Abstract
Background
Carbapenemase-producing Klebsiella pneumoniae (CPKP) strains have emerged as a major problem for healthcare systems. The aim of this study was to determine the circulating clones and analyze the clinical and molecular characteristics of CPKP in our hospital.
Methods
A total of 74 carbapenemase producers collected from our hospital from 2012 to 2014 were analyzed for the prevalence of extended-spectrum β-lactamase (ESBLs), plasmid-mediated quinolone resistance genes (PMQRs), exogenously acquired 16S rRNA methyltransferase (16S-RMTase), and plasmid-mediated AmpC enzyme (pAmpCs) by PCR and DNA sequencing. The sequence types (STs) of the carbapenemase producers were analyzed by multi-locus sequence typing (MLST). And Pulsed-field gel electrophoresis (PFGE) was performed to investigate the genetic relationship of KPC-2 producing strains. Clinical data were retrieved from the medical records.
Results
KPC-2 (n = 72) was the predominant enzyme followed by NDM-1 (n = 2); The genes blaCTX-M, blaSHV, blaTEM-1, blaDHA-1, rmtB, armA, oqxA, oqxB, and qnrB were present in 29 (39.2 %), 27 (36.5 %), 46 (62.2 %), 2 (2.7 %), 25 (33.8 %), 1 (1.4 %), 60 (81.1 %) and 56 (75.7 %), 6 (8.1 %) isolates, respectively. MLST analysis revealed 10 different STs. The most dominant ST was ST11 (78.4 %, 58/74), followed by ST15 (8.1 %, 6/74). PFGE patterns of the KPC-2 producing K. pneumoniae isolates exhibited clonal dissemination of ST11 and ST15 clones as well as a genetic diversity of the remaining strains.
Conclusion
The intra- and inter-hospital cross-transmission of KPC-2-producing K. pneumoniae ST11 co-carrying oqxAB and rmtB in our hospital strongly suggested that rapid identification of colonized or infected patients and screening of carriers is quite necessary to prevent a scenario of endemicity.
Keywords: KPC-2, ST11, OqxAB, MLST, Klebsiella pneumoniae
Background
Klebsiella pneumoniae are part of the human gut microbiota and frequently cause infections from mild urinary and gastrointestinal infections to severe bacteremia and pneumonia with a high rate of mortality and morbidity [1]. An increasing emergence of carbapenem resistance among K. pneumoniae isolates worldwide has limited the therapeutic options for treatment of these infections [2].
Carbapenem resistance in K. pneumoniae is frequently caused by the predominant production of KPC carbapenemase worldwide [3], the production of OXA-48-like carbapenemase is currently spreading most rapidly in many European countries with IMP and VIM being occasionally detected [4]. And transmission of blaNDM are potentially global health issues [5]. In addition, Carriage of blaESBL and/or blaAmpC in combination with active efflux or porin loss has also been shown to be responsible for carbapenem-resistance in Enterobacteriaceae [6].
Carbapenem resistant K. pneumoniae (CRKP) is reported to be caused either by epidemic clones or by the horizontal dissemination of mobile elements such as plasmids and insertion sequences. In Europe and American countries, KPC-producing ST258 K. pneumoniae can undeniably be regarded as one of the most successful multidrug-resistant nosocomial pathogens [7]. And the international high-risk clone of K. pneumoniae ST11 is frequently reported as a successful pathogen at infections in Asian and European countries because of the association of important co-resistance and virulence factors [8]. And the quick expansion and evolution of this multi-resistant and virulent clone may lead to an evolving crisis of global dimensions [9].
In recent years, CRKP has been emerging with an increasing frequency worldwide [2]. All these isolates were resistant to currently available antimicrobial agents except tigecycline. Extended-spectrum β-lactamase (ESBLs), exogenously acquired 16S rRNA methyltransferase (16S-RMTase) genes, and plasmid-mediated AmpC enzyme (pAmpCs) has been frequently detected among the clinical multidrug-resistant Enterobacteriaceae [10]. In addition, the prevalence of PMQR genes has been reported to be associated with the increasing quinolone resistance development in Enterobacteriaceae which involved an increase in the diversity of the PMQR genes [11]. Nevertheless, data on the distribution of these resistant determinants among carbapenemase-producing K. pneumoniae (CPKP) isolates collected during a long time period is still limited, and the dissemination of the internationally predominant CPKP isolate is still unclear.
The aim of the present study was to determine the circulating clones of CPKP and our study focused on a hospital-wide screening of ESBLs, 16S-RMTase, PMQRs and pAmpCs in CPKP strains collected during a 3-year-old period.
Methods
Bacterial isolates
Nanjing Drum Tower hospital is comprehensive tertiary hospital which manages 32 clinical departments, maintains 1460 patients, and admits more than 42,000 inpatients. A total of 108 non-replicate and consecutive K. pneumoniae isolates including 107 CRKP and 1 strains intermediate to carbapenem were collected from our hospital from January, 2012 to December, 2014. The Vitek 2 system (bioMérieux. Firenze, Italy) was used for isolate identification and antimicrobial drug susceptibility testing. Clinical specimens consisted of sputum (n = 44), blood (n = 18), urine (n = 12), secretion (n = 11), ascites (n = 9), bile (n = 6) catheter tips (n = 6), and others (n = 2). Among them, carbapenemase producers were further analyzed in this study. Clinical data such as the clinical features, laboratory results, and treatment were retrieved from the medical records department. The study protocol was approved by the Ethics Committee of Nanjing Drum Tower Hospital and written informed consent was obtained from all patients included in the study.
Carba NP confirmatory test
For carbapenemase detection, Carba NP Confirmatory test was performed on 108 K. pneumoniae isolates according to the guidelines of Nordmann et al. [12] and CLSI 2015 [13]. Briefly, 2 microcentrifuge tubes for each patient isolate were labelled as “a” and “b” with 100 μL of bacterial protein extraction reagent (Tris–HCl 20 mmol/L lysis buffer) being added into each tube. A 1-μL loopful of tested bacteria from an overnight blood agar plate was emulsified in both tubes and vortexed for 5 s. Carba NP Solution A (2 mL 0.5 % phenol red solution and 180 μL 10 mM zinc sulfate solution were added into 16.6 mL clinical laboratory reagent water, pH 7.8 ± 0.1) was added into tube “a” and Carba NP Solution B (solution A + 6 mg/mL imipenem) into tube “b”, respectively. After the tubes were well vortexed, An incubation at 35 ± 2 °C for up to 2 h was needed. Test results were interpreted as follows: carbapenemase producers were reported when tube “b” became into light-orange, dark yellow, or yellow before 2 h, and tube “a” kept red or red–orange.
ESBLs confirmatory testing
ESBL production was detected by using the 2015 CLSI recommendations for phenotypic confirmatory test [13]. In brief, disks containing 30 μg of cefotaxime and ceftazidime, either alone or coupled with 10 μg of clavulanate (Oxoid Ltd, Cambridge, UK), were placed at distances of 20 mm (center to center). When the inhibition zone differed by ≥5 mm, between at least 1 of the combination disks and its corresponding single antibiotic disk, the strain was identified as an ESBL producer. E. coli ATCC 25922 and K. pneumoniae ATCC 700603 were used as quality control strains.
Detection of antimicrobial resistance determinants
DNA templates were prepared by the boiling method. All the 108 strains were screened for blaOXA-48 and blaNDM genes by PCR as previously described [14]. The 74 carbapenemase-producers identified by Carba NP Confirmatory test were further analyzed for carbapenemase encoding genes (blaKPC, blaIMI, blaSME, blaGES, blaVIM, blaIMP, blaNDM, blaGIM) by PCR-based techniques with subsequent sequencing according to the protocols as previously described [15], PMQRs (qnrA, qnrB, qnrC, qnrD, qnrS, aac (6′)-Ib-cr, and qepA) were detected by simplex PCR, followed by sequencing [16]. Among the 74 carbapenemase-producing strains, the presence of blaESBLs (blaCTX, blaTEM, blaSHV) were checked for 42 strains positive for the ESBLs confirmatory testing according to the previous protocol [17], The 50 strains resistant to amikacin were further analyzed for the prevalence of 16S-RMTases (armA, npmA, rmtA, rmtB, rmtC, rmtD, and rmtE) [18] by multiplex PCR followed by single PCR for gene confirmation. Positive products were further purified with a DNA purification kit and then sent to the Majorbio Company (Shanghai, China) for sequencing. Sequences were analyzed by using the Chromas-Pro application and BLAST (http://www.ncbi.nlm.nih.gov/BLAST), and the subtypes of β-lactamase genes were confirmed by referring to the Lahey system (http://www.lahey.org/studies/).
Multi-locus sequence typing
The sequence types of the 74 carbapenemase producers were determined by multi-locus sequence typing (MLST) analysis with 7 housekeeping genes including gapA, infB, mdh, pgi, phoE, rpoB, and tonB being amplified and sequenced according to Diancourt et al. [19]. Alleles and sequence types (STs) were assigned by using the MLST database (http://www.pasteur.fr/mlst/Kpneumoniae.html).
Pulsed-field gel electrophoresis
Clonal relatedness of the 72 producing KPC-2 K. pneumoniae strains was analyzed by pulsed-field electrophoresis (PFGE) as previously described [20]. The restriction endonuclease XbaI (Fermentas, ABI, Germany) was used to digest the prepared genomic DNA and resultant DNA fragments were separated in a PFGE CHEF-DR III system (Bio-Rad Laboratories, Hercules, CA, USA) in 0.5× Tris–borate-EDTA buffer at 120 V for 19 h, with pulse times ranging from 2.2 to 54.2 s. The banding patterns were analyzed by the BioNumerics software (Applied Math, Sint-Maten-Latem, Belgium). Cutoff lines at 80 % were used to analyze genetic relatedness.
Results
Prevalence of carbapenemase and Antimicrobial susceptibility of CPKP isolates
Carba NP Confirmatory test showed that 74 out of 108 carbapenem resistant K. pneumoniae isolates were confirmed to be carbapenemase producers. PCR and molecular typing identified KPC-2 among 72 out of 74 carbapenemase producers and the other 2 isolates was found to produce NDM-1. Other carbapenemase-encoding genes were not detected. All the 74 carbapenemase producers found in our study were resistant to all of the β-lactams tested with 67.6 % (50/74) exhibiting resistance to amikacin and 72.73 % (54/74) exhibiting resistance to quinolones.
Clinical characteristics of CPKP isolates
A total of 15 departments in our hospital have been involved with the clonal dissemination of KPC-2 producing K. pneumoniae isolates in our hospital with the intensive care units (ICU) being the commonest department (n = 21), followed by Emergency (n = 10) and Department of Neurosurgery (n = 9). Departments of Geriatrics and General Surgery, Infectious Diseases, Otolaryngology, Urology, Respiratory, Hematology, Neurology, Cardio-thoracic, and Cardiology have also been included. The isolates were mainly from the respiratory tract (n = 35), blood (n = 12) and urine (n = 8). Overall, the spread of KPC-2 producing K. pneumoniae ST11 clones co-carrying other resistance genes in hospital wards during 2012–2014 (Table 1) showed that compared to the data in 2012, such strains rapidly increased with more hospital wards being involved, especially in ICU. And a rapid decrease of such strains was observed during 2014 resulting from the strengthening implementation of hand hygiene in our hospital at the beginning of 2014 (personal communication). In addition, the Retrieval of NDM-producing K. pneumoniae strains showed that one strain was isolated from the blood of a 61-year-old male diagnosed as electrolyte imbalance after colorectal cancer surgery since he was admitted into ICU, persistent fever and blood culture showed severe septicemia caused by a NDM-producing K. pneumoniae strain, after a combination therapy of imipenem, tigecycline, and cefepime for 20 days, the patient died; the other NDM-producing K. pneumoniae was recovered from the secretion of a 41-year-old male who had undergone a serious burns on his whole body. No fever was observed although NDM-producing K. pneumoniae was isolated from the burn wounds of the patients. After a series of Rinse debridement treatment, the patient got a good wound healing and discharged.
Table 1.
The spread of KPC-2 producing K. pneumonia ST11 clones associated with other resistance genes in hospital wards during 2012–2014
| Isolation year | KPK-ST11 | Antimicrobial Resistance determinants | Ward | ||||
|---|---|---|---|---|---|---|---|
| ESBLs | PMQRs | 16S-RMTase | Carbapenemase | AmpC | |||
| 2012 | 10 | 8 | 10 | 7 | 10 | 0 | ER(3), GS(2), NL(1), GA(1), NS(1), RP(1), VS(1) |
| 2013 | 36 | 33 | 32 | 13 | 36 | 1 | ICU(11), NS(6), ER(6), GS(3) GA(2), IDS(1), OG(1), UL(1), RP(1) |
| 2014 | 11 | 8 | 9 | 1 | 11 | 1 | ICU(4), CS(3), AD(1), GA(1), NS(1) |
KPK-ST11: KPC-2 producing K. pneumonia ST11
ER emergency room, ICU Intensive Care Unit, CS cardiothoracic surgery, NL neurology, AD andrology, UL urology, VS vascular surgery, NS neurosurgery, GS general surgery, GA geriatrics, RP respiratory, IDS infectious diseases, OG Otolaryngology
Prevalence of antimicrobial resistant determinants
ESBLs confirmatory testing showed that 42 strains were positive for ESBLs among the 74 carbapenemase-producing strains. PCR and DNA sequencing identified CTX-M, SHV, and TEM-1 type β-lactamases in 29 (39.2 %), 27 (36.5 %), 46 (62.2 %) isolates, respectively. CTX-M variants included 24 CTX-M-14, 3 CTX-M-15, 2 CTX-M-38; one SHV-1 and multiple SHV variants including SHV-11 (n = 2), SHV-12 (n = 13), SHV-28 (n = 2), SHV-31 (n = 6), SHV-160 (n = 2), and SHV-36a (n = 1) were observed. In addition, 2 DHA-1 were identified. 16S-RMTase including RmtB (n = 25) and Arm (n = 1) were identified among 26 out of the 50 carbapenemase-producers displaying resistance to amikacin. Fifty out of 74 carbapenemase producers were found to carry oqxAB. And the prevalence of both oqxA and oqxB detected in K. pneumoniae was high: 83.3 and 77.8 %, respectively. Of them, 48 oqxAB, 58 oqxA and 56 oqxB were detected among 69 carbapenemase producers highly resistant to fluoroquinolones. Whereas, 2 oqxAB, 2 oqxA and 2 oqxB were found among the 3 carbapenemase producers susceptible to fluoroquinolones.
In addition, one novel mutants named oqxA11 and 3 novel oqxB mutants named oqxB13, oqxB27, and oqxB28 were identified. And 3 kinds of qnrB variants including qnrB6 (n = 1), qnrB10 (n = 3) and qnrB66 (n = 2) were identified. Other PMQRs were not detected.
Of particular concern, qnrB6, oqxA, blaTEM-1, and blaCTX-M-15 were identified in one NDM-producing isolate, albeit only oqxB, NDM-1 was found in the other one.
MLST
MLST analysis of 74 carbapenemase producers revealed 10 different STs (Fig. 1). The most dominant ST was ST11 (78.4 %, 58/74), followed by ST15 (8.1 %, 6/74). The remaining isolates were typed as ST23, ST218, ST362, ST412, ST420, ST606, ST690, and ST709. MLST analysis of 2 NDM-1 producing K. pneumoniae isolates identified as ST826 and ST307, respectively.
Fig. 1.
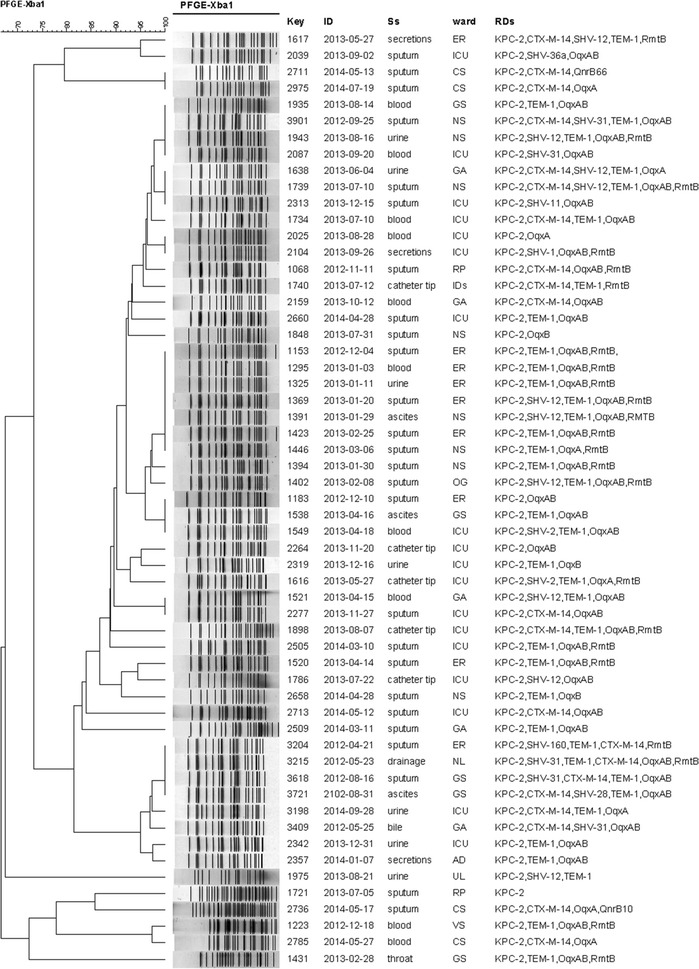

Dendrogram based on PFGE profiles of 57 KPC-2 producing K. pneumoniae ST11 isolates. The dendrogram was produced by the UPGMA algorithm based on the Dice similarity coefficient included five PFGE groups as defined based on 80 % similarity of PFGE profiles. ID isolation date, Ss samples, RDs resistant determinants, ER emergency room, ICU Intensive Care Unit, CS cardiothoracic surgery, NL neurology, AD andrology, UL urology, VS vascular surgery, NS neurosurgery, GS general surgery, GA geriatrics, RP respiratory, IDS infectious diseases, OG otolaryngology
Clonal relatedness
According to the PFGE patterns of the isolates (Fig. 1), the 57 KPC-2 producing K. pneumoniae ST11 isolates exhibited 7 PFGE profiles. One major clonal groups composed of 47 closely related strains was observed among all the ST11 clones according to the 80 % similarity level. The PFGE data of the other 13 KPC-2 producing K. pneumoniae displayed a genetic diversity (Fig. 2). Among them, 2 PFGE profiles composed of 5 ST15 clones were observed. In addition, 2 strains including 1 ST11 clone and 1 ST15 one were non-typable by PFGE.
Fig. 2.
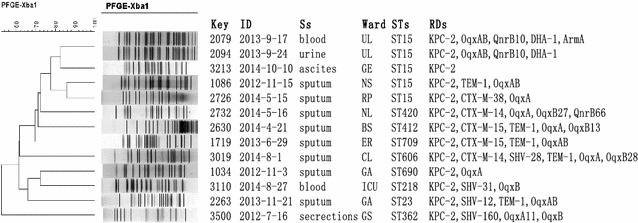
Dendrogram based on PFGE profiles of 13 KPC-2 producing K. pneumoniae isolates. The dendrogram was produced by the UPGMA algorithm based on the Dice similarity coefficient included five PFGE groups as defined based on 80 % similarity of PFGE profiles. ID isolation date, Ss samples, RDs resistant determinants, ER emergency room, UL urology, GE gastroenterology, NS neurosurgery, RP respiratory, NL neurology, BS burn surgery, ER emergency room, CL cardiology, GA geriatrics, ICU Intensive Care Unit, GS general surgery
Discussion
Clonal dissemination of CRKP in the health care settings frequently cause a potential threat to the public health with the increasing use of carbapenems in the clinical treatments worldwide. And we provided the first report on the clonal dissemination of K. pneumoniae ST11 with a high prevalence of OqxAB in a tertiary hospital in China.
In this study, 72 out of 74 carbapenemase producers carried KPC-2, indicating that KPC is the major CHBLs accounting for the carbapenem resistance and KPC-2 is the most prevalent enzyme which was in accordance with the previous studies both at home and abroad [3]. OqxAB, encoded by both chromosome and plasmids as a quinolone and olaquindox efflux pump, has become increasingly prevalent among members of Enterobacteriaceae over the past decade [21, 22]. It has been demonstrated that acquisition of the oqxAB encoding plasmids may facilitate the selection of CIP resistant Salmonella typhimuriu [23], and high expression of this pump contributes to reduced susceptibility to quinolones in clinical isolates of ESBL-producing K. pneumoniae [24]. A more recently published paper also described that overexpression of the oqxAB efflux pump in K. pneumoniae has been linked to an antibiotic cross-resistance phenotype [25] and OqxAB may contribute to tigecycline resistance in K. pneumoniae isolates [26]. Furthermore, a previous analysis of the genotypic and phenotypic characteristics of K. pneumoniae harboring oqxAB-like elements found that chromosomal oqxAB elements in K. pneumoniae strains exhibited cross resistance to olaquindox, chloramphenicol and the quinolones, whereas, transposition of the oqxAB operon from bacterial chromosome to plasmids results in more than 80-fold increase in the level of expression of the oqxAB pump, confirming that oqxAB efflux system is constitutively expressed when located in bacterial mobile elements [27]. The resistance mediated by the mutations in the quinolone-resistance determining regions in DNA gyrase and DNA topoisomerase IV of K. pneumoniae makes it difficult for us to speculate whether oqxAB in our study locates on plasmids or on the chromosome, and further study is needed to analyze the locations of these oqxAB operons and the role that oqxAB, oqxA and oqxB played in the fluoroquinolone development.
All together, the observed high prevalence of pump gene oqxA and oqxB in combination or alone in the KPC-2 producing K. pneumoniae ST11 represents a potential reservoir for the spread of these genes.
Of note, it’s the first time that we identified a new variant qnrB66 as well as the novel mutant named oqxA11 and 3 novel oqxB mutants named oqxB13, oqxB27, and oqxB28 in K. pneumoniae which indicate that these PMQRs have been rapidly evolving under the selection pressure of antimicrobial agents in the healthcare settings.
Furthermore, the high prevalence of rmtB among KPC-2 producing K. pneumoniae isolates has been reported in China [28], it has also been characterized that rmtB and blaKPC-2 were located in the same plasmid and frequently disseminated among K. pneumoniae isolates under the selection pressure of antimicrobial agents [29].
Up to date, ST11 strains has been frequently reported to be the carriage of multiple resistant determinants such as CTX-M, KPC-2, as well as RmtB [8]. And the co-carriage of different ESCs (extended-spectrum cephalosporinaes) and 16S-RMTases, alone or in combination, were also noted among the KPC-2 isolates in our study, indicating that the clone dissemination of ST11 strain may carry different types of plasmids or more than one plasmid. Additionally, recent reports showed that ST11 is the dominant strain associated with carbapenem resistance in K. pneumoniae [30] and even also the predominant clone of tigecycline-resistant K. pneumoniae strains [26], indicating that K. pneumoniae ST11 has been developing as major high-risk clone. Taken together, these data indicated rapid evolution and expansion of the KPC-2 producing ST11 isolates with acquisition and spread of transferable resistance determinants in our hospital [31].
Noticeably, there is a potential dissemination of ST15 clone secondary to the ST11 clones in our study, indicating that ST15 is another major high-risk clones. This is consistent with a recent study [32]. ST23 has been frequently reported to be the carbapenem resistant strains [33], whereas, the other clones including ST218, ST362, ST412, ST420, ST606, ST690, and ST709 has not been reported to be involved with carbapenem resistance.
Of particular concern, NDM has been frequently found in K. pneumoniae isolates and involved with the healthcare associated outbreaks in China [34, 35] and the emergence of NDM-producing K. pneumoniae isolates in our hospital showed a spread potential of these strains and infection control measures should be emergently implemented to prevent the healthcare associated outbreaks since the choice of antimicrobial agents is quite limited once the patients were infected with NDM-producing K. pneumoniae isolates. Additionally, to the best of our knowledge, it’s the first time that we provide data on the K. pneumoniae ST826 and ST307 clones carrying NDM.
In recent years, the clonal dissemination of CRKP ST11 among Chinese hospitals has been frequently reported [32] and the wide distribution of the KPC-2-producing K. pneumoniae ST11 isolates co-carrying oqxAB and rmtB in our study indicate an intra- and inter- hospital cross-transmission of such strains in our hospital, which further strongly suggested that these isolates have a high potential to cause healthcare-associated outbreaks. Careful monitoring of carbapenem susceptibilities and rapid identification of colonized or infected patients as well as screening of carriers is quite necessary for implementation of infection control measures to prevent a scenario of endemicity.
Authors’ contributions
LC and XC performed experimental work and drafted manuscript; ZZ and MN analyzed the study data; WZ, JC and JZ provided interpretation of data; KZ, HS conceived the study and provided data interpretation. All authors read and approved the final manuscript.
Acknowledgements
This study was supported by the Youth Fund of Jiangsu Province (Grant no. BK2014099) and Nanjing Medical Technology Development Fund (ZKX13028).
Competing interests
The authors declare that they have no competing interests.
Abbreviations
- CPKP
carbapenemase-producing Klebsiella pneumoniae
- ESBLs
extended-spectrum β-lactamase
- PMQRs
plasmid-mediated quinolone resistance genes
- 16S-RMTase
16S rRNA methyltransferase
- pAmpCs
plasmid-mediated AmpC enzyme
- STs
sequence types
- MLST
multi-locus sequence typing
- PFGE
pulsed-field gel electrophoresis
- ID
isolation date
- Ss
samples
- RDs
resistant determinants
- ER
emergency room
- ICU
Intensive Care Unit
- CS
general surgery
- NL
neurology
- AD
andrology
- UL
urology
- VS
vascular surgery
- NS
neurosurgery
- GS
general surgery
- GA
geriatrics
- RP
respiratory
- IDs
infectious diseases
- OG
otolaryngology
- GE
gastroenterology
- BS
burn surgery
- CL
cardiology
Footnotes
Li Cheng and Xiao-Li Cao contributed equally to this work
Contributor Information
Li Cheng, Email: chenglimed@163.com.
Xiao-Li Cao, Email: cao-xiao-li@163.com.
Zhi-Feng Zhang, Email: njzhangzf@hotmail.com.
Ming-zhe Ning, Email: nmz780714@sina.com.
Xue-Jing Xu, Email: xxuejing@hotmail.com.
Wanqing Zhou, Email: zwq_096@163.com.
Jun-Hao Chen, Email: nj_cjh@aliyun.com.
Jin-hua Zhang, Email: zhangjinhua0908@163.com.
Han Shen, Phone: + 86 25 83 10 53 60, Email: shenhan10366@sina.com.
Kui Zhang, Phone: + 86 25 83 10 53 60, Email: 13505151066@163.com.
References
- 1.Hussein K, Raz-Pasteur A, Finkelstein R, Neuberger A, Shachor-Meyouhas Y, Oren I, Kassis I. Impact of carbapenem resistance on the outcome of patients’ hospital-acquired bacteraemia caused by Klebsiella pneumoniae. J Hosp Infect. 2013;83(4):307–313. doi: 10.1016/j.jhin.2012.10.012. [DOI] [PubMed] [Google Scholar]
- 2.Hussein K, Sprecher H, Mashiach T, Oren I, Kassis I, Finkelstein R. Carbapenem resistance among Klebsiella pneumoniae isolates: risk factors, molecular characteristics, and susceptibility patterns. Infect Control Hosp Epidemiol. 2009;30(7):666–671. doi: 10.1086/598244. [DOI] [PubMed] [Google Scholar]
- 3.Munoz-Price LS, Poirel L, Bonomo RA, Schwaber MJ, Daikos GL, Cormican M, Cornaglia G, Garau J, Gniadkowski M, Hayden MK, Kumarasamy K, Livermore DM, Maya JJ, Nordmann P, Patel JB, Paterson DL, Pitout J, Villegas MV, Wang H, Woodford N, Quinn JP. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect Dis. 2013;13(9):785–796. doi: 10.1016/S1473-3099(13)70190-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Canton R, Akova M, Carmeli Y, Giske CG, Glupczynski Y, Gniadkowski M, Livermore DM, Miriagou V, Naas T, Rossolini GM, Samuelsen Ø, Seifert H, Woodford N, Nordmann P. Rapid evolution and spread of carbapenemases among Enterobacteriaceae in Europe. Clin Microbiol Infect. 2012;18(5):413–431. doi: 10.1111/j.1469-0691.2012.03821.x. [DOI] [PubMed] [Google Scholar]
- 5.Wei WJ, Yang HF, Ye Y, Li JB. New Delhi metallo-beta-lactamase-mediated carbapenem resistance: origin, diagnosis, treatment and public health concern. Chin Med J (Engl) 2015;128(14):1969–1976. doi: 10.4103/0366-6999.160566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shin SY, Bae IK, Kim J, Jeong SH, Yong D, Kim JM, Lee K. Resistance to carbapenems in sequence type 11 Klebsiella pneumoniae is related to DHA-1 and loss of OmpK35 and/or OmpK36. J Med Microbiol. 2012;61(Pt 2):239–245. doi: 10.1099/jmm.0.037036-0. [DOI] [PubMed] [Google Scholar]
- 7.Tzouvelekis LS, Markogiannakis A, Psichogiou M, Tassios PT, Daikos GL. Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: an evolving crisis of global dimensions. Clin Microbiol Rev. 2012;25(4):682–707. doi: 10.1128/CMR.05035-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Andrade LN, Vitali L, Gaspar GG, Bellissimo-Rodrigues F, Martinez R, Darini AL. Expansion and evolution of a virulent, extensively drug-resistant (polymyxin B-resistant), QnrS1-, CTX-M-2-, and KPC-2-producing Klebsiella pneumoniae ST11 international high-risk clone. J Clin Microbiol. 2014;52(7):2530–2535. doi: 10.1128/JCM.00088-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li H, Zhang J, Liu Y, Zheng R, Chen H, Wang X, Wang Z, Cao B, Wang H. Molecular characteristics of carbapenemase-producing Enterobacteriaceae in China from 2008 to 2011: predominance of KPC-2 enzyme. Diagn Microbiol Infect Dis. 2014;78(1):63–65. doi: 10.1016/j.diagmicrobio.2013.10.002. [DOI] [PubMed] [Google Scholar]
- 10.Cao X, Xu X, Zhang Z, Shen H, Chen J, Zhang K. Molecular characterization of clinical multidrug-resistant Klebsiella pneumoniae isolates. Ann Clin Microbiol Antimicrob. 2014;13:16. doi: 10.1186/1476-0711-13-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Carattoli A. Plasmids and the spread of resistance. Int J Med Microbiol. 2013;303(6–7):298–304. doi: 10.1016/j.ijmm.2013.02.001. [DOI] [PubMed] [Google Scholar]
- 12.Nordmann P, Poirel L, Dortet L. Rapid detection of carbapenemase-producing Enterobacteriaceae. Emerg Infect Dis. 2012;18(9):1503–1507. doi: 10.3201/eid1809.120355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Clinical Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; 25th informational supplement. M100-S25. CLSI, Wayne, PA. 2015.
- 14.Zowawi HM, Sartor AL, Balkhy HH, Walsh TR, Al Johani SM, AlJindan RY, Alfaresi M, Ibrahim E, Al-Jardani A, Al-Abri S, Al Salman J, Dashti AA, Kutbi AH, Schlebusch S, Sidjabat HE, Paterson DL. Molecular characterization of carbapenemase-producing Escherichia coli and Klebsiella pneumoniae in the countries of the Gulf cooperation council. Dominance of OXA-48 and NDM producers. Antimicrob Agents Chemother. 2014;58(6):3085–3090. doi: 10.1128/AAC.02050-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Poirel L, Dortet L, Bernabeu S, Nordmann P. Genetic features of blaNDM-1-positive Enterobacteriaceae. Antimicrob Agents Chemother. 2011;55(11):5403–5407. doi: 10.1128/AAC.00585-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Elgorriaga-Islas E, Guggiana-Nilo P, Dominguez-Yevenes M, González-Rocha G, Mella-Montecinos S, Labarca-Labarca J, García-Cañete P, Bello-Toledo H. Prevalence of plasmid-mediated quinolone resistance determinant aac(6’)-Ib-cr among ESBL producing enterobacteria isolates from Chilean hospitals. Enferm Infecc Microbiol Clin. 2012;30(8):466–468. doi: 10.1016/j.eimc.2012.01.024. [DOI] [PubMed] [Google Scholar]
- 17.Dallenne C, Da Costa A, Decre D, Favier C, Arlet G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J Antimicrob Chemother. 2010;65(3):490–495. doi: 10.1093/jac/dkp498. [DOI] [PubMed] [Google Scholar]
- 18.Doi Y, Arakawa Y. 16S ribosomal RNA methylation: emerging resistance mechanism against aminoglycosides. Clin Infect Dis. 2007;45(1):88–94. doi: 10.1086/518605. [DOI] [PubMed] [Google Scholar]
- 19.Diancourt L, Passet V, Verhoef J, Grimont PA, Brisse S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J Clin Microbiol. 2005;43(8):4178–4182. doi: 10.1128/JCM.43.8.4178-4182.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Carrer A, Lassel L, Fortineau N, Mansouri M, Anguel N, Richard C, Nordmann P. Outbreak of CTX-M-15-producing Klebsiella pneumoniae in the intensive care unit of a French hospital. Microb Drug Resist. 2009;15(1):47–54. doi: 10.1089/mdr.2009.0868. [DOI] [PubMed] [Google Scholar]
- 21.Yuan J, Xu X, Guo Q, Zhao X, Ye X, Guo Y, Wang M. Prevalence of the oqxAB gene complex in Klebsiella pneumoniae and Escherichia coli clinical isolates. J Antimicrob Chemother. 2012;67(7):1655–1659. doi: 10.1093/jac/dks086. [DOI] [PubMed] [Google Scholar]
- 22.Perez F, Rudin SD, Marshall SH, Coakley P, Chen L, Kreiswirth BN, Rather PN, Hujer AM, Toltzis P, van Duin D, Paterson DL, Bonomo RA. OqxAB, a quinolone and olaquindox efflux pump, is widely distributed among multidrug-resistant Klebsiella pneumoniae isolates of human origin. Antimicrob Agents Chemother. 2013;57(9):4602–4603. doi: 10.1128/AAC.00725-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wong MH, Chan EW, Liu LZ, Chen S. PMQR genes oqxAB and aac(6′)Ib-cr accelerate the development of fluoroquinolone resistance in Salmonella typhimurium. Front Microbiol. 2014;5:521. doi: 10.3389/fmicb.2014.00521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rodriguez-Martinez JM, Diaz de Alba P, Briales A, Machuca J, Lossa M, Fernández-Cuenca F, Rodríguez Baño J, Martínez-Martínez L, Pascual Á. Contribution of OqxAB efflux pumps to quinolone resistance in extended-spectrum-beta-lactamase-producing Klebsiella pneumoniae. J Antimicrob Chemother. 2013;68(1):68–73. doi: 10.1093/jac/dks377. [DOI] [PubMed] [Google Scholar]
- 25.Bialek-Davenet S, Lavigne JP, Guyot K, Mayer N, Tournebize R, Brisse S, Leflon-Guibout V, Nicolas-Chanoine MH. Differential contribution of AcrAB and OqxAB efflux pumps to multidrug resistance and virulence in Klebsiella pneumoniae. J Antimicrob Chemother. 2015;70(1):81–88. doi: 10.1093/jac/dku340. [DOI] [PubMed] [Google Scholar]
- 26.Zhong X, Xu H, Chen D, Zhou H, Hu X, Cheng G. First emergence of acrAB and oqxAB mediated tigecycline resistance in clinical isolates of Klebsiella pneumoniae pre-dating the use of tigecycline in a Chinese hospital. PLoS One. 2014;9(12):e115185. doi: 10.1371/journal.pone.0115185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wong MH, Chan EW, Chen S. Evolution and dissemination of OqxAB-like efflux pumps, an emerging quinolone resistance determinant among members of Enterobacteriaceae. Antimicrob Agents Chemother. 2015;59(6):3290–3297. doi: 10.1128/AAC.00310-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li JJ, Sheng ZK, Deng M, Bi S, Hu FS, Miao HF, Ji ZK, Sheng JF, Li LJ. Epidemic of Klebsiella pneumoniae ST11 clone coproducing KPC-2 and 16S rRNA methylase RmtB in a Chinese University Hospital. BMC Infect Dis. 2012;12:373. doi: 10.1186/1471-2334-12-373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sheng JF, Li JJ, Tu S, Sheng ZK, Bi S, Zhu MH, Shen XM, Li LJ. blaKPC and rmtB on a single plasmid in Enterobacter amnigenus and Klebsiella pneumoniae isolates from the same patient. Eur J Clin Microbiol Infect Dis. 2012;31(7):1585–1591. doi: 10.1007/s10096-011-1481-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sun K, Chen X, Li C, Yu Z, Zhou Q, Yan Y. Clonal dissemination of multilocus sequence type 11 Klebsiella pneumoniae carbapenemase—producing K. pneumoniae in a Chinese teaching hospital. APMIS. 2015;123(2):123–127. doi: 10.1111/apm.12313. [DOI] [PubMed] [Google Scholar]
- 31.Wang JT, Wu UI, Lauderdale TL, Chen MC, Li SY, Hsu LY, Chang SC. Carbapenem-nonsusceptible enterobacteriaceae in taiwan. PLoS ONE. 2015;10(3):e0121668. doi: 10.1371/journal.pone.0121668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yan JJ, Wang MC, Zheng PX, Tsai LH, Wu JJ. Associations of the major international high-risk resistant clones and virulent clones with specific ompK36 allele groups in Klebsiella pneumoniae in Taiwan. New Microbes New Infect. 2015;5:1–4. doi: 10.1016/j.nmni.2015.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Qu TT, Zhou JC, Jiang Y, Shi KR, Li B, Shen P, Wei ZQ, Yu YS. Clinical and microbiological characteristics of Klebsiella pneumoniae liver abscess in East China. BMC Infect Dis. 2015;15(1):161. doi: 10.1186/s12879-015-0899-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Jin Y, Shao C, Li J, Fan H, Bai Y, Wang Y. Outbreak of multidrug resistant NDM-1-producing Klebsiella pneumoniae from a Neonatal Unit in Shandong Province, China. PLoS One. 2015;10(3):e0119571. doi: 10.1371/journal.pone.0119571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu Y, Wan LG, Deng Q, Cao XW, Yu Y, Xu QF. First description of NDM-1-, KPC-2-, VIM-2- and IMP-4-producing Klebsiella pneumoniae strains in a single Chinese teaching hospital. Epidemiol Infect. 2015;143(2):376–384. doi: 10.1017/S0950268814000995. [DOI] [PMC free article] [PubMed] [Google Scholar]


