Abstract
We previously reported that actin-depolymerizing agents promote the alkalization of the Golgi stack and the trans-Golgi network. The main determinant of acidic pH at the Golgi is the vacuolar-type H+-translocating ATPase (V-ATPase), whose V1 domain subunits B and C bind actin. We have generated a GFP-tagged subunit B2 construct (GFP-B2) that is incorporated into the V1 domain, which in turn is coupled to the V0 sector. GFP-B2 subunit is enriched at distal Golgi compartments in HeLa cells. Subcellular fractionation, immunoprecipitation, and inversal FRAP experiments show that the actin depolymerization promotes the dissociation of V1-V0 domains, which entails subunit B2 translocation from Golgi membranes to the cytosol. Moreover, molecular interaction between subunits B2 and C1 and actin were detected. In addition, Golgi membrane lipid order disruption by d-ceramide-C6 causes Golgi pH alkalization. We conclude that actin regulates the Golgi pH homeostasis maintaining the coupling of V1-V0 domains of V-ATPase through the binding of microfilaments to subunits B and C and preserving the integrity of detergent-resistant membrane organization. These results establish the Golgi-associated V-ATPase activity as the molecular link between actin and the Golgi pH.
Keywords: actin, Golgi, intracellular trafficking, lipid raft, pH regulation, vacuolar ATPase
Introduction
The secretory pathway is characterized by progressive lumen acidification of its organelles, from almost neutral in the endoplasmic reticulum (ER)4 (pH ≈7.1–7.2), along the cis-to-trans Golgi stack (pH ≈6.7–6.0), to more acidic in the trans-Golgi network (TGN) and secretory vesicles/granules (pH ≈5.0) (1–3). This pH gradient is crucial for post-translational modifications and membrane trafficking events (4, 5). The main molecular determinant of the progressive fall in pH along the secretory pathway is the vacuolar [H+]ATPase (V-ATPase) (6–8). V-ATPase is a multisubunit complex composed of two large domains, V0 and V1. The V0 domain is a 260-kDa integral membrane complex made up of five different subunits (a, b, c, c′, c″, d, and e), which mediates proton translocation; the V1 domain is a 600–650-kDa peripheral complex composed of eight different subunits (A, B, C, D, E, F, G, and H), which is responsible for the ATP hydrolysis that provides the mechanical force necessary for proton (H+) translocation (7, 9–11). Whereas they are the primary source of proton delivery to endomembranes consuming ATP, the final steady-state pH in the secretory pathway is the result of the balance between active H+ pumping by the V-ATPase, passive H+ efflux through organelle endogenous H+ permeability, and differences in counter-ion conductance (3, 12).
How differences in the pH of individual secretory compartments are generated is not well understood. Differential V-ATPase density and/or local regulatory mechanisms in secretory organelles and subcompartments are possible (13). In this respect, V-ATPase-dependent proton translocation could be regulated by several mechanisms, which include the following: (a) differential V-ATPase subunit expression; (b) intracellular targeting and recycling of V-ATPase-containing transport carriers to and from the plasma membrane and endomembranes (mainly late endosomes and lysosomes); (c) modulation of the coupling ratio between ATP hydrolysis and proton pumping; and (d) the reversible association of the V0 and V1 domains (14). This latter phenomenon is characterized by the release of the V1 domain into the cytosol, leading to the inhibition of ATPase and proton transport activities (15). This mechanism was first identified in tobacco hornworm and yeast after glucose deprivation (16), which was subsequently reported in mammalian cells (17, 18). When cells are cultured in the absence of glucose, a rapid dissociation of V1 and V0 domains occurs, which is completely reassembled after glucose re-addition, which obviates the need for de novo protein synthesis (19–21). Importantly, the cytoskeleton also seems to play a role in V-ATPase assembly and activity. In particular, microtubule integrity is necessary for the reversible dissociation of the two domains, because their disruption with nocodazole blocked the V0-V1 dissociation in response to glucose depletion in yeast (22). In contrast, actin interacts with the B and C subunits of the V1 domain, both of which contain at least one actin-binding domain (23–25). Filamentous actin (F-actin) binds to the amino terminus of the kidney and brain B1 and B2 subunit isoforms, respectively (26). This actin interaction is necessary for the transport of the V-ATPase to the plasma membrane during osteoclast inactivation (27). Recombinant subunit C binds both filamentous (F) actin and globular (G) actin (25, 28). Notice that these studies were focused on the V-ATPase located in the plasma membrane or assayed in vitro utilizing purified proteins. Little is known about the functional relevance of such interaction and even less about whether it also takes place in endomembrane systems.
The presence of an H+-translocating ATPase in Golgi membranes of animal and plant cells has long been known (29–34), and it is particularly enriched in lipid rafts (35). Besides V-ATPase, other ion channels have been reported in the Golgi, such as the Golgi chloride channel pH regulator (36), Golgi anion channels GOLAC-1 and GOLAC-2 (37), and Na+/H+ exchanger isoforms NHE7 and NHE8 (38). The presence of all these ion channels, including V-ATPase, has contributed to our understanding of the importance of Golgi pH homeostasis in health and disease. In particular, both glycosylation and protein-sorting events are known to be highly sensitive to changes in intra-Golgi pH (39–46). In contrast, much less is known about the subcellular localization and regulatory mechanisms of V-ATPase in the Golgi.
Our group previously reported significant similarities between events occurring after microfilament disruption (with actin-depolymerizing agents) (47–49) and those seen after the pharmacological inhibition of V-ATPase (with bafilomycin A1 and concanamycin A) (43, 44, 49). These similarities include the following: (a) membrane trafficking alterations in the Golgi-to-ER and post-Golgi protein transports; (b) alkalization of the Golgi complex, and (c) strong dilatation of cisternae, observed under the electron microscope. Knowing that B and C subunits of the V1 domain bind to F- and/or G-actin (25, 26, 28, 50), we hypothesized that actin could participate in Golgi pH homeostasis through the regulation of V-ATPase activity. In particular, we hypothesized that microfilaments are crucial to the maintenance of V0 and V1 domain association (49, 51). In this study, we provide experimental evidence that microfilaments do indeed maintain V0 and V1 domain association via two mechanisms as follows: one through the interaction between actin and subunits B2 and C1, and other based on the actin-dependent integrity of detergent-resistant membranes (DRMs) where V-ATPase localizes.
Experimental Procedures
Antibodies and Reagents
Mouse monoclonal antibodies to GM130 and Golgin97 were from Transduction Laboratories (Lexington, KY) and Molecular Probes, Life Technologies, Inc. (Paisley, UK), respectively. Sheep polyclonal anti-TGN46 was from Abcam (Cambridge, UK). Mouse monoclonal anti-transferrin receptor (TfR), rabbit polyclonal anti-Rho-GDI, rabbit polyclonal anti-actin, and mouse monoclonal anti-caveolin1 were from Sigma, Santa Cruz Biotechnology (Santa Cruz, CA), Sigma, and BD Transduction Laboratories, respectively. Mouse monoclonal anti-GFP was from Molecular Probes. Mouse monoclonal and polyclonal antibodies against subunits A or B2 from V-ATPase V1 domain were from Abnova (Taipei, Taiwan) and Abcam, respectively. Rabbit polyclonal against subunit a1 from the V0 domain was from Santa Cruz Biotechnology. Rabbit polyclonal against subunit C1 was from Abcam. Mouse monoclonal antibody against subunit B1 was from OriGene (Rockville, MD). Mouse monoclonal antibody against actin used for immunoprecipitation experiments was agarose-conjugated (Santa Cruz Biotechnology). Cy3-conjugated rabbit secondary antibodies were from Jackson ImmunoResearch (West Grove, PA), and Alexa Fluor 488-conjugated or Alexa Fluor 546-conjugated anti-mouse, anti-rabbit, and anti-sheep antibodies were from Molecular Probes, Life Technologies, Inc. Peroxidase-conjugated secondary antibodies were from Promega (Eugene, OR). Cholera toxin B subunit, from Vibrio cholerae-HRP, was from Sigma. Protein A-agarose beads were from Santa Cruz Biotechnology. Cycloheximide, cytochalasin D, nigericin, monensin, and FITC-phalloidin were from Sigma. DAPI Fluoromount G was from SouthernBiotech. Latrunculin B and Mowiol were from Calbiochem (Darmstadt, Germany). Jasplakinolide was from Invitrogen, Life Technologies, Inc., and dithiobis[succinimidylpropionate] was from Thermo Scientific, Pierce. 35S-Labeled protein labeling mix (35S-EXPRESS) was from PerkinElmer Life Sciences. N-Hexanoyl-d-erythrosphingosine (d-ceramide-C6) and N-hexanoyl-l-erythrosphingosine (l-ceramide-C6) were from Matreya, and dissolved in ethanol as a stock solution.
Transfection of Plasmids and siRNA
Plasmids were transfected with FuGENE® HD transfection reagent (Promega, Eugene, OR) following the manufacturer's recommendations. siRNA pool for subunit B2 was purchased from Dharmacon (ON-TARGETplus SMARTpool, reference number L-011589-01-0005). siRNAs (20 nm) were transfected in cells in suspension using Lipofectamine® 2000 reagent (Invitrogen) following the manufacturer's instructions. Experiments were performed 72 h after treatment. All siRNA-mediated knockdown experiments were validated with a pool of four non-targeting siRNAs (Dharmacon, reference number D-001810-10−05).
Plasmids
Plasmids encoding pHluorin and pHluorin-TGN constructs (52) were kindly provided by Juan L. Llopis (Universidad de Castilla-La Mancha, Albacete, Spain). Plasmid encoding Cherry-ts045VSV-G was kindly provided by K. Simons (Max Planck Institute, Dresden, Germany). The GFP-tagged subunit B2 in amino-terminal plasmid (GFP-B2) was amplified from untagged subunit B2 plasmid (Origene Technologies, SC119083, Rockville, MD) with the forward and reverse primers as follows: 5′-ACGTAAGCTTATGGCGCTGCGGGCG-3′ and 5′-ACGTGGATCCCTAATGCTTTGCAGA-3′. The PCR product was then subcloned into pEGFP-C3 vector after cutting with HindIII and BamHI restriction enzymes.
Immunofluorescence Microscopy
HeLa cells were fixed and processed as described previously (53). Working dilutions of antibodies were as follows: anti-GM130 (1:1000); anti-Golgin97 (1:300); anti-TGN46 (1:500). GFP-B2 was visualized directly by the fluorescent emission of GFP. The images were processed using ImageJ software. Colocalization quantitative analysis was done by calculating the number of green pixels that colocalize with red pixels divided by the total number of green pixels of each cell. Background subtraction was performed before the analysis.
Inversal FRAP
iFRAP experiments were carried out using a Leica TCS SL with argon and HeNe lasers attached to a Leica DMIRE2 inverted microscope equipped with an incubation system with temperature and CO2 control as reported previously (48). All experiments were performed at 37 °C and 5% CO2. Cells were grown on 35-mm dishes, transfected with GFP-subunit B2, and incubated for 24 h at 37 °C. For visualization of GFP, images were acquired using a ×63 oil immersion objective lens (NA 1.32), 488-nm laser line (20% powered), excitation beam splitter RSP 500, and emission range detection as follows: 500–600 nm with the confocal pinhole set at 4.94 Airy units to minimize changes in fluorescence due to protein-GFP moving away from the plane of focus. The whole cytoplasm staining (with the exception of the Golgi complex) of the GFP-subunit B2-transfected cell was photobleached using 50–80 scans with the 488-nm laser line at full power. Post-bleach images were monitored at 5-s intervals for 15 min. The excitation intensity was attenuated to ∼5% of the half-laser power to avoid significant photobleaching. To evaluate the results, the observed fluorescence equilibration in the unbleached region (the Golgi complex) was quantified using ImageJ software. For each time point, the loss of total fluorescent intensity in the unbleached region of interest was calculated as shown in Equation 1,
where It is the average intensity of unbleached region of interest at point t, and I0 is the average pre-bleach intensity of the region of interest. Fitting of iFRAP curves was performed with Graphpad Prism software version 5.0 (Graphpad Software, San Diego) and modeled assuming two-phase exponential decay iFRAP, whereas they were equally well modeled with the one-phase exponential decay Equation 2,
where Y started at span + plateau and decayed to plateau with the rate constant K. Half-time was calculated as 0.69/K. Afterward, data were plotted as fluorescence intensity that remained in the Golgi versus time. Mobile fraction (MF) was calculated as shown in Equation 3,
where Fpre was the initial fluorescence intensity and Fend the final recovered fluorescence intensity. Statistical analysis was performed by one-tailed Student's t test.
Golgi and TGN pH Measurements
To measure the pH of the Golgi stack and the TGN, cells were grown on 35-mm dishes and transfected with GalT-pHluorin (for the Golgi stack) or TGN38-pHluorin (for the TGN) and incubated overnight at 37 °C. Golgi-associated Golgi-pHluorin or TGN-associated TGN-pHluorin ratiometric fluorescence intensities were measured. Thereafter, cells were incubated with seven different pH calibration buffers containing 70 mm NaCl, 70 mm KCl, 1.5 mm K2HPO4, 1 mm MgSO4, 10 mm HEPES, 10 mm MES, 2 mm CaCl2, 10 mm glucose, 0.01 mm nigericin, and 0.01 mm monensin. These seven ratio measurements established a linear regression from which initial in situ Golgi or TGN pH measurements were extrapolated. Measurements were obtained with a Leica TCS-SP5 confocal microscope (Leica Microsystems Heidelberg, Manheim, Germany). The excitation was performed with 470- and 405-nm filters, and emission was detected at 508 nm with the confocal pinhole set at 1 Airy unit.
LifeAct Assay
Cells were grown on 35-mm dishes and cotransfected with LifeAct-RFP and GFP subunit B2 plasmids for 24 h at 37 °C. A single cell was recorded with a Leica TCS-SP5 confocal microscope. Images were taken at 0.5-s intervals for 2.5 min with ×63 glycerol immersion objective lens at 1 Airy unit with 512 × 512 pixel frame size. The microscope was equipped with temperature and CO2 control.
Protein Transport Assays
To examine the VSV-G protein transport, HeLa cells were grown on 35-mm dishes, transfected with the respective pools of siRNAs, and incubated at 37 °C for 72 h. The cells were then transfected with Cherry-ts045VSV-G, seeded on coverslips, and incubated overnight at 40 °C. For post-Golgi transport assay, cells were incubated for 2 h at 20 °C and then shifted to 32 °C to synchronize VSV-G transport. For the ER to the Golgi pathway, cells were shifted directly to 32 °C. Cycloheximide (100 μg/ml) was added 30 min before the temperature shift. At the indicated times, cells were fixed and processed for immunofluorescence analysis.
To examine the VSV-G protein transport biochemically, stable HeLa cells that constitutively express VSV-G-GFP were grown on 35-mm dishes, transfected with the respective pools of siRNAs, and incubated at 37 °C for 72 h. Cells were then incubated at 40 °C for 24 h. Cycloheximide was added to a final concentration of 100 μg/ml (for the last 30 min), and cells were then shifted to 32 °C to allow protein transport. At the indicated times, cells were solubilized with 0.5% SDS and 1% 2-mercaptoethanol (0.1 ml/dish) and heated to 100 °C for 10 min. A portion of lysate was digested with Endo H following the manufacturer's protocol and then subjected to SDS-PAGE on 8% gels. VSV-G-GFP was visualized by immunoblotting with a polyclonal anti-GFP.
For the soluble protein secretion experiments, HeLa cells were starved for 30 min in Met/Cys-free medium and then pulse-labeled with 20 μCi of [35S]Met/Cys mix per well in a six-well plate for 10 min. Cells were then rinsed in cold complete medium and shifted to 19 °C for 3 h to allow accumulation at the Golgi of 35S-labeled protein synthesized de novo. Thereafter, cells were washed twice with 5% BSA in PBS and then transferred to 37 °C. To determine the kinetics of secretion, culture supernatants were collected at the indicated times. 35S-Labeled secreted proteins were precipitated with 20% trichloroacetic acid (TCA), washed in cold acetone, and quantified by scintillation counting. To determine total incorporation of [35S]Met/Cys into cellular proteins, cells were lysed with 0.1 n sodium hydroxide in 0.1% sodium deoxycholate, treated with TCA, and processed as above. As a positive control, cells were treated with 5 μg/ml of brefeldin A.
Isolation of Golgi Membranes
Golgi-enriched fractions from rat liver were obtained as reported previously (54) Adult female Wistar rats were starved for 24 h. Rat livers were extracted after surgery and placed into 200 ml of cold 0.5 m phosphate buffer, pH 6.7, with 0.5 m sucrose. Livers were swirled, squeezed, and cut into several pieces; excess buffer was removed, and liver pieces were homogenized by gently pressing them through a 150-μm mesh stainless sieve with the bottom of a conical flask. 13 ml of this homogenate was added to a discontinuous gradient and centrifuged in an SW-28 rotor at 28,000 rpm for 1 h at 4 °C. 2–3 ml of the Golgi fractions were collected from between the 0.5 and 0.86 m sucrose fractions using a Pasteur pipette. Golgi fractions were diluted to 0.25 m sucrose and centrifuged at 7,000 rpm for 30 min at 4 °C in an SW-28 rotor. Pellets were resuspended in 2 ml of phosphate buffer containing 0.25 m sucrose and centrifuged again at 7,000 rpm for 30 min at 4 °C. The final pellet was resuspended in 4.5 ml of phosphate buffer with 0.25 m sucrose, aliquoted, and frozen in liquid nitrogen. The purity of the Golgi-enriched fractions was checked by Western blot by enrichment of Golgi marker GM130.
Subcellular (Membrane and Cytosol) Fractionation
Cells were grown on 100 mm dishes. After treatments, cells were washed twice in PBS and scraped into XB buffer (20 mm HEPES, 150 mm KCl, pH 7.7) supplemented with protease inhibitors. Extracts were mechanically cracked with an insulin syringe and centrifuged at 1,000 × g for 10 min to remove cell debris and nuclei. The supernatant was subsequently subjected to 60 min of ultracentrifugation at 60,000 rpm using an MLA-130 rotor (Beckman Coulter Inc., Brea, CA) at 4 °C. The resulting supernatant was the cytosolic fraction. The pellet was resuspended in the same volume as the cytosolic fraction in XB buffer and again subjected to 60,000 rpm centrifugation for 60 min at 4 °C. The resulting pellet (membrane fraction) was solubilized in radioimmunoprecipitation assay buffer (50 mm Tris-HCl, pH 7.4, 150 mm NaCl, 1% Triton X-100, 1% sodium deoxycholate, 0.1% SDS, 1 mm EDTA plus protease inhibitors). All subcellular fractions were subjected to 10% (v/v) SDS-PAGE.
Immunoprecipitation and Western Blotting Experiments
Whole cell extracts (total lysates) were obtained by adding 200 μl of lysis buffer (20 mm Tris-HCl, pH 7.4, 0.6% CHAPS, 1 mm EDTA, 1 mm EGTA, 1.5% octyl β-d-glucopyranoside, and 10% glycerol) containing protein and phosphatase inhibitors (aprotinin, leupeptin, and pepstatin A and sodium orthovanadate and phenylmethylsulfonyl fluoride, respectively). Samples were passed 10 times through a 25-gauge needle, incubated for 30 min on ice, and centrifuged (14,000 × g for 10 min at 4 °C).
For immunoprecipitation experiments, equal amounts of total lysates (100 μg) were incubated overnight at 4 °C with 30 μl of protein A-Sepharose beads (pre-cleaned lysates). In parallel, 30 μl of protein A-Sepharose beads was incubated for 2 h at 4 °C with the antibody of interest (10 μg) to generate the immunobeads, which were subsequently mixed with pre-cleaned lysates and incubated overnight at 4 °C. The next day, beads were rinsed three times in TA buffer (20 mm Tris-HCl, pH 7.5, 5 mm sodium azide, 1 mm PMSF, 1 mm EGTA). Proteins were eluted from Sepharose beads by adding 20 μl of loading buffer 5× (containing 10% β-mercaptoethanol). Subsequently, samples were processed for Western blotting (18).
For Western blotting experiments, 20 μg of protein were loaded per well. Dilutions of the primary antibodies used were as follows: for anti-subunit A, 1:1,000; anti-subunit B2, 1:4,000; anti-subunit C1, 1:1,1000; anti-subunit a1, 1:4,000; anti-RhoGDI, 1:1,000; anti-TfR, 1:5,000; anti-GFP, 1:25,000; anti-actin, 1:25,000; and anti-caveolin, 1:1,000. Band intensities were measured by densitometry scanning of the film using ImageJ software.
Filamentous/Globular-Actin Isolation
After treatment, cells (grown on 100-mm dishes) were washed twice in PBS and incubated for 30 min in PBS containing 1 mm dithiobis[succinimidylpropionate] containing 250 nm TRITC-phalloidin, rinsed in PBS, and blocked for 15 min in 20 mm Tris-HCl, pH 7.5, diluted in PBS containing 250 nm TRITC-phalloidin. Subsequently, cells were homogenized in lysis buffer (50 mm PIPES, pH 6.9, 50 mm KCl, 5 mm MgCl2, 5% (v/v) glycerol, 0,1% Nonidet P-40, 0.1% Triton X-100, 0.1% Tween 200, 1% 2-mercaptoethanol, and 0.001% antifoam C) containing 250 nm TRITC-phalloidin and protease inhibitors. Lysates were centrifuged at 1,000 × g for 10 min at room temperature to remove cell debris and nuclei. Supernatants were collected and ultracentrifuged at 45,000 rpm for 1 h at room temperature using an MLA-130 rotor (Beckman Coulter Inc., Brea, CA). The pellet contained the F-actin fraction and the supernatant the G-actin fraction. Samples were subsequently analyzed by immunoblotting.
DRM Isolation
Cells were grown on five 100-mm plates per condition. After treatments, cells were washed three times in PBS, collected in 1 ml of HES buffer (20 mm HEPES, 1 mm EDTA, 250 mm sucrose, pH 7.4) containing protease and phosphatase inhibitors, and homogenized by passing 10 times through a 22-gauge needle. Samples were ultracentrifuged at 67,000 rpm for 90 min at 4 °C using an MLA-130 rotor (Beckman Coulter Inc., Brea, CA). The pellets were resuspended with 1 ml of MBS buffer (25 mm MES, 150 mm NaCl, 1% Triton X-100, pH 6.5) containing protease inhibitors, incubated for 20 min at 4 °C, and passed 10 times through a 22-gauge needle. A discontinuous sucrose gradient was then prepared as follows. At the bottom, 1 ml of resuspended pellet was mixed with 1 ml of MBS buffer, 80% sucrose. Subsequently, 2 ml of MBS buffer, 30% sucrose was added. On the top, 1 ml of MBS buffer, 5% sucrose was added. The gradients were ultracentrifuged at 35,000 rpm for 17 h at 4 °C using an SW-55 Ti rotor (Beckman Coulter Inc.). The next day, fractions (500 μl each) were collected from the top to the bottom. Protein was precipitated from each fraction by adding the same volume of acetone and incubated overnight at −20 °C. Thereafter, precipitated protein was resuspended with 100 μl of loading buffer 1× (containing 10% β-mercaptoethanol) and processed by Western blot.
Cell Viability Assay
HeLa cells were transfected with the indicated siRNA (siRNA non-targeting or siRNA against subunit B2). After transfection (72 h), the same number of cells were cultured and fixed. Cell viability was assessed by nuclear DNA staining with Hoechst 33258 as described previously (55). Twenty fields were counted by condition and experiment, comprising a total of at least 200 cells.
Results
V-ATPase Revealed by GFP-tagged Subunit B2 of the V1 Domain Is Enriched in Distal Golgi Compartments andSupports Post-Golgi Protein Transport
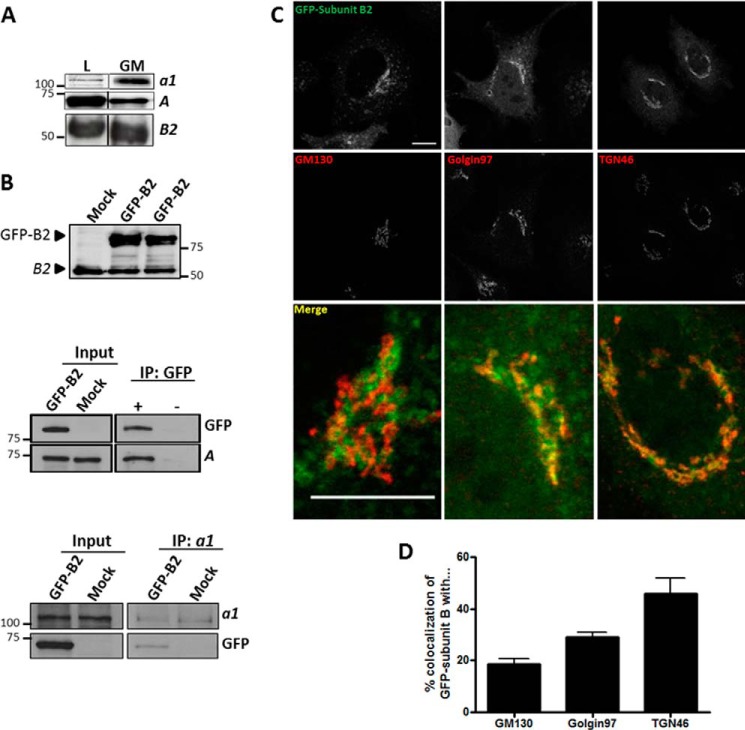
We first examined whether V-ATPases indeed localized in the Golgi complex as reported previously. We first tested for the presence of V-ATPase in the Golgi complex of HeLa cells, as reported previously (31), using a variety of commercially available antibodies against the subunits of V0 and V1 domains. Anti-V1 subunits A and B2 and anti-V0 subunit a1 antibodies recognized bands of the appropriate molecular weight in Golgi fractions isolated from rat liver (Fig. 1A), but unfortunately these antibodies were not suitable for immunocytochemistry. To overcome this drawback, we decided to clone subunits B2 and C1 tagged to GFP, because they both contain actin-binding domains (25, 27). We only succeeded for subunit B2, which was tagged to GFP at the NH2 or COOH terminus (B2-GFP and GFP-B2, respectively). As shown in Fig. 1B (upper panel), anti-subunit B2 antibody recognizes both the endogenous and overexpressed (GFP-B2) proteins at their expected molecular weights. We next examined whether GFP-tagged subunit B2 protein was integrated into V-ATPase holoenzyme. To this end, GFP-B2 or B2-GFP plasmids were expressed in HeLa cells, and cell extracts were immunoprecipitated with anti-GFP antibodies and revealed for subunit A by Western blotting. Endogenous subunit A coimmunoprecipitated with the expressed GFP-B2 form (NH2-tagged) (Fig. 1B, middle panel), but not with the B2-GFP form (COOH-tagged) (data not shown). Consequently, thereafter we used only the GFP-B form. Next, we investigated whether the V1 domain containing GFP-B2 interacts with the V0 domain to form the V-ATPase complex. To this aim, lysates of cells expressing GFP-B2 were immunoprecipitated with antibodies to the V0 subunit a1 and subsequently subjected to Western blotting to GFP. We observed that subunit a1 coimmunoprecipitated with expressed GFP-B2 (Fig. 1B, lower panel), which indicates that the incorporation of GFP-B2 into the V1 domain does not interfere with its interaction with the V0 domain.
FIGURE 1.
V-ATPase is present in Golgi membranes and enriched in distal Golgi compartments. A, Western blotting analysis for the presence of subunits A and B2 of V1 domain and subunit a1 of V0 domain in HeLa cells lysates (L) and in Golgi-enriched fractions (GM) from rat liver. B, cells expressing for 24 h the subunit B2 tagged to GFP in carboxyl terminus (GFP-B2) and non-transfected cells (Mock) were lysed, subjected to Western blotting, and stained with specific antibodies to subunit B2 to see the endogenous and overexpressed subunits (upper panel). GFP-B2-transfected cells were immunoprecipitated (IP) with antibodies to GFP and subsequently subjected to Western blotting for the presence of the V1 subunit A domain (middle panel). Cells expressing GFP-B2 were equally lysed but immunoprecipitated with antibodies to subunit a1 and subsequently subjected to Western blotting for the presence of the GFP tag (lower panel). C, HeLa cells transfected with GFP-subunit B2 construct (expressed for 24 h) were fixed and stained with antibodies against cis- (GM130), trans-Golgi (Golgin97), or TGN (TGN46) markers. Bar, 10 μm. D, quantitative analysis of results shown in B from at least 45 cells/marker of three independent experiments.
We next examined the subcellular localization of expressed GFP-B2 in HeLa cells. Cells were transfected with GFP-B2 and fixed after several expression times (from 6 to 24 h). GFP-B2 was localized both to cytoplasmic vesicular structures that may correspond to endosomes and lysosomes (identified by their annular fluorescence staining) and to a juxtanuclear and reticular structure identified as the Golgi complex by its colocalization with a variety of Golgi markers. Quantitative confocal image analysis showed that GFP-B2 mainly localized in distal Golgi cisternae, as indicated by its strong colocalization with trans (Golgi97) and trans-Golgi network (TGN46) protein markers, in comparison with other more proximal Golgi markers (GM130) (Fig. 1, C and D). This subcellular Golgi distribution of the V-ATPase correlates well with the diminishing pH gradient recorded in the Golgi of HeLa cells (Table 1).
TABLE 1.
Depletion of subunit B2 raises Golgi and TGN pH
Ratiometric measurements of pH in the Golgi stack and in the TGN in HeLa cells expressing (72 h) non-targeting siRNA (NT siRNA) or siRNA to subunit B2 (siRNA-B2). Thereafter, cells were transfected with Golgi-pHluorin or TGN-pHluorin constructs for 18 h. Data represent means ± S.D. of at least three independent experiments. Significant differences with respect to NT siRNA (***, p ≤ 0.001) and to intra-Golgi pH (¶, p ≤ 0.001) using Student's t test are shown. n is number of measured cells.
| Golgi pH | TGN pH | |
|---|---|---|
| NT siRNA | 6.88 ± 0.07 (n = 85) | 6.16 ± 0.06 (n = 32)¶ |
| siRNA-B2 | 7.50 ± 0.07 (n = 94)*** | 6.70 ± 0.07 (n = 37)¶*** |
The pharmacological inhibition of V0 domain blocks V-ATPase activity and in turn alters the pH (49) as well as the Golgi-associated protein transport (44, 46, 56), but little is known about the specific contribution of each V1 subunit in V-ATPase activity. We then knocked down the expression of subunit B2 using siRNA technology to see whether the subunit is essential for V-ATPase activity and its role in Golgi-associated protein transport. The silencing of subunit B2 (∼80% without alteration in the subunit B1 expression; Fig. 2A) did not perturb cell viability (Fig. 2B). HeLa cells transiently transfected with the transmembrane Cherry-VSV-G protein and silenced for subunit B2 showed a large accumulation of Cherry-VSV-G at the Golgi 90 min after the temperature shift. In comparison, in non-targeting siRNA (NT siRNA) transfected cells, the viral glycoprotein was observed in transit to the plasma membrane at 90 min (Fig. 2C). In contrast, ER-to-Golgi protein transport of VSV-G was not altered in subunit B2-depleted cells (Fig. 2, D and E), which was in accordance with previous results obtained using V-ATPase pharmacological inhibitors (44). In addition, the secretion of radioactive-labeled soluble proteins transported from the Golgi to the extracellular medium was also significantly blocked in knockdown cells (Fig. 2F). These impairments in protein transport caused by the depletion of subunit B2 could be attributable to the disruption of Golgi pH, because V-ATPase is the main supplier of protons in endomembranes, including the Golgi complex (6, 7). To explore this, we measured the pH both in the Golgi stack (Golgi pH) and in the TGN (TGN pH) using pHluorin-tagged sensors (52). As expected, control cells showed that the pH in the TGN was lower than in the Golgi (Table 1), and knockdown cells displayed severe alkalization of both Golgi compartments. Curiously, the alkalization effect was stronger in the Golgi stack than in the TGN (Table 1).
FIGURE 2.
V-ATPase subunit B2 depletion blocks post-Golgi trafficking of transmembrane and soluble secretory proteins. A, total cell lysates from HeLa cells previously transfected (72 h) with non-targeting siRNA pool (NT siRNA) or specific siRNA pool to subunit B2 were subjected to immunoblot analysis using monospecific antibodies to subunit B2 and subunit B1. On the right, quantitative analysis of immunoblots is also shown. Values are the mean ± S.D. from four independent experiments. Statistical analysis versus control (NT siRNA) using Student's t test, ***, p ≤ 0.001, is shown. A.U., arbitrary units. B, after transfection with NT siRNA or siRNA-B2, HeLa cells were stained with DAPI to study cell viability. Apoptotic cells were counted. Graph represents the mean of percentage of apoptotic cells ± S.D. of four independent experiments. C, NT siRNA- and siRNA-B2-expressing HeLa cells were transfected with Cherry-ts045VSV-G and incubated overnight at 40 °C and then at 20 °C for 2 h, showing the viral protein accumulated in the ER and in the Golgi, respectively. When cells were shifted from 20 to 32 °C for 90 min, VSV-G is seen in transport carriers and at the plasma membrane in the case of cells transfected with NT siRNA but not to those cells with siRNA-B2. Bar, 10 μm. Graphs on the right correspond to quantitative analysis of results shown in B. D, representative images of HeLa cells transfected with Cherry-ts045VSV-G and non-targeting (NT siRNA) or subunit B2 (siRNA-B2) siRNAs. After overnight incubation of cells at 40 °C, temperature was shifted to 32 °C for 30 min and VSV-G protein was transported to the Golgi. Bar, 10 μm. E, biochemical transport assay for VSV-G-GFP using Endo H assay. HeLa cells constitutively expressing VSV-G-GFP were transfected for 72 h with NT siRNA or siRNA-B2 and incubated at 40 °C for the last 24 h. The cells were then shifted at 32 °C to induce the transport of VSV-G from the ER, lysed at indicated times, and subjected to Endo H treatment. R and S indicate Endo H-resistant and Endo H-sensitive forms, respectively. The ratio of the amount of Endo H-resistant form to that of total (R + S) amount is plotted. Values are represented as the mean ± S.D. of three independent experiments, and no significant differences were found. F, HeLa cells transfected with NT siRNA or siRNA-B2 for 72 h were pulse-labeled with [35S]Met/Cys, incubated at 19 °C for 3 h, and shifted to 37 °C for 60 min. Secreted proteins of the culture medium and cell lysates were precipitated and quantified by scintillation counting. As positive control, the secretion assay was performed in cells treated with brefeldin A (BFA, 5 μg/ml). Results are the mean ± S.D. from three independent experiments. Significant differences with respect to the control (NT siRNA) using Student's t test; *, p ≤ 0.05, and **, p ≤ 0.01.
Overall, these results show that V-ATPase revealed by GFP-tagged subunit B2 localized in the Golgi, particularly in distal compartments, where it is necessary for post-Golgi trafficking. GFP-B2 subunit is a tool that facilitates the study of the dynamics and molecular interactions of V-ATPase at the Golgi complex.
Actin-depolymerizing Agents Induce the V1-V0 Disassociation of the V-ATPase at the Golgi Complex
On the one hand, we previously reported that actin depolymerization raises the pH in the Golgi and alters Golgi-to-ER and post-Golgi trafficking (47–49). These results are similar to those recorded after the pharmacological inhibition of V-ATPase using bafilomycin and concanamycin (44). On the other hand, it has been reported that subunits B and C of V-ATPase V1 domain interact with F-actin and/or G-actin (26, 28). Taking all these observations into account, we hypothesized that V-ATPase activity at the Golgi could be regulated by actin dynamics (49, 51, 57). To test this hypothesis, we first examined whether the V1-V0 association is perturbed in the presence of actin toxins that either depolymerize (latrunculin B (LtB); cytochalasin D (CytD)) or stabilize (jasplakinolide (JpK)) actin. As a positive control, we cultured HeLa cells in the absence of glucose and fetal bovine serum (−Glu/FBS), because this treatment reversibly uncouples V-ATPase domains both in yeast (16) and in mammalian cells (17). Cells were incubated with LtB (500 nm/90 min), CytD (1 μm/90 min), JpK (500 nm/90 min), or −Glu/FBS (4 h), lysed, and immunoprecipitated with anti-subunit A antibodies and subjected to Western blotting for subunit a1 (which forms part of V0 complex). The ratio between a1/A (Fig. 3A) subunits was significantly lower in LtB- and in −Glu/FBS-treated cells (but not in JpK-treated cells) when compared with untreated cells (control) (Fig. 3A). Similar results were obtained with CytD (data not shown). Therefore, taking these results into account, we postulated that actin depolymerization would lead to an enrichment of V1 subunits in the cytosol. We obtained highly pure membrane and cytosol fractions (respectively identified by the presence of transferrin receptor/TfR and RhoGDI markers) from LtB- or JpK-treated cells, and we evaluated the presence of subunits A and B2 in both fractions by Western blotting. Unlike JpK, LtB significantly enriched both subunits in the cytosol (Fig. 3B).
FIGURE 3.
Actin depolymerization induces the disassociation of V0 and V1 domains. A, HeLa cells treated with LtB (500 nm for 90 min), JpK (500 nm for 90 min), or cells starved of glucose and FBS (−Glu/FBS; for 4 h) were lysed and subjected to Western blotting (WB) (input) or immunoprecipitation (IP) with anti-subunit A antibodies and Western blotting to subunits a1 or A. Supernatant (SN) from immunoprecipitation. A representative experiment is shown. Quantitative analysis showing the ratio between V0 and V1 subunits (ratio a1/A) from four independent experiments. Results are represented as means ± S.D. Statistical analysis using Student's t test versus control; **, p ≤ 0.01. C, control; Lt, total lysate; P, pellet. B, post-nuclear supernatants of HeLa cells untreated (C) or treated with LtB (1 μm for 90 min) or Jpk (1 μm for 90 min) were ultracentrifuged, and the pellet (membrane fraction) and the supernatant (cytosol fraction) were subjected to SDS-PAGE. Western blotting was revealed with antibodies to TfR (membrane protein marker), Rho guanidine dissociation inhibitor (RhoGDI; a cytosolic protein marker), and to subunits A and B2. Quantitative results are represented as means ± S.D. from five independent experiments. Amounts of subunits A and B2 in cytosol and membrane fractions were normalized to RhoGDI and TfR, respectively, before the cytosol/membrane ratio was determined. Statistical analysis using Student's t test versus control; *, p ≤ 0.05, is shown.
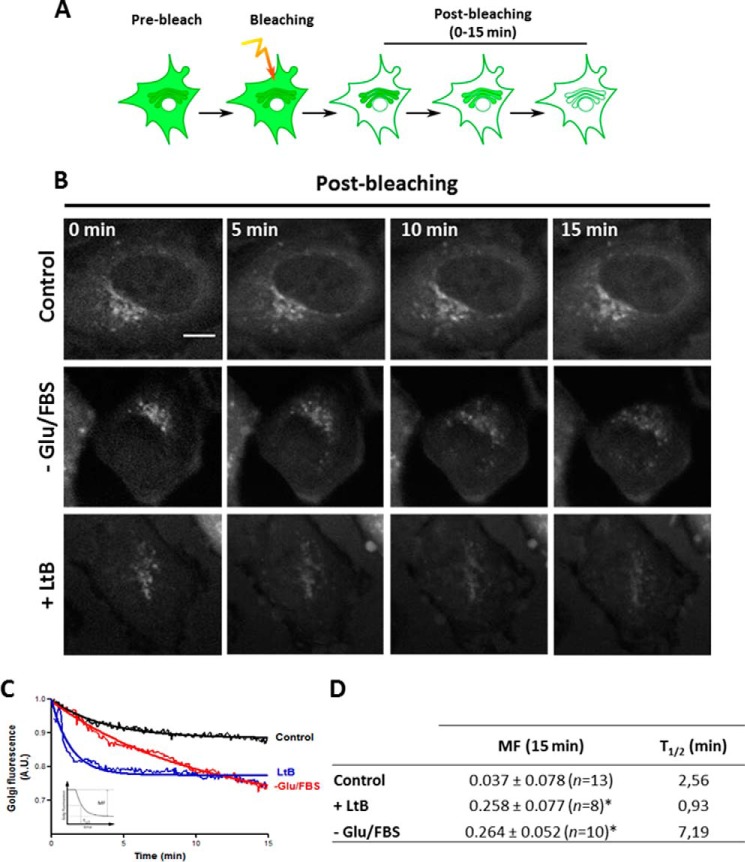
Because we observed that the GFP-B2 subunit localizes in the Golgi complex (Fig. 1B), we next used iFRAP microscopy to visualize whether actin depolymerization induces the release of the GFP-B2 subunit from the Golgi, as a demonstration of the dissociation of V1-V0 domains in vivo. To this end, in cells expressing GFP-B2, we bleached the cytoplasm, except the Golgi, and we measured the dissociation of the V1 domain by the loss of fluorescence at the Golgi (Fig. 4A). Sequential pictures obtained from supplemental movies 1–3 indeed showed the loss of the Golgi-associated GFP-B2 fluorescence after incubation of cells in the absence of glucose and FBS (−Glu/FBS) and with LtB (Fig. 4B; also compare supplemental movie 2/−Glu/FBS, and supplemental movie 3/LtB with supplemental movie 1/control). Quantitative image analysis showed that LtB and −Glu/FBS caused a similar loss of the total GFP-B2 fluorescence at the Golgi at 15 min (Fig. 4C and mobile fraction/MF values shown in D). However, their respective kinetics was different, the loss being much faster for LtB than for −Glu/FBS, as shown by their respective t½ (Fig. 4D). Therefore, actin depolymerization promotes the V0-V1dissociation of Golgi-associated V-ATPase and therefore alterations in its activity and the resulting pH.
FIGURE 4.
Actin depolymerization promotes the release of GFP-B2 subunit from the Golgi to the cytosol. A, schematic representation of the iFRAP experiment, whose results are shown in B. The cytoplasm of HeLa cells expressing GFP-B2 was bleached except for the Golgi area, and subsequently the fluorescence of the Golgi was recorded. B, representative sequential images of time-lapse microscopy of GFP-B2 expressing HeLa cells treated with LtB (500 nm) or cells that were grown in normal culture medium (control) or in medium without glucose and FBS (−Glu/FBS; 4 h). Bar, 10 μm. C, inversal FRAP decay curves of the Golgi fluorescence from GFP-B2 shown in B. Golgi fluorescence values were obtained every 5 s and for a total period of 15 min. Inset, corresponds to a graphic representation of parameters measured in the table. A.U., arbitrary units. D, mobile fraction (MF) and t½(min) for GFP-B2 fluorescence at the Golgi. Statistical analysis versus control cells using the one-tail Student's t test; *, p ≤ 0.05.
V1 Domain Subunits B2 and C1 Interact with Actin in Vivo
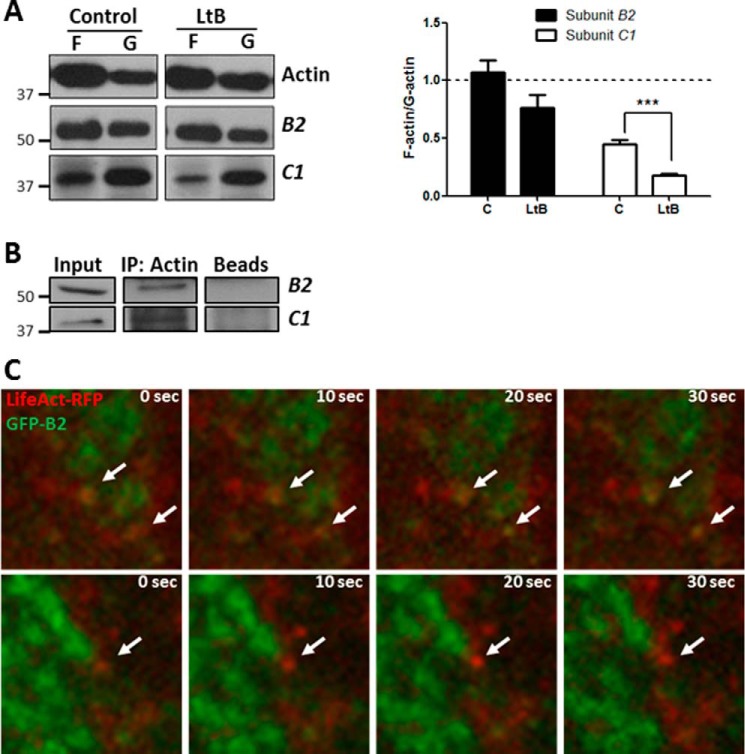
We next approached how microfilaments regulate V-ATPase activity at the Golgi. It is important to note that in HeLa cells V-ATPase is not present at the plasma membrane, which so far is the only molecular interaction where actin and V-ATPase have been described (25, 27). Therefore, to examine whether microfilaments interact with subunits B2 and C1 at endomembranes in HeLa cells, we obtained F- and G-actin-enriched fractions from untreated and LtB-treated cells, and we subsequently used Western blotting to examine the presence of both subunits. In untreated cells, B2 and C1 were indeed present in F- and G-actin fractions, with the former almost equally localized in both fractions and the latter more enriched in the G-actin fraction. After LtB treatment, unlike subunit B2, subunit C1 showed a significant reduction of the F-/G-actin ratio and was thus more enriched in the G-actin fraction (Fig. 5A). These results suggest that subunit C1 shows more affinity for G-actin, unlike subunit B2 that does not show any preference for F- or G-actin. Immunoprecipitation experiments with anti-actin antibodies and subsequent Western blotting for subunits B2 or C1 showed that indeed both V1 subunits interacted with actin, with apparently stronger affinity for C1 (Fig. 5B). Overall, our results suggest that in HeLa cells the molecular interaction between actin and V-ATPase in endomembranes indeed occurs through subunits B2 and C1.
FIGURE 5.
Actin interacts with V-ATPase V1 domain subunits B2 and C1. A, F-/G-actin fractionation of HeLa cells untreated (control, C) or treated with LtB (1 μm; 90 min). Samples were subjected to Western blotting analysis for actin and subunits B2 and C1. On the right, quantitative analysis of results shown on the left. Statistical analysis using Student's t test of results versus respective control from three independent experiments; ***, p ≤ 0.001. B, coimmunoprecipitation experiments with anti-actin (IP: actin) antibody beads conjugated from HeLa cell lysates and subsequent Western blotting analysis for the presence of subunits B2 and C1. C, high magnification of Golgi region snapshots of a 10-s interval time-lapse confocal supplemental movie 4 (upper panel) and supplemental movie 5 (bottom panel) from HeLa cells co-expressing GFP-B2 (green) and the F-actin probe Lifeact-RFP (red). Discrete accumulations of F-actin colocalizing with GFP-B2 at the Golgi are indicated by arrows. Scale bar, 1 μm.
We next analyzed whether F-actin colocalized with V-ATPase at the Golgi complex. The difficulty to visualize F-actin at the Golgi is well known, although there is both functional and morphological evidence of its presence (57–59). To this end, we used the LifeAct-RFP probe (Fig. 5C) (60). HeLa cells co-expressing LifeAct and GFP-B2 showed at the Golgi a small fraction of discrete actin-positive punctae adjacent to GFP-B2-positive zones (Fig. 5C, arrows; supplemental movies 4 and 5), which suggests that F-actin could transiently associate with V-ATPase in Golgi membranes.
Actin Depolymerization-induced V1-V0 Domain Disassociation also Occurs by Disruption of DRMs
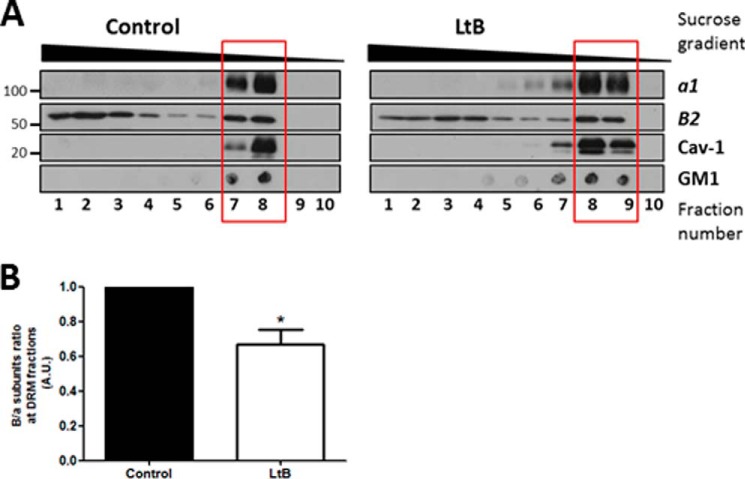
It has been reported that several V-ATPase subunits are major components in Golgi membrane-derived detergent-insoluble complexes, which are the Golgi equivalent to sphingomyelin- and cholesterol-enriched domains isolated as detergent-resistant membranes from total cell extracts (DRMs) (35). In contrast, actin filaments are reported to maintain the integrity of DRMs (61). Then, we reasoned that besides its interaction with subunits B and C, actin could also indirectly regulate association of V1-V0 domains by maintaining the integrity of DRMs. To test this possibility, we isolated total DRMs from control (untreated) and LtB-treated HeLa cells and examined the presence of V0 and V1 subunits a1 and B2, respectively, in DRMs and detergent-sensitive membrane fractions (Fig. 6A). Unlike polarized epithelial cells (14), HeLa cells do not express V-ATPase at the plasma membrane, and therefore, we assumed that the putative source of V-ATPase in DRMs would be endomembranes, mainly from the Golgi (35) and late endosomes and lysosomes (62). DRMs were identified by the presence of caveolin-1 and the GM1 ganglioside. Subunit B2 distributed both in detergent-sensitive membranes and in DRMs (red square) (Fig. 6A), but subunit a1 was exclusively located in DRMs. This difference in distribution is explained because subunit B2 is synthesized in the cytosol and is not always coupled to the V0 domain. When HeLa cells were treated with LtB, DRMs were partially disrupted, as revealed by the presence of DRMs markers in heavier sucrose fractions. The ratio of B2/a1 (V1–V0) subunits in DRMs was significantly reduced (Fig. 6B). To confirm that the presence of cholesterol- and sphingomyelin-enriched domains in Golgi membranes are necessary for V-ATPase function, we manipulated lipid homeostasis in this organelle. Thus, we exogenously added short chain ceramide-C6, the non-metabolizable enantiomer l-ceramide-C6 (N-hexanoyl-l-erythrosphingosine) as a control, and the enantiomer d-ceramide-C6 (N-hexanoyl-d-erythrosphingosine), which is incorporated into Golgi membranes and locally increases levels of short-chain sphingomyelin, short-chain glucosylceramide, and diacylglycerol. As a consequence, there is a reduction in the formation of cholesterol/sphingomyelin-enriched domains into the Golgi (63). Thereafter, we measured Golgi and TGN pH in cells transfected with Golgi- or TGN-pHluorin constructs, and we subsequently cells treated with d- or l-ceramide-C6 (20 μm for 30 min each). d-Ceramide-C6, but not l-ceramide-C6, significantly increased both Golgi and TGN pH (Table 2). Knowing that V-ATPase is the main source of H+ in the Golgi lumen, our results strongly suggest that the integrity of DRMs in Golgi membranes is essential for V-ATPase activity. Altogether, these results suggest that actin could also indirectly regulate V1-V0 association of V-ATPase in Golgi membranes by maintaining the integrity of DRMs.
FIGURE 6.
Disassociation V1-V0 domains by actin depolymerization also occurs by the disruption of DRMs. A, DRMs were obtained from HeLa cells untreated or treated with LtB (1 μm; 90 min) submitted to solubilization in 1% Triton X-100 at 4 °C. The Triton X-100 fractions were subsequently loaded at the bottom of a sucrose gradient as indicated under “Experimental Procedures.” After centrifugation, fractions were collected from the top (from 10/top to 1/bottom) and analyzed by SDS-PAGE followed by Western blotting to detect caveolin 1 (Cav-1) and subunits B2 and a1. For GM1, dot blot was carried out from each fraction collected and before the SDS-PAGE using cholera toxin-HRP. B, quantitative analysis examining the subunits B2/a ratio present in DRMs fractions (indicated with a red box in A). Statistical analysis using Student's t test of results versus control from three independent experiments; *, p ≤ 0.01.
TABLE 2.
Reduction of Golgi membrane lipid order increases Golgi and TGN pH
Ratiometric measurements of Golgi and TGN pH in HeLa cells treated with 20 μm of either l-ceramide-C6 or d-ceramide-C6 during 30 min. Thereafter, cells were transfected with Golgi-pHluorin or TGN-pHluorin constructs for 18 h. Data represent means ± S.D. of three independent experiments. Significant differences with respect l-ceramide-C6 (***, p ≤ 0.001) using Student's t test is shown. n, number of measured cells.
| Golgi pH | TGN pH | |
|---|---|---|
| l-Ceramide-C6 | 6.12 ± 0.04 (n = 48) | 6.56 ± 0.08 (n = 35) |
| d-Ceramide-C6 | 6.47 ± 0.05 (n = 44)*** | 7.07 ± 0.10 (n = 34)*** |
Discussion
We report here that actin regulates the activity of the V-ATPase in the secretory pathway by controlling the association of V1-V0 domains. We suggest that actin carries out such regulation through two non-mutually exclusive mechanisms as follows: (a) its direct interaction with subunits B2 and C1; and (b) indirectly maintaining the organization of lipid rafts. Although V-ATPase localization at the Golgi complex is well known (29–34), we provide here a more detailed picture of V-ATPase distribution along the Golgi stack. Because commercially available antibodies were unsuitable for immunocytochemistry, we cloned the V1 domain subunit B2, which, like subunit C, contains actin-binding sites (27). Importantly, expressed GFP-B2 (GFP tagged to its amino end) was effectively incorporated into the V1 domain without interfering with its interaction with the V0 domain, which indicates that GFP-B2 is indeed a representative tool for studying V-ATPase localization and dynamics in living cells. Expressed GFP-B2 localized V-ATPase in the Golgi complex, where it was visualized along all the compartments, although it was more enriched in distal ones. This Golgi distribution correlates well with the pH gradient in the Golgi (1). Because the mechanisms that generate this pH gradient are unclear (3), our results suggest that a progressive increase in V-ATPase along the cis-to-trans-Golgi axis is the main factor responsible for the progressive fall in pH.
Our group previously reported that the interference in actin dynamics induced by a variety of actin toxins blocks Golgi-to-ER and post-Golgi trafficking alkalinizes the Golgi and induces cisternae swelling. All these were similar to the effects caused by the interference of V-ATPase activity with specific pharmacological agents targeted to V0 domain, such as bafilomycin and concanamycin A (5, 47, 48). Therefore, we postulated that one of the potential mechanisms by which actin participates in Golgi protein transport and flat cisternae morphology, as well as in the maintenance of pH, might be through the regulation of V-ATPase activity, taking into account that both V1 domain subunits B and C contain actin-binding domains (25, 27). Studies on actin interaction with subunit B have focused on V-ATPase of the plasma membrane in osteoclasts and on the role of actin in the recycling of the proton pump to and from intracellular membranes (50, 64). In contrast, the interaction of subunit C with actin cytoskeleton was studied only in vitro using recombinant proteins (25, 28). However, nothing is known about the potential interaction of B and C subunits with actin filaments in the secretory pathway endomembranes and in the Golgi in particular. Our results show an interaction between actin and V1 subunits B2 and C1, because both coimmunoprecipitated with actin. Besides this biochemical evidence for subunit B2-actin interaction, we furthermore visualized this interaction at the Golgi in living cells expressing LifeAct-RFP and GFP-B2. Some spots showed colocalization with both fluorescent probes, although it is true that they were not the majority. It is reasonable to postulate that the interaction between F-actin (revealed by LifeAct) and V-ATPase (revealed by GFP-B2) could be highly dynamic and/or transient. In respect of actin-subunit C1, immunoprecipitation results support the previously suggested role of subunit C directly connecting and modulating the interaction between V1 and V0 domains of the V-ATPase (24), and actin could regulate such a function. We show here that the interaction of the two domains is reduced by LtB, and the V1 domain is concomitantly enriched in the cytosol, which clearly indicates that F-actin maintains the association of the two domains. These biochemical observations were morphologically confirmed by iFRAP experiments, where the LtB treatment GFP-B2 is shifted from the Golgi to the cytoplasm. Taken together, our results strongly suggest that actin could stabilize and/or reinforce functional V-ATPase V1-V0 domain association in vivo through its interaction with both subunits.
Although the B2 and C1 subunits do interact with actin, F/G-actin fractionation and immunoprecipitation experiments suggest more affinity and sensitivity to actin for subunit C1 than for subunit B2. The reason for this is unknown, but it could be related to the functional relevance that both could have in vivo. F-actin interaction to B2 could interfere with its ATP-binding role and consequently with the ATPase activity of subunit A. We cannot discard that such differences might be attributable to the different affinity of antibodies used or even to the presence of an intermediate protein that connects actin with subunit B2. In contrast, F-actin interaction with C1 could promote or stabilize the complex formed by the V0-V1 domains, which is essential for the function of V-ATPase. The question then arises as to the functional relevance of actin-binding sites of both Golgi-associated V1 subunits. As expected, subunit B2 is crucial for the activity of V-ATPase at the Golgi. This conclusion is based on the finding that silencing of subunit B2 raised Golgi and TGN pH values and significantly impaired post-Golgi transport (but not that between the ER to the Golgi) of membrane VSV-G glycoprotein and luminal secretory cargo. The physiological significance of this blockade can be explained by the reduction of subunit B-associated ATP consumption of the V1 domain (9, 65). Similar results were obtained after knockdown of subunit a of the V0 domain (45).
Furthermore, it has been described that V-ATPase also localizes to the lipid rafts in Golgi membranes (35). Our results demonstrate that the LtB treatment induced a partial disorganization of cholesterol and sphingomyelin enriched domains. Moreover, actin depolymerization induces V1-V0 domain dissociation in these domains, which could result from the lipid raft disorganization primarily caused by the disassembly of microfilaments. The role of the actin cytoskeleton in the organization of lipid rafts has been described only in the plasma membrane (61). Taking into account that V-ATPase (revealed by GFP-B2) in HeLa cells is in the Golgi and not in the plasma membrane, and that the disruption of lipid rafts also affects the V0 domain subunit a1, we suggest that actin filaments also participate in the functional association of V1-V0 domain thanks to their role in maintaining lipid raft organization in Golgi membranes. This postulate is supported by results obtained with d-ceramide-C6. This lipid derivative causes the disruption of cholesterol- and sphingomyelin-enriched domains at the Golgi (63), which is accompanied by a swelling of cisternae and alterations in N-glycosylation (63, 66), both strikingly coincidental with impairments at the Golgi occurring after actin depolymerization and pharmacological V-ATPase inhibition (49, 57, 58). Importantly, d-ceramide-C6 treatment also caused Golgi and TGN pH alkalization, which also occurs after actin-depolymerizing toxins (49). Therefore, we establish a functional link at the Golgi between actin dynamics and lipid raft integrity, and consequently V-ATPase activity, which help to maintain pH homeostasis and secretory transport activity (Fig. 7). This new role of actin in the activity of the V-ATPase holoenzyme is compatible with other direct roles of actin in Golgi membranes, such as membrane fission (59) and calcium import (67). It is clear that actin participates in the Golgi architecture and transport functions at different levels (57).
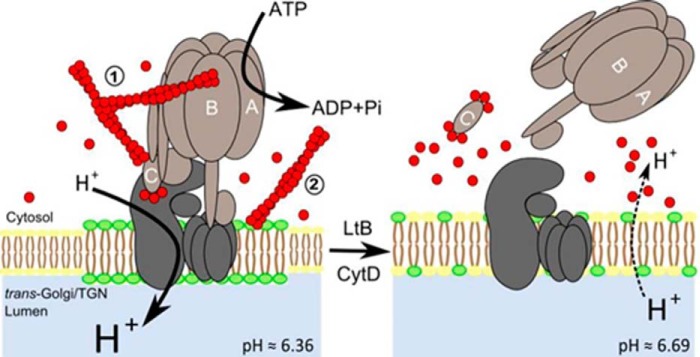
FIGURE 7.
Model of the mechanisms of action by which actin depolymerization promotes dissociation of V1-V0 domains of V-ATPase in Golgi membranes. We suggest two mechanisms, not mutually exclusive, for the regulation of V0-V1 coupling in distal compartments of the Golgi complex. One of them was through the direct interaction between F-actin and V1 subunits B2 and C1; the other one was through the actin-dependent maintenance of the lipid raft domain integrity, which is essential to support V-ATPase activity. When Golgi-associated F-actin is depolymerized through the physiological actin dynamics cycle or widely by the action of actin toxins (LtB or CytD as indicated in the figure), the F-actin-subunit C interaction is not maintained anymore and V1 uncouples from V0, which leads to the rise of the Golgi pH (values are indicated; see Ref. 49). Under physiological conditions, this uncoupling would be very transient thanks to the fast Golgi-associated actin dynamics. However, after massive actin depolymerization caused by actin toxins, a large G-actin pool is generated, and the higher affinity of C subunit for G-actin prevents that the subunits could be reutilized for recoupling both V domains, leading to the permanent dysfunction of V-ATPase at the Golgi and its alkalization. This mechanistic dysfunction could be additionally enhanced by the concomitant disruption of lipid rafts (represented in green) of Golgi membranes where V-ATPase resides. These mechanisms could well occur in other endomembrane systems such endo/lysosomal compartments where V-ATPase is present. The different size of letter H+ indicates different proton concentration consistently to its size. C subunit binds F- or G-actin at both extremes (26). Leakage of luminal Golgi protons to the cytosol is indicated by dashed lines.
Author Contributions
C. S. P. and G. E. designed and coordinated the study and wrote the paper. C. S. P. performed and analyzed all the experiments. A. S. designed, performed, and analyzed experiments shown in Fig. 2D. J. L. provided technical assistance and DNA constructs for experiment shown in Table 1. All the authors reviewed the results and approved the final version of the manuscript.
Supplementary Material
Acknowledgments
We thank Francisco Lázaro-Diéguez for critical reading of the manuscript; Miriam Esgles-Izquierdo and Darya Gorbenko del Blanco (University of Barcelona School of Medicine, Barcelona, Spain) and Agustín Hernández (Universidad Pablo Olavide, Sevilla, Spain) and Yuri Y. Sautin (University of Florida College of Medicine) for technical advice in cloning and immunoprecipitation experiments, respectively; Stephanie Miserey-Lenkei (Institut Curie, Paris) for LifeAct-RFP; Félix Campelo and Juan M. Durán (Centre of Genomic Regulation, Barcelona) for advice with d-/l-ceramide C6; Maite Muñoz for general technical support; and the personnel in charge of Plataformes Tecnològiques (Campus Casanova) for technical support with confocal microscopy and pH measurements. We also thank Robin Rycroft for editorial assistance.
This work was supported in part by Grants BFU2009-07186 and 2012-33932 from Ministerio de Economía y Competitividad del Gobierno de España (to G. E.). The authors declare that they have no conflicts of interest with the contents of this article.

This article contains supplemental Movies S1–S5.
- ER
- endoplasmic reticulum
- TGN
- trans-Golgi network
- V-ATPase
- vacuolar H+-ATPase
- DRM
- detergent-resistant membrane
- F-actin
- filamentous actin
- G-actin
- globular actin
- TfR
- transferrin receptor
- TRITC
- tetramethylrhodamine isothiocyanate
- FRAP
- fluorescence recovery after photobleaching
- iFRAP
- inversal FRAP
- LtB
- latrunculin B
- CytD
- cytochalasin D
- JpK
- jasplakinolide
- Endo H
- endoglycosidase H
- GDI
- guanidine dissociation inhibitor
- NT siRNA
- non-targeting siRNA.
References
- 1. Wu M. M., Grabe M., Adams S., Tsien R. Y., Moore H. P., and Machen T. E. (2001) Mechanisms of pH regulation in the regulated secretory pathway. J. Biol. Chem. 276, 33027–33035 [DOI] [PubMed] [Google Scholar]
- 2. Paroutis P., Touret N., and Grinstein S. (2004) The pH of the secretory pathway: measurement, determinants, and regulation. Physiology 19, 207–215 [DOI] [PubMed] [Google Scholar]
- 3. Casey J. R., Grinstein S., and Orlowski J. (2010) Sensors and regulators of intracellular pH. Nat. Rev. Mol. Cell Biol. 11, 50–61 [DOI] [PubMed] [Google Scholar]
- 4. Weisz O. A. (2003) Organelle acidification and disease. Traffic 4, 57–64 [DOI] [PubMed] [Google Scholar]
- 5. Maeda Y., and Kinoshita T. (2010) The acidic environment of the Golgi is critical for glycosylation and transport. Methods Enzymol. 480, 495–510 [DOI] [PubMed] [Google Scholar]
- 6. Forgac M. (2007) Vacuolar ATPases: rotary proton pumps in physiology and pathophysiology. Nat. Rev. Mol. Cell Biol. 8, 917–929 [DOI] [PubMed] [Google Scholar]
- 7. Jefferies K. C., Cipriano D. J., and Forgac M. (2008) Function, structure and regulation of the vacuolar (H+)-ATPases. Arch. Biochem. Biophys. 476, 33–42 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Kane P. M. (2006) The where, when, and how of organelle acidification by the yeast vacuolar H-ATPase. Microbiol. Mol. Biol. Rev. 70, 177–191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. MacLeod K. J., Vasilyeva E., Baleja J. D., and Forgac M. (1998) Mutational analysis of the nucleotide binding sites of the yeast vacuolar proton-translocating ATPase. J. Biol. Chem. 273, 150–156 [DOI] [PubMed] [Google Scholar]
- 10. Drory O., and Nelson N. (2006) The emerging structure of vacuolar ATPases. Physiology 21, 317–325 [DOI] [PubMed] [Google Scholar]
- 11. Inoue T., Wang Y., Jefferies K., Qi J., Hinton A., and Forgac M. (2005) Structure and regulation of the V-ATPases. J. Bioenerg. Biomembr. 37, 393–398 [DOI] [PubMed] [Google Scholar]
- 12. Demaurex N. (2002) pH homeostasis of cellular organelles. News Physiol. Sci. 17, 1–5 [DOI] [PubMed] [Google Scholar]
- 13. Marshansky V., and Futai M. (2008) The V-type H+-ATPase in vesicular trafficking: targeting, regulation and function. Curr. Opin. Cell Biol. 20, 415–426 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Breton S., and Brown D. (2007) New insights into the regulation of V-ATPase-dependent proton secretion. Am. J. Physiol. Renal. Physiol. 292, F1–F10 [DOI] [PubMed] [Google Scholar]
- 15. Kane P. M. (2012) Targeting reversible disassembly as a mechanism of controlling V-ATPase activity. Curr. Protein Pept. Sci. 13, 117–123 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kane P. M. (1995) Disassembly and reassembly of the yeast vacuolar H+-ATPase in vivo. J. Biol. Chem. 270, 17025–17032 [PubMed] [Google Scholar]
- 17. Nakamura S. (2004) Glucose activates H+-ATPase in kidney epithelial cells. Am. J. Physiol. Cell Physiol. 287, C97–C105 [DOI] [PubMed] [Google Scholar]
- 18. Sautin Y. Y., Lu M., Gaugler A., Zhang L., and Gluck S. L. (2005) Phosphatidylinositol 3-kinase-mediated effects of glucose on vacuolar H+-ATPase assembly, translocation, and acidification of intracellular compartments in renal epithelial cells. Mol. Cell. Biol. 25, 575–589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Parra K. J., and Kane P. M. (1998) Reversible association between the V1 and V0 domains of yeast vacuolar H+-ATPase is an unconventional glucose-induced effect. Mol. Cell. Biol. 18, 7064–7074 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Rubenstein E. M., and Schmidt M. C. (2010) The glucose signal and metabolic p[H+]lux. EMBO J. 29, 2473–2474 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Lu M., Sautin Y. Y., Holliday L. S., and Gluck S. L. (2004) The glycolytic enzyme aldolase mediates assembly, expression, and activity of vacuolar H+-ATPase. J. Biol. Chem. 279, 8732–8739 [DOI] [PubMed] [Google Scholar]
- 22. Xu T., and Forgac M. (2001) Microtubules are involved in glucose-dependent dissociation of the yeast vacuolar [H+]ATPase in vivo. J. Biol. Chem. 276, 24855–24861 [DOI] [PubMed] [Google Scholar]
- 23. Zuo J., Vergara S., Kohno S., and Holliday L. S. (2008) Biochemical and functional characterization of the actin-binding activity of the B subunit of yeast vacuolar H+-ATPase. J. Exp. Biol. 211, 1102–1108 [DOI] [PubMed] [Google Scholar]
- 24. Pérez-Sayáns M., Suárez-Peñaranda J. M., Barros-Angueira F., Diz P. G., Gándara-Rey J. M., and García-García A. (2012) An update in the structure, function, and regulation of V-ATPases: the role of the C subunit. Braz. J. Biol. 72, 189–198 [DOI] [PubMed] [Google Scholar]
- 25. Vitavska O., Wieczorek H., and Merzendorfer H. (2003) A novel role for subunit C in mediating binding of the H+-V-ATPase to the actin cytoskeleton. J. Biol. Chem. 278, 18499–18505 [DOI] [PubMed] [Google Scholar]
- 26. Holliday L. S., Lu M., Lee B. S., Nelson R. D., Solivan S., Zhang L., and Gluck S. L. (2000) The amino-terminal domain of the B subunit of vacuolar H+-ATPase contains a filamentous actin binding site. J. Biol. Chem. 275, 32331–32337 [DOI] [PubMed] [Google Scholar]
- 27. Lee B. S., Gluck S. L., and Holliday L. S. (1999) Interaction between vacuolar H+-ATPase and microfilaments during osteoclast activation. J. Biol. Chem. 274, 29164–29171 [DOI] [PubMed] [Google Scholar]
- 28. Vitavska O., Merzendorfer H., and Wieczorek H. (2005) The V-ATPase subunit C binds to polymeric F-actin as well as to monomeric G-actin and induces cross-linking of actin filaments. J. Biol. Chem. 280, 1070–1076 [DOI] [PubMed] [Google Scholar]
- 29. Zhang F., and Schneider D. L. (1983) The bioenergetics of Golgi apparatus function: evidence for an ATP-dependent proton pump. Biochem. Biophys. Res. Commun. 114, 620–625 [DOI] [PubMed] [Google Scholar]
- 30. Chanson A., and Taiz L. (1985) Evidence for an ATP-dependent proton pump on the Golgi of corn coleoptiles. Plant Physiol. 78, 232–240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Moriyama Y., and Nelson N. (1989) H+-translocating ATPase in golgi apparatus. Characterization as vacuolar H+-ATPase and subunit structures. J. Biol. Chem. 264, 18445–18450 [PubMed] [Google Scholar]
- 32. Hurley D., and Taiz L. (1989) Immunocytochemical localization of the vacuolar H+-ATPase in maize root tip cells. Plant Physiol. 89, 391–395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Young G. P., Qiao J. Z., and Al-Awqati Q. (1988) Purification and reconstitution of the proton-translocating ATPase of Golgi-enriched membranes. Proc. Natl. Acad. Sci. U.S.A. 85, 9590–9594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Glickman J., Croen K., Kelly S., and Al-Awqati Q. (1983) Golgi membranes contain an electrogenic H+ pump in parallel to a chloride conductance. J. Cell Biol. 97, 1303–1308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Gkantiragas I., Brügger B., Stüven E., Kaloyanova D., Li X. Y., Löhr K., Lottspeich F., Wieland F. T., and Helms J. B. (2001) Sphingomyelin-enriched microdomains at the Golgi complex. Mol. Biol. Cell. 12, 1819–1833 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Maeda Y., Ide T., Koike M., Uchiyama Y., and Kinoshita T. (2008) GPHR is a novel anion channel critical for acidification and functions of the Golgi apparatus. Nat. Cell Biol. 10, 1135–1145 [DOI] [PubMed] [Google Scholar]
- 37. Thompson R. J., Nordeen M. H., Howell K. E., and Caldwell J. H. (2002) A large-conductance anion channel of the Golgi complex. Biophys. J. 83, 278–289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Nakamura N., Tanaka S., Teko Y., Mitsui K., and Kanazawa H. (2005) Four Na+/H+ exchanger isoforms are distributed to Golgi and post-Golgi compartments and are involved in organelle pH regulation. J. Biol. Chem. 280, 1561–1572 [DOI] [PubMed] [Google Scholar]
- 39. Ono M., and Hakomori S. (2004) Glycosylation defining cancer cell motility and invasiveness. Glycoconj. J. 20, 71–78 [DOI] [PubMed] [Google Scholar]
- 40. Zhao Y.-Y., Takahashi M., Gu J.-G., Miyoshi E., Matsumoto A., Kitazume S., and Taniguchi N. (2008) Functional roles of N-glycans in cell signaling and cell adhesion in cancer. Cancer Sci. 99, 1304–1310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Kornak U., Reynders E., Dimopoulou A., van Reeuwijk J., Fischer B., Rajab A., Budde B., Nürnberg P., Foulquier F., ARCL Debré-type Study Group, Lefeber D., Urban Z., Gruenewald S., Annaert W., Brunner H. G., et al. (2008) Impaired glycosylation and cutis laxa caused by mutations in the vesicular H+-ATPase subunit ATP6V0A2. Nat. Genet. 40, 32–34 [DOI] [PubMed] [Google Scholar]
- 42. Rivinoja A., Pujol F. M., Hassinen A., and Kellokumpu S. (2012) Golgi pH, its regulation and roles in human disease. Ann. Med. 44, 542–554 [DOI] [PubMed] [Google Scholar]
- 43. Muroi M., Shiragami N., Nagao K., Yamasaki M., and Takatsuki A. (1993) Folimycin (concanamycin A), a specific inhibitor of V-ATPase, blocks intra-cellular translocation of the glycoprotein of vesicular stomatitis virus before arrival to the Golgi apparatus. Cell Struct. Funct. 18, 139–149 [DOI] [PubMed] [Google Scholar]
- 44. Palokangas H., Metsikkö K., and Väänänen K. (1994) Active vacuolar H+ATPase is required for both endocytic and exocytic processes during viral infection of BHK-21 cells. J. Biol. Chem. 269, 17577–17585 [PubMed] [Google Scholar]
- 45. Sobota J. A., Bäck N., Eipper B. A., and Mains R. E. (2009) Inhibitors of the V0 subunit of the vacuolar H+-ATPase prevent segregation of lysosomal- and secretory-pathway proteins. J. Cell Sci. 122, 3542–3553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Yilla M., Tan A., Ito K., Miwa K., and Ploegh H. L. (1993) Involvement of the vacuolar H(+)-ATPases in the secretory pathway of HepG2 cells. J. Biol. Chem. 268, 19092–19100 [PubMed] [Google Scholar]
- 47. Valderrama F., Luna A., Babía T., Martinez-Menárguez J. A., Ballesta J., Barth H., Chaponnier C., Renau-Piqueras J., and Egea G. (2000) The Golgi-associated COPI-coated buds and vesicles contain β/γ-actin. Proc. Natl. Acad. Sci. U.S.A. 97, 1560–1565 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Lázaro-Diéguez F., Colonna C., Cortegano M., Calvo M., Martínez S. E., and Egea G. (2007) Variable actin dynamics requirement for the exit of different cargo from the trans-Golgi network. FEBS Lett. 581, 3875–3881 [DOI] [PubMed] [Google Scholar]
- 49. Lázaro-Diéguez F., Jiménez N., Barth H., Koster A. J., Renau-Piqueras J., Llopis J. L., Burger K. N., and Egea G. (2006) Actin filaments are involved in the maintenance of Golgi cisternae morphology and intra-Golgi pH. Cell Motil. Cytoskeleton 63, 778–791 [DOI] [PubMed] [Google Scholar]
- 50. Chen S.-H., Bubb M. R., Yarmola E. G., Zuo J., Jiang J., Lee B. S., Lu M., Gluck S. L., Hurst I. R., and Holliday L. S. (2004) Vacuolar H+-ATPase binding to microfilaments: regulation in response to phosphatidylinositol 3-kinase activity and detailed characterization of the actin-binding site in subunit B. J. Biol. Chem. 279, 7988–7998 [DOI] [PubMed] [Google Scholar]
- 51. Egea G., Lázaro-Diéguez F., and Vilella M. (2006) Actin dynamics at the Golgi complex in mammalian cells. Curr. Opin. Cell Biol. 18, 168–178 [DOI] [PubMed] [Google Scholar]
- 52. Miesenböck G., De Angelis D. A., and Rothman J. E. (1998) Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature 394, 192–195 [DOI] [PubMed] [Google Scholar]
- 53. Gutiérrez-Martínez E., Fernández-Ulibarri I., Lázaro-Diéguez F., Johannes L., Pyne S., Sarri E., and Egea G. (2013) Lipid phosphate phosphatase 3 participates in transport carrier formation and protein trafficking in the early secretory pathway. J. Cell Sci. 126, 2641–2655 [DOI] [PubMed] [Google Scholar]
- 54. Balch W. E., Dunphy W. G., Braell W. A., and Rothman J. E. (1984) Reconstitution of the transport of protein between successive compartments of the Golgi measured by the coupled incorporation of N-acetylglucosamine. Cell 39, 405–416 [DOI] [PubMed] [Google Scholar]
- 55. Xifró X., García-Martínez J. M., Del Toro D., Alberch J., and Pérez-Navarro E. (2008) Calcineurin is involved in the early activation of NMDA-mediated cell death in mutant huntingtin knock-in striatal cells. J. Neurochem. 105, 1596–1612 [DOI] [PubMed] [Google Scholar]
- 56. Huang C., and Chang A. (2011) pH-dependent cargo sorting from the Golgi. J. Biol. Chem. 286, 10058–10065 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Egea G., Serra-Peinado C., Salcedo-Sicilia L., and Gutiérrez-Martínez E. (2013) Actin acting at the Golgi. Histochem. Cell Biol. 140, 347–360 [DOI] [PubMed] [Google Scholar]
- 58. Valderrama F., Durán J. M., Babià T., Barth H., Renau-Piqueras J., and Egea G. (2001) Actin microfilaments facilitate the retrograde transport from the Golgi complex to the endoplasmic reticulum in mammalian cells. Traffic 2, 717–726 [DOI] [PubMed] [Google Scholar]
- 59. Miserey-Lenkei S., Chalancon G., Bardin S., Formstecher E., Goud B., and Echard A. (2010) Rab and actomyosin-dependent fission of transport vesicles at the Golgi complex. Nat. Cell Biol. 12, 645–654 [DOI] [PubMed] [Google Scholar]
- 60. Riedl J., Crevenna A. H., Kessenbrock K., Yu J. H., Neukirchen D., Bista M., Bradke F., Jenne D., Holak T. A., Werb Z., Sixt M., and Wedlich-Soldner R. (2008) Lifeact: a versatile marker to visualize F-actin. Nat. Methods 5, 605–607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Chichili G. R., and Rodgers W. (2009) Cytoskeleton-membrane interactions in membrane raft structure. Cell. Mol. Life Sci. 66, 2319–2328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Lafourcade C., Sobo K., Kieffer-Jaquinod S., Garin J., and van der Goot F. G. (2008) Regulation of the V-ATPase along the endocytic pathway occurs through reversible subunit association and membrane localization. PLoS ONE 3, 2758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Duran J. M., Campelo F., van Galen J., Sachsenheimer T., Sot J., Egorov M. V., Rentero C., Enrich C., Polishchuk R. S., Goñi F. M., Brügger B., Wieland F., and Malhotra V. (2012) Sphingomyelin organization is required for vesicle biogenesis at the Golgi complex. EMBO J. 31, 4535–4546 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Holliday L. S., Bubb M. R., Jiang J., Hurst I. R., and Zuo J. (2005) Interactions between vacuolar H+-ATPases and microfilaments in osteoclasts. J. Bioenerg. Biomembr. 37, 419–423 [DOI] [PubMed] [Google Scholar]
- 65. Vasilyeva E., Liu Q., MacLeod K. J., Baleja J. D., and Forgac M. (2000) Cysteine scanning mutagenesis of the noncatalytic nucleotide binding site of the yeast V-ATPase. J. Biol. Chem. 275, 255–260 [DOI] [PubMed] [Google Scholar]
- 66. van Galen J., Campelo F., Martínez-Alonso E., Scarpa M., Martínez-Menárguez J. Á., and Malhotra V. (2014) Sphingomyelin homeostasis is required to form functional enzymatic domains at the trans-Golgi network. J. Cell Biol. 206, 609–618 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. von Blume J., Alleaume A.-M., Cantero-Recasens G., Curwin A., Carreras-Sureda A., Zimmermann T., van Galen J., Wakana Y., Valverde M. A., and Malhotra V. (2011) ADF/cofilin regulates secretory cargo sorting at the TGN via the Ca2+ ATPase SPCA1. Dev. Cell. 20, 652–662 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.