Abstract
The explosive accumulation of protein sequences in the wake of large-scale sequencing projects is in stark contrast to the much slower experimental determination of protein structures. Improved methods of structure prediction from the gene sequence alone are therefore needed. Here, we report a substantial increase in both the accuracy and quality of secondary-structure predictions, using a neural-network algorithm. The main improvements come from the use of multiple sequence alignments (better overall accuracy), from "balanced training" (better prediction of beta-strands), and from "structure context training" (better prediction of helix and strand lengths). This method, cross-validated on seven different test sets purged of sequence similarity to learning sets, achieves a three-state prediction accuracy of 69.7%, significantly better than previous methods. In addition, the predicted structures have a more realistic distribution of helix and strand segments. The predictions may be suitable for use in practice as a first estimate of the structural type of newly sequenced proteins.
Full text
PDF
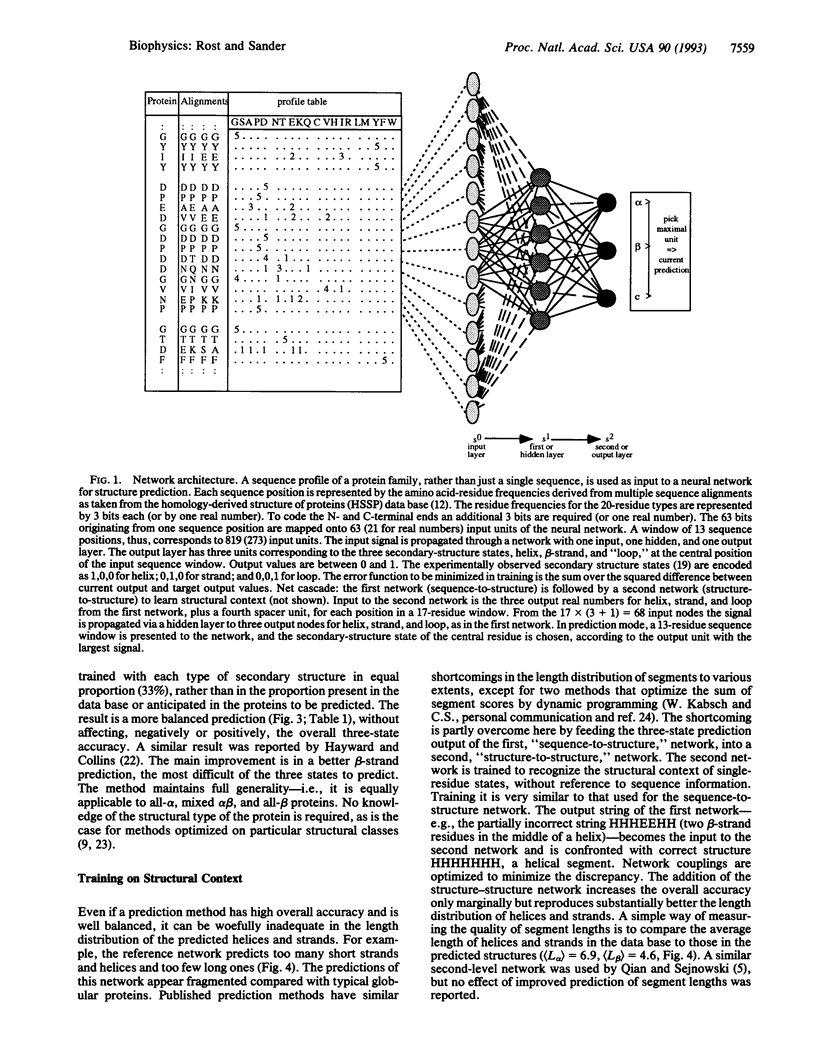


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barton G. J., Newman R. H., Freemont P. S., Crumpton M. J. Amino acid sequence analysis of the annexin super-gene family of proteins. Eur J Biochem. 1991 Jun 15;198(3):749–760. doi: 10.1111/j.1432-1033.1991.tb16076.x. [DOI] [PubMed] [Google Scholar]
- Benner S. A., Gerloff D. Patterns of divergence in homologous proteins as indicators of secondary and tertiary structure: a prediction of the structure of the catalytic domain of protein kinases. Adv Enzyme Regul. 1991;31:121–181. doi: 10.1016/0065-2571(91)90012-b. [DOI] [PubMed] [Google Scholar]
- Bohr H., Bohr J., Brunak S., Cotterill R. M., Lautrup B., Nørskov L., Olsen O. H., Petersen S. B. Protein secondary structure and homology by neural networks. The alpha-helices in rhodopsin. FEBS Lett. 1988 Dec 5;241(1-2):223–228. doi: 10.1016/0014-5793(88)81066-4. [DOI] [PubMed] [Google Scholar]
- Chothia C., Lesk A. M. The relation between the divergence of sequence and structure in proteins. EMBO J. 1986 Apr;5(4):823–826. doi: 10.1002/j.1460-2075.1986.tb04288.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garnier J., Levin J. M. The protein structure code: what is its present status? Comput Appl Biosci. 1991 Apr;7(2):133–142. doi: 10.1093/bioinformatics/7.2.133. [DOI] [PubMed] [Google Scholar]
- Hayward S., Collins J. F. Limits on alpha-helix prediction with neural network models. Proteins. 1992 Nov;14(3):372–381. doi: 10.1002/prot.340140306. [DOI] [PubMed] [Google Scholar]
- Hobohm U., Scharf M., Schneider R., Sander C. Selection of representative protein data sets. Protein Sci. 1992 Mar;1(3):409–417. doi: 10.1002/pro.5560010313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holley L. H., Karplus M. Protein secondary structure prediction with a neural network. Proc Natl Acad Sci U S A. 1989 Jan;86(1):152–156. doi: 10.1073/pnas.86.1.152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holm L., Ouzounis C., Sander C., Tuparev G., Vriend G. A database of protein structure families with common folding motifs. Protein Sci. 1992 Dec;1(12):1691–1698. doi: 10.1002/pro.5560011217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kabsch W., Sander C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers. 1983 Dec;22(12):2577–2637. doi: 10.1002/bip.360221211. [DOI] [PubMed] [Google Scholar]
- Kneller D. G., Cohen F. E., Langridge R. Improvements in protein secondary structure prediction by an enhanced neural network. J Mol Biol. 1990 Jul 5;214(1):171–182. doi: 10.1016/0022-2836(90)90154-E. [DOI] [PubMed] [Google Scholar]
- Levin J. M., Garnier J. Improvements in a secondary structure prediction method based on a search for local sequence homologies and its use as a model building tool. Biochim Biophys Acta. 1988 Aug 10;955(3):283–295. doi: 10.1016/0167-4838(88)90206-3. [DOI] [PubMed] [Google Scholar]
- Matthews B. W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim Biophys Acta. 1975 Oct 20;405(2):442–451. doi: 10.1016/0005-2795(75)90109-9. [DOI] [PubMed] [Google Scholar]
- Maxfield F. R., Scheraga H. A. Improvements in the prediction of protein backbone topography by reduction of statistical errors. Biochemistry. 1979 Feb 20;18(4):697–704. doi: 10.1021/bi00571a023. [DOI] [PubMed] [Google Scholar]
- Muggleton S., King R. D., Sternberg M. J. Protein secondary structure prediction using logic-based machine learning. Protein Eng. 1992 Oct;5(7):647–657. doi: 10.1093/protein/5.7.647. [DOI] [PubMed] [Google Scholar]
- Overington J., Johnson M. S., Sali A., Blundell T. L. Tertiary structural constraints on protein evolutionary diversity: templates, key residues and structure prediction. Proc Biol Sci. 1990 Aug 22;241(1301):132–145. doi: 10.1098/rspb.1990.0077. [DOI] [PubMed] [Google Scholar]
- Ptitsyn O. B., Finkelstein A. V. Theory of protein secondary structure and algorithm of its prediction. Biopolymers. 1983 Jan;22(1):15–25. doi: 10.1002/bip.360220105. [DOI] [PubMed] [Google Scholar]
- Qian N., Sejnowski T. J. Predicting the secondary structure of globular proteins using neural network models. J Mol Biol. 1988 Aug 20;202(4):865–884. doi: 10.1016/0022-2836(88)90564-5. [DOI] [PubMed] [Google Scholar]
- Richardson J. S. The anatomy and taxonomy of protein structure. Adv Protein Chem. 1981;34:167–339. doi: 10.1016/s0065-3233(08)60520-3. [DOI] [PubMed] [Google Scholar]
- Russell R. B., Breed J., Barton G. J. Conservation analysis and structure prediction of the SH2 family of phosphotyrosine binding domains. FEBS Lett. 1992 Jun 8;304(1):15–20. doi: 10.1016/0014-5793(92)80579-6. [DOI] [PubMed] [Google Scholar]
- Sacchettini J. C., Gordon J. I., Banaszak L. J. Refined apoprotein structure of rat intestinal fatty acid binding protein produced in Escherichia coli. Proc Natl Acad Sci U S A. 1989 Oct;86(20):7736–7740. doi: 10.1073/pnas.86.20.7736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salzberg S., Cost S. Predicting protein secondary structure with a nearest-neighbor algorithm. J Mol Biol. 1992 Sep 20;227(2):371–374. doi: 10.1016/0022-2836(92)90892-n. [DOI] [PubMed] [Google Scholar]
- Sander C., Schneider R. Database of homology-derived protein structures and the structural meaning of sequence alignment. Proteins. 1991;9(1):56–68. doi: 10.1002/prot.340090107. [DOI] [PubMed] [Google Scholar]
- Stolorz P., Lapedes A., Xia Y. Predicting protein secondary structure using neural net and statistical methods. J Mol Biol. 1992 May 20;225(2):363–377. doi: 10.1016/0022-2836(92)90927-c. [DOI] [PubMed] [Google Scholar]
- Zhang X., Mesirov J. P., Waltz D. L. Hybrid system for protein secondary structure prediction. J Mol Biol. 1992 Jun 20;225(4):1049–1063. doi: 10.1016/0022-2836(92)90104-r. [DOI] [PubMed] [Google Scholar]
- Zvelebil M. J., Barton G. J., Taylor W. R., Sternberg M. J. Prediction of protein secondary structure and active sites using the alignment of homologous sequences. J Mol Biol. 1987 Jun 20;195(4):957–961. doi: 10.1016/0022-2836(87)90501-8. [DOI] [PubMed] [Google Scholar]



