Abstract
Host and viral proteinases are believed to be required for the production of at least nine hepatitis C virus (HCV)-specific polyprotein cleavage products. Although several cleavages appear to be catalyzed by host signal peptidase or the HCV NS3 serine proteinase, the enzyme responsible for cleavage at the 2/3 site has not been identified. In this report, we have defined the 2/3 cleavage site and obtained evidence which suggests that this cleavage is mediated by a second HCV-encoded proteinase, located between aa 827 and 1207. This region encompasses the C-terminal portion of the 23-kDa NS2 protein, the 2/3 cleavage site, and the serine proteinase domain of NS3. Efficient processing at the 2/3 site was observed in mammalian cells, Escherichia coli, and in plant or animal cell-free translation systems in the absence of microsomal membranes. Cleavage at the 2/3 site was abolished by alanine substitutions for NS2 residues His-952 or Cys-993 but was unaffected by several other substitution mutations, including those that inactivate NS3 serine proteinase function. Mutations abolishing cleavage at the 2/3 site did not block cleavage at other sites in the HCV polyprotein. Cotransfection experiments indicate that the 2/3 site can be cleaved in trans, which should facilitate purification and further characterization of this enzyme.
Full text
PDF
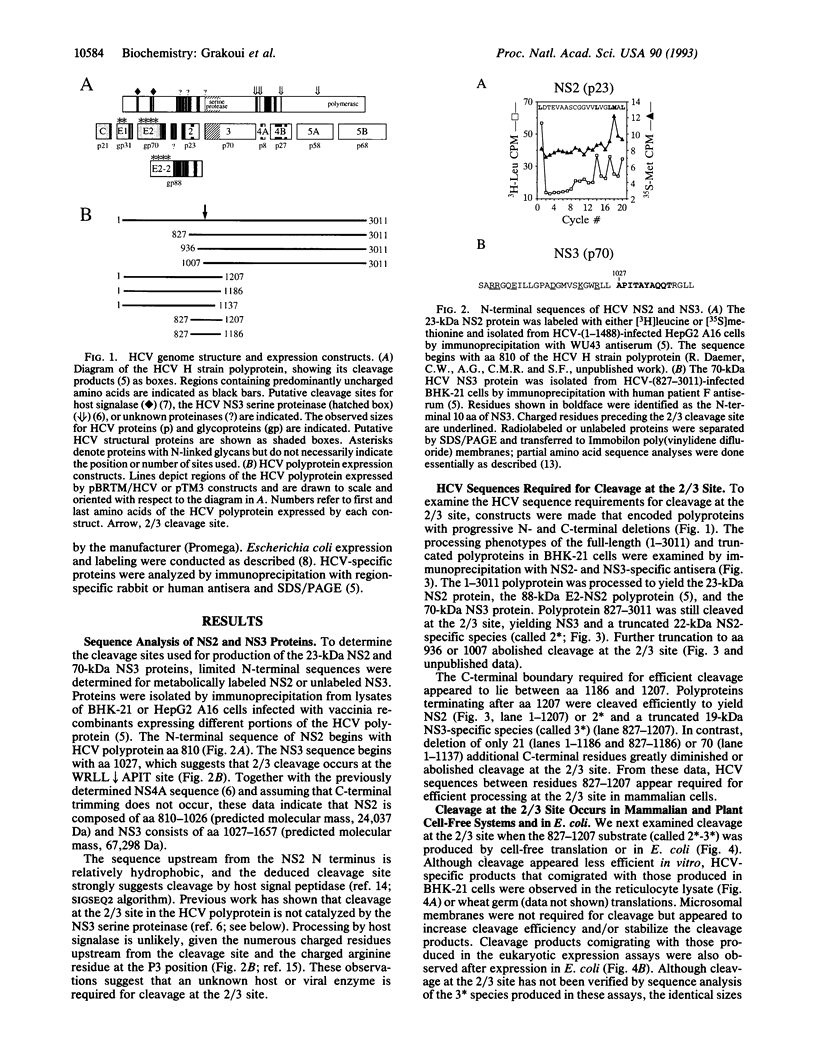


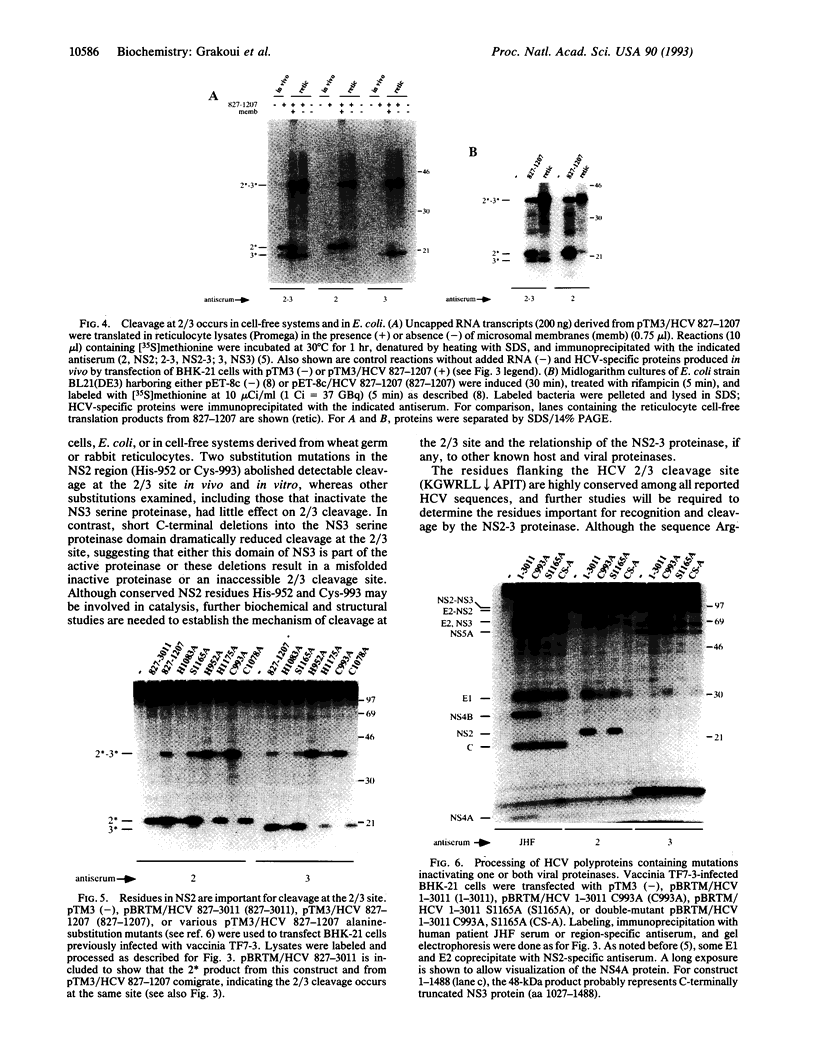

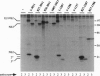
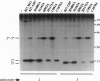
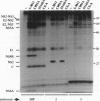
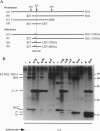
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arias C. F., Preugschat F., Strauss J. H. Dengue 2 virus NS2B and NS3 form a stable complex that can cleave NS3 within the helicase domain. Virology. 1993 Apr;193(2):888–899. doi: 10.1006/viro.1993.1198. [DOI] [PubMed] [Google Scholar]
- Cha T. A., Beall E., Irvine B., Kolberg J., Chien D., Kuo G., Urdea M. S. At least five related, but distinct, hepatitis C viral genotypes exist. Proc Natl Acad Sci U S A. 1992 Aug 1;89(15):7144–7148. doi: 10.1073/pnas.89.15.7144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers T. J., Grakoui A., Rice C. M. Processing of the yellow fever virus nonstructural polyprotein: a catalytically active NS3 proteinase domain and NS2B are required for cleavages at dibasic sites. J Virol. 1991 Nov;65(11):6042–6050. doi: 10.1128/jvi.65.11.6042-6050.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers T. J., McCourt D. W., Rice C. M. Yellow fever virus proteins NS2A, NS2B, and NS4B: identification and partial N-terminal amino acid sequence analysis. Virology. 1989 Mar;169(1):100–109. doi: 10.1016/0042-6822(89)90045-7. [DOI] [PubMed] [Google Scholar]
- Chambers T. J., Weir R. C., Grakoui A., McCourt D. W., Bazan J. F., Fletterick R. J., Rice C. M. Evidence that the N-terminal domain of nonstructural protein NS3 from yellow fever virus is a serine protease responsible for site-specific cleavages in the viral polyprotein. Proc Natl Acad Sci U S A. 1990 Nov;87(22):8898–8902. doi: 10.1073/pnas.87.22.8898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collett M. S. Molecular genetics of pestiviruses. Comp Immunol Microbiol Infect Dis. 1992 Jul;15(3):145–154. doi: 10.1016/0147-9571(92)90087-8. [DOI] [PubMed] [Google Scholar]
- Falgout B., Pethel M., Zhang Y. M., Lai C. J. Both nonstructural proteins NS2B and NS3 are required for the proteolytic processing of dengue virus nonstructural proteins. J Virol. 1991 May;65(5):2467–2475. doi: 10.1128/jvi.65.5.2467-2475.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Folz R. J., Gordon J. I. Computer-assisted predictions of signal peptidase processing sites. Biochem Biophys Res Commun. 1987 Jul 31;146(2):870–877. doi: 10.1016/0006-291x(87)90611-5. [DOI] [PubMed] [Google Scholar]
- Gorbalenya A. E., Koonin E. V., Donchenko A. P., Blinov V. M. Two related superfamilies of putative helicases involved in replication, recombination, repair and expression of DNA and RNA genomes. Nucleic Acids Res. 1989 Jun 26;17(12):4713–4730. doi: 10.1093/nar/17.12.4713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grakoui A., McCourt D. W., Wychowski C., Feinstone S. M., Rice C. M. Characterization of the hepatitis C virus-encoded serine proteinase: determination of proteinase-dependent polyprotein cleavage sites. J Virol. 1993 May;67(5):2832–2843. doi: 10.1128/jvi.67.5.2832-2843.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grakoui A., Wychowski C., Lin C., Feinstone S. M., Rice C. M. Expression and identification of hepatitis C virus polyprotein cleavage products. J Virol. 1993 Mar;67(3):1385–1395. doi: 10.1128/jvi.67.3.1385-1395.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hijikata M., Kato N., Ootsuyama Y., Nakagawa M., Shimotohno K. Gene mapping of the putative structural region of the hepatitis C virus genome by in vitro processing analysis. Proc Natl Acad Sci U S A. 1991 Jul 1;88(13):5547–5551. doi: 10.1073/pnas.88.13.5547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Houghton M., Weiner A., Han J., Kuo G., Choo Q. L. Molecular biology of the hepatitis C viruses: implications for diagnosis, development and control of viral disease. Hepatology. 1991 Aug;14(2):381–388. [PubMed] [Google Scholar]
- Kunkel T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc Natl Acad Sci U S A. 1985 Jan;82(2):488–492. doi: 10.1073/pnas.82.2.488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin C., Chambers T. J., Rice C. M. Mutagenesis of conserved residues at the yellow fever virus 3/4A and 4B/5 dibasic cleavage sites: effects on cleavage efficiency and polyprotein processing. Virology. 1993 Feb;192(2):596–604. doi: 10.1006/viro.1993.1076. [DOI] [PubMed] [Google Scholar]
- Meyers G., Tautz N., Dubovi E. J., Thiel H. J. Viral cytopathogenicity correlated with integration of ubiquitin-coding sequences. Virology. 1991 Feb;180(2):602–616. doi: 10.1016/0042-6822(91)90074-L. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers G., Tautz N., Stark R., Brownlie J., Dubovi E. J., Collett M. S., Thiel H. J. Rearrangement of viral sequences in cytopathogenic pestiviruses. Virology. 1992 Nov;191(1):368–386. doi: 10.1016/0042-6822(92)90199-Y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss B., Elroy-Stein O., Mizukami T., Alexander W. A., Fuerst T. R. Product review. New mammalian expression vectors. Nature. 1990 Nov 1;348(6296):91–92. doi: 10.1038/348091a0. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Warrener P., Tamura J. K., Collett M. S. RNA-stimulated NTPase activity associated with yellow fever virus NS3 protein expressed in bacteria. J Virol. 1993 Feb;67(2):989–996. doi: 10.1128/jvi.67.2.989-996.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wengler G., Wengler G., Nowak T., Castle E. Description of a procedure which allows isolation of viral nonstructural proteins from BHK vertebrate cells infected with the West Nile flavivirus in a state which allows their direct chemical characterization. Virology. 1990 Aug;177(2):795–801. doi: 10.1016/0042-6822(90)90552-3. [DOI] [PubMed] [Google Scholar]
- Wengler G., Wengler G. The carboxy-terminal part of the NS 3 protein of the West Nile flavivirus can be isolated as a soluble protein after proteolytic cleavage and represents an RNA-stimulated NTPase. Virology. 1991 Oct;184(2):707–715. doi: 10.1016/0042-6822(91)90440-m. [DOI] [PubMed] [Google Scholar]
- Wiskerchen M., Belzer S. K., Collett M. S. Pestivirus gene expression: the first protein product of the bovine viral diarrhea virus large open reading frame, p20, possesses proteolytic activity. J Virol. 1991 Aug;65(8):4508–4514. doi: 10.1128/jvi.65.8.4508-4514.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Heijne G. A new method for predicting signal sequence cleavage sites. Nucleic Acids Res. 1986 Jun 11;14(11):4683–4690. doi: 10.1093/nar/14.11.4683. [DOI] [PMC free article] [PubMed] [Google Scholar]