Abstract
Exogenous and endogenous sources of chemical species can react, directly or after metabolic activation, with DNA to yield DNA adducts. If not repaired, DNA adducts may compromise cellular functions by blocking DNA replication and/or inducing mutations. Unambiguous identification of the structures and accurate measurements of the levels of DNA adducts in cellular and tissue DNA constitute the first and important step towards understanding the biological consequences of these adducts. The advances in mass spectrometry (MS) instrumentation in the past 2–3 decades have rendered MS an important tool for structure elucidation, quantification, and revelation of the biological consequences of DNA adducts. In this review, we summarized the development of MS techniques on these fronts for DNA adduct analysis. We placed our emphasis of discussion on sample preparation, the combination of MS with gas chromatography-or liquid chromatography (LC)-based separation techniques for the quantitative measurement of DNA adducts, and the use of LC-MS along with molecular biology tools for understanding the human health consequences of DNA adducts. The applications of mass spectrometry-based DNA adduct analysis for predicting the therapeutic outcome of anti-cancer agents, for monitoring the human exposure to endogenous and environmental genotoxic agents, and for DNA repair studies were also discussed.
1. Introduction
DNA adducts can be induced by a variety of endogenous or exogenous sources of chemical species, such as byproducts of lipid peroxidation,1,2 chemotherapeutic drugs,3–5 tobacco carcinogens,6–8 and environmental contaminants.9,10 If not repaired correctly or timely, these adducts may lead to myriads of alterations within the cell, including, but are not limited to, cell death, mutations in the genome, and aberrant cell cycle control.11
Genetic instability introduced by mutated genes may lead to cell dysregulation and dysfunction.12 Multiple lines of evidence support that mutations in relevant target genes, particularly oncogenes and tumor suppressor genes, are associated with the carcinogenic process.13,14 However, given that the repair efficiency and the adverse effects on the transmission of genetic information of a specific DNA lesion also depend on its chemical structure,15 the biological significance and health risks of DNA adducts may vary from one adduct to another.16–17 If aberrant DNA adduct formation is indispensable for genotoxicity and cancer initiation, quantitative assessment of DNA adduct accumulation would be of great importance in toxicological tests. The first key question to be addressed in such studies is to confirm the existence of covalent DNA adducts with the chemical or its metabolite in question. Secondly, direct measurement of DNA adducts is necessary for providing more accurate and reliable answers about whether the toxicity process has a safety margin and for assessing their consequences on human health. In this vein, DNA adducts can serve as biomarkers for environmental exposure and pathological conditions (e.g. oxidative stress), and form the basis for improving our understanding of the mechanisms of carcinogenesis as well as for enhancing the accuracy of risk assessment. Thirdly, it is important to reveal the location of adduct sites in DNA and to understand the site selectivity for DNA adduct formation, which are particularly needed for understanding the origins of mutational hotspots that are frequently found in some tumor suppressor genes in human cancers.18
As the interest in understanding the biological effects of DNA adducts on living organisms increases, the investigation has proceeded on the basis of more sophisticated test systems, from isolated DNA and cultured cells treated in vitro to experimental animals and humans exposed to environmental or therapeutic DNA damaging agents. Since DNA adducts are often present at very low levels in biological systems, considerable emphasis has been placed on the development of sensitive and specific methods for the detection and quantification of DNA adducts. Especially for in vivo studies with a limited amount of tissue sample available, the analytical method of choice must have adequate sensitivity for DNA adduct measurements.
Over the last three decades, a variety of analytical techniques have been used for DNA adduct analysis, including 32P-postlabeling assay,19 immunoassays,20 and electrochemical detection.21 In this vein, the development of the 32P-postlabeling method greatly enhanced the ability in detecting chemically modified DNA, owing to its high sensitivity (1 adduct in 1010 nucleotides) and low sample requirement (1–10 µg DNA).19,22 However, 32P-postlabeling, like any other analytical methods, has some disadvantages. The assay uses a radioactive isotope that poses health risks; thus, the method requires extra caution and a dedicated, isolated experimental environment. More importantly, the assay provides very limited structural information, which may lead to ambiguity in the identities of the DNA adducts under investigation. Recent developments in mass spectrometric methods for DNA adduct analysis have overcome these shortcomings. Despite possessing poorer sensitivity than the 32P-postlabeling method at the very beginning, the sensitivity of MS methods has improved rapidly as a consequence of continuous development in instrumentation and sample preparation methods. In light of these continuing advances, MS techniques have gained widespread applications in the qualitative and quantitative analysis of DNA adducts by providing sufficient sensitivity, specificity, and detailed structural information. For a detailed account of the advantages of MS over 32P-postlabeling assay for DNA adduct analysis, the readers should consult a recent review of this subject by Vouros and co-workers.23
The overall objective of this review is to summarize mass spectrometric methods for the structure elucidation, quantification, and the assessment about the biological consequences of DNA adducts. Focus will be placed on the utilization of MS for the characterization and quantification of DNA adducts, benefiting from two unique attributes of MS - unambiguous specificity and high sensitivity. While the subject of MS for the analysis of structurally modified DNA was recently reviewed,24 we placed more emphasis on sample preparation and the use of MS for understanding the biological consequences of DNA adducts.
2. Overview of reactive sites in DNA
Being held by complementary base pairing, the stable structure of duplex DNA relies mainly on the shape of nucleobases and the molecular electrostatic potentials of the nucleophilic sites on nucleobases and base pairs. The unique chemical properties of nucleobases also permit sequence-specific recognition of DNA by cellular proteins.25 Yet, for the same reason, the functional groups on nucleotides render DNA susceptible to modifications by electrophilic agents, which lie at the heart of understanding mutagenesis.11,26,27 Therefore, a better interpretation of the reactive sites in DNA provides a strong reference for the identification of potential DNA adducts and builds a solid foundation for the assessment of toxicity and health risk. Fig. 1 displays the reactive sites on nucleobases, 2-deoxyribose, and phosphate backbone of DNA where adducts are preferably formed. In this vein, it is worth noting that many DNA damaging agents can also induce damage to other types of biomolecules, including proteins.28,29
Fig. 1.
A summary of the reactive sites in DNA.
2.1 Reactive sites on nucleobases
The most extensively studied DNA adducts are generated through the covalent reactions between DNA bases and alkylating agents, the mode of which usually involves transferring alkyl groups from these agents to nucleophilic sites on nucleobases.30 So far numerous types of alkylating agents have been investigated, including methylating agents,15 ethylating agents,31 polycyclic aromatic hydrocarbons (PAHs),32 nitrosamines,33 aflatoxins,34 mustards,35 and haloalkanes.36 Studies of these DNA alkylation-inducing chemicals uncovered most ring nitrogen atoms as well as exocyclic oxygen and nitrogen atoms of nucleobases as targets for alkylation. These include N7, N3, N2, N1, and O6 positions of guanine, the N7, N6, N3, and N1 positions of adenine, the N3, N4, and O2 positions of cytosine, and the N3, O2, and O4 positions of thymine. In this context, alkylation may also result in the simultaneous modifications of two nucleophilic sites in a single nucleobase. For instance, etheno derivatives of guanine, adenine, and cytosine can arise from either the metabolic activation of exogenous chemicals (e.g. vinyl chloride37 or ethyl carbamate38) or endogenous metabolic processes (e.g. lipid peroxidation).1 Their targeting sites are almost all localized between the exocyclic nitrogen and one of its neighboring ring nitrogen atoms, which give rise to the formation of 1,N2- and 3,N2-ethenoguanine (εGua), 1,N6-ethenoadenine (εAde), and 3,N4-ethenocytosine (εCyt).39–41 In addition, as one of the most abundant products formed from lipid peroxidation, malondialdehyde (MDA) can also react with the N1 and N2 atoms of guanine to produce an exocyclic adduct, pyrimido[1,2-a]purin-10(3H)-one (M1G, structure shown in Fig. 2).42,43
Fig. 2.
The structures of some representative DNA adducts discussed in this review. “dR”; represents 2-deoxyribose, and the phosphate linking the 2-deoxyriboses in the CP-d(GpG) is omitted.
Therapeutic drugs benefit from their conjugation with reactive sites in DNA to produce monoadducts (MAs) and/or cross-links that can inhibit critical cellular processes in abnormally proliferating cells.44,45 For example, nitrogen mustards bind to the N7 position of guanine to yield monoadducts, and a second reaction with the N7 of guanine on the opposing DNA strand yields interstrand cross-links (ICLs).46 On the other hand, mitomycin C reacts almost exclusively with the N2 of guanines to form either MAs or ICLs at CpG sites.47 More complex cross-links are produced upon the exposure of DNA to platinum derivatives, a group of bioreductive prodrugs that target the N7 positions of guanine and adenine (the structure of a cisplatin-induced guanine adduct is shown in Fig. 2).48 Another example for therapeutic DNA binding occurs between psoralen derivatives and thymine, whose C5 and C6 atoms frequently serve as the reactive sites in the formation of cross-links.49
Different from DNA alkylation, a heterogeneous group of carcinogens can modify DNA via arylamination. This group includes, for example, aromatic amines (e.g. 4-aminobiphenyl, 4-ABP),50 nitroaromatic compounds (e.g. 2-nitrofluorene),51 and heterocyclic aromatic amines found in cooked fish and meats, such as 2-amino-3-methylimidazo[4,5-f]quinoline (IQ),52 2-amino-1-methyl-6-phenylimidazo[4,5-b]pyridine (PhIP),53 and 2-amino-3,8-dimethylimidazo-[4,5-f]quinoxaline (MeIQx).54 Although the sites of substitution on nucleobases vary substantially, it appears that the C8 atom and the exocyclic amino nitrogen of the purine bases, particularly of guanine, are the major targets for arylamination. Such a pattern is in stark contrast to that of alkylating agents which displayed lower efficiency in targeting the aforementioned two reactive sites.
The hydroxyl radical is one of the most reactive species formed from aerobic metabolism or from exposure to ionizing radiation. This radical can bind to pyrimidine bases with a preference for the C5 and C6 positions, generating C5–OH– and C6–OH-adduct radicals, respectively; it can also conjugate with purine bases, giving rise to C4–OH–, C5–OH–, and C8–OH-adduct radicals for both guanine and adenine55,56 as well as a C2–OH-adduct radical for adenine.57,58 Examples of hydroxyl radical-induced DNA adducts include 5-hydroxyuracil (5-OH-Ura), 5-hydroxycytosine (5-OH-Cyt), 5,6-dihydroxy-5,6-dihydrothymine (a.k.a. thymine glycol), and 8-oxo-7,8-dihydroguanine (8-oxo-Gua).59–62 Additionally, the hydroxyl radical can abstract a hydrogen atom from the 5-methyl group of 5-methylcytosine and thymine to yield the corresponding 5-hydoxymethyl, 5-formyl and 5-carboxyl derivatives of the two nucleobases.63–67 In this context, these oxidations of the 5-methyl group of 5-methylcytosine, mediated by the ten-eleven translocation (Tet) family of dioxygenases, have attracted great attention in recent years owing to their potential involvement in active cytosine demethylation in mammals.68–70
2.2 Reactive sites on the phosphate backbone and 2-deoxyribose
Whilst the majority of DNA adduct studies have focused on the interaction of reactive chemicals with nucleobases, there are also a number of investigations showing that the oxygen of the backbone phosphate group can serve as a reactive site, esterification of which leads to the formation of a phosphotriester. Phosphate adducts are known to be induced in DNA by a wide range of alkylating agents.31 By using two-dimensional NMR spectroscopy and mass spectrometry, Yates et al.71 characterized 2-cyano-2-hydroxyethyl phosphotriester (~25% modifications) as a product from cyanoethylene oxide-induced phosphate alkylation. Deforce et al.72 studied the formation of phosphotriester dinucleotide adducts following in vitro treatment of calf thymus DNA with the phenyl glycidyl ether, a compound used in the paint and resin industry. Haglund et al.73 reported the detection of ten different ethyl phosphotriester adducts on the basis of their accurate masses and product-ion spectra after treating DNA with N-ethyl-N-nitrosourea in vitro. Similar modifications of the phosphate backbone in DNA have also been detected in cells treated with cyclophosphamide.74
In contrast to nucleobases and the phosphate group, the 2-deoxyribose moiety is inert toward alkylating and arylaminating agents. However, the hydroxyl radical can abstract a hydrogen atom from each of the five carbon atoms on 2-deoxyribose, thereby giving rise to oxidized derivatives of 2-deoxyribose, epimeric lesions on 2-deoxyribose, DNA strand breaks, and/or DNA–protein cross-links.75–78
3. Qualitative analysis of DNA adducts
As a consequence of the various reactive sites present in DNA, genotoxic chemicals can bind covalently with DNA to give a diverse array of DNA adducts. Structural elucidation represents the initial step towards determining their biological consequences. Among the available methods for DNA adduct analyses, the needs of low microgram quantities of DNA and the superior sensitivity render the 32P-postlabeling method amenable for the detection of DNA adducts that are present at extremely low levels.19,22 The enzyme-linked immunosorbent assay (ELISA) is most suited for large-scale epidemiological studies or clinical evaluations as it is a rapid and cost-effective method that obviates the needs of radioisotopes.79 However, a significant disadvantage of these methods is the lack of structural specificity. Mass spectrometry, owing to its capability in measuring the m/z values for parent and fragment ions, offers information about the elemental compositions and structures of DNA adducts. The development process in the use of MS for DNA adduct identification began with the employment of MS coupled with electron impact (EI) ionization of DNA-derived samples by direct probe injection. Later, an improved method using gas chromatography (GC)-MS with EI or electron capture-negative ion chemical ionization (EC-NICI) after derivatization was developed. In recent years, electrospray ionization-MS (ESI-MS) and, to a lesser degree, matrix-assisted laser desorption/ionization (MALDI)-MS were frequently used. In this section, we briefly summarize these methods, with the focus of discussion being placed on those for the identification of nucleoside and nucleotide adducts, and for locating DNA adducts in oligodeoxyribonucleotides (ODNs).
3.1 Methodology
3.1.1 GC-MS
To confirm the identity of a DNA adduct, MS is often hyphenated with chromatographic techniques, GC and LC, two techniques that are complementary in separating DNA adducts with a broad range of polarity and volatility. Prior to the introduction of LC-MS, GC-MS has received widespread applications for the characterizations of target DNA adducts. The capillary column of online GC provides excellent resolution in separating volatile samples as they migrate through the column. Following EI or CI, the ionized molecules and their fragment ions with specific m/z are recorded by the mass analyzer. GC-MS offers rich structural information by providing mass measurements for fragment ions, and sometimes, the molecular ion.80 When combined with a high-resolution time-of-flight (TOF) mass analyzer, it can also offer exact masses for the ions detected.81 In this vein, the hyphenation of GC with MS allows for robust identification of DNA adducts. An early investigation employed GC-EI-MS to confirm the presence of benzo[a]pyrene diol epoxide (BPDE)–DNA adducts in 10 out of 28 human placentas, illustrating the capability of GC-MS in the structural elucidation of DNA adducts.82
Compared to the positive-ion EI mode, NICI possesses higher selectivity and superior sensitivity toward compounds with high electron affinities. Hence, it later overwhelmed EI in measuring such compounds using GC-MS. Methods using GC-NICI-MS are available for DNA adducts arising from exposure to lipid peroxidation by-products,83,84 bactericidal agents,85 tobacco smoke,86,87 and dietary intake such as PhIP.88 However, despite the advances made for GC-NICI-MS, its applications for carcinogen–DNA adduct measurements in biological matrices have been limited. This is mainly attributed to the need of derivatization to increase analyte volatility (vide infra).
3.1.2 LC-ESI-MS
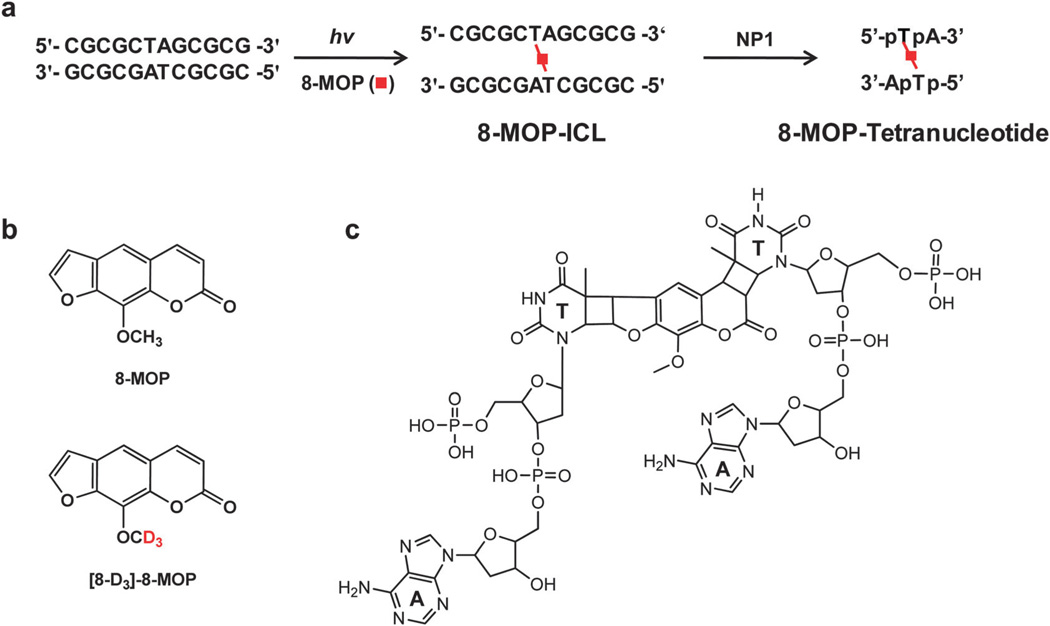
LC shares the same separation principle as GC with respect to eluting a pressurized sample mixture through a column, where the polarity of the sample affects elution time. Nevertheless, the separation achieved in the gas or liquid phase has brought in significant differences in the applications of these two techniques. Due to the use of a liquid mobile phase and a much shorter column mostly at room temperature, LC is more suitable for separating DNA adducts that are relatively more polar, less volatile and/or thermally labile without any derivatization. Thus, it distinguishes from GC as an efficient and cost-effective separation method. When coupled to MS, LC is particularly beneficial for providing an elution time characteristic of the analyte during separation as well as the capability in interfacing with various ionization methods, including the most commonly used ESI. Being a soft ionization method, ESI, mostly generating protonated or deprotonated molecules without complicated fragment ions, constitutes a straightforward method for the ionization of DNA adducts. Apart from modified nucleobases, LC-ESI-MS also allows for the detection of sugar–phosphate adducts and modified ODNs that are not amenable to GC-MS analysis.89–91 Owing to the highly acidic nature of the phosphate backbone, such analyses are commonly performed in the negative-ion mode. Moreover, the ability of ESI in producing multiply charged ions, in combination with improved mass range of modern mass analyzers, has facilitated the characterization of large analytes like ODNs. As an example depicted in Fig. 3a and 4a, the negative-ion LC-ESI-MS of a self-complementary duplex ODN d(CGCGCTAGCGCG) harboring an 8-methoxypsoralen (8-MOP)-ICL revealed ions of m/z 937.7, 1071.8, 1250.6, 1500.8, and 1876.3 as the intact duplex at different charge states (i.e. from 8- to 4-, respectively). In addition, the LC-ESI-MS of its nuclease P1 digestion product showed the [M − 2H]2− and [M − 3H]2− ions of a tetranucleotide consisting of two dinucleotides of d(pTpA) with the two thymines being bridged with an 8-MOP (Fig. 3c and 4b).92
Fig. 3.
Experimental outline of the enzymatic digestion of 8-MOP-ICL in ODNs (a) and chemical structures of 8-MOP and [8-D3]-8-MOP (b) and the tetranucleotide generated from nuclease P1 (NP1) digestion (c).
Fig. 4.
MS for 8-MOP-ICL-containing duplex ODN (a) and its digestion products (b) and MS/MS for 8-MOP–MA-housing single-stranded ODN (c).
3.1.3 CE-ESI-MS
While LC is the dominant chromatographic technique coupled to ESI-MS, capillary electrophoresis (CE), owing to its unparalleled separation efficiency and short separation time, has also found its use in coupling with ESI-MS for DNA adduct analysis. For instance, Deforce et al.72,93 and Barry et al.94 employed capillary zone electrophoresis (CZE)-MS for the analyses of DNA adducts formed from in vitro reactions with phenyl glycidyl ethers and BPDE, respectively. Despite with superb mass-detection limits, the major drawback of CZE-MS lies in the poor concentration-detection limits; this is mainly attributed to the very limited sample loading volume imposed by the CZE method (typically 5–10 nL), though this could be partially improved by using the sample stacking technique.95 With the resurging interest in CE-MS, we expect that the method will become more widely employed in DNA adduct analysis.
3.1.4 Tandem MS
As the molecular ion peaks exhibited in ESI-MS only offer molecular weight information about a DNA adduct, additional structural information of the adduct is often obtained through the analysis of fragment ions produced by collision-induced dissociation (CID). In CID, molecular ions with high kinetic energy collide with a neutral target gas, which converts some of the kinetic energy to internal energy and induces bond cleavages in the molecular ion to yield smaller fragments. While CID can occur in the electrostatic lens area, it is more widely conducted by incorporating a collision chamber to generate the so-called tandem MS. Tandem MS is a powerful method for identifying modified nucleosides. Chen et al.96–98 developed a series of tandem MS methods for the simultaneous characterization and determination of exocyclic DNA adducts derived from exogenous industrial chemicals and endogenous by-products of lipid peroxidation in human DNA. As an example, the LC-MS data indicated that two peaks eluting at 21–22 min on the LC system had a predictable m/z of 338, which correspond to the protonated ions of two diastereomers of 1,N2-propano-2’-deoxyguanosine adducts, namely, (6S,8S)-and (6R,8R)-3-(2′-deoxyribos-1′-yl)-5,6,7,8-tetrahydro-8-hydroxy-6-methylpyrimido[1,2-a]purine-10(3H)one (CrodG, Fig. 2). The identities of these two modified nucleosides were further confirmed by MS/MS analysis, where collisional activation of the molecular ion yielded fragment ions of [M + H – deoxyribose]+ ([BH]+), [BH – H2O]+, [BH – C2H4O]+, and [Gua + H]+.96
It is noteworthy that LC-MS and MS/MS can be used to verify the structures of unknown DNA adducts. For instance, because 2-hydroxyethyl-dA (HE-dA) adducts can be produced in the reaction of ethylene oxide (EtO) with dA at the N1 and N6 positions (structures shown in Fig. 2), two peaks eluting at ~10 and 16 min were observed as different HE-dA adducts, but both yielded [M + H]+ ions at m/z 296 and generated the same product-ion spectra.99 To distinguish these two adducts, Tompkins and colleagues99 first monitored the conversion of two peaks on LC after incubation of the sample in a NaOH solution, given that a strong base can promote the Dimroth rearrangement of an alkyl group from the N1 to N6 position. In addition, a more common method was used together with tandem MS analysis, where the structure could be established by comparison of chromatographic retention times and tandem mass spectra between a target analyte and its corresponding isotope-labeled standard. In this respect, the isotope-labeled standard for one of the HE-dA adducts, i.e. [15N5]-N1-HE-2′-dA, was synthesized and detected together with all the analytes. The similar retention time and tandem mass spectrum to those of N1-HE-dA can unambiguously distinguish it from N6-HE-dA. Alternatively, if the isotope-labeled standards are not available, a putative DNA adduct standard can be synthesized and analyzed in parallel with the analyte to establish whether the analyte exhibits identical retention time as well as similar MS and tandem MS as the synthetic standard.6,100 However, special attention needs to be paid in this case to avoid misidentification of analytes.
MS/MS in the typical product-ion scan mode provides a full-scan tandem mass spectrum which is very useful for the structural elucidation of modified nucleosides and nucleotides. However, MS/MS can also be conducted in a constant neutral-loss scan mode on a triple quadrupole or triple quadrupole linear ion trap mass spectrometer.101 The latter mode of analysis is particularly attractive for interrogating mixtures of DNA adducts because protonated ions of most 2′-deoxynucleosides exhibit facile neutral loss of a 2-deoxyribose during collisional activation, which may facilitate an “omics”; approach for DNA adduct analysis.102 This subject was discussed extensively in a recent review.103
In addition to analyzing single nucleoside or nucleotide adducts, the MS/MS method has also been widely employed for the characterizations of ODNs. With CID, sequence information can be derived from MS/MS acquired on various instrument platforms.34,104,105 While the generation of complementary fragment ion pairs renders simple interpretation of MS/MS data, the gentle activation induced by CID also minimizes further fragmentation of product ions. Under the collisional activation conditions, multiply charged ODNs are shown to have a strong tendency to cleave at the N-glycosidic bond between the nucleo-base and 2-deoxyribose and the 3′ C–O bond of the same nucleoside to form [an − base] and wn series of fragment ions (Fig. 5).106 In an approach for confirming the presence of an 8-MOP-DNA adduct in a synthesized ODN, a single-stranded ODN d(CGCGCTAGCGCG) harboring an 8-MOP–MA was subjected to LC-ESI-MS/MS analysis and the resulting MS/MS of the [M − 3H]3− ion (m/z 1286.2, Fig. 4c) provided valuable information that unambiguously validated the location of 8-MOP on the single thymine residue in the ODN.4 The m/z value observed for the w5 ion (m/z 1582) was identical to that found for the corresponding unmodified ODN, whereas the w72− ion (m/z 1207) displayed a mass increase of 216 Da as compared to that found in the spectrum of the unmodified ODN, suggesting the conjugation of 8-MOP with the thymine at the 6th position or the adenine at the 7th position counting from the 5′ terminus of the ODN. Meanwhile, the observed m/z value for [a5-C] (m/z 1333) is the same as that found for the corresponding unmodified ODN, whereas the [a7-A]2− ion (m/z 1071) displays an increase in mass that is consistent with the incorporation of an 8-MOP modification, supporting that 8-MOP is linked with the 5th cytosine or the 6th thymine in this ODN. Taken together, a conclusion can be made that 8-MOP is covalently bonded with the 6th thymine in this ODN. Apart from the 3′- and 5′-terminal (wn and [an – base]) ions resulting from CID, nozzle-skimmer and infrared multi-photon dissociation can also be employed to produce internal fragment ions and MS3 ions by further fragmentation of the wn or [an – base] ions. The combination of these dissociation methods with ESI-Fourier transform MS (FTMS) facilitated complete sequencing of a 50-mer DNA and extensive sequence verification for DNA containing up to 108 bases.107
Fig. 5.
Nomenclature of fragment ions for oligodeoxyribonucleotides.106 ‘B1’, ‘B2’, and ‘B3’ represent nucleobases, and ‘dR’ designates 2-deoxyribose.
ESI-MS/MS is not only useful for characterizing purified ODNs carrying a site-specifically incorporated DNA adduct, the method, when coupled with LC separation, has also been used for mixture analysis, particularly for the identification of the positional isomers of DNA adduct-containing ODNs108,109 and for the relative quantifications of the levels of adducts formed at different sites in synthetic ODNs.109 In addition, software tools have been developed for the automated processing of LC-MS/MS data for determining the sites of DNA adduction.110,111
Aside from the MS/MS methods described above, ESI-MS combined with exonuclease digestion has also been used to confirm the sequence and to identify the sites of modification of adducted ODNs. Several studies have used the so-called “ladder”; sequencing through coupling phosphodiesterase digestion with MS analysis. These exonucleases are able to hydrolyze the phosphodiester bonds in DNA sequentially from the 3′- or 5′ -terminus (Fig. 6a and b), and the m/z values of the gradually shortened products, an indicator of sequence information, can be obtained from MS analysis.112–114 For instance, Gupta et al.113 performed the partial enzymatic digestion of purified cisplatin–ODN adducts prior to LC-ESI-MS analysis to confirm the bifunctional binding sites of the metal to expected nucleobase sequences. As the period of digestion by 3′ → 5′ phosphodiesterase I prolonged (from 30 min to >5 h), ions were observed for the ODN remnants from the sequential loss of up to seven nucleotides from the 3′ end of cisplatin-adducted 16mer ODN (5′-CCTCTCCGGTCCTTCC-3′, where the sites of cisplatin conjugation were underlined), while other ions arising from the loss of more than seven nucleotides were absent. This indicated that the metal is covalently linked to the 9th guanine in this ODN and such DNA-platination prevents further hydrolysis of DNA at and beyond the modification sites. However, since ladder sequencing mainly relies on the activity of phosphodiesterases in digesting adducted DNA, it is often necessary to optimize the digestion conditions so as to obtain sequence ladders suitable for MS analysis. Critical factors affecting enzymatic digestion may include the length and base composition of DNA sequence of interest, the types of phosphodiesterases, the enzyme/substrate ratio, digestion time, and the pH of digestion buffers. Additionally, the digestion efficiency of adducted DNA also depends on the structure of adduct-inducing agents and the conformation of the adduct-containing DNA. Bulky DNA adducts that cannot fit into the active sites of enzymes may result in incomplete hydrolysis of the sequence, thereby providing mass spectral information that is different from the unmodified DNA sequence. In the aforementioned investigation by Gupta et al.,113 the inhibitory effect of the cisplatin adduct on 5′ → 3′ phosphodiesterase II was found to be so pronounced that it stopped hydrolysis before encountering the platination sites. After enzymatic digestion for up to 24 h, only ions produced by the removal of first 5–6 nucleotides from the 5′ side of cisplatin-conjugated 16mer ODN (5′-CCTCTCCGGTCCTTCC-3′) were observed in mass spectra, while the metal was bound to the 8th and 9th guanines on the remaining ODNs.
Fig. 6.
Application of exonuclease ladder digestion together with MALDI-MS to locate the site of a photoproduct in an ODN. Shown in (a) and (b) are schematic representations of ODN sequentially digested by 5′ - and 3′-exonucleases; and displayed in (c) and (d) are the time-dependent MALDI-MS of the corresponding digestion products. Panels (c) and (d) were adapted from Fig. 2 and 1 from Zhang et al117
3.1.5 MALDI-MS
MALDI, a technique using a pulsed laser to ablate and desorb molecules that are co-crystallized with a suitable light-absorbing matrix,115 is among the most common ionization methods for nucleic acid studies. Similar to ESI, MALDI is a soft ionization method, but it provides far fewer multiply charged ions in the gas phase than ESI. This latter attribute simplifies data interpretation and better facilitates mixture analysis. Additionally, it offers high-throughput and affords accurate mass measurements when coupled to a high-resolution TOF mass analyzer. While the advantages of MALDI-TOF-MS render it a useful tool for the analysis of high-molecular weight biomolecules such as proteins,116 it also allows for rapid determination of the accurate mass of modified ODNs as well as their sequences. For instance, Zhang and co-workers117 reported the combined application of MALDI-TOF-MS with exonuclease ladder digestion to characterize a 14mer ODN carrying a DNA photoproduct (Fig. 6). Interpretation of mass spectra obtained from independent digestion reactions by 5′- and 3′-phosphodiesterases enabled the unequivocal determination of the site of modification in this ODN. Likewise, MALDI-TOF-MS, together with exonuclease digestion, was employed to identify the sites of modifications in ODNs containing a site-specifically incorporated 2′-O-methyladenosine or various types of modified guanine derivatives.118,119
In addition to mapping an adduct-harboring DNA sequence, MALDI-MS analysis has also been employed to gain insights into the cleavage of a 33mer 5-OH-Cyt-containing ODN mediated by formamidopyrimidine DNA glycosylase (Fpg), an enzyme involved in base excision repair.60 The mass spectrum of the products resulting from Fpg cleavage showed fragments corresponding to ODNs generated by the expected β-δ-elimination mechanism of Fpg rather than by a β-elimination mechanism.
Together, the exonuclease ladder digestion, along with ESI- or MALDI-MS, constitutes a useful method that is alternative to the aforementioned MS/MS approach for locating the sites of modifications in synthetic ODNs. Because MALDI mainly produces singly charged ions, MALDI-MS is advantageous over ESI-MS in characterizing mixtures arising from the reaction with exonucleases or DNA repair enzymes, as the former provides simpler mass spectra that can be readily interpreted. Nevertheless, the MS/MS approach is more direct and efficient than exonuclease digestion coupled with the MS method.
3.2 Applications of qualitative MS analysis in biological studies
The last few decades have witnessed significant progress in our understanding of the biological consequences of DNA adducts. In addition to advances in molecular biology, this may also arise, at least partly, from the impressive improvement in analytical tools aimed at characterizing DNA lesions at nucleoside or nucleotide levels and in DNA segments. The structure elucidation of an unknown nucleoside or a nucleotide adduct achieved by chromatography coupled with tandem MS provides a solid foundation for further quantitative analysis of the adduct.53,120,121 In this vein, stable isotope-labeled standards often play an instrumental role in tandem MS-based qualitative analysis by possessing identical elution time with the analyte on chromatography as well as a similar fragmentation pattern in MS/MS, as detailed below.33,53,122 On the other hand, selective stable isotope incorporation into unique sites in the modified nucleosides, together with multi-stage MS analysis, facilitates the unambiguous elucidation of fragmentation pathways of modified nucleosides.61 Qualitative analysis for single nucleoside or nucleotide adducts are also used for the characterization of synthetic analytes and their corresponding minor modifications that may occur unexpectedly during synthesis and purification. For instance, Wu et al.123 carried out tandem MS analysis together with nuclease P1 digestion to identify unanticipated chemical modifications generated from the synthesis of fluorescein-labeled ODNs. The mass spectra of all four unexpected products indicated changes in the thiourea linkage [–NH–C(═S)–NH–] to the fluorescein moiety and the adjacent phosphate group, and accurate mass measurements with FTMS further confirmed their identities.
Driven by the optimal sensitivity and simple fragmentation pattern, DNA adducts are frequently detected as nucleosides or nucleotides. However, sequence information that is closely correlated with DNA adduct formation is lost upon digestion to individual nucleosides or nucleotides. It has been revealed that the reaction of carcinogens with DNA often exhibits sequence selectivity, the revelation of which could be important for understanding their mutagenic potentials.46,124 As discussed above, improved mass range in modern mass spectrometers and the capability in forming multiply charged ions during ESI have made possible the detection and sequencing of larger DNA adduct-bearing ODN fragments.
Apart from sequence confirmation and verification of site-specific incorporation of adducts into ODNs via MS and MS/MS, the combination of MS/MS-based ODN sequencing with molecular biology tools also allows for the examination of the biological consequences of DNA adducts, particularly how they compromise the efficiency and fidelity of DNA replication and transcription. For instance, LC-ESI-MS/MS was employed for studying in vitro translesion synthesis across various DNA lesions mediated by purified DNA polymerases.125–128
The study has also been extended to examine how DNA lesions inhibit DNA replication and transcription as well as induce mutations in cells, where LC-MS and MS/MS are employed to identify, and sometimes quantify, the mutagenic products arising from in vivo replicative or transcriptional bypass of DNA adducts.129–135 As illustrated in Fig. 7, a double-stranded plasmid harboring a site-specifically incorporated and structurally defined lesion is allowed to replicate in cultured human cells. The resulting progeny plasmid is isolated, amplified by polymerase chain reaction (PCR), and the PCR products are digested by two restriction enzymes and an alkaline phosphatase. The digestion products are then subjected to LC-MS and MS/MS analyses to sequence the ODN fragment housing the initially incorporated damage site, thereby facilitating the identification of mutations arising from replication across the lesion site. A representative MS/MS revealing the presence of the restriction fragment with a G → A mutation at the original S6-methylthioguanine (S6mG) site is shown in Fig. 8a.131
Fig. 7.
Restriction digestion coupled with LC-MS/MS for the identification of mutagenic products arising from replicative bypass of DNA lesions in cultured human cells. A lesion, designated with ‘X’, is placed downstream of an SV40 replication origin in a double-stranded plasmid, and the plasmid is allowed to replicate in a large T antigen-transformed cells. After replication, the progeny genome is purified, amplified by PCR, and digested with two restriction enzymes (SacI and FspI in this example) and alkaline phosphatase. The restriction recognition sites for SacI and FspI are highlighted in bold and restriction cleavage sites are indicated by a solid triangle. To distinguish the replication products from the lesion-containing strand and its complementary strand, an A:A mismatch (underlined) is placed two nucleotides away from the lesion site. MS/MS of the ‘M’- and/or ‘N’-containing restriction fragments allowed for the identification of the replication products of ‘X’ arising from cellular DNA replication, where ‘M’ designates the nucleobase inserted at the lesion site after cellular DNA replication, and ‘N’ denotes the nucleobase that is opposite to ‘M’ in the complementary strand. This figure was adapted from Fig. 3A of Yuan et al.131
Fig. 8.
Representative LC-MS and MS/MS for monitoring the restriction fragments of the S6mG-bearing substrate with or without a G → A mutation from in vivo replication and transcription. (a) MS/MS for the [M − 4H]4− ions of the wild-type sequence d(GCAAAGCTAGAGCT) (m/z 1072.7) and the mutant sequence d(GCAAAACTAGAGCT) (m/z 1068.7) arising from replication in 293T human kidney epithelial cells. Depicted in the insets are schemes summarizing the observed fragment ions. (b) High-resolution “ultra zoom-scan”; ESI-MS of the [M − 3H]3− ions of the wild-type sequence d(GCAAAGCTAGAGCT) (m/z 1430.5) and the mutant sequence d(GCAAAACTAGAGCT) (m/z 1425.2) arising from transcription in GM04429 human skin fibroblast cell lines. Panel (a) was adapted from Fig. S3 of Yuan et al.,131 and panel (b) was adapted from Fig. 3 of You et al135
Building upon the aforementioned replication and adduct bypass assay, You et al.134 recently developed a competitive transcription and adduct bypass assay, together with LC-MS, to investigate transcriptional alterations induced by DNA lesions in mammalian cells. In this respect, runoff transcripts generated from in vivo transcription of lesion-containing substrates are first reverse transcribed to produce cDNA and amplified by PCR. The resultant RT-PCR products are subsequently digested by appropriate restriction enzymes and analyzed on MS. As displayed in Fig. 8b, the identity of a G → A mutation is confirmed by high-resolution “ultra-zoom scan”; ESI-MS analysis of the restriction fragments released from the RT-PCR products arising from the transcription template housing an S6mG. Supported by this observation, the authors proposed that, when situated on the transcribed strand, S6mG exhibited strong mutagenic potential (i.e. with uridine misincorporation) during transcription in human fibroblast cells.135 In addition to qualitative analysis, these MS-based assays are also amenable for determining quantitatively the frequencies of mutations arising from the cellular DNA replication and transcription past the damage site. In this vein, it is worth noting that calibration curves of the relative quantifications of ODNs in the restriction digestion mixture are required since the ionization and fragmentation efficiencies vary with the sequences of the ODNs.127
4. Quantitative analysis of DNA adducts
Endogenous DNA adducts are often present at a level that is lower than 1/106 canonical nucleosides. This necessitates the development of highly sensitive and specific methods that require only a small amount of DNA. As technique improves, MS has played an increasingly important role in taking up this analytical challenge. In comparison with earlier successful platforms such as 32P-postlabeling, MS has advantages including the use of standards and capability of providing structural information on DNA adducts. To achieve highly sensitive detection, analytes are generally separated and/or enriched prior to MS analysis. The combination of MS with other techniques also enabled sensitive quantification of trace amounts of DNA adducts in a relatively small amount of DNA isolated from living organisms. Herein, we focus on sample preparation and mass spectrometric analysis, two important steps in the quantitative analyses of DNA adducts.
4.1 Sample preparation
4.1.1 DNA hydrolysis
The analysis of DNA adducts has been performed for DNA samples isolated from cultured cells,132,136 tissues,136–142 and biofluids.41,98,143 Apart from DNA, these samples also contain large amounts of salts, RNA, and proteins in the matrices that may participate in the metabolism and/or interfere with the detection of targeted DNA adducts. Thus, it is crucial to lyse the cells or tissues, isolate DNA, and break up long DNA molecules to analytes that are amenable for MS analysis. In this respect, the analysis of adducted DNA in the form of nucleobase, mononucleoside, or mononucleotide by MS offers the highest sensitivity and accuracy for the quantification of the majority of DNA adducts.
For nucleobase adducts with a destabilized N-glycosidic bond, neutral thermal hydrolysis is a simple method for their release from the DNA backbone. For instance, the N-glycosidic bonds of nucleosides carrying an N7-alkylguanine, N3-alkyladenine, or O2-alkylcytosine are susceptible toward hydrolysis.144,145 Therefore, selective cleavage of the N-glycosidic bonds in these labile alkylated nucleosides can be induced by simply heating the DNA at neutral pH. This sample preparation method has been employed for the quantification of a variety of DNA adducts, including N3-ethyladenine (N3-EtAde) and N7-ethylguanine (N7-EtGua) produced by ethylating agents in cigarette smoke,146 7-[4-(3-pyridyl)-4-oxobut-1-yl]guanine (7-POB-Gua) and O2-[4-(3-pyridyl)-4-oxobut-1-yl]cytosine (O2-POB-Cyt) arising from tobacco-specific nitrosamines,33 and N7-HE-Gua adducts formed from exposure to EtO.99 In addition, given that neutral thermal hydrolysis only breaks the N-glycosidic bonds between structurally altered nucleobases and 2-deoxyribose moieties, it provides a hydrolysate with the nucleobase adducts that can be simply isolated for subsequent determination by MS analysis. In this vein, if adduct recovery is acceptable with neutral thermal hydrolysis, the method is advantageous owing to its simplicity and specificity. Nevertheless, potential analyte loss from spontaneous elimination of the alkylated nucleobases during DNA extraction may compromise the accuracy of measurements.
The aforementioned neutral thermal hydrolysis has limited scope of applications. For the majority of DNA adducts that are chemically stable, quantification is often conducted after complete digestion of DNA to 2′-deoxynucleosides. In general, adduct-carrying DNA is first digested down to 2′-deoxynucleoside-5′-monophosphates. Among a number of digestion enzymes in this category, DNase I and nucleases are non-specific endonucleases that can cleave DNA into ODNs and mononucleotides.53,147,148 The subsequent use of phosphodiesterases results in further cleavage of ODNs into mononucleotides. The commonly used phosphodiesterases include spleen phosphodiesterase (phosphodiesterase II) and snake venom phosphodiesterase (phosphodiesterase I), which are 5′- and 3′-exonucleases, respectively.149,150 To obtain better sensitivity through analysis by MS in the positive-ion mode, conversion of nucleotides to nucleosides is often necessary, where alkaline phosphatase is frequently used.151–153
It should be noted that precautions are often needed during enzymatic hydrolysis to minimize errors if DNA lesions are unstable or can be produced artificially. For instance, N2-ethylidene-dG is an acetaldehyde-induced DNA lesion that is quite stable in DNA but rapidly decomposes at the nucleoside level.121 By adding NaCNBH3 during DNA extraction and enzymatic digestion, Wang et al.121 quantitatively converted N2-ethylidene-dG to the more stable N2-ethyl-dG and indirectly demonstrated the presence of this endogenous lesion in human liver DNA. Similarly, reduction with NaCNBH3 was employed during the quantification of formaldehyde-induced N2-hydroxymethyl-dG154 as the corresponding stable N2-methyl-dG. Among the primary or secondary DNA lesions produced by reactive oxygen species (ROS) and reactive nitrogen species (RNS), 8-oxo-7,8-dihydro-2′-deoxyguanosine (8-oxodG) has been studied most extensively in the past several decades. However, it is no longer broadly accepted as a biomarker for oxidative stress due to the fact that the levels of 8-oxodG reported in many previous studies may result from artifacts during DNA isolation and sample processing rather than from endogenous sources of ROS.155,156 Apart from ROS, nitric oxide (NO•), an important physiological messenger involved in cell signalling and an environmental pollutant, can induce deamination of nucleobases in vivo.157 The addition of radical scavengers, such as TEMPO (2,2,6,6-tetramethylpiperidine 1-oxyl), appears to inhibit artifactual deamination throughout sample preparation processes. Problems are also encountered in LC-MS analysis of deamination or other modification products of 2′-deoxyadenosine when deaminase is present in DNA samples158 or in commercial preparation of enzymes used for DNA hydrolysis.159 To avoid the adventitious deamination, a deaminase inhibitor [i.e. erythro-9-(2-hydroxy-3-nonyl)adenine] is often added during DNA digestion.140 Altogether, in cases where some adducts could form as artifacts during sample preparation, methods for DNA adduct measurement should be carefully validated, and the artificial formation or degradation of the analyte should be reduced to a level below the detection limit of the method for a successful analysis.
The efficiency of enzymes in hydrolyzing adducted DNA is also affected by the structure of adduct-inducing agents and the conformation of adduct-bearing DNA. Bulky DNA adducts that cannot fit into the active site of digestion enzymes may lead to incomplete hydrolysis. For instance, owing to distortion to the DNA helical structure, PhIP adducts (Fig. 2) are more efficiently hydrolyzed by nuclease P1 than micrococcal nuclease, where the active site of the latter enzyme may not be spacious enough to accommodate the lesion.160 Other DNA adducts, including dimeric DNA photoproducts,161,162 thymidine glycol,163 and DNA interstrand cross-link lesions,164 cannot be fitted into the active site of nuclease P1, which prevents the cleavage of the phosphodiester bond on the 3′ side of the modified nucleoside. While caution needs to be exerted to ensure complete digestion by enzymes during sample preparation, researchers also employed selective enzymatic digestion to produce analytes suitable for more sensitive and selective detection of DNA adducts by MS and for the assessment of isomeric DNA lesions produced at different sites. To improve the separation and detection of DNA ICLs introduced by nitrogen mustard melphalan, Mohamed and Linscheid165 used a combination of benzonase and nuclease S1, together with careful control of the digestion time, to release the cross-link moiety as a trinucleotide. Wang and co-workers4,92,166,167 also employed nuclease P1 to degrade ICL-containing DNA induced by psoralen derivatives to a tetranucleotide remnant for LC-MS and MS/MS analyses (Fig. 3a). Unlike releasing ICL-containing DNA as free base or nucleoside adducts that cannot be distinguished from intrastrand crosslinks, the formation of the ICL-bearing tetranucleotide provides unequivocal chemical specificity for the subsequent quantitative analysis as well as information about the identities of the flanking nucleosides of the cross-linked nucleosides. Based on the fact that nuclease P1 is incapable of cleaving the phosphodiester bond on the 3′ side of dimeric DNA photoproducts,162 Gross, Taylor, and co-workers168,169 assessed the DNA photoproduct formation at adjacent dithymidine sites and photo-crosslinking in human telomeric G-quadruplex DNA. In another approach, the ethyl phosphotriester dinucleotide adducts were selectively released from DNA by using nuclease P1 since the enzyme is incapable of hydrolyzing internucleotide bonds adjacent to a completely esterified phosphate group.73
An exception to DNA hydrolysis is shown by electrophilic adduct-inducing agents that can bind to nucleobases in DNA and destabilize the N-glycosidic bond. When the N-glycosidic bonds in these nucleosides become labile, depurination or depyrimidination occurs spontaneously on the adducted nucleosides; this hydrolysis process could also be mediated by base excision repair enzymes (i.e. DNA glycosylases) in vivo, resulting in the loss of the adducted base from DNA.170–172 To meet the requirement for detecting such compounds in biological specimens, sensitive MS methods for the measurement of unstable DNA adducts excreted in urine have been developed.173–175 These adducts include alkylated and oxidized 2′ -deoxynucleosides and nucleobases.176,177 In this vein, such a quantification obviates the need for DNA extraction and hydrolysis, but there are other steps to be particularly concerned about when handling urine samples, including sample storage, clean-up, and pre-concentration.178
4.1.2 Enrichment
Sample enrichment has been widely employed for the quantitative measurement of DNA adducts. Apart from adducts of interest, DNA hydrolysates also contain the bulk of unmodified nucleosides, proteins, inorganic salts, and other components which can interfere with MS analysis. As a result, extensive clean-up steps prior to MS analysis are typically required to further improve detection sensitivity. Meanwhile, following DNA hydrolysis, adduct isolation could greatly influence the ability of MS methods to detect DNA adducts. Examples of sample enrichment methods for DNA adduct analyses encompass liquid–liquid extraction (LLE), ultrafiltration, solid-phase extraction (SPE), on-line column switching, off-line high-performance LC (HPLC), and immunoaffinity chromatography. In LLE, DNA adducts are extracted with organic solvents, while other components of the biological matrix remain in the aqueous phase. For instance, N7-HE-Gua could be extracted using 1-butanol.179 This step is mainly used for the separation of analytes that are not polar or exhibit low polarity. A study has also found it essential for minimizing matrix suppression and achieving a lower limit of quantitation.180 However, the enrichment factor achieved by LLE is generally not high, nor is it good enough for enriching extremely low levels of DNA adducts. As a consequence of these limitations, the application of LLE is not as broad as the use of other enrichment methods, or it has to be combined with other purification procedures.
Ultrafiltration is the simplest pre-treatment tool that is often employed for purification and concentration of nucleic acids in a sample. By filtering DNA hydrolysate through a semi-permeable membrane, the disposable filtration unit only allows molecules of certain molecular weights to pass through. As filters with different molecular weight cut-offs (MWCOs) are commercially available, the use of filters with a lower MWCO can physically separate low molecular-weight species from proteins, DNA hydrolysis enzymes, and other macromolecules present in a given sample, and is therefore a fast and efficient method for enriching adducted nucleic acid. For example, Olsen et al.181 employed ultrafiltration to separate 2′-deoxyribonucleosides from the enzymatic digestion mixture of calf thymus DNA after exposure to glyoxal for sensitive determination of a glyoxal– DNA adduct. Aside from proteins and digestion enzymes, ultrafiltration can also remove the partially depurinated DNA backbone from which adducted bases are selectively released following the aforementioned neutral thermal hydrolysis.182 Despite being simple and rapid, ultrafiltration is always coupled with further clean-up procedures, such as SPE and off-line HPLC because of its relatively low efficiency and specificity in sample purification.
SPE has unique advantages in processing samples with high throughput and concentrating multiple types of DNA adducts simultaneously. Depending on the ratio of the highest sample volume applied on the cartridge over the lowest volume of an eluent, significant pre-concentration of the target adducts can be achieved. Although the resolution of the SPE column is lower than that of HPLC, the disposable SPE column is free of cross-contamination, and many samples can be processed in parallel. Thus far, SPE has been utilized for the enrichment of a number of adducted 2′-deoxyribonucleosides from DNA isolated from cells and tissues [e.g. N2-ethylidene-dG,121 M1G,183 and trans-2-hexenal-derived exocyclic 1,N2-propano-2′-deoxyguanosine (Hex-PdG)184] and adducted bases in urine (e.g. εAde).170 For more sensitive measurements of DNA adducts, the validation of SPE methods (with regard to the recovery, reproducibility, and clean-up efficiency) is necessary prior to mass spectrometric analyses. For example, method validation indicated that the recovery of Hex-PdG via SPE enrichment was not affected by the presence of DNA and no significant suppression of ionization was observed during the ESI process.184 In spite of a wide range of applications of SPE, this enrichment method has been restricted by sample quantity and adduct types. Chen et al.185 suggested that the recovery of glyoxal-induced cross-link decreases with reducing analyte quantity, even though the elution order on the SPE column does not change. For cisplatin DNA adducts, analyte recovery with SPE enrichment was ~20%, whereas clean-up utilizing HPLC increased the recovery to >90%.5
Column switching is another method that enables the trapping, enrichment and desalting of analytes prior to LC-MS/MS analysis. This is particularly important for nanoLC-MS analysis because column switching facilitates the injection of a relatively large volume of sample for on-line LC-MS analysis in the nano flow range.186 Special attention often needs to be paid toward the choice of trapping materials for the trapping column, where the analyte of interest should be efficiently trapped and subsequently eluted for online LC-MS analysis, as shown previously for analyses of peptides and modified ribonucleosides.187,188
Further separation of adducts from biological samples is also frequently achieved by off-line HPLC. When a DNA sample is digested to nucleosides and subjected to HPLC analysis, even if the adducts can escape detection by a conventional UV detector, the fractions known to contain specific adducts can be collected and analyzed by MS. A number of investigators have employed HPLC enrichment of DNA adducts (e.g. HE–DNA adducts99,179 and cisplatin 1,2-intrastrand guanine-guanine adduct5) for subsequent MS detection. Wang and co-workers132,136,140,189,190 also employed HPLC enrichment to improve the MS detection sensitivity and specificity of a number of DNA adducts, including carboxyalkylated DNA lesions, oxidatively induced 8,5′-cyclopurine-2′-deoxynucleosides, and oxidized derivatives of 5-methyl-2′-deoxycytidine and thymidine. Shown in Fig. 9 is a representative HPLC trace amount for the enrichment of 8,5′-cyclopurine-2′-deoxynucleosides in a nucleoside mixture of genomic DNA isolated from a rat liver. The (5′R) and (5′S) diastereomers of 8,5′-cyclo-2′-deoxyguanosine (cdG) and 8,5′-cyclo-2′-deoxyadenosine (cdA) can be readily resolved from each other and from the four unmodified canonical nucleosides. The elution orders of cdG, cdA, and the natural nucleosides on a reverse-phase column also reveal that these modified nucleosides cannot be simply enriched from the nucleoside mixture with the use of SPE. This offline HPLC enrichment is considered as one of the best options for isolating trace amounts of DNA adducts from DNA hydrolysates prior to MS analysis as it affords nearly quantitative analyte recovery, avoids the introduction of solid phase particles, which can clog nano-HPLC columns,191 and removes most unmodified canonical nucleosides as well as buffer salts employed in enzymatic digestion. However, when higher amounts of analytes are first injected to establish their elution times, the HPLC system can become contaminated. To avoid problems such as analyte carryover and cross-contamination, HPLC enrichment blank must be included prior to sample enrichment.
Fig. 9.
A representative HPLC trace for the enrichment of oxidatively induced cyclopurine lesions from the nucleoside mixture of genomic DNA isolated from mouse liver tissue. The (5′R) and (5′S) diastereomers of 8,5′-cyclo-2′-deoxyadenosine (R-cdA and S-cdA) and 8,5′-cyclo-2′-deoxyguanosine (R-cdG and S-cdG) can be resolved from each other and from the four canonical nucleosides. This figure was adapted from Fig. S3 of Wang et al140
Like HPLC, immunoaffinity chromatography is attractive for sample clean-up since a high separation resolution can be achieved with little or no effort required for optimizing the conditions. Antibodies developed against DNA adducts can be covalently bound to a matrix and the resulting material immobilized in columns is capable of binding specific adducts in a DNA digest. The combination of this technique with online LC-MS offers excellent selectivity for three reasons. First, the immunoaffinity purification step takes advantage of antibody’s specificity for target DNA adducts. Second, online LC analysis provides retention time that is characteristic of the DNA adducts. Finally, the mass spectrometer detects the precursor and product ions of m/z values that are unique for the analyte. The successful identification of different DNA adducts, e.g. εGua,37 alkyl-DNA,176 M1G,192 and ABP-C8-dG,193 supports the need for highly specific assays. Nevertheless, the development of this powerful technique has been hampered by the limited availability of antibodies that are suitable for target DNA adducts.
4.2 Mass spectrometric analysis
4.2.1 Standards for quantitative analysis
Reliable quantification of DNA adducts by MS requires accurate calibration of MS data using external or internal standards. For quantitation using external standards, the calibration normally involves a simple comparison of MS response (i.e. peak area or peak height) from a target DNA adduct with that from an exactly measured quantity of the identical analyte.194,195 Simplicity and applicability to a wide variety of methods constitute the main benefits of external standard calibration; however, its use is severely restricted by the drawback that this calibration could be greatly influenced by the stability of the MS detector as well as the sample matrix that may affect chromatographic separation, analyte ionization and precursor ion selection. As an example, Leclercq et al.196 constructed an external calibration curve by acquiring the LC-ESI-MS data for various quantities of N7-HE-Gua and later proceeding to signal integrations. They observed good linear correlation from 1 fmol to 1000 pmol of N7-HE-Gua; however, peak integration values corresponding to N7-HE-Gua exhibited a 20% day-to-day variation in the low pmol range. This reproducibility problem necessitated complete calibration prior to every LC-MS session. Chen et al.185 encountered a similar problem when they used external calibration to measure glyoxal-induced DNA cross-links on a capillary LC-ESI-MS/MS system. To overcome this, the authors constructed a standard curve each day before analyzing the samples and determined adduct levels of a sample on different days. Furthermore, they confirmed the accuracy of the assay by assessing the recovery of standard cross-links added to the hydrolysate of a known amount of DNA sample.185
The development of LC-MS has also facilitated the use of internal and co-chromatography standards, which is necessary for monitoring analyte recovery during the assay procedures. Additionally, the use of an internal standard removes the effects of a sample matrix on signal suppression and variation in equipment response among runs. Among a variety of approaches, incorporation of a stable isotope-labeled form of the analyte as the internal standard provides accurate and precise quantification by MS. Because these standards are chromatographically identical to, but different in mass from the DNA adducts of interest, their retention time on chromatogram and fragmentation pattern in tandem MS offer unequivocal specificity for the identification of DNA adducts. Stable isotope-labeled internal standards of nucleosides containing 2H, 15N, and/or 13C in nucleosides have been widely used for the quantification of DNA adducts.33,121,184,197 Alternatively, stable isotopes can also be incorporated into the chemical entity that is adducted to DNA. For instance, Cao et al.92 employed isotope-labeled [8-D3]-8-MOP-modified ODNs as the internal standard for quantitative analysis of 8-MOP-induced DNA ICLs in mammalian cells (the structure of [8-D3]-8-MOP is shown in Fig. 3b). Overall, the use of stable isotope-labeled standard provides unambiguous identification and accurate quantification of DNA adducts, and the method should be used whenever possible. Nevertheless, the isotope-dilution method also has some drawbacks. First, isotopically labeled adducts must be conveniently available, which limits the range of analytes that can be measured. Caution is also required for the use of some 2H-labeled standards due to the possible exchange of deuterium for hydrogen.198
In cases where the stable isotope-labeled DNA adducts are not used due to the low yield in their synthesis or the high cost of the isotope-labeled materials, a surrogate internal standard bearing a similar structure as the analyte can be employed for quantitative analysis. Along this line, Wang and co-workers199 recently developed an assay for the quantification of β-d-glucosyl-5-hydroxymethyl-2′-deoxyuridine (dJ) in Trypanosoma brucei DNA, where a surrogate standard, p-D-glucosyl-5-hydroxymethyl-2′-deoxycytidine, was employed to avoid pursuing a multi-step synthesis of an isotopically labeled dJ. From the data acquired on three separate days, excellent linearity (R2 = 0.998) was observed for the calibration curve. In another approach to detect methyl phosphotriester DNA adducts [dTp(Me)dT], thymidylyl(3′-5′)-thymidine ethyl phosphotriester [dTp(Et)dT] was selected as an internal standard for the same reason.200 Their data also revealed that the calibration curve obtained by LC-ESI-MS/MS was highly linear (R2 > 0.999) over the concentration range of 2–1000 ng mL−1.
4.2.2 Gas chromatography-mass spectrometry
Most online combinations of methods use conventional chromatographic separation as the first step to improve the specificity of detection. Since the introduction of GC in the 1950s, it has become a powerful separation technique which, when combined with MS, also provides excellent resolution, peak capacity, and structural information for volatile molecules. Even though GC is only compatible with volatile molecules, GC-MS has achieved widespread use for the analysis of different classes of DNA adducts. In the study of DNA alkylating agents, EtO is an important industrial chemical that forms DNA adducts without metabolic activation.201 By using a highly sensitive and specific GC/EC-CI-MS method together with the isotope-dilution technique, Kao et al.179 and Yong et al.202 reported the detection of a major EtO-induced adduct, N7-HE-Gua, in human granulocyte DNA. GC-MS in the NICI mode was also employed for the measurement of 4-hydroxy-1-(3-pyridyl)-1-butanone (4-HPB)-releasing DNA adducts formed from metabolites of the tobacco-specific nitrosamines 4-(methyl-nitrosamino)-1-(3-pyridyl)-1-butanone (NNK) and N′-nitrosonornicotine (NNN).8,87,203 Samples of human urinary bladder and lung DNA, hydrolyzed and subjected to GC-NICI-MS analysis, were shown to contain 4-ABP-DNA adducts at levels that are in keeping with 32P-postlabeling analysis of the same samples using appropriate standards.204,205 Additionally, the levels of MDA-induced M1dG were determined to be as low as 52 and 6.2 adducts per 108 nucleosides in rat liver83 and human leukocyte DNA,192 respectively, by the GC/EC-NICI-MS method.
Compared to coupling of MS with another conventional chromatographic separation technique – LC-MS, the growth of GC-MS for the detection of DNA adducts has been painfully slow. The primary factor limiting such growth is the indirect methods by which DNA adducts must be measured. Given that DNA adducts lacking volatility and thermal stability are not suitable for direct detection by GC-MS, one or more chemical derivatization steps are necessary as part of the sample preparation procedures. Chen and Chang206 compared the use of GC-MS and LC-ESI-MS/MS for the quantification of εAde in human urine samples. Their result revealed comparable levels of urinary εAde obtained by LC-ESI-MS/MS and GC-NICI-MS using the same isotopomer standard. Although the sensitivity of this GC-NICI-MS method was proven higher, partly due to the intrinsically lower separation efficiency of LC relative to GC with the use of capillary columns, a substantial decrease in time and cost of the reagents as well as the SPE columns required for the GC-NICI-MS analysis renders the LC-ESI-MS/ MS assay a more attractive method for measuring urinary εAde in a large number of samples. In addition, attention is needed toward a set of problems, including artifact production, interferences and sample-to-sample variation during derivatization.207
4.2.3 Liquid chromatography-mass spectrometry
LC-MS attempts to achieve three desirable attributes for DNA adduct analysis, namely, high sensitivity, low sample requirements, and structural identification. Since conventional-flow LC-MS was employed for the analysis of DNA adducts, a number of publications have demonstrated the increasing role that LC-MS would play in the characterization and detection of DNA adducts. Given that ESI produces ions in the gas-phase directly from a flowing liquid solution at a broad range of flow rates (typically in the range of 100–200 000 nL min−1), it became an ideal ionization technique used to interface with liquid-phase separation techniques. In the early 1990s, the hyphenation of LC with ESI-MS was first introduced for the measurement of DNA adducts induced by PhIP,208 chloroethylene oxide,207 bisphenol A,89 cyclic nitrosamine,209 etc. The set-up of LC-ESI-MS methodology opened a new door to DNA adduct analysis, leading to a dramatic increase in the utilization of MS for the detection and quantification of DNA adducts.
A summary highlighting sample requirements and sensitivities of various LC-MS platforms for the analyses of different DNA adducts is shown in Table 1. Overall, the development of LC-MS in the field of DNA adduct analyses has a trend towards a decline in sample consumption and an improvement in detection sensitivity. Tandem MS, as discussed above, constitutes a powerful tool for providing specific structural information about DNA adducts via diagnostic fragment ions. Such a specificity of adduct identification further helps increase the sensitivity of the method by lowering the background noise. Consequently, LC-MS/MS has become the method of choice for the unambiguous and simultaneous structural identification during quantification. An earlier platform used LC-MS/MS with conventional flow on 1–4.6 mm internal diameter (i.d.) columns for in vivo animal studies. For instance, Chen et al.120 used LC-ESI-MS/MS to confirm the endogenous presence of εAde in human placental DNA at a level of 2.3 adducts per 106 Ade bases. Gamboa da Costa et al.210 employed LC-ESI-MS/MS with a 2 × 150 mm reversed-phase column and a flow rate of 200 mL min−1 to determine two glycidamide (GA)-induced DNA adducts, N7-(2-carbamoyl-2-hydroxyethyl)guanine and N3-(2-carbamoyl-2-hydroxyethyl)adenine, in mouse tissues. The analytical method enabled the detection of adducts in liver, lung, and kidney of adult mice with the levels of the two lesions being approximately 200 and 2 adducts per 107 nucleotides, respectively.
Table 1.
Examples of LC-MS-based detection methods for DNA adducts
| Platform | DNA adduct | Sample req. | Sensitivity (on column) | Sensitivity (in DNA sample) | Year[Ref] |
|---|---|---|---|---|---|
| LC-ESI-MS | εGua | LOD (s/n = 2.5), 5 fmol | LOD (s/n = 3), 50 fmol | 1996207 | |
| LC-ESI-MS | N7-HE-Gua | 200 µg | LOD (s/n = 2), 1 fmol | 3 adducts/108 nucleotides | 1997196 |
| LC-ESI-MS | εAde | 0.1–0.3 mL urine | LOD (s/n = 3), 270 fmol | — | 199840 |
| LC-ESI-MS/MS | εAde | 1000 µg | 2.2 pmol (0.6 ng) | — | 1999120 |
| capLC-µESI-MS | IQ-C8-dG | 300 µg | LOD (s/n = 3), 6 fmol | — | 2001231 |
| LC-ESI-MS/MS | N3-GA-Ade | 100 µg | — | LOQ(s/n = 10), 1.5 adducts/ 108 nucleotides |
2003210 |
| LC-ESI-MS/MS | εAde | 1.0 mL urine | LOD (s/n = 3), 2 pg | 10 pg mL−1 | 2004206 |
| µLC-ESI-MS/MS | O6-POB-dG | 20–200 µg | LOD, 5 fmol | 50 fmol/1.5 mg DNA | 2004122 |
| LC-ESI-MS/MS | dG-N2-TAM | 10–160 µg | — | 8 adducts/108 nucleotides | 2005232 |
| LC-ESI-MS/MS | dTp(Me)dT | 300 µg | LOD (s/n = 3), 1.0 ng mL−1 | — | 2005200 |
| LC-ESI-MS/MS | Hex-PdG | 200 µg | — | 0.015 fmol mg−1 DNA (5 adducts/10 nucleotides) |
2006184 |
| capLC-ESI-MS/MS |
N2-Ethylidene-dG (N2-ethyl-dG by NaBH3CN) |
40–480 µg | LOD (s/n = 6), 0.05 fmol | 10 fmol mg−1 DNA | 2007233 |
| LC-ESI-MS/MS | HE–DNA adducts | 50–100 µg | LOD (s/n = 6), 0.5–25 fmol | — | 200899 |
| UPLC-ESI-MS/MS | CP-d(GpG) | 25 µg | LOQ, 3 fmol | 3.7 adducts/108 nucleotides | 20095 |
| capLC-nESI-MS/MS | Glyoxal–DNA adducts | 20 µg | LOD, 12–75 amol | — | 2009185 |
| capLC-nESI-MS/MS | ABP-C8-dG | 5 µg | LOD, 20 amol | 5 adducts/109 nucleosides | 2010215 |
| Column-switching | PhIP-C8-dG | 50 µg | LOD (s/n = 3), 2.5 fmol | 1.5 adducts/108 nucleosides | 201053 |
| LC-ESI-MS/MS | |||||
| nUPLC-MS/MS | N2-HmdG | 30 µg | LOD, 20 amol | — | 2011154 |
| nLC-nESI-HRMS/MS | N7-Ethyl-Gua | 188 ± 114 µg | LOD, 10 amol | 8 fmol µmol−1 guanine | 2011197 |
| UPLC-ESI-MS/MS | 7-OE-Gua | 50 µg | LOD, 1 fmol | — | 2012234 |
| UPLC-ESI-MS/MS | Acrolein–DNA adducts | 20 µg | LOD (s/n = 3), 40–80 amol | — | 2013235 |
| nLC-nESI-MS/MS | N3-Ethyl-Ade | 20 µg | LOD (s/n = 3), 15 fg (92 amol) | 9.4 adducts/109 nucleotides | 2014146 |
| N7-Ethyl-Gua | LOD (s/n = 3), 10 fg (56 amol) | 8.6 adducts/109 nucleotides | |||
| nLC-nESI-HRMS/MS/MS | EB-GII | 3–76 µg | LOD (s/n = 3), 0.05 fmol | 0.1 adducts/109 nucleotides | 2014182 |
Although tandem MS is highly specific, sensitivity of the assay is sometimes sacrificed due to the fragmentation of the parent ion to daughter ions. Moreover, the use of this technique in DNA adduct analysis has been limited because the levels of some DNA adducts in vivo are far below the limits of detection attainable by standard LC-ESI-MS/MS. To increase assay sensitivity for analytes of low abundance, it is a trend to use LC with a small diameter column at a low flow rate along with micro- and nano-ESI (µESI and nESI) sources.211,212 The recent emergence of these changes has resulted in profound decrease in the limits of detection for LC-MS/MS (Table 1). For instance, in analyses conducted on conventional bore 2 mm i.d. column, a well-studied nucleoside adduct, ABP-C8-dG, was reported at the levels of 4.9–30 adducts per 107 nucleosides in hepatic DNA isolated from mice treated with 4-ABP.50 By enzymatically hydrolyzing 100 µg of DNA, this assay achieved a detection limit of ~ 10 pg on-column, equivalent to 0.7 ABP-C8-dG in 107 normal nucleotides. Compared to columns with small i.d., the relatively large bore column used in this LC-MS method has limited analytical sensitivity. For the purpose of improving trace adduct detection in biological DNA, a capillary LC-µESI-MS/MS technique was later carried out on a 320 µm i.d. column to investigate the presence of ABP-C8-dG adducts in human pancreatic DNA, which yielded an LOQ approaching 1 adduct per 108 nucleotides by using only 13.3 µg of DNA per analysis.213 This amount was further decreased to 2.5 µg of DNA per analysis in a subsequent study using a capillary column with a smaller i.d. (75 µm i.d.) and a lower flow rate of 200 nL min−1 compared to the previous 20 µL min−1.214 Furthermore, Randall et al215 reported an integration of a 75 µm i.d. analytical column with online sample enrichment on a trapping column for sensitive quantification of ABP-C8-dG in 4-ABP-exposed human bladder cells and rat bladder tissues. This improved quantification method has a detection level of 5 adducts per 109 nucleosides with the use of 5 µg of DNA and the equivalent of only 1.25 µg of DNA per analysis. However, as significant as the improvement in sensitivity of LC-MS technique will be, there are going to be some drawbacks. In particular, nano-LC (nLC) separation has the limitations of lower optimum injection volume, lower column capacity, and shorter column lifetime as a consequence of having a high back pressure on the column after extended use. Therefore, extra care should be taken during sample handling.
Impressive improvement in overall sensitivity for the detection of trace levels of DNA adducts has also been achieved by coupling nLC-based separation methods with high-resolution MS (HRMS) and multi-stage MS (MSn). Sangaraju et al.182 found that the sensitivity of nLC-nESI-MS/MS was insufficient for the detection of 1,3-butadiene (BD)-DNA adducts in human samples (typical LOD, 0.5 adducts/106 nucleosides). To overcome this, the authors developed an nLC-nESI-HRMS3 method to decrease detection limits to the low fmol to amol range (1–10 per 109 nucleosides with 3–76 µg DNA) for their in vivo studies. The use of high-resolution MS for the analysis of DNA adducts in complex samples could dramatically reduce the matrix background, leading to greatly improved signal-to-noise (s/n) ratios.197,216 Likewise, the MSn scan mode also affords better s/n ratio owing to its high specificity, especially when there are co-eluting isobaric interferences. Meanwhile, the method allows for unambiguous identification of analytes of interest as detailed structural information is obtained from additional fragmentation pathways. Attracted by the analytical merits of multi-stage MS, LC-ESI-MS3 has been employed for the quantification of a number of DNA lesions including tobacco carcinogen- and cooked-meat carcinogen-induced DNA adducts, N2-(1-carboxyethyl)-2′-deoxyguanosine (N2-CEdG), 5-HmdU, 5-FodU, and purine cyclonucleoside in cellular and tissue DNA, which was shown to be more reliable than the MS2 mode for the analyses of these types of lesions.132,139,140,217,218 To illustrate this, Fig. 10 shows the comparison of MS/MS with MS/MS/MS for the quantification of an ionizing radiation-induced DNA lesion d(G[8-5m]T), where the C8 of the guanine is covalently bonded with the methyl carbon of its neighboring 3′ thymine.129 The selected-ion chromatogram obtained from MS/MS/MS provides much better s/n than that from MS/MS. In addition, aside from fragment ion expected for the analyte (m/z 472) and the stable isotope-labeled internal standard (m/z 475), MS/MS display many other fragment ions from co-eluting interferences. The MS/MS/MS, however, only show fragment ions anticipated for the analyte and internal standard.
Fig. 10.
LC-MS/MS/MS provides more robust quantification of an ionizing radiation-induced d(G[8-5m]T) lesion in DNA isolated from HeLa cells. Shown in (a) and (b) are the selected-ion chromatograms for monitoring the m/z 570 → 472 and m/z 573 → 475 transitions for the analyte and the stable isotope-labeled internal standard, respectively. Displayed in (c) and (d) are the selected-ion chromatograms for monitoring the m/z 570 → 472 → 276 and m/z 573 → 475 → 278 transitions for the analyte and the stable isotope-labeled internal standard, respectively. The corresponding MS/MS and MS/MS/MS are shown in the insets. The structures of the analyte and stable isotope-labeled standard and the cleavage pathways for the major fragment ions are shown in (e).
Among the several advantages, LC-tandem MS also permits the simultaneous detection of a large series of compounds and the separation of complex mixtures of diastereomers. Aldehydic compounds released during lipid peroxidation, e.g. 4-hydroxy-2-nonenal (HNE), can react predominantly with strongly nucleophilic ring nitrogens of DNA bases to yield adducts. In order to gain insights into the biological relevance of HNE-induced DNA lesions, Douki et al.195 used enzymatic hydrolysis combined with LC-MS/MS to determine the distribution of different possible exocyclic adducts [i.e. 1,N2-propano-2′-deoxyguanosine adduct of HNE (HNE-dG), M1dG, 1,N6-etheno-2′-deoxyadenosine (1,N6-εdA), 1,N2-etheno-2′-deoxyguanosine (1,N2-εdG), the (1,2-dihydroxy-heptyl)-substituted derivatives of 1,N6-εdA and 1,N2-εdG] in DNA isolated from HNE-treated human monocytes. Their quantification data revealed that the diastereomers of the HNE-dG adducts are the major DNA lesions induced in cells.
Other assays also used ultra-performance liquid chromatography (UPLC) coupled with tandem MS for rapid and sensitive analysis of DNA adducts (Table 1). UPLC operates with the use of sub-2 µm particles as packing materials in the analytical columns and specially designed instrumentation to withstand high pressure (6000–15 000 psi).219 In this vein, the UPLC-MS technique has gained increased popularity in DNA adduct analysis in recent years for the reason that it can provide a wider range of flow rates while maintaining good chromatographic resolution and therefore more rapid analysis compared to conventional HPLC. For instance, Feng and co-workers220 measured four diastereomers of anti-BPDE-N2-dG adducts on a UPLC-MS/MS system, where the separation time was shortened to 2–4 min and a low limit of detection of <0.7 fmol (s/n = 3) was achieved. Likewise, Baskerville-Abraham et al.5 reported the utilization of isotope-dilution UPLC-MS/MS for the accurate quantification of cisplatin-derived 1,2-guanine–guanine intrastrand cross-link [CP-d(GpG)] (structure shown in Fig. 2). Using a 2.1 × 100 mm UPLC column packed with 1.8 µm particles and a flow rate of 200 µL min−1, an LOQ of 3.7 adducts per 108 nucleosides was achieved starting with 25 µg of DNA. Although there have been a limited number of DNA adduct studies based on UPLC-MS, its importance will certainly grow owing to its advantages over conventional HPLC in terms of sensitivity, analytical speed, and sample consumption.
It is of note that, while LC-MS is advantageous over GC-MS in detecting DNA adducts directly, derivatization can still be employed to improve the ionization efficiency and/or the chromatographic separation of analytes during LC-MS analysis. In this regard, Hong et al.218 employed LC-ESI-MS/MS together with chemical derivatization for the highly sensitive detection of an oxidatively induced thymidine lesion, 5-formyl-2′-deoxyuridine (5-FodU). Given that the detection limit of 5-FodU was relatively poor compared with other oxidative DNA base damage, Hong et al.218 first derivatized 5-FodU with Girard’s reagent T (GirT) to form a hydrazone conjugate harboring a precharged quaternary ammonium moiety (Fig. 11a). This enabled the facile detection of the resulting conjugate by positive-ion ESI-MS with a pronounced increase in detection sensitivity (by ~20-fold compared with direct analysis of the underivatized compound). In addition, Tang et al.221 developed selective derivatization of cytosine moieties with 2-bromo-1-(4-dimethylamino-phenyl)-ethanone (BDAPE) coupled with LC-ESI-MS/MS for the determination of 5-methyl-2′-deoxycytidine (5-mdC), 5-hydroxymethyl-2′-deoxycytidine (5-HmdC), 5-formyl-2′-deoxycytidine (5-FodC), and 5-carboxyl-2′-deoxycytidine (5-CadC) in genomic DNA isolated from human colorectal carcinoma tissues (Fig. 11b). It turned out that the chemical derivatization notably improved the LC separation, and the detection sensitivities of 5-mdC, 5-HmdC, 5-FodC, and 5-CadC conjugated with BDAPE were increased by 35, 93, 89, and 123 fold, respectively. Although chemical derivatization has not been widely employed for the detection of DNA adducts by LC-MS, we expect to see more applications of this method for the quantitative assessment of certain DNA adducts, which may dramatically improve the detection sensitivity of trace-level analytes in a limited amount of DNA.
Fig. 11.
Representative derivatization reactions used in LC-MS analysis for DNA adducts. (a) Conjugation of 5-FodU with Girard reagent T. (b) Derivatization of 5-mdC, 5-hmdC, 5-fodC, and 5-cadC by 2-bromo-1-(4-dimethylamino-phenyl)-ethanone. In (b), R = CH3, CH2OH, CHO, and COOH.
4.3 Applications of quantitative MS analysis in biological studies
4.3.1 Quantitative analysis for biomarker discovery
Chemical modifications in DNA followed by cell proliferation can lead to mutations that may eventually result in cancer development. Biomarkers of carcinogenesis are essential for assessing the human health consequences of exogenous or endogenous exposure to DNA damaging agents. Such an exogenous exposure is hardly determined by external doses owing to the potential disposition of toxicants within an organism. Modulated by genetic and environmental factors, the patterns of absorption, distribution, metabolism, and excretion vary among different toxicants. To better predict the mutagenic and carcinogenic potentials of genotoxic agents, more suitable biomarkers are needed. Signatures of gene expression may provide insights into the mechanisms of toxicity and serve as biomarkers for toxicant classification, exposure monitoring, and risk assessment.222 However, to identify signatures of toxicity, mutagenicity or carcinogenicity, gene expression signatures must be anchored to phenotypic changes that are associated with a particular endpoint. On the contrary, as a well-established key step towards the onset of disease, DNA adducts are more proximal indicators for genotoxicity because their in vivo levels reflect exposure, metabolic activation, DNA adduction, and repair. Hence, the analysis of adducted DNA can elucidate mechanisms of action of chemical carcinogens in humans and identify risk-associated biomarkers for prediction of cancer development. Additionally, DNA adducts formed from endogenous sources are generally considered as end points of reaction; thus, monitoring their formation may provide a useful index of exposure to endogenous DNA damaging agents.183,206
The analytical challenge in the application of DNA adducts as biomarkers is the validation of a sensitive and selective method which could allow for the measurement of trace levels of modifications in a very low quantity of DNA. The method should also provide information about the adduct structure because the potential cytotoxic and mutagenic properties of a DNA adduct depend on its chemical structure. With the development of MS methodologies, which have enabled the routine detection of DNA adducts at a level of <1 lesion per 106 nucleosides, DNA adduct measurements have become more and more important in understanding the risk of carcinogen exposure. Furthermore, capabilities in quantitative analysis also form the basis for the implementation of new biomarkers.
Corroborative evidence pointing to the importance of DNA adducts in tobacco carcinogenesis includes numerous studies on establishing the association of tobacco smoke exposure with the induction of DNA adducts in humans. Ethylated DNA adducts could be induced by ethylating agents in cigarette smoke. By using nLC-nESI-MS/MS, Chen et al.146 reported a significantly higher level of N7-EtGua, but not N3-EtAde, in DNA from saliva of smokers than in non-smokers, suggesting that N7-EtGua in human salivary DNA might serve as a non-invasive biomarker for DNA damage induced by cigarette smoking. Being established as a carcinogen in humans, the tobacco-specific N-nitrosamine NNK, after metabolic activation, can add a POB moiety to DNA and the resulting unstable product can be further degraded to release 4-HPB.7 Quantitative analysis of 4-HPB-releasing DNA adducts uncovered these adducts as another robust biomarker to indicate the individual susceptibility to tobacco-induced cancers in humans.216
DNA adducts also serve as useful indicators of the relative contributions of different types of DNA adducts to carcinogenesis and the roles of metabolizing enzymes in carcinogenesis. In this respect, Monien et al.141 recently employed enrichment followed by UPLC-MS/MS analysis for monitoring simultaneously multiple DNA adducts derived from different genotoxic agents in DNA isolated from human lung biopsy specimens. Adducts derived from by-products of lipid peroxidation, furfuryl alcohol, and methyleugenol could be detected.141 As discussed previously, pyridyloxobutylating agents generated from metabolic activation of NNN and NNK by cytochrome P450 enzymes can alkylate DNA to produce POB–DNA adducts. An LC-ESI-MS/MS study revealed that O6-POB-dG was more abundant than other POB–DNA adducts in rat lung tissues, suggesting its important role in NNK-induced lung tumor in rats.33 Apart from serving as tissue-specific biomarkers, the presence of DNA adducts also reflects directly the metabolic activation of toxicants to form DNA-binding intermediates. In a UPLC-MS/MS-based approach, the stereoisomers of anti-BPDE-N2-dG were selected as biomarkers to assess the stereoselectivity of metabolic activation of B[a]P in human lung cells.220 Using LC-MS/MS, Nelson et al.223 confirmed the bioactivation of PhIP to yield N-hydroxy-PhIP (N-OH-PhIP) as well as the formation of PhIP–DNA adducts in human prostate and human prostate-derived cells, which provided insights into the potential role of PhIP exposure in early-stage prostate carcinogenesis. The observation of a lower PhIP-adduct level in glutathione S-transferase π-overexpressing cells also provided insights into the role of this enzyme in protecting against early-stage prostate carcinogenesis in humans.
The concentration of a drug-induced DNA adduct in cancer cells can also be used as a biomarker for the prognosis of therapeutic outcome in cancer patients. In this respect, accelerator MS (AMS) distinguishes itself from other MS methods owing to its extremely high sensitivity, specificity, and precision for measuring rare isotopes, such as 14C, in biological samples.224 Although limited by the need for radiolabeled precursors, this technique has demonstrated its capability in monitoring drug– DNA adducts and in determining drug sensitivity in different models at a nontoxic, subpharmacological concentration of the drug. For example, HPLC separation followed by AMS analysis was employed for determining the distribution of [14C]oxaliplatin-induced DNA adducts in drug-sensitive testicular and drug-resistant breast cancer cells at a clinically relevant drug concentration.225,226 The observation of differential adduction of the drug in cellular DNA supported the marked differences in abilities of different types of cancer cells in tolerating platinum chemotherapy. The development of such biomarkers for determining tissue-specific molecular dosimetry during treatment will lead to a more complete understanding of the therapeutic efficacy and side effects of chemotherapeutic drugs.5
4.3.2 Quantitative analysis for DNA repair study
There are a variety of DNA repair pathways for removing DNA adducts from the genome.11 Failure in repair or inaccurate repair may result in adverse biological consequences. A predominant pathway leading to mutation is structural modifications resulting in aberrant base pairing during DNA replication. Structural modifications of DNA bases (e.g. DNA adducts caused by ROS) bring not only immediate, but also long-lasting impacts on the cell. Slow but constant accrual of non-lethal genetic changes in DNA has been considered as key events leading to chronic genetic diseases, including cancer and aging. While the formation of only a few unrepaired DNA adducts among millions of normal DNA bases can result in significant biological consequences, measuring such a small difference requires analytical methods with high accuracy and sensitivity.
Improvements in MS-based analytical methods for measuring trace levels of DNA adducts, together with genetic manipulation, have offered an opportunity to assess the DNA repair process by monitoring directly the levels of DNA adducts. By using HPLC-ESI-MS/MS, Liu et al.167 recently studied the repair of DNA ICL and MAs introduced by 8-MOP in cultured mammalian cells. The detection limit was found to be as low as 3 fmol, which facilitated the observation of a rapid clearance of 8-MOP-ICL from cellular DNA 24 h post-irradiation. When combined with genetic depletion of DNA repair proteins, MS-based DNA adduct quantification also provides a method for pinpointing the role of a specific DNA repair pathway associated with the repair factors in removing the DNA adduct of interest. For instance, the authors also found that the progressive decrease in the 8-MOP-induced ICL and MAs were abolished in cultured mammalian cells deficient in XPA or ERCC1, suggesting that 8-MOP photoadducts are substrates for nucleotide excision repair (NER) in mammalian cells.167 Likewise, Wang and co-workers137,139 found that deficiency in ERCC1 led to markedly elevated accumulation of cdA and cdG in mouse tissues, thereby providing direct evidence to support the involvement of ERCC1, and thus the NER pathway, in repairing the endogenously induced purine cyclonucleosides in vivo. Similarly, Malayappan et al.227 used LC-MS/MS to follow the formation and repair of cyclophosphamide-induced N,N-bis[2-(N7-guanyl)ethyl]amine cross-link in human blood. The use of a LC-MS/MS-based assay also led to the observation that ICLs induced by mitomycin C could be partially removed in mouse mammary tumor cells.228
5. Concluding remarks and future perspectives
In this review, we summarized the commonly used MS methods for DNA adduct analysis and discussed the applications of these methods. The detection sensitivity of MS has greatly increased over the past 25 years, and additional improvements are expected. Initially GC-MS constituted the most sensitive method for DNA adduct detection; the method, however, necessitates derivatization to render the analytes volatile and thus amenable for GC separation. The development of ESI facilitated the direct coupling of HPLC with MS and enabled the direct measurement of polar compounds without derivatization. GC- and LC-MS techniques are particularly advantageous in elucidating the chemical structures of DNA adducts being analyzed and achieving highly accurate quantification with the addition of standards for calibration.
Furthermore, the recent application of LC with a small diameter column coupled with µESI- or nESI-MS systems facilitates quantitative analysis of DNA adducts with the use of low microgram quantities of DNA. While DNA adducts are widely characterized and quantified at the nucleoside or nucleotide level, sequence information significant in determining the selectivity of adduct formation in synthetic ODNs and in confirming site-specific incorporation of DNA adduct into ODNs can also be acquired using tandem MS or, when combined with exonuclease digestion, MALDI-TOF-MS.
In the coming years, developments in MS instrumentation and online separation methods are expected to improve the detection sensitivity and specificity while reducing the amount of sample required. Additionally, both sample preparation methods and instrumentation need to be pushed to achieve the lower detection limit and higher specificity, especially for extreme trace analysis of DNA adducts in vivo. In this vein, the recently introduced quadrupole-Orbitrap type of hybrid mass analyzer (i.e. the QExactive series of instruments) has demonstrated its success in the highly sensitive and specific quantification of peptides, lipids, and small molecules. Owing to its capability in accurate mass measurements in both MS and MS/MS and its high sensitivity, we expect to observe the increased application of these types of instruments in DNA adduct analysis. In addition, because of its high separation speed and resolution, UPLC is anticipated to be more widely used as the front-end separation technique for LC-MS-based DNA adduct analysis. The improvement in detection limit provided by the new sample preparation methods and new instrumentation will enable the monitoring of DNA adducts in very limited amounts of cell or tissue samples, e.g. in cancer stem cells or in specific regions of the brain or other tissues. Such an improvement will also facilitate future quantitative assessment of those “minor”; DNA adducts that are formed at low frequency but may foster more adverse biological consequences.
Future improvements will also extend MS techniques to the identification and/or quantification of DNA adducts emanating from a broader range of novel environmental and dietary carcinogens, the discovery of which may provide useful biomarkers for toxicological and/or epidemiological studies as well as for biomonitoring and early diagnosis of disease in humans. While the majority of DNA adduct analyses has focused on DNA samples from cultured cells and frozen or fresh tissues, not much work has been conducted for formalin-fixed human tissues. It was demonstrated recently that DNA could be retrieved from such tissue samples for DNA adduct analysis.142 Since large collections of such tissues are available, further developments in DNA isolation from fixed tissues will enable systematic investigation about the implications of DNA adducts in the development of human diseases.
MS, in conjunction with the molecular biology-based method, has recently been employed for understanding, at the molecular level, how structurally defined DNA adducts compromise the flow of genetic information by inhibiting DNA replication and transcription and elicit mutations in these processes. Likewise, the combination of DNA adduct quantification with the manipulations of the expression or activities of DNA repair genes has been demonstrated to be capable of offering important insights about how structurally defined DNA lesions are repaired in mammalian cells and tissues. Further applications of MS for studying the biological consequences and repair of DNA adducts will drive the DNA adduct field forward by providing molecular underpinnings of the implications of DNA adducts in the etiology of cancer and other human diseases, e.g. by providing a better understanding about the relationship between DNA adduct formation and mutation induction. Such knowledge may also give rise to the development of novel strategies for cancer chemoprevention (i.e. through inhibition of DNA adduct formation or stimulation of DNA repair).
Considering that DNA damaging agents constitute a major class of agents for cancer therapy,44,45 a better understanding about how structurally defined DNA adducts perturb the transmission of genetic information and how they are repaired will also guide the development of better cancer chemotherapeutic agents. In this vein, it is desirable to develop chemotherapeutic agents that are cytotoxic, but induce minimal mutagenic effects so that the therapeutic benefits can be maximized whereas the side effects can be minimized. In addition, understanding the molecular mechanisms of repair of DNA adducts induced by chemotherapeutic agents may also promote the development of agents that can inhibit DNA repair, which may facilitate the development of novel combination therapy for cancer treatment. Along this line, inhibitors of a DNA repair protein, poly(ADP-ribose) polymerase (PARP), are known to improve the therapeutic outcome of cancer patients carrying mutations in BRCA1 or BRCA2 gene.229,230
Together, MS has played a significant role in many aspects of DNA adduct research, including structural elucidation, quantification, as well as the revelation of the biological consequences and repair of DNA adducts. We expect that MS will continue to assume a crucial role in this important area of research.
Abbreviations
- MS
Mass spectrometry
- EI
Electron impact
- EC
Electron capture
- NICI
Negative ion chemical ionization
- CI
Chemical ionization
- ESI
Electrospray ionization
- GC
Gas chromatography
- LC
Liquid chromatography
- UPLC
Ultra-performance liquid chromatography
- TOF
Time-of-flight
- CID
Collision-induced dissociation
- FTMS
Fourier transform
- MS
SPE Solid-phase extraction
- µESI
Micro-flow electrospray ionization
- nESI
Nano-flow electrospray ionization
- AMS
Accelerator MS
Biographies

Shuo Liu
Shuo Liu completed her BS degree (2007) and MS degree (2010) in Environmental Science at Nankai University, China. She is currently a PhD candidate in the Environmental Toxicology Graduate Program at the University of California, Riverside, under the supervision of Prof. Yinsheng Wang. At the time of writing, Shuo is close to completing her dissertation research project, which involves the development of mass spectrometric methods for the characterization and quantification of different types of structurally modified DNA. She has also been focusing on the application of these methods in assessing the biological effects of DNA adducts in various living organisms.

Yinsheng Wang
Yinsheng Wang was born in 1971. He joined the faculty of the University of California Riverside after receiving his PhD in Chemistry from Washington University in St. Louis in 2001, under the guidance of Professors Michael L. Gross and John-Stephen A. Taylor. His main research interests include the assessment of the occurrence, biological consequences and repair of DNA damage along with the functional characterization of post-translational modifications of proteins. He was the recipient of the inaugural Chemical Research in Toxicology Young Investigator Award in 2012 and the 2013 Biemann Medal, and he was elected as an AAAS Fellow in 2012.
References
- 1.Sun X, Nair J, Linseisen J, Owen RW, Bartsch H. Int. J. Cancer. 2012;131:1983–1990. doi: 10.1002/ijc.27479. [DOI] [PubMed] [Google Scholar]
- 2.Nair U, Bartsch H, Nair J. Free Radical Biol. Med. 2007;43:1109–1120. doi: 10.1016/j.freeradbiomed.2007.07.012. [DOI] [PubMed] [Google Scholar]
- 3.Gamboa da Costa G, Pereira PC, Churchwell MI, Beland FA, Marques MM. Chem. Res. Toxicol. 2007;20:300–310. doi: 10.1021/tx600275d. [DOI] [PubMed] [Google Scholar]
- 4.Lai C, Cao H, Hearst JE, Corash L, Luo H, Wang Y. Anal. Chem. 2008;80:8790–8798. doi: 10.1021/ac801520m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Baskerville-Abraham IM, Boysen G, Troutman JM, Mutlu E, Collins L, Dekrafft KE, Lin W, King C, Chaney SG, Swenberg JA. Chem. Res. Toxicol. 2009;22:905–912. doi: 10.1021/tx800481j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Upadhyaya P, Sturla SJ, Tretyakova N, Ziegel R, Villalta PW, Wang M, Hecht SS. Chem. Res. Toxicol. 2003;16:180–190. doi: 10.1021/tx0256376. [DOI] [PubMed] [Google Scholar]
- 7.Sturla SJ, Scott J, Lao Y, Hecht SS, Villalta PW. Chem. Res. Toxicol. 2005;18:1048–1055. doi: 10.1021/tx050028u. [DOI] [PubMed] [Google Scholar]
- 8.Heppel C, Heling AK, Richter E. Anal. Bioanal. Chem. 2009;393:1525–1530. doi: 10.1007/s00216-008-2566-y. [DOI] [PubMed] [Google Scholar]
- 9.Lai Y, Lu M, Gao X, Wu H, Cai Z. Environ. Sci. Technol. 2011;45:10720–10727. doi: 10.1021/es203068f. [DOI] [PubMed] [Google Scholar]
- 10.Leuratti C, Jones NJ, Marafante E, Kostiainen R, Peltonen K, Waters R. Carcinogenesis. 1994;15:1903–1910. doi: 10.1093/carcin/15.9.1903. [DOI] [PubMed] [Google Scholar]
- 11.Friedberg EC, Walker GC, Siede W, Wood RD, Schultz RA, Ellenberger T. DNA Repair and Mutagenesis. Washington, D.C: ASM Press; 2006. [Google Scholar]
- 12.Bohr VA. Carcinogenesis. 1995;16:2885–2892. doi: 10.1093/carcin/16.12.2885. [DOI] [PubMed] [Google Scholar]
- 13.Poirier MC, Beland FA. Environ. Health Perspect. 1994;102(suppl 6):161–165. doi: 10.1289/ehp.94102s6161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wiencke JK. Oncogene. 2002;21:7376–7391. doi: 10.1038/sj.onc.1205799. [DOI] [PubMed] [Google Scholar]
- 15.Wyatt MD, Pittman DL. Chem. Res. Toxicol. 2006;19:1580–1594. doi: 10.1021/tx060164e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Nestmann ER, Bryant DW, Carr CJ. Regul. Toxicol. Pharmacol. 1996;24:9–18. doi: 10.1006/rtph.1996.0059. [DOI] [PubMed] [Google Scholar]
- 17.Vineis P, Perera F. Int. J. Cancer. 2000;88:325–328. [PubMed] [Google Scholar]
- 18.Pfeifer GP, Tang M, Denissenko MF. Curr. Top. Microbiol. Immunol. 2000;249:1–19. doi: 10.1007/978-3-642-59696-4_1. [DOI] [PubMed] [Google Scholar]
- 19.Randerath K, Reddy MV, Gupta RC. Proc. Natl. Acad. Sci. U. S. A. 1981;78:6126–6129. doi: 10.1073/pnas.78.10.6126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hsu IC, Poirier MC, Yuspa SH, Grunberger D, Weinstein IB, Yolken RH, Harris CC. Cancer Res. 1981;41:1091–1095. [PubMed] [Google Scholar]
- 21.Park JW, Cundy KC, Ames BN. Carcinogenesis. 1989;10:827–832. doi: 10.1093/carcin/10.5.827. [DOI] [PubMed] [Google Scholar]
- 22.Phillips DH, Arlt VM. Nat. Protoc. 2007;2:2772–2781. doi: 10.1038/nprot.2007.394. [DOI] [PubMed] [Google Scholar]
- 23.Klaene JJ, Sharma VK, Glick J, Vouros P. Cancer Lett. 2013;334:10–19. doi: 10.1016/j.canlet.2012.08.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tretyakova N, Villalta PW, Kotapati S. Chem. Rev. 2013;113:2395–2436. doi: 10.1021/cr300391r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rohs R, Jin X, West SM, Joshi R, Honig B, Mann RS. Annu. Rev. Biochem. 2010;79:233–269. doi: 10.1146/annurev-biochem-060408-091030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Burcham PC. Mutagenesis. 1998;13:287–305. doi: 10.1093/mutage/13.3.287. [DOI] [PubMed] [Google Scholar]
- 27.Marnett LJ. Carcinogenesis. 2000;21:361–370. doi: 10.1093/carcin/21.3.361. [DOI] [PubMed] [Google Scholar]
- 28.Berlett BS, Stadtman ER. J. Biol. Chem. 1997;272:20313–20316. doi: 10.1074/jbc.272.33.20313. [DOI] [PubMed] [Google Scholar]
- 29.Liebler DC. Chem. Res. Toxicol. 2008;21:117–128. doi: 10.1021/tx700235t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sedgwick B, Bates PA, Paik J, Jacobs SC, Lindahl T. DNA Repair. 2007;6:429–442. doi: 10.1016/j.dnarep.2006.10.005. [DOI] [PubMed] [Google Scholar]
- 31.Singer B. Environ. Health Perspect. 1985;62:41–48. doi: 10.1289/ehp.856241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Rybicki BA, Nock NL, Savera AT, Tang D, Rundle A. Cancer Lett. 2006;239:157–167. doi: 10.1016/j.canlet.2005.07.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lao Y, Villalta PW, Sturla SJ, Wang M, Hecht SS. Chem. Res. Toxicol. 2006;19:674–682. doi: 10.1021/tx050351x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Marzilli LA, Wang D, Kobertz WR, Essigmann JM, Vouros P. J. Am. Soc. Mass Spectrom. 1998;9:676–682. doi: 10.1016/S1044-0305(98)00039-7. [DOI] [PubMed] [Google Scholar]
- 35.Batal M, Boudry I, Clery-Barraud C, Mouret S, Douki T. Toxicol. Environ. Chem. 2013;95:260–276. [Google Scholar]
- 36.Huang H, Jemal A, David C, Barker SA, Swenson DH, Means JC. Anal. Biochem. 1998;265:139–150. doi: 10.1006/abio.1998.2891. [DOI] [PubMed] [Google Scholar]
- 37.Morinello EJ, Ham AJL, Ranasinghe A, Nakamura J, Upton PB, Swenberg JA. Cancer Res. 2002;62:5189–5195. [PubMed] [Google Scholar]
- 38.Fernando RC, Nair J, Barbin A, Miller JA, Bartsch H. Carcinogenesis. 1996;17:1711–1718. doi: 10.1093/carcin/17.8.1711. [DOI] [PubMed] [Google Scholar]
- 39.Morinello EJ, Ham AJL, Ranasinghe A, Sangaiah R, Swenberg JA. Chem. Res. Toxicol. 2001;14:327–334. doi: 10.1021/tx0002076. [DOI] [PubMed] [Google Scholar]
- 40.Yen TY, Holt S, Sangaiah R, Gold A, Swenberg JA. Chem. Res. Toxicol. 1998;11:810–815. doi: 10.1021/tx9800194. [DOI] [PubMed] [Google Scholar]
- 41.Chen HJC, Lin TC, Hong CL, Chiang LC. Chem. Res. Toxicol. 2001;14:1612–1619. doi: 10.1021/tx015551x. [DOI] [PubMed] [Google Scholar]
- 42.Rouzer CA, Chaudhary AK, Nokubo M, Ferguson DM, Reddy GR, Blair IA, Marnett LJ. Chem. Res. Toxicol. 1997;10:181–188. doi: 10.1021/tx9601216. [DOI] [PubMed] [Google Scholar]
- 43.Hoberg AM, Otteneder M, Marnett LJ, Poulsen HE. J. Mass Spectrom. 2004;39:38–42. doi: 10.1002/jms.547. [DOI] [PubMed] [Google Scholar]
- 44.Helleday T, Petermann E, Lundin C, Hodgson B, Sharma RA. Nat. Rev. Cancer. 2008;8:193–204. doi: 10.1038/nrc2342. [DOI] [PubMed] [Google Scholar]
- 45.Gerson SL. Nat. Rev. Cancer. 2004;4:296–307. doi: 10.1038/nrc1319. [DOI] [PubMed] [Google Scholar]
- 46.Hartley JA, Bingham JP, Souhami RL. Nucleic Acids Res. 1992;20:3175–3178. doi: 10.1093/nar/20.12.3175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Warren AJ, Hamilton JW. Chem. Res. Toxicol. 1996;9:1063–1071. doi: 10.1021/tx960070c. [DOI] [PubMed] [Google Scholar]
- 48.Jamieson ER, Lippard SJ. Chem. Rev. 1999;99:2467–2498. doi: 10.1021/cr980421n. [DOI] [PubMed] [Google Scholar]
- 49.Stern RS. N. Engl. J. Med. 2007;357:682–690. doi: 10.1056/NEJMct072317. [DOI] [PubMed] [Google Scholar]
- 50.Doerge DR, Churchwell MI, Marques MM, Beland FA. Carcinogenesis. 1999;20:1055–1061. doi: 10.1093/carcin/20.6.1055. [DOI] [PubMed] [Google Scholar]
- 51.Malejka-Giganti D, Parkin DR, Decker RW, Niehans GA, Bliss RL, Churchwell MI, Beland FA. Int. J. Cancer. 2008;122:1958–1965. doi: 10.1002/ijc.23352. [DOI] [PubMed] [Google Scholar]
- 52.Choi JY, Stover JS, Angel KC, Chowdhury G, Rizzo CJ, Guengerich FP. J. Biol. Chem. 2006;281:25297–25306. doi: 10.1074/jbc.M605699200. [DOI] [PubMed] [Google Scholar]
- 53.Singh R, Arlt VM, Henderson CJ, Phillips DH, Farmer PB, Gamboa da Costa G. J. Chromatogr. B: Anal. Technol. Biomed. Life Sci. 2010;878:2155–2162. doi: 10.1016/j.jchromb.2010.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Bendaly J, Zhao S, Neale JR, Metry KJ, Doll MA, States JC, Pierce WM, Jr, Hein DW. Cancer Epidemiol, Biomarkers Prev. 2007;16:1503–1509. doi: 10.1158/1055-9965.EPI-07-0305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Steenken S. Chem. Rev. 1989;89:503–520. [Google Scholar]
- 56.Breen AP, Murphy JA. Free Radical Biol. Med. 1995;18:1033–1077. doi: 10.1016/0891-5849(94)00209-3. [DOI] [PubMed] [Google Scholar]
- 57.Cheng Q, Gu J, Compaan KR, Schaefer HF., 3rd Chemistry. 2012;18:4877–4886. doi: 10.1002/chem.201102415. [DOI] [PubMed] [Google Scholar]
- 58.Kamiya H. Nucleic Acids Res. 2003;31:517–531. doi: 10.1093/nar/gkg137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Dizdaroglu M, Bergtold DS. Anal. Biochem. 1986;156:182–188. doi: 10.1016/0003-2697(86)90171-5. [DOI] [PubMed] [Google Scholar]
- 60.D’Ham C, Romieu A, Jaquinod M, Gasparutto D, Cadet J. Biochemistry. 1999;38:3335–3344. doi: 10.1021/bi981982b. [DOI] [PubMed] [Google Scholar]
- 61.Cao H, Wang Y. J. Am. Soc. Mass Spectrom. 2006;17:1335–1341. doi: 10.1016/j.jasms.2006.05.019. [DOI] [PubMed] [Google Scholar]
- 62.Hua YS, Wainhaus SB, Yang YN, Shen LX, Xiong YS, Xu XY, Zhang FG, Bolton JL, van Breemen RB. J. Am. Soc. Mass Spectrom. 2001;12:80–87. doi: 10.1016/S1044-0305(00)00191-4. [DOI] [PubMed] [Google Scholar]
- 63.Cao H, Wang Y. Nucleic Acids Res. 2007;35:4833–4844. doi: 10.1093/nar/gkm497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Brierley DJ, Martin SA. Antioxid. Redox Signaling. 2013;18:2420–2428. doi: 10.1089/ars.2012.4994. [DOI] [PubMed] [Google Scholar]
- 65.Madugundu GS, Cadet J, Wagner JR. Nucleic Acids Res. 2014;42:7450–7460. doi: 10.1093/nar/gku334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zhang Q, Wang Y. J. Am. Chem. Soc. 2003;125:12795–12802. doi: 10.1021/ja034866r. [DOI] [PubMed] [Google Scholar]
- 67.Romieu A, Bellon S, Gasparutto D, Cadet J. Org. Lett. 2000;2:1085–1088. doi: 10.1021/ol005643y. [DOI] [PubMed] [Google Scholar]
- 68.He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, Ding J, Jia Y, Chen Z, Li L, Sun Y, Li X, Dai Q, Song CX, Zhang K, He C, Xu GL. Science. 2011;333:1303–1307. doi: 10.1126/science.1210944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Ito S, Shen L, Dai Q, Wu SC, Collins LB, Swenberg JA, He C, Zhang Y. Science. 2011;333:1300–1303. doi: 10.1126/science.1210597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Wu SC, Zhang Y. Nat. Rev. Mol. Cell Biol. 2010;11:607–620. doi: 10.1038/nrm2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Yates JM, Fennell TR, Turner MJ, Jr, Recio L, Sumner SC. Carcinogenesis. 1994;15:277–283. doi: 10.1093/carcin/15.2.277. [DOI] [PubMed] [Google Scholar]
- 72.Deforce DL, Lemiere F, Hoes I, Millecamps RE, Esmans EL, De Leenheer A, Van den Eeckhout EG. Carcinogenesis. 1998;19:1077–1086. doi: 10.1093/carcin/19.6.1077. [DOI] [PubMed] [Google Scholar]
- 73.Haglund J, Van Dongen W, Lemiere F, Esmans EL. J. Am. Soc. Mass Spectrom. 2004;15:593–606. doi: 10.1016/j.jasms.2003.12.012. [DOI] [PubMed] [Google Scholar]
- 74.Maccubbin AE, Caballes L, Riordan JM, Huang DH, Gurtoo HL. Cancer Res. 1991;51:886–892. [PubMed] [Google Scholar]
- 75.Dedon PC. Chem. Res. Toxicol. 2008;21:206–219. doi: 10.1021/tx700283c. [DOI] [PubMed] [Google Scholar]
- 76.Amato NJ, Wang Y. Chem. Res. Toxicol. 2014;27:470–479. doi: 10.1021/tx400430g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Amato NJ, Zhai Q, Navarro D, Niedernhofer LJ, Wang Y. Nucleic Acids Res. 2015 doi: 10.1093/nar/gkv725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Gates KS. Chem. Res. Toxicol. 2009;22:1747–1760. doi: 10.1021/tx900242k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Harris CC, Vahakangas K, Newman MJ, Trivers GE, Shamsuddin A, Sinopoli N, Mann DL, Wright WE. Proc. Natl. Acad. Set U. S. A. 1985;82:6672–6676. doi: 10.1073/pnas.82.19.6672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Silvestri C, Brodbelt JS. Mass Spectrom. Rev. 2013;32:247–266. doi: 10.1002/mas.21363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Mueller DC, Piller M, Niessner R, Scherer M, Scherer G. J. Proteome Res. 2014;13:1602–1613. doi: 10.1021/pr401099r. [DOI] [PubMed] [Google Scholar]
- 82.Manchester DK, Weston A, Choi JS, Trivers GE, Fennessey PV, Quintana E, Farmer PB, Mann DL, Harris CC. Proc. Natl. Acad. Sci. U. S. A. 1988;85:9243–9247. doi: 10.1073/pnas.85.23.9243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Chandhary AK, Nokubo M, Marnett LJ, Blair IA. Biol. Mass Spectrom. 1994;23:457–464. doi: 10.1002/bms.1200230802. [DOI] [PubMed] [Google Scholar]
- 84.Chen HJC, Zhang L, Cox J, Cunningham JA, Chung FL. Chem. Res. Toxicol. 1998;11:1474–1480. doi: 10.1021/tx980107o. [DOI] [PubMed] [Google Scholar]
- 85.Chen HJC, Row SW, Hong CL. Chem. Res. Toxicol. 2002;15:262–268. doi: 10.1021/tx015578g. [DOI] [PubMed] [Google Scholar]
- 86.Oreffo V, Singh R, Rich KJ, Shuker DEG, Carthew P, Cordero R, Van Delft JHM, Farmer PB. Biomarkers. 2000;5:323–340. doi: 10.1080/135475000424348. [DOI] [PubMed] [Google Scholar]
- 87.Schlobe D, Holzle D, Hatz D, von Meyer L, Tricker AR, Richter E. Toxicology. 2008;245:154–161. doi: 10.1016/j.tox.2007.12.021. [DOI] [PubMed] [Google Scholar]
- 88.Friesen MD, Kaderlik K, Lin DX, Garren L, Bartsch H, Lang NP, Kadlubar FF. Chem. Res. Toxicol. 1994;7:733–739. doi: 10.1021/tx00042a004. [DOI] [PubMed] [Google Scholar]
- 89.Vanhoutte K, VanDongen W, Esmans EL, VandenEeckhout E, VanOnckelen H. Eur. Mass Spectrom. 1996;2:181–192. [Google Scholar]
- 90.Van den Driessche B, Lemiere F, van Dongen W, Esmans EL. J. Am. Soc. Mass Spectrom. 2004;15:568–579. doi: 10.1016/j.jasms.2003.12.005. [DOI] [PubMed] [Google Scholar]
- 91.Huber CG, Oberacher H. Mass Spectrom. Rev. 2001;20:310–343. doi: 10.1002/mas.10011. [DOI] [PubMed] [Google Scholar]
- 92.Cao H, Hearst JE, Corash L, Wang Y. Anal. Chem. 2008;80:2932–2938. doi: 10.1021/ac7023969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Deforce DL, Ryniers FP, van den Eeckhout EG, Lemiere F, Esmans EL. Anal. Chem. 1996;68:3575–3584. doi: 10.1021/ac9600013. [DOI] [PubMed] [Google Scholar]
- 94.Barry JP, Norwood C, Vouros P. Anal. Chem. 1996;68:1432–1438. doi: 10.1021/ac9510245. [DOI] [PubMed] [Google Scholar]
- 95.Apruzzese WA, Vouros P. J. Chromatogr. A. 1998;794:97–108. doi: 10.1016/s0021-9673(97)00820-0. [DOI] [PubMed] [Google Scholar]
- 96.Chen HJ, Lin WP. Anal. Chem. 2009;81:9812–9818. doi: 10.1021/ac9019472. [DOI] [PubMed] [Google Scholar]
- 97.Chen HJ, Lin GJ, Lin WP. Anal. Chem. 2010;82:4486–4493. doi: 10.1021/ac100391f. [DOI] [PubMed] [Google Scholar]
- 98.Chen HJ, Lin WP. Anal. Chem. 2011;83:8543–8551. doi: 10.1021/ac201874d. [DOI] [PubMed] [Google Scholar]
- 99.Tompkins EM, Jones DJ, Lamb JH, Marsden DA, Farmer PB, Brown K. Rapid Commun. Mass Spectrom. 2008;22:19–28. doi: 10.1002/rcm.3328. [DOI] [PubMed] [Google Scholar]
- 100.Wang M, Lao Y, Cheng G, Shi Y, Villalta PW, Hecht SS. Chem. Res. Toxicol. 2007;20:625–633. doi: 10.1021/tx600332p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Hopfgartner G, Varesio E, Tschappat V, Grivet C, Bourgogne E, Leuthold LA. J. Mass Spectrom. 2004;39:845–855. doi: 10.1002/jms.659. [DOI] [PubMed] [Google Scholar]
- 102.Spilsberg B, Rundberget T, Johannessen LE, Kristoffersen AB, Hoist-Jensen A, Berdal KG. J. Agric. Food Chem. 2010;58:6370–6375. doi: 10.1021/jf903065a. [DOI] [PubMed] [Google Scholar]
- 103.Balbo S, Turesky RJ, Villalta PW. Chem. Res. Toxicol. 2014;27:356–366. doi: 10.1021/tx4004352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Wang Y, Zhang Q, Wang Y. J. Am. Soc. Mass Spectrom. 2004;15:1565–1571. doi: 10.1016/j.jasms.2004.07.012. [DOI] [PubMed] [Google Scholar]
- 105.Ni JS, Pomerantz SC, Rozenski J, Zhang YH, McCloskey JA. Anal. Chem. 1996;68:1989–1999. doi: 10.1021/ac960270t. [DOI] [PubMed] [Google Scholar]
- 106.Mcluckey SA, Vanberkel GJ, Glish GL. J. Am. Soc. Mass Spectrom. 1992;3:60–70. doi: 10.1016/1044-0305(92)85019-G. [DOI] [PubMed] [Google Scholar]
- 107.Little DP, Aaserud DJ, Valaskovic GA, McLafferty FW. J. Am. Chem. Soc. 1996;118:9352–9359. [Google Scholar]
- 108.Xiong W, Glick J, Lin Y, Vouros P. Anal. Chem. 2007;79:5312–5321. doi: 10.1021/ac0701435. [DOI] [PubMed] [Google Scholar]
- 109.Chowdhury G, Guengerich FP. Angew. Chem., Int. Ed. 2008;47:381–384. doi: 10.1002/anie.200703942. [DOI] [PubMed] [Google Scholar]
- 110.Liao Q, Shen C, Vouros P. J. Mass Spectrom. 2009;44:549–560. doi: 10.1002/jms.1532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Sharma VK, Glick J, Liao Q, Shen C, Vouros P. J. Mass Spectrom. 2012;47:490–501. doi: 10.1002/jms.2054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Gao L, Zhang L, Cho BP, Chiarelli MP. J. Am. Soc. Mass Spectrom. 2008;19:1147–1155. doi: 10.1016/j.jasms.2008.04.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Gupta R, Beck JL, Sheil MM, Ralph SF. J. Inorg. Biochem. 2005;99:552–559. doi: 10.1016/j.jinorgbio.2004.11.001. [DOI] [PubMed] [Google Scholar]
- 114.Wu H, Aboleneen H. Anal. Biochem. 2000;287:126–135. doi: 10.1006/abio.2000.4827. [DOI] [PubMed] [Google Scholar]
- 115.Gassner C, Meyer S, Frey BM, Vollmert C. Transfus. Med. Rev. 2013;27:2–9. doi: 10.1016/j.tmrv.2012.10.001. [DOI] [PubMed] [Google Scholar]
- 116.Reiber DC, Grover TA, Brown RS. Anal. Chem. 1998;70:673–683. doi: 10.1021/ac971157l. [DOI] [PubMed] [Google Scholar]
- 117.Zhang LK, Ren Y, Rempel D, Taylor JS, Gross ML. J. Am. Soc. Mass Spectrom. 2001;12:1127–1135. doi: 10.1016/S1044-0305(01)00291-4. [DOI] [PubMed] [Google Scholar]
- 118.Pieles U, Zurcher W, Schar M, Moser HE. Nucleic Acids Res. 1993;21:3191–3196. doi: 10.1093/nar/21.14.3191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Tretyakova N, Matter B, Ogdie A, Wishnok JS, Tannenbaum SR. Chem. Res. Toxicol. 2001;14:1058–1070. doi: 10.1021/tx010062i. [DOI] [PubMed] [Google Scholar]
- 120.Chen HJC, Chiang LC, Tseng MC, Zhang LL, Ni JS, Chung FL. Chem Res. Toxicol. 1999;12:1119–1126. doi: 10.1021/tx990074s. [DOI] [PubMed] [Google Scholar]
- 121.Wang M, Yu N, Chen L, Villalta PW, Hochalter JB, Hecht SS. Chem. Res. Toxicol. 2006;19:319–324. doi: 10.1021/tx0502948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Thomson NM, Mijal RS, Ziegel R, Fleischer NL, Pegg AE, Tretyakova NY, Peterson LA. Chem. Res. Toxicol. 2004;17:1600–1606. doi: 10.1021/tx0498298. [DOI] [PubMed] [Google Scholar]
- 123.Wu H, Skrzypczynski Z, Cornwell MJ, Aboleneen H. Rapid Commun. Mass Spectrom. 2000;14:26–32. doi: 10.1002/(SICI)1097-0231(20000115)14:1<26::AID-RCM827>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- 124.Koehl P, Valladier P, Lefevre JF, Fuchs RP. Nucleic Acids Res. 1989;17:9531–9541. doi: 10.1093/nar/17.23.9531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Andersen N, Wang P, Wang Y. Chem. Res. Toxicol. 2013;26:1730–1738. doi: 10.1021/tx4002995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Andersen N, Wang J, Wang P, Jiang Y, Wang Y. Chem. Res. Toxicol. 2012;25:2523–2531. doi: 10.1021/tx300325q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Gu C, Wang Y. Nucleic Acids Res. 2007;35:3693–3704. doi: 10.1093/nar/gkm247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Choi JY, Chowdhury G, Zang H, Angel KC, Vu CC, Peterson LA, Guengerich FP. J. Biol. Chem. 2006;281:38244–38256. doi: 10.1074/jbc.M608369200. [DOI] [PubMed] [Google Scholar]
- 129.Hong H, Cao H, Wang Y. Nucleic Acids Res. 2007;35:7118–7127. doi: 10.1093/nar/gkm851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Yuan B, You C, Andersen N, Jiang Y, Moriya M, O’Connor TR, Wang Y. J. Biol. Chem. 2011;286:17503–17511. doi: 10.1074/jbc.M111.232835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Yuan B, O’Connor TR, Wang Y. ACS Chem. Biol. 2010;5:1021–1027. doi: 10.1021/cb100214b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Yuan B, Cao H, Jiang Y, Hong H, Wang Y. Proc. Natl. Acad. Sci. U. S. A. 2008;105:8679–8684. doi: 10.1073/pnas.0711546105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Zhai Q, Wang P, Wang Y. Carcinogenesis. 2014;35:2002–2006. doi: 10.1093/carcin/bgu085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.You C, Dai X, Yuan B, Wang J, Brooks PJ, Niedernhofer LJ, Wang Y. Nat. Chem. Biol. 2012;8:817–822. doi: 10.1038/nchembio.1046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.You C, Dai X, Yuan B, Wang Y. J. Biol. Chem. 2012;287:40915–40923. doi: 10.1074/jbc.M112.418681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Liu S, Wang J, Su Y, Guerrero C, Zeng Y, Mitra D, Brooks PJ, Fisher DE, Song H, Wang Y. Nucleic Acids Res. 2013;41:6421–6429. doi: 10.1093/nar/gkt360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Tilstra JS, Robinson AR, Wang J, Gregg SQ, Clauson CL, Reay DP, Nasto LA, St Croix CM, Usas A, Vo N, Huard J, Clemens PR, Stolz DB, Guttridge DC, Watkins SC, Garinis GA, Wang Y, Niedernhofer LJ, Robbins PD. J. Clin. Invest. 2012;122:2601–2612. doi: 10.1172/JCI45785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Wang J, Cao H, You C, Yuan B, Bahde R, Gupta S, Nishigori C, Niedernhofer LJ, Brooks PJ, Wang Y. Nucleic Acids Res. 2012;40:7368–7374. doi: 10.1093/nar/gks357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Wang J, Clauson CL, Robbins PD, Niedernhofer LJ, Wang Y. Aging Cell. 2012;11:714–716. doi: 10.1111/j.1474-9726.2012.00828.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Wang J, Yuan B, Guerrero C, Bahde R, Gupta S, Wang Y. Anal. Chem. 2011;83:2201–2209. doi: 10.1021/ac103099s. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Monien BH, Schumacher F, Herrmann K, Glatt H, Turesky RJ, Chesne C. Anal. Chem. 2014;87:641–648. doi: 10.1021/ac503803m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Yun BH, Rosenquist TA, Nikolic J, Dragicevic D, Tomic K, Jelakovic B, Dickman KG, Grollman AP, Turesky RJ. Anal. Chem. 2013;85:4251–4258. doi: 10.1021/ac400612x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Chung YT, Chen CL, Wu CC, Chan SA, Chi CW, Liu TY. Toxicol. Lett. 2008;183:21–27. doi: 10.1016/j.toxlet.2008.09.013. [DOI] [PubMed] [Google Scholar]
- 144.Tarun M, Rusling JF. Anal. Chem. 2005;77:2056–2062. doi: 10.1021/ac048283r. [DOI] [PubMed] [Google Scholar]
- 145.Hecht SS, Villalta PW, Sturla SJ, Cheng G, Yu N, Upadhyaya P, Wang M. Chem. Res. Toxicol. 2004;17:588–597. doi: 10.1021/tx034263t. [DOI] [PubMed] [Google Scholar]
- 146.Chen HJ, Lin CR. Toxicol. Lett. 2014;225:27–33. doi: 10.1016/j.toxlet.2013.11.032. [DOI] [PubMed] [Google Scholar]
- 147.Lao Y, Villalta PW, Sturla SJ, Wang M, Hecht SS. Chem. Res. Toxicol. 2006;19:674–682. doi: 10.1021/tx050351x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Doerge DR, Churchwell MI, Fang JL, Beland FA. Chem. Res. Toxicol. 2000;13:1259–1264. doi: 10.1021/tx0001575. [DOI] [PubMed] [Google Scholar]
- 149.Delatour T, Mally A, Richoz J, Ozden S, Dekant W, Ihmels H, Otto D, Gasparutto D, Marin-Kuan M, Schilter B, Cavin C. Mol. Nutr. Food Res. 2008;52:472–482. doi: 10.1002/mnfr.200700276. [DOI] [PubMed] [Google Scholar]
- 150.Paini A, Punt A, Viton F, Scholz G, Delatour T, Marin-Kuan M, Schilter B, van Bladeren PJ, Rietjens IM. Toxicol. Appl. Pharmacol. 2010;245:57–66. doi: 10.1016/j.taap.2010.01.016. [DOI] [PubMed] [Google Scholar]
- 151.Pan SS, Iracki T. Mol. Pharmacol. 1988;34:223–228. [PubMed] [Google Scholar]
- 152.Marsch GA, Mundkowski RG, Morris BJ, Manier ML, Hartman MK, Guengerich FP. Chem. Res. Toxicol. 2001;14:600–608. doi: 10.1021/tx010006h. [DOI] [PubMed] [Google Scholar]
- 153.Frandsen H, Grivas S, Turesky RJ, Andersson R, Dragsted LO, Larsen JC. Carcinogenesis. 1994;15:2553–2558. doi: 10.1093/carcin/15.11.2553. [DOI] [PubMed] [Google Scholar]
- 154.Lu K, Moeller B, Doyle-Eisele M, McDonald J, Swenberg JA. Chem. Res. Toxicol. 2011;24:159–161. doi: 10.1021/tx1003886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Lunec J. Free Radical Res. 1998;29:601–608. doi: 10.1080/10715769800300651. [DOI] [PubMed] [Google Scholar]
- 156.Collins AR, Cadet J, Moller L, Poulsen HE, Vina J. Arch. Biochem. Biophys. 2004;423:57–65. doi: 10.1016/j.abb.2003.12.022. [DOI] [PubMed] [Google Scholar]
- 157.Wink DA, Kasprzak KS, Maragos CM, Elespuru RK, Misra M, Dunams TM, Cebula TA, Koch WH, Andrews AW, Allen JS, et al. Science. 1991;254:1001–1003. doi: 10.1126/science.1948068. [DOI] [PubMed] [Google Scholar]
- 158.Van der Weyden MB, Kelley WN. J. Biol. Chem. 1976;251:5448–5456. [PubMed] [Google Scholar]
- 159.Dong M, Wang C, Deen WM, Dedon PC. Chem. Res. Toxicol. 2003;16:1044–1055. doi: 10.1021/tx034046s. [DOI] [PubMed] [Google Scholar]
- 160.Neale JR, Smith NB, Pierce WM, Hein DW. Polycyclic Aromat. Compd. 2008;28:402–417. doi: 10.1080/10406630802377773. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Wang Y, Taylor JS, Gross ML. Chem. Res. Toxicol. 2001;14:738–745. doi: 10.1021/tx010009u. [DOI] [PubMed] [Google Scholar]
- 162.Wang Y, Taylor JS, Gross ML. Chem. Res. Toxicol. 1999;12:1077–1082. doi: 10.1021/tx9900831. [DOI] [PubMed] [Google Scholar]
- 163.Wang Y. Chem. Res. Toxicol. 2002;15:671–676. doi: 10.1021/tx0155855. [DOI] [PubMed] [Google Scholar]
- 164.Wang Y, Wang Y. Anal. Chem. 2003;75:6306–6313. doi: 10.1021/ac034683n. [DOI] [PubMed] [Google Scholar]
- 165.Mohamed D, Linscheid M. Anal. Bioanal Chem. 2008;392:805–817. doi: 10.1007/s00216-008-2236-0. [DOI] [PubMed] [Google Scholar]
- 166.Wang Y. Anal. Chem. 2003;75:6306–6313. doi: 10.1021/ac034683n. [DOI] [PubMed] [Google Scholar]
- 167.Liu S, Wang Y. Anal. Chem. 2013;85:6732–6739. doi: 10.1021/ac4012232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168.Wang Y, Gross ML, Taylor JS. Biochemistry. 2001;40:11785–11793. doi: 10.1021/bi0111552. [DOI] [PubMed] [Google Scholar]
- 169.Su DG, Fang H, Gross ML, Taylor JS. Proc. Natl. Acad. Sci. U. S. A. 2009;106:12861–12866. doi: 10.1073/pnas.0902386106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170.Chen HJ, Chiu WL. Chem. Res. Toxicol. 2003;16:1099–1106. doi: 10.1021/tx034057l. [DOI] [PubMed] [Google Scholar]
- 171.Fang M, Edwards RJ, Bartlet-Jones M, Taylor GW, Murray S, Boobis AR. Carcinogenesis. 2004;25:1053–1062. doi: 10.1093/carcin/bgh084. [DOI] [PubMed] [Google Scholar]
- 172.Da Pieve C, Sahgal N, Moore SA, Velasco-Garcia MN. Rapid Commun. Mass Spectrom. 2013;27:2493–2503. doi: 10.1002/rcm.6709. [DOI] [PubMed] [Google Scholar]
- 173.Knutson CG, Skipper PL, Liberman RG, Tannenbaum SR, Marnett LJ. Chem. Res. Toxicol. 2008;21:1290–1294. doi: 10.1021/tx800049v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174.Huang CC, Shih WC, Wu CF, Chen MF, Chen YL, Lin YH, Wu KY. Rapid Commun. Mass Spectrom. 2008;22:706–710. doi: 10.1002/rcm.3414. [DOI] [PubMed] [Google Scholar]
- 175.Chen HJ, Wu CF, Huang JL. Toxicol. Lett. 2005;155:403–410. doi: 10.1016/j.toxlet.2004.11.009. [DOI] [PubMed] [Google Scholar]
- 176.Shuker DE, Bartsch H. Mutat. Res. 1994;313:263–268. doi: 10.1016/0165-1161(94)90055-8. [DOI] [PubMed] [Google Scholar]
- 177.Sharma RA, Farmer PB. Clin. Cancer Res. 2004;10:4901–4912. doi: 10.1158/1078-0432.CCR-04-0098. [DOI] [PubMed] [Google Scholar]
- 178.Fernandez-Peralbo MA, de Castro MDL. TrAC, Trends Anal. Chem. 2012;41:75–85. [Google Scholar]
- 179.Kao CY, Giese RW. Chem. Res. Toxicol. 2005;18:70–75. doi: 10.1021/tx049854d. [DOI] [PubMed] [Google Scholar]
- 180.Leuthold LA, Grivet C, Allen M, Baumert M, Hopfgartner G. Rapid Commun. Mass Spectrom. 2004;18:1995–2000. doi: 10.1002/rcm.1587. [DOI] [PubMed] [Google Scholar]
- 181.Olsen R, Ovrebo S, Thorud S, Lundanes E, Thomassen Y, Greibrokk T, Molander P. Analyst. 2008;133:802–809. doi: 10.1039/b719842f. [DOI] [PubMed] [Google Scholar]
- 182.Sangaraju D, Villalta PW, Wickramaratne S, Swenberg J, Tretyakova N. J. Am. Soc. Mass Spectrom. 2014;25:1124–1135. doi: 10.1007/s13361-014-0916-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183.Otteneder MB, Knutson CG, Daniels JS, Hashim M, Crews BC, Remmel RP, Wang H, Rizzo C, Marnett LJ. Proc. Natl. Acad. Sci. U. S A. 2006;103:6665–6669. doi: 10.1073/pnas.0602017103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184.Stout MD, Jeong YC, Boysen G, Li Y, Sangaiah R, Ball LM, Gold A, Swenberg JA. Chem. Res. Toxicol. 2006;19:563–570. doi: 10.1021/tx050346t. [DOI] [PubMed] [Google Scholar]
- 185.Chen HJ, Chen YC. Chem. Res. Toxicol. 2009;22:1334–1341. doi: 10.1021/tx900129e. [DOI] [PubMed] [Google Scholar]
- 186.Vanhoutte K, Van Dongen W, Hoes I, Lemiere F, Esmans EL, Van Onckelen H, Van den Eeckhout E, van Soest RE, Hudson AJ. Anal. Chem. 1997;69:3161–3168. doi: 10.1021/ac970121q. [DOI] [PubMed] [Google Scholar]
- 187.Tang H, Fang H, Yin E, Brasier AR, Sowers LC, Zhang K. Anal. Chem. 2014;86:5526–5534. doi: 10.1021/ac500972x. [DOI] [PubMed] [Google Scholar]
- 188.Fu L, Amato NJ, Wang P, McGowan SJ, Niedernhofer LJ, Wang Y. Anal. Chem. 2015 doi: 10.1021/acs.analchem.5b00951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189.Wang H, Cao H, Wang Y. Chem. Res. Toxicol. 2010;23:74–81. doi: 10.1021/tx900286c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190.Liu S, Dunwell TL, Pfeifer GP, Dunwell JM, Ullah I, Wang Y. PLoS One. 2013;8:e84620. doi: 10.1371/journal.pone.0084620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 191.Tretyakova N, Goggin M, Sangaraju D, Janis G. Chem. Res. Toxicol. 2012;25:2007–2035. doi: 10.1021/tx3002548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192.Rouzer CA, Chaudhary AK, Nokubo M, Ferguson DM, Reddy GR, Blair IA, Marnett LJ. Chem. Res. Toxicol. 1997;10:181–188. doi: 10.1021/tx9601216. [DOI] [PubMed] [Google Scholar]
- 193.Zayas B, Stillwell SW, Wishnok JS, Trudel LJ, Skipper P, Yu MC, Tannenbaum SR, Wogan GN. Carcinogenesis. 2007;28:342–349. doi: 10.1093/carcin/bgl142. [DOI] [PubMed] [Google Scholar]
- 194.Inagaki S, Esaka Y, Goto M, Deyashiki Y, Sako M. Biol. Pharm. Bull. 2004;27:273–276. doi: 10.1248/bpb.27.273. [DOI] [PubMed] [Google Scholar]
- 195.Douki T, Odin F, Caillat S, Favier A, Cadet J. Free Radical Biol. Med. 2004;37:62–70. doi: 10.1016/j.freeradbiomed.2004.04.013. [DOI] [PubMed] [Google Scholar]
- 196.Leclercq L, Laurent C, De Pauw E. Anal. Chem. 1997;69:1952–1955. doi: 10.1021/ac9607673. [DOI] [PubMed] [Google Scholar]
- 197.Balbo S, Villalta PW, Hecht SS. Chem. Res. Toxicol. 2011;24:1729–1734. doi: 10.1021/tx200262d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 198.Singh R, Farmer PB. Carcinogenesis. 2006;27:178–196. doi: 10.1093/carcin/bgi260. [DOI] [PubMed] [Google Scholar]
- 199.Liu S, Ji D, Cliffe L, Sabatini R, Wang Y. J. Am. Soc. Mass Spectrom. 2014;25:1763–1770. doi: 10.1007/s13361-014-0960-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 200.Zhang F, Bartels MJ, Pottenger LH, Gollapudi BB, Schisler MR. Rapid Commun. Mass Spectrom. 2005;19:2767–2772. doi: 10.1002/rcm.2125. [DOI] [PubMed] [Google Scholar]
- 201.Walker VE, Fennell TR, Upton PB, Skopek TR, Prevost V, Shuker DEG, Swenberg JA. Cancer Res. 1992;52:4328–4334. [PubMed] [Google Scholar]
- 202.Yong LC, Schulte PA, Kao CY, Giese RW, Boeniger MF, Strauss GHS, Petersen MR, Wiencke JK. Am. J. Ind. Med. 2007;50:293–302. doi: 10.1002/ajim.20443. [DOI] [PubMed] [Google Scholar]
- 203.Foiles PG, Akerkar SA, Carmella SG, Kagan M, Stoner GD, Resau JH, Hecht SS. Chem. Res. Toxicol. 1991;4:364–368. doi: 10.1021/tx00021a017. [DOI] [PubMed] [Google Scholar]
- 204.Lin DX, Lay JO, Bryant MS, Malaveille C, Friesen M, Bartsch H, Lang NP, Kadlubar FF. Environ. Health Perspect. 1994;102:11–16. doi: 10.1289/ehp.94102s611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205.Airoldi L, Orsi F, Magagnotti C, Coda R, Randone D, Casetta G, Peluso M, Hautefeuille A, Malaveille C, Vineis P. Carcinogenesis. 2002;23:861–866. doi: 10.1093/carcin/23.5.861. [DOI] [PubMed] [Google Scholar]
- 206.Chen HJ, Chang CM. Chem. Res. Toxicol. 2004;17:963–971. doi: 10.1021/tx0341963. [DOI] [PubMed] [Google Scholar]
- 207.Yen TY, ChristovaGueoguieva NI, Scheller N, Holt S, Swenberg JA, Charles MJ. J. Mass Spectrom. 1996;31:1271–1276. doi: 10.1002/(SICI)1096-9888(199611)31:11<1271::AID-JMS420>3.0.CO;2-J. [DOI] [PubMed] [Google Scholar]
- 208.Rindgen D, Turesky RJ, Vouros P. Chem. Res. Toxicol. 1995;8:1005–1013. doi: 10.1021/tx00050a003. [DOI] [PubMed] [Google Scholar]
- 209.Liu Z, YoungSciame R, Hecht SS. Chem. Res. Toxicol. 1996;9:774–780. doi: 10.1021/tx950206r. [DOI] [PubMed] [Google Scholar]
- 210.Gamboa da Costa G, Churchwell MI, Hamilton LP, Von Tungeln LS, Beland FA, Marques MM, Doerge DR. Chem. Res. Toxicol. 2003;16:1328–1337. doi: 10.1021/tx034108e. [DOI] [PubMed] [Google Scholar]
- 211.Emmett MR, Caprioli RM. J. Am. Soc. Mass Spectrom. 1994;5:605–613. doi: 10.1016/1044-0305(94)85001-1. [DOI] [PubMed] [Google Scholar]
- 212.Shevchenko A, Wilm M, Vorm O, Mann M. Anal. Chem. 1996;68:850–858. doi: 10.1021/ac950914h. [DOI] [PubMed] [Google Scholar]
- 213.Ricicki EM, Soglia JR, Teitel C, Kane R, Kadlubar F, Vouros P. Chem. Res. Toxicol. 2005;18:692–699. doi: 10.1021/tx049692l. [DOI] [PubMed] [Google Scholar]
- 214.Ricicki EM, Luo W, Fan WH, Zhao LP, Zarbl H, Vouros P. Anal. Chem. 2006;78:6422–6432. doi: 10.1021/ac0607360. [DOI] [PubMed] [Google Scholar]
- 215.Randall KL, Argoti D, Paonessa JD, Ding Y, Oaks Z, Zhang YS, Vouros P. J. Chromatogr. A. 2010;1217:4135–4143. doi: 10.1016/j.chroma.2009.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216.Stepanov I, Muzic J, Le CT, Sebero E, Villalta P, Ma B, Jensen J, Hatsukami D, Hecht SS. Chem. Res. Toxicol. 2013;26:37–45. doi: 10.1021/tx300282k. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 217.Bessette EE, Goodenough AK, Langouet S, Yasa I, Kozekov ID, Spivack SD, Turesky RJ. Anal. Chem. 2009;81:809–819. doi: 10.1021/ac802096p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 218.Hong H, Wang Y. Anal. Chem. 2007;79:322–326. doi: 10.1021/ac061465w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 219.Zhao YY, Wu SP, Liu S, Zhang Y, Lin RC. Chem.-Biol. Interact. 2014;220C:181–192. doi: 10.1016/j.cbi.2014.06.029. [DOI] [PubMed] [Google Scholar]
- 220.Feng F, Wang X, Yuan H, Wang H. J. Chromatogr. B: Anal. Technol. Biomed. Life Sci. 2009;877:2104–2112. doi: 10.1016/j.jchromb.2009.05.054. [DOI] [PubMed] [Google Scholar]
- 221.Tang Y, Zheng SJ, Qi CB, Feng YQ, Yuan BF. Anal. Chem. 2015;87:3445–3452. doi: 10.1021/ac504786r. [DOI] [PubMed] [Google Scholar]
- 222.Iwata M, Graf L, Awaya N, Torok-Storb B. Blood. 2002;100:1318–1325. doi: 10.1182/blood-2002-01-0062. [DOI] [PubMed] [Google Scholar]
- 223.Nelson CP, Kidd LC, Sauvageot J, Isaacs WB, De Marzo AM, Groopman JD, Nelson WG, Kensler TW. Cancer Res. 2001;61:103–109. [PubMed] [Google Scholar]
- 224.Hah SS, Henderson PT, Turteltaub KW. J. Biomed. Sci. 2009;16:54. doi: 10.1186/1423-0127-16-54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225.Hah SS, Henderson PT, Turteltaub KW. Bioorg. Med. Chem. Lett. 2010;20:2448–2451. doi: 10.1016/j.bmcl.2010.03.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 226.Hah SS, Sumbad RA, de Vere White RW, Turteltaub KW, Henderson PT. Chem. Res. Toxicol. 2007;20:1745–1751. doi: 10.1021/tx700376a. [DOI] [PubMed] [Google Scholar]
- 227.Malayappan B, Johnson L, Nie B, Panchal D, Matter B, Jacobson P, Tretyakova N. Anal. Chem. 2010;82:3650–3658. doi: 10.1021/ac902923s. [DOI] [PubMed] [Google Scholar]
- 228.Paz MM, Ladwa S, Champeil E, Liu Y, Rockwell S, Boamah EK, Bargonetti J, Callahan J, Roach J, Tomasz M. Chem. Res. Toxicol. 2008;21:2370–2378. doi: 10.1021/tx8002615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 229.Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C, Martin NM, Jackson SP, Smith GC, Ashworth A. Nature. 2005;434:917–921. doi: 10.1038/nature03445. [DOI] [PubMed] [Google Scholar]
- 230.Fong PC, Boss DS, Yap TA, Tutt A, Wu P, Mergui-Roelvink M, Mortimer P, Swaisland H, Lau A, O’Connor MJ, Ashworth A, Carmichael J, Kaye SB, Schellens JH, de Bono JS. N. Engl. J. Med. 2009;361:123–134. doi: 10.1056/NEJMoa0900212. [DOI] [PubMed] [Google Scholar]
- 231.Soglia JR, Turesky RJ, Paehler A, Vouros P. Anal. Chem. 2001;73:2819–2827. doi: 10.1021/ac010218j. [DOI] [PubMed] [Google Scholar]
- 232.Schild LJ, Phillips DH, Osborne MR, Hewer A, Beland FA, Churchwell MI, Brown K, Gaskell M, Wright E, Poirier MC. Mutagenesis. 2005;20:115–124. doi: 10.1093/mutage/gei015. [DOI] [PubMed] [Google Scholar]
- 233.Chen L, Wang M, Villalta PW, Luo X, Feuer R, Jensen J, Hatsukami DK, Hecht SS. Chem. Res. Toxicol. 2007;20:108–113. doi: 10.1021/tx060232x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234.Mutlu E, Jeong YC, Collins LB, Ham AJ, Upton PB, Hatch G, Winsett D, Evansky P, Swenberg JA. Chem. Res. Toxicol. 2012;25:391–399. doi: 10.1021/tx200447w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235.Yin RC, Liu SQ, Zhao C, Lu ML, Tang MS, Wang HL. Anal. Chem. 2013;85:3190–3197. doi: 10.1021/ac3034695. [DOI] [PubMed] [Google Scholar]