold: first, SNF5 directly downregulates EZH2 gene expression in tumor cells [88]; in addition, the SWI/SNF complex restricts the recruitment and/or spreading of PRC2 at its critical target genes such as the tumor-suppressive locus p16INK4a and numerous lineage differentiation genes [67,88]. PRC2-mediated repression of p16INK4a is counteracted by SNF5, and occupancy of H3K27me3 was elevated at p16INK4a in SNF5-deficient lymphomas [67,88]; restoration of SNF5 in MRT cells caused eviction of polycomb proteins and loss of H3K27me3 at the p16INK4a gene, as well as concomitant increased occupancy of transactivators, leading to its elevated transcription [67] (Fig. 5B). A similar phenomenon was seen at differentiation genes in these SNF5-deficient lymphomas [88]. Therefore, in tumors carrying SNF5 deficiency such as T-cell lymphomas, PRC2 complexes abnormally enforce repression of genes critical for tumor suppression and cell differentiation, keeping cancer cells in a proliferative, undifferentiated state. Indeed, knockdown or deletion of EZH2 in the SNF5-deficient cells reversed p16INK4a repression [88]. These findings indicate PRC2 dependence in SNF5-deficient cancers. A recent study found that blockade of PRC2 activity by an EZH2 inhibitor specifically suppressed growth of SNF5-deleted and not wild-type MRTs, by inducing apoptosis and cell differentiation in the mutant MRTs [45]. As mutations of other SWI/SNF complex components such as BRG1, ARID1A, and PBRM1 (Figs. 4 and 5B, stars) were found in various cancers and are generally less well understood, it would be intriguing to examine if these mutations also confer SWI/SNF loss-of-function and, thus, PRC2/EZH2 dependence. Indeed, two recent studies have reported that BRG1-mutated non–small-cell lung cancer (NSCLC) cells [92] and ARID1A-mutated ovarian cancer cells [93] exhibit sensitivity to GSK126 (Table 1), an EZH2-selective inhibitor, in comparison to cancer cells without such a lesion. Hematopoietic cancers bearing the similar mutations might be equally sensitive to PRC2 inhibition.
MLL gene rearrangement
The mammalian mixed lineage leukemia (MLL) gene encodes an H3K4-specific methyltransferase [3,94]. Rearrangement of MLL is responsible for ~70% of infant acute myeloid, lymphoid, or mixed-lineage leukemias and ~7%–10% of adult cases [94]. Acute leukemia with MLL rearrangement has a poor prognosis with low survival rates, highlighting the need for novel molecularly targeted interventions [95,96]. It has been found that MLL fusion oncoproteins produced by MLL gene rearrangements recruit epigenetic factors and/or transcriptional elongation-promoting complexes such as DOT1L and bromodomain proteins (BRD4) to enforce abnormal activation of oncogenes such as HOX, MEIS1, and c-MYC [3,94–96]. In addition, studies have also found that MLL fusion oncoproteins are recruited to the promoter of EZH2 and promote its transcription [97] (Fig. 5B), indicating a role for PRC2 in MLL-rearranged leukemia. Indeed, several works have found that PRC2 acts in parallel with MLL rearrangements by controlling a distinctive program to sustain leukemogenicity [50,51,98]. In detail, PRC2 directly represses genes that are critical for myeloid differentiation such as EGR1, as well as genes that limit proliferation and self-renewal of hematopoietic cells such as CDKN2A [50,51,98]. In addition, EZH2 was also found to physically interact with C/EBPα, a differentiation-promoting transcription factor, and repressed the C/EBPα-mediated prodifferentiation program [97]. In the MLL-rearranged acute leukemia, both EZH2 and EZH1 are expressed and compensate one another to promote acute leukemogenesis [50,51]. Disruption of both enzymes is required to inhibit growth of leukemia carrying MLL-AF9, a common form of MLL rearrangements [50,51]. UNC1999, the most panactive inhibitor of both EZH1 and EZH2 to date (Table 1), indeed demonstrated a unique growth-suppressing effect on a panel of MLL-rearranged leukemia cells in vitro by inhibiting their repopulating ability and promoting cell differentiation and apoptosis, whereas GSK126, the EZH2-selective inhibitor with much less activity against EZH1 (Table 1), failed to efficiently inhibit H3K27me3 or suppress proliferation of MLL-rearranged leukemia cells [52]. Genomic profiling by ChIP-seq revealed the molecular events following PRC2 enzymatic inhibition: UNC1999 preferentially “erases” H3K27me3 associated with distal regulatory elements such as enhancers and remodels the landscape of H3K27me3 versus H3K27ac at proximal promoters, leading to derepression of numerous PRC2 target genes that include CDKN2A and development/differentiation-related genes [52]. Oral administration of UNC1999 prolonged survival of a well-defined murine leukemia model bearing MLL-AF9, a common form of MLL rearrangement [52]. Therefore, the targeting of both PRC2-EZH2 and PRC2-EZH1 with small-molecule compounds such as UNC1999 represents a new way of treating MLL-rearranged leukemia; a similar tumor-suppressing phenomenon was also observed in MLL-rearranged leukemia using a hydrocarbon-stapled peptide recently developed to disrupt physical interactions of EZH2 and EZH1 with its cofactor EED, which leads to the proteasome-mediated degradation of PRC2 complexes [99].
Translocation and activating mutation of MMSET/NSD2
MMSET (also known as NSD2 or WHSC1) is a histone methyltransferase that catalyzes dimethylation of histone H3 at lysine 36 (H3K36me2), a histone modification associated with transcriptional initiation and/or elongation [100]. Reciprocal t(4;14) translocation, which results in MMSET fusion to the immunoglobulin heavy chain locus and, hence, overexpression of MMSET, was reported in 15%–20% of multiple myeloma patients [101]. This genetic event is believed to drive tumorigenesis of myeloma and is associated with lower survival rates of myeloma patients [102–105]. Consistently, a recurrent gain-of-function mutation of MMSET (E1099 K) was also identified in a subset of ALL patients [74,75] and specifically targets the catalytic SET domain of MMSET. In both cases, the activated MMSET reprograms the landscape of histone modifications, inducing a global increase in H3K36me2 and concurrent decrease in H3K27me3, caused by the antagonistic effect of H3K36me2 on PRC2 and, thus, H3K27 methylation (Figs. 4 and 5C) [74,106,107]. However, despite a genomewide net loss of H3K27me3, this silencing mark is maintained and even enriched further at certain specific loci [106]. Motif analysis suggested that CTCF, a known genome organizer and insulator, is likely to be responsible for preventing these PRC2/H3K27me3 domains from the intrusion of MMSET [106]. This study indicates that although the hyperactivated MMSET restricts the binding and activity of PRC2 at H3K36me2-demarcated loci, it also induces a collateral effect at putative “CTCF-protected” loci because the free pool of PRC2 has nowhere else to bind [106]. As a result, the latter loci end up with enhanced PRC2 binding and a concomitant elevated level of H3K27me3, leading to abnormal repression of the embedded genes [106] (Fig. 5C). Further genomic profiling revealed that genes embedded within the retained PRC2 domains include the GC B cell-associated gene signatures, as well as critical microRNAs such as miR-126, a negative regulator of c-MYC [106] (Fig. 5C), suggesting a critical role for certain newly acquired PRC2-binding sites in promotion of growth and aggressiveness of myeloma cells with MMSET hyperactivation. Indeed, myeloma cells carrying t(4;14) MMSET translocations exhibit more sensitivity to GSK343 (Table 1) than t(4;14)-negative myeloma cells [106]. GSK343 treatment also led to elevated miR-126 levels and concomitant decreased MYC levels. Therefore, although additional studies need to be carried out, PRC2 inhibitors represent a promising therapeutic agent for treatment of myeloma patients with MMSET hyperactivity.
Inactivating mutations of WT1
The Wilms’ tumor 1 (WT1) gene encodes a sequence-specific zinc finger transcription factor that was originally identified as a tumor suppressor in Wilms’ tumor, a rare kidney cancer [108]. WT1 has been linked to regulation of critical differentiation genes in various biological processes, particularly nephrogenesis and hematopoiesis [109]. Heterozygous somatic mutations of WT1 were identified in approximately 10% of cases of acute myeloid leukemia (AML), with the hotspot mutations centered within the DNA-binding zinc finger domains [110,111]. These loss-of-function mutations of WT1 correlate with poor prognosis and chemotherapy resistance in AML patients, but the role of WT1 and oncogenic mechanisms that underlie WT1-mutated AMLs is unclear [111]. Recent studies have linked WT1 mutations to regulation of DNA methylation [112–114] (Figs. 4 and 5D). WT1 physically interacts with the methylcytosine dioxygenases TET2 and TET3, and serves as one of the mechanisms responsible for recruitment of TET2/3 to target loci (Fig. 4), where TET proteins mediate DNA demethylation via conversion of 5-methylcytosine (5 mC) to 5-hydroxymethylcytosine (5hmC) and subsequent oxidative derivatives [112,113]. Inactivated mutants of WT1 in AMLs lack the critical DNA binding domains, leading to a DNA hypermethylation phenotype in the affected AMLs [112–114]. Indeed, introduction of mutant WT1 into WT1–wild-type AML cells is sufficient in inducing DNA hypermethylation, indicating a causal role for WT1 mutation in regulating DNA methylation [114]. Intriguingly, the DNA hypermethlylated loci were found enriched with signature of known PRC2 targets such as differentiation genes [114], indicating cooperation between DNA hypermethylation and PRC2 (Fig. 5D). Consistently, the mutant and not wild-type WT1 blocked myeloid differentiation of hematopoietic stem cells [114]. Moreover, EZH2 is significantly overexpressed in WT1-mutated AMLs compared with WT1–wild-type cases [114] (Fig. 5D). EZH2 knockdown or its blockade with the inhibitor GSK126 promoted myeloid differentiation in AML cells carrying WT1 mutations [114]. These studies have provided rationale for using EZH2 inhibitors in the treatment of WT1-mutated AMLs.
Potential adverse, side, and toxic effects of PRC2 or EZH2 inhibition
Despite enthusiasm shared by the community for EZH2/1 inhibitors as a future cancer therapy, recent studies have raised concerns over the potentially side, toxic, or even adverse effects caused by PRC2/EZH2 blockade.
First, in contrast to the oncogenic function of PRC2 and EZH2 described above, inactivating mutation of EZH2 and other PRC2 components has emerged as a recurrent theme in a range of human cancers, which demonstrates a dichotomous role for PRC2 in different biological contexts and raises concerns over PRC2 and EZH2 inhibition in clinical treatment. Specifically, various damaging mutations, such as biallelic deletion and missense, frameshifting, or truncation mutations, that “hit” either the PRC2 core component (EZH2, EED, and SUZ12) [115–117] or cofactor genes (JARID2 [118] and ASXL1 [119]) are recurrent in a range of myeloid neoplasms such as myelodysplastic syndromes (MDSs), primary myelofibrosis (PMF), myeloproliferative neoplasms (MPNs), and chronic myelomonocytic leukemia (CMML); similar PRC2-inactivating somatic mutations also occur frequently among human T-ALL [120–122] and solid tumors such as malignant peripheral nerve sheath tumors (MPNSTs) [123–126]. These PRC2-inactivating mutations are associated with adverse prognostic outcomes of MDS and PMF patients [127]. The tumor-suppressive role of PRC2 has been verified with several murine cancer models: deletion of Ezh2 alone in hematopoietic systems is sufficient to cause spontaneous T-ALL with an average latency of 150 days in mice [122]; Ezh2 loss also led to development of an MDS/MPN-like phenotype following serial bone marrow transplantation [128]; loss of PRC2 caused aberrant gene transcription programs and significantly enhanced in vivo tumorigenesis driven by mutant RAS signaling [123–126]. These recent findings have collectively indicated that EZH2 and PRC2 act as bona fide tumor suppressors in certain cell lineages or biological contexts.
In addition, EZH2/1 or PRC2 was also reported to be important in normal development. In particular, EZH1 is documented as a crucial factor in maintaining the self-renewal capacity of adult HSCs by protecting them from senescence [13]; similar developmental roles for EZH2 or EZH1 were reported as well in other tissue-specific stem and progenitor cells [129,130]. A dual EZH2 and EZH1 inhibitor may therefore cause greater toxicity by impairing normal functions of the PRC2 assembled by both enzymes.
Collectively, these recent advances have urged the field to re-evaluate the clinical application of PRC2 and EZH2 inhibitors with more caution. A safe therapeutic window needs to be determined to eradicate cancer cells over generation of the unwanted toxic or even adverse effect during treatment. Furthermore, efforts are required to explore the cancer genetic backgrounds that define either beneficial or adverse effects of PRC2/EZH2 inhibition. Indeed, a recent study of NSCLCs indicated that usage of EZH2 inhibitors can cause differential or even opposite effects among NSCLC patients carrying different genetic backgrounds: although EZH2 inhibition sensitizes the BRG1-mutated NSCLCs to etoposide, a topoisomerase II inhibitor, and improves cancer treatment, the same EZH2 inhibitor causes an opposite effect by conferring BRG1 wild-type NSCLC resistance on topoisomerase II inhibitors [92].
Combination therapy and drug resistance
PRC2 inhibitors can be potentially used with other Food and Drug Administration-approved agents, either sequentially or in a combinational therapeutic strategy, to further enhance their therapeutic value. Indeed, pretreatment with EZH2 inhibitor sensitized BRG1-mutated NSCLCs to the topoisomerase II inhibitors, and dual treatment with EZH2 and topoisomerase II inhibitors provided synergy in reversing oncogenicity [92]. A similar and dramatic synergistic antitumor effect was observed when combining EZH2 inhibitors with Food and Drug Administration-approved glucocorticoid receptor agonists such as prednisolone and dexamethasone in the treatment of lymphomas, regardless of EZH2 mutation status [131]. However, the molecular basis for these described synergistic effects remains unclear. Use of a PRC2 inhibitor in combination therapy with other developed agents, for example, inhibitor of histone deacetylase, will also be explored (Fig. 4), because PRC2 and histone deacetylase are known to interact physically in their coordinated action in mediating gene silencing [26].
In addition, it is probably not of a total surprise that tumor cells may develop strategies such as the acquired additional mutations that confer resistance to EZH2/1 inhibitors, because clinically relevant and inhibitor-resistant mutations were found during treatment with inhibitors of oncogenic kinase such as BCR-ABL [132] and B-RAF [133]. Indeed, a very recent cell line–based study identified the two missense mutations of EZH2 that affect compound binding and therefore confer resistance to several tested EZH2 inhibitors [134]. One of such acquired mutations, Y661D, occurs cis in the EZH2 Y641-mutated allele and is in the vicinity of the SAM pocket where these inhibitors are believed to bind (Fig. 2A); however, the other mutation, Y111L, is found in the wild-type EZH2 allele, thus providing additional support for the model of cooperativity between wild-type and mutant EZH2 enzymes. The fact that Y111L is far from the SAM-binding pocket (located between the EED-binding domain and the SANT domain) suggests a long-range interaction potentially via formation of a high-order PRC2 complex for mediating regulation or formation of an active methyltransferase domain of EZH2. Research is required to learn the structural details of EZH2/1 inhibitors bound to EZH2 or the PRC2 complex, which will enable the rational design of new strategies for inhibiting the inhibitor-resistant mutations that tumors can acquire during treatment.
Conclusions
Therapeutic targeting of PRC2 initially emerged from studies reporting that overexpression or hyperactive mutation of EZH2 can initiate, promote, or maintain oncogenesis. The already developed small-molecule inhibitors of PRC2 or EZH2/1 have achieved early success in the laboratory and are currently under clinical evaluation for the treatment of germinal-center B-cell lymphomas bearing EZH2 mutations, which account for about 15%–20% of all cases. Studies have revealed the genetic interaction and functional crosstalk (either cooperative or antagonistic) of PRC2 with a variety of other chromatin modifiers and epigenetic regulators such as SWI/SNF, UTX, and MMSET. Somatic mutations that affect these epigenetic factors have been identified as a recurrent genetic event in a range of hematopoietic malignancies, which cause a collateral dependence of PRC2 and an increased sensitivity to pharmacologic inhibition of PRC2 or EZH2. In addition, increasing evidence has started to reveal the PRC2-independent roles of EZH2/1 in various cancer and biological contexts [135–137], and further investigations shall be extended to studies of their nonhistone substrates [138] and/or noncanonical gene-activation roles [139,140]. Thus, we expect to see an increasing list of such PRC2 or EZH2 dependencies in the future. For example, the trithorax group proteins, such as the MLL family of H3K4-specific methyltransferases and histone acetyltransferases CBP/p300, are known to antagonize PRC2 (Fig. 4) [3], and inactivating somatic mutations of MLL2, MLL4, CBP, and p300 are frequent in lymphomas [23,24,141]. In addition to EZH2 gain-of-function mutations, identification of collateral PRC2 or EZH2 dependency in various hematopoietic malignancies has expanded the potential application of the already developed PRC2 inhibitors and will promote the precision medicine and personalized therapy. However, considering the fact that PRC2 or EZH2 has a tumor-suppressive role with loss-of-function mutations recurrently identified in various human cancers, we need to avoid treating cancer patients with certain genetic lesions and backgrounds with EZH2 or PRC2 inhibitors because such blockade could accelerate cancer progression, as reported in recent studies [92,123–126]. Although ongoing efforts continue to optimize the pharmacokinetic and pharmacodynamic properties of the currently existing PRC2 and EZH2/1 inhibitors, as well as development of second-generation inhibitors to target additional mutations of EZH2 that tumors acquire to produce inhibitor resistance, we remain optimistic and excited about their future clinical applications in hematologic cancer, as well as a broader spectrum of human cancers being unraveled to show PRC2 dependency and, thus, PRC2 inhibitor sensitivity.
Acknowledgments
KDK is supported by an American Chemical Society Medicinal Chemistry Predoctoral Fellowship. JJ is supported by National Institutes of Health Institute of General Medical Sciences Grant R01GM103893. GGW is supported by a National Cancer Institute “Pathway to Independence” Award in Cancer Research (CA151683), a Department of Defense Career Development award (CA130247), and grants from Gabrielle’s Angel Foundation and Concern Foundation. GGW is also a Kimmel Scholar of Sidney Kimmel Foundation for Cancer Research and an American Society of Hematology (ASH) Scholar in Basic Science.
Footnotes
Conflict of interest disclosure
The authors have no conflicting financial interests to disclose.
References
- 1.Strahl BD, Allis CD. The language of covalent histone modifications. Nature. 2000;403:41–45. doi: 10.1038/47412. [DOI] [PubMed] [Google Scholar]
- 2.Schubeler D. Function and information content of DNA methylation. Nature. 2015;517:321–326. doi: 10.1038/nature14192. [DOI] [PubMed] [Google Scholar]
- 3.Chi P, Allis CD, Wang GG. Covalent histone modifications—Miswritten, misinterpreted and mis-erased in human cancers. Nat Rev Cancer. 2010;10:457–469. doi: 10.1038/nrc2876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang GG, Allis CD. ChiChromatin remodeling and cancer: Part II. ATP-dependent chromatin remodeling. Trends Mol Med. 2007;13:373–380. doi: 10.1016/j.molmed.2007.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Weber CM, Henikoff S. Histone variants: Dynamic punctuation in transcription. Genes Dev. 2014;28:672–682. doi: 10.1101/gad.238873.114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shih AH, Abdel-Wahab O, Patel JP, Levine RL. The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer. 2012;12:599–612. doi: 10.1038/nrc3343. [DOI] [PubMed] [Google Scholar]
- 7.Dawson MA, Kouzarides T. Cancer epigenetics: From mechanism to therapy. Cell. 2012;150:12–27. doi: 10.1016/j.cell.2012.06.013. [DOI] [PubMed] [Google Scholar]
- 8.Rodriguez-Paredes M, Esteller M. Cancer epigenetics reaches mainstream oncology. Nat Med. 2011;17:330–339. doi: 10.1038/nm.2305. [DOI] [PubMed] [Google Scholar]
- 9.Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M. Epigenetic protein families: A new frontier for drug discovery. Nat Rev Drug Discov. 2012;11:384–400. doi: 10.1038/nrd3674. [DOI] [PubMed] [Google Scholar]
- 10.Margueron R, Reinberg D. The polycomb complex PRC2 and its mark in life. Nature. 2011;469:343–349. doi: 10.1038/nature09784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Di Croce L, Helin K. Transcriptional regulation by polycomb group proteins. Nat Struct Mol Biol. 2013;20:1147–1155. doi: 10.1038/nsmb.2669. [DOI] [PubMed] [Google Scholar]
- 12.Mochizuki-Kashio M, Mishima Y, Miyagi S, et al. Dependency on the polycomb gene Ezh2 distinguishes fetal from adult hematopoietic stem cells. Blood. 2011;118:6553–6561. doi: 10.1182/blood-2011-03-340554. [DOI] [PubMed] [Google Scholar]
- 13.Hidalgo I, Herrera-Merchan A, Ligos JM, et al. Ezh1 is required for hematopoietic stem cell maintenance and prevents senescence-like cell cycle arrest. Cell Stem Cell. 2012;11:649–662. doi: 10.1016/j.stem.2012.08.001. [DOI] [PubMed] [Google Scholar]
- 14.Caganova M, Carrisi C, Varano G, et al. Germinal center dysregulation by histone methyltransferase EZH2 promotes lymphomagenesis. J Clin Invest. 2013;123:5009–5022. doi: 10.1172/JCI70626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Beguelin W, Popovic R, Teater M, et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell. 2013;23:677–692. doi: 10.1016/j.ccr.2013.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Musselman CA, Gibson MD, Hartwick EW, et al. Binding of PHF1 Tudor to H3K36me3 enhances nucleosome accessibility. Nat Commun. 2013;4:2969. doi: 10.1038/ncomms3969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cai L, Rothbart SB, Lu R, et al. An H3K36 methylation-engaging Tudor motif of polycomb-like proteins mediates PRC2 complex targeting. Mol Cell. 2013;49:571–582. doi: 10.1016/j.molcel.2012.11.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ballare C, Lange M, Lapinaite A, et al. Phf19 links methylated Lys36 of histone H3 to regulation of polycomb activity. Nat Struct Mol Biol. 2012;19:1257–1265. doi: 10.1038/nsmb.2434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pasini D, Cloos PA, Walfridsson J, et al. JARID2 regulates binding of the polycomb repressive complex 2 to target genes in ES cells. Nature. 2010;464:306–310. doi: 10.1038/nature08788. [DOI] [PubMed] [Google Scholar]
- 20.Li G, Margueron R, Ku M, Chambon P, Bernstein BE, Reinberg D. Jarid2 and PRC2, partners in regulating gene expression. Genes Dev. 2010;24:368–380. doi: 10.1101/gad.1886410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Brockdorff N. Noncoding RNA and polycomb recruitment. RNA. 2013;19:429–442. doi: 10.1261/rna.037598.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Morin RD, Johnson NA, Severson TM, et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet. 2010;42:181–185. doi: 10.1038/ng.518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Morin RD, Mendez-Lago M, Mungall AJ, et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature. 2011;476:298–303. doi: 10.1038/nature10351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Okosun J, Bodör C, Wang J, et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet. 2014;46:176–181. doi: 10.1038/ng.2856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Van Galen JC, Dukers DF, Giroth C, et al. Distinct expression patterns of polycomb oncoproteins and their binding partners during the germinal center reaction. Eur J Immunol. 2004;34:1870–1881. doi: 10.1002/eji.200424985. [DOI] [PubMed] [Google Scholar]
- 26.Wang GG, Konze KD, Tao J. Polycomb genes, miRNA, and their deregulation in B-cell malignancies. Blood. 2015;125:1217–1225. doi: 10.1182/blood-2014-10-606822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Van Kemenade FJ, Raaphorst FM, Blokzijl T, et al. Coexpression of BMI-1 and EZH2 polycomb-group proteins is associated with cycling cells and degree of malignancy in B-cell non-Hodgkin lymphoma. Blood. 2001;97:3896–3901. doi: 10.1182/blood.v97.12.3896. [DOI] [PubMed] [Google Scholar]
- 28.Visser HP, Gunster MJ, Kluin-Nelemans HC, et al. The polycomb group protein EZH2 is upregulated in proliferating, cultured human mantle cell lymphoma. Br J Haematol. 2001;112:950–958. doi: 10.1046/j.1365-2141.2001.02641.x. [DOI] [PubMed] [Google Scholar]
- 29.Abd Al Kader L, Oka T, Takata K, et al. In aggressive variants of non-Hodgkin lymphomas, Ezh2 is strongly expressed and polycomb repressive complex PRC1.4 dominates over PRC1.2. Virchows Arch. 2013;463:697–711. doi: 10.1007/s00428-013-1428-y. [DOI] [PubMed] [Google Scholar]
- 30.Van Galen JC, Muris JJF, Oudejans JJ, et al. Expression of the polycomb-group gene BMI1 is related to an unfavourable prognosis in primary nodal DLBCL. J Clin Pathol. 2007;60:167–172. doi: 10.1136/jcp.2006.038752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McCabe MT, Graves AP, Ganji G, et al. Mutation of A677 in histone methyltransferase EZH2 in human B-cell lymphoma promotes hyper-trimethylation of histone H3 on lysine 27 (H3K27) Proc Natl Acad Sci U S A. 2012;109:2989–2994. doi: 10.1073/pnas.1116418109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Majer CR, Lei J, Scott MP, et al. A687V EZH2 is a gain-of-function mutation found in lymphoma patients. FEBS Lett. 2012;586:3448–3451. doi: 10.1016/j.febslet.2012.07.066. [DOI] [PubMed] [Google Scholar]
- 33.Bodör C, Grossmann V, Popov N, et al. EZH2 mutations are frequent and represent an early event in follicular lymphoma. Blood. 2013;122:3165–3168. doi: 10.1182/blood-2013-04-496893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yap DB, Chu J, Berg T, et al. Somatic mutations at EZH2 Y641 act dominantly through a mechanism of selectively altered PRC2 catalytic activity, to increase H3K27 trimethylation. Blood. 2011;117:2451–2459. doi: 10.1182/blood-2010-11-321208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sneeringer CJ, Scott MP, Kuntz KW, et al. Coordinated activities of wild-type plus mutant EZH2 drive tumor-associated hypertrimethylation of lysine 27 on histone H3 (H3K27) in human B-cell lymphomas. Proc Natl Acad Sci U S A. 2010;107:20980–20985. doi: 10.1073/pnas.1012525107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Swalm BM, Knutson SK, Warholic NM, et al. Reaction coupling between wild-type and disease-associated mutant EZH2. ACS Chem Biol. 2014;9:2459–2464. doi: 10.1021/cb500548b. [DOI] [PubMed] [Google Scholar]
- 37.Ott HM, Graves AP, Pappalardi MB, et al. A687V EZH2 is a driver of histone H3 lysine 27 (H3K27) hypertrimethylation. Mol Cancer Ther. 2014;13:3062. doi: 10.1158/1535-7163.MCT-13-0876. [DOI] [PubMed] [Google Scholar]
- 38.Antonysamy S, Condon B, Druzina Z, et al. Structural context of disease-associated mutations and putative mechanism of autoinhibition revealed by X-ray crystallographic analysis of the EZH2-SET domain. PLoS One. 2013;8:e84147. doi: 10.1371/journal.pone.0084147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wu H, Zeng H, Dong A, et al. Structure of the catalytic domain of EZH2 reveals conformational plasticity in cofactor and substrate binding sites and explains oncogenic mutations. PLoS One. 2013;8:e83737. doi: 10.1371/journal.pone.0083737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ciferri C, Lander GC, Maiolica A, Herzog F, Aebersold R, Nogales E. Molecular architecture of human polycomb repressive complex 2. ELife. 2012;1:e00005. doi: 10.7554/eLife.00005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.McCabe MT, Ott HM, Ganil G, et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature. 2012;492:108–112. doi: 10.1038/nature11606. [DOI] [PubMed] [Google Scholar]
- 42.Knutson SK, Wigle TJ, Warholic NM, et al. A selective inhibitor of EZH2 blocks H3K27 methylation and kills mutant lymphoma cells. Nat Chem Biol. 2012;8:890–896. doi: 10.1038/nchembio.1084. [DOI] [PubMed] [Google Scholar]
- 43.Qi W, Chan HM, Teng L, et al. Selective inhibition of Ezh2 by a small molecule inhibitor blocks tumor cells proliferation. Proc Natl Acad Sci U S A. 2012;109:21360–21365. doi: 10.1073/pnas.1210371110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Konze KD, Ma A, Li FL, et al. An orally bioavailable chemical probe of the lysine methyltransferases EZH2 and EZH1. ACS Chem Biol. 2013;8:1324–1334. doi: 10.1021/cb400133j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Knutson SK, Warholic NM, Wigle TJ, et al. Durable tumor regression in genetically altered malignant rhabdoid tumors by inhibition of methyltransferase EZH2. Proc Natl Acad Sci U S A. 2013;110:7922–7927. doi: 10.1073/pnas.1303800110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Garapaty-Rao S, Nasveschuk C, Gagnon A, et al. Identification of EZH2 and EZH1 small molecule inhibitors with selective impact on diffuse large B cell lymphoma cell growth. Chem Biol. 2013;20:1329–1339. doi: 10.1016/j.chembiol.2013.09.013. [DOI] [PubMed] [Google Scholar]
- 47.Bradley WD, Arora S, Busby J, et al. EZH2 Inhibitor efficacy in non-Hodgkin’s lymphoma does not require suppression of H3K27 monomethylation. Chem Biol. 2014;21:1463–1475. doi: 10.1016/j.chembiol.2014.09.017. [DOI] [PubMed] [Google Scholar]
- 48.Shen X, Liu YC, Hsu YJ, et al. EZH1 mediates methylation on histone H3 lysine 27 and complements EZH2 in maintaining stem cell identity and executing pluripotency. Mol Cell. 2008;32:491–502. doi: 10.1016/j.molcel.2008.10.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Margueron R, Li G, Sarma K, et al. Ezh1 and Ezh2 maintain repressive chromatin through different mechanisms. Mol Cell. 2008;32:503–518. doi: 10.1016/j.molcel.2008.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Neff T, Sinha AU, Kluk MJ, et al. Polycomb repressive complex 2 is required for MLL-AF9 leukemia. Proc Natl Acad Sci U S A. 2012;109:5028–5033. doi: 10.1073/pnas.1202258109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Shi J, Wang E, Zuber J, et al. The polycomb complex PRC2 supports aberrant self-renewal in a mouse model of MLL-AF9;Nras(G12D) acute myeloid leukemia. Oncogene. 2013;32:930–938. doi: 10.1038/onc.2012.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Xu B, On DM, Ma A, et al. Selective inhibition of EZH2 and EZH1 enzymatic activity by a small molecule suppresses MLL-rearranged leukemia. Blood. 2015;125:346–357. doi: 10.1182/blood-2014-06-581082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Velichutina I, Shaknovich R, Geng H, et al. EZH2-mediated epigenetic silencing in germinal center B cells contributes to proliferation and lymphomagenesis. Blood. 2010;116:5247–5255. doi: 10.1182/blood-2010-04-280149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Berg T, Thoene S, Yap D, et al. A transgenic mouse model demonstrating the oncogenic role of mutations in the polycomb-group gene EZH2 in lymphomagenesis. Blood. 2014;123:3914–3924. doi: 10.1182/blood-2012-12-473439. [DOI] [PubMed] [Google Scholar]
- 55.Knutson SK, Kawano S, Minoshima Y, et al. Selective inhibition of EZH2 by EPZ-6438 leads to potent antitumor activity in EZH2 mutant non-Hodgkin lymphoma. Mol Cancer Ther. 2014;13:842–854. doi: 10.1158/1535-7163.MCT-13-0773. [DOI] [PubMed] [Google Scholar]
- 56.Zhang X, Zhao X, Fiskus W, et al. Coordinated silencing of MYC-mediated miR-29 by HDAC3 and EZH2 as a therapeutic target of histone modification in aggressive B-cell lymphomas. Cancer Cell. 2012;22:506–523. doi: 10.1016/j.ccr.2012.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 57.Copeland RA, Keilhack H, Italiano A, et al. EZH2 inhibitor EPZ-6438 (E7438) in non-hodgkin lymphoma: Pre-clinical models and early clinical observations. Presented at the annual American Society of Hematology (ASH) meeting; 12 August 2014. [Google Scholar]
- 58.Mosammaparast N, Shi Y. Reversal of histone methylation: Biochemical and molecular mechanisms of histone demethylases. Annu Rev Biochem. 2010;79:155–179. doi: 10.1146/annurev.biochem.78.070907.103946. [DOI] [PubMed] [Google Scholar]
- 59.Schmitges FW, Prusty AB, Faty M, et al. Histone methylation by PRC2 is inhibited by active chromatin marks. Mol Cell. 2011;42:330–341. doi: 10.1016/j.molcel.2011.03.025. [DOI] [PubMed] [Google Scholar]
- 60.Yuan W, Xu M, Huang C, Liu N, Chen S, Zhu B. H3K36 methylation antagonizes PRC2-mediated H3K27 methylation. J Biol Chem. 2011;286:7983–7989. doi: 10.1074/jbc.M110.194027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yuan W, Wu T, Fu H, et al. Dense chromatin activates polycomb repressive complex 2 to regulate H3 lysine 27 methylation. Science. 2012;337:971–975. doi: 10.1126/science.1225237. [DOI] [PubMed] [Google Scholar]
- 62.Barski A, Cuddapah S, Cui K, et al. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129:823–837. doi: 10.1016/j.cell.2007.05.009. [DOI] [PubMed] [Google Scholar]
- 63.Pasini D, Malatesta M, Jung HR, et al. Characterization of an antagonistic switch between histone H3 lysine 27 methylation and acetylation in the transcriptional regulation of polycomb group target genes. Nucleic Acids Res. 2010;38:4958–4969. doi: 10.1093/nar/gkq244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Tie F, Banerjee R, Stratton CA, et al. CBP-mediated acetylation of histone H3 lysine 27 antagonizes Drosophila polycomb silencing. Development. 2009;136:3131–3141. doi: 10.1242/dev.037127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kennison JA, Tamkun JW. Dosage-dependent modifiers of polycomb and antennapedia mutations in Drosophila. Proc Natl Acad Sci U S A. 1988;85:8136–8140. doi: 10.1073/pnas.85.21.8136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Wilson BG, Roberts CW. SWI/SNF nucleosome remodellers and cancer. Nat Rev Cancer. 2011;11:481–492. doi: 10.1038/nrc3068. [DOI] [PubMed] [Google Scholar]
- 67.Kia SK, Gorski MM, Giannakopoulos S, Verrijzer CP. SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol Cell Biol. 2008;28:3457–3464. doi: 10.1128/MCB.02019-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Van der Vlag J, Otte AP. Transcriptional repression mediated by the human polycomb-group protein EED involves histone deacetylation. Nat Genet. 1999;23:474–478. doi: 10.1038/70602. [DOI] [PubMed] [Google Scholar]
- 69.Vire E, Brenner C, Deplus R, et al. The polycomb group protein EZH2 directly controls DNA methylation. Nature. 2006;439:871–874. doi: 10.1038/nature04431. [DOI] [PubMed] [Google Scholar]
- 70.Ohm JE, McGarvey KM, Yu X, et al. A stem cell-like chromatin pattern may predispose tumor suppressor genes to DNA hypermethylation and heritable silencing. Nat Genet. 2007;39:237–242. doi: 10.1038/ng1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Schlesinger Y, Straussman R, Keshet I, et al. Polycomb-mediated methylation on Lys27 of histone H3 pre-marks genes for de novo methylation in cancer. Nat Genet. 2007;39:232–236. doi: 10.1038/ng1950. [DOI] [PubMed] [Google Scholar]
- 72.Van Haaften G, Dalgliesh GL, Davies H, et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat Genet. 2009;41:521–523. doi: 10.1038/ng.349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Mar BG, Bullinger L, Basu E, et al. Sequencing histone-modifying enzymes identifies UTX mutations in acute lymphoblastic leukemia. Leukemia. 2012;26:1881–1883. doi: 10.1038/leu.2012.56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Oyer JA, Huang X, Zheng Y, et al. Point mutation E1099K in MMSET/NSD2 enhances its methyltranferase activity and leads to altered global chromatin methylation in lymphoid malignancies. Leukemia. 2014;28:198–201. doi: 10.1038/leu.2013.204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Jaffe JD, Wang Y, Chan HM, et al. Global chromatin profiling reveals NSD2 mutations in pediatric acute lymphoblastic leukemia. Nat Genet. 2013;45:1386–1391. doi: 10.1038/ng.2777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Agger K, Cloos PA, Christensen J, et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature. 2007;449:731–734. doi: 10.1038/nature06145. [DOI] [PubMed] [Google Scholar]
- 77.Lan F, Bayliss PE, Rinn JL, et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature. 2007;449:689–694. doi: 10.1038/nature06192. [DOI] [PubMed] [Google Scholar]
- 78.Lee MG, Villa R, Trojer P, et al. Demethylation of H3K27 regulates polycomb recruitment and H2A ubiquitination. Science. 2007;318:447–450. doi: 10.1126/science.1149042. [DOI] [PubMed] [Google Scholar]
- 79.Greenfield A, Carrel L, Pennisi D, et al. The UTX gene escapes X inactivation in mice and humans. Hum Mol Genet. 1998;7:737–742. doi: 10.1093/hmg/7.4.737. [DOI] [PubMed] [Google Scholar]
- 80.Van der Meulen J, Sanghvi V, Mavrakis K, et al. The H3K27me3 demethylase UTX is a gender-specific tumor suppressor in T-cell acute lymphoblastic leukemia. Blood. 2015;125:13–21. doi: 10.1182/blood-2014-05-577270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Ntziachristos P, Tsirigos A, Welstead GG, et al. Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia. Nature. 2014;514:513–517. doi: 10.1038/nature13605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Jankowska AM, Makishima H, Tiu RV, et al. Mutational spectrum analysis of chronic myelomonocytic leukemia includes genes associated with epigenetic regulation: UTX, EZH2, and DNMT3A. Blood. 2011;118:3932–3941. doi: 10.1182/blood-2010-10-311019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Kar SA, Jankowska A, Makishima H, et al. Spliceosomal gene mutations are frequent events in the diverse mutational spectrum of chronic myelomonocytic leukemia but largely absent in juvenile myelomonocytic leukemia. Haematologica. 2013;98:107–113. doi: 10.3324/haematol.2012.064048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Ezponda T, Popovic R, Zheng Y. Loss of the histone demethylase UTX contributes to multiple myeloma and sensitizes cells to EZH2 inhibitors. Blood. 2014;124:611. [Google Scholar]
- 85.De la Serna IL, Ohkawa Y, Imbalzano AN. Chromatin remodelling in mammalian differentiation: Lessons from ATP-dependent remodellers. Nat Rev Genet. 2006;7:461–473. doi: 10.1038/nrg1882. [DOI] [PubMed] [Google Scholar]
- 86.Kadoch C, Hargreaves DC, Hodges C, et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nat Genet. 2013;45:592–601. doi: 10.1038/ng.2628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Shain AH, Pollack JR. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS One. 2013;8:e55119. doi: 10.1371/journal.pone.0055119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Wilson BG, Wang X, Shen X, et al. Epigenetic antagonism between polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell. 2010;18:316–328. doi: 10.1016/j.ccr.2010.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Versteege I, Sévenet N, Lange J, et al. Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature. 1998;394:203–206. doi: 10.1038/28212. [DOI] [PubMed] [Google Scholar]
- 90.Biegel JA, Zhou JY, Rorke LB, Stenstrom C, Wainwright LM, Fogelgren B. Germ-line and acquired mutations of INI1 in atypical teratoid and rhabdoid tumors. Cancer Res. 1999;59:74–79. [PubMed] [Google Scholar]
- 91.Yuge M, Nagai H, Uchida T, et al. HSNF5/INI1 gene mutations in lymphoid malignancy. Cancer Genet Cytogenet. 2000;122:37–42. doi: 10.1016/s0165-4608(00)00274-0. [DOI] [PubMed] [Google Scholar]
- 92.Fillmore CM, Xu C, Desai PT, et al. EZH2 inhibition sensitizes BRG1 and EGFR mutant lung tumours to TopoII inhibitors. Nature. 2015;520:239–242. doi: 10.1038/nature14122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Bitler BG, Aird KM, Garipov A, et al. Synthetic lethality by targeting EZH2 methyltransferase activity in ARID1A-mutated cancers. Nat Med. 2015;21:231–238. doi: 10.1038/nm.3799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Dou Y, Hess JL. Mechanisms of transcriptional regulation by MLL and its disruption in acute leukemia. Int J Hematol. 2008;87:10–18. doi: 10.1007/s12185-007-0009-8. [DOI] [PubMed] [Google Scholar]
- 95.Krivtsov AV, Armstrong SA. MLL translocations, histone modifications and leukaemia stem-cell development. Nat Rev Cancer. 2007;7:823–833. doi: 10.1038/nrc2253. [DOI] [PubMed] [Google Scholar]
- 96.Slany RK. The molecular biology of mixed lineage leukemia. Haematologica. 2009;94:984–993. doi: 10.3324/haematol.2008.002436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Thiel AT, Feng Z, Pant DK, et al. The trithorax protein partner menin acts in tandem with EZH2 to suppress C/EBPalpha and differentiation in MLL-AF9 leukemia. Haematologica. 2013;98:918–927. doi: 10.3324/haematol.2012.074195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Tanaka S, Miyagi S, Sashida G, et al. Ezh2 augments leukemogenicity by reinforcing differentiation blockage in acute myeloid leukemia. Blood. 2012;120:1107–1117. doi: 10.1182/blood-2011-11-394932. [DOI] [PubMed] [Google Scholar]
- 99.Kim W, Bird GH, Neff T, et al. Targeted disruption of the EZH2–EED complex inhibits EZH2-dependent cancer. Nat Chem Biol. 2013;9:643–650. doi: 10.1038/nchembio.1331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Wagner EJ, Carpenter PB. Understanding the language of Lys36 methylation at histone H3. Nat Rev Mol Cell Biol. 2012;13:115–126. doi: 10.1038/nrm3274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Keats JJ, Reiman T, Belch AR, Pilarski LM. Ten years and counting: so what do we know about t(4;14)(p16;q32) multiple myeloma. Leuk Lymphoma. 2006;47:2289–2300. doi: 10.1080/10428190600822128. [DOI] [PubMed] [Google Scholar]
- 102.Keats JJ, Reiman T, Maxwell CA, et al. In multiple myeloma, t(4; 14)(p16;q32) is an adverse prognostic factor irrespective of FGFR3 expression. Blood. 2003;101:1520–1529. doi: 10.1182/blood-2002-06-1675. [DOI] [PubMed] [Google Scholar]
- 103.Santra M, Zhan F, Tian E, Barlogie B, Shaughnessy J., Jr A subset of multiple myeloma harboring the t(4;14)(p16;q32) translocation lacks FGFR3 expression but maintains an IGH/MMSET fusion transcript. Blood. 2003;101:2374–2376. doi: 10.1182/blood-2002-09-2801. [DOI] [PubMed] [Google Scholar]
- 104.Avet-Loiseau H, Facon T, Grosbois B, et al. Oncogenesis of multiple myeloma: 14q32 and 13q chromosomal abnormalities are not randomly distributed, but correlate with natural history, immunological features, and clinical presentation. Blood. 2002;99:2185–2191. doi: 10.1182/blood.v99.6.2185. [DOI] [PubMed] [Google Scholar]
- 105.Fonseca R, Blood E, Rue M, et al. Clinical and biologic implications of recurrent genomic aberrations in myeloma. Blood. 2003;101:4569–4575. doi: 10.1182/blood-2002-10-3017. [DOI] [PubMed] [Google Scholar]
- 106.Popovic R, Martinez-Garcia E, Giannopoulou EG, Zhang QW, Zhang QY, Exponda T. Histone methyltransferase MMSET/NSD2 alters EZH2 binding and reprograms the myeloma epigenome through global and focal changes in H3K36 and H3K27 methylation. PLoS Genet. 2014;10:e1004566. doi: 10.1371/journal.pgen.1004566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Zheng Y, Sweet SM, Popovic R, et al. Total kinetic analysis reveals how combinatorial methylation patterns are established on lysines 27 and 36 of histone H3. Proc Natl Acad Sci U S A. 2012;109:13549–13554. doi: 10.1073/pnas.1205707109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Haber DA, Buckler AJ, Glaser T, et al. An internal deletion within an 11p13 zinc finger gene contributes to the development of Wilms’ tumor. Cell. 1990;61:1257–1269. doi: 10.1016/0092-8674(90)90690-g. [DOI] [PubMed] [Google Scholar]
- 109.Huff V. Wilms’ tumours: About tumour suppressor genes, an oncogene and a chameleon gene. Nat Rev Cancer. 2011;11:111–121. doi: 10.1038/nrc3002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.King-Underwood L, Pritchard-Jones K. Wilms’ tumor (WT1) gene mutations occur mainly in acute myeloid leukemia and may confer drug resistance. Blood. 1998;91:2961–2968. [PubMed] [Google Scholar]
- 111.Virappane P, Gale R, Hills R, et al. Mutation of the Wilms’ tumor 1 gene is a poor prognostic factor associated with chemotherapy resistance in normal karyotype acute myeloid leukemia: The United Kingdom Medical Research Council Adult Leukaemia Working Party. J Clin Oncol. 2008;26:5429–5435. doi: 10.1200/JCO.2008.16.0333. [DOI] [PubMed] [Google Scholar]
- 112.Wang Y, Xiao M, Chen X, et al. WT1 recruits TET2 to regulate Its target gene expression and suppress leukemia cell proliferation. Mol Cell. 2015;57:662–673. doi: 10.1016/j.molcel.2014.12.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Rampal R, Alkalin A, Madzo J, et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep. 2014;9:1841–1855. doi: 10.1016/j.celrep.2014.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Sinha S, Thomas D, Yu L, et al. Mutant WT1 is associated with DNA hypermethylation of PRC2 targets in AML and responds to EZH2 inhibition. Blood. 2015;125:316–326. doi: 10.1182/blood-2014-03-566018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Ernst T, Chase AJ, Score J, et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet. 2010;42:722–726. doi: 10.1038/ng.621. [DOI] [PubMed] [Google Scholar]
- 116.Nikoloski G, Langemeijer SMC, Kuiper RP, et al. Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet. 2010;42:665–667. doi: 10.1038/ng.620. [DOI] [PubMed] [Google Scholar]
- 117.Score J, Hidalgo-Curtis C, Jones AV, et al. Inactivation of polycomb repressive complex 2 components in myeloproliferative and myelodysplastic/myeloproliferative neoplasms. Blood. 2012;119:1208–1213. doi: 10.1182/blood-2011-07-367243. [DOI] [PubMed] [Google Scholar]
- 118.Puda A, Milosevic JD, Berg T, et al. Frequent deletions of JARID2 in leukemic transformation of chronic myeloid malignancies. Am J Hematol. 2012;87:245–250. doi: 10.1002/ajh.22257. [DOI] [PubMed] [Google Scholar]
- 119.Abdel-Wahab O, Adli M, LaFave LM, et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell. 2012;22:180–193. doi: 10.1016/j.ccr.2012.06.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Zhang J, Ding L, Holmfeldt L, et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature. 2012;481:157–163. doi: 10.1038/nature10725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Ntziachristos P, Tsirigos A, van Vlierberghe P, et al. Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat Med. 2012;18:298–301. doi: 10.1038/nm.2651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Simon C, Chagraoui J, Krost J, et al. A key role for EZH2 and associated genes in mouse and human adult T-cell acute leukemia. Genes Dev. 2012;26:651–656. doi: 10.1101/gad.186411.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Baude A, Lindroth AM, Plass C. PRC2 loss amplifies Ras signaling in cancer. Nat Genet. 2014;46:1154–1155. doi: 10.1038/ng.3124. [DOI] [PubMed] [Google Scholar]
- 124.Lee W, Teckie S, Wiesner T, et al. PRC2 is recurrently inactivated through EED or SUZ12 loss in malignant peripheral nerve sheath tumors. Nat Genet. 2014;46:1227–1232. doi: 10.1038/ng.3095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Zhang M, Wang Y, Jones S, et al. Somatic mutations of SUZ12 in malignant peripheral nerve sheath tumors. Nat Genet. 2014;46:1170–1172. doi: 10.1038/ng.3116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.De Raedt T, Beert E, Pasmant E, et al. PRC2 loss amplifies Ras-driven transcription and confers sensitivity to BRD4-based therapies. Nature. 2014;514:247–251. doi: 10.1038/nature13561. [DOI] [PubMed] [Google Scholar]
- 127.Woods BA, Levine RL. The role of mutations in epigenetic regulators in myeloid malignancies. Immunol Rev. 2015;263:22–35. doi: 10.1111/imr.12246. [DOI] [PubMed] [Google Scholar]
- 128.Muto T, Santida G, Oshima M, et al. Concurrent loss of Ezh2 and Tet2 cooperates in the pathogenesis of myelodysplastic disorders. J Exp Med. 2013;210:2627–2639. doi: 10.1084/jem.20131144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Bardot ES, Valdes VJ, Zhang J, et al. Polycomb subunits Ezh1 and Ezh2 regulate the Merkel cell differentiation program in skin stem cells. EMBO J. 2013;32:1990–2000. doi: 10.1038/emboj.2013.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Pal B, Boouras T, Shi W, et al. Global changes in the mammary epigenome are induced by hormonal cues and coordinated by Ezh2. Cell Rep. 2013;3:411–426. doi: 10.1016/j.celrep.2012.12.020. [DOI] [PubMed] [Google Scholar]
- 131.Knutson SK, Warholic NM, Johnston LD, et al. Synergistic anti-tumor activity of EZH2 inhibitors and glucocorticoid receptor agonists in models of germinal center non-Hodgkin lymphomas. PLoS One. 2014;9:e111840. doi: 10.1371/journal.pone.0111840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Weisberg E, Manley PW, Coean-Jacob SW, Hochhaus A, Griffin JD. Second generation inhibitors of BCR-ABL for the treatment of imatinib-resistant chronic myeloid leukaemia. Nat Rev Cancer. 2007;7:345–356. doi: 10.1038/nrc2126. [DOI] [PubMed] [Google Scholar]
- 133.Lito P, Rosen N, Solit DB. Tumor adaptation and resistance to RAF inhibitors. Nat Med. 2013;19:1401–1409. doi: 10.1038/nm.3392. [DOI] [PubMed] [Google Scholar]
- 134.Gibaja V, Shen F, Haran J, et al. Development of secondary mutations in wild-type and mutant EZH2 alleles cooperates to confer resistance to EZH2 inhibitors. Oncogene. 2015 doi: 10.1038/onc.2015.114. Article in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Lee ST, Li Z, Wu Z, et al. Context-specific regulation of NF-kappaB target gene expression by EZH2 in breast cancers. Mol Cell. 2011;43:798–810. doi: 10.1016/j.molcel.2011.08.011. [DOI] [PubMed] [Google Scholar]
- 136.Jung HY, Jun S, Lee M, et al. PAF and EZH2 induce Wnt/beta-catenin signaling hyperactivation. Mol Cell. 2013;52:193–205. doi: 10.1016/j.molcel.2013.08.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Xu K, Wu ZJ, Groner AC, et al. EZH2 oncogenic activity in castration-resistant prostate cancer cells is polycomb-independent. Science. 2012;338:1465–1469. doi: 10.1126/science.1227604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.He A, Shen X, Ma Q, et al. PRC2 directly methylates GATA4 and represses its transcriptional activity. Genes Dev. 2012;26:37–42. doi: 10.1101/gad.173930.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Xu J, Shao Z, Li D, et al. Developmental control of polycomb subunit composition by GATA factors mediates a switch to non-canonical functions. Mol Cell. 2015;57:304–316. doi: 10.1016/j.molcel.2014.12.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Konski A, Weiss C, Rosier R, et al. The use of postoperative irradiation for the prevention of heterotopic bone after total hip replacement with biologic fixation (porous coated) prosthesis: An animal model. Int J Radiat Oncol Biol Phys. 1990;18:861–865. doi: 10.1016/0360-3016(90)90408-c. [DOI] [PubMed] [Google Scholar]
- 141.Pasqualucci L, Dominguez-Sola D, Chiarenza A, et al. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature. 2011;471:189–195. doi: 10.1038/nature09730. [DOI] [PMC free article] [PubMed] [Google Scholar]