Significance
The recent development of the clustered regularly interspaced short palindromic repeats (CRISPR)-associated (Cas)9 system has greatly simplified the process of genomic editing. However, it has remained difficult to induce mutations in postnatal animals due to delivery challenges of the CRISPR/Cas9 components. Here, we report the generation of a transgenic mouse line that expresses Cas9 exclusively in cardiomyocytes. By using Adeno-Associated Virus 9 to deliver single-guide RNA (sgRNA) this method can rapidly induce genomic insertions and deletions in the heart. As proof of concept, administration of sgRNA against the Myh6 gene induced Myh6 editing, resulting in cardiomyopathy and heart failure in the cardiac-specific Cas9 mouse. This transgenic mouse model offers a valuable tool for cardiovascular research, as a straightforward strategy to edit genes of interest in the heart.
Keywords: gene knockdown, cardiovascular pathology, CRISPR-associated endonuclease, transgenic mouse, cardioediting
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR)-associated (Cas)9 genomic editing has revolutionized the generation of mutant animals by simplifying the creation of null alleles in virtually any organism. However, most current approaches with this method require zygote injection, making it difficult to assess the adult, tissue-specific functions of genes that are widely expressed or which cause embryonic lethality when mutated. Here, we describe the generation of cardiac-specific Cas9 transgenic mice, which express high levels of Cas9 in the heart, but display no overt defects. In proof-of-concept experiments, we used Adeno-Associated Virus 9 (AAV9) to deliver single-guide RNA (sgRNA) that targets the Myh6 locus exclusively in cardiomyocytes. Intraperitoneal injection of postnatal cardiac-Cas9 transgenic mice with AAV9 encoding sgRNA against Myh6 resulted in robust editing of the Myh6 locus. These mice displayed severe cardiomyopathy and loss of cardiac function, with elevation of several markers of heart failure, confirming the effectiveness of this method of adult cardiac gene deletion. Mice with cardiac-specific expression of Cas9 provide a tool that will allow rapid and accurate deletion of genes following a single injection of AAV9-sgRNAs, thereby circumventing embryonic lethality. This method will be useful for disease modeling and provides a means of rapidly editing genes of interest in the heart.
The ability to generate mice with either gain or loss-of-function mutations has allowed the identification of genetic regulators of many aspects of development, physiology, and disease (1). Historically, however, the generation of mutant mice has been time-consuming and labor-intensive. The recent identification of the clustered regularly interspaced short palindromic repeats (CRISPR)-associated (Cas)9 system has revolutionized the field of genetics and has greatly facilitated the generation of genetically modified animals (2).
CRISPRs were first identified as part of the bacterial immune system, playing a role in viral defense (3). The CRISPR-associated endonuclease Cas9 can be targeted to specific locations in the genome via an RNA-guided system involving single-guide (sg) RNAs to induce double-strand breaks in regions of interest (4–7). The double-strand breaks induced by Cas9 cleavage are preferentially repaired by Non-Homologous End Joining (NHEJ), an error-prone form of DNA repair (8, 9). Consequently, short insertions or deletions (indels) are frequently introduced at the site of Cas9 cleavage, leading to frameshift mutations and the induction of a premature stop codon. Subsequently, translation of the protein of interest is terminated, resulting in degradation of the transcript by nonsense-mediated decay and protein loss (10, 11). As a result, CRISPR/Cas9 has been increasingly used to generate loss-of-function mutations in genes of interest in a variety of organisms, including zebrafish (12, 13), mice (14, 15), and nonhuman primates (16).
Despite the ease with which CRISPR/Cas9 can be used to induce genetic mutations, most applications of the technology have relied upon germline genomic editing in zygotes, rather than in postnatal or adult animals. As a result, difficulties remain with using the technology to analyze the function of genes that cause embryonic lethality when mutated. Similarly, as many genes are widely expressed in different tissues, most current applications of CRISPR technology are less amenable to tissue-specific analysis of genetic function.
Here, we describe the generation of transgenic mice that express Cas9 exclusively in cardiomyocytes. In proof-of-concept experiments using Adeno-Associated Virus to deliver single-guide RNA (sgRNA) against Myh6, we demonstrate robust cardiac-specific genomic editing, termed cardioediting, at the Myh6 locus. Ensuing cardiac failure in these mice confirms the effectiveness of this model for cardiac-specific genetic loss of function. These cardiac Cas9-expressing animals will be useful for disease modeling, cardiac gene editing, and exploring potential gene therapies in the context of cardiac disease and dysfunction.
Results
Generation of Myh6-Cas9 Transgenic Mice.
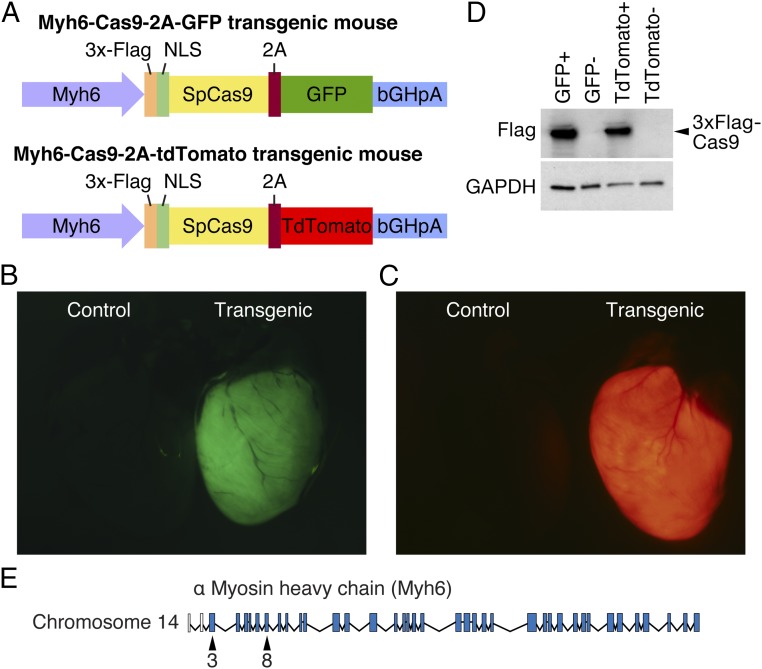
To perform cardiac-specific genome editing with CRISPR/Cas9, we modified a construct that expressed Cas9 from Streptococcus pyogenes, together with a GFP tag, pSpCas9-2A-GFP (11), by replacing the CBh promoter with the promoter for Myh6 (17), allowing expression of Cas9 exclusively in cardiomyocytes. In addition, the 2A-GFP fluorescent tag was replaced with a 2A-TdTomato construct, allowing use of either GFP or TdTomato as a fluorescent reporter for monitoring Cas9 expression (Fig. 1A). These constructs were injected into mouse zygotes to generate transgenic Myh6-Cas9 animals. We identified founder lines that robustly expressed Cas9 in cardiomyocytes in both the Myh6-Cas9-2A-GFP (Fig. 1B) and Myh6-Cas9-2A-TdTomato lines (Fig. 1C).
Fig. 1.
Generation of Myh6-Cas9 transgenic mice. (A) Constructs encoding Myh6-Cas9-2A-GFP and Myh6-Cas9-2A-TdTomato were generated and injected in murine zygotes to generate transgenic mouse lines that express Cas9 exclusively in cardiomyocytes. (B) Fluorescent microscopy confirms that GFP is robustly expressed in the heart of Myh6-Cas9-2A-GFP transgenic animals. (Left) A negative littermate control. (C) Myh6-Cas9-2A-TdTomato animals display strong expression of TdTomato in the heart by fluorescent microscopy, whereas a negative littermate control (Left) had no TdTomato expression. (D) Western blot confirms that 3x-Flag–tagged Cas9 is correctly produced in both Myh6-Cas9-2A-GFP and Myh6-Cas9-2A-TdTomato lines. No band was observed in transgene negative littermates, whereas GAPDH was expressed. (E) Gene structure indicating that exon 3 and exon 8 of murine Myh6 were targeted.
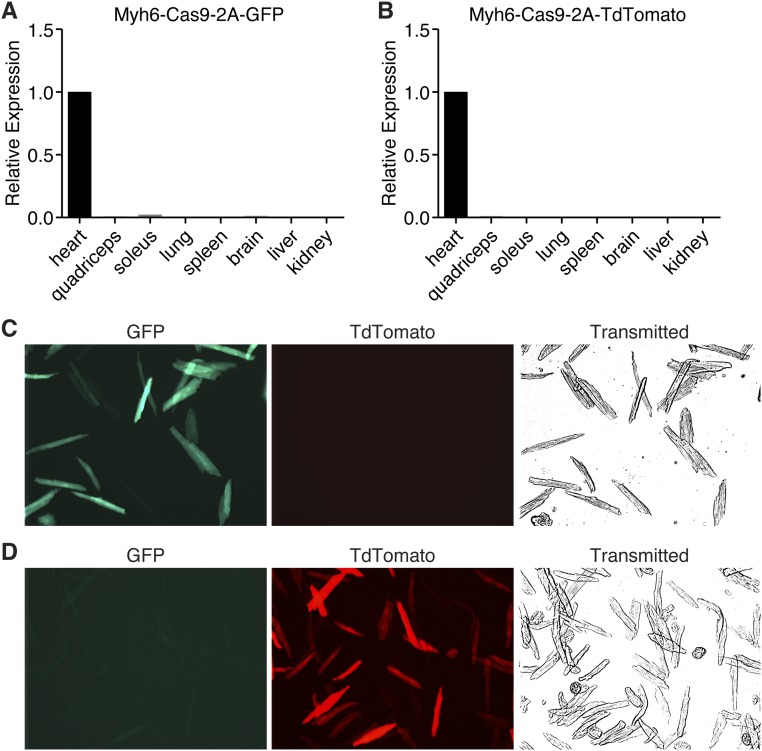
To confirm that Cas9 was correctly produced in these animals, we examined expression of Cas9 by both real-time quantitative PCR (RT-qPCR) (Fig. S1 A and B) and Western blot analysis for Flag-tagged Cas9 (Fig. 1D). Cas9 was robustly expressed in the heart, but was not detected in any other tissue examined by qPCR (Fig. S1 A and B) consistent with the cardiac specificity of the Myh6 promoter. In addition, we isolated cardiomyocytes and examined expression of the Cas9 fluorescent reporter in these cells. GFP or TdTomato was expressed in all cardiomyocytes, suggesting robust expression of the transgene (Fig. S1 C and D). Importantly, we have observed no overt defects in Cas9 transgenic animals, suggesting that high expression of Cas9 is not harmful or toxic, consistent with previous reports (18).
Fig. S1.
Cardiac expression of Cas9. (A) Cas9 expression was observed only in the heart and not in any other tissues examined by qPCR of Myh6-Cas9-2A-GFP animals. (B) Cas9 is correctly targeted to the cardiomyocytes of Myh6-Cas9-2A-TdTomato animals, as no Cas9 was observed in any other tissue by qPCR. (C) Myh6-Cas9-2A-GFP animals show robust expression of GFP in all isolated cardiomyocytes, indicating strong Cas9 expression. (D) Cardiomyocyte isolation of Myh6-Cas9-2A-TdTomato animals confirmed that Cas9 is strongly expressed in cardiomyocytes, as assessed by TdTomato fluorescence.
AAV9-Driven Expression of sgRNA Against Myh6.
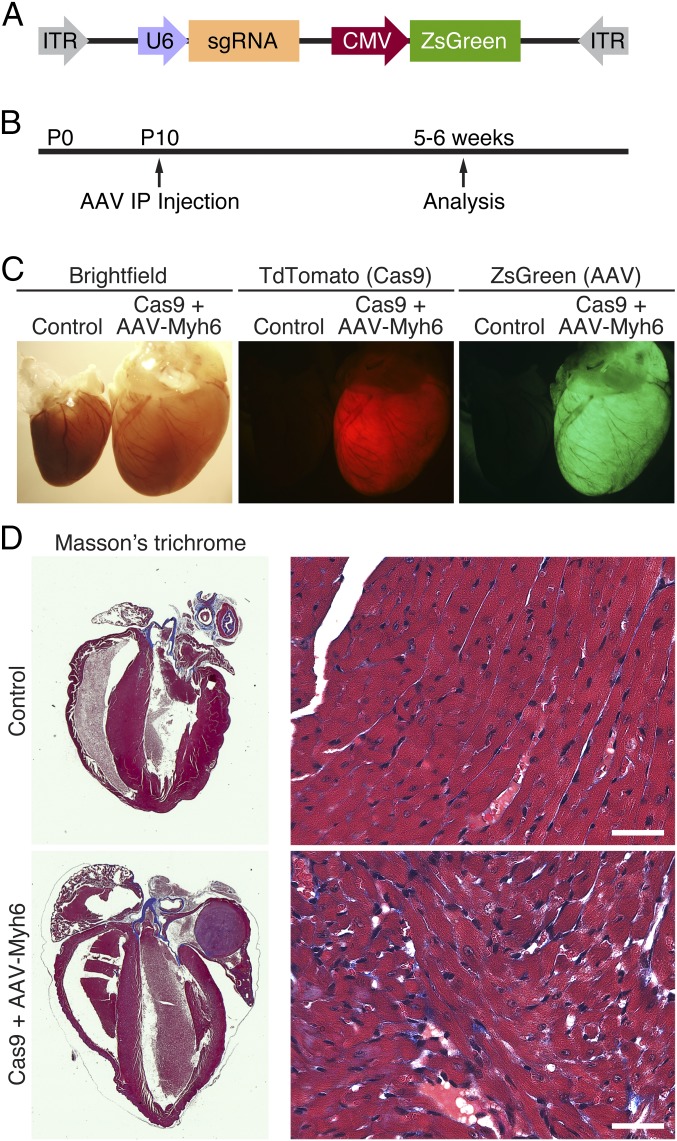
To determine the utility of this mouse model, we chose to use Adeno-Associated Virus 9 (AAV9) to drive expression of an sgRNA against Myh6 (Fig. 1E). We identified and tested sgRNAs against exon 3 and exon 8 of Myh6, confirming in tissue culture using 10T1/2 cells that they exhibited activity against the correct locus. These guides were selected based on their specificity for the Myh6 locus, as well as minimization of potential off-target sites (12 potential off-targets in coding regions for Myh6 exon 3 sgRNA and 10 potential off-targets in coding regions for Myh6 exon 8 sgRNA). Subsequently, the U6 promoter followed by the sgRNA against Myh6 and the guide RNA scaffold were cloned into an AAV backbone (Fig. 2A). This AAV vector also contained the CMV-promoter–driving expression of the fluorescent protein ZsGreen, allowing monitoring of cardiomyocyte transduction after AAV administration. The vector plasmid was packaged into an AAV9 capsid serotype known to efficiently transduce the heart upon systemic infusion (19).
Fig. 2.
AAV9-driven expression of Myh6 sgRNA. (A) Myh6 sgRNA under the control of the U6 promoter was cloned into an AAV9 backbone, together with a CMV-driven ZsGreen reporter. (B) Animals were injected intraperitoneally at postnatal day 10 (P10) and subsequently analyzed 5–6 wk later. (C) An example of a Myh6-Cas9-2A-TdTomato heart (red, Center) that also received AAV-sgRNA against Myh6 exon 3 (green, Right). Compared with a littermate control animal, hearts from animals that received both Cas9 and sgRNA against Myh6 displayed extreme cardiac dilation and hypertrophy. (D) Histological section of a control heart and a heart that contained both Cas9 and AAV-sgRNA against Myh6 exon 3. Edited hearts displayed thinning of the ventricular walls and massive dilation of both the atria and ventricles. Minimal fibrosis was observed by Masson’s trichrome staining. (Scale bar, 40 μm.)
Myh6 Expression Is Decreased After Knockdown via AAV in Cas9 Animals.
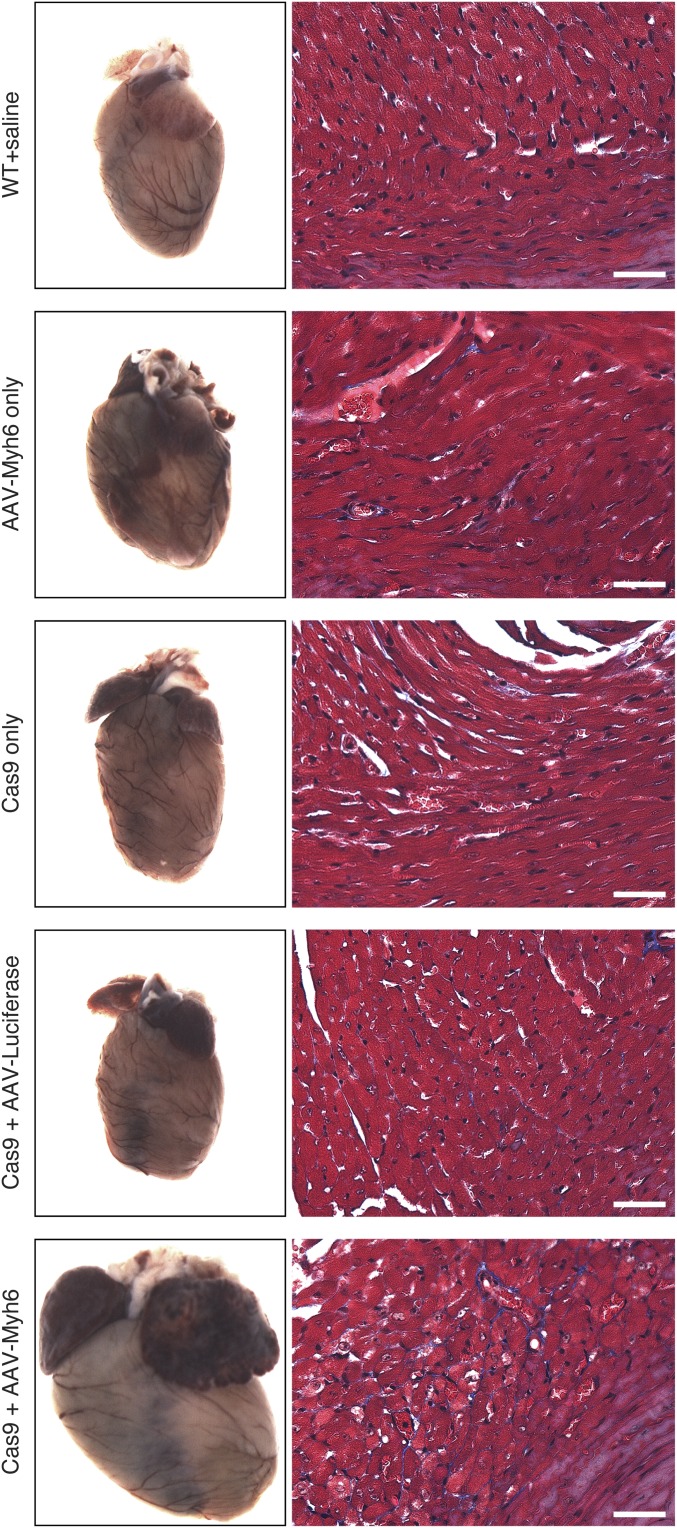
Animals were injected intraperitoneally with a single dose of 1 × 1012 viral genomes of AAV9-Myh6 sgRNA targeting exon 3 or a control injection of saline at postnatal day 10 (P10) (Fig. 2B). Five weeks later, robust expression of ZsGreen in the hearts of these animals was observed (Fig. 2C), with ∼75% of isolated cardiomyocytes showing some degree of ZsGreen expression, confirming that AAV was effectively transduced into the heart. Consistent with effective transduction of AAV and knockdown of Myh6, we observed extreme cardiac hypertrophy in animals that were cardioedited (Fig. 2C). Histological analysis of Cas9+ animals with AAV-Myh6 sgRNA demonstrated extensive dilation of both atria and ventricles, as well as thinning of the ventricle walls, although minimal fibrosis was observed (Fig. 2D). Importantly, we also injected AAV encoding sgRNA against luciferase and observed no changes in heart morphology 12 wk after injection (Fig. S2), confirming that the cardiac dilation observed after Myh6 knockdown was not a result of toxicity from AAV or an AAV/sgRNA complex. Furthermore, hearts of control mice at 3 mo of age, including Cas9+ alone, Cas9+ injected with AAV-luciferase sgRNA, and injected with AAV-Myh6 sgRNA, showed similar histology as WT mice, as seen by Masson’s trichrome stain, suggesting that at this age none of these components alone contribute to cardiac pathology. In contrast, knockdown of Myh6 via AAV-Myh6 in a Cas9+ background resulted in fibrosis, indicative of the cardiomyopathy observed in edited animals (Fig. S2).
Fig. S2.
Knockdown of Myh6 with Cas9 results in cardiac failure and hypertrophy. No overt cardiac abnormalities were observed in animals that (i) received AAV-sgRNA against Myh6 only; (ii) were Cas9+ only; or (iii) were both Cas9+ and received AAV-sgRNA against Luciferase, confirming that there is no detectable toxicity associated with either Cas9 or AAV in the heart. In contrast, animals that were Cas9+ and received AAV-sgRNA against Myh6 displayed massive cardiac enlargement characteristic of heart failure. Tissues were isolated 12 wk after AAV delivery. Fibrosis, as assessed by Masson’s trichrome stain (Right), was observed only when both Cas9 and AAV-sgRNA against Myh6 were present. (Scale bar, 40 μm.)
Cardioediting of Myh6 Results in Cardiac Failure and Hypertrophy.
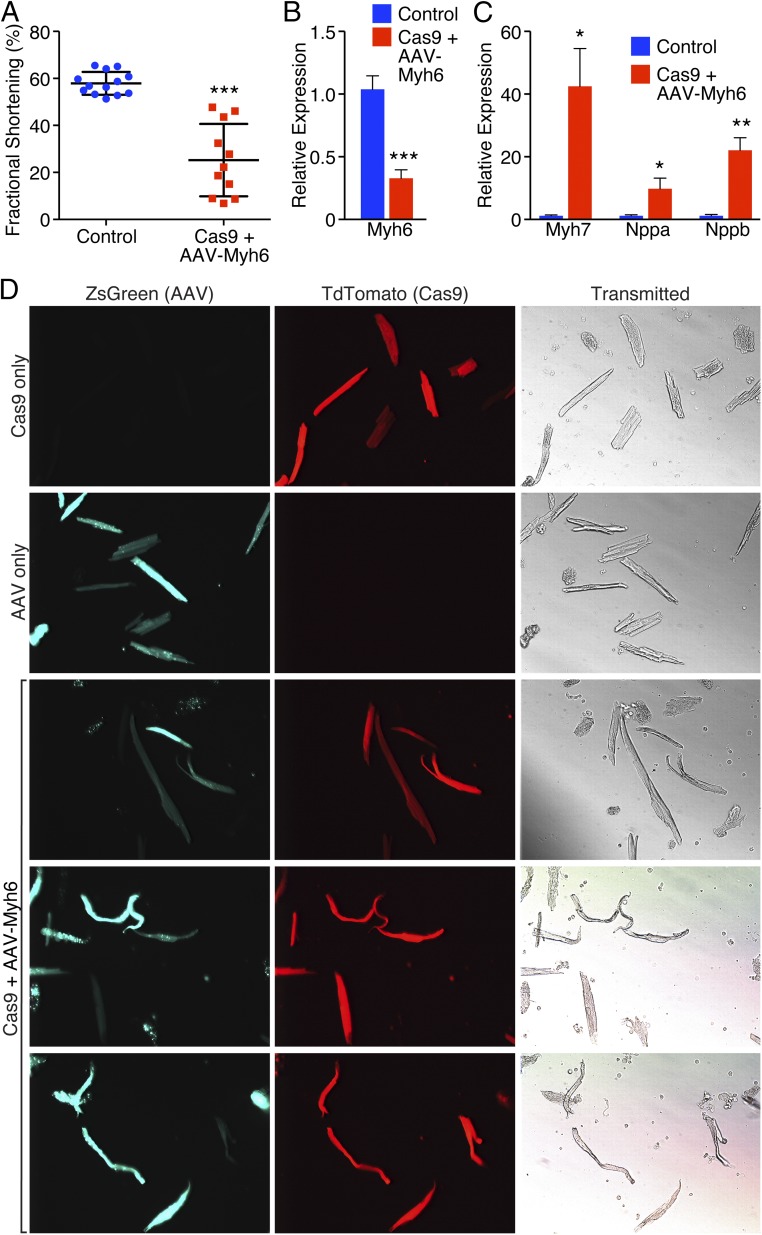
Consistent with the cardiac dilation noted (Fig. 2 C and D), we also observed a marked reduction in fractional shortening in Myh6 edited animals, suggesting that loss of Myh6 severely impaired cardiac performance (Fig. 3A). Similarly, by qPCR, we observed a strong decrease in expression of Myh6 (Fig. 3B), concomitant with an up-regulation of Myh7 expression (Fig. 3C), suggesting that these animals had undergone compensatory myosin switching, consistent with the presence of heart failure (20, 21). In addition, natriuretic peptide A (Nppa) and natriuretic peptide B (Nppb), sensitive markers of cardiac stress (22), were also up-regulated as measured by qPCR, suggesting cardiac dysfunction (Fig. 3C). Isolation of cardiomyocytes from animals that were both Cas9+ and injected with AAV-Myh6 sgRNA revealed the presence of cardiomyocytes that appeared both elongated and enlarged, indicative of heart failure (Fig. 3D) (23). In addition, we noted cells that were fragile, misshapen, bent, and floppy, lacking the characteristic rod-like, striated appearance of adult cardiomyocytes, suggesting that the lack of myosin perturbed cytoskeletal integrity. Using genomic DNA obtained from isolated cardiomyocytes, we observed robust editing at the Myh6 exon 3 locus using a T7 Endonuclease I assay, used to assess genomic editing (Fig. S3A), indicating the correct genomic location was targeted for cleavage by Cas9.
Fig. 3.
Cardioediting of Myh6 results in cardiac failure. (A) Echocardiography revealed a significant decrease in fractional shortening of animals that were Cas9+ and received AAV-sgRNA against Myh6 compared to littermate controls. ***P < 0.001; n = 13 control animals, 11 edited animals. (B) qPCR revealed that Myh6 expression was significantly down-regulated following knockdown of Myh6. ***P < 0.001; n = 5 control animals, 7 edited animals. (C) Myh7, Nppa, and Nppb were significantly up-regulated as detected by qPCR after Myh6 knockdown. *P < 0.05, **P < 0.01; n = 5 control animals, 7 edited animals. (D) Isolated cardiomyocytes from a (i) Myh6-Cas9-2A-TdTomato animal (top panel), (ii) wild-type animal that received AAV-sgRNA only (second panel), and (iii) Myh6-Cas9-2A-TdTomato positive animal that received AAV-sgRNA (bottom three panels). Edited animals display cardiomyocyte elongation characteristic of that seen in heart failure, as well as cardiomyocytes that are bent and floppy, suggesting loss of sarcomeric integrity and rigidity.
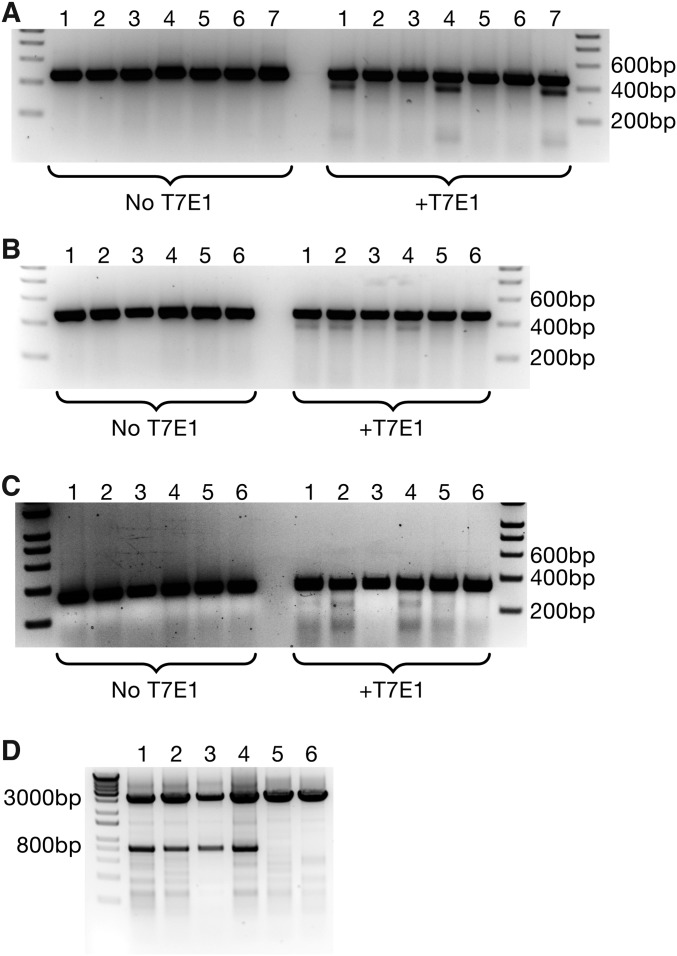
Fig. S3.
T7 Endonuclease I assays on isolated cardiomyocytes. (A) Cardiomyocytes isolated from edited animals displayed a positive T7 assay at the targeted locus of Myh6, confirming the induction of indels by Cas9. Lanes 1, 4, and 7 contained genomic DNA from edited animals, whereas lanes 2, 3, 5, and 6 were from control animals. (B) T7 Endonuclease I Assay on Myh6 exon 3 from animals that received dual AAV-sgRNA against Myh6 exons 3 and 8. Lanes 1–4 were from edited animals, and all show induction of indels, whereas lanes 5–6 were from controls. (C) A T7 Endonuclease I Assay on Myh6 exon 8 using genomic DNA from double–AAV-sgRNA animals revealed the presence of indels at the Myh6 exon 8 locus. Lanes 1–4 were from edited animals, whereas lanes 5–6 were from controls. (D) PCR using a forward primer that anneals upstream of Myh6 exon 3 and a reverse primer that anneals downstream of Myh6 exon 8 indicates that animals that were both Cas9+ and received AAV-sgRNA against both exons 3 and 8 of Myh6 have large genomic deletions between the two guide sites. The lower band indicates deletion of the intervening fragment, whereas the upper band is unedited genomic DNA. Lanes 1–4 were from edited animals, whereas lanes 5–6 were from controls.
Double-Guide Knockdown of Myh6.
As use of a single guide limits the types of indels that can be obtained in a target gene, we sought to identify ways to increase the types of mutations we could recover. We hypothesized that injecting virus that targeted two separate exons might increase the types of mutations induced, both by doubling the Myh6 loci that could be edited via induction of indels and by allowing for either deletion or inversion of the intervening fragment of genomic DNA found between the two guide sites (24).
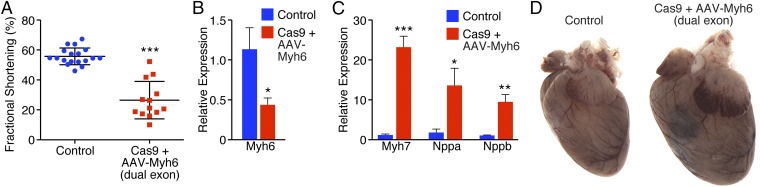
To test this theory, animals were intraperitoneal (i.p.)-injected with a mixture of virus at P10 that contained sgRNAs against both exon 3 and exon 8 of Myh6 (Fig. 1E). Within 3 wk of virus injection, a strong reduction in fractional shortening was observed in animals that were both Cas9+ and received dual AAV-Myh6 sgRNA (Fig. 4A). Similarly, by 4 wk postinjection, myosin switching by qPCR (Fig. 4 B and C) and induction of Nppa and Nppb (Fig. 4C) were observed. Furthermore, extensive cardiac dilation and hypertrophy consistent with heart failure were evident (Fig. 4D).
Fig. 4.
Double-guide knockdown of Myh6 results in cardiac failure and hypertrophy. (A) Animals that received AAV9-sgRNA against both Myh6 exon 3 and exon 8 displayed a robust decrease in fractional shortening within 3 wk of AAV delivery. ***P < 0.001; n = 18 control animals, 13 edited animals. (B) Four weeks after AAV delivery of dual sgRNAs, animals showed decreased expression of Myh6 by qPCR. *P < 0.05; n = 7 control animals, 6 edited animals. (C) After Myh6 knockdown with two AAV-sgRNAs, animals showed elevated levels of Myh7, Nppa, and Nppb by qPCR within 4 wk of AAV delivery. *P < 0.05, **P < 0.01, ***P < 0.001; n = 7 control animals, 6 edited animals. (D) Animals that were Cas9+ and received dual AAV-sgRNA against Myh6 exhibited cardiac dilation within 4 wk of AAV delivery.
To confirm that both exons 3 and 8 of Myh6 were targeted with this strategy, T7 Endonuclease I assays were performed on isolated cardiomyocytes. We observed editing at both the Myh6 exon 3 (Fig. S3B) and exon 8 (Fig. S3C) loci, consistent with the delivery of sgRNA targeting both genomic locations. Importantly, using primers that flank the sgRNA sites of both exon 3 (Forward Primer) and exon 8 (Reverse Primer), strong deletion of the intervening genomic DNA was observed, as reflected in the smaller ∼800 base pair PCR product that is visible in edited animals (Fig. S3D). Together, these data suggest that the delivery of sgRNA targeting two locations could be used to increase the types of mutations induced via CRISPR-mediated genomic editing.
Discussion
Here, we have generated a transgenic Cas9 system that enables robust genomic editing exclusively in cardiomyocytes. Through proof-of-concept experiments, we demonstrate that the knockdown of Myh6 quickly and efficiently induces massive cardiac dilation and heart failure, within 3 wk of AAV delivery of sgRNA. We anticipate that cardioediting can be extended to many other genes, enabling the rapid assessment of gene functions in the heart. This system will be an especially valuable tool for investigators interested in studying the function of genes that cause embryonic lethality when mutated or are expressed in a broad range of tissues. Although no overt toxicity due to constitutive expression of Cas9 was observed, it remains unclear if high levels of Cas9 could be problematic over long periods of time, similar to the cardiotoxic effects reported with both Cre (25) and GFP (26). However, the use of a tamoxifen-inducible Myh6 promoter driving Cas9 expression could help alleviate this potential problem by allowing for temporally controlled induction of Cas9. Additionally, the use of newer versions of Cas9 that exhibit strong reductions in off-target cleavage (27) will aid in increasing the fidelity of Cas9-mediated gene knockdown, helping reduce the likelihood of nonspecific genomic editing.
The double-sgRNA approach used here increases the number of potential ways in which to disrupt the coding sequence of the target gene. As a result, we suggest that simultaneous editing with two guides that target proximal regions of the same gene could enhance the knockdown of genes using postnatal delivery of sgRNA by AAV9 in a Cas9+ transgenic animal. Furthermore, the use of two guide RNAs enables the deletion of enhancers, lncRNAs, or miRNA clusters, which are not as easily targeted with the use of one sgRNA, thus expanding the types of genomic editing that can be performed with this system.
The Cas9-mediated knockdown of Myh6 demonstrated here confirms the essentiality of MYH6 in cardiac function and its role in the development and progression of heart failure (28). Importantly, the phenotype observed following Myh6 knockdown is more severe than that observed in adult mice that are heterozygous for Myh6, which display modest adult onset heart failure (29), suggesting that this approach leads to the loss of more than 50% of Myh6. It is likely that the more severe phenotype is due to the loss of both copies of Myh6 in a fraction of myocytes as opposed to the loss of one allele in all myocytes in the heterozygous Myh6 animals. Alternatively, it is possible that the enhanced phenotype is due to the inability of the animal to adequately compensate for a postnatal loss of myosin rather than a loss from early development, as in the full-body heterozygous mice.
CRISPR/Cas9 has revolutionized the ease with which genomic editing can be completed. Since mammalian CRISPR-based genetic editing was first reported (6), the system has been used to correct mutations in a variety of disease models, including Muscular Dystrophy (30), Cystic Fibrosis (31), and Tyrosinemia (32). Together, these studies are indicative of the broad application and uses of the technology for genetic manipulation and correction. Future uses of CRISPR are likely to involve attempted correction of diseases in humans, after safety concerns are addressed (33, 34).
Despite the rapid progress that has been made in the CRISPR/Cas9 field, challenges remain. One major limitation has been the ability to deliver components of the CRISPR/Cas9 complex, particularly via AAV, due to the packaging limit of the virus (35). Here, we circumvent this limitation by constitutively expressing Cas9 in cardiomyocytes, allowing us to exclusively use AAV9 to deliver sgRNA, which is well within the packaging limit of the virus. In principle, cardioediting will enable rapid and efficient analysis of function of any coding gene in the heart by delivering a single dose of AAV9 virus encoding an sgRNA against a gene of interest.
Materials and Methods
Generation of Transgenic Mice.
All animal experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of University of Texas Southwestern Medical Center. Addgene plasmid #48138, pSpCas9(BB)-2A-GFP (11), a gift from Feng Zhang, Massachusetts Institute of Technology, was digested overnight with EcoRI to remove the 2A-GFP–coding region. The digestion product was gel-purified using a Promega SV Gel and PCR Cleanup System. The 2A-TdTomato construct was generated by cloning the T2A self-cleaving peptide (EGRGSLLTCGDVEENPGP) upstream of the coding sequence of the red fluorescent protein TdTomato. The 2A-TdTomato construct was subsequently PCR-amplified and recombined in-frame with Cas9 in the PX458 vector using infusion cloning (Clontech), with the following primers:
F: 5′-AAAGAAAAAGGAATTCGCTAGCGAGGGCAGAGGA-3′
R: 5′-GCTCTAGTTAGAATTCATCGATTTACTTGTACAGCTC-3′.
To add the Myh6 promoter to both the Cas9-2A-GFP and -TdTomato reporter constructs, the plasmids were digested overnight with XbaI and AgeI to remove the CBh promoter. The Myh6 promoter was amplified from the BamHI to SalI sites using Phusion Taq Polymerase off the Myh6 PBS2-Sk+ vector (17). The Myh6 promoter was recombined into the digested PX458 vector using infusion cloning (Clontech) and the following primers:
F: 5′-GACAAATGGCTCTAGAGGATCCTGCAAGGTCACACA-3′
R: 5′-CCATGGTGGCACCGGTGTCGACTCAAACTCTTATGG-3′
All constructs were confirmed by sequencing. The Myh6-Cas9-2A-GFP construct was digested from the backbone overnight with BamHI and NotI, whereas the Myh6-Cas9-2A-TdTomato construct was digested overnight with BamHI and NarI. Both digestion products were gel-purified before injection. The transgenic constructs were injected in the pronucleus of murine embryos followed by implantation in surrogate dams. Founder animals (F0) were identified by tail biopsy followed by PCR for the presence of either GFP or TdTomato and subsequently bred to wild-type animals.
Genotyping of Transgenic Mice.
Transgenic mice were genotyped based on the presence of either the GFP or TdTomato construct. Tail biopsies were digested in 100 μL of 25-mM NaOH, 0.2-mM EDTA (pH 12) for 20 min at 95 °C. Tails were briefly centrifuged followed by addition of 100 μL of 40-mM Tris⋅HCl (pH 5) and mixed to homogenize. Two microliters of this reaction was used for subsequent PCR reactions with the primers below, followed by gel electrophoresis:
GFP_F: 5′-GGTGAACTTCAAGATCCGCC-3′
GFP_R: 5′-CTTGTACAGCTCGTCCATGC-3′
TdTomato_F: 5′-ACATGGCCGTCATCAAAGA-3′
TdTomato_R: 5′-CTTGTACAGCTCGTCCATGC-3′
sgRNA Identification and Cloning.
Myh6 exon 3, Myh6 exon 8, and Luciferase guide RNAs were identified using crispr.mit.edu. Guide sequences were cloned into Addgene plasmid #42230 (6), a gift from Feng Zhang, using the following primers:
Myh6 exon 3_F: 5′- CACCGTTAAGGCCAAGGTCGTGTCC -3′
Myh6 exon 3_R: 5′- AAACGGACACGACCTTGGCCTTAAC-3′
Myh6 exon 8_F: 5′- CACCGTATCCAGGCTAACCCCGCTC-3′
Myh6 exon 8_R: 5′- AAACGAGCGGGGTTAGCCTGGATAC-3′
Luciferase_F: 5′- CACCGTCCAGCGGATAGAATGGCGC-3′
Luciferase_R: 5′- AAACGCGCCATTCTATCCGCTGGAC-3′
Guide sequences were tested in culture using 10T1/2 cells before cloning into the AAV backbone.
AAV Production and Delivery.
Guide sequences and guide RNA scaffold were digested using Nde I, which cut twice in the donor plasmid backbone, and subcloned into the corresponding sites of pZac2.1-U6- CMV-ZsGreen (University of Pennsylvania) containing the AAV vector backbone or into the pZac2.1-U6- CMV-ZsGreen backbone with the ZsGreen coding sequence removed between the Sal1 sites.
Infectious recombinant AAV vector particles were generated in HEK293T cells cultured in roller bottles by a cross-packaging approach whereby the vector genome was packaged into AAV capsid serotype-9 (36). Viral stocks were obtained by PEG precipitation and CsCl2 gradient centrifugation. Full viral particles obtained from the gradient were extensively dialyzed in PBS and stored in aliquots at −80 °C until use. The physical titer of recombinant AAVs was determined by quantifying vector genomes (vg) packaged into viral particles by real-time PCR against a standard curve of a plasmid containing the vector genome (37). Values obtained were in the range of 5 × 1012 to 5 × 1013 vg per milliliter.
Postnatal day 10 (P10) mice were administered 1 × 1012 viral genomes of AAV (single-guide), diluted in 50 μL of saline. For double-guide injections, 5 × 1011 viral genomes of each AAV guide were mixed, diluted in 50 µL of saline, and delivered via i.p. injection at P10. Control animals received 50 µL of saline.
Quantitative Real-Time PCR.
RNA was isolated from whole-heart tissue using TRIzol (Invitrogen) according to manufacturer’s instructions. cDNA was synthesized using BioRad iScript Reverse Transcription Supermix. qPCR was performed using Applied Biosystems TaqMan probes for Myh6, Myh7, Nppa, and Nppb and using Sybr Green for Cas9 (F Primer: 5′-GGACTCCCGGATGAACACTA-3′; R Primer:5′-TCGCTTTCCAGCTTAGGGTA-3′). All reactions were normalized using eukaryotic 18s rRNA endogenous control (Applied Biotechnologies). Experiments were performed on a StepOne Real-Time PCR System (Life Technologies) and analyzed using the delta–delta Ct method.
Western Blot Analysis.
Isolated hearts were snap-frozen in liquid nitrogen. Lysates were generated by pulverizing cardiac tissue on ice in RIPA Buffer (150 mM NaCl, 1% vol/vol Ipegal CA-630, 50 mM Tris-Cl pH 8.3, 0.5% wt/vol sodium deoxycholate, 0.1% wt/vol SDS) together with additional protease inhibitors (complete ULTRA mini tablet, Roche). BCA Protein Assay Kit (Pierce) was used to determine protein concentrations. Lysates were separated using any kD Mini-Protean TGX precast gels (BioRad). Samples were transferred onto Immobilon P membranes (Millipore) and blocked for 1 h at room temperature in 5% (wt/vol) nonfat dry milk in Tris-buffered saline with Tween 20 (TBST). Primary antibodies were hybridized overnight at 4 °C [GAPDH (Millipore), 1:1,000 dilution; Flag-HRP–conjugated (Sigma), 1:1,000 dilution]. Membranes were washed three times for 5 min each in TBST. Blots for GAPDH were incubated with HRP-conjugated secondary antibody (BioRad) at a 1:20,000 dilution for 1 h at room temperature. Chemiluminescent substrate was used to develop blots, which were then exposed to audioradiographic film.
Echocardiography.
Cardiac function was determined by echocardiography on conscious animals at either 3 wk post-AAV delivery (double-guide animals) or 5–6 wk post-AAV delivery (single-guide animals). This analysis was performed using a Visual SonicsVevo 2100 system equipped with a 35-MHz transducer. Fractional shortening (FS) of the left ventricle was determined using Left Ventricle Internal Dimension at Diastole (LVIDd) and Left Ventricle Internal Dimension at Systole (LVIDs). FS was calculated according to the formula FS(%)= [(LVIDd − LVIDs)/LVIDd] × 100.
Cardiomyocyte Isolation.
Cardiomyocytes were isolated from adult animals as previously described (38, 39). Animals were initially anesthetized with 3% isoflurane and subsequently maintained with 1% isoflurane. Hearts were removed from the animals. Aortic cannulation was performed using a Langendorff Apparatus with constant flow. Cardiac digestion was achieved by perfusion with a Tyrode’s solution consisting of 0.2 mg/mL Liberase DH (Roche), 0.14 mg/mL Trypsin (Gibco/Invitrogen), 0.02 mM CaCl2, 10 mM glucose, 5 mM Hepes, 5.4 mM KCl, 1.2 mM MgCl2, 150 mM NaCl, and 2 mM sodium pyruvate (pH 7.4). Subsequent to tissue softening, the heart was isolated and delicately minced, filtered, and equilibrated at room temperature using Tyrode’s solution also containing 200 μM CaCl2 and 1% BSA.
Histological Analysis.
Hearts were isolated and fixed in 4% paraformaldehyde in PBS for 48 h at 4 °C followed by paraffin embedding and sectioning. Masson’s trichrome stain was performed according to standard protocols.
T7 Endonuclease Assay.
Genomic DNA was isolated from either whole-heart or isolated cardiomyocytes using a DNeasy Blood and Tissue Kit (Qiagen). Exons were PCR-amplified using Taq Polymerase (New England Biolabs) and the following primers:
Myh6 exon 3 F Primer: 5′-AGGCACCCTTACCCCACATA-3′
Myh6 exon 3 R Primer: 5′-CAACCCCTTTCCCTAAGCCG-3′
Myh6 exon 8 F Primer: 5′-GGTCGGTGCAGACAATCTTT-3′
Myh6 exon 8 R Primer: 5′-CCATTTTCTTGACACATTTGAGG-3′
PCR products were hybridized and subsequently digested with T7 Endonuclease I for 30 min at 37 °C. Digestion products were separated using standard agarose gels (1.5%).
To detect deletion of the genomic DNA between the exon 3 and exon 8 guide sites, the following primers were used:
F: 5′-AGGCACCCTTACCCCACATA-3′
R: 5′-GGGAAAACAGCTACCAGGCT-3′
Statistical Analysis.
For all experiments, pooled data consist of studies using both Myh6-Cas9-2A-GFP and Myh6-Cas9-2A-TdTomato animals. Controls represent a compilation of data from animals that were (i) wild type and received a saline injection, (ii) wild type and received injection of AAV, and (iii) Cas9+ and received a saline injection. All qPCR statistics are displayed as mean ± SEM while echocardiograms are displayed as mean ± SD. An unpaired t test with Welch’s correction was used to determine statistical significance. Any result with a P value <0.05 was considered statistically significant.
Acknowledgments
We thank the members of the E.N.O. laboratory for helpful discussions, Jose Cabrera for help with images, and Wei Tan for echocardiography assistance. This work was supported by grants from the National Institutes of Health (Grants HL-077439, HL-111665, HL-093039, DK-099653, and U01-HL-100401), Foundation Leducq Networks of Excellence (Grant 14CVD04 to E.N.O. and M.G.), Cancer Prevention and Research Institute of Texas, the Robert A. Welch Foundation (Grant 1-0025 to E.N.O.), and by Grant PRIN 2010RNXM9C from the Ministero Istruzione Universita Ricera, Italy (to M.G.).
Footnotes
The authors declare no conflict of interest.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1523918113/-/DCSupplemental.
References
- 1.Capecchi MR. Gene targeting in mice: Functional analysis of the mammalian genome for the twenty-first century. Nat Rev Genet. 2005;6(6):507–512. doi: 10.1038/nrg1619. [DOI] [PubMed] [Google Scholar]
- 2.Doudna JA, Charpentier E. Genome editing. The new frontier of genome engineering with CRISPR-Cas9. Science. 2014;346(6213):1258096. doi: 10.1126/science.1258096. [DOI] [PubMed] [Google Scholar]
- 3.Barrangou R, et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science. 2007;315(5819):1709–1712. doi: 10.1126/science.1138140. [DOI] [PubMed] [Google Scholar]
- 4.Jinek M, et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337(6096):816–821. doi: 10.1126/science.1225829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jinek M, et al. RNA-programmed genome editing in human cells. eLife. 2013;2:e00471. doi: 10.7554/eLife.00471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Cong L, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–823. doi: 10.1126/science.1231143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mali P, et al. RNA-guided human genome engineering via Cas9. Science. 2013;339(6121):823–826. doi: 10.1126/science.1232033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bibikova M, Golic M, Golic KG, Carroll D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics. 2002;161(3):1169–1175. doi: 10.1093/genetics/161.3.1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell. 2014;157(6):1262–1278. doi: 10.1016/j.cell.2014.05.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cong L, Zhang F. Genome engineering using CRISPR-Cas9 system. Methods Mol Biol. 2015;1239:197–217. doi: 10.1007/978-1-4939-1862-1_10. [DOI] [PubMed] [Google Scholar]
- 11.Ran FA, et al. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8(11):2281–2308. doi: 10.1038/nprot.2013.143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hwang WY, et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol. 2013;31(3):227–229. doi: 10.1038/nbt.2501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hwang WY, et al. Heritable and precise zebrafish genome editing using a CRISPR-Cas system. PLoS One. 2013;8(7):e68708. doi: 10.1371/journal.pone.0068708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Shen B, et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 2013;23(5):720–723. doi: 10.1038/cr.2013.46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wang H, et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell. 2013;153(4):910–918. doi: 10.1016/j.cell.2013.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Niu Y, et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell. 2014;156(4):836–843. doi: 10.1016/j.cell.2014.01.027. [DOI] [PubMed] [Google Scholar]
- 17.Gulick J, Subramaniam A, Neumann J, Robbins J. Isolation and characterization of the mouse cardiac myosin heavy chain genes. J Biol Chem. 1991;266(14):9180–9185. [PubMed] [Google Scholar]
- 18.Platt RJ, et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell. 2014;159(2):440–455. doi: 10.1016/j.cell.2014.09.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zacchigna S, Zentilin L, Giacca M. Adeno-associated virus vectors as therapeutic and investigational tools in the cardiovascular system. Circ Res. 2014;114(11):1827–1846. doi: 10.1161/CIRCRESAHA.114.302331. [DOI] [PubMed] [Google Scholar]
- 20.Nakao K, Minobe W, Roden R, Bristow MR, Leinwand LA. Myosin heavy chain gene expression in human heart failure. J Clin Invest. 1997;100(9):2362–2370. doi: 10.1172/JCI119776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gupta MP. Factors controlling cardiac myosin-isoform shift during hypertrophy and heart failure. J Mol Cell Cardiol. 2007;43(4):388–403. doi: 10.1016/j.yjmcc.2007.07.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nishikimi T, Maeda N, Matsuoka H. The role of natriuretic peptides in cardioprotection. Cardiovasc Res. 2006;69(2):318–328. doi: 10.1016/j.cardiores.2005.10.001. [DOI] [PubMed] [Google Scholar]
- 23.Kehat I, Molkentin JD. Molecular pathways underlying cardiac remodeling during pathophysiological stimulation. Circulation. 2010;122(25):2727–2735. doi: 10.1161/CIRCULATIONAHA.110.942268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Canver MC, et al. Characterization of genomic deletion efficiency mediated by clustered regularly interspaced palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J Biol Chem. 2014;289(31):21312–21324. doi: 10.1074/jbc.M114.564625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pugach EK, Richmond PA, Azofeifa JG, Dowell RD, Leinwand LA. Prolonged Cre expression driven by the α-myosin heavy chain promoter can be cardiotoxic. J Mol Cell Cardiol. 2015;86:54–61. doi: 10.1016/j.yjmcc.2015.06.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Huang WY, Aramburu J, Douglas PS, Izumo S. Transgenic expression of green fluorescence protein can cause dilated cardiomyopathy. Nat Med. 2000;6(5):482–483. doi: 10.1038/74914. [DOI] [PubMed] [Google Scholar]
- 27.Slaymaker IM, et al. Rationally engineered Cas9 nucleases with improved specificity. Science. 2015 doi: 10.1126/science.aad5227. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Carniel E, et al. Alpha-myosin heavy chain: A sarcomeric gene associated with dilated and hypertrophic phenotypes of cardiomyopathy. Circulation. 2005;112(1):54–59. doi: 10.1161/CIRCULATIONAHA.104.507699. [DOI] [PubMed] [Google Scholar]
- 29.Jones WK, et al. Ablation of the murine alpha myosin heavy chain gene leads to dosage effects and functional deficits in the heart. J Clin Invest. 1996;98(8):1906–1917. doi: 10.1172/JCI118992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Long C, et al. Prevention of muscular dystrophy in mice by CRISPR/Cas9-mediated editing of germline DNA. Science. 2014;345(6201):1184–1188. doi: 10.1126/science.1254445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Schwank G, et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell. 2013;13(6):653–658. doi: 10.1016/j.stem.2013.11.002. [DOI] [PubMed] [Google Scholar]
- 32.Yin H, et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat Biotechnol. 2014;32(6):551–553. doi: 10.1038/nbt.2884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Baltimore D, et al. Biotechnology. A prudent path forward for genomic engineering and germline gene modification. Science. 2015;348(6230):36–38. doi: 10.1126/science.aab1028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lanphier E, Urnov F, Haecker SE, Werner M, Smolenski J. Don’t edit the human germ line. Nature. 2015;519(7544):410–411. doi: 10.1038/519410a. [DOI] [PubMed] [Google Scholar]
- 35.Senís E, et al. CRISPR/Cas9-mediated genome engineering: an adeno-associated viral (AAV) vector toolbox. Biotechnol J. 2014;9(11):1402–1412. doi: 10.1002/biot.201400046. [DOI] [PubMed] [Google Scholar]
- 36.Inagaki K, et al. Robust systemic transduction with AAV9 vectors in mice: Efficient global cardiac gene transfer superior to that of AAV8. Mol Ther. 2006;14(1):45–53. doi: 10.1016/j.ymthe.2006.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zentilin L, Marcello A, Giacca M. Involvement of cellular double-stranded DNA break binding proteins in processing of the recombinant adeno-associated virus genome. J Virol. 2001;75(24):12279–12287. doi: 10.1128/JVI.75.24.12279-12287.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Makarewich CA, et al. Transient receptor potential channels contribute to pathological structural and functional remodeling after myocardial infarction. Circ Res. 2014;115(6):567–580. doi: 10.1161/CIRCRESAHA.115.303831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Jaleel N, et al. Ca2+ influx through T- and L-type Ca2+ channels have different effects on myocyte contractility and induce unique cardiac phenotypes. Circ Res. 2008;103(10):1109–1119. doi: 10.1161/CIRCRESAHA.108.185611. [DOI] [PMC free article] [PubMed] [Google Scholar]