Abstract
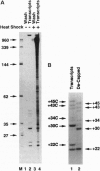
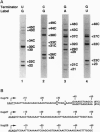
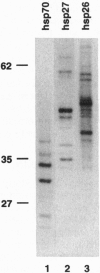
The regulation of many eukaryotic genes occurs at the level of transcriptional elongation. On the uninduced hsp70 gene of Drosophila melanogaster, for example, an RNA polymerase II complex has initiated transcription but has paused early in elongation. In this study, we examine pausing on hsp70 and two of the small heat shock genes (hsp27 and hsp26) at high resolution, using a technique that utilizes paramagnetic particle-mediated selection of terminated run-on transcripts. This technique provides precise information on the distribution of RNA polymerase within each transcription unit. It also details the progression of 5' cap formation on the elongating transcripts. For each gene, we find polymerases paused over a relatively narrow promoter-proximal region. The regions are generally around 20 nucleotides wide, with two preferred pausing positions spaced roughly 10 nucleotides apart or about one turn of the helix. The bulk of capping occurs as transcripts pass between 20 and 30 nucleotides in length. Interestingly, in the three genes examined here, elongational pausing and 5' cap formation appear largely coincident.
Full text
PDF
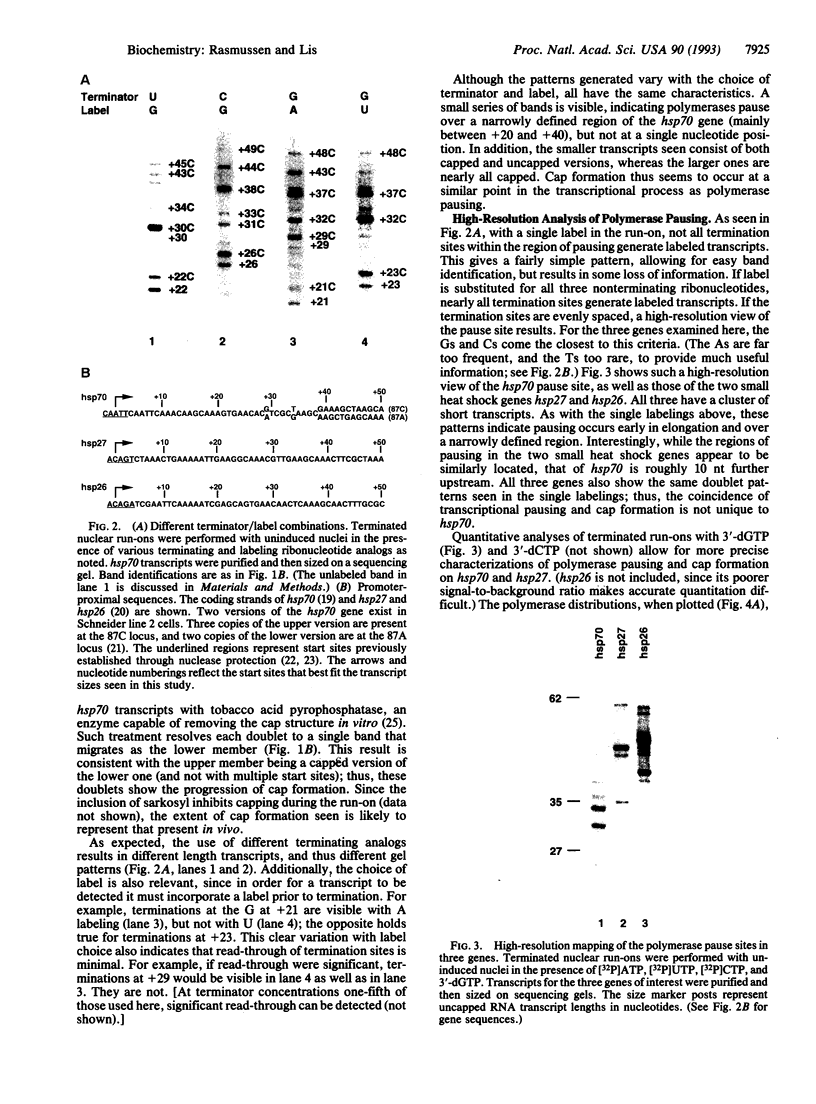


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Banerjee A. K. 5'-terminal cap structure in eucaryotic messenger ribonucleic acids. Microbiol Rev. 1980 Jun;44(2):175–205. doi: 10.1128/mr.44.2.175-205.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biggin M. D., Tjian R. Transcription factors that activate the Ultrabithorax promoter in developmentally staged extracts. Cell. 1988 Jun 3;53(5):699–711. doi: 10.1016/0092-8674(88)90088-8. [DOI] [PubMed] [Google Scholar]
- Chinsky J. M., Maa M. C., Ramamurthy V., Kellems R. E. Adenosine deaminase gene expression. Tissue-dependent regulation of transcriptional elongation. J Biol Chem. 1989 Aug 25;264(24):14561–14565. [PubMed] [Google Scholar]
- Coppola J. A., Field A. S., Luse D. S. Promoter-proximal pausing by RNA polymerase II in vitro: transcripts shorter than 20 nucleotides are not capped. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1251–1255. doi: 10.1073/pnas.80.5.1251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Echalier G., Ohanessian A. In vitro culture of Drosophila melanogaster embryonic cells. In Vitro. 1970 Nov-Dec;6(3):162–172. doi: 10.1007/BF02617759. [DOI] [PubMed] [Google Scholar]
- Edery I., Sonenberg N. Cap-dependent RNA splicing in a HeLa nuclear extract. Proc Natl Acad Sci U S A. 1985 Nov;82(22):7590–7594. doi: 10.1073/pnas.82.22.7590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ensinger M. J., Moss B. Modification of the 5' terminus of mRNA by an RNA (guanine-7-)-methyltransferase from HeLa cells. J Biol Chem. 1976 Sep 10;251(17):5283–5291. [PubMed] [Google Scholar]
- Georgiev O., Mous J., Birnstiel M. L. Processing and nucleo-cytoplasmic transport of histone gene transcripts. Nucleic Acids Res. 1984 Nov 26;12(22):8539–8551. doi: 10.1093/nar/12.22.8539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giardina C., Pérez-Riba M., Lis J. T. Promoter melting and TFIID complexes on Drosophila genes in vivo. Genes Dev. 1992 Nov;6(11):2190–2200. doi: 10.1101/gad.6.11.2190. [DOI] [PubMed] [Google Scholar]
- Gilmartin G. M., McDevitt M. A., Nevins J. R. Multiple factors are required for specific RNA cleavage at a poly(A) addition site. Genes Dev. 1988 May;2(5):578–587. doi: 10.1101/gad.2.5.578. [DOI] [PubMed] [Google Scholar]
- Gilmour D. S., Lis J. T. RNA polymerase II interacts with the promoter region of the noninduced hsp70 gene in Drosophila melanogaster cells. Mol Cell Biol. 1986 Nov;6(11):3984–3989. doi: 10.1128/mcb.6.11.3984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilmour D. S., Thomas G. H., Elgin S. C. Drosophila nuclear proteins bind to regions of alternating C and T residues in gene promoters. Science. 1989 Sep 29;245(4925):1487–1490. doi: 10.1126/science.2781290. [DOI] [PubMed] [Google Scholar]
- Hackett R. W., Lis J. T. DNA sequence analysis reveals extensive homologies of regions preceding hsp70 and alphabeta heat shock genes in Drosophila melanogaster. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6196–6200. doi: 10.1073/pnas.78.10.6196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagler J., Shuman S. A freeze-frame view of eukaryotic transcription during elongation and capping of nascent mRNA. Science. 1992 Feb 21;255(5047):983–986. doi: 10.1126/science.1546295. [DOI] [PubMed] [Google Scholar]
- Hart R. P., McDevitt M. A., Nevins J. R. Poly(A) site cleavage in a HeLa nuclear extract is dependent on downstream sequences. Cell. 1985 Dec;43(3 Pt 2):677–683. doi: 10.1016/0092-8674(85)90240-5. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., Roeder R. G. Separation and partial characterization of three functional steps in transcription initiation by human RNA polymerase II. J Biol Chem. 1985 Jul 5;260(13):8163–8172. [PubMed] [Google Scholar]
- Ingolia T. D., Craig E. A. Primary sequence of the 5' flanking regions of the Drosophila heat shock genes in chromosome subdivision 67B. Nucleic Acids Res. 1981 Apr 10;9(7):1627–1642. doi: 10.1093/nar/9.7.1627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inoue K., Ohno M., Sakamoto H., Shimura Y. Effect of the cap structure on pre-mRNA splicing in Xenopus oocyte nuclei. Genes Dev. 1989 Sep;3(9):1472–1479. doi: 10.1101/gad.3.9.1472. [DOI] [PubMed] [Google Scholar]
- Izban M. G., Luse D. S. The RNA polymerase II ternary complex cleaves the nascent transcript in a 3'----5' direction in the presence of elongation factor SII. Genes Dev. 1992 Jul;6(7):1342–1356. doi: 10.1101/gad.6.7.1342. [DOI] [PubMed] [Google Scholar]
- Johnson P. F., McKnight S. L. Eukaryotic transcriptional regulatory proteins. Annu Rev Biochem. 1989;58:799–839. doi: 10.1146/annurev.bi.58.070189.004055. [DOI] [PubMed] [Google Scholar]
- Kao S. Y., Calman A. F., Luciw P. A., Peterlin B. M. Anti-termination of transcription within the long terminal repeat of HIV-1 by tat gene product. Nature. 1987 Dec 3;330(6147):489–493. doi: 10.1038/330489a0. [DOI] [PubMed] [Google Scholar]
- Karch F., Török I., Tissières A. Extensive regions of homology in front of the two hsp70 heat shock variant genes in Drosophila melanogaster. J Mol Biol. 1981 May 25;148(3):219–230. doi: 10.1016/0022-2836(81)90536-2. [DOI] [PubMed] [Google Scholar]
- Konarska M. M., Padgett R. A., Sharp P. A. Recognition of cap structure in splicing in vitro of mRNA precursors. Cell. 1984 Oct;38(3):731–736. doi: 10.1016/0092-8674(84)90268-x. [DOI] [PubMed] [Google Scholar]
- Krumm A., Meulia T., Brunvand M., Groudine M. The block to transcriptional elongation within the human c-myc gene is determined in the promoter-proximal region. Genes Dev. 1992 Nov;6(11):2201–2213. doi: 10.1101/gad.6.11.2201. [DOI] [PubMed] [Google Scholar]
- Krummel B., Chamberlin M. J. RNA chain initiation by Escherichia coli RNA polymerase. Structural transitions of the enzyme in early ternary complexes. Biochemistry. 1989 Sep 19;28(19):7829–7842. doi: 10.1021/bi00445a045. [DOI] [PubMed] [Google Scholar]
- Krummel B., Chamberlin M. J. Structural analysis of ternary complexes of Escherichia coli RNA polymerase. Deoxyribonuclease I footprinting of defined complexes. J Mol Biol. 1992 May 20;225(2):239–250. doi: 10.1016/0022-2836(92)90918-a. [DOI] [PubMed] [Google Scholar]
- Lattier D. L., States J. C., Hutton J. J., Wiginton D. A. Cell type-specific transcriptional regulation of the human adenosine deaminase gene. Nucleic Acids Res. 1989 Feb 11;17(3):1061–1076. doi: 10.1093/nar/17.3.1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee H., Kraus K. W., Wolfner M. F., Lis J. T. DNA sequence requirements for generating paused polymerase at the start of hsp70. Genes Dev. 1992 Feb;6(2):284–295. doi: 10.1101/gad.6.2.284. [DOI] [PubMed] [Google Scholar]
- Lis J. T., Neckameyer W., Dubensky R., Costlow N. Cloning and characterization of nine heat-shock-induced mRNAs of Drosophila melanogaster. Gene. 1981 Oct;15(1):67–80. doi: 10.1016/0378-1119(81)90105-0. [DOI] [PubMed] [Google Scholar]
- Luo Y., Hagler J., Shuman S. Discrete functional stages of vaccinia virus early transcription during a single round of RNA synthesis in vitro. J Biol Chem. 1991 Jul 15;266(20):13303–13310. [PubMed] [Google Scholar]
- Nepveu A., Marcu K. B. Intragenic pausing and anti-sense transcription within the murine c-myc locus. EMBO J. 1986 Nov;5(11):2859–2865. doi: 10.1002/j.1460-2075.1986.tb04580.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Brien T., Lis J. T. RNA polymerase II pauses at the 5' end of the transcriptionally induced Drosophila hsp70 gene. Mol Cell Biol. 1991 Oct;11(10):5285–5290. doi: 10.1128/mcb.11.10.5285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riddihough G., Pelham H. R. Activation of the Drosophila hsp27 promoter by heat shock and by ecdysone involves independent and remote regulatory sequences. EMBO J. 1986 Jul;5(7):1653–1658. doi: 10.1002/j.1460-2075.1986.tb04408.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rougvie A. E., Lis J. T. Postinitiation transcriptional control in Drosophila melanogaster. Mol Cell Biol. 1990 Nov;10(11):6041–6045. doi: 10.1128/mcb.10.11.6041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rougvie A. E., Lis J. T. The RNA polymerase II molecule at the 5' end of the uninduced hsp70 gene of D. melanogaster is transcriptionally engaged. Cell. 1988 Sep 9;54(6):795–804. doi: 10.1016/s0092-8674(88)91087-2. [DOI] [PubMed] [Google Scholar]
- Ryner L. C., Manley J. L. Requirements for accurate and efficient mRNA 3' end cleavage and polyadenylation of a simian virus 40 early pre-RNA in vitro. Mol Cell Biol. 1987 Jan;7(1):495–503. doi: 10.1128/mcb.7.1.495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawadogo M., Sentenac A. RNA polymerase B (II) and general transcription factors. Annu Rev Biochem. 1990;59:711–754. doi: 10.1146/annurev.bi.59.070190.003431. [DOI] [PubMed] [Google Scholar]
- Schneider I. Cell lines derived from late embryonic stages of Drosophila melanogaster. J Embryol Exp Morphol. 1972 Apr;27(2):353–365. [PubMed] [Google Scholar]
- Shinshi H., Miwa M., Kato K., Noguchi M., Matsushima T., Sugimura T. A novel phosphodiesterase from cultured tobacco cells. Biochemistry. 1976 May 18;15(10):2185–2190. doi: 10.1021/bi00655a024. [DOI] [PubMed] [Google Scholar]
- Shuman S., Broyles S. S., Moss B. Purification and characterization of a transcription termination factor from vaccinia virions. J Biol Chem. 1987 Sep 5;262(25):12372–12380. [PubMed] [Google Scholar]
- Sisodia S. S., Sollner-Webb B., Cleveland D. W. Specificity of RNA maturation pathways: RNAs transcribed by RNA polymerase III are not substrates for splicing or polyadenylation. Mol Cell Biol. 1987 Oct;7(10):3602–3612. doi: 10.1128/mcb.7.10.3602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southgate R., Ayme A., Voellmy R. Nucleotide sequence analysis of the Drosophila small heat shock gene cluster at locus 67B. J Mol Biol. 1983 Mar 25;165(1):35–57. doi: 10.1016/s0022-2836(83)80241-1. [DOI] [PubMed] [Google Scholar]
- Spencer C. A., Groudine M. Transcription elongation and eukaryotic gene regulation. Oncogene. 1990 Jun;5(6):777–785. [PubMed] [Google Scholar]
- Strobl L. J., Eick D. Hold back of RNA polymerase II at the transcription start site mediates down-regulation of c-myc in vivo. EMBO J. 1992 Sep;11(9):3307–3314. doi: 10.1002/j.1460-2075.1992.tb05409.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas G. H., Elgin S. C. Protein/DNA architecture of the DNase I hypersensitive region of the Drosophila hsp26 promoter. EMBO J. 1988 Jul;7(7):2191–2201. doi: 10.1002/j.1460-2075.1988.tb03058.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Török I., Karch F. Nucleotide sequences of heat shock activated genes in Drosophila melanogaster. I. Sequences in the regions of the 5' and 3' ends of the hsp 70 gene in the hybrid plasmid 56H8. Nucleic Acids Res. 1980 Jul 25;8(14):3105–3123. doi: 10.1093/nar/8.14.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Usheva A., Maldonado E., Goldring A., Lu H., Houbavi C., Reinberg D., Aloni Y. Specific interaction between the nonphosphorylated form of RNA polymerase II and the TATA-binding protein. Cell. 1992 May 29;69(5):871–881. doi: 10.1016/0092-8674(92)90297-p. [DOI] [PubMed] [Google Scholar]
- Venkatesan S., Gershowitz A., Moss B. Modification of the 5' end of mRNA. Association of RNA triphosphatase with the RNA guanylyltransferase-RNA (guanine-7-)methyltransferase complex from vaccinia virus. J Biol Chem. 1980 Feb 10;255(3):903–908. [PubMed] [Google Scholar]
- Weber J., Jelinek W., Darnell J. E., Jr The definition of a large viral transcription unit late in Ad2 infection of HeLa cells: mapping of nascent RNA molecules labeled in isolated nuclei. Cell. 1977 Apr;10(4):611–616. doi: 10.1016/0092-8674(77)90093-9. [DOI] [PubMed] [Google Scholar]
- Wu C. Two protein-binding sites in chromatin implicated in the activation of heat-shock genes. Nature. 1984 May 17;309(5965):229–234. doi: 10.1038/309229a0. [DOI] [PubMed] [Google Scholar]
- Zimarino V., Wu C. Induction of sequence-specific binding of Drosophila heat shock activator protein without protein synthesis. 1987 Jun 25-Jul 1Nature. 327(6124):727–730. doi: 10.1038/327727a0. [DOI] [PubMed] [Google Scholar]