Abstract
Adenoid Cystic Carcinoma (ACC), the second most common malignancy of salivary glands, is a rare tumor with bleak prognosis for which therapeutic targets are unavailable. We used RNA-sequencing (RNA-seq) to analyze low-quality RNA from archival, formaldehyde-fixed, paraffin-embedded samples. In addition to detecting the most common ACC translocation, t(6;9) fusing the MYB proto-oncogene to NFIB, we also detected previously unknown t(8;9) and t(8;14) translocations fusing the MYBL1 gene to the NFIB and RAD51B genes, respectively. RNA-seq provided information about gene fusions, alternative RNA splicing and gene expression signatures. Interestingly, tumors with MYB and MYBL1 translocations displayed similar gene expression profiles, and the combined MYB and MYBL1 expression correlated with outcome, suggesting that the related Myb proteins are interchangeable oncogenic drivers in ACC. Our results provide important details about the biology of ACC and illustrate how archival tissue samples can be used for detailed molecular analyses of rare tumors.
Keywords: MYB-NFIB, MYBL1-NFIB, FFPE, transcriptome
INTRODUCTION
Adenoid Cystic Carcinoma (ACC) is the second most frequent malignancy of the minor and major salivary glands and has poor long-term prognosis (1). ACC tumors display heterogeneous morphology, making them difficult to diagnose and characterize, and have a variable clinical course, in part due to their slow growth and tendency for perineural invasion. Standard therapy, including surgery plus intensive radiation treatment, often requires extensive reconstructive surgery and leads to significant morbidity (2), exacerbated by the fact that many ACC patients develop distant, lethal metastases after 10 or more years (3). Molecular studies of ACC tumors have been complicated by the relative rarity of the tumors, differences in diagnosis and characterization (4), and the lack of bona fide ACC cell lines (5). While the development of xenograft models has overcome some of these obstacles (6), there is an urgent need to develop comprehensive molecular approaches to characterize ACC tumors and to explore their development, progression and heterogeneity.
A large majority of ACC tumors, which occur mainly in salivary glands but also in breast and other tissues (7–9), contain a recurrent t(6;9) translocation, which fuses the MYB proto-oncogene on chromosome 6q to the NFIB gene on chromosome 9p, potentially resulting in the expression of novel MYB-NFIB fusion oncogenes (1, 7, 8, 10, 11). The MYB gene encodes the c-Myb (MYB) transcription factor, which has a highly conserved, N-terminal DNA binding domain, a central domain required for transcriptional activation and conserved C-terminal domains controlling specificity and negative regulation (12, 13). Although overexpression of c-Myb has been implicated in several types of human tumors (14–16), deletion of the C-terminal domains is required to unmask its oncogenic potential in transformation assays (12, 13). In leukemias, alternative RNA splicing has been implicated in the production of truncated versions of c-Myb with altered C-terminal domains and increased oncogenic activity (17, 18). The fusions in ACC tumors could work similarly, leading to over-expression of c-Myb variants with the C-terminal domains deleted (1). In addition to c-Myb, vertebrates also express the related A-Myb (MYBL1) protein, encoded by the MYBL1 gene on chromosome 8q, which has a nearly identical DNA binding domain and a similar overall structure (12). The c-Myb and A-Myb proteins bind and activate the same reporter genes in transfection assays (19), but regulate different endogenous target genes when expressed in human cells (20–22), due to their unique C-terminal specificity domains (12, 23). Recently, recurrent rearrangements involving the MYBL1 gene have been found in gliomas (24, 25), suggesting that it may also be a human oncogene. The NFIB gene also encodes a transcription factor, nuclear factor I/B (NFI-B), a key regulator in hematopoietic and epithelial cells (26, 27), raising the possibility that the MYB-NFIB fusion genes encode novel regulators with the DNA binding domain of c-Myb fused to the specificity and activation domains of NFI-B.
Detailed molecular analyses, including whole exome sequencing (28, 29), have identified a spectrum of cancer-relevant mutations in ACC samples, but have also shown that the translocation t(6;9) is the only highly recurrent genetic alteration in these tumors, suggesting that the product of the MYB-NFIB fusion gene is the key driver mutation in the development of ACC. Therefore, identification of the specifically regulated downstream target genes is critical to better understand ACC and to design novel therapeutic approaches. Recently, detailed microarray analyses of high-quality RNA isolated from a selected panel of frozen ACC tumor samples provided the first information about the transcriptional profiles of ACC (30). However, high-quality RNA for ACC and other rare tumors is very difficult to obtain, especially for older samples for which clinical outcomes are available. Here, we utilized RNA-sequencing (RNA-seq) to analyze low-quality RNA isolated from archival Formaldehyde-Fixed, Paraffin-Embedded (FFPE) pathology samples. Our results show that even low-quality RNA can be used for detailed molecular analyses to identify novel fusion genes, explore differential gene expression profiles and alternative RNA splicing and to uncover complexities in groups of tumor samples. These results provide important information about the biology of ACC and illustrate how archival tissue samples can be used for detailed molecular analyses of rare tumors.
RESULTS
RNA-seq From Poor Quality, Archival, FFPE-Derived RNA
We investigated the usefulness of performing genome-wide RNA-sequencing (RNA-seq) on a set of FFPE-derived ACC tumor samples or normal salivary gland tissue obtained from tissue repositories (Supplementary Table S1). As shown in Figure 1A, the histology of normal salivary gland (top row, N3477 and N4200) is quite different from ACC tumor samples (T115, T399, T440 and T485), which can display varied and heterogeneous morphologies. For RNA-seq, tissue samples were scraped from baked, unstained slides, deparaffinized and RNA was extracted and purified. The archival FFPE tissue blocks yielded RNA that was uniformly of poor quality (RNA Integrity Numbers, RIN < 4), regardless of sample age, tissue type, or tumor stage (Fig 1B, Table S1). Nevertheless, we were able to generate random primed RNA-Seq libraries that were sufficient for whole transcriptome analysis. Libraries were generated from total RNA extracted from FFPE tissue samples from 20 ACC salivary gland tumors and five normal salivary glands, which were then subjected to Ion Proton sequencing. On average, ~16 × 106 mapped reads were obtained for each sample (range 5.9–27 × 106) with an average mean read length of 117 nucleotides (range 97–137; Table S1) and the vast majority of reads mapped to unique sites in the human genome (range 74–98%; Table S1). An average of 9% of the reads (range 7–12%) mapped to exon features, which is somewhat higher than expected for samples without ribosomal RNA depletion.
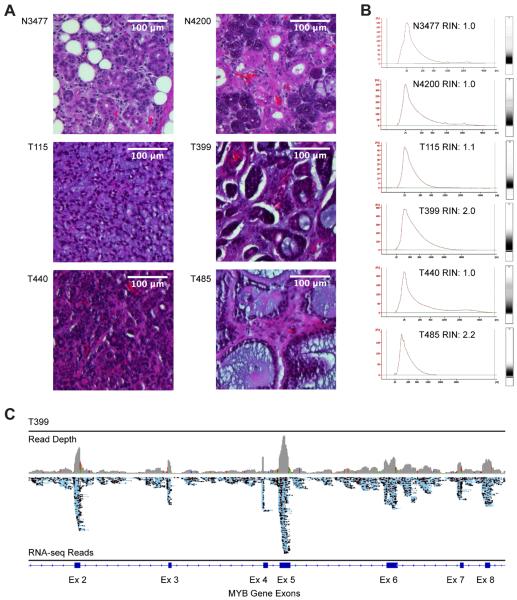
Figure 1. RNA Sequencing From Low Quality RNA.
A. Hematoxylin and eosin stained, Formaldehyde-Fixed, Paraffin-Embedded (FFPE) sections of normal salivary gland samples (top row: N3477 and N4200) and Adenoid Cystic Carcinoma samples (bottom rows: T115, T399, T440 and T485). Scale bars = 100 μm.
B. Bioanalyzer scans of RNA isolated from FFPE samples. X- and Y-axes show RNA size and absorbance, respectively. RNA Integrity Numbers (RIN) are all less than 4, indicating degraded, poor quality RNA.
C. Genome browser view of raw RNA-seq reads from ACC tumor sample T399. The map of the exon 2–8 region of the MYB gene is shown at the bottom, raw reads are in the middle track (blue) and read depth is at the top (gray).
Figure 1C shows a typical Integrative Genomics Viewer (31) depiction of the resulting RNA-seq data, in this case from ACC tumor sample T399, across a region of the MYB gene on chromosome 6. Although most of the reads pile up over the exons, some reads did map to intron regions, probably because the random priming also captures unspliced nuclear RNA. There were relatively few reads mapped to intergenic regions, suggesting there was little contamination of genomic DNA in these libraries (data not shown).
RNA-seq Identifies Novel Chromosomal Translocations in ACC Tumors
A distinctive feature of ACC is the recurrent translocation t(6;9), resulting in fusion of the MYB gene on chromosome 6 to the NFIB gene on chromosome 9, which has previously been observed in 60% to 80% of patient samples (7, 8, 10, 11, 32, 33). We performed a visual analysis of the reads that mapped to the MYB gene in all of the ACC samples. Four samples (T263, T364, T406, T440) were fully transcribed with reads spanning the entire MYB gene and with read pileups corresponding to each exon and the 3'UTR (e.g., T263; Fig 2A). However, eight samples (T013, T042, T074, T087, T192, T349, T399 and T430) displayed clear evidence of an abrupt break in MYB gene transcription, with very few reads in the 3'-half of the gene (e.g., T349; Fig 2A). This uneven pattern of RNA-seq reads is consistent with loss of the 3'-half of the gene due to a translocation. These results also suggest that only the MYB gene involved in the fusion is expressed in these tumors – the other allele appears to be silent. Another eight samples (T060, T072, T115, T229, T411, T441, T452, and T485) demonstrated no, or very low, MYB transcription with very few reads mapping to any MYB exons. However, three of these samples (T072, T115 and T452) displayed abrupt breaks in the transcription of the MYBL1 gene on chromosome 8 (e.g., T452; Fig 2A). Similar to the breaks seen in MYB, these samples displayed substantial expression from the 5' exons but not the 3' exons of the MYBL1 gene. These results provide the first evidence of novel translocations in ACC tumors involving a related, but distinct, Myb-family transcription factor. Thus, RNA-seq data provides a means of identifying genes that are involved in translocations, such as MYB and MYBL1, even when using poor quality RNA extracted from FFPE samples.
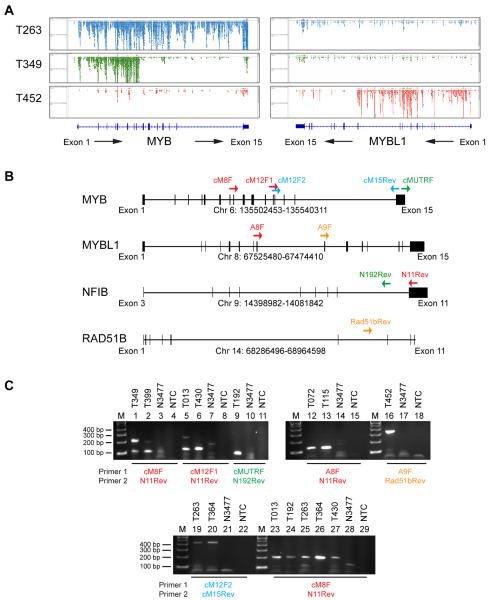
Figure 2. Detection of MYB and MYBL1 Fusions by RNA-seq.
A. Genome browser view of raw RNA-seq reads from the MYB (left) and MYBL1 (right) genes in ACC tumor samples T263 (top row, blue), T349 (second row, green) and T452 (third row, red). The exon maps of the MYB and MYBL1 genes are shown at the bottom, with directions of transcription indicated by arrows.
B. PCR primers used for validating novel MYB and MYBL1 fusions in ACC tumor samples. The maps show the exon structures of the MYB, MYBL1, NFIB and RAD51B genes, with direction of transcription from left to right. Colored arrows show the positions of PCR primers. See Table S3 for details and primer sequences.
C. PCR validation of MYB and MYBL1 fusions. RNA was extracted and PCR amplified as described in the Supplementary Material and Methods section, and products were separated on 2% agarose gels. The top panels verify visually apparent translocations, while the bottom panel demonstrates the presence of cryptic translocations. The presence of multiple bands in lanes 1, 2, 5, 19 and 25 indicate the presence of multiple splice products in several samples. All PCR bands were Topo TA cloned and Sanger sequenced for verification (See Supplementary Table S3).
Chimeric reads were identified at the apparent break points for 8 of the 11 samples with observed MYB or MYBL1 translocations (Supplementary Table S2). In several cases, e.g., T013, chimeric reads appeared to be the products of RNA splicing from an exon in MYB or MYBL1 to a downstream exon in NFIB (Supplementary Table S2), suggesting that the MYB-NFIB or MYBL1-NFIB fusion transcripts are spliced normally and encode fusion proteins. Analysis of the fusion reads in sample T452 identified a novel fusion of MYBL1 to an intron of the RAD51B gene on chromosome 14q23-q24.2. The fusion leads to anti-sense transcription of part of the RAD51B intron, so the A-Myb protein is truncated but no part of the RAD51B protein is included in the predicted fusion protein (for details see Supplementary Table S2).
We used Reverse-Transcription coupled to PCR (RT-PCR) followed by cloning and conventional Sanger sequencing to confirm the novel fusions that were identified by RNA-seq (Figure 2B and Supplementary Table S3). For example, when RNA from tumor sample T349, which had an apparent break in transcription between MYB exons 8 and 9 (Figure 2A) was tested with a forward primer from MYB exon 8 (cM8F) and a reverse primer from NFIB exon 11 (N11Rev), RT-PCR generated products of 220 and 110 bp (Figure 2C, lane 1). These products were not formed when the same primers were used with RNA from normal salivary sample N3477 (lane 3) or in the no template control (NTC, lane 4). Cloning and sequencing confirmed that the PCR products were derived from transcripts correctly spliced from MYB exon 8 either to an alternative exon 10 in NFIB, which was further spliced to NFIB exon 11, to generate the larger PCR product, or directly to NFIB exon 11 to generate the smaller product (for details see Supplementary Table S3). The MYB-NFIB fusion transcripts in tumor samples T013, T192, T349, T399 and T430 were similarly confirmed (Figure 2C, lanes 1–11). MYBL1 fusions to NFIB were confirmed in samples T072 and T115, and a MYBL1-RAD51B fusion was confirmed in sample T452 (Figure 2C, lanes 12–18, Table S3).
To confirm the presence of the chromosome fusions that were inferred from the RNA sequencing data, we also performed specific tests using genomic DNA. PCR amplification across the chromosome breakpoints was performed using RNAase-treated genomic DNA from samples T115, T349 and T452 as well as a control normal salivary gland sample, N3477. Primers designed to amplify across the predicted chromosome breakpoints yielded PCR products of the expected sizes, but only from the specific samples containing those fusions. Cloning and Sanger sequencing of the products confirmed the presence of a MYB-NFIB fusion in sample T349 and the novel MYBL1 fusions with NFIB and RAD51B in samples T115 and T452 (Supplementary Figure S1, Table S3).
Some ACC Tumors Generate Both Full Length and Truncated MYB Forms
Alternative RNA splicing of MYB transcripts has been implicated in the expression of truncated c-Myb variants that may play a role in leukemogenesis (17, 18). We investigated whether the ACC tumor samples that appeared to express full-length MYB transcripts might harbor cryptic chromosomal translocations and/or express truncated variants of Myb proteins via alternative RNA splicing. We used RT-PCR assays followed by cloning and Sanger sequencing to probe two tumors that appeared to have full-length transcription of the MYB gene, T263 and T364. Using a forward primer from MYB exon 12 (cM12F2) and a reverse primer from MYB exon 15 (cM15Rev), we confirmed (Figure 2C, lanes 19–20, Table S3) that both tumors produced full-length MYB transcripts. These transcripts were not detected in the normal salivary gland sample N3477 or in the no template control (lanes 21–22). However, these two tumors were also positive using a forward primer from MYB exon 8 (cM8F) and a reverse primer from NFIB exon 11 (N11Rev, lanes 25–26, Table S3). We conclude that these two ACC tumors are either heterogeneous mixtures of cells that express full length or truncated MYB transcripts or that the individual tumor cells harbor cryptic fusions of the MYB and NFIB genes and produce both full-length, normal c-Myb protein as well as fusion products encoded by a MYB-NFIB fusion gene. Additional experiments with single-cell assays may be needed to resolve these two possibilities.
Employing a similar strategy, we also detected evidence of alternative RNA splicing in the MYB transcripts in ACC tumor samples. Using a forward primer from MYB exon 8 (cM8F) and a reverse primer from NFIB exon 11 (N11Rev) produced two RT-PCR products from tumors T349, T399 (Figure 2C, lanes 1–2) and T263 (lane 25). Cloning and Sanger sequencing confirmed that the larger 220 bp product was produced from the normal exon 8 splice donor site while the smaller 110 bp product was spliced from an alternative splice donor site (Table S3), effectively shortening MYB exon 8 into what has been referred to elsewhere as exon 8s (17, 18). Furthermore, samples T013 and T430, which have confirmed MYB exon 12 to NFIB translocations (Figure 2C, lanes 5–6) and sample T192, which has a confirmed MYB 3'UTR to NFIB translocation (Figure 2C, lane 9), also were positive using the MYB exon 8 (cM8F) and NFIB exon 11 (N11) primer set (Figure 2C, lanes 23–24, 27, Table S3), suggesting that they also produced alternatively-spliced transcripts involving exon 8.
In summary, a majority of the ACC tumor samples analyzed displayed frank MYB-NFIB or MYBL1-NFIB fusions, detected by abrupt breaks in the transcription of the MYB or MYBL1 genes, or displayed cryptic fusions that were detected by PCR (Table S4). The results suggest that nearly all ACC tumors have some type of chromosomal rearrangement involving either the MYB or MYBL1 genes, consistent with the conclusion that activated MYB or MYBL1 genes are the dominant “driver” mutation in these tumors. Some tumors also displayed alternative RNA splicing of MYB transcripts, which could produce truncated, activated forms of c-Myb protein.
MYB / MYBL1 Fusion Proteins Display Increased Transcriptional Activities
Several previous studies have demonstrated that the transcriptional and oncogenic activities of c-Myb increase as C-terminal regulatory domains are mutated or altered through alternative RNA splicing (12). We used the RNA-seq information described above to predict the structures of the transcripts that would be encoded by each tumor, and then the MYB- or MYBL1-fusion proteins that would be produced (Table S2). Plasmid vectors were constructed that express the normal c-Myb or A-Myb proteins, the 9s/10 splice variant of c-Myb, which has a truncated C-terminal domain and has been implicated in leukemogenesis (18), or one of several of the c-Myb or A-Myb fusion proteins predicted to be encoded by ACC tumors (Figure 3A). All of the plasmids encoded c-Myb or A-Myb proteins of the expected sizes when transfected into HEK293T cells (Figure S2), and activated Myb-responsive reporter genes in co-transfection assays, suggesting that all the variants could bind to promoters and activate transcription. This suggests that the fusion proteins are active transcription factors that likely regulate specific genes in the ACC tumors. However, the different proteins displayed distinct activities when tested with two different Myb-responsive reporter constructs. As shown in Figure 3B, the normal c-Myb and A-Myb proteins activated a synthetic promoter engineered to contain 5 high-affinity Myb binding sites (5xMRE-Luc) approximately 100-fold, as did the longest fusion proteins, C12:N8 (c-Myb exon 12 fused to NFIB exon 8) and C12:N10 encoded by tumors T013 and T430. In contrast, the C8:N10, C8Ni10 and A8:N10 proteins encoded by tumors T399, T349 and T115 activated the reporter gene more than 150-fold. The truncated protein encoded by the MYB 9s/10 splice variant activated the promoter about 600-fold and the A9:Ri8 protein encoded by tumor T452 activated it more than 2000-fold. Although some of the fusion proteins were expressed at different levels in the transfected cells (Figure S2), there was no simple correlation between higher steady state protein levels and transcriptional activity in this assay.
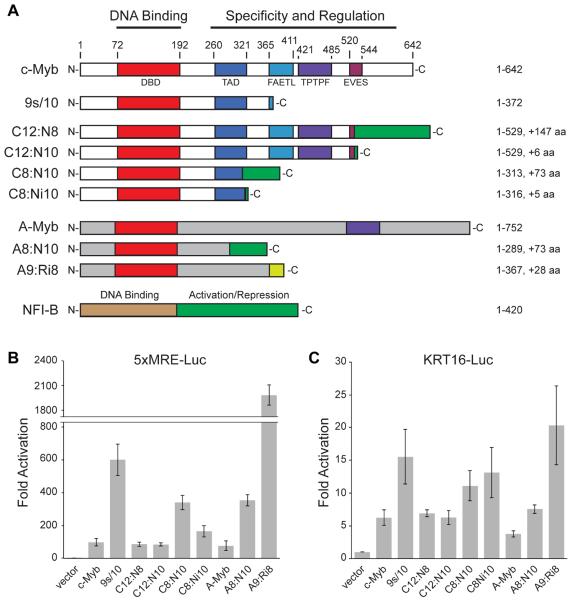
Figure 3. Transcriptional Activities of c-Myb and A-Myb Fusion Proteins.
A. The diagrams show the normal and variant forms of c-Myb (top, white), A-Myb (gray) and NFI-B (bottom, brown/green) proteins, encoded by the MYB, MYBL1 and NFIB genes, respectively, with N-termini at left and C-termini at right. Conserved domains are shaded, including Myb DNA binding domain (red) and other domains involved in transcriptional activation (TAD), transformation (FAETL) and negative regulation (TPTPF, EVES) (for review see (12)). Numbers at top and right indicate amino acid residues. The A-Myb DNA binding (red) and TPTPF (purple) domains are also labeled. NFI-B protein has an N-terminal DNA binding domain (brown) and C-terminal (green) transcriptional activation domain. Also shown are diagrams of the 9s/10 variant produced by alternative RNA splicing in leukemias(18) and the fusion proteins from ACC tumor samples T013 (C12:N8), T430 (C12:N10), T349 (C8:N10), T399 (C8:Ni10) and T115 (A8:N10), which have c-Myb or A-Myb proteins fused to parts of NFI-B, and tumor T452 (A9:Ri8), which has A-Myb fused to a fragment of the RAD51B gene. Letters (C, N, A, R) and numbers refer to the fused proteins (c-Myb, NFI-B, A-Myb, and Rad51b, respectively) and exons (for details see Table S2).
B. and C. Reporter gene assays performed in HEK293T cells with a synthetic minimal promoter engineered to have 5 high-affinity Myb binding sites (panel B, 5xMRE-luc) or with the human KRT16 gene promoter (panel C, KRT16-luc). Bars indicate fold activation compared to an empty vector control. The c-Myb and A-Myb variants that were tested are indicated at bottom. Error bars indicate standard deviation of triplicate assays. The results shown are representative of several independent experiments.
Each of the c-Myb or A-Myb proteins was also tested using a reporter gene derived from the promoter of the Keratin 16 gene (KRT16-luc), a previously identified Myb target gene (20–22). Although all of the c-Myb and A-Myb proteins were able to activate the KRT16 reporter gene, there were some differences in the relative activities compared to the synthetic reporter gene (Figure 3C). For example, the C8:Ni10 protein encoded by tumor T399 was amongst the best activators of the KRT16 promoter and one of the worst activators of the synthetic 5xMRE promoter. These results suggest that the minor differences in the structures of the c-Myb and A-Myb proteins encoded by the ACC tumors could alter their specificities, leading to activation or repression of specific genes that contribute to the development or progression of ACC tumors. To fully understand how the c-Myb and A-Myb proteins function in tumorigenesis, it will be necessary to identify the genes that are activated by the fusion proteins in ACC tumors.
Myb-Expressing ACC Tumors Display Unique Gene Expression Signatures
The RNA-seq results provided a means of analyzing gene expression signatures in the ACC tumors. Briefly, RNA-seq reads that mapped to exons of known genes were used for exon counts that were summed to generate gene-level data. By using only the reads that mapped to exons, we eliminated interference or complications from reads that mapped to introns or intergenic regions. After filtering out genes with little or no expression, 11,278 detectable genes remained for analysis. As shown in Figure 4A, Principal Components Analysis (PCA) clearly separated the five normal salivary gland samples (black) from the 20 ACC tumor samples (red). Although the latter showed significant heterogeneity and scatter in the PCA plot, they did not overlap with the normal samples.
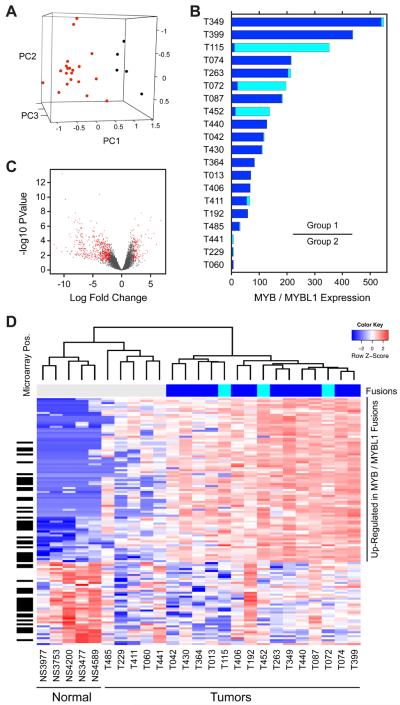
Figure 4. Gene Expression Signatures of ACC Tumors.
A. Principal Components Analysis (PCA) of RNA-seq data from 20 ACC tumors (red) and 5 normal salivary gland samples (black).
B. Gene expression levels of MYB (dark blue) and MYBL1 (cyan) in ACC tumor samples, which are ordered by the sum of MYB + MYBL1. Three samples had very low expression of both genes and were placed in a separate group for differential expression analysis (indicated by bar at right).
C. Volcano plot showing distribution of gene expression changes in ACC tumors samples, with genes showing >2-fold change and adjusted p-value < 0.05 shaded red.
D. Heatmap summarizing differential gene expression in ACC tumors vs. normal samples and illustrating gene expression signatures for ACC tumors. Blue and red shading indicate down- and up-regulation, respectively, as shown in the color key at upper right. Sample names are indicated along the bottom (see Supplementary Table S1 for details). The color bar at top indicates the samples with detected MYB (dark blue) and MYBL1 (cyan) fusions. The black bars at left indicate genes that were reported by others to be differentially expressed in ACC tumors using high quality RNA and microarray assays (30). A larger version of the heatmap with genes labeled is provided in the Supplement.
Because of the biological importance of the MYB and MYBL1 fusions in ACC tumors, we calculated and ranked the samples by the combined expression of MYB and MYBL1 (Figure 4B). The high (Group 1) and low (Group 2) MYB/MYBL1 expression groups were used for differential expression analysis compared to the normal samples using the Bioconductor (34, 35) package edgeR (36–38) to develop a gene-wise negative binomial log-linear model, which identified a total of 1812 differentially expressed genes, as shown in the Volcano Plot (Figure 4C). Requiring a 2-fold change and a false discovery rate-adjusted p-value cutoff of 0.05 yielded a total of 1494 differentially-expressed genes comparing Group 1 to normal, and 219 differentially expressed genes comparing Group 2 to normal (Supplementary Tables S5, S6).
The results of the differential expression analysis are summarized in the heatmap in Figure 4D, which shows the gene expression signatures of the most differentially expressed genes in these samples (a larger version with the genes labeled is in the supplementary materials, Figure S3). The color bar at the top of the heatmap indicates the samples with MYB and MYBL1 fusions (dark blue and cyan, respectively). Perhaps the most striking feature of the analysis is that samples with MYBL1 fusions did not form their own cluster, but were instead scattered throughout the group of samples with MYB fusions. Thus, it appears that, at least in terms of gene expression signatures, the MYB and MYBL1 fusions are interchangeable. This suggests that the biology and gene expression patterns of ACC tumors are driven by the presence of a rearranged and activated transcription factor in the Myb family, either MYB or MYBL1, but not specific to either one.
Along the left edge of the heatmap in Figure 4D are black marks indicating the genes that were recently reported as differentially expressed when high quality ACC tumor RNA samples were analyzed using microarrays (30). Of the 127 genes displayed in the heatmap, 52 were also identified in the published microarray analysis (30). Gene Set Enrichment Analysis (39) indicated that the gene list identified previously was highly correlated to the list identified here (FDR=0.077, see supplementary data, Figure S4). The marked overlap suggests that the analysis of poor-quality RNA by RNA-seq is, at the least, a viable alternative to microarrays, especially when high-quality RNA is not available. In addition, the unbiased RNA-seq approach may be identifying additional genes that were missed in the microarray assays.
The heatmap is dominated by a large group of up-regulated genes (shaded red, in the upper right corner of the heatmap and labeled at right), which contains several genes involved in controlling differentiation, such as transcription factors EN1 (engrailed homeobox 1), POU3F2 and SOX11 and genes that have been identified previously as direct targets of c-Myb or A-Myb regulation, including CCNB1, which c-Myb binds and regulates only in the G2/M phase of the cell cycle (40). Gene Ontology (41, 42) analysis of the genes in that block, which are up-regulated in the ACC tumors, shows enrichment of genes involved in several biological processes linked to cell growth, cell cycle and cell division (Table S7). There are also genes that could be used in targeted immunotherapies for ACC tumors, such as PRAME, which is highly up-regulated in ACC tumors but not expressed in the normal tissues. The PRAME gene is also up-regulated in other tumors and is under testing as a target for immunotherapeutic approaches (43, 44). Notably absent is KIT, which has been implicated as a biomarker in ACC tumors (45, 46), which was expressed in both normal and ACC tumor samples but was not sufficiently differentially expressed to be included in our gene lists.
We also analyzed our expression data for genes whose expression either positively or negatively correlated with combined MYB and MYBL1 expression (Figure S5). Positively correlated genes included important regulators EN1, PRAME, SOX4, SOX11, SMO and CDK6. Gene Ontology analysis of the genes most and least correlated with MYB and MYBL1 expression identified the biological processes of cell differentiation in both epithelial and neuronal cells and transcriptional regulation (Table S8), consistent with a role of Myb proteins in regulation of differentiation and proliferation (18).
Combined Expression of MYB and MYBL1 Correlates with Poor Outcome
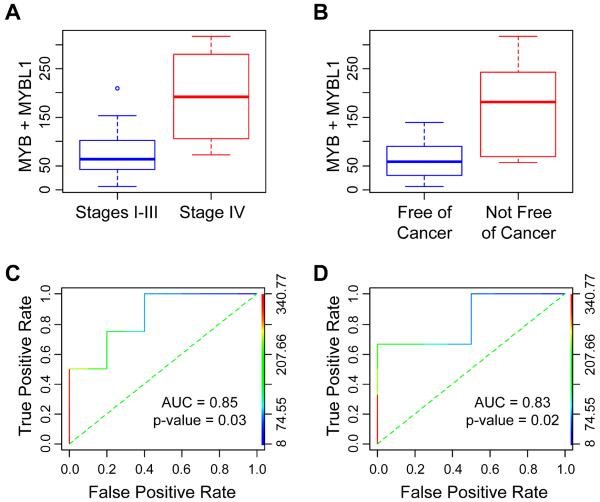
High expression of c-Myb protein, measured by immuno-staining in tissue samples, has been linked to worse outcome in ACC tumors (1). Although clinical outcome information was only available for 14 of the samples that we analyzed, we tested whether the combined expression of MYB plus MYBL1, measured by RNA-seq, would also correlate with poor outcome. The boxplots in Figure 5 (panels A and B) show that the combined MYB and MYBL1 expression is significantly higher in patients diagnosed as Stage IV compared to patients diagnosed as Stages I – III (panel A) and also in patients described as not free of cancer compared to free of cancer (panel B). The Receiver Operating Characteristic (ROC) curves (Figure 5, Panels C and D) show the combined expression as the predictors of stage and outcome. Despite the small sample size, the areas under the ROC curves were 0.85 and 0.83 and p-values of the one-sided Mann-Whitney test were .03 and .02, indicating that the combined MYB and MYBL1 expression was predictive of tumor stage (Figure 5, panel C) and outcome (Figure 5, panel D). Although these results will need to be confirmed in larger cohorts, the results suggest that the RNA-seq approach, even from poor-quality samples, can provide important information about gene expression patterns in ACC tumors that correlate with clinical outcome.
Figure 5. MYB and MYBL1 Expression Correlates with Tumor Stage and Outcome.
Boxplots (panels A and B) indicate that combined MYB and MYBL1 expression is significantly higher in patients with tumor Stage IV than in those with Stages I – III (panel A, red and blue, respectively) and is also significantly higher in patients who are not free of cancer than in those who are (panel B, red and blue, respectively). P-values of the one-sided Mann-Whitney test are .03 and .02. ROC analysis indicates that the combined MYB and MYBL1 expression is predictive of tumor stage (panel C) and outcome (panel D). The ROC accuracies, i.e., the areas under the ROC curves, are 0.85 and 0.83, respectively.
DISCUSSION
Salivary gland ACC is a tumor type with known driver mutations but little molecular information, such as gene expression patterns, for classifying the tumors or for predicting clinical outcome. By utilizing sensitive RNA-seq methods with RNA isolated from archived pathology samples of ACC tumors, we were able to identify new translocation breakpoints and fusion gene products and to classify and test the activities of truncated c-Myb and A-Myb fusion proteins produced by ACC tumors. Our results show that both MYB and MYBL1 fusions are likely to be driver mutations in ACC. Interestingly, the truncated fusion proteins produced in the tumors lack most of the C-terminal domains of the c-Myb and A-Myb proteins normally encoded by these genes, which are the regions of the proteins that give them unique activities and that enable them to activate different sets of human genes (19–22). These results fit with models of Myb protein function, in which the C-terminal domains are thought to form protein-protein interactions that direct the transcription factor to specific subsets of target genes (12, 13, 23). In ACC tumors, it seems likely that the translocations lead to the expression of fusion transcripts and truncated c-Myb or A-Myb proteins that have lost specificity and gained oncogenic activity.
The RNA-seq results, and subsequent validation by PCR and conventional Sanger sequencing (Figure 2), confirmed that nearly all of the ACC tumors analyzed expressed correctly spliced MYB or MYBL1 fusion transcripts that were likely to encode truncated c-Myb or A-Myb proteins. As we were preparing this manuscript for submission, we learned that another laboratory independently identified MYBL1 fusions in ACC tumors (47). These results suggest that ACC tumors may be similar to some gliomas, which also harbor frequent rearrangements in the MYB and MYBL1 genes (24, 25). The fusion products included widely different amounts of NFI-B protein at their C-termini, ranging from 0 to 147 amino acids, and one fusion had 28 amino acids derived from anti-sense transcription of the RAD51B gene. Thus, it is difficult to conclude that the oncogenic activities of these proteins stem from anything contributed by specific residues donated by NFI-B. In some samples we detected multiple variants of MYB transcripts expressed within a single tumor. This is reminiscent of Acute Lymphocytic Leukemias, which display elevated levels of MYB alternative RNA splicing and produce a number of transcript variants, some of which correlate with poor prognosis (18). Thus, mutations or activated signaling pathways that affect alternative RNA splicing could be important for tumor development, if the result is the production of oncogenic isoforms of proteins like c-Myb or A-Myb.
The gene expression studies identified several subgroups of ACC tumors. Interestingly, the tumors with MYB or MYBL1 fusions did not segregate into their own clusters or display unique gene expression signatures. Instead, the truncated MYB and MYBL1 gene products appear to be equally oncogenic and interchangeable. This fits with previous studies showing that the A-Myb and c-Myb DNA binding domains could be swapped without significantly affecting the genes that the proteins activated (20, 22). We identified a large number of genes that were similarly up-regulated in the tumors with MYB or MYBL1 fusions. Some of these genes are likely to be direct targets that are regulated by the truncated c-Myb or A-Myb proteins. We also identified several tumors that formed a distinct cluster, displayed a significantly different gene expression pattern, and that did not contain evidence of MYB or MYBL1 fusions. These “Group 2” tumors may have different, as yet unidentified driver mutations, or may be slightly different tumor types. Future studies will need to analyze larger sets of tumors with associated clinical outcome data in order to correlate these different gene expression patterns with relevant prognostic features.
These studies show that poor-quality RNA isolated from archived pathology material can be used in RNA-seq experiments to obtain important information about rare tumors like ACC. Our results were validated by others who found many of the same genes using microarrays to study gene expression patterns in high-quality RNA isolated from frozen ACC tumors (30). This opens the door for future studies in ACC and other tumor types for which properly frozen material or high quality RNA is difficult to obtain.
Finally, the RNA-seq results raise several questions for future studies. Other than the recurrent translocations, there is very little information about underlying genomic changes in ACC. It seems likely that the development of ACC tumors require a rearranged MYB or MYBL1 oncogene as well as additional cooperating mutations. Although two studies have previously addressed this through exome sequencing (28, 29), no candidates for recurrent mutations that might cooperate with activated Myb proteins were identified. This raises the question of whether mutations in regulatory regions or non-coding RNAs could play a role. For example, the recurrent translocations observed in ACC tumors could bring enhancers to the MYB or MYBL1 genes to increase their expression. Mutations in non-coding regions have been shown to play a role in other tumors that involve activated or over-expressed Myb proteins (48), so similar mechanisms could also be at work in ACC tumors. Additional studies, including an analysis of epigenetic events and non-coding changes, will likely be necessary to answer these questions.
METHODS
Human Salivary Gland FFPE Samples
Twenty de-identified salivary gland adenoid cystic carcinoma FFPE tumor samples were obtained from the Salivary Gland Tumor Biorepository (MD Anderson Cancer Center, Houston, TX, Table S1). Five de-identified normal salivary gland FFPE samples were obtained from the Human Tissue Repository and Cell Analysis Shared Resource at the University of New Mexico (Department of Pathology and UNM Cancer Center, Albuquerque, NM). All samples were collected with informed consent of the donors and studies were conducted in accordance with the principles of the Declaration of Helsinki. All studies were performed under Institutional Review Board-approved protocols.
RNA Isolation and Sequencing
Total RNA was isolated from single 10 μM slide-mounted FFPE sections using the RNeasy FFPE kit from Qiagen. After isolation, total RNA was mixed with ERCC Spike-In Mix 1 control RNA (Life Technologies) and converted to cDNA using the SMARTer Universal Low Input RNA Kit for Sequencing (Clontech), which uses random primers for the conversion. The Ion Plus Fragment Library kit (Life Technologies) was used to add barcodes, amplify and size-select the final library. Four individually barcoded samples were sequenced together on Ion Proton P1v2 chips by the Analytical and Translational Genomics Shared Resource at the University of New Mexico Cancer Center. See Supplementary Methods for details. RNA sequencing data is available for download from the NCBI Sequence Read Archive using SRA study accession number SRP059557. Nine of the ACC samples were analyzed twice to confirm the reproducibility of the assays (data not shown).
Data Analysis
RNA-seq reads for each library were mapped to the human genome (GRCh37; hg19) using TMAP (v4.0.6). Gene counts were calculated using HT-Seq with the FeatureCounter plugin (v1.0.6, Mode = union) against a RefSeq hg19 exons BED file, with overlapping exons trimmed to remove duplicates. Exon counts for each gene were summed to produce gene counts and normalized to ERCC standard curves determined using the ERCC Analysis plugin (v4.2-r87667). Normalized data was further analyzed using R packages edgeR, DESeq, plot3D, topGO and pathview (49, 50).
Verification of Translocations
Translocations were verified by RT-PCR amplification of RNA isolated from FFPE slices. Fresh RNA was isolated from a new FFPE slice using the same method described above. The SuperScript III One-Step RT-PCR System (Life Technologies) was used to both create and amplify cDNA using gene specific primers (Supplementary Table S3). PCR fragments were then cloned (Topo TA for sequencing, Life Technologies) and insert-positive plasmids were analyzed by Sanger sequencing (Eurofins) to confirm translocations.
Cell Culture and Luciferase Assays
HEK293TN Human Kidney Producer cells (HEK293TN; purchased from System Biosciences in July 2013 and not further authenticated) were cultured at 37 °C in 5% CO2 in Dulbecco's Modified Eagle's Medium (DMEM; ATCC) supplemented with 5% (v/v) fetal bovine serum (Life Technologies), 5% (v/v) newborn calf serum (Rocky Mountain Biologicals, Inc.), and 1% Antibiotic-Antimycotic (Life Technologies). For transfections, cells were seeded in 24 well plates with approximately 4–6×104 cells per well. After 24 hours of growth, cells were transiently co-transfected with 50 ng of luciferase reporter plasmid and 5 ng of activator plasmid (MYB or MYBL1 fusions cloned into pcDNA3.0). Transfections were performed in duplicate using the TransIT-2020 transfection (Mirus) reagent according to manufacturer instructions. Cells were harvested and firefly luciferase activity was measured after 48 hours using the Luciferase Assay System (Promega). Background subtracted data was normalized to cells transfected with empty pcDNA3.0 and the reporter plasmid. Reporter gene assays were performed in triplicate.
Supplementary Material
STATEMENT OF SIGNIFICANCE.
Using RNA-seq to perform whole-transcriptome analysis of archival ACC tumor samples, we identified novel, recurrent gene fusions, detected alternative RNA splicing and established gene expression signatures that provide detailed information about the biology of ACC tumors.
ACKNOWLEDGMENTS
The authors acknowledge the outstanding technical support from Jennifer Woods, Maggie Cyphery, Jamie Padilla, Jason Byars and Gavin Pickett. We are also grateful to the support from Adenoid Cystic Carcinoma Research Foundation, to the Salivary Gland Tumor Biorepository and Dr. Adel El-Naggar at the University of Texas MD Anderson Cancer Center.
GRANT SUPPORT This work was supported by NIH grant R01DE023222 (to SAN). Some experiments used the facilities or services provided by the Analytical and Translational Genomics Shared Resource, the Fluorescence Microscopy and Cell Imaging Shared Resource and the Human Tissue Repository and Cell Analysis Shared Resource, which are supported by the State of New Mexico and the UNM Cancer Center P30CA118100.
Financial Support, By Authors: K.J. Brayer, C.A. Frerich: NIH grant 5R01DE023222
H. Kang, S.A. Ness: NIH grants 5R01CA170250, 5R01DE023222, 5P30CA118100
Footnotes
Disclosure of Potential Conflicts of Interest: No potential conflicts of interest for any author.
REFERENCES
- 1.Mitani Y, Rao PH, Futreal PA, Roberts DB, Stephens PJ, Zhao YJ, et al. Novel chromosomal rearrangements and break points at the t(6;9) in salivary adenoid cystic carcinoma: association with MYB-NFIB chimeric fusion, MYB expression, and clinical outcome. Clin Cancer Res. 2011;17:7003–14. doi: 10.1158/1078-0432.CCR-11-1870. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fordice J, Kershaw C, El-Naggar A, Goepfert H. Adenoid cystic carcinoma of the head and neck: predictors of morbidity and mortality. Arch Otolaryngol Head Neck Surg. 1999;125:149–52. doi: 10.1001/archotol.125.2.149. [DOI] [PubMed] [Google Scholar]
- 3.Hunt JL. An update on molecular diagnostics of squamous and salivary gland tumors of the head and neck. Arch Pathol Lab Med. 2011;135:602–9. doi: 10.5858/2010-0655-RAIR.1. [DOI] [PubMed] [Google Scholar]
- 4.Coca-Pelaz A, Rodrigo JP, Bradley PJ, Vander Poorten V, Triantafyllou A, Hunt JL, et al. Adenoid cystic carcinoma of the head and neck - An update. Oral Oncol. 2015;51:652–61. doi: 10.1016/j.oraloncology.2015.04.005. [DOI] [PubMed] [Google Scholar]
- 5.Zhao M, Sano D, Pickering CR, Jasser SA, Henderson YC, Clayman GL, et al. Assembly and initial characterization of a panel of 85 genomically validated cell lines from diverse head and neck tumor sites. Clin Cancer Res. 2011;17:7248–64. doi: 10.1158/1078-0432.CCR-11-0690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Moskaluk CA, Baras AS, Mancuso SA, Fan H, Davidson RJ, Dirks DC, et al. Development and characterization of xenograft model systems for adenoid cystic carcinoma. Lab Invest. 2011;91:1480–90. doi: 10.1038/labinvest.2011.105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Brill LB, Kanner WA, Fehr A, Andrén Y, Moskaluk CA, Löning T, et al. Analysis of MYB expression and MYB-NFIB gene fusions in adenoid cystic carcinoma and other salivary neoplasms. Mod Pathol. 2011;24:1169–76. doi: 10.1038/modpathol.2011.86. [DOI] [PubMed] [Google Scholar]
- 8.Persson M, Andrén Y, Mark J, Horlings HM, Persson F, Stenman G. Recurrent fusion of MYB and NFIB transcription factor genes in carcinomas of the breast and head and neck. Proc Natl Acad Sci U S A. 2009;106:18740–4. doi: 10.1073/pnas.0909114106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Roden AC, Greipp PT, Knutson DL, Kloft-Nelson SM, Jenkins SM, Marks RS, et al. Histopathologic and Cytogenetic Features of Pulmonary Adenoid Cystic Carcinoma. J Thorac Oncol. 2015;10:1570–75. doi: 10.1097/JTO.0000000000000656. [DOI] [PubMed] [Google Scholar]
- 10.Fehr A, Kovács A, Löning T, Frierson H, van den Oord J, Stenman G. The MYB-NFIB gene fusion-a novel genetic link between adenoid cystic carcinoma and dermal cylindroma. J Pathol. 2011;224:322–7. doi: 10.1002/path.2909. [DOI] [PubMed] [Google Scholar]
- 11.Mitani Y, Li J, Rao PH, Zhao YJ, Bell D, Lippman SM, et al. Comprehensive analysis of the MYB-NFIB gene fusion in salivary adenoid cystic carcinoma: Incidence, variability, and clinicopathologic significance. Clin Cancer Res. 2010;16:4722–31. doi: 10.1158/1078-0432.CCR-10-0463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.George OL, Ness SA. Situational awareness: regulation of the myb transcription factor in differentiation, the cell cycle and oncogenesis. Cancers (Basel) 2014;6:2049–71. doi: 10.3390/cancers6042049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhou Y, Ness SA. Myb proteins: angels and demons in normal and transformed cells. Front Biosci. 2011;16:1109–31. doi: 10.2741/3738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Biroccio A, Benassi B, D'Agnano I, D'Angelo C, Buglioni S, Mottolese M, et al. c-Myb and Bcl-x overexpression predicts poor prognosis in colorectal cancer: clinical and experimental findings. Am J Pathol. 2001;158:1289–99. doi: 10.1016/S0002-9440(10)64080-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Clappier E, Cuccuini W, Kalota A, Crinquette A, Cayuela JM, Dik WA, et al. The C-MYB locus is involved in chromosomal translocation and genomic duplications in human T-cell acute leukemia (T-ALL), the translocation defining a new T-ALL subtype in very young children. Blood. 2007;110:1251–61. doi: 10.1182/blood-2006-12-064683. [DOI] [PubMed] [Google Scholar]
- 16.Ramsay RG, Thompson MA, Hayman JA, Reid G, Gonda TJ, Whitehead RH. Myb expression is higher in malignant human colonic carcinoma and premalignant adenomatous polyps than in normal mucosa. Cell Growth Differ. 1992;3:723–30. [PubMed] [Google Scholar]
- 17.O'Rourke JP, Ness SA. Alternative RNA splicing produces multiple forms of c-Myb with unique transcriptional activities. Mol Cell Biol. 2008;28:2091–101. doi: 10.1128/MCB.01870-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhou YE, O'Rourke JP, Edwards JS, Ness SA. Single molecule analysis of c-myb alternative splicing reveals novel classifiers for precursor B-ALL. PLoS One. 2011;6:e22880. doi: 10.1371/journal.pone.0022880. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rushton JJ, Ness SA. The conserved DNA binding domain mediates similar regulatory interactions for A-Myb, B-Myb, and c-Myb transcription factors. Blood Cells Mol Dis. 2001;27:459–63. doi: 10.1006/bcmd.2001.0405. [DOI] [PubMed] [Google Scholar]
- 20.Lei W, Rushton JJ, Davis LM, Liu F, Ness SA. Positive and negative determinants of target gene specificity in myb transcription factors. J Biol Chem. 2004;279:29519–27. doi: 10.1074/jbc.M403133200. [DOI] [PubMed] [Google Scholar]
- 21.Liu F, Lei W, O'Rourke JP, Ness SA. Oncogenic mutations cause dramatic, qualitative changes in the transcriptional activity of c-Myb. Oncogene. 2006;25:795–805. doi: 10.1038/sj.onc.1209105. [DOI] [PubMed] [Google Scholar]
- 22.Rushton JJ, Davis LM, Lei W, Mo X, Leutz A, Ness SA. Distinct changes in gene expression induced by A-Myb, B-Myb and c-Myb proteins. Oncogene. 2003;22:308–13. doi: 10.1038/sj.onc.1206131. [DOI] [PubMed] [Google Scholar]
- 23.Ness SA. Myb protein specificity: evidence of a context-specific transcription factor code. Blood Cells Mol Dis. 2003;31:192–200. doi: 10.1016/s1079-9796(03)00151-7. [DOI] [PubMed] [Google Scholar]
- 24.Ramkissoon LA, Horowitz PM, Craig JM, Ramkissoon SH, Rich BE, Schumacher SE, et al. Genomic analysis of diffuse pediatric low-grade gliomas identifies recurrent oncogenic truncating rearrangements in the transcription factor MYBL1. Proc Natl Acad Sci U S A. 2013;110:8188–93. doi: 10.1073/pnas.1300252110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhang J, Wu G, Miller CP, Tatevossian RG, Dalton JD, Tang B, et al. Whole-genome sequencing identifies genetic alterations in pediatric low-grade gliomas. Nat Genet. 2013;45:602–12. doi: 10.1038/ng.2611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chen L, Kostadima M, Martens JH, Canu G, Garcia SP, Turro E, et al. Transcriptional diversity during lineage commitment of human blood progenitors. Science. 2014;345:1251033. doi: 10.1126/science.1251033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Grabowska MM, Elliott AD, DeGraff DJ, Anderson PD, Anumanthan G, Yamashita H, et al. NFI transcription factors interact with FOXA1 to regulate prostate-specific gene expression. Mol Endocrinol. 2014;28:949–64. doi: 10.1210/me.2013-1213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ho AS, Kannan K, Roy DM, Morris LG, Ganly I, Katabi N, et al. The mutational landscape of adenoid cystic carcinoma. Nat Genet. 2013;45:791–8. doi: 10.1038/ng.2643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stephens PJ, Davies HR, Mitani Y, Van Loo P, Shlien A, Tarpey PS, et al. Whole exome sequencing of adenoid cystic carcinoma. J Clin Invest. 2013;123:2965–8. doi: 10.1172/JCI67201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gao R, Cao C, Zhang M, Lopez MC, Yan Y, Chen Z, et al. A unifying gene signature for adenoid cystic cancer identifies parallel MYB-dependent and MYB-independent therapeutic targets. Oncotarget. 2014;5:12528–42. doi: 10.18632/oncotarget.2985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Thorvaldsdottir H, Robinson JT, Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 2013;14:178–92. doi: 10.1093/bib/bbs017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Stenman G, Andersson MK, Andrén Y. New tricks from an old oncogene: gene fusion and copy number alterations of MYB in human cancer. Cell Cycle. 2010;9:2986–95. doi: 10.4161/cc.9.15.12515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.West RB, Kong C, Clarke N, Gilks T, Lipsick JS, Cao H, et al. MYB expression and translocation in adenoid cystic carcinomas and other salivary gland tumors with clinicopathologic correlation. Am J Surg Pathol. 2011;35:92–9. doi: 10.1097/PAS.0b013e3182002777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5:R80. doi: 10.1186/gb-2004-5-10-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Huber W, Carey VJ, Gentleman R, Anders S, Carlson M, Carvalho BS, et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat Methods. 2015;12:115–21. doi: 10.1038/nmeth.3252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.McCarthy DJ, Chen Y, Smyth GK. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 2012;40:4288–97. doi: 10.1093/nar/gks042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Robinson MD, Smyth GK. Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics. 2008;9:321–32. doi: 10.1093/biostatistics/kxm030. [DOI] [PubMed] [Google Scholar]
- 39.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Quintana AM, Zhou YE, Pena JJ, O'Rourke JP, Ness SA. Dramatic repositioning of c-Myb to different promoters during the cell cycle observed by combining cell sorting with chromatin immunoprecipitation. PLoS One. 2011;6:e17362. doi: 10.1371/journal.pone.0017362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Gene Ontology C. Gene Ontology Consortium: going forward. Nucleic Acids Res. 2015;43:D1049–56. doi: 10.1093/nar/gku1179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Babiak A, Steinhauser M, Gotz M, Herbst C, Dohner H, Greiner J. Frequent T cell responses against immunogenic targets in lung cancer patients for targeted immunotherapy. Oncol Rep. 2014;31:384–90. doi: 10.3892/or.2013.2804. [DOI] [PubMed] [Google Scholar]
- 44.Yao J, Caballero OL, Yung WK, Weinstein JN, Riggins GJ, Strausberg RL, et al. Tumor subtype-specific cancer-testis antigens as potential biomarkers and immunotherapeutic targets for cancers. Cancer Immunol Res. 2014;2:371–9. doi: 10.1158/2326-6066.CIR-13-0088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Bell D, Hanna EY. Salivary gland cancers: biology and molecular targets for therapy. Curr Oncol Rep. 2012;14:166–74. doi: 10.1007/s11912-012-0220-5. [DOI] [PubMed] [Google Scholar]
- 46.Tang YL, Fan YL, Jiang J, Li KD, Zheng M, Chen W, et al. C-kit induces epithelial-mesenchymal transition and contributes to salivary adenoid cystic cancer progression. Oncotarget. 2014;5:1491–501. doi: 10.18632/oncotarget.1606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.El-Naggar A, et al. Unknown. Clin Cancer Res. 2015 [Google Scholar]
- 48.Mansour MR, Abraham BJ, Anders L, Berezovskaya A, Gutierrez A, Durbin AD, et al. Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science. 2014;346:1373–7. doi: 10.1126/science.1259037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Alexa A, Rahnenfuhrer J, Lengauer T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics. 2006;22:1600–7. doi: 10.1093/bioinformatics/btl140. [DOI] [PubMed] [Google Scholar]
- 50.Luo W, Brouwer C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics. 2013;29:1830–1. doi: 10.1093/bioinformatics/btt285. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.