Abstract
Tcra/Tcrd includes 2 genes with distinct developmental programs controlled by 2 distant enhancers, Eα and Eδ. These enhancers work as a developmental switch during thymocyte development and they are essential for generation of αβ and γδ T-lymphocytes. Tcra and Tcrd transit from an unrearranged configuration to a rearranged configuration during T-cell development. Eα and Eδ are responsible for transcription of their respective unrearranged genes in thymocytes but are dispensable for such functions in the context of the rearranged genes in mature T-cells. Interestingly, Eα activates transcription of the rearranged Tcrd in γδ T-lymphocytes but it is inactive in αβ T-lymphocytes.
Keywords: chromatin, enhancer, promoter; transcription; T-lymphocytes; thymocytes; T-cell development; T-cell receptor
Abbreviations
- bp
base pairs
- Cα
Tcra constant region
- Cδ
Tcrd constant region
- D
diversity
- DN
CD4−CD8− double negative
- DP
CD4+CD8+ double-positive
- Eα
Tcra enhancer
- Eδ
Tcrd enhancer
- Eγ
Tcrg enhancer
- eDP
early DP
- IL-7
interleukin 7
- ISP
immature single positive
- H3K4me1
histone 3 lysine 4 monomethylation
- H3K4me3
histone 3 lysine 4 trimethylation
- J
joining
- lDP
late DP
- RNAP II
RNA polymerase II
- SP
single-positive
- TCR
T-cell receptor
- TF
transcription factor
- V
variable.
Developmental Regulation of Transcription by Distant Enhancers
Stem cells from multicellular organisms must differentiate to generate a broad range of specialized cells containing the same genome. This process is possible because of the presence of regulatory DNA elements throughout the genome known as enhancers that dictate the precise spatiotemporal patterns of gene expression during development through the binding of cell lineage or signal-dependent transcription factors (TFs).1 Enhancers are composed of clusters of TF binding sites and typically span a few hundred base pairs (bp) located in intergenic regions, introns and exons far away from their target genes. Enhancers can be classified as inactive, primed, poised, or active. Among all of the millions of potential enhancers that are present in the mammalian genome, only a small subset of them is active in a given cell type. Inactive enhancers are buried in compact chromatin and are devoid of TFs and histone modifications. Primed enhancers are non-functional elements that are characterized by closely bound TFs in a nucleosome-free region and can be identified by specific epigenetic marks such as the enrichment of histone H3 lysine 4 monomethylation (H3K4me1) and the depletion of H3K4 trimethylation (H3K4me3), compared with promoters. The priming of enhancers is triggered by the binding of lineage-determining or pioneer TFs that, together with other TFs, form a compact nucleoprotein complex that binds to DNA or enhanceosome. Poised enhancers are primed enhancers that also contain repressive epigenetic marks such as histone H3 lysine 27 trimethylation and are commonly found in early stem cells. Primed enhancers can become activated by the recruitment of signal-dependent TFs. Binding of signal-dependent TFs to primed enhancers enables broadly expressed inducible TFs to regulate gene expression in a cell type-specific fashion. Active enhancers have been associated with histone 3 lysine 27 acetylation and the presence of transcribing RNA polymerase 2 (RNAP II), which triggers the transcription of enhancers. During differentiation, specific primed enhancers can be rapidly turned on while others can be rapidly turned off at specific time points in particular cell lineages, acting as independent gene switches through the recruitment of signal-dependent TFs to control unique sets of genes in complex spatiotemporal patterns. Active enhanceosomes are able to recruit the mediator complex, cohesin, TFIID, histone-modifying enzymes, or ATP-dependent chromatin-modifying complexes to interact with specific promoters to recruit RNAP II to the promoter and activate transcription initiation and elongation. Here, the current knowledge of the pivotal role of the 2 distant enhancers present in the Tcra/Tcrd locus during the development of αβ and γδ T-cell lineages is reviewed.
V(D)J Recombination of T-Cell Receptor (TCR) Loci During Thymocyte Development
During thymic T-cell development (Fig. 1), T-cell progenitors mature progressively through different thymocyte populations that can be distinguished based on the expression of CD4 and CD8 surface markers: CD4−CD8− double-negative (DN) thymocytes, immature single-positive (ISP) CD8+ cells in mouse and CD4+ cells in human thymocytes, CD4+CD8+ double-positive (DP) thymocytes, and CD4+ or CD8+ single-positive (SP) thymocytes.2 Among the DN population, 4 subpopulations can be further distinguished based on the expression of CD25 and CD44 surface markers: DN1 (CD44+CD25−), DN2 (CD44+ CD25+), DN3 (CD44−CD25+), and DN4 (CD44−CD25−). In addition, 2 DN3 thymocyte subpopulations can be distinguished based on the expression of the CD27 surface marker: DN3a (CD27low) and DN3b (CD27high).3 DN3a thymocytes are quiescent small cells whereas DN3b are large cycling cells. The early DN1-DN3a thymocytes are precursors of the 2 existing mature T-lymphocyte populations: αβ and γδ T-lymphocytes. γδ T-lymphocytes derive directly from DN2 and DN3a thymocytes, whereas αβ T-lymphocytes derive from the transit of DN3a thymocytes through the DN3b, DN4, ISP, early DP (eDP), late DP (lDP), and SP thymocyte populations before migrating to the periphery as mature αβ T-cells. During DN3a thymocytes to DP thymocyte differentiation, cells undergo extensive proliferation. Two DP populations can be distinguished based on the expression of CD71: early DP (eDP) thymocytes (CD71+) and late DP (lDP) thymocytes (CD71−).4,5 eDP thymocytes are large cells that have recently differentiated from ISP cells and constitute approximately 10% of total DP thymocytes, whereas lDP thymocytes are small resting cells (Fig. 1).4
Figure 1.
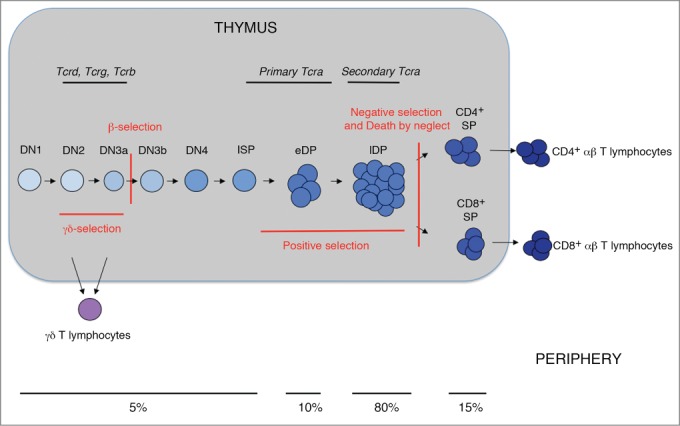
V(D)J rearrangement at TCR loci during the development and generation of αβ and γδ T-lymphocytes. Schematic diagram of thymocyte maturation depicting the various developmental stages and periods of TCR locus rearrangements. β- and γδ-selection, which depends on the expression of a pre-TCR or a TCRγδ, respectively, are indicated in red. The positive selection, negative selection, and death by neglect events are also indicated in red. Commitment to the T-cell lineage is indicated by the transition from light blue to dark blue (αβ T-lymphocytes) or purple (γδ T-lymphocytes). The approximate percentage of thymocytes present in the different populations is indicated.
Thymocytes receive different signals during T-cell development.2 During the first stages of thymocyte maturation, Notch signaling promotes the development of DN1 thymocytes to DN3a thymocytes, committing the cells to the T-cell lineage, whereas interleukin 7 (IL-7) signaling is essential for DN2 and DN3a thymocyte survival and cell protection against apoptosis. The generation of αβ and γδ T-lymphocytes depends on the successful recombination of dispersed V (variable), D (diversity) and J (joining) gene segments at Tcra and Tcrb or Tcrg and Tcrd loci during thymocyte development, respectively, to express functional T-cell antigen receptors, TCRαβ or TCRγδ, at the cell membrane (Fig. 1).6-8 V(D)J recombination at Tcrg and Tcrd is completed in DN2 and DN3a thymocytes, at Tcrb in DN3a thymocytes, and at Tcra in DP thymocytes (Fig. 1).2,6,7 Successful Tcrg and Tcrd rearrangements permit the expression of TCRγδ, which drives cell differentiation to γδ T-lymphocytes in a process known as γδ-selection.9 Successful Tcrb rearrangements in DN3a thymocytes permit the expression of a pre-TCR formed by the assembly of a TCRβ chain with the invariant pre-Tα chain. Signaling transduced through the pre-TCR, together with those transduced by Notch receptor and IL-7, activates the differentiation of DN3a to DN3b, DN4, eDP, and lDP thymocytes in a process known as β-selection.10,11 Successful Tcra rearrangement in DP thymocytes permits the expression of a TCRα chain that associates with the previously expressed TCRβ to form TCRαβ. The expression of a TCRαβ in DP thymocytes allows for positive and negative selection events to permit the survival of a few DP thymocytes that will differentiate into CD4+ and CD8+ SP thymocytes and migrate to the periphery as mature αβ T-lymphocytes.
Transcriptional Regulation of The Tcra/Tcrd Locus During Thymocyte Development: Roles for Enhancers and Promoters
Tcra and Tcrd are organized in a single genetic locus, Tcra/Tcrd, which spans 1.7 Mb with a conserved genomic structure and location between the olfactory receptor genes and Dad1 in chromosome 14 in mice and humans (Fig. 2).6,8,12 The linkage of Tcra and Tcrd in vertebrates is present in teleost fish, indicating that these genes have been together for more than 400 million years.13 In mammals and birds, Tcra/Tcrd presents a nested organization that precludes the coexistence of Tcrd and Tcra rearrangements on the same chromosome because Tcra rearrangements result in the deletion of Tcrd in an extra-chromosomal circle.14,15 The 1.6-Mb 5′ locus region of Tcra/Tcrd includes 132 V gene segments, while the remaining approximately 100-kb 3′ locus region contains the Tcrd Dδ and Jδ gene segments, constant Tcrd region (Cδ), Jα gene segments and constant Tcra region (Cα).6,8 Among the V gene segments, some only rearrange with the Dδ gene segments, some only with the Jα gene segments, and some can rearrange with either Dδ or Jα, contributing to both the TCRδ and TCRα repertoires.16 Although the nested organization of Tcra/Tcrd prevents αβ T-lymphocyte expression of a TCRδ chain, the chromosome excision and gene expression of the productively rearranged Tcrd are dispensable for αβ T-cell lineage maturation.17
Figure 2.
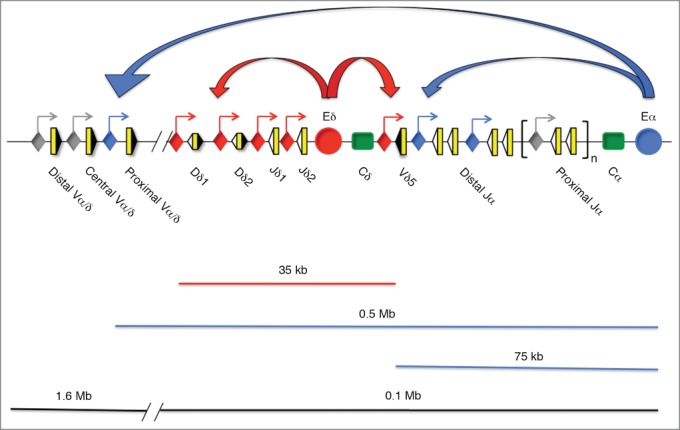
Representation of the genomic structure of the Tcra/Tcrd locus and active Eδ- and Eα-promoter interactions. The V, D, and J segments are represented by yellow rectangles, and the recombination signal sequences are shown as black or white triangles depending on the presence of 23-bp or 12-bp spacers, respectively. The Cδ and Cα regions are represented as green rectangles. The 1.6-Mb 5′-region includes the Vα and Vδ gene segments, and the 0.1-Mb 3′-region includes the Tcrd locus, the Jα gene segments, the Cα region and Eα. A red circle and a blue circle represent Eδ and Eα, respectively. The blue lines represent the chromatin area of Eα influence, whereas the red line represents that of Eδ. Promoters are represented as diamonds: red diamonds represent promoters that are responsive to Eδ, blue diamonds represent promoters that are responsive to Eα, and gray diamonds represent promoters that are not responsive to either Eδ or Eα. Straight arrows represent active sites of transcription associated with promoters. Curved arrows represent enhancer-promoter interactions: red curved arrows represent Eδ-promoter interactions, and blue curved arrows represent Eα-promoter interactions.
Each Tcra and Tcrd gene is equipped with one transcriptional enhancer, Eα and Eδ, located at the 3′ end of Cα and Cδ, respectively, and numerous promoters that are associated with the V, D, and J gene segments (Fig. 2).6,8 Eδ functions as a local enhancer with a range of influence that is limited to a 10 to 20-kb region that includes Dδ, Jδ, and Vδ5 in adult DN3a thymocytes and to a 55-kb region that also includes Vδ4 in fetal DN3a thymocytes, in comparison with the broad area influenced by Eα, which affects over 500 kb including the proximal Vα/δ gene segments that occupy 1/3 of all Vα/δ gene segments, Tcrd, and the whole Jα cluster (Fig. 2).18,19
Antigen receptor gene promoters function to drive sterile (or germline) transcription at unrearranged genes, and to activate the recombination of their associated gene segments during thymocyte development.8 Promoters associated with V gene segments also drive transcription of the rearranged genes. Pioneering experiments have demonstrated that the germline transcription of specific gene segments correlates with their suitability to undergo rearrangement,20-22 suggesting that transcription and V(D)J recombination are linked processes. This notion was supported by the observation that Eα and Eδ, as well as Tcra/Tcrd promoters, function by targeting V(D)J recombination to specific Tcra or Tcrd gene segments in a developmentally regulated fashion.23-28 Eα and Eδ are responsible for the developmental stage specificity of V(D)J recombination through the general regulation of chromatin structure at multiple gene segments that are separated by large distances, whereas promoters locally mediate the accessibility of the recombinase machinery to a gene or group of gene segments.8 In addition, transcriptional elongation is required to open the chromatin of gene segments that are positioned far away from a promoter, as occurs for several Jα gene segments.29,30
For VδDδJδ rearrangements and the subsequent expression of a mature TCRδ chain in DN2/3a thymocytes, Tcrd germline transcription initiates from Vδ-, Dδ-, and δ-associated promoters, and this process is, at least in part, Eδ-dependent.23 During these early thymocyte stages, Eα is not functionally active but is primed through the assembly of multiple constitutive TFs that bind cooperatively to enhancer DNA and the presence of the H3K4me1 histone mark.31-35 During β-selection, Eα is activated and Eδ is inactivated during the transition from DN3a to DP thymocytes.32 Pre-TCR signaling triggers the formation of a chromatin hub through physical interactions among TFs bound to the Eα enhanceosome and Cα-proximal Vα- and Cα-distal Jα promoters in a genomic region of approximately 500 kb.19,36 In addition to Eα-bound TFs, the CCCTC binding factor, CTCF, which binds closely to Eα and the most proximal Vα and distal Jα promoters, and interacting cohesin contribute to the stability of these long-range chromatin looping interactions.4,36 Eα-dependent activation of the proximal Vα and distal Jα promoters within the context of this chromatin hub triggers germline transcription to open the chromatin structure and provide accessibility to the recombination machinery in these regions in eDP thymocytes and promote proximal Vα to distal Jα rearrangements.19,28-30 These initial VαJα recombinations involving the proximal Vα and the distal Jα gene segments are known as primary rearrangements (Fig. 1).8,16,28 If the primary rearrangements are not productive, then secondary VαJα rearrangements involving central and distal Vα gene segments (more 5′ Vα gene segments) and Cα-proximal (more 3′) Jα gene segments occur in lDP thymocytes (Fig. 1).8,16,37 This strategy of successive Vα-to-Jα rearrangements using further 5′ Vα and 3′ Jα gene segments permits multiple rearrangements of the Tcra allele to assure the expression of a productive TCRα in all DP thymocytes providing greater likelihood that positive selection can occur.16
Hence, Eα and Eδ are responsible for the developmental stage specificity of V(D)J recombination at Tcra/Tcrd through the regulation of multiple promoters to drive germline transcription and gene segment accessibility to the recombinase machinery along the locus.8 Interestingly, these 2 enhancers are never simultaneously active during thymocyte development: in DN2/3a thymocytes, Eδ is on and Eα is off, whereas in DP thymocytes, Eδ is off and Eα is on.32 These data suggest that these 2 enhancers work as a developmental switch in the regulation of V(D)J recombination at Tcrd or Tcra through a common mechanism of Eδ inactivation and Eα activation during the transition of DN2/3a to DP thymocytes. The strict control of Eα activation up to the eDP thymocyte stage is critical to prevent premature Tcrd deletion in early thymocyte stages that could preclude the correct development of γδ T-cells. In addition, early TCRα expression in DN3a thymocytes generates TCRαγ complexes that signal the DN3a-to-DP transition independently of pre-TCR and impair further thymocyte development due to the silencing of Tcrg in DP thymocytes.38,39
Regulation of Eδ Function in Thymocytes and Mature T-Cells
Eδ activity depends critically on the binding of Runx and c-Myb TFs within a 30-bp protein-bound element known as δE3 (Fig. 3).40-42 The binding of these 2 TFs is absolutely required to activate both Tcrd germline transcription and VδDδJδ recombination.43,44 In addition to these 2 essential TFs for Eδ function, GATA-3 seems to contribute also to enhancer activity by binding to the immediately adjacent δE4 element (Fig. 3).42 Eδ is inactivated during the transition from DN3a to DP thymocytes through the dissociation of enhancer-bound TFs (Fig. 3).32
Figure 3.
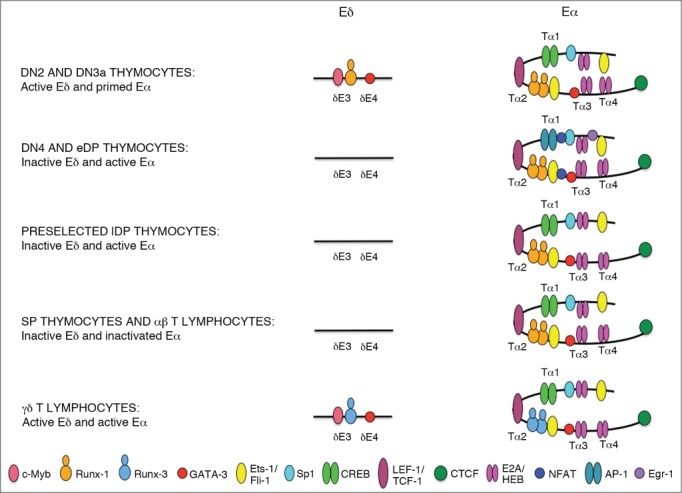
Model for the structure of Eδ and Eα enhanceosomes during T-cell development. The diagrams depict the TFs that are presumably bound to Eδ and Eα in thymocytes and T-lymphocytes. The locations of the defined δE3 and δE4 Eδ elements and of the Tα1, Tα2, Tα3, and Tα4 Eα elements are indicated. The identity of the different TFs is indicated by color-coded ovals. The strength of TF binding is represented by the positions of the colored ovals representing TFs on enhancer DNA: a more centered position represents stronger binding. Different TF-bound complexes are assembled on Eδ and Eα during T-cell development. In DN2/3a thymocytes, Eδ is active and occupied by c-Myb, Runx-1, and GATA-3, whereas Eα is primed and occupied by CREB, TCF-1/LEF-1, Runx-1, Ets-1, Sp1, GATA-3, and E2A/HEB; in DN4 and eDP thymocytes, Eδ becomes inactive and unoccupied by TFs, whereas Eα is activated through the recruitment of the pre-TCR-induced NFAT, Egr-1, and AP-1 TFs; in preselected lDP thymocytes, Eδ remains inactive and unoccupied, whereas Eα is activated through the strong binding of E2A and Ets-1; in SP thymocytes and αβ T-lymphocytes, Eδ remains inactive and presumably unoccupied, whereas Eα is inactive and demonstrates low binding of E2A and HEB TFs; and in γδ T-lymphocytes, Eδ is active and occupied by c-Myb, Runx-3, and GATA-3, whereas Eα is also active and presumably occupied by the same TFs assembled in the preselected lDP thymocyte enhanceosome, with the exception of Runx-3 that it might be exchanged for Runx-1.
Runx, c-Myb, and GATA-3 TFs bind in a highly hierarchical fashion to Eδ: Runx TF binds first to permit the access of c-Myb and GATA-3 to enhancer chromatin.45 Three Runx proteins are present in the thymus with presumably redundant activity but different expression levels in DN2/DN3a thymocytes: Runx-1 (also known as AML-1), Runx-2 (also known as AML-3), and Runx-3 (also known as AML-2).46 and www.immgen.org Three findings support Runx-1 as the relevant Runx TF for the function of Eδ during thymocyte development: (1) Runx-1 is able to open chromatin to form stable regulatory regions for important hematopoietic regulator genes,47 (2) Runx1 transcription exceeds that of Runx2 by 5–10 fold in DN2 thymocytes and by 8–20 fold in DN3a thymocytes and that of Runx3 by 2–5 fold in DN2 thymocytes and by 6–10 fold in DN3a thymocytes.3,46 www.immgen.org (3) Runx1 transcription is very strongly downregulated by pre-TCR signaling.3,46, www.immgen.org These data indicate that Runx-1 might act as a pioneer TF in the assembly of the active Eδ enhanceosome that is present in DN2/3a thymocytes and that its downregulation during β-selection might cause the TF dissociation and enhancer inactivation observed during the transition from DN3a to DP thymocytes.32 Interestingly, the downregulation of Runx TF expression during β-selection has been proposed to be the molecular mechanism underlying the inhibition of Eγs Tcrg enhancer activity in DP thymocytes.48 In fact, Eδ and Eγs are regulated very similarly; both are dependent on the essential recruitment of Runx and c-Myb TFs.40,43,44,48,49 Interestingly, Runx3 transcripts are approximately 4–15 fold more abundant than those of Runx1 and Runx2 in γδ T lymphocytes,3, www.immgen.org suggesting that Runx-1 and Runx-3 might have redundant roles for Eδ and Eγs function and that a potential exchange of Runx-3 for Runx-1 at the Eδ and Eγs enhanceosomes during DN2/DN3a thymocyte to γδ T cell development might occur (Fig. 3).
Conditional deletions of Runx1 and cMyb during early stages of thymocyte development provide results that are in agreement with their potential roles in the regulation of Eδ.50-52 Initial Lck-Cre-induced conditional deletion of Runx1 during thymocyte development revealed a developmental block in DN3a to DN4 maturation due to inefficient rearrangement of Tcrb, whereas deletion of Runx3 resulted in the defective generation of CD8+ SP thymocytes,50 with no obvious γδ T-cell-related phenotype in either case. However, conditional Runx1 deficiency in mouse bone marrow cells revealed defective T-lymphocyte development with a significant block in the maturation of DN2 to DN3a thymocytes.51 Because γδ T-lymphocytes can be derived directly from DN2 thymocytes, the failure of Runx1-deficient DN2 thymocytes to generate γδ T-cells could be the consequence of an essential role for Runx-1 in the regulation of Tcrd recombination. However, effective cMyb deletion in DN3a thymocytes resulted in a drastic decrease in the γδ T-cell population, supporting a role for this TF in the regulation of Tcrd.52 cMyb deletion also resulted in defective Tcra and Tcrb rearrangements.52 It is possible that the defective rearrangement of antigen receptor loci might be due, in part, to a role for c-Myb in regulating the transcription of the recombination-activating gene Rag2.53 In the case of Gata3 inhibition, its essential role in DN1 to DN3 thymocyte differentiation precludes a clear analysis of its potential role in regulating the transcription of T-cell antigen receptor genes.54
Although they are essential for the function of Eδ, δE3-bound Runx and c-Myb TFs alone or in combination with collaborating δE4-bound GATA-3 are not sufficient to activate Tcrd transcription and recombination in the context of a transgenic recombination reporter construct.44 δE3- and δE4-interacting TFs need to collaborate functionally with additional TFs bound to the 380-bp Eδ fragment, which contains 7 protein-binding elements, δE1 to δE7, to establish Jδ gene segment accessibility to the V(D)J recombinase machinery of a transgenic recombination reporter construct in mice.44,55 Furthermore, the 380-bp Eδ fragment itself is not able to activate either transcription or recombination in transgenic lines containing only one copy of the recombination reporter construct, requiring the functional collaboration with 2 flanking matrix attachment regions.56 At present, the identities of the TFs that collaborate with δE3-δE4-binding proteins for Eδ function are unknown.
Regulation of Eα Function in Thymocytes and Mature T-Cells
Eα is part of a locus control region that is located between Tcra and the ubiquitously expressed Dad1.57 It is formed by 4 protein-bound elements known as Tα1 to Tα4 (Fig. 3).58 The 116-bp Tα1-Tα2 enhancer fragment constitutes the core enhancer with essential binding sites for constitutive TFs such as CREB/ATF, TCF-1/LEF-1, Runx and Ets-1 that bind cooperatively in an all-or-none fashion.31, 59 Similarly to the situation in DN2a/DN3 thymocytes, Runx1 transcripts exceed that of Runx2 transcripts by 5 fold and that of Runx3 transcripts by 4 fold in DP thymocytes, www.immgen.org suggesting that Runx-1 is the Runx TF involved in the assembly of the Eα enhanceosome in these cells. All Eα-binding TF are all required for normal thymocyte development and the generation of αβ T-lymphocytes.60-62 These TFs are bound to a primed Eα prior to its activation in DN3a thymocytes and also when the enhancer is fully active in preselected lDP thymocytes.32-34,63 Although Tα1-Tα2 can activate transcription and V(D)J recombination at short distances in the context of a transgenic recombination reporter construct, it does not display proper Eα developmental regulation because it is activated prematurely in DN3a thymocytes.59 In addition, Tα1-Tα2 is not sufficient to activate Vα-to-Jα recombination at the endogenous Tcra locus.64 For proper Eα developmental regulation of V(D)J recombination, the 275-bp Tα1-Tα4 enhancer fragment is required.65 Additional binding sites in Tα3-Tα4 include those for the constitutive TFs Sp1, GATA-3, and the E proteins E2A and HEB.34 E2A and HEB together are essential for β-selection.66 To identify the molecular mechanism underlying Eα activation during β-selection, comparative analyses of enhancer occupancy during the transition from DN3a to DP thymocytes were performed by genomic footprinting and chromatin immunoprecipitation.32-34 Although the initial analyses did not reveal clear differences in enhancer occupancy by constitutive TFs between DN3a and preselected lDP thymocytes, more recent experiments demonstrated that Eα activation during β-selection is transitory and dependent on the binding of pre-TCR-inducible TFs, such as NFAT, AP-1 and Egr-1, in DN4/eDP thymocytes (Fig. 3).63 Furthermore, the binding of these signal-dependent TFs to Tα1-Tα4 in DN4/eDP thymocytes induces additional strong binding of constitutive E2A and Ets-1 in preselected lDP thymocytes.63 It was recently shown that after a functional TCRαβ is assembled in lDP thymocytes to positively select and induce further cell differentiation to SP thymocytes and αβ T-lymphocytes, Eα is inactivated.67 Inactivation of Eα was evidenced by the analysis of enhancer-dependent Vα and Jα germline transcripts, the disruption of long-range enhancer-promoter interactions analyzed by chromatin conformation capture assays, and the loss of activating histone marks such as H3K4me3 and H3K4me1, and it is associated with the loss of E2A and HEB binding to the enhancer.67 It is interesting to note that in contrast to Eα inactivation in mature αβ T-lymphocytes, Eα remains active and functionally relevant for transcription of the rearranged Tcrd in γδ T-cells.24
Hence, 5 Eα enhanceosomes with different activities are assembled during T-cell development: 2 inactive enhanceosomes that are assembled in DN2/3 thymocytes and αβ T-lymphocytes, and 3 active enhanceosomes that are assembled in DN4/eDP thymocytes, preselected lDP thymocytes, and γδ T-lymphocytes (Fig. 3).24,34,63,67 Both inactive Eα enhanceosomes display diminished E protein binding, suggesting that strong binding of E2A and HEB might be important for enhancer activation. Although the active Eα enhanceosome assembled in γδ T-lymphocytes has not yet been characterized, it is tempting to speculate that it might resemble the one that is present in preselected lDP thymocytes because it should be active in resting cells. In addition, because Runx3 transcription exceeds very dramatically that of Runx1 and Runx2 in γδ T lymphocytes,3, www.immgen.org it is predictable that Runx-3 will be exchanged for Runx-1 at the Eα enhanceosome during DN2/DN3a thymocyte to γδ T cell development (Fig. 3).
Considering all of the data reported for Eδ and Eα functions during T-cell development, 4 different scenarios for Tcrd and Tcra regulation by these enhancers can be distinguished: DN2/3a thymocytes, DN4/eDP/preselected lDP thymocytes, γδ T-lymphocytes, and SP thymocytes/αβ T-lymphocytes (Fig. 4). In DN2/3a thymocytes, Eδ is active to drive germline Tcrd transcription and VδDδJδ rearrangements, whereas Eα is primed but inactive.23,32,34,35 In DN4, eDP, and preselected lDP thymocytes, Eα is activated by pre-TCR signal-dependent TFs to drive germline Tcra transcription that is initiated at the promoters associated with the proximal Vα and distal Jα gene segments that trigger primary VαJα rearrangements, whereas Eδ becomes inactivated through the dissociation of its bound TFs.24,32,35,63 Hence, during the transition from DN2/3a to preselected lDP thymocytes, Eδ and Eα function as a developmental switch in the regulation of the Tcra/Tcrd locus, suggesting a common a signaling pathway for the activation of Eα and inactivation of Eδ during β-selection. Pre-TCR signaling is responsible for Runx1 transcriptional downregulation and the induction of NFAT, Egr-1, and AP-1 in the cell nucleus, and hence, it is the obvious signaling candidate to regulate the Eα/Eδ functional switch during β-selection. However, other signaling pathways associated with β-selection, such as those transduced by the Notch and IL-7 receptors, might also be involved.
Figure 4.
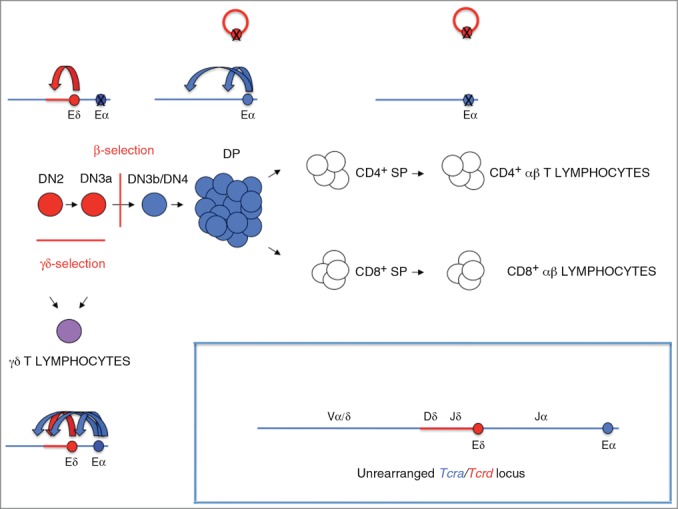
Regulation of Tcra/Tcrd transcription by Eδ and Eα during T-cell development. Schematic diagram of T-cell development depicting the various maturation stages. β- and γδ-selection are indicated. Eδ and Eα are represented a red and blue crossed ovals, respectively. The Tcrd locus and Eδ-activated cell stages are represented in red, the Tcra locus and Eα-activated cell stages are represented in blue, the Eδ- and Eα-activated stage corresponding to γδ T-lymphocytes is represented in purple, and the cell stages during which both Eα and Eδ are inactive is represented in white. Crossed circles represent inactive enhancers. A red empty circle represents deleted Tcrd. Curved arrows represent enhancer-promoter interactions: red curved arrows represent Eδ-promoter interactions, and blue curved arrows represent Eα-promoter interactions. In DN2/3a thymocytes, Eδ drives germline Tcrd transcription and VδDδJδ rearrangements, whereas Eα is primed but inactive. In DN4, eDP, and preselected lDP thymocytes, Eα drives germline Tcra transcription and primary VαJα rearrangements, whereas Eδ becomes inactive. In SP thymocytes and αβ T-lymphocytes, both Eα and Eδ are inactive. In γδ T-lymphocytes, Eα and Eδ are both active to drive the transcription of the rearranged Tcrd locus.
The patterns of Eα and Eδ activity are also drastically distinct in γδ and αβ T-cells.23,24,67 In γδ T-lymphocytes, both enhancers are active to drive the transcription of the rearranged Tcrd, whereas both are inactive in αβ T-lymphocytes. Transcription of the rearranged Tcrd in γδ T-cells depends on the strong activity of Eα whereas the contribution of Eδ to the generation of mature Tcrd transcripts is negligible.23 Hence, Eδ is essential for germline Tcrd transcription and recombination during thymocyte development but not for rearranged Tcrd transcription in γδ T-lymphocytes.23 During αβ T-lymphocyte differentiation, Eδ is deleted from the chromosome by Tcra rearrangement, and Eα is inactivated during the transition from preselected lDP to SP thymocytes.67 Hence, Eα is essential for germline Tcra transcription and recombination during the maturation of thymocytes but not for rearranged Tcra transcription in SP thymocytes and αβ T-lymphocytes.24,67 Thus, Eδ and Eα seem to constitute a type of enhancer that is specifically relevant to dictate the patterns of Tcrd and Tcra gene recombination and expression during T-cell development but not the expression of their respective rearranged genes in mature lymphocytes.
Insights Regarding the Molecular Mechanism Underlying the Transcription of the Rearranged Tcra
The inhibition of Eα in SP thymocytes and peripheral αβ T-lymphocytes does not preclude the strong transcription of the rearranged Tcra locus in these cells, implying that an Eα-independent molecular mechanism directs the transcription of the rearranged locus. Attempts to re-activate Eα in peripheral αβ T-lymphocytes through cell activation or T-helper differentiation have been unsuccessful, which suggests that enhancer inactivation is quite stable and permanent during T-cell homeostasis.67 The molecular basis for the transcription of the rearranged Tcra, which determines that the same Eα-dependent Vα promoters that are in an unrearranged configuration at the locus become Eα-independent following rearrangement, is currently unknown. It is possible that the different conformation of the rearranged locus compared with the unrearranged locus due to the deletion of intergenic sequences might facilitate the intrinsic activities of the rearranged Vα promoters or might unmask a novel enhancer in the locus that must be located upstream of Vα1 or downstream of Jα2 to ensure its retention following VαJα recombination. This Eα-independent mechanism for the transcriptional activation of the rearranged Tcra locus in αβ T-lymphocytes might be activated by tonic signals that are transduced by the TCRαβ-CD3 complex in resting cells because basal LAT-diacylglycerol-RasGRP1 signals are necessary to maintain the normal transcription of the rearranged locus.68 In support of this hypothesis, rearranged Tcra transcription responds positively to treatments with phorbol esters.69-73
It is interesting to note that in contrast to the positive effect of pre-TCR-transduced signaling in DN3a thymocytes, TCRαβ-transduced signaling in lDP thymocytes has a negative effect on the function of Eα. In support of the involvement of different signaling pathways in Tcra germline transcription by Eα activation during β-selection compared with the transcription of the rearranged Tcra during later stages of development, treatments with ionomycin synergistically activate the former in DN3a thymocytes but inhibit the latter in postselected lDP thymocytes.63,72,74,75
Future experiments will be directed toward deciphering the molecular pathways involved in regulating enhancer and promoter functions during thymocyte development, the detailed characterization of the enhanceosomes assembled during the different cell stages, and elucidation of the mechanism responsible for the transcription of the rearranged Tcra in αβ T-lymphocytes. These studies are of broad interest not only to understand the regulation of an essential locus for T-cell development and function but also because they might contribute to our understanding of the molecular mechanisms that control the dynamic regulation of distant enhancers. In addition, an understanding of the mechanism(s) by which Eα and Eδ control the process of recombination during normal development is crucial to our understanding of those cases in which their deregulation causes aberrant differentiation and leukemic transformation due to chromosomal translocations of the Tcra/Tcrd locus.76,77
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Funding
This work was supported by the Spanish government (Grant BFU2013-44660R) and the Andalusian government (Grant CTS-6587), which is financed in part by the European Regional Development Fund (ERDF/FEDER).
References
- 1.Heinz S, Romanoski CE, Benner C, Glass CK. The selection and functions of cell type-specific enhancers. Nat Rev 2015; 16:144-53; PMID:25650801; http://dx.doi.org/ 10.1038/nrm3949 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rothenberg EV, Tanghon T. Molecular genetics of T cell development. Annu Rev Immunol 2005; 23:601-49; PMID:15771582; http://dx.doi.org/ 10.1146/annurev.immunol.23.021704.115737 [DOI] [PubMed] [Google Scholar]
- 3.Taghon T, Yui MA, Pant R, Diamond RA, Rothenberg EV. Developmental and molecular characterization of emerging β- and γδ-selected pre-T cells in the adult mouse thymus. Immunity 2006; 24:53-64; PMID:16413923; http://dx.doi.org/ 10.1016/j.immuni.2005.11.012 [DOI] [PubMed] [Google Scholar]
- 4.Seitan VC, Hao B, Tachibana-Konwalski K, Lavagnolli T, Mira-Bontenbal H, Brown KE, Teng G, Carroll T, Terry A, Horan K, et al.. A role for cohesin in T-cell-receptor rearrangement and thymocyte differentiation. Nature 2011; 476:467-71; PMID:21832993; http://dx.doi.org/ 10.1038/nature10312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brekelmans P, van Soest P, Voerman J, Platenburg PP, Leenen PJ, van Ewijk W. Transferrin receptor expression as a marker of immature cycling thymocytes in the mouse. Cell Immunol 1994; 159:331-9; PMID:7994765; http://dx.doi.org/ 10.1006/cimm.1994.1319 [DOI] [PubMed] [Google Scholar]
- 6.Cobb RM, Oestreich KJ, Osipovich OA, Oltz EM. Accessibility control of V(D)J recombination. Adv Immunol 2006; 91:45-109; PMID:16938538; http://dx.doi.org/ 10.1016/S0065-2776(06)91002-5 [DOI] [PubMed] [Google Scholar]
- 7.Osipovich O, Oltz EM. Regulation of antigen receptor gene assembly by genetic-epigenetic crosstalk. Semin Immunol 2010; 22:313-22; PMID:20829065; http://dx.doi.org/ 10.1016/j.smim.2010.07.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.del Blanco B, Angulo Ú, Hernández-Munain C. Epigenetic control of T cell receptor locus rearrangements in normal and aberrant conditions. Epigenetic and Human Health 2014; Transcriptional and Epigenetic Mechanisms Regulating Normal and Aberrant Blood Cell Development 12:295-329; http://dx.doi.org/ 10.1007/978-3-642-45198-0_12 [DOI] [Google Scholar]
- 9.Prinz I, Sansoni A, Kissenpfennig A, Ardouin L, Malissen M, Malissen B. Visualization of the earliest steps of γδ T cell development in the adult thymus. Nat Immunol 2006; 7:995-1003; PMID:16878135; http://dx.doi.org/ 10.1038/ni1371 [DOI] [PubMed] [Google Scholar]
- 10.Ciofani M, Schmitt TM, Ciofani A, Michie AM, Cuburu N, Aublin A, Maryanski JL, Zúñiga-Pflucker JC. Obligatory role for cooperative signaling by pre-TCR and Notch during thymocyte differentiation. J Immunol 2004; 172:5230-9; PMID:15100261; http://dx.doi.org/ 10.4049/jimmunol.172.9.5230 [DOI] [PubMed] [Google Scholar]
- 11.Boudil A, Matei IR, Shih HY, Bogdanoski G, Yuan JS, Chang SG, Montpellier B, Kowalski PE, Voisin V, Bashir S, et al.. IL-7 coordinates proliferation, differentiation and Tcra recombination during thymocyte β-selection. Nat Immunol 2015; 16:397-405; PMID:25729925; http://dx.doi.org/ 10.1038/ni.3122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Glusman G, Rowen L, Lee I, Boysen C, Roach JC, Smit AFA, Wang K, Koop BF, Hood L. Comparative genomics of the human and mouse T cell receptor loci. Immunity 2001; 15:337-49; PMID:11567625; http://dx.doi.org/ 10.1016/S1074-7613(01)00200-X [DOI] [PubMed] [Google Scholar]
- 13.Fischer C, Bouneau L, Ozouf-Costaz C, Crnogorac-Jurcevic T, Weissenbach J, Bernot A. Conservation of the T-cell receptor α/δ linkage in the teleost fish Tetraodon nigroviridis. Genomics 2002; 79:241-8; PMID:11829494; http://dx.doi.org/ 10.1006/geno.2002.6688 [DOI] [PubMed] [Google Scholar]
- 14.Koop BF, Rowen L, Wang K, Kuo CL, Seto D, Lenstra JA, Howard S, Shan W, Deshpande P, Hood L. The human T-cell receptor TCRAC/TCRDC (Cα/Cδ) region: organization, sequence, and evolution of 97.6 kb of DNA. Genomics 1994; 19:478-93; PMID:8188290; http://dx.doi.org/ 10.1006/geno.1994.1097 [DOI] [PubMed] [Google Scholar]
- 15.Kubota T, Wang J, Gobel TW, Hockett RD, Cooper MD, Chen CH. Characterization of an avian (Gallus gallus domesticus) TCR α/δ gene locus. J Immunol 1999; 163:3858-66; PMID:10490985 [PubMed] [Google Scholar]
- 16.Krangel MS, Carabana J, Abarrategui I, Schlimgen R, Hawwari A. Enforcing order within a complex locus: current perspectives on the control of V(D)J recombination at the murine T-cell receptor α/δ locus. Immun Rev 2004; 200:224-32; PMID:15242408; http://dx.doi.org/ 10.1111/j.0105-2896.2004.00155.x [DOI] [PubMed] [Google Scholar]
- 17.Khor B, Wehrly TD, Sleckman BP. Chromosomal excition of TCRδ chain genes is dispensable for αβ T cell lineage commitment. Int Immunol 2005; 17:225-32; PMID:15642954; http://dx.doi.org/ 10.1093/intimm/dxh202 [DOI] [PubMed] [Google Scholar]
- 18.Hao B, Krangel MS. Long-distance regulation of fetal Vδ gene segment TRDV4 by the Tcrd enhancer. J Immunol 2011; 187:2484-91; PMID:21784972; http://dx.doi.org/ 10.4049/jimmunol.1100468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hawwari A, Krangel MS. Regulation of TCR δ and α repertoires by local and long-distance control of variable gene segment chromatin structure. J Exp Med 2005; 202:467-72; PMID:16087716; http://dx.doi.org/ 10.1084/jem.20050680 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Capone M, Hockett RD Jr, Zlotnik A. Kinetics of T cell receptor β, γ, and δ rearrangements during adult thymic development: T cell receptor rearrangements are present in CD44+CD25+ Pro-T thymocytes. Proc Natl Acad Sci U S A 1998; 95:12522-7; PMID:9770518; http://dx.doi.org/ 10.1073/pnas.95.21.12522 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Livak F, Tourigny M, Schatz DG, Petrie HT. Characterization of TCR gene rearrangements during adult murine T cell development. J Immunol 1999; 162:2575-80; PMID:10072498 [PubMed] [Google Scholar]
- 22.Dik WA, Pike-Overzet K, Weerkamp F, de Ridder D, de Hass EFE, Baert MRM, van der Spek P, Koster EEL, Reinders MJT, van Dongen JJM, et al.. New insights on human T cell development by quantitative T cell receptor gene rearrangement studing and gene expression profiling. J Exp Med 2005; 201:1715-23; PMID:15928199; http://dx.doi.org/ 10.1084/jem.20042524 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Monroe RJ, Sleckman BP, Monroe BC, Khor B, Claypool S, Ferrini R, Davidson L, Alt FW. Developmental regulation of TCR δ locus accessibility and expression by the TCR δ enhancer. Immunity 1999; 10:503-13; PMID:10216098; http://dx.doi.org/ 10.1016/S1074-7613(00)80050-3 [DOI] [PubMed] [Google Scholar]
- 24.Sleckman BP, Bardon CG, Ferrini R, Davidson L, Alt FW. Function of the TCR α enhancer in αβ and γδ T cells. Immunity 1997; 7:505-15; http://dx.doi.org/ 10.1016/S1074-7613(00)80372-6 [DOI] [PubMed] [Google Scholar]
- 25.Lauzurica P, Krangel MS. Temporal and lineage-specific control of T cell receptor α/δ gene rearrangement by T cell receptor α and δ enhancers. J Exp Med 1994; 179:1913-21; PMID:8195717; http://dx.doi.org/ 10.1084/jem.179.6.1913 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Villey I, Caillol D, Selz F, Ferrier P, de Villartay JP. Defect in rearrangement of the most 5′ TCR-Jα following targeted deletion of T early α (TEA): implications for TCRα locus accessibility. Immunity 1996; 5:331-42; PMID:8885866; http://dx.doi.org/ 10.1016/S1074-7613(00)80259-9 [DOI] [PubMed] [Google Scholar]
- 27.Naik AK, Hawwari A, Krangel MS. Specification of Vδ and Vα usage by Tcra/Tcrd locus V gene segment promoters. J Immunol 2015; 194:790-4; PMID:25472997; http://dx.doi.org/ 10.4049/jimmunol.1402423 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hawwari A, Bock C, Krangel MS. Regulation of T cell receptor α gene assembly by a complex hierarchy of germline Jα promoters. Nat Immunol 2005; 6:481-9; PMID:15806105; http://dx.doi.org/ 10.1038/ni1189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Abarrategui I, Krangel MS. Regulation of T cell receptor-α gene recombination by transcription. Nat Immunol 2006; 7:1109-15; PMID:16936730; http://dx.doi.org/ 10.1038/ni1379 [DOI] [PubMed] [Google Scholar]
- 30.Abarrategui I, Krangel MS. Noncoding transcription controls downstream promoters to regulate T-cell receptor α recombination. EMBO J 2007; 26:4380-90; PMID:17882258; http://dx.doi.org/ 10.1038/sj.emboj.7601866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hernández-Munain C, Roberts JL, Krangel MS. Cooperation among multiple transcription factors is required for access to minimal T-cell receptor α-enhancer chromatin in vivo. Mol Cell Biol 1998; 18:3223-33; PMID:9584163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hernández-Munain C, Sleckman BP, Krangel MS. A developmental switch from TCRδ enhancer to TCRα enhancer function during thymocyte maturation. Immunity 1999; 10:723-33; PMID:10403647; http://dx.doi.org/ 10.1016/S1074-7613(00)80071-0 [DOI] [PubMed] [Google Scholar]
- 33.Spicuglia S, Payet D, Tripathi RK, Rameil P, Verthuy C, Imbert J, Ferrier P, Hempel WM. TCRα enhancer activation occurs via a conformational change of a pre-assembled nucleo-protein complex. EMBO J 2000; 19:2034-45; PMID:10790370; http://dx.doi.org/ 10.1093/emboj/19.9.2034 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.del Blanco B, Roberts JL, Zamarreño N, Balmelle-Devaux N, Hernández-Munain C. Flexible stereospecific interactions and composition within nucleoprotein complexes assembled on the TCRα gene enhancer. J Immunol 2009; 183:1871-83; PMID:19596981; http://dx.doi.org/ 10.4049/jimmunol.0803351 [DOI] [PubMed] [Google Scholar]
- 35.Pekowska A, Benoukraf T, Zacarias-Cabeza J, Belhocine M, Koch F, Holota H, Imbert J, Andrau JC, Ferrier P, Spicuglia S. H3K4 tri-methylation provides an epigenetic signature of active enhancers. EMBO J 2011; 30:4198-210; PMID:21847099; http://dx.doi.org/ 10.1038/emboj.2011.295 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shih HY, Verma-Gaur J, Torkamani A, Feeney AJ, Galjart N, Krangel MS. Tcra gene recombination is supported by a Tcra enhancer- and CTCF-dependent chromatin hub. Proc Natl Acad Sci USA 2012; 109:3493-502; PMID:23169622; http://dx.doi.org/ 10.1073/pnas.1214131109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Hawwari A, Krangel MS. Role for rearranged variable gene segments in directing secondary T cell receptor α recombination. Proc Natl Acad Sci USA 2007; 104:903-7; PMID:17210914; http://dx.doi.org/ 10.1073/pnas.0608248104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Erman B, Feigenbaum L, Coligan JE, Singer A. Early TCRα expression generated TCRαγ complexes that signal the DN-to-DP transition and impairs development. Nat Immunol 2002; 3:564-9; PMID:12021779; http://dx.doi.org/ 10.1038/ni800 [DOI] [PubMed] [Google Scholar]
- 39.Ferrero I, Mancini SJ, Grosjean F, Wilson A, Otten L, MacDonald HR. TCRγ silencing during αβ T cell development depends upon pre-TCR-induced proliferation. J Immunol 2006; 177:6038-43; PMID:17056529; http://dx.doi.org/ 10.4049/jimmunol.177.9.6038 [DOI] [PubMed] [Google Scholar]
- 40.Hernández-Munain C, Krangel MS. Regulation of the T-cell receptor δ enhancer by functional cooperation between c-Myb and core-binding factors. Mol Cell Biol 1994; 14:473-83; PMID:8264615 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Hernández-Munain C, Krangel MS. c-Myb and core-binding factor/PEBP2 display functional synergy but bind independently to adjacent sites in the T-cell receptor δ enhancer. Mol Cell Biol 1995; 15:3090-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Redondo JM, Pfohl JL, Krangel MS. Identification of an essential site for transcriptional activation within the human T-cell receptor δ enhancer. Mol Cell Biol 1991; 11:5671-80; PMID:1833636 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hernández-Munain C, Lauzurica P, Krangel MS. Regulation of T cell receptor δ gene rearrangement by c-Myb. J Exp Med 1996; 183:289-93; http://dx.doi.org/ 10.1084/jem.183.1.289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lauzurica P, Zhong XP, Krangel MS, Roberts JL. Regulation of T cell receptor δ gene rearrangement by CBF/PEBP2. J Exp Med 1997; 185:1193-201; http://dx.doi.org/ 10.1084/jem.185.7.1193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hernández-Munain C, Krangel MS. Distinct roles for c-Myb and core binding factor/polyoma enhancer-binding protein 2 in the assembly and function of a multiprotein complex on the TCR δ enhancer in vivo. J Immunol 2002; 169:4362-9; PMID:12370369; http://dx.doi.org/ 10.4049/jimmunol.169.8.4362 [DOI] [PubMed] [Google Scholar]
- 46.David-Fung ES, Butler R, Buzi G, Yui MA, Diamond RA, Anderson MK, Rowen L, Rothenberg EV. Transcription factor expression dynamics of early-T lymphocyte specification and commitment. Dev Biol 2009; 325:444-67; PMID:19013443; http://dx.doi.org/ 10.1016/j.ydbio.2008.10.021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lichtinger M, Hoogenkamp M, Krysinka H, Ingram R, Bonifer C. Chromatin regulation by RUNX1. Blood Cell Mol Dis 2010; 44:287-90; PMID:20194037; http://dx.doi.org/ 10.1016/j.bcmd.2010.02.009 [DOI] [PubMed] [Google Scholar]
- 48.Tani-ichi S, Satake M, Ikuta K. The pre-TCR signal induces transcriptional silencing of the TCRγ locus by reducing the recruitment of STAT5 and Runx to transcriptional enhancers. Int Immunol 2011; 25:553-63; PMID:21750145; http://dx.doi.org/ 10.1093/intimm/dxr055 [DOI] [PubMed] [Google Scholar]
- 49.Hsiang YH, Goldman JP, Raulet DH. The role of c-Myb or a related factor in regulating the T cell receptor γ gene enhancer. J Immunol 1995; 154:5195-204; PMID:7730626 [PubMed] [Google Scholar]
- 50.Taniuchi I, Osato M, Egawa T, Sunshine MJ, Bae S-C, Komori T, Ito Y, Littman DR. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 2002; 111:621-33; PMID:12464175; http://dx.doi.org/ 10.1016/S0092-8674(02)01111-X [DOI] [PubMed] [Google Scholar]
- 51.Ichikawa M, Asai T, Saito T, Seo S, Yamazaki I, Yamagata T, Mitani K, Chiba S, Ogawa S, Kurokawa M, Hirai H, AML-1 is required for megakaryocytic maturation and lymphocytic differentiation, but not for maintenance of hematopoietic stem cells in adult hematopoiesis. Nat Med 2014; 10:299-304; PMID:14966519; http://dx.doi.org/ 10.1038/nm997 [DOI] [PubMed] [Google Scholar]
- 52.Bender TP, Kremer CS, Kraus M, Buch T, Rajewsky K. Critical functions for c-Myb at three checkpoints during thymocyte development. Nat Immunol 2004; 5:721-9; PMID:15195090; http://dx.doi.org/ 10.1038/ni1085 [DOI] [PubMed] [Google Scholar]
- 53.Wang QF, Lauring J, Schlissel MS. c-Myb binds to a sequence in the proximal region of RAG-2 promoter and is essential for promoter activity in T-lineage cells. Mol Cell Biol 2000; 20:9203-11; PMID:11094072; http://dx.doi.org/ 10.1128/MCB.20.24.9203-9211.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Scripture-Adams DD, Danmle SS, Li L, Elihu KJ, Qin S, Arias AM, Butler RR 3rd, Champhekar A, Zhang JA, Rothenberg EV. GATA-3 dose-dependent checkpoints in early cell commitment. J Immunol 2014; 193:3470-91; PMID:25172496; http://dx.doi.org/ 10.4049/jimmunol.1301663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Redondo JM, Hata S, Brocklehurst C, Krangel MS. A T cell-specific transcriptional enhancer within the human T cell receptor δ locus. Science 1990; 247:1225-9; PMID:2156339; http://dx.doi.org/ 10.1126/science.2156339 [DOI] [PubMed] [Google Scholar]
- 56.Zhong XP, Carabana J, Krangel MS. Flanking nuclear matrix attachment regions synergize with the T cell receptor δ enhancer to promote V(D)J recombination. Proc Natl Acad Sci U S A 1999; 96:11970-5; PMID:10518560; http://dx.doi.org/ 10.1073/pnas.96.21.11970 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Diaz P, Cado D, Winoto A. A locus control region in the T cell receptor α/δ locus. Immunity 1994; 1:207-17; PMID:7889409; http://dx.doi.org/ 10.1016/1074-7613(94)90099-X [DOI] [PubMed] [Google Scholar]
- 58.Ho IC, Leiden JM. Regulation of the human T-cell receptor α gene enhancer: multiple ubiquitous and T-cell-specific nuclear proteins interact with four hypomethylated enhancer elements. Mol Cell Biol 1990; 10:4720-7; PMID:2388624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Roberts JL, Lauzurica P, Krangel MS. Developmental regulation of VDJ recombination by the core fragment of the T cell receptor α enhancer. J Exp Med 1997; 185:131-40; PMID:8996249; http://dx.doi.org/ 10.1084/jem.185.1.131 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Eyquem S, Chemin K, Fasseu M, Bories J-C. The Ets-1 transcription factor is required for complete pre-T cell receptor function and allelic exclusion at the T cell receptor β locus. Proc Natl Acad Sci U S A 2004; 101:15712-7; PMID:15496469; http://dx.doi.org/ 10.1073/pnas.0405546101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yu S, Zhou X, Steinke FC, Liu C, Chen S-C, Zagorodna O, Jing X, Yokota Y, Meyerholz DK, Mullighan CG, et al.. The TCF-1 and LEF-1 transcription factors have cooperative and opposing roles in T cell development and malignancy. Immunity 2012; 37:813-26; PMID:23103132; http://dx.doi.org/ 10.1016/j.immuni.2012.08.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Rudolph D, Tafuri A, Gass P, Hammerling GJ, Arnold B, Schutz G. Impaired fetal T cell development and perinatal lethality in mice lacking the cAMP response element binding protein. Proc Natl Acad Sci USA 1998; 95:4481-6; PMID:9539763; http://dx.doi.org/ 10.1073/pnas.95.8.4481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.del Blanco B, García-Mariscal A, Wiest DL, Hernández-Munain C. Tcra enhancer activation by inducible transcription factors downstream of pre-TCR signaling. J Immunol 2012; 188:3278-93; PMID:22357628; http://dx.doi.org/ 10.4049/jimmunol.1100271 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Bassing CH, Tillman RE, Woodman BB, Canty D, Monroe BC, Sleckman BP, Alt FW. T cell receptor (TCR) α/δ locus enhancer identity and position are critical for the assembly of TCR δ and α variable region genes. Proc Natl Acad Sci U S A 2003; 100:2598-603; PMID:12604775; http://dx.doi.org/ 10.1073/pnas.0437943100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Balmelle N, Zamarreño N, Krangel MS, Hernández-Munain C. Developmental activation of the TCR α enhancer requires functional collaboration among proteins bound inside and outside the core enhancer. J Immunol 2004; 173:5054-63; PMID:15470049; http://dx.doi.org/ 10.4049/jimmunol.173.8.5054 [DOI] [PubMed] [Google Scholar]
- 66.Jones ME, Zhuang Y. Stage-specific functions of E-proteins at the β-selection and T-cell receptor checkpoints during thymocyte development. Immunol Res 2011; 49:202-15; PMID:21128008; http://dx.doi.org/ 10.1007/s12026-010-8182-x [DOI] [PubMed] [Google Scholar]
- 67.del Blanco B, Angulo Ú, Krangel MS, Hernández-Munain C. The Tcra enhancer is inactivated in αβ T lymphocytes. Proc Natl Acad Sci USA 2015; 112:1744-53; PMID:25831496; http://dx.doi.org/ 10.1073/pnas.1406551112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Markegard E, Trager E, Ou Yang C-W, Zhang W, Weiss A, Roose JP. Basal LAT-diacylglycerol-RasGRP1 signals in T cells maintain TCRα gene expression. Plos One 2011; 6:1-9; PMID:21966541; http://dx.doi.org/ 10.1371/journal.pone.0025540 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Mallory MJ, Jackson J, Weber B, Chi A, Heyd F, Lynch KW. Signal- and development-dependent alternative splicing of LEF1 in T cells is controlled by CELF2. Mol Cell Biol 2011; 31:2184-95; PMID:21444716; http://dx.doi.org/ 10.1128/MCB.05170-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Roose JP, Diehn M, Tomlinson MG, Lin J, Alizadeh AA, Botstein D, Brown PO, Weiss A. T cell receptor-independent basal signaling via Erk and Abl kinases supresses RAG gene expression. Plos Biol 2003; 1:271-87; PMID:14624253; http://dx.doi.org/ 10.1371/journal.pbio.0000053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Lindsten T, June CH, Thompson CB. Transcription of T cell antigen receptor genes is induced by protein kinase C activation. J Immunol 1988; 141:1769-74; PMID:3261760 [PubMed] [Google Scholar]
- 72.Martinez-Valdez H, Doherty PJ, Thompson E, Benedict SH, Gelfand EW, Cohen A. Antagonistic effects of calcium ionophores and phorbol esters on T cell receptor mRNA levels in human thymocytes. J Immunol 1988; 140:361-6; PMID:3257235 [PubMed] [Google Scholar]
- 73.Martinez-Valdez H, Thompson E, Cohen A. Coordinate transcriptional regulation of α and δ chains of T-cell antigen receptors by phorbol esters and cyclic adenosine 5′-monophosphate in human thymocytes. J Biol Chem 1988; 263:9561-4 [PubMed] [Google Scholar]
- 74.Wilkinson MF, MacLeod CL. Induction of T-cell receptor-α and -β mRNA in SL12 cells can occur by transcriptional and post-transcriptional mechanisms. EMBO J 1988; 7:101-9; PMID:3129288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Herrick T, Qian L, Wilkinson MF. TcR-α mRNA accumulation does not dictate cell surface TcR/CD3 expression. Mol Immunol 1992; 29:531-6; PMID:1533011; http://dx.doi.org/ 10.1016/0161-5890(92)90011-L [DOI] [PubMed] [Google Scholar]
- 76.Aifantis I, Raetz E, Buonamici S. Molecular pathogenesis of T-cell leukemia and lymphoma. Nat Immunol 2008; 8:380-90; PMID:18421304; http://dx.doi.org/ 10.1038/nri2304 [DOI] [PubMed] [Google Scholar]
- 77.Dadi S, Le Noir S, Payet-Bornet D, Lhermitte L, Zacarias-Cabeza J, Bergeron J, Villarese P, Vachez E, Dik WA, Millien C, et al.. TLX homoedomain oncogenes mediate T cell maturation arrest in T-ALL via interaction with ETS1 and suppresion of TCRα gene expression. Cancer Cell 2012; 21:563-76; PMID:22516263; http://dx.doi.org/ 10.1016/j.ccr.2012.02.013 [DOI] [PubMed] [Google Scholar]


