Abstract
Background
Trichophyton rubrum is a cosmopolitan filamentous fungus that can infect human keratinized tissue (skin, nails and, rarely, hair) and is the major agent of all chronic and recurrent dermatophytoses. The dermatophyte infection process is initiated through the release of arthroconidial adhesin, which binds to the host stratum corneum. The conidia then germinate, and fungal hyphae invade keratinized skin structures through the secretion of proteases. Although arthroconidia play a central role in pathogenesis, little is known about the dormancy and germination of T. rubrum conidia and the initiation of infection. The objective of this study was to evaluate the transcriptional gene expression profile of T. rubrum conidia during growth on keratin- or elastin-containing medium, mimicking superficial and deep dermatophytosis, respectively.
Results
A transcriptional profiling analysis was conducted using a custom oligonucleotide-based microarray by comparing T. rubrum conidia grown on elastin and keratin substrates. This comparison shows differences according to protein source used, but consisted of a very small set of genes, which could be attributed to the quiescent status of conidia. The modulated genes were related to the dormancy, survival and germination of conidia, including genes involved in the respiratory chain, signal transduction and lipid metabolism. However, an induction of a great number of proteases occurred when T. rubrum was grown in the presence of keratin such as the subtilisin family of proteases (Sub 1 and Sub 3) and leucine aminopeptidase (Lap 1 and Lap 2). Interestingly, keratin also promoted the up-regulation of a gene encoding an adhesin-like protein with a tandem repeat sequence. In silico analysis showed that the protein contains a domain related to adhesin that may play a role in host-pathogen interactions. The expression of this adhesin-like gene was also induced during the co-culture of T. rubrum with a human keratinocyte cell line, confirming its role in fungal-host interactions.
Conclusion
These results contribute to the discovery of new targets involved in the adhesion of conidia and the maintenance of conidial dormancy, which are essential for triggering the process of infection and the chronicity of dermatophytosis.
Keywords: Adhesin-like protein, Conidia, Elastin, Keratin, Transcriptional gene expression, Trichophyton rubrum
Background
Trichophyton rubrum is the main aetiological agent of human dermatophytoses, as well as all chronic and recurrent fungal infections in the world [1, 2]. Dermatophytes are adapted to infect keratinized tissues such as skin, hair and nails due to their ability to use keratin as a nutrient [3]. Although dermatophytes rarely penetrate beyond the epidermis, deeper penetration and systemic infections can occur in immunocompromised hosts [4]. Currently, T. rubrum has become an important public health problem due to an increase in invasive infections in immunocompromised patients [5, 6]. Analysis of the gene expression profile of fungi grown on culture medium containing protein substrates such as keratin and elastin, which mimic superficial and deep infections, respectively, can be used to understand fungal-host interactions [4, 7]. Additionally, the gene expression response of T. rubrum co-cultured on human keratinocytes can be evaluated.
The dermatophyte infection process is initiated through the release of arthroconidia adhesins, which bind to the host stratum corneum [8]. Most fungal adhesins contain an N-terminal carbohydrate or peptide-binding domain, central Ser- and Thr-rich domains, commonly in tandem repeats, and a C-terminal region that mediates covalent cross-linking to the wall through modified glycosylphosphatidylinositol (GPI) anchors [9]. Tandem repeats are adjacent DNA sequences 2–200 nucleotides in length. Some tandem repeats are involved in the pathogenicity of microorganisms and adaptation to a new environment [10]. Adhesins are considered the first line of a pathogen’s stratagem of host-cell invasion, and differences in adhesion have been associated with the greater pathogenicity/virulence of one strain over another [11]. Adhesins participate in mating, colony morphology changes, biofilm formation, fruiting body development, and interactions with mammalian and plant hosts. However, very few adhesins have been identified thus far in filamentous fungi [12].
After adhesion to the host’s skin, quiescent arthroconidia begin to germinate, leading to the formation of fungal hyphae that invade keratinized skin structures through the secretion of endo- and exoproteases [13]. Elucidation of this response of T. rubrum to the host may reveal new molecular targets that could be explored for the development of novel antifungal agents. These targets may be involved in the establishment and maintenance of fungal infection, and they may include genes that participate in the adhesion, dormancy and onset of the germination of conidia. The vast majority of studies on the gene expression of the fungal-host relationship in T. rubrum have used grown mycelium and then added skin fragments or protein substrates to the culture medium [14, 15]. However, arthroconidia are considered the primary infectious propagules that reach the skin and nails during infection in humans, and their germination is a crucial step in this process [16]. Therefore, the aim of the present study was to evaluated and compare the transcriptional profile of T. rubrum conidia during growth on keratin and elastin substrates by that contributing to the understanding of the infectious process of dermatophytes.
Results
The transcriptome profile of T. rubrum after growth on protein substrates was analysed using a microarray custom slide containing 6,091 sequences, which correspond to approximately 70 % of T. rubrum protein coding genes (according to the latest update released by the Broad Institute on 02/12/2014, available at www.broadinstitute.org/annotation/genome/dermatophyte_comparative). We identified 215 differentially expressed transcripts (P < 0.05, fold change ≥ 4) when the two growth conditions (keratin and elastin) were compared each one against the control (Cove’s minimal medium). The transcripts were mapped according to the Broad Institute database, and we found 145 and 142 transcripts that were modulated in the presence of elastin and keratin, respectively. Seventy-two differentially expressed transcripts were shared in both conditions (Fig. 1).
Fig. 1.
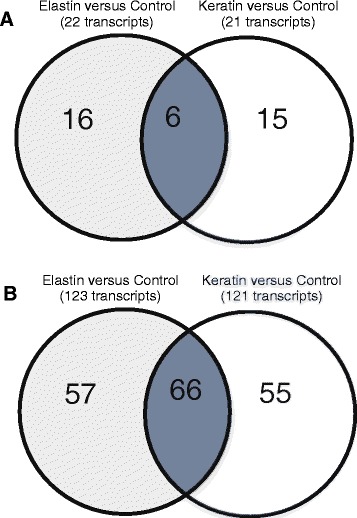
Venn diagrams showing the number of transcripts which are specifically up-regulated (a) and down-regulated (b) during the growth of T. rubrum on keratin or elastin compared to glucose minimal medium, respectively
Functional categorization of the genes differentially expressed on the two protein substrates identified genes involved in signal transduction, transport, drug resistance mechanisms, regulation of biological processes, response to stress, protease activity, fatty acid and lipid metabolism, the cell wall, and metabolic processes. Regarding to protease activity, the growth of T. rubrum conidia on keratin induced six protease genes that encoded respectively: leucine aminopeptidases Lap 1 and Lap 2; subtilisin-like proteins Sub 1, Sub3 and Sub 6; and metalloproteinases Mep3. Exclusive induction of the gene encoding Mep 4 was observed for conidia grown on keratin and elastin. The repression of 40 genes involved in metabolic processes was observed during the growth of T. rubrum conidia on protein substrates (keratin and elastin). On the other hand, genes related to the respiratory chain (NADP-dependent leukotriene b4 12-hydroxydehydrogenase) and tricarboxylic acid cycle (acyl enzyme) were induced. Besides genes differentially expressed in the two conditions, 30-34 % were found to be unclassified (Figs. 2a and b). Those genes exclusively modulated on keratin or elastin and those commonly modulated on both substrates are shown in Tables 1, 2 and 3, respectively. A subset of genes involved in different biological processes, such as adhesion (adhesin-like protein), dormancy (phosphatidyl synthase, polarized growth protein, Ras-guanyl exchange factor), protease secretion (subtilisins 1, 3 and alkaline phosphatase), and adaptation to nutritional stress (sugar MFS transporter and glutathione synthase), were validated by quantitative PCR (Fig. 3a), and shows a strong positive correlation.
Fig. 2.
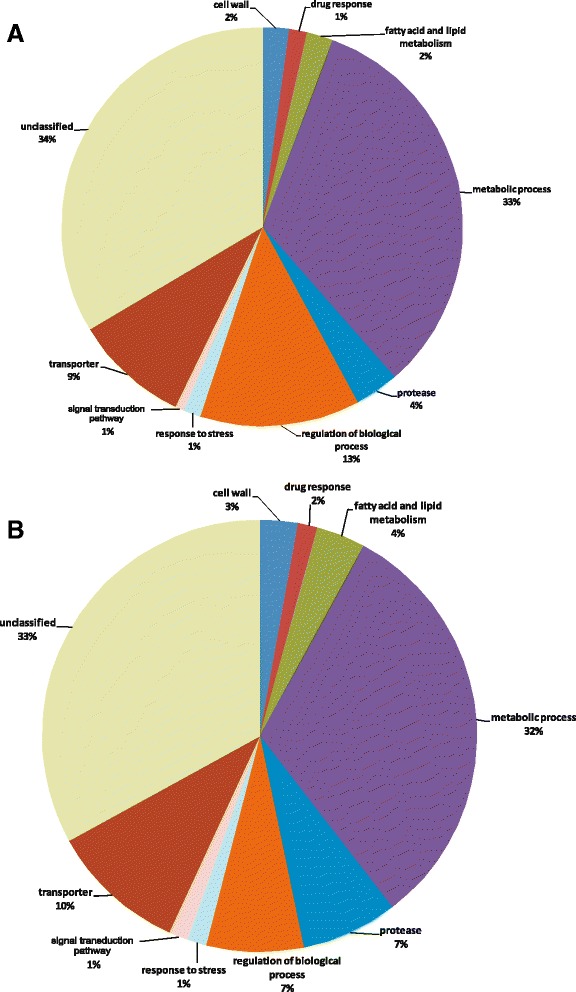
Functional annotation of genes modulated on keratin (a) and elastin (b) based on Gene Ontology
Table 1.
Genes exclusively modulated on keratin substrates
| Gene ID | Tentative annotation | Expression change (n.fold)a |
|---|---|---|
| Metabolic process | ||
| TERG_00499 | Hypothetical protein | −4.88 |
| TERG_02958 | Assimilatory sulfite reductase | −6.51 |
| TERG_03258 | Alkaline phosphatase | −5.57 |
| TERG_03339 | Alternative oxidase | −4.69 |
| TERG_03705 | Cytochrome p450 | −11.11 |
| TERG_03706 | Cytochrome p450 | −5.68 |
| TERG_05504 | Thiol-specific antioxidant | −4.40 |
| TERG_05628 | GPI ethanolamine phosphate transferase 1 | −5.17 |
| TERG_06614 | Formate dehydrogenase | −4.67 |
| TERG_07017 | NADP-dependent leukotriene b4 12-hydroxydehydrogenase | +4.47 |
| TERG_07477 | Copper-sulfate regulated protein 1 | −4.17 |
| TERG_07236 | Hypothetical protein | −4.20 |
| TERG_08353 | Cytochrome p450 | −8.40 |
| TERG_07796 | Short-chain dehydrogenase reductase family | −4.15 |
| TERG_08140 | 2-Heptaprenyl- -naphthoquinone | −6.92 |
| TERG_07777 | O-acetylhomoserine –lyase | −4.58 |
| TERG_12680 | Alpha-mannosidase | −5.26 |
| Regulation of biological process | ||
| TERG_00222 | CP2 transcription | +5.17 |
| TERG_01360 | C6 transcription factor | +4.79 |
| TERG_08771 | Adhesin like protein | +4.17 |
| TERG_11518 | Glucokinase | −4.25 |
| TERG_07957 | AP-1-like transcription factor | −6.07 |
| TERG_08139 | NAD dependent epimerase dehydratase family protein | −5.21 |
| Transporter | ||
| TERG_01994 | OPT family oligopeptide transporter | −6.25 |
| TERG_02616 | Integral membrane protein | −6.15 |
| TERG_03174 | Siderochrome-iron transporter sit1 | −5.95 |
| TERG_03250 | Monocarboxylate permease-like protein | −4.35 |
| TERG_03928 | Zinc-regulated transporter 1 | −4.45 |
| TERG_05199 | MFS gliotoxin efflux transporter | −6.08 |
| Protease activity | ||
| TERG_02199 | Glutamate carboxypeptidase | −4.70 |
| TERG_02990 | Subtilisin-like protease 6 | +12.45 |
| TERG_03248 | Extracellular metalloproteinase 3 | +5.81 |
| TERG_03400 | Subtilisin-like protease 1 | +13.73 |
| TERG_03815 | Subtilisin-like protease 3 | +9.66 |
| TERG_05652 | Leucine aminopeptidase 1 | +6.62 |
| TERG_08405 | Leucine aminopeptidase 2 | +10.74 |
| Fatty acid and lipid metabolism | ||
| TERG_01901 | Glycerol kinase | −6.79 |
| TERG_02984 | Cytochrome p450 51 | −5.13 |
| TERG_05518 | Short chain dehydrogenase | −5.35 |
| TERG_11671 | Phosphatidylserine synthase | −4.76 |
| Cell wall | ||
| TERG_04234 | Hypothetical protein | −4.97 |
| TERG_08178 | Endoglucanase | +4.16 |
| Signal transduction pathway | ||
| TERG_04867 | Sam and pH domain-containing protein | −5.17 |
| TERG_05744 | GTP-binding protein | −4.41 |
| Drug response | ||
| TERG_04952 | Multidrug resistance protein | +4.78 |
| Response to stress | ||
| TERG_07058 | HSP70 family | +4.47 |
a Only genes with a fold change of four or higher are indicated. + induction; − repression
Table 2.
Genes exclusively modulated on elastin substrates
| Gene Id | Tentative annotation | Expression change (n.fold)a |
|---|---|---|
| Metabolic process | ||
| TERG_00058 | Oxidoreductase | −4.18 |
| TERG_00563 | GNAT family n-acetyltransferase | −5.67 |
| TERG_00831 | Gamma-glutamyltranspeptidase | −4.22 |
| TERG_00852 | Phytanoyl- dioxygenase | −4.57 |
| TERG_01164 | Beta-alanine synthase | −4.82 |
| TERG_02340 | Polysaccharide deacetylase | −4.85 |
| TERG_02839 | Nacht and ankyrin domain protein | −5.83 |
| TERG_02842 | 6-Hydroxy-d-nicotine oxidase | −4.43 |
| TERG_03695 | Pyrroline-5-carboxylate reductase | +4.38 |
| TERG_04310 | Alcohol dehydrogenase | −4.62 |
| TERG_04543 | Classes i and ii family protein | −4.08 |
| TERG_05299 | Glutathione s- | −4.99 |
| TERG_06147 | Rhodocoxin reductase | −4.00 |
| TERG_06160 | Nitrite copper-containing | −4.69 |
| TERG_06741 | Ubiquitin c-terminal hydrolase | −4.54 |
| TERG_07943 | Hypothetical protein | −4.33 |
| TERG_00830 | Cytochrome p450 | −4.05 |
| TERG_01578 | NB-ARC and ankyrin domain protein | −4.03 |
| TERG_07083 | Hypothetical protein | −4.04 |
| Regulation of biological process | ||
| TERG_00487 | Hypothetical protein | −8.50 |
| TERG_01003 | 37 s ribosomal protein rsm22 | −4.90 |
| TERG_01198 | Pre-mRNA-splicing factor rse1 | −5.29 |
| TERG_02418 | Translation initiation factor sui1 | −31.53 |
| TERG_05380 | Protein kinase regulator ste50 | −8.12 |
| TERG_05655 | An1 zinc finger protein | −4.46 |
| TERG_05963 | WD repeat protein | +4.40 |
| TERG_06059 | Helicase swr1 | −7.58 |
| TERG_06159 | Hypothetical protein | −4.72 |
| TERG_06729 | Taz1-interacting factor 1 | −4.49 |
| TERG_06822 | Polarized growth protein | +13.44 |
| TERG_06891 | C6 transcription factor | −5.52 |
| TERG_08611 | E3 Ubiquitin ligase complex scf subunit sconc | −4.33 |
| Transporter | ||
| TERG_01336 | MFS transporter | −4.33 |
| TERG_01353 | V-type c subunit family protein | −9.94 |
| TERG_02333 | Acetyl-coenzyme A transporter 1 | +4.23 |
| TERG_02545 | MFS monocarboxylate transporter | −4.89 |
| TERG_02654 | MFS amine transporter | −4.70 |
| TERG_03907 | Amino acid transporter | +5.60 |
| TERG_04093 | K+ homeostasis protein kha1 | −4.21 |
| TERG_12078 | FMN-binding split barrel-like protein | −4.31 |
| TERG_12574 | Tmem1 family | −6.62 |
| Protease activity | ||
| TERG_02988 | Asparaginase | −4.30 |
| Fatty acid and lipid metabolism | ||
| TERG_01347 | Phosphatidyl synthase | +5.40 |
| TERG_12530 | 3-Ketoacyl- thiolase | +4.07 |
| Cell wall | ||
| TERG_03843 | Chitin synthase b | +4.25 |
| Signal transduction pathway | ||
| TERG_12191 | Ras guanyl-nucleotide exchange factor | +4.33 |
| TERG_07570 | G-protein signaling- receptor signaling pathway | −4.11 |
| Drug response | ||
| TERG_01820 | MFS drug transporter | −4.61 |
| TERG_05575 | MFS multidrug transporter | −4.01 |
| Response to stress | ||
| TERG_02795 | Thiazole biosynthetic mitochondrial | −7.22 |
a Only genes with a fold change of four or higher are indicated. + induction; −repression
Table 3.
T. rubrum genes modulated on both keratin and elastin substrates
| Gene Id | Expression change (n-fold)a | Tentative annotation | |
|---|---|---|---|
| Keratin vs control | Elastin vs control | ||
| Metabolic Process | |||
| TERG_00072 | −6.70 | −6.73 | Hypothetical protein |
| TERG_00073 | −5.98 | −7.40 | NADPH dehydrogenase |
| TERG_00374 | −6.72 | −5.09 | Metallophosphoesterase domain-containing protein 2 |
| TERG_00881 | −5.18 | −5.39 | Reticulon-4-interacting protein 1 |
| TERG_01338 | −6.84 | −5.30 | Hydantoinase |
| TERG_02078 | −7.57 | −5.84 | Thiamine biosynthesis protein |
| TERG_02132 | −8.95 | −10.39 | 5-Histidylcysteine sulfoxide synthase |
| TERG_02133 | −4.79 | −6.16 | Flug protein |
| TERG_02134 | −10.00 | −10.03 | Indoleamine -dioxygenase-like protein |
| TERG_02197 | −4.85 | −4.49 | Aliphatic nitrilase |
| TERG_02217 | −5.99 | −4.87 | GNAT family protein |
| TERG_02538 | −4.95 | −4.85 | Carboxylesterase family protein |
| TERG_02712 | −44.76 | −43.66 | Glutamyl-tRNA amidotransferase |
| TERG_03707 | −12.95 | −5.49 | Fusicoccadiene synthase |
| TERG_04073 | −4.98 | −6.64 | Glutathione synthetase |
| TERG_06261 | −6.17 | −5.18 | Phosphoric ester hydrolase |
| TERG_07159 | −4.91 | −4.72 | Prenyltransferase alpha subunit |
| TERG_07504 | −5.71 | −6.65 | Carbohydrate-binding protein |
| TERG_07821 | −8.24 | −11.57 | Hypothetical protein |
| TERG_08054 | −6.35 | −7.18 | Homoserine acetyltransferase family protein |
| TERG_08261 | −4.47 | −5.79 | Glutamate decarboxylase |
| TERG_08554 | −4.37 | −6.20 | Riboflavin-specific deaminase |
| TERG_08787 | −7.01 | −5.61 | Aminotransferase family protein |
| TERG_08868 | −5.02 | −4.38 | FKBP-type peptidyl-prolyl |
| TERG_06540 | −4,10 | −7.32 | Glutathione transferase |
| Regulation of Biological Process | |||
| TERG_01762 | −7.45 | −4.30 | Sulfite reductase beta-component |
| TERG_03972 | −4.43 | −5.42 | Elongation factor g |
| TERG_04862 | −11.93 | −7.97 | C6 sexual development transcription factor |
| TERG_05617 | −6.98 | −8.13 | Hypothetical protein |
| TERG_08437 | −8.34 | −6.36 | C2H2 transcription factor |
| TERG_11890 | −8.78 | −6.75 | Hypothetical protein |
| Transporter | |||
| TERG_04308 | −20.09 | −13.39 | MFS sugar transporter |
| TERG_06954 | −10.53 | −4.10 | Hypothetical protein |
| Protease Activity | |||
| TERG_02001 | −17.10 | −14.96 | Dipeptidyl-peptidase 5 |
| TERG_03104 | −7.94 | −10.17 | Signal peptidase i |
| TERG_04324 | +14.97 | +4.77 | Extracellular metalloproteinase 4 |
| TERG_05842 | −4,08 | −5.55 | Peptidase |
| Fatty acid and lipid metabolism | |||
| TERG_02704 | −12.05 | −19.34 | Short-chain dehydrogenase |
| TERG_05484 | +6.56 | +5.30 | Acyl- dehydrogenase |
| TERG_11720 | −4.39 | −4.65 | Acyl- dehydrogenase |
| Cell Wall | |||
| TERG_00060 | −9.15 | −7.25 | Bys1 domain |
| TERG_05625 | −15.77 | −7.94 | Glycoside hydrolase family 18 protein |
| TERG_11657 | −6.60 | −9.36 | Glycoside hydrolase family 18 protein |
| Drug Response | |||
| TERG_05309 | −5.85 | −6.39 | Puromycin resistance protein pur8 |
| Response to stress | |||
| TERG_01122 | +6.72 | +6.75 | Chaperone heat shock protein |
aOnly genes with a fold change of four or higher are indicated. + induction; −repression
Fig. 3.
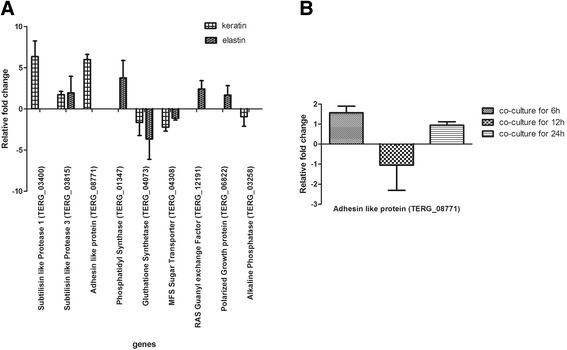
Real-time RT-PCR of selected genes from the microarray hybridization of T. rubrum genes during growth on keratin and elastin compared to control (a). Modulation of the adhesin-like protein gene of T. rubrum co-cultured with keratinocytes compared to fungal conidia (b)
Among those, one gene drew our attention, an adhesin-like protein upregulated in T. rubrum conidia grown in keratin substrate and containing an interesting pattern of tandem repeat sequences related to adhesion and virulence. This gene was also induced when T. rubrum conidia were co-cultured with a human keratinocyte cell line for 6 h and 24 h (Fig. 3b). Importantly, the induction of this gene was not observed in a microarray analysis using T. rubrum mycelium grown on the same protein substrate (data not shown), suggesting that this gene might play a role in the early stages of infection. The function of an adhesin-like protein was predicted using FaaPred software, with the gene showing a high confidence score (0.997).
Besides the in silico characterization of this gene indicated a sequence of 3,030 bp (GenBank Database under the accession number: 327302703), containing a tandem repeat sequence. The tandem repeat pattern is a minisatellite type that shows high variability among dermatophyte species and strains and is located between positions 1,382 and 2,425, with a consensus region of 45 bp and a total length of 1,044 bp. The tandem repeat sequence encodes 348 amino acids; has a repeat unit of glycine, glutamine and proline; and is characterized by the presence of a collagen triple helix domain preceded by a mucin-like glycoprotein domain and flocculin type 3 domain (Fig. 4a). In addition the similarities between MAD1 (Metarhizium anisopliae adhesin) and the T. rubrum adhesin-like protein was verified and exist at the N-terminus, starting in the glycine-rich region. Furthermore, both proteins share a predicted GPI cell wall anchor site at the C-terminus and exhibit a tandem repeat sequence in the mid-region (Figs. 4b and c). Similarly, the findings showed similarities between the T. rubrum adhesin-like protein and a cell surface protein of Aspergillus fumigatus (cspA - Afu3g08990), which is characterized by a 188-amino acid serine/threonine/proline-rich N-terminus followed by a large, variable, six-amino acid serine/proline [PGQPS (A/V)] tandem repeat region (Fig. 4d). In the last case, besides the tandem repeats, both proteins also contain collagen and flocculin domains and a GPI anchor site.
Fig. 4.
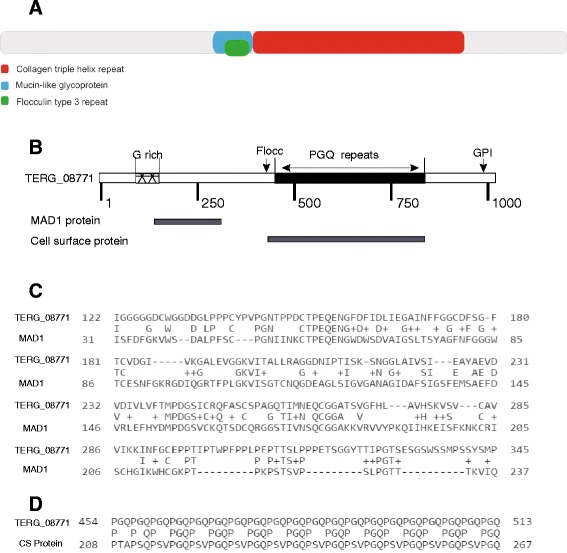
Structural features of TERG_08771 adhesin-like protein a) Main domains in the adhesin-like protein sequence according to the NCBI’s conserved domain database. b Schematic structure of TERG_08771 showing a glycine rich region, tandem repeats of proline, glycine and glutamine, and a glycosylphosphatidylinositol anchor site (GPI). c Alignment of the homologous region between TERG_08771 and MAD1. d Conservation of tandem repeat regions between TERG_08771 and cell surface protein (CS protein) of Aspergillus fumigatus
The adhesin-like protein gene of T. rubrum has homologous genes in dermatophyte species, which indicate that the repetitive units are conserved between species. Variation of the extent of the tandem repeats can be observed among species and strains (Fig. 5).
Fig. 5.
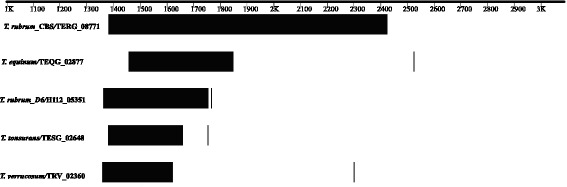
Variable numbers of tandem repeat sequences in the adhesion-like protein gene of different dermatophytes. The vertical bars represent the end of the transcribed genes of the different species and strains
Discussion
The establishment of T. rubrum infection initiates by adhesion to the tissue surface mediated through the release of carbohydrate-binding adhesins by arthroconidia that bind to surface of host receptors [8, 17, 18]. The conidia in the dormant stage begins to germinate and then hyphae readily penetrate the stratum corneum, preventing the fungus to be disconnected from the skin due to flaking of the keratinized epithelium [13]. After adhesion, dermatophytes secrete a wide range of enzymes such as proteases, keratinases, lipase, elastase, collagenase, phosphatases and esterases, which are important factors during the infectious process [19–21]. The keratinase secreted by dermatophytes catalyze the degradation of keratin present in the host tissue into oligopeptides or amino acids, which can then be assimilated by the fungi [22].
In this study, the use of different protein sources such as keratin and elastin did not induce a profile of gene expression which would be characteristic of the superficial or deep infection, respectively. This could be attributed to the dormant stage of conidia, because the status of quiescence may be responsible for slow transcriptional profile due to starvation of nutrients. However, it is believed that there is a specific induction of proteases with respect to the protein source used [23]. In this work it was observed in fact that fungal growth in the presence of keratin promoted the induction of a greater number of proteases, specifically of the subtilisin family of proteases (Sub 1 and Sub 3) and leucine aminopeptidase (Lap 1 and Lap 2) compared to growth of the fungus in elastin.
In silico identification and prediction of a gene coding an adhesin-like protein induced by keratin
In our microarray data an adhesin like protein was modulated during the growth of T. rubrum on keratin and also in co-culture in a keratinocyte cell. Adhesins are also required for the early stages of infection in dermatophytes [8]. The gene expression modulation of T. rubrum adhesin like-protein during the co-culture of conidia in keratinocytes cell line assessed by qPCR showed an increase in gene expression 6 h followed by a decrease at 12 h and a subsequent increase at 24 h. Liu et al. [16] demonstrated that dormant conidia of T. rubrum show a fluctuation on gene expression. During the germination process, the same work followed a different time course of conidia development and the morphological transitions promoted during the time was also evaluated indicating that at 6 h the conidia is brighter and swollen and after 12 h the hyphae begin to be developed. As the adhesins are mainly expressed in germinating conidia, as occoured for MAD1 in M. anisopliae [24], the result obtained by qPCR for the expression of T. rubrum adhesin-like protein was even expected for the incubation of 6 h. Furthermore, Aljabre et al. [25] studying the co-culture of Trichophyton mentagrophytes in corneocytes cells showed that the adherence of the arthroconidia requires 6 h and more than 4 h for germination. Regarding to the time of 24 h, some works also demonstrated the adherence of spores to corneum stratum for this time [26] . Thus, our assumption is that the adhesin-like protein has a fluctuation on gene expression according to the cell cycle process.
In silico analysis of this adhesin like protein showed the presence of collagen triple helix domains, mucin-like glycoprotein domain and flocullin domain, as described previously. The collagen triple helix domains are currently being investigated because of their role in host-pathogen interactions and bacterial adhesins [27]. The mucin-like glycoprotein domain of these proteins plays a role in the adhesion and pathogenicity of microorganisms, including biofilm formation [28]. The flocculin domain is present in many cell wall proteins (CWPs) with tandem repeats that are responsible for mediating the host-pathogen interaction by cell-cell adhesion, and it is related to the biofilm formation observed in Aspergillus fumigatus and Saccharomyces cerevisiae [29, 30]. The fungal adhesion process has been extensively studied in Candida spp. [31], but it has only begun to be addressed in other pathogenic fungi. In this respect, an adhesin called MAD1 has been characterized in the entomopathogenic fungus Metarhizium anisopliae. The disruption of MAD1 adhesin delays germination, suppresses blastospore formation, and reduces virulence against caterpillars [24]. Also, the cell surface protein of Aspergillus fumigatus, which shows features of adhesin-like protein was investigated, and a null mutant for the cspA gene showed a phenotype characterized by rapid conidial germination and reduced adhesion to the extracellular matrix [32]. Tandem repeats are more commonly found in cell wall proteins, and the number of repeats, as well as the length of the tandem repeat unit, can vary among different species and within isolates, promoting diversity and improving adhesion capacity [29].
Expression of genes involved in dormancy and germination of conidia
Genes involved in the specific germination processes of conidia are interesting targets for the development of new antifungal compounds. Furthermore, the chronic infections caused by these fungi may be related to conidial dormancy because an important factor underlying chronicity is the ability of T. rubrum to survive as conidia inside the human body [33]. The pattern of low metabolic activity reflected by the repression of a high number of genes involved in the metabolism process, also known as quiescent status, seems to be related to conidial dormancy [16, 20]. While at the same time the induction of a few genes in energy metabolism probably are important for maintaining dormancy and initiating germination [20].
The modulation of genes encoding signal transduction system proteins and regulation of biologic process that are involved in conidial dormancy and the early stages of spore germination was also observed in the present study [34]. Moreover, we found that elastin promoted modulation of a higher percentage of genes involved in the regulation of biological processes, which may be related to the establishment of deep infections. Indeed, studies have shown that genes coding proteins required for polarized growth and WD-repeat proteins (related to the interaction of signaling molecules) appear to be important for the processes of systemic fungal infections caused by A. fumigatus and C. albicans [35, 36]. However, further studies are needed to better understand the role of signal transduction systems and regulation of biological process in the maintenance of dormancy in T. rubrum conidia.
Expression of genes coding proteases
Proteases play a central role in pathogenesis, as they are widely implicated to have proteolytic activity [37, 38]. At least 20 protease gene that belong to the metalloprotease and serine protease families of proteolytic enzymes are found in the genome of T. rubrum and other dermatophytes [39]. Among endoproteases, there are five metalloproteases (fungalysins) and seven serine proteases (subtilisins). The exoproteases are represented by two metalloproteases, leucine aminopeptidases Lap1 and Lap2, and two serine proteases, dipeptidyl-peptidases DppIV and DppV [38]. In addition to these proteases, T. rubrum was also found to secrete a metallocarboxypeptidase (McpA) and to produce two membrane-anchored serine carboxypeptidases when cultured in medium containing protein as the sole nitrogen and carbon source [40]. The proteases modulated in this work, especially during the growth of T. rubrum on keratin, are relevant to better understand the role of these proteases for the conidia infection process.
Conclusion
The present results broaden the knowledge of the molecular features of infection with T. rubrum conidia. Genes involved in conidial adhesion and dormancy seem to be important for the infection process and could be explored as potential targets for the development of new antifungal agents.
Methods
Strain, media and growth conditions
T. rubrum strain CBS118892 was cultured on Sabouraud dextrose agar (Oxoid, Hampshire, England) for 15 days at 28 °C to induce full sporulation. The conidial solution was filtered twice through glass wool to remove any hyphal fragments and was inspected by microscopy. The number of conidia was counted with a hemocytometer under a Nikon microscope. Approximately 2.6 x 106 conidia/mL were added to 10 mL of three different media in triplicate: i) Cove’s minimal medium (control) containing 70 mM nitrate (Sigma Aldrich, St. Louis, MO, USA) and 50 mM glucose (Sigma Aldrich); ii) Cove’s minimal medium supplemented with 0.5 % bovine keratin; and iii) Cove’s medium supplemented with 0.25 % elastin (Sigma Aldrich). Cultures ii and iii received 3.5 mM nitrate and 2.7 mM glucose. T. rubrum cultures (i, ii, and iii) were incubated for 24, 36 and 72 h at 28 °C under shaking (130 rpm) and were collected by centrifugation at 1,000 g for 10 min.
Co-culture conditions
The human keratinocyte cell line HaCat was grown in RPMI medium (Sigma Aldrich) supplemented with 10 % fetal bovine serum (FBS) (Sigma Aldrich) at 37 °C in 5 % CO2. Keratinocytes were collected, washed, and counted with a hemocytometer. A total of 2.5 x 105 cells/mL were plated in 250-mL tissue culture flasks containing RPMI supplemented with 2 % FBS and grown for 24 h at 37 °C in 5 % CO2. The T. rubrum solution containing 1 x 107 conidia/mL was resuspended in RPMI medium containing 2 % FBS. The solution of conidia was added to the keratinocyte cultures and incubated for 6 h, 12 h and 24 h at 37 °C in 5 % CO2. Fungi and human cells were collected by scraping with a rubber scraper, transferred to 1.5-mL microtubes, and centrifuged at 1,730 g for 10 min.
Total RNA extraction
T. rubrum cultures (i, ii, and iii) grown for 24 h or co-cultured with keratinocytes were treated with lysis solution (20 mg/mL lysozyme, 0.7 M KCl and 1 M MgSO4, pH 6.8) for 1 h at 28 °C while shaking (130 rpm) and were collected by centrifugation at 1,000 g for 10 min. The cells were ground with a mortar and pestle and pulverized in liquid nitrogen. Total RNA was extracted using the Illustra RNAspin Mini RNA Isolation Kit (GE Healthcare-Little Chalfont, Buckinghamshire, UK). RNA preparations were confirmed to be free of protein and phenol by UV spectrophotometry. RNA degradation was assessed by microfluidic electrophoresis using Agilent 6000 RNA Nano chips and an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Only RNA samples that were free of protein and phenol and had an RNA integrity number (RIN) ≥ 9.0 were used.
Microarray hybridization
Twenty-five nanograms of RNA from each incubation time (24, 36 and 72 h) and treatment condition (control, keratin and elastin) were pooled. Finally, 75 ng of RNA from each condition was used to synthesize double-stranded cDNA and cyanine 3 (Cy3)-CTP labelled complementary amplified RNA (cRNA) using the Agilent Low Input Amplification Kit (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s instructions. Agilent 4 × 44 K High-Density Oligonucleotide custom microarray slides were designed with the e-array tool (Agilent Technology Genomics). The ESTs (expressed sequence tags) selected were obtained from the NCBI database (www.ncbi.nlm.nih.gov/), the mapping of these ESTs with the genome of T. rubrum retrieved in 6,091 encoding genes. The cyanine-labelled complementary RNA was hybridized to microarrays slides (Agilent Technologies) in SureHyb chambers (Agilent) in a rotator oven for 18 h at 60 °C; two biological replicates were used for each condition. Internal control probes were included in addition to the functional genes of T. rubrum. The arrays were washed according to the manufacturer’s instructions and scanned with an Agilent DNA Microarray Scanner.
Microarray data analysis
The oligo-mRNA array slides were scanned with a DNA microarray scanner (Agilent Technologies), and Agilent Feature Extraction 10.5 software was used to extract the hybridization signals. The analysis was performed by pairwise comparison of keratin x control or elastin x control. The quantitative microarray data were normalized with a 95th percentile expression filter and were analysed using the Gene Spring GX 12.6 Bioinformatics Platform (http://www.agilent.com/chem/genespring) according to the manufacturer’s instructions. Statistical analysis was performed by ANOVA (P < 0.05) using a fold change ≥ 4.0. The mapping of each EST with a protein coding gene in T. rubrum genome was obtained through alignments performed with Blastx (e-value 1e-5). Also, the putative annotations were retrieved using Blastx according to ncbi bank, then the biological function of putative protein was assessed through GO terms obtained with BLAST2GO 2.4.8 software. Some additional information of relevant genes was obtained from NCBI’s conserved domain database. After the Blast2GO analysis, some genes did not present any GO associated term and in these cases these genes were described as “Unclassified”. The raw data are deposited in the Gene Expression Omnibus (GEO) (www.ncbi.nlm.nih.gov/geo) database under accession number GSE 69305.
In silico identification and prediction of the adhesin-like protein gene and protein sequence analysis
Annotation of the TERG_08771 gene occurred in May 2014 through the Broad Institute’s Dermatophyte Comparative Database, and the gene was assigned as a hypothetical protein. In silico identification of this gene was performed using the following tools: Blast2GO [41], Funcat [42], NCBI Blast, and CDD (Conserved Domain Database) [43]. Blastx from NCBI with an e-value of 1e-05 was used to identify homologies. Additionally, the TERG_08771 gene was submitted to the FaaPred prediction method for fungal adhesins and adhesin-like proteins (http://bioinfo.icgeb.res.in/faap/query.html) [9]. A tandem repeat analysis of the TERG_08771 gene between dermatophytes was performed with Tandem Repeat Finder [44] using the following parameters: matching weight, 2; mismatching penalty, 5; indel penalty, 5; match probability, 0.8; indel probability, 0.1; score ≥ 40; and maximum period, 500. Variability analysis was performed using the SERV algorithm [45].
Quantitative RT-PCR
A set of nine genes (Table 4) was selected for quantitative RT-PCR to validate the microarray expression data. Additionally, the expression of TERG_08771 (adhesion-like protein) was evaluated in a co-culture with the keratinocyte cell line. Complementary DNA was synthesized from 500 ng of total RNA in a 20-μL reaction volume using the RevertAID H Minus First Strand cDNA Synthesis Kit (Fermentas®). The quantitative RT-PCR experiments were performed in triplicate using the SYBR Taq Ready Mix Kit (Sigma) on an Mx3300 QPCR system (Stratagene) according to Bitencourt et al. [46]. The cycling conditions were as follows: initial denaturation at 94 °C for 10 min, followed by 40 cycles at 94 °C for 2 min, 60 s at 60 °C and 1 min at 72 °C. A dissociation curve was constructed at the end of each PCR cycle to verify if a single product was amplified. Expression levels were calculated by the comparative Ct method using beta-tubulin for normalization. The reference for validation of the microarray data was Cove’s minimal medium, and for the co-culture assay the reference used was the dormant conidia solution according to Komoto et al. [47] with some modifications. The results are reported as the mean ± standard deviation of three independent experiments.
Table 4.
Primers used in RT-PCR
| Primer | Sequence | GI number | Size (bp) | Reference |
|---|---|---|---|---|
| Adhesin like protein | F:5′- CTGCGCAGTTGTTATCAAGAAG-3′ | 327302703 | 98 | This paper |
| R: 5′- GTAGGCTGGTAGTTGGGAATG-3′ | ||||
| Subtilin 1 | F: 5′- GCTGGCTCCAATCTACTCATAC-3′ | 327303325 | 105 | This paper |
| R:5′- CGCTGTATCCCTTCATCTTGT-3′ | ||||
| Subtilisin 3 | F: 5′- AGGTTAGTCCTGAAGCCCTCT-3′ | 38146042 | 105 | This paper |
| R: 5′- GCGGTCGTGCTCTACATAGT-3′ | ||||
| Phosphatidyl synthase | F: 5′- CCAAGAGTCCGCCGTCTATC-3′ | 327309347 | 179 | This paper |
| R: 5′- GGTGTGACTTCGGCAGATGA-3′ | ||||
| Glutathione synthetase | F: 5′- ACTGACTGGCTGGGAGAGAT-3′ | 327300656 | 124 | This paper |
| R: 5′- ACAAGCCAAGTGAGAGAGGC-3′ | ||||
| MFS sugar transport | F: 5′- AAACCACCGCCTCGTTATGT-3′ | 327301116 | 127 | This paper |
| R: 5′- GATGGCCAAAAGACCCGGTA-3′ | ||||
| Tubulin beta chain | F: 5′- AACATGATGGCTGCCACTGA-3′ | 10371186 | 253 | [48] |
| R: 5′ - AAGATGGCAGAGCAGGTAAGGT-3′ |
Availability of data and materials
The dataset supporting the conclusions of this article is available in the Expression Omnibus (GEO) in http://www.ncbi.nlm.nih.gov/geo database under accession number GSE 69305.
Acknowledgements
This study was supported by grants from Fundação de Amparo à Pesquisa do Estado de São Paulo (Process numbers 2014/23841-3 and 2012/03845-9), a Fapesp doctoral fellowship granted to TAB (Process number 2012/02920-7), a CAPES doctoral fellowship granted to MEF and a master’s degree fellowship granted to TTK. We thank the staff of the Biotechnology Unit, UNAERP, Instituto Federal do Sul de Minas - Campus Machado for their general support and Professor Nilce M. Martinez-Rossi for kindly providing the T. rubrum strain CBS.
Footnotes
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
TAB performed the laboratory experiments and bioinformatics analysis and wrote the manuscript. CM hybridized the microarray. MEF performed the bioinformatics analysis and the in silico identification of the adhesin-like protein gene. AFA performed the bioinformatics analysis. TTK performed the co-culture experiments and the analysis of adhesin-like protein gene expression by RT-PCR. EGS and ROB discussed the manuscript. IM customized the T. rubrum microarray slide and wrote the manuscript. MM supervised the research and contributed reagents and materials. ALF designed the project, supervised the research, contributed reagents/materials/analysis tools and wrote the manuscript. All the authors have read and approved the final manuscript.
References
- 1.Borman AM, Campbell CK, Fraser M, Johnson EM. Analysis of the dermatophyte species isolated in the British Isles between 1980 and 2005 and review of worldwide dermatophyte trends over the last three decades. Med Mycol. 2007;45(2):131–141. doi: 10.1080/13693780601070107. [DOI] [PubMed] [Google Scholar]
- 2.Nenoff P, Kruger C, Ginter-Hanselmayer G, Tietz HJ, Nenoff P, Kruger C, Ginter-Hanselmayer G, Tietz HJ. Mycology - an update. Part 1: Dermatomycoses: causative agents, epidemiology and pathogenesis. J Dtsch Dermatol Ges. 2014;12(3):188–209. doi: 10.1111/ddg.12245. [DOI] [PubMed] [Google Scholar]
- 3.Cafarchia C, Iatta R, Latrofa MS, Graser Y, Otranto D. Molecular epidemiology, phylogeny and evolution of dermatophytes. Infect Genet Evol. 2013;20:336–351. doi: 10.1016/j.meegid.2013.09.005. [DOI] [PubMed] [Google Scholar]
- 4.Nir-Paz R, Elinav H, Pierard GE, Walker D, Maly A, Shapiro M, Barton RC, Polacheck I. Deep infection by Trichophyton rubrum in an immunocompromised patient. J Clin Microbiol. 2003;41(11):5298–5301. doi: 10.1128/JCM.41.11.5298-5301.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.da Silva BCM, Paula CR, Auler ME, Ruiz LS, dos Santos JI, Yoshioka MCN, Fabris A, Castro LGM. 9 Duarte AJdS, Gambale W. Dermatophytosis and immunovirological status of HIV-infected and AIDS patients from Sao Paulo city. Brazil Mycoses. 2014;57(6):371–6. doi: 10.1111/myc.12169. [DOI] [PubMed] [Google Scholar]
- 6.Quatresooz P, Piérard-Franchimont C, Arrese JE, Piérard GE. Clinicopathologic presentations of dermatomycoses in cancer patients. J Eur Acad Dermatol Venereol. 2008;22(8):907–917. doi: 10.1111/j.1468-3083.2008.02774.x. [DOI] [PubMed] [Google Scholar]
- 7.Kaufman G, Berdicevsky I, Woodfolk JA, Horwitz BA. Markers for host-induced gene expression in Trichophyton dermatophytosis. Infect Immun. 2005;73(10):6584–6590. doi: 10.1128/IAI.73.10.6584-6590.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Peres NT, Sanches PR, Falcao JP, Silveira HC, Paiao FG, Maranhao FC, Gras DE, Segato F, 17 Cazzaniga RA, Mazucato M et al. Transcriptional profiling reveals the expression of novel genes in response to various stimuli in the human dermatophyte Trichophyton rubrum. BMC Microbiol. 2010;10:39. [DOI] [PMC free article] [PubMed]
- 9.Ramana J, Gupta D. FaaPred: a SVM-based prediction method for fungal adhesins and adhesin-like proteins. PLoS One. 2010;5(3):e9695. doi: 10.1371/journal.pone.0009695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kashi Y, King DG. Simple sequence repeats as advantageous mutators in evolution. Trends Genet. 2006;22(5):253–259. doi: 10.1016/j.tig.2006.03.005. [DOI] [PubMed] [Google Scholar]
- 11.Verstrepen KJ, Klis FM. Flocculation, adhesion and biofilm formation in yeasts. Mol Microbiol. 2006;60(1):5–15. doi: 10.1111/j.1365-2958.2006.05072.x. [DOI] [PubMed] [Google Scholar]
- 12.Dranginis AM, Rauceo JM, Coronado JE, Lipke PN. A biochemical guide to yeast adhesins: glycoproteins for social and antisocial occasions. Microbiol Mol Biol Rev. 2007;71(2):282–294. doi: 10.1128/MMBR.00037-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Baldo A, Monod M, Mathy A, Cambier L, Bagut ET, Defaweux V, Symoens F, Antoine N, Mignon 30 B. Mechanisms of skin adherence and invasion by dermatophytes. Mycoses. 2012;55(3):218–23. [DOI] [PubMed]
- 14.Liu T, Xu X, Leng W, Xue Y, Dong J, Jin Q. Analysis of gene expression changes in Trichophyton rubrum after skin interaction. J Med Microbiol. 2014;63(Pt 5):642–648. doi: 10.1099/jmm.0.059386-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zaugg C, Monod M, Weber J, Harshman K, Pradervand S, Thomas J, Bueno M, Giddey K, Staib P. Gene expression profiling in the human pathogenic dermatophyte Trichophyton rubrum during growth on proteins. Eukaryotic Cell. 2009;8(2):241–250. doi: 10.1128/EC.00208-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Liu T, Zhang Q, Wang L, Yu L, Leng W, Yang J, Chen L, Peng J, Ma L, Dong J, et al. The use of global transcriptional analysis to reveal the biological and cellular events involved in distinct development phases of Trichophyton rubrum conidial germination. BMC Genomics. 2007;8:100. doi: 10.1186/1471-2164-8-100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Esquenazi D, de Souza W, Alviano CS, Rozental S. The role of surface carbohydrates on the interaction of microconidia of Trichophyton mentagrophytes with epithelial cells. FEMS Immunol Med Microbiol. 2003;35(2):113–123. doi: 10.1016/S0928-8244(03)00007-5. [DOI] [PubMed] [Google Scholar]
- 18.Esquenazi D, Alviano CS, de Souza W, Rozental S. The influence of surface carbohydrates during in vitro infection of mammalian cells by the dermatophyte Trichophyton rubrum. Res Microbiol. 2004;155(3):144–153. doi: 10.1016/j.resmic.2003.12.002. [DOI] [PubMed] [Google Scholar]
- 19.Brouta F, Descamps F, Monod M, Vermout S, Losson B, Mignon B. Secreted metalloprotease gene family of Microsporum canis. Infect Immun. 2002;70(10):5676–5683. doi: 10.1128/IAI.70.10.5676-5683.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Leng W, Liu T, Li R, Yang J, Wei C, Zhang W, Jin Q. Proteomic profile of dormant Trichophyton rubrum conidia. BMC Genomics. 2008;9:303. doi: 10.1186/1471-2164-9-303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Vermout S, Tabart J, Baldo A, Mathy A, Losson B, Mignon B. Pathogenesis of dermatophytosis. Mycopathologia. 2008;166(5–6):267–275. doi: 10.1007/s11046-008-9104-5. [DOI] [PubMed] [Google Scholar]
- 22.Lechenne B, Reichard U, Zaugg C, Fratti M, Kunert J, Boulat O, Monod M. Sulphite efflux pumps in Aspergillus fumigatus and dermatophytes. Microbiology. 2007;153(Pt 3):905–913. doi: 10.1099/mic.0.2006/003335-0. [DOI] [PubMed] [Google Scholar]
- 23.Chen J, Yi J, Liu L, Yin S, Chen R, Li M, Ye C, Zhang YQ, Lai W. Substrate adaptation of Trichophyton rubrum secreted endoproteases. Microb Pathog. 2010;48(2):57–61. doi: 10.1016/j.micpath.2009.12.001. [DOI] [PubMed] [Google Scholar]
- 24.Wang C, St Leger RJ. The MAD1 adhesin of Metarhizium anisopliae links adhesion with blastospore production and virulence to insects, and the MAD2 adhesin enables attachment to plants. Eukaryotic cell. 2007;6(5):808–816. doi: 10.1128/EC.00409-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Aljabre SH, Richardson MD, Scott EM, Rashid A, Shankland GS. Adherence of arthroconidia and germlings of anthropophilic and zoophilic varieties of Trichophyton mentagrophytes to human corneocytes as an early event in the pathogenesis of dermatophytosis. Clin Exp Dermatol. 1993;18(3):231–235. doi: 10.1111/j.1365-2230.1993.tb02176.x. [DOI] [PubMed] [Google Scholar]
- 26.Duek L, Kaufman G, Ulman Y, Berdicevsky I. The pathogenesis of dermatophyte infections in human skin sections. J Infect. 2004;48(2):175–180. doi: 10.1016/j.jinf.2003.09.008. [DOI] [PubMed] [Google Scholar]
- 27.Berisio R, Vitagliano L. Polyproline and triple helix motifs in host-pathogen recognition. Curr Protein Pept Sci. 2012;13(8):855–865. doi: 10.2174/138920312804871157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Karunanithi S, Vadaie N, Chavel CA, Birkaya B, Joshi J, Grell L, Cullen PJ. Shedding of the mucin-like flocculin Flo11p reveals a new aspect of fungal adhesion regulation. Curr Biol. 2010;20(15):1389–1395. doi: 10.1016/j.cub.2010.06.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Levdansky E, Sharon H, Osherov N. Coding fungal tandem repeats as generators of fungal diversity. Fungal Biol Rev. 2008;22(3–4):85–96. doi: 10.1016/j.fbr.2008.08.001. [DOI] [Google Scholar]
- 30.Verstrepen KJ, Jansen A, Lewitter F, Fink GR. Intragenic tandem repeats generate functional variability. Nat Genet. 2005;37(9):986–990. doi: 10.1038/ng1618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sheppard DC, Yeaman MR, Welch WH, Phan QT, Fu Y, Ibrahim AS, Filler SG, Zhang M, Waring AJ, Edwards JE., Jr Functional and structural diversity in the Als protein family of Candida albicans. J Biol Chem. 2004;279(29):30480–30489. doi: 10.1074/jbc.M401929200. [DOI] [PubMed] [Google Scholar]
- 32.Levdansky E, Kashi O, Sharon H, Shadkchan Y, Osherov N. The Aspergillus fumigatus cspA gene encoding a repeat-rich cell wall protein is important for normal conidial cell wall architecture and interaction with host cells. Eukaryotic Cell. 2010;9(9):1403–1415. doi: 10.1128/EC.00126-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Dahl MV, Grando SA. Chronic dermatophytosis: what is special about Trichophyton rubrum? Adv Dermatol. 1994;9:97–109. [PubMed] [Google Scholar]
- 34.Osherov N, May GS. The molecular mechanisms of conidial germination. FEMS Microbiol Lett. 2001;199(2):153–160. doi: 10.1111/j.1574-6968.2001.tb10667.x. [DOI] [PubMed] [Google Scholar]
- 35.Brand A. Hyphal growth in human fungal pathogens and its role in virulence. Int J Microbiol. 2012;2012:517529. doi: 10.1155/2012/517529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Liu X, Nie X, Ding Y, Chen J. Asc1, a WD-repeat protein, is required for hyphal development and virulence in Candida albicans. Acta Biochim Biophys Sin. 2010;42(11):793–800. doi: 10.1093/abbs/gmq093. [DOI] [PubMed] [Google Scholar]
- 37.Krappmann S, Braus GH. Nitrogen metabolism of Aspergillus and its role in pathogenicity. Med Mycol. 2005;43(Suppl 1):S31–40. doi: 10.1080/13693780400024271. [DOI] [PubMed] [Google Scholar]
- 38.Monod M. Secreted proteases from dermatophytes. Mycopathologia. 2008;166(5–6):285–294. doi: 10.1007/s11046-008-9105-4. [DOI] [PubMed] [Google Scholar]
- 39.Rawlings ND, Salvesen G: Handbook of Proteolytic Enzymes vol. 1, 2 and 3. London, UK. Elsevier; 2013.
- 40.Zaugg C, Jousson O, Léchenne B, Staib P, Monod M. Trichophyton rubrum secreted and membrane-associated carboxypeptidases. Int J Med Microbiol. 2008;298(7–8):669–682. doi: 10.1016/j.ijmm.2007.11.005. [DOI] [PubMed] [Google Scholar]
- 41.Conesa A, Götz S, Garcia-Gomez JM, Terol J, Talón M, Robles M. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics. 2005;21:3674–3676. doi: 10.1093/bioinformatics/bti610. [DOI] [PubMed] [Google Scholar]
- 42.Ruepp A, Zollner A, Maier D, Albermann K, Hani J, Mokrejs M. The FunCat, a functional annotation scheme for systematic classification of proteins from whole genomes. Nucleic Acids Res. 2004;32:5539–5545. doi: 10.1093/nar/gkh894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Bryant SH. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2011;39:D225–D229. doi: 10.1093/nar/gkq1189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Benson G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 1999;27(2):573–580. doi: 10.1093/nar/27.2.573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Legendre M, Pochet N, Park T, Verstrepen K. Sequence-based estimation of minisatellite and microsatellite repeat variability. Genome Res. 2007;17:1787–1796. doi: 10.1101/gr.6554007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Bitencourt TA, Komoto TT, Massaroto BG, Miranda CE, Beleboni RO, Marins M, Fachin AL. Trans-chalcone and quercetin down-regulate fatty acid synthase gene expression and reduce ergosterol content in the human pathogenic dermatophyte Trichophyton rubrum. BMC Complement Altern Med. 2013;13:229. doi: 10.1186/1472-6882-13-229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Komoto TT, Bitencourt TA, Silva G, Beleboni RO, Marins M, Fachin AL. Gene Expression Response of Trichophyton rubrum during Coculture on Keratinocytes Exposed to Antifungal Agents. Evid Based Complement Alternat Med. 2015;2015:7. doi: 10.1155/2015/180535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jacob TR, Peres NTA, Persinoti GF, Silva LG, Mazucato M, Rossi A, Martinez-Rossi NM. Rpb2 is a reliable reference gene for quantitative gene expression analysis in the dermatophyte Trichophyton rubrum. Medical Mycology 2012;50(4):368–377. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The dataset supporting the conclusions of this article is available in the Expression Omnibus (GEO) in http://www.ncbi.nlm.nih.gov/geo database under accession number GSE 69305.


