Abstract
Human pluripotent stem cells (PSCs) represent an opportunity to study human development in vitro, to model diseases in a dish, to screen drugs as well as to provide an unlimited and ethically unimpeded source of therapeutic cells. Cortical GABAergic interneurons, which are generated from Medial Ganglionic Eminence (MGE) cells and Caudal Ganglionic Eminence (CGE) cells during embryonic development, regulate cortical neural networks by providing inhibitory inputs. Their malfunction, resulting in failure to intricately regulate neural circuit balance, has been implicated in brain diseases, such as schizophrenia, autism and epilepsy. In this study, using combinatorial and temporal modulation of developmentally relevant dorsoventral and rostrocaudal signaling pathways, we efficiently generated MGE cells vs. CGE cells from human PSCs, which predominantly generate Parvalbumin-expressing or Somatostatin-expressing interneurons vs. Calretinin-expressing interneurons, respectively. Efficient generation of specific differentiated progenies of hPSCs as shown in this study will be a pivotal step to realize the full potential of hPSCs for regenerative medicine, developmental studies, disease modeling, bioassay, and drug screening.
Keywords: Pluripotent stem cells, Medial Ganglionic Eminence, Caudal Ganglionic Eminence, Interneurons, Differentiation
1. Introduction
During early neurodevelopment, cortical interneuron progenitors arise from the ventral telencephalic area, such as Medial Ganglionic Eminence (MGE) and Caudal Ganglionic Eminence (CGE) [1]. Secreted signaling molecules from nearby organizers, such as SHH [2,3], Wnt [4] and FGF8 [5,6], play a crucial role during this early phenotype specification of ventral telencephalon. These signals trigger the regulatory cascades that lead to cortical interneuron development by inducing key transcription factors, such as Nkx2.1 in MGE [7–9] and CoupTFII in CGE [8,9]. Once phenotypically specified in the ventral area, MGE- or CGE-derived GABAergic interneurons migrate tangentially to their target sites, where they form local synaptic connections and critically modulate cortical circuitry. MGE cells mostly mature to generate Parvalbumin+ or Somatostatin+ interneurons, which comprise more than half of the entire cortical interneuron population, while CGE cells mostly mature to generate Calretinin+ interneurons [10].
Dysfunction of interneurons has been implicated in various brain diseases, such as epilepsy, schizophrenia and autism [11], for which more effective treatments are desperately needed. Human pluripotent stem cells (hPSCs), especially human induced pluripotent stem cells (hiPSCs), provide an unprecedented opportunity to study disease mechanisms and develop novel therapeutics for these brain diseases [12–17], as long as specific neuronal subtypes can be efficiently derived from hiPSCs. We and others have recently reported on the efficient generation of MGE-type interneurons from hPSCs [18–21] by employing developmentally relevant signaling pathway activation, and the derivation of CGE-type interneurons using various methods [21–23].
In this study, we provide a detailed description of the method for the stepwise and combinatorial treatment of developmentally relevant signaling molecules to efficiently generate MGE vs. CGE cells from hPSCs. In this protocol, we first ventralize early neuroectoderm using SHH activation and Wnt inhibition, followed by further rostralization and caudalization of ventralized neuroectoderm using FGF8 vs. FGF19 signaling activation to induce MGE vs. CGE phenotype. This highly efficient method, which recapitulates early developmental processes, will help accelerate the realization of the full potential of induced pluripotent stem cell (iPSC) technology in the study of disease mechanisms and the development of novel therapeutics.
2. Materials
2.1. Reagents
2.1.1. Cells
Human embryonic stem cells (ESCs): H9 cells, WA09 (WiCell, Madison, WI, passage 45–55)
Human ESCs: H7 cells, WA07 (WiCell, Madison, WI, passage 41–51)
Human iPSC2497 (a kind gift from Dr. Fred Gage, [15], passage 30–40)
2.1.2. Growth media and supplements
DMEM (Life Technologies, Carlsbad, CA, cat. no. 11965–118)
Essential 8 medium kit (E8 medium; Life Technologies, Carlsbad, CA, cat. no. A1517001)
ROCK inhibitor Y-27632 (Selleck Chemicals, Houston, TX, cat. no. S1049)
Knockout serum replacement (KSR; Life Technologies, Carlsbad, CA, cat. no. 10828–028)
Penicillin-streptomycin (Life Technologies, Carlsbad, CA, cat. no. 15140–163)
2-Mercaptoethanol (Sigma-Aldrich, St. Louis, MO, cat. no. M3148)
l-Glutamine, 200 mM (Life Technologies, Carlsbad, CA, cat. no. 25030–164)
LDN193189 (Stemgent, Cambridge, MA, cat. no. 04–0074)
SB431542 (Selleck Chemicals, Houston, TX, cat. no. S1067)
IWP-2 (Selleck Chemicals, Houston, TX, cat. no. S7085)
SAG (Cayman Chemical, Ann Arbor, MI, 11914)
Recombinant human FGF-8 (PeproTech, Rocky Hill, NJ, cat. no. 100–25)
Recombinant human FGF-19 (PeproTech, Rocky Hill, NJ, cat. no. 100–32)
N-2 supplement (Life Technologies, Carlsbad, CA, cat. no. 17502–048)
DMEM-F12 (Life Technologies, Carlsbad, CA, cat. no. 11320– 082)
Recombinant human GDNF (PeproTech, Rocky Hill, NJ, cat. no. 450–10)
Recombinant human BDNF (PeproTech, Rocky Hill, NJ, cat. no. 450–02)
HBSS, calcium, magnesium, no phenol red (Life Technologies, Carlsbad, CA, cat. no. 14025–134)
Boc-Asp(OMe) fluoromethyl ketone (BAF; Sigma-Aldrich, St. Louis, MO, cat. no. B2682)
2.1.3. Enzymes and other reagents
Matrigel, hESC-qualified matrix (BD Biosciences, San Jose, CA, cat. no. 354277)
DPBS, without calcium and magnesium (Life Technologies, Carlsbad, CA, cat. no. 14190–250)
EDTA (Sigma-Aldrich, St. Louis, MO, cat. no. E6758)
Poly-l-ornithine solution (PLO; Sigma-Aldrich, St. Louis, MO, cat. no. P4957)
Fibronectin (FN; Sigma-Aldrich, St. Louis, MO, cat. no. F0895)
Trypsin-EDTA solution, 0.5% (Life Technologies, Carlsbad, CA, cat. no. 15400-054)
Fixation/permeabilization solution kit (BD Biosciences, San Jose, CA, cat. no. 554714)
Bovine serum albumin (BSA; Sigma-Aldrich, St. Louis, MO, cat. no. A9418)
Saponin (Sigma-Aldrich, St. Louis, MO, cat. no. 47036)
TRIzol reagent (Life Technologies, Carlsbad, CA, cat. no. 15596-026)
PureLink RNA mini kit (Life Technologies, Carlsbad, CA, cat. no. 12183018A)
SuperScript II reverse transcriptase (Life Technologies, Carlsbad, CA, cat. no. 18064-014)
Oligo d(T)12–18 (Gene Link, Hawthorne, NY, cat. no. 26–4000-05)
SYBR green I nucleic acid gel stain (Sigma-Aldrich, St. Louis, MO, cat. no. S9430)
Formaldehyde, 37% (Electron Microscopy Sciences, Hatfield, PA, cat. no. 15686)
Normal donkey serum (NDS; Sigma-Aldrich, St. Louis, MO, cat. no. D9663)
Triton X-100 (Sigma-Aldrich, St. Louis, MO, cat. no. X100)
Hoechst 33342 (Life Technologies, Carlsbad, CA, cat. no. H3570)
Fluoromount-G (SouthernBiotech, Birmingham, AL, cat. no. 0100-01)
Isoflurane (Sigma-Aldrich, St. Louis, MO, cat. no. 792632)
Buprenorphine hydrochloride (Sigma-Aldrich, St. Louis, MO, cat. no. B9275)
Triple antibiotic ointment (Fisher Scientific, Pittsburgh, PA, cat. no. 19–075–383)
Pentobarbital (Sigma-Aldrich, St. Louis, MO, cat. no. P0500000)
Heparin sodium (Sigma-Aldrich, St. Louis, MO, cat. no. 1304005)
Sodium chloride standard, aqueous solution (Fisher Scientific, Pittsburgh, PA, cat. no. 7213095)
Sucrose (Fisher Scientific, Pittsburgh, PA, cat. no. S6–12)
2.1.4. Antibodies
Table 1.
List of antibodies used in these experiments.
| Antibody | Host | Protein concentration/dilution | Supplier | Catalog number |
|---|---|---|---|---|
| Anti-Goat, Alexa Fluor 488 | Chicken | 2 µg/ml | Life Technologies | A-21467 |
| Anti-Goat, Alexa Fluor 594 | Donkey | 2 µg/ml | Life Technologies | A-11058 |
| Anti-Mouse, Alexa Fluor 488 | Chicken | 2 µg/ml | Life Technologies | A-21200 |
| Anti-Rabbit, Alexa Fluor 488 | Donkey | 2 µg/ml | Life Technologies | A-21206 |
| Anti-Rabbit, Alexa Fluor 594 | Chicken | 2 µg/ml | Life Technologies | A-21442 |
| Anti-Rabbit, Alexa Fluor 647 | Goat | 2 µg/ml | Life Technologies | A-21245 |
| Anti-Rat, Alexa Fluor 594 | Goat | 2 µg/ml | Life Technologies | A-11007 |
| Anti-Sheep, Alexa Fluor 488 | Donkey | 2 µg/ml | Life Technologies | A-11015 |
| Anti-Sheep, Alexa Fluor 594 | Donkey | 2 µg/ml | Life Technologies | A-11016 |
| Anti-Sheep, Alexa Fluor 647 | Donkey | 2 µg/ml | Life Technologies | A-21448 |
| β-Tubulin | Rabbit | 0.5 µg/ml | Covance | MRB-435P |
| β-Tubulin | Mouse | 0.5 µg/ml | Covance | MMS-435P |
| Calbindin | Rabbit | 1/10000 | Swant | CB38 |
| Calretinin | Goat | 1/5000 | Swant | CG1 |
| CoupTFII | Mouse | 1 µg/ml | Perseus Proteomics | PP-H7147-00 |
| FoxG1 | Rabbit | 1.4–2 µg/ml | Abcam | ab18259 |
| GABA | Rabbit | 2.5–5 µg/ml | Sigma–Aldrich | A2052 |
| GAD 65/67 | Rabbit | 1/1000 | EMD Millipore | AB1511 |
| GFAP | Rabbit | 1/1000 | DAKO | Z0334 |
| Human nuclei | Mouse | 1/1000 | EMD Millipore | MAB1281 |
| Lhx6 | Rabbit | 1/1000 | Gift from Dr. Pachnis | – |
| Neuropeptide Y | Sheep | 1/1000 | EMD Millipore | AB1583 |
| Nkx2.1 (TTF1) | Rabbit | 1/2000 | Abcam | ab133737 |
| Olig2 | Rabbit | 1/500 | EMD Millipore | AB9610 |
| Parvalbumin | Mouse | 1/5000 | EMD Millipore | MAB1572 |
| Pax6 | Mouse | 2–5 µg/ml | Developmental Studies Hybridoma Bank | PAX6 |
| Somatostatin | Rat | 1/5000 | EMD Millipore | MAB354 |
| Sox6 | Rabbit | 0.1–1 µg/ml | EMD Millipore | AB5805 |
| VIP | Rabbit | 1/1000 | ImmunoStar | 20077 |
2.1.5. Primers
Table 2.
List of primers used in these experiments.
| cDNA | Direction | Sequence |
|---|---|---|
| Nanog | Forward | TTGGGACTGGTGGAAGAATC |
| Reverse | GATTTGTGGGCCTGAAGAAA | |
| Oct | Forward | TCTCCAGGTTGCCTCTCACT |
| Reverse | GTGGAGGAAGCTGACAACAA | |
| Nkx2.1 | Forward | ACTGACACAAAGGAAGTGCCCC |
| Reverse | GCCGCAAATACCAAACTGCC | |
| Gsx2 | Forward | ATGTCGCGCTCCTTCTATGTC |
| Reverse | CAAGCGGGATGAAGAAATCCG | |
| Dlx2 | Forward | ACCAGACCTCGGGATCCGCC |
| Reverse | CTGCGGGGTCTGAGTGGGGT | |
| Lhx6 | Forward | GGTCTATTCCTTGCGTGGATTATG |
| Reverse | TCCGTGTGTGTGTTTTCCCC |
2.2. Equipment and software
Laminar flow hood
CO2 incubator (Sanyo, Osaka, Japan)
Refrigerator, 4 °C (Sanyo, Osaka, Japan)
Inverted microscope with phase contrast (Olympus, Center Valley, PA, CK40)
Centrifuge with swing-bucket rotor (Eppendorf, Hauppauge, NY, 5810)
Pipet-aid (Drummond Scientific, Broomall, PA, cat. no. 4-000–101)
Serological pipettes (25, 10 and 5 ml; BD Biosciences, San Jose, CA, cat. nos. 357535, 357551 and 357543, respectively)
Pipetman (P1000, P200, P20 and P10; Gilson, Middleton, WI)
Sterile pipette tips (1 ml, 200 and 10 µl; Fisher Scientific, Pittsburgh, PA, cat. nos. 02–681-165, 02–681-457 and 02–681-440, respectively)
Sterile filter units, 500 ml (Thermo Scientific, Waltham, MA, cat. no. 566-0020)
Tissue culture dishes, 100 mm (Fisher Scientific, Pittsburgh, PA, cat. no. 08–772-E)
Conical tubes (50 and 15 ml; Fisher Scientific, Pittsburgh, PA, cat. nos. 14–959-49A and 14–959-49B, respectively)
Ultra low attachment 6-well tissue culture plates (Corning, Corning, NY, cat. no. 3471)
Cell lifters (Fisher Scientific, Pittsburgh, PA, cat. no. 08–100–240)
Cover slips, 12 mm (Chemglass life Sciences, Vineland, NJ, cat. no. CLS-1760-012)
Standard 24-well tissue culture plates (Corning, Corning, NY, cat. no. 353047)
FACSAria III (BD Biosciences, San Jose, CA)
FlowJo software (Tree Star, Ashland, OR, http://www.flowjo.com)
Low-profile PCR tube strips, 0.2 ml (Bio-Rad Laboratories, Inc., Hercules, CA, cat. no. TLS-0801)
Cap strips for PCR tubes (Bio-Rad Laboratories, Inc., Hercules, CA, cat. no. TCS-0803)
DNA Engine Opticon (MJ Research, Waltham, MA)
MacVector software (MacVector, Cary, NC, http://macvector.com)
Inverted microscope with DSU spinning disk confocal (Olympus, Center Valley, PA IX81)
Stereo Investigator image capture equipment and software (MBF Bioscience, Williston, VT, http://www.mbfbioscience.com/stereo-investigator.)
Prism 6 software (GraphPad Software, La Jolla, CA, http://www.graphpad.com/scientific-software/prism)
Angle Two stereotaxic for mouse (Leica Biosystems, Buffalo Grove, IL, cat. no. 39464610)
Cunningham mouse stereotaxic adaptor (Harvard Apparatus, Holliston, MA, cat. no. 724794)
Mobile isoflurane vaporstick system with small induction box (Harvard Apparatus, Holliston, MA, cat. no. 726422)
Electric heating pad (Fisher Scientific, Pittsburgh, PA, cat. no. NC0331246)
Cryostat (Leica Biosystems, Buffalo Grove, IL, CM1950)
MetaMorph software (Molecular Devices, Sunnyvale, CA, http://www.moleculardevices.com/)
3. Methods
3.1. PSC culture
3.1.1. Preparation of Matrigel-coated plates
Matrigel was used as a substrate to support the expansion of hPSCs. At least 1 day before the experiment, a bottle of Matrigel was thawed overnight at 4 °C. The bottle was transferred on ice to a tissue culture hood and kept cold during its preparation as Matrigel solidifies at RT. The Matrigel was diluted 1:50 with DMEM by repetitively pipetting in cold DMEM taken directly from the refrigerator. Immediately prior to passaging the hPSCs, a 10 cm plate was coated with 1–2 ml of diluted Matrigel and shaken well to cover the entire surface. After incubation at RT for at least 30 min, the Matrigel was aspirated from the plate.
3.1.2. Passaging and maintenance of PSCs
Human PSCs were passaged using 0.5 mM EDTA every 4–5 days when confluence reached ~80%. To prepare working solution of EDTA for passaging, we added 500 µl of 0.5 M EDTA into 500 ml of PBS, filter sterilized the solution, and stored it at 4 °C for up to 6 months [24].
Prior to passaging, the cells were inspected under an inverted microscope. In the rare case that any differentiating colony was detected, those colonies were marked and aspirated before passaging. The medium from the hPSCs was aspirated. The plate was washed twice with 1–2 ml of PBS (without Ca++ and Mg++) to remove the remaining magnesium and calcium from the medium. When washing, PBS was added to the wall of the plate slowly to avoid washing the colonies off the plate. After two washes, the cells were incubated with 1 ml of EDTA at RT for 5 min within the hood. After 5 min, 1–2 ml of KSR medium (20% knockout serum replacement, DMEM, 2 mM l-Glutamine, 10 µM β-Mercaptoethanol and penicillin–streptomycin) with ROCK inhibitor (Y-27632, 5 µM) was added to neutralize the EDTA. The cells were resuspended with a P1000 pipette to lift the colonies off the plate and collect at the bottom of the plate, which was tilted at a 30° angle. A cell lifter was also used to detach the remaining cells by gentle mechanical dislodgement. The collected pool of colonies was dispersed and broken into smaller clusters by triturating the cells with the plate continuously tilted. Vigorous mixing or pipetting was avoided to prevent the cells from shearing. The homogeneous cell suspension was plated at about 2.5 × 106 cells per Matrigel-coated 10 cm plate with E8 medium. Rock inhibitor (Y-27632, 5 µM) was added on the first day of passaging to prevent dissociation-induced death of hPSCs [25]. The plate was immediately shaken back and forth and side to side to evenly distribute the cells and prevent the cells from aggregating densely in one area. The plate was then placed in the incubator to allow the cells to attach and expand. Human PSCs were maintained in E8 medium (Fig. 1a), which was changed every day.
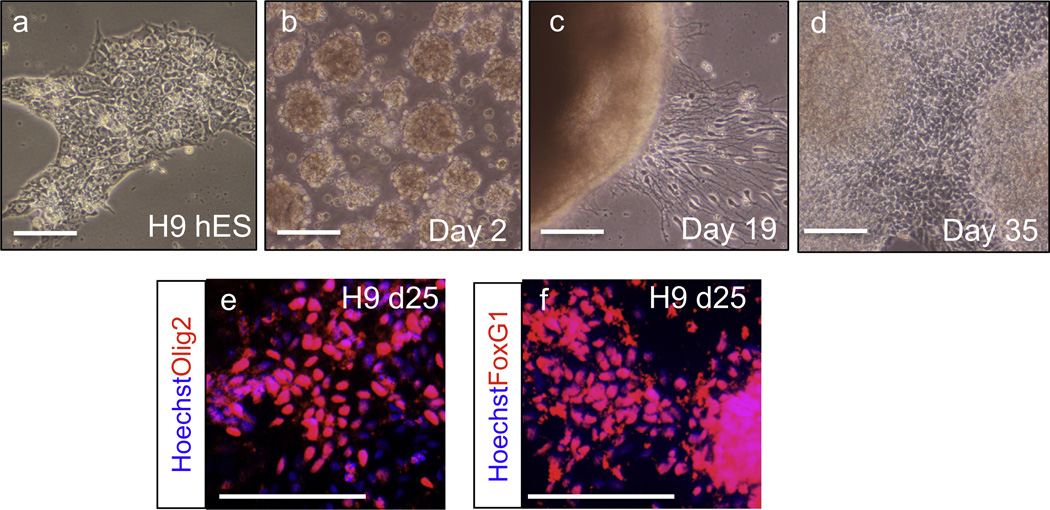
Fig. 1.
(a–d) Bright-field images taken during in vitro differentiation of H9 cells. (e and f) H9-derived MGE cells, assayed after 25 days of differentiation, highly expressed independent ventral telencephalic marker Olig2 and telencephalic marker FoxG1 (adapted from [46]). Scale bar: 100 µm.
3.2. Differentiation into ventral telencephalic lineage
3.2.1. Neural induction of floating spheres
For differentiation, PSCs were passaged and grown as floating aggregates (Fig. 1b) in low adherent plates. The passaged cells were transferred to a 15 ml conical tube and centrifuged at 1000 rpm for 1 min, followed by careful removal of the supernatant. The cell pellet was resuspended in KSR medium (20% knockout serum replacement, DMEM, 2 mM l-Glutamine, 10 µM β-mercaptoethanol and penicillin-streptomycin) with ROCK inhibitor (Y-27632, 5 µM) and gently triturated using a P1000 pipette to obtain a homogeneous cell suspension. About 2.5 × 106 cells were added to each well of a 6-well low adherent plate with 3 ml of KSR medium in each well. High cell density per well optimized the survival of cells. Rock inhibitor (Y-27632, 5 µM) was added on the first day of differentiation to prevent dissociation-induced death of hPSCs [25]. For neural induction, cells were treated with LDN193189 (100 nM) from day 0 to day 14 and with SB431542 (10 µM) from day 0 to day 7 [26].
3.2.2. Induction of ventral telencephalic lineage
For ventral telencephalic lineage induction, cells were treated with IWP-2 (5 µM) from day 0 to day 7 to prevent dorsalization and rostralization, and with SAG (0.1 µM) from day 0 to day 21 to induce ventralization. This treatment resulted in cell populations enriched in ventral telencephalic tissues (Fig. 1e,f and Table 4). After 2 weeks of floating culture, cells were transferred to Poly-l-ornithine- (PLO; 15 mg/ml) and fibronectin- (FN; 1 mg/ml) coated surfaces for adherent culture (Fig. 1c).
Table 4.
Summary of cell counting analysis in vitro. N = independent in vitro differentiation.
| Cell line | Induction | Day of diff. |
% Total cells (Mean ± SEM) |
N | |
|---|---|---|---|---|---|
| Olig2 | H9 | MGE | 25 | 71.8 + 3.7 | 4 |
| FoxG1 | H9 | MGE | 25 | 89.8 + 2.0 | 4 |
| Nkx2.1 | H9 | MGE | 25 | 82.5 + 2.7 | 4 |
| Nkx2.1 | H7 | MGE | 25 | 84.1 + 1.4 | 4 |
| Nkx2.1 | iPSC 2497 |
MGE | 25 | 81.6 + 6.7 | 4 |
| GABA | H7 | MGE | 42 | 76.3 + 2.1 | 4 |
| GAD | H9 | MGE | 42 | 69.1 + 4.1 | 4 |
| Lhx6 | H9 | MGE | 42 | 71.4 + 6.4 | 4 |
| Lhx6 | H7 | MGE | 42 | 66.9 + 3.4 | 4 |
| Sox6 | H9 | MGE | 42 | 77.4 + 2.2 | 4 |
| Calb | H9 | MGE | 42 | 22.2 + 6.0 | 4 |
| PV | H9 | MGE | 42 | 0.87 + 0.32 | 4 |
| SST | H9 | MGE | 42 | 1.53 + 0.57 | 4 |
| VIP | H9 | MGE | 60 | 0.0 + 0.0 | 4 |
| Calr | H9 | MGE | 60 | 0.57 + 0.33 | 4 |
| VIP | H9 | CGE | 60 | 10.46 + 1.35 | 4 |
| Calr | H9 | CGE | 60 | 16.77 + 5.36 | 4 |
| PV | H9 | CGE | 60 | 0.0 + 0.0 | 4 |
| SST | H9 | CGE | 60 | 0.0 + 0.0 | 4 |
3.3. Further specification of MGEvs. CGE phenotype
3.3.1. Induction of MGE vs. CGE phenotype
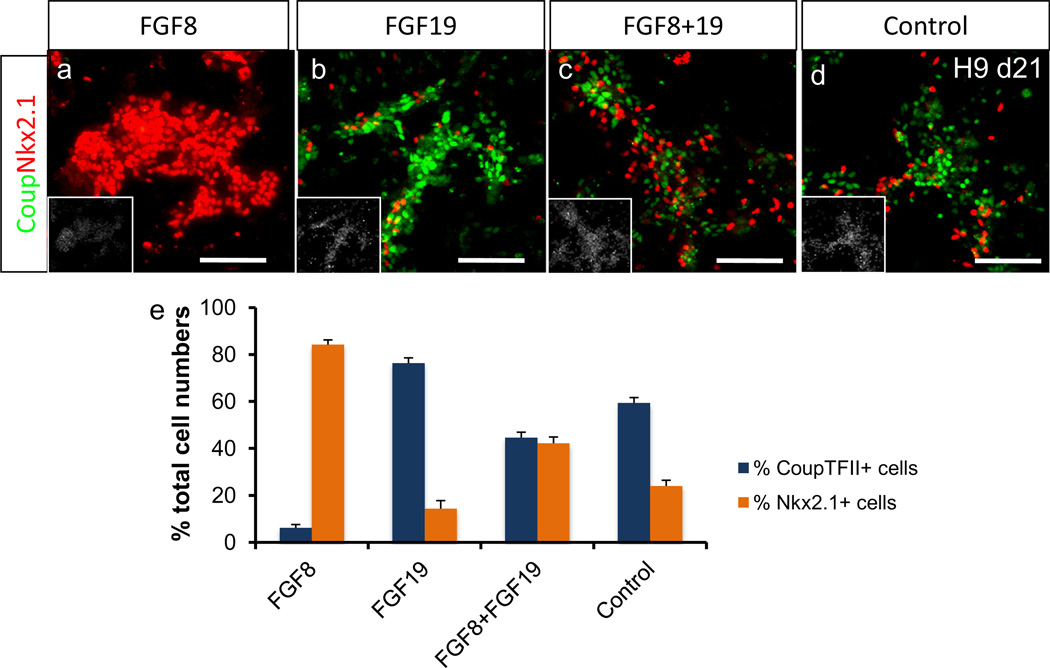
For induction of MGE phenotype, differentiating spheres were initially treated with FGF8 (100 ng/ml) in KSR medium in the second week followed by FGF8 in N2 medium (DMEM-F12 medium supplemented with N2) in the third week. This treatment resulted in cultures enriched in MGE cells, as shown by the high proportion of Nkx2.1+ cells (Fig. 2a and e). For induction of CGE phenotype, differentiating spheres were initially treated with FGF19 (100 ng/ml) in KSR medium in the second week followed by FGF19 in N2 medium in the third week. This resulted in cultures enriched in CGE cells, as indicated by the high proportion of CoupTFII+ cells (Fig. 2b and e).
Fig. 2.
FGF8 and FGF19 regulate rostral/caudal identity of ventral telencephalic cells (adapted from [46]). FGF8 treatment induced MGE phenotype while FGF19 induced CGE phenotype, as shown by immunocytochemistry and cell counting analysis after 3 weeks of differentiation of H9 cells (Mean ± SEM; n = 3, P < 0.05, two tailed t-test). The image in the insets is Hoechst-labeled nuclei of the same microscope field. Scale bar: 100 µm.
3.3.2. Maturation of interneurons
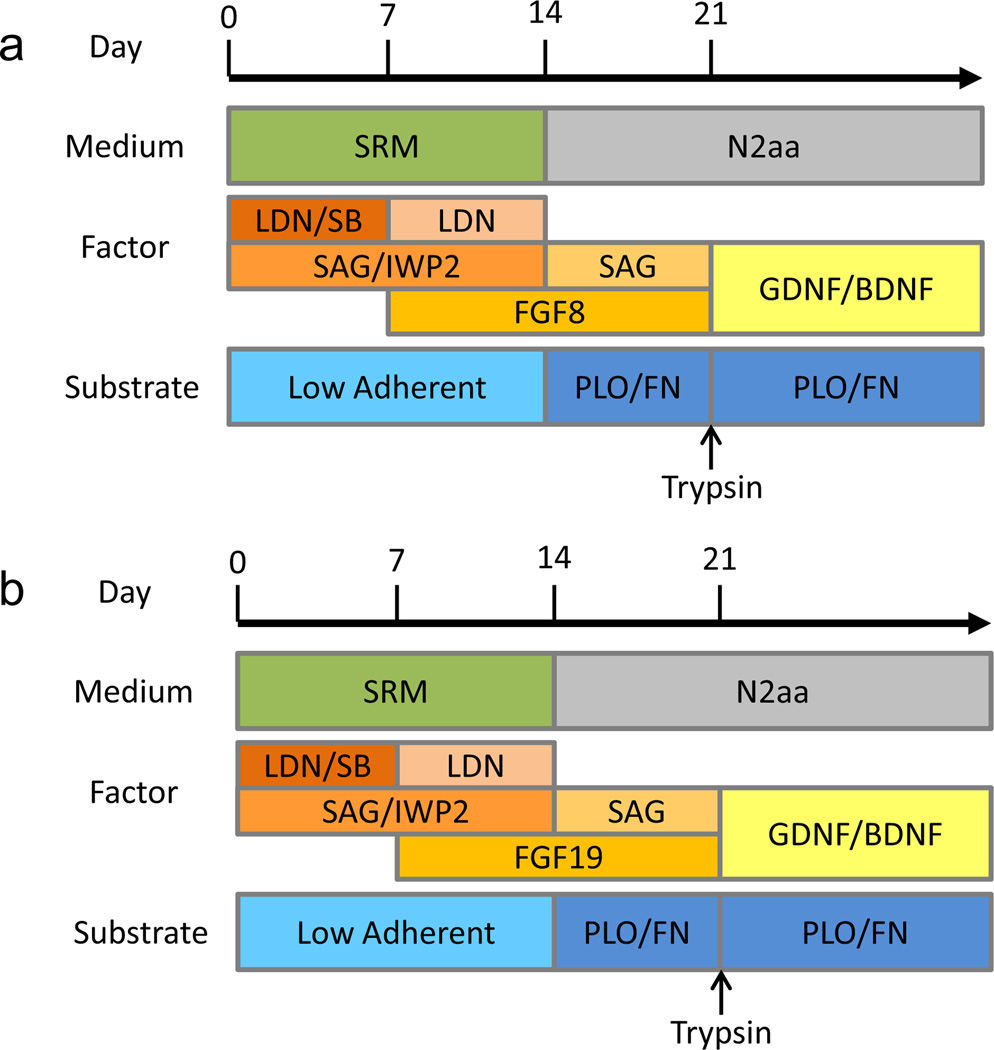
After 3 weeks of differentiation, cells were trypsinized, and droplets of 100,000 cells in 5 µl were transferred to PLO/FN-coated coverslips. After attachment of spotted cells to the coverslips, 0.5 ml of differentiation medium (N2 medium [27] supplemented with 10 ng/ml GDNF and 10 ng/ml BDNF) was added per well of a 24-well plate for further differentiation and maturation (Fig. 1d). Schematic diagrams of the differentiation procedures are presented in Fig. 3a (MGE differentiation) and b (CGE differentiation).
Fig. 3.
(a) Overview of the optimized MGE derivation protocol. (b) Overview of the optimized CGE derivation protocol.
3.4. FACS analysis of fixed cells
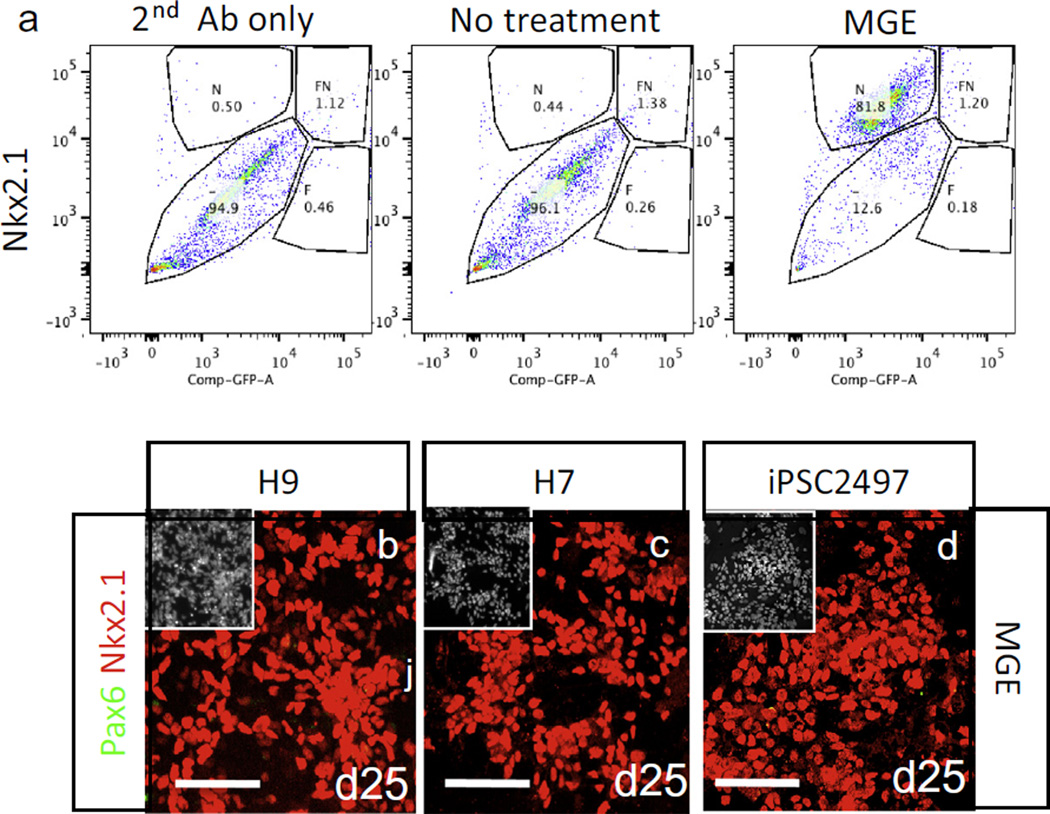
To quantitatively analyze the phenotype of generated cells, we performed FACS analysis of stained cells at day 21 of differentiation. Differentiated cells were trypsinized and fixed in Fix/Perm solution for 30 min, and incubated with blocking buffer (PBS with 0.1 mg/ml BSA and 0.1% Saponin) for 10 min. Blocked cells were incubated with primary antibody (anti-Nkx2.1) in blocking buffer for 30 min. After washing with PBS, we incubated the cells with Alexa 647-conjugated secondary antibodies (1:1000) for 15 min. As a control, some samples were incubated only with secondary antibody. After washing with PBS, we suspended the cells in blocking buffer and analyzed them using the FACSAria. Flowjo software was used to analyze raw data. Ten thousand cells were recorded per analysis. As shown in Fig. 4a, more than 80% of the total cells were induced into an MGE phenotype by MGE induction conditions, as indicated by Nkx2.1 positivity (79.5 ± 1.8% Nkx2.1+ cells, n = 3). These results were in accordance with immunocytochemical characterization of cells, obtained from three different hPSC lines, using the same antibody. All PSC lines that were tested generated enriched MGE cell populations (Fig. 4b–d, Table 4).
Fig. 4.
FACS analysis of fixed cells (adapted from [46]). (a) FACS analysis of MGE cells generated from H9 cells after Nkx2.1 staining at day 21 of differentiation. (b–d) Combined and temporal treatment with IWP-2, SAG and FGF8 resulted in robust induction of MGE cells from H9 and H7 hESCs as well as iPSCs, assayed after 25 days of differentiation. The image in the insets is Hoechst-labeled nuclei of the same microscope field. Scale bar: 100 µm.
3.5. Real-time PCR analysis of time-course gene expression
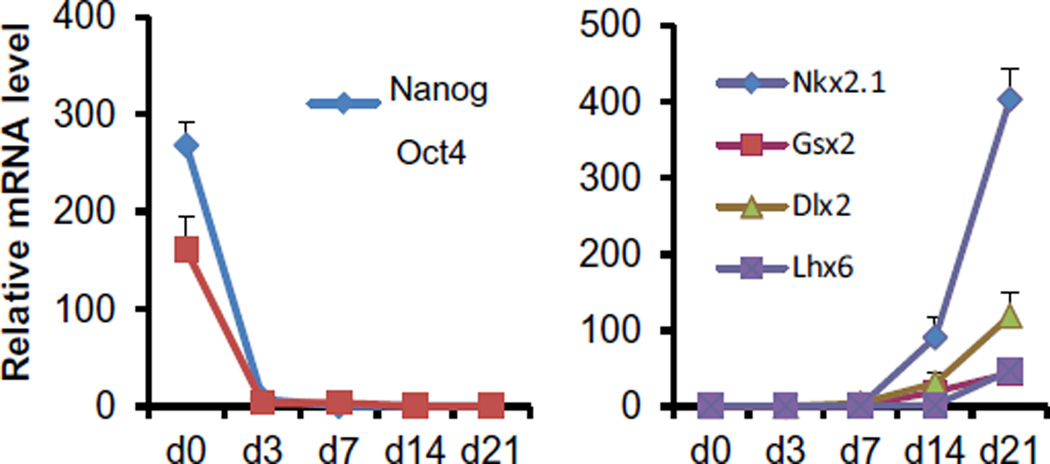
To quantitatively analyze multiple marker gene expressions at multiple time points during differentiation (days 0, 3, 7, 14 and 21 of differentiation), we performed real-time PCR analysis. Total RNA was prepared using TRIzol reagent and PureLink RNA mini kit. cDNA from total RNA was generated using SuperScript II reverse transcriptase and Oligo d(T) primers. For quantitative analysis of the expression level of mRNAs, real-time PCR analyses were performed using the DNA Engine Opticon system and SYBR Green I assays. Primers were designed using the MacVector software (primer sequences are available upon request). PCR was carried out in a 25 µl volume containing 0.5 mM of each primer, 0.5 × SYBR Green I, and 1 µl of cDNA and performed with conditions of a denaturation of 95 °C for 2 min, followed by 50 cycles of amplification (95 °C for 30 s, 55 °C for 30 s, 72 °C for 30 s, and 79 °C for 5 s). Primer dimers were melted at 79 °C before the fluorescent signals after each cycle were measured. The mRNA expression level of each gene was normalized using the housekeeping gene GAPDH as an internal control. The relative values were calculated by setting the normalized value of control as 1. Real-time PCR analysis under these optimal conditions showed that expression of pluripotent stem cell markers Nanog and Oct4 was down-regulated by day 3, and expression of ventral telencephalic markers Nkx2.1, GSX2, DLX2 became evident starting on day 14, followed by expression of postmitotic MGE marker Lhx6 on day 21 (Fig. 5).
Fig. 5.
Gene expression analysis during MGE derivation of H9 cells (adapted from [46]), assayed by real-time PCR (Mean ± SEM; n = 3) on days 0, 3, 7, 14 and 21 of differentiation. The expression level of Nanog, Oct4, Nkx2.1, Gsx2, Dlx2 and Lhx6 was analyzed at each time point after differentiation. The expression level of each gene was normalized to that of GAPDH.
3.6. Immunocytochemistry after neuronal differentiation
For more detailed characterization of phenotype maturation, we performed immunocytochemical analysis at day 21 or 25 of differentiation for precursor phenotype analysis and at day 42 or 60 of differentiation for neuronal phenotype analysis, followed by cell counting analysis. Fixed cells were permeabilized in blocking buffer (PBS, 10% normal donkey serum; NDS) containing 0.1% Triton for 10 min. Cells were then incubated overnight at 4 °C with primary antibodies diluted in PBS containing 2% NDS. The primary and secondary antibody list can be found in Table 1. As secondary antibodies, Alexa 488- Alexa 594- or Alexa 647-labeled IgG was added in a 1:1000 dilution in PBS containing 2% NDS for 30 min at RT. Cell nuclei were stained with Hoechst 33342 (4 mg/ml), and cover slips/tissue sections were mounted onto slides with Fluoromount-G. Confocal analysis was performed using an Olympus DSU Spinning Disk Confocal on an IX81 inverted microscope. Cells were counted using Stereo Investigator image capture equipment and software. For cell counting on cover slips, an optical fractionator probe was used with a 500 µm × 500 µm grid size and 100 µm × 100 mm counting frame (>40 counting sites with >1000 total cells counted per sample) at 40 × magnification. Cover slips from 3–4 independent differentiations were used for analysis. For statistical analysis, we performed a t-test (alpha = 0.05) for comparison of control vs. sample using GraphPad Prism 6 software.
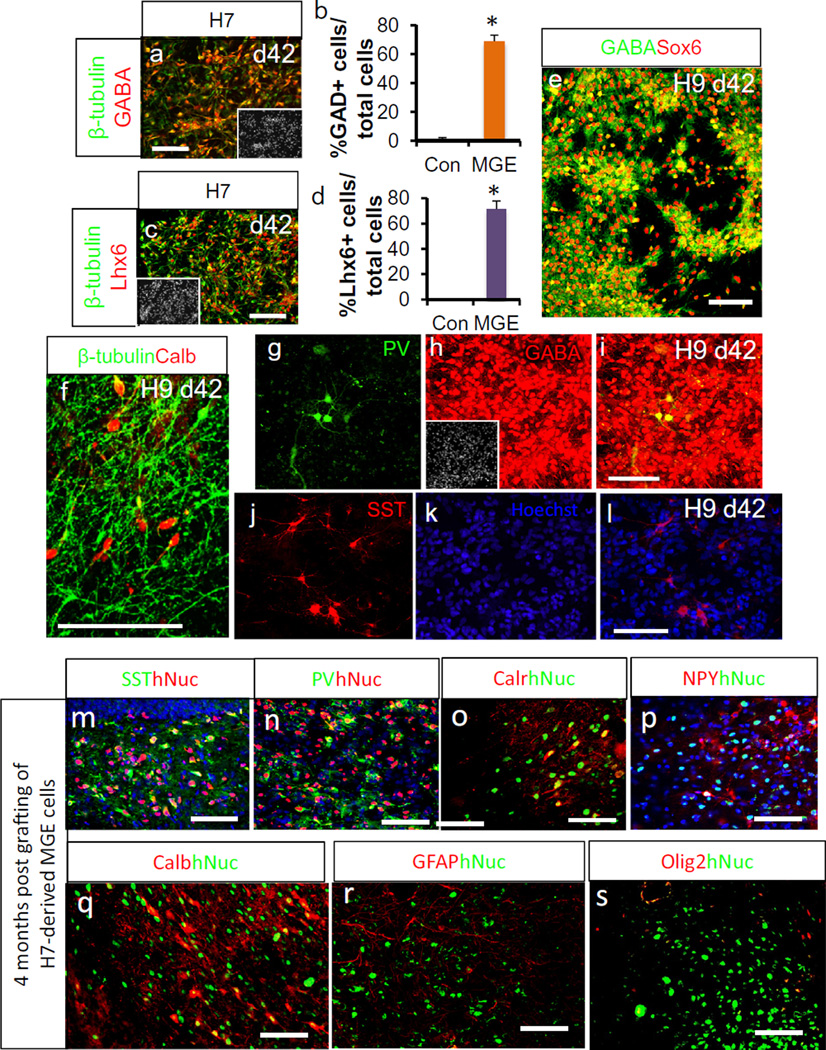
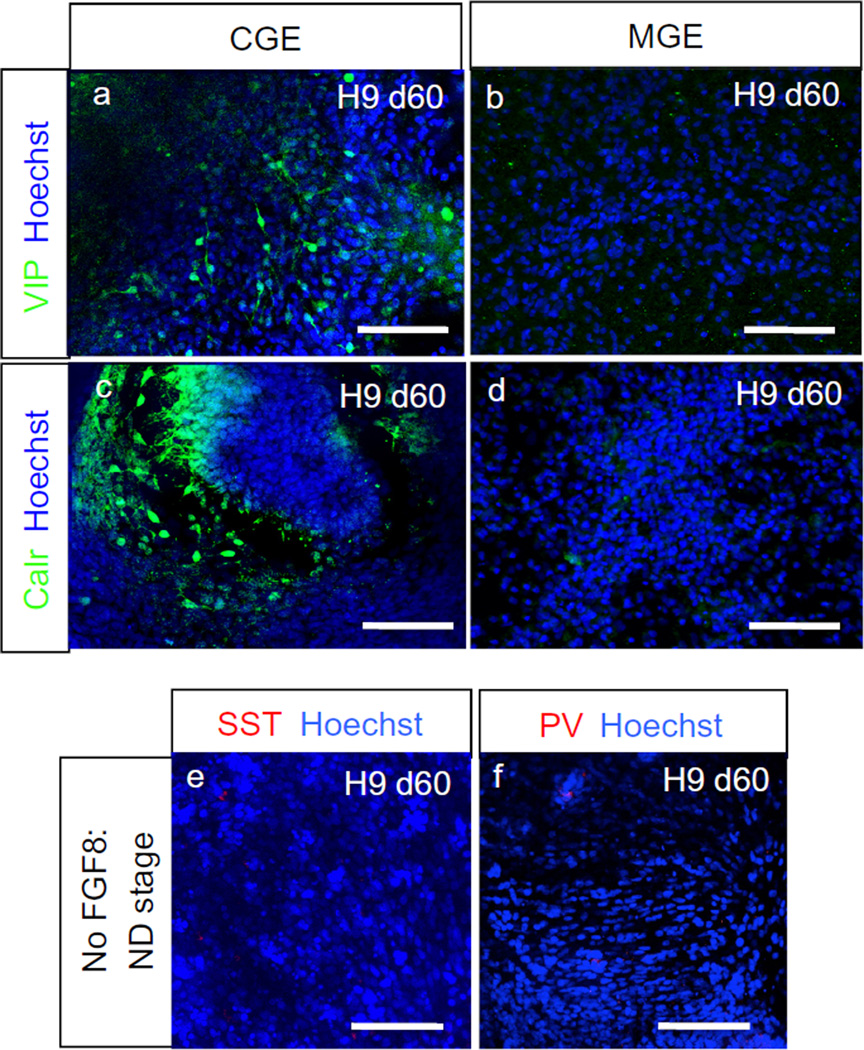
Robust induction of GABAergic neurons was also observed after further maturation of MGE cells (Fig. 6a,b and Table 4). These GABAergic neurons expressed Lhx6, indicating their MGE origin (Fig. 6c,d and Table 4). In addition, these GABAergic neurons co-expressed Sox6 (Fig. 6e and Table 4), which has been recently shown to be a very specific marker for human MGE-derived interneurons [8]. Some of these cells expressed Calbindin (Fig. 6f and Table 4), which was expressed in some of the migrating interneurons. Some of these cells expressed more mature interneuron markers Parvalbumin (PV; Fig. 6g – i and Table 4) and Somatostatin (SST; Fig. 6j – l and Table 4) at this time point, as would be predicted from their MGE origin. CGE cells generated neural populations enriched with VIP+ or Calretinin+ neurons (Fig. 7a – d and Table 4), whereas few SST+ or PV+ cells were observed (Fig. 7e,f and Table 4).
Fig. 6.
Human MGE cells generated GABAergic interneurons (adapted from [46] and [29]). (a–l) Immunocytochemistry and cell counting analysis after 6 weeks of differentiation (Mean ± SEM; n = 4, P < 0.05, two tailed t-test), showing the expression of β-tubulin, GABA, Lhx6, GAD, Sox6, Calbindin (Calb), Parvalbumin (PV) and Somatostatin (SST). Immunostaining was performed on H9-derived cells except a and c, in which H7-derived cells were analyzed. The image in the insets is Hoechst-labeled nuclei of the same microscope field. (m–s) Immunohistochemical analysis of transplanted H7-derived MGE cells 4 months after transplantation, showing the expression of SST, PV, Calretinin (Calr), Neuropeptide Y (NPY), Calb among transplanted human nuclei (hNuc)+ cells. Scale bar: 100 µm.
Fig. 7.
Human CGE cells generated VIP+ or Calretinin+ neurons (adapted from [46]). Immunocytochemistry after 60 days of differentiation of H9 cells. Scale bar: 100 µm.
3.7. Immunohistochemistry after transplantation
3.7.1. Transplantation of MGE cells into hippocampus of mice
Human ESC-derived MGE cells in transplantation medium (HBSS with 10ng/ml GDNF, 10ng/ml BDNF and 20 µM Boc-Asp (OMe) fluoromethyl ketone (BAF)) were injected into hippocampus of mice using a Leica Angle Two digital stereotaxic instrument fitted with a Cunningham Mouse Adaptor. Nod-Scid mice were anesthetized using an induction chamber supplied with 4–5% isoflurane mixed with 0.8-1 L/min oxygen using a calibrated vaporizer. Animals were then administered continuous isoflurane (1–2%) mixed with oxygen (0.8–1.0 L/min) for the duration of anesthesia. Cells were transplanted into the following coordinates: AP 1.75 mm, L ±2.3 mm, V −1.7 mm for the rostral CA3 site; AP 3.25 mm, L ±3.0 mm, V −3.65 mm, −2.9 mm and −2.0 mm for the three caudal sites along the dorso-ventral axis of the hippocampus in this coronal plane. A total of 5 × 104 MGE cells in a 0.5 µl volume were delivered to each target coordinate (0.5 µl/1 min). Animals were removed from the stereotaxic instrument, and an intraperitoneal injection (ip) of buprenorphine (0.03 mg/kg) was administered as an analgesic medication. Topical triple antibiotic ointment was applied to the incision site to prevent infection. During recovery, animals were placed on a heating pad and monitored until recovery of motor function.
3.7.2. Immunohistochemistry and cell counting
Transplanted mice were terminally anesthetized with an ip overdose of pentobarbital (150 mg/kg) and perfused transcardially with heparin saline (0.1% heparin in saline) followed by paraformaldehyde (4%) 4 months post grafting. Brains were removed, postfixed in 4% paraformaldehyde for 12 h, equilibrated in 20% sucrose/PBS solution, and then sectioned coronally at 40-µm using a cryostat. For immunofluorescence staining, tissue sections were incubated with blocking buffer (PBS, 10% NDS) containing 0.1% Triton for 10 min. Cells were then incubated overnight at 4 °C with primary antibodies diluted in PBS containing 2% NDS. After rinsing with PBS, samples were incubated with fluorescent dye-labeled secondary antibodies in PBS containing 2% NDS for 30 min at RT. After rinsing with PBS, Hoechst 33342 (4 µg/ml) was used for counterstaining, and tissue sections were mounted onto slides with Fluoromount-G. Confocal analysis was performed using an Olympus DSU Spinning Disc Confocal on an IX81 inverted microscope, installed with MetaMorph software. Stereo Investigator image capture equipment and software were used for cell counting and estimation of total cell number in the graft using the optical fractionator workflow from every 12th section.
Four months after transplantation, some of grafted cells expressed more mature cortical interneuron markers Somato-statin, Parvalbumin, Calretinin, Neuropeptide Y and Calbindin (Fig. 6m – q and Table 5). Transplanted cells generated very small numbers of astrocytes (GFAP+; Fig. 6r and Table 5) or oligodendrocyte lineage cells at this time point (Olig2+; Fig. 6s and Table 5).
Table 5.
Summary of cell counting analysis of grafted MGE cells. N = animal numbers.
| Months post-txp | % +Cells/grafted cells (Mean ± SEM) | N | |
|---|---|---|---|
| SST | 4 | 30.7 + 2.9 | 3 |
| PV | 4 | 8.0 + 1.7 | 3 |
| Calr | 4 | 5.2 + 0.7 | 3 |
| NPY | 4 | 2.9 + 0.3 | 3 |
| Calb | 4 | 26.7 + 2.0 | 3 |
| β-tubulin | 4 | 89.0 + 1.5 | 3 |
| GFAP | 4 | 0.99 + 0.10 | 3 |
| Olig2 | 4 | 0.83 + 0.42 | 3 |
4. Conclusion
To realize the full potential of hPSC technology, it is critical to efficiently generate homogeneous populations of specific differentiated progenies, as hPSC differentiation tends to be heterogeneous and stochastic [13]. In this study, employing stepwise regulation of developmentally relevant signaling pathways that recapitulates in vivo embryonic development, we established a differentiation method that allowed highly efficient and homogeneous differentiation of hPSCs to MGE cells or CGE cells and then to GABAergic interneurons. Recent studies have shown the potential of GABAergic interneurons as sources for novel cellular therapies for epilepsy [28,29], Parkinson’s disease [30], schizophrenia [31] and injury-induced neuropathic pain [32], yet optimal cell sources for such therapies are limiting. Therefore, the generation of a homogeneous population of GABAergic interneurons from hPSCs is an important step towards clinical translation of this experimental therapeutics. Furthermore, iPSC technology offers the potential to create disease models in a dish, thus facilitating disease mechanistic studies from human disease neurons, again, as long as disease-specific neuronal subtypes can be generated at high efficiency. This method for generating specific subtypes of GABAergic interneurons will play a pivotal role in reliable disease modeling of interneuron-associated diseases, such as epilepsy, autism and schizophrenia [11].
To generate very homogeneous populations of ventral telencephalic neuronal subtypes, we have used temporal and combined modulation of developmentally relevant signaling pathways. Specifically, we have employed (i) induction of neuroectoderm formation by dual SMAD inhibition [26], (ii) inhibition of Wnt signaling, which otherwise caudalizes [33,34] and dorsalizes [35] differentiating neuroectoderms, (iii) strong activation of SHH signaling, which ventralizes neuroectoderms [2], and (iv) activation of FGF8 or FGF19 signaling, which induces the MGE or CGE phenotype, respectively, at the expense of the other phenotype. According to our previous study, a more efficient ventral telencephalic phenotype induction is achieved by early modulation of Wnt and SHH signaling pathways, even before neuroectoderm formation is completed [21]. This is consistent with previous developmental studies, which have shown that early inhibition of Wnt signaling is important for telencephalic induction of the neural plate [33,34], and early SHH signaling in the anterior neural plate at the gastrula stage induces the prospective ventral telencephalon, [36] even prior to neural tube formation.
Recently, a number of protocols for MGE induction from hPSCs was published [4,18,19,21,37], as summarized in Table 6. Most of these protocols use inhibition of BMP (or dual SMAD inhibition [26]) to inhibit non-neural lineage induction, with the exception of the protocol of Liu et al., which employs low density culture for neural induction without any added inhibitor. Though a side-by-side comparison will be needed, there appears to be a tendency for sphere cultures to generate MGE cells more efficiently than adherent cultures (Table 6). These protocols commonly employ a high concentration of SHH (or SHH pathway activator), sometimes along with an inhibitor of the dorsalizing Wnt pathway, resulting in ventralization of the neuroectoderm (Table 6). Strong SHH signaling used to ventralize telencephalic tissues can induce mutually inhibitory downstream signaling of FGF8 and FGF19, which generate MGE cells vs. CGE cells, respectively [21]. In our protocol, by employing exogenous FGF8 and FGF19 signaling after initial ventralization of neuroepithelium, we can efficiently induce either MGE cells or CGE cells, allowing us to avoid any stochastic events during ventral telencephalic induction.
Table 6.
Comparison of recently published protocols for MGE induction (Pur: Purmorphamine, SB: SB431542, LDN: LDN193189, BF: BMPR1A–Fc, XV: XAV939, SAG: smoothened agonist).
| Author | Initial culture | Neural induction | Ventralization | Rostralization | %Nkx2.1+ (%) |
|---|---|---|---|---|---|
| Liu et al. | Sphere | – | d11–25 1.5 µM Pur | – | 90 |
| Germain et al. | Adherent | 500 ng/ml noggin | d6–16 500 ng/ml SHH | – | 46.70 |
| Nicholas et al. | Sphere | 10 µM SB 1.5 µg/ml BF | d0–14 1 µg/ml DKK1 d0–35 1–2 µM Pur | – | 90.80 |
| Maroof et al. | Adherent | 10 µM SB 100 nM LDN | d0–10 2 µM XV d10–18 1µM Pur 5 nM SHH | – | ~50 |
| Kim et al. | Sphere | 10 µM SB 100 nM LDN | d0–14 5µM IWP2 d0–21 100 nM SAG | 100 ng/ml FGF8 | 81.80 |
MGE cells spontaneously differentiate into Lhx6- and Sox6-expressing GABAergic interneurons, indicating that they are phenotype-specified neural progenitors with intrinsic properties to become specific subtypes of GABAergic interneurons. CGE cells generated VIP+ and Calretinin+ neurons, again demonstrating their developmental potential. However, within the time frame of this study, there were still many interneurons that did not show mature marker expressions, such as Somatostatin or Parvalbumin. Previous studies have reported that such markers are expressed in the long-term maturation process of human interneurons that follow their internal clock during the course of human brain development [18,38]. This long-term maturation could pose a major problem for downstream application that requires mature interneurons. However, such early developing interneurons before full maturation exhibit great potential for cell replacement therapy with their migratory function and ability to integrate into adult brains [39]. Additionally, they are more amenable to cell culture manipulations, such as trypsinization, compared to mature cortical interneurons with elaborate branching patterns. In fact, we have shown that these maturing interneurons demonstrate efficacy in ameliorating seizure activity even before attaining full maturation [29]. These developmental interneurons could also provide valuable resources for modeling neurodevelopmental diseases, such as schizophrenia, enabling us to model early developmental time points, during which time the developing brains are known to be more susceptible to environmental challenges [40–42]. This is especially relevant, since many known schizophrenia risk genes are highly expressed during early development [43–45].
In this study, we describe the method for the efficient generation of authentic human MGE and CGE cells as well as GABAergic cortical interneurons by developmentally relevant dorsoventral and rostrocaudal modulation, which well recapitulates in vivo ventral telencephalic development. This will help accelerate the realization of the full potential of iPSC technology. In particular, this will provide cell sources for regenerative medicine, for which numerous proof-of-principle studies have been performed to treat epilepsy, Parkinson’s disease, schizophrenia and neuropathic pain [28,30–32]. This homogeneous and authentic cell population will also be beneficial for human developmental studies, where use of embryo-derived human tissues is unethical and limited. Furthermore, efficient generation of different subtypes of cortical interneurons will prove critical to studying disease mechanisms in vitro from patient-derived cells and to screening novel small molecules for treatment of associated diseases. (See Table 3).
Table 3.
Summary of media composition.
| Name | Composition |
|---|---|
| KSR medium | DMEM medium 20% knockout serum replacement 2 mM l-Glutamine 10 µM β-Mercaptoethanol penicillin–streptomycin |
| E8 medium | E8 basal medium 1 × E8 supplement |
| N2 medium | DMEM-F12 medium 1 × N2 supplement |
| Differentiation medium | DMEM-F12 medium 1 × N2 supplement 10 ng/ml GDNF 10 ng/ml BDNF |
Acknowledgements
We thank Dr. Pachnis for kindly providing us with anti-Lhx6 antibodies. This study was supported by NIH Grants (NS079977, MH107884, NS084869, and NS070577) and the Harvard Stem Cell Institute Seed Grant.
Abbreviations
- PSCs
pluripotent stem cells
- hPSCs
human pluripotent stem cells
- MGE
Medial Ganglionic Eminence
- CGE
Caudal Ganglionic Eminence
- hiPSCs
human induced pluripotent stem cells
- iPSC
induced pluripotent stem cell
- ESCs
embryonic stem cells
- E8
Essential 8
- KSR
knockout serum replacement
- OMe
Boc-Asp
- BAF
fluoromethyl ketone
- PLO
Poly-l-ornithine
- FN
fibronectin
- BSA
bovine serum albumin
- NDS
normal donkey serum
- PV
Parvalbumin
- SST
Somatostatin
- ip
intraperitoneal injection
- Calb
Calbindin
- Calr
Calretinin
- NPY
Neuropeptide Y
- hNuc
human nuclei
Footnotes
Disclosure of potential conflict of interest
The authors declare no potential conflicts of interest.
References
- 1.Wonders CP, Anderson SA. Nat. Rev. Neurosci. 2006;7:687–696. doi: 10.1038/nrn1954. [DOI] [PubMed] [Google Scholar]
- 2.Chiang C, Litingtung Y, Lee E, Young KE, Corden JL, Westphal H, Beachy PA. Nature. 1996;383:407–413. doi: 10.1038/383407a0. [DOI] [PubMed] [Google Scholar]
- 3.Fuccillo M, Rallu M, McMahon AP, Fishell G. Development. 2004;131:5031–5040. doi: 10.1242/dev.01349. [DOI] [PubMed] [Google Scholar]
- 4.Li XJ, Zhang X, Johnson MA, Wang ZB, Lavaute T, Zhang SC. Development. 2009;136:4055–4063. doi: 10.1242/dev.036624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fukuchi-Shimogori T, Grove EA. Science. 2001;294:1071–1074. doi: 10.1126/science.1064252. [DOI] [PubMed] [Google Scholar]
- 6.Garel S, Huffman KJ, Rubenstein JL. Development. 2003;130:1903–1914. doi: 10.1242/dev.00416. [DOI] [PubMed] [Google Scholar]
- 7.Sussel L, Marin O, Kimura S, Rubenstein JL. Development. 1999;126:3359–3370. doi: 10.1242/dev.126.15.3359. [DOI] [PubMed] [Google Scholar]
- 8.Ma T, Wang C, Wang L, Zhou X, Tian M, Zhang Q, Zhang Y, Li J, Liu Z, Cai Y, Liu F, You Y, Chen C, Campbell K, Song H, Ma L, Rubenstein JL, Yang Z. Nat. Neurosci. 2013;16:1588–1597. doi: 10.1038/nn.3536. [DOI] [PubMed] [Google Scholar]
- 9.Hansen DV, Lui JH, Flandin P, Yoshikawa K, Rubenstein JL, Alvarez-Buylla A, Kriegstein AR. Nat. Neurosci. 2013;16:1576–1587. doi: 10.1038/nn.3541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tamamaki N, Yanagawa Y, Tomioka R, Miyazaki J, Obata K, Kaneko T. J. Comp. Neurol. 2003;467:60–79. doi: 10.1002/cne.10905. [DOI] [PubMed] [Google Scholar]
- 11.Arber CE, Li M. Front. Cell. Neurosci. 2013;7 doi: 10.3389/fncel.2013.00010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Castiglioni V, Onorati M, Rochon C, Cattaneo E. Neurobiol. Dis. 2012;46:30–40. doi: 10.1016/j.nbd.2011.12.032. [DOI] [PubMed] [Google Scholar]
- 13.Bellin M, Marchetto MC, Gage FH, Mummery CL. Nat. Rev. Mol. Cell Biol. 2012;13:713–726. doi: 10.1038/nrm3448. [DOI] [PubMed] [Google Scholar]
- 14.Matsui T, Akamatsu W, Nakamura M, Okano H. Exp. Neurol. 2012 doi: 10.1016/j.expneurol.2012.09.016. [DOI] [PubMed] [Google Scholar]
- 15.Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, Li Y, Mu Y, Chen G, Yu D, McCarthy S, Sebat J, Gage FH. Nature. 2011;473:221–225. doi: 10.1038/nature09915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lee G, Ramirez CN, Kim H, Zeltner N, Liu B, Radu C, Bhinder B, Kim YJ, Choi IY, Mukherjee-Clavin B, Djaballah H, Studer L. Nat. Biotechnol. 2012;30:1244–1248. doi: 10.1038/nbt.2435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Egawa N, Kitaoka S, Tsukita K, Naitoh M, Takahashi K, Yamamoto T, Adachi F, Kondo T, Okita K, Asaka I, Aoi T, Watanabe A, Yamada Y, Morizane A, Takahashi J, Ayaki T, Ito H, Yoshikawa K, Yamawaki S, Suzuki S, Watanabe D, Hioki H, Kaneko T, Makioka K, Okamoto K, Takuma H, Tamaoka A, Hasegawa K, Nonaka T, Hasegawa M, Kawata A, Yoshida M, Nakahata T, Takahashi R, Marchetto MC, Gage FH, Yamanaka S, Inoue H. Sci. Transl. Med. 2012;4:145ra104. doi: 10.1126/scitranslmed.3004052. [DOI] [PubMed] [Google Scholar]
- 18.Nicholas CR, Chen J, Tang Y, Southwell DG, Chalmers N, Vogt D, Arnold CM, Chen YJ, Stanley EG, Elefanty AG, Sasai Y, Alvarez-Buylla A, Rubenstein JL, Kriegstein AR. Cell Stem Cell. 2013;12:573–586. doi: 10.1016/j.stem.2013.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Maroof AM, Keros S, Tyson JA, Ying SW, Ganat YM, Merkle FT, Liu B, Goulburn A, Stanley EG, Elefanty AG, Widmer HR, Eggan K, Goldstein PA, Anderson SA, Studer L. Cell Stem Cell. 2013;12:559–572. doi: 10.1016/j.stem.2013.04.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu Y, Weick JP, Liu H, Krencik R, Zhang X, Ma L, Zhou GM, Ayala M, Zhang SC. Nat. Biotechnol. 2013;31:440–447. doi: 10.1038/nbt.2565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kim T-G, Yao R, Cho J-H, Vasudevan A, Koh A, Moon M, Datta D, Bolshakov V, Kim K-S, Chung S. Stem Cells. 2014;32(7):1789–1804. doi: 10.1002/stem.1704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Nestor MW, Jacob S, Sun B, Pre D, Sproul AA, Hong SI, Woodard C, Zimmer M, Chinchalongporn V, Arancio O, Noggle SA. Am. J. Physiol. Cell Physiol. 2015;308:C209–C219. doi: 10.1152/ajpcell.00263.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cambray S, Arber C, Little G, Dougalis AG, de Paola V, Ungless MA, Li M, Rodriguez TA. Nat. Commun. 2012;3:841. doi: 10.1038/ncomms1817. [DOI] [PubMed] [Google Scholar]
- 24.Beers J, Gulbranson DR, George N, Siniscalchi LI, Jones J, Thomson JA, Chen G. Nat. Protoc. 2012;7:2029–2040. doi: 10.1038/nprot.2012.130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Watanabe K, Ueno M, Kamiya D, Nishiyama A, Matsumura M, Wataya T, Takahashi JB, Nishikawa S, Nishikawa S, Muguruma K, Sasai Y. Nat. Biotechnol. 2007;25:681–686. doi: 10.1038/nbt1310. [DOI] [PubMed] [Google Scholar]
- 26.Chambers SM, Fasano CA, Papapetrou EP, Tomishima M, Sadelain M, Studer L. Nat. Biotechnol. 2009;27:275–280. doi: 10.1038/nbt.1529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chung S, Moon J-I, Leung A, Aldrich D, Lukianov S, Kitayama Y, Park S, Li Y, Bolshakov VY, Lamonerie T, Kim K-S. Proc. Natl. Acad. Sci. 2011;108:9703–9708. doi: 10.1073/pnas.1016443108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Baraban SC, Southwell DG, Estrada RC, Jones DL, Sebe JY, Alfaro-Cervello C, Garcia-Verdugo JM, Rubenstein JL, Alvarez-Buylla A. Proc. Natl. Acad. Sci. USA. 2009;106:15472–15477. doi: 10.1073/pnas.0900141106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Cunningham M, Cho J-H, Leung A, Savvidis G, Ahn S, Moon M, Lee KJ Paula, Han Jason J, Azimi N, Kim K-S, Bolshakov Vadim Y, Chung S. Cell Stem Cell. 2014;15:559–573. doi: 10.1016/j.stem.2014.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Martinez-Cerdeno V, Noctor SC, Espinosa A, Ariza J, Parker P, Orasji S, Daadi MM, Bankiewicz K, Alvarez-Buylla A, Kriegstein AR. Cell Stem Cell. 2010;6:238–250. doi: 10.1016/j.stem.2010.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tanaka DH, Toriumi K, Kubo K, Nabeshima T, Nakajima K. J. Neurosci. 2011;31:14116–14125. doi: 10.1523/JNEUROSCI.2786-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Braz JM, Sharif-Naeini R, Vogt D, Kriegstein A, Alvarez-Buylla A, Rubenstein JL, Basbaum AI. Neuron. 2012;74:663–675. doi: 10.1016/j.neuron.2012.02.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Houart C, Caneparo L, Heisenberg C, Barth K, Take-Uchi M, Wilson S. Neuron. 2002;35:255–265. doi: 10.1016/s0896-6273(02)00751-1. [DOI] [PubMed] [Google Scholar]
- 34.Nordstrom U, Jessell TM, Edlund T. Nat. Neurosci. 2002;5:525–532. doi: 10.1038/nn0602-854. [DOI] [PubMed] [Google Scholar]
- 35.Gunhaga L, Marklund M, Sjodal M, Hsieh JC, Jessell TM, Edlund T. Nat. Neurosci. 2003;6:701–707. doi: 10.1038/nn1068. [DOI] [PubMed] [Google Scholar]
- 36.Gunhaga L, Jessell TM, Edlund T. Development. 2000;127:3283–3293. doi: 10.1242/dev.127.15.3283. [DOI] [PubMed] [Google Scholar]
- 37.Germain ND, Banda EC, Becker S, Naegele JR, Grabel LB. Stem Cells Dev. 2013 doi: 10.1089/scd.2012.0264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Johnson MA, Weick JP, Pearce RA, Zhang S-C. J. Neurosci. 2007;27:3069–3077. doi: 10.1523/JNEUROSCI.4562-06.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Southwell DG, Nicholas CR, Basbaum AI, Stryker MP, Kriegstein AR, Rubenstein JL, Alvarez-Buylla A. Science. 2014;344:1240622. doi: 10.1126/science.1240622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Aguilar-Valles A, Luheshi GN. Psychoneuroendocrinology. 2011;36:634–648. doi: 10.1016/j.psyneuen.2010.09.006. [DOI] [PubMed] [Google Scholar]
- 41.Piper M, Beneyto M, Burne TH, Eyles DW, Lewis DA, McGrath JJ. Psychiatr. Clin. North Am. 2012;35:571–584. doi: 10.1016/j.psc.2012.06.002. [DOI] [PubMed] [Google Scholar]
- 42.Li Q, Cheung C, Wei R, Hui ES, Feldon J, Meyer U, Chung S, Chua SE, Sham PC, Wu EX, McAlonan GM. PLoS ONE. 2009;4:e6354. doi: 10.1371/journal.pone.0006354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Xu B, Ionita-Laza I, Roos JL, Boone B, Woodrick S, Sun Y, Levy S, Gogos JA, Karayiorgou M. Nat. Genet. 2012;44:1365–1369. doi: 10.1038/ng.2446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lin M, Pedrosa E, Shah A, Hrabovsky A, Maqbool S, Zheng D, Lachman HM. PLoS ONE. 2011;6:e23356. doi: 10.1371/journal.pone.0023356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Gilman SR, Chang J, Xu B, Bawa TS, Gogos JA, Karayiorgou M, Vitkup D. Nat. Neurosci. 2012;15:1723–1728. doi: 10.1038/nn.3261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kim TG, Yao R, Monnell T, Cho JH, Vasudevan A, Koh A, Peeyush KT, Moon M, Datta D, Bolshakov VY, Kim KS, Chung S. Stem Cells. 2014;32:1789–1804. doi: 10.1002/stem.1704. [DOI] [PMC free article] [PubMed] [Google Scholar]