Abstract
When aptamers first emerged almost two decades ago, most were RNA species that bound and tagged or inhibited simple target ligands. Very soon after, the ‘selectionologists’ developing aptamer technology quickly realized more potential for the aptamer. In recent years, advances in aptamer techniques have enabled the use of aptamers as small molecule inhibitors, diagnostic tools and even therapeutics. Aptamers are now being employed in novel applications. We review, herein, some of the recent and exciting applications of aptamers in cell-specific recognition and delivery.
Keywords: aptamers, in vitro selection, SELEX, targeted delivery, siRNA delivery
Introduction
Aptamers are nucleic acid binding species capable of tightly binding to and discreetly distinguishing target ligands. In fact, they have often been described as nucleic acid versions of antibodies. However, unlike antibodies, aptamers have yet to elicit immunogenicity in vivo.1–5 Moreover, these molecules are readily amenable to chemical synthesis (thereby decreasing production costs) and can be easily modified during the synthesis process, making them more adaptable for different applications.
Early aptamer selections focused on RNA or DNA that bound small molecule ligands, such as dyes,6 ATP,7 or soluble proteins such as thrombin8 or polymerases9, 10. Since those initial selections, hundreds of targets have been successfully selected against, with the targets for selection ranging from simple organic molecules to proteins, complexes, and, more recently, whole cells and organisms (see the Aptamer Database at http://aptamer.icmb.utexas.edu for a comprehensive compilation11). By varying key parameters of a selection, aptamers with finely tuned physical and functional properties have been selected, and the applications appear almost limitless. For example, RNA aptamers can and have been encoded in expression cassettes and expressed in vivo for use in gene therapy applications (reviewed in 12–14), whereas DNA aptamers, due to their ease of production, are well suited for diagnostic applications.15, 16 Additionally, the inclusion of modified RNA containing 2′deoxy, 2′F, 2′NH3 or 2′OMe into selections (reviewed in 17), have allowed for stabilized aptamers that can now be used in complex biological solutions, such as blood and serum, paving the way for in vivo use of aptamers.
In the current review, we will summarize some advances in aptamer selections and some of the newer applications of aptamers in the most recent few years. In particular, we will focus on developments in targeting cells and cell surface receptors and delivery to cells using aptamers which target the cell surface. For more details about aptamers for gene therapy, aptazymes, riboswitch-type aptamers or other earlier generation aptamers, there are several reviews already available.12, 14, 16, 18
The process of in vitro selection (SELEX)
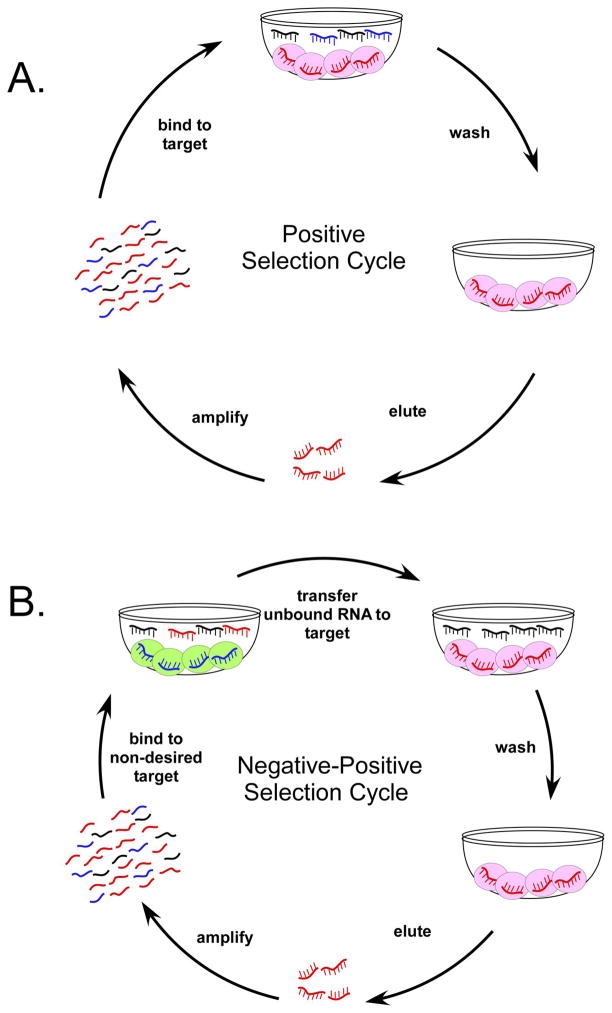
Aptamers are nucleic acid binding species generated by iterative rounds of in vitro selection, or SELEX (Figure 1A).6, 9 Briefly, pools of random-sequence RNA or ssDNA are incubated with target molecules under carefully chosen selection conditions. Binding species are partitioned away from non-binders, amplified to generate a new pool, and the process is repeated until a desired ‘phenotype’ is achieved or until sequence diversity is significantly diminished.
Figure 1.
The process of in vitro selection (SELEX). A). In a traditional selection, random sequence pools (squiggly lines) are incubated with a target (pink orbs). Non-binding species are washed away, and the bound species are eluted and amplified to regenerate the pool for the next round. B). In more complex target selections, the pool is first incubated with a non-target (green orbs) in a negative selection step. Non-bound species from the negative step are added to the target, and the selection proceeds as usual. Over many cycles of selection, high-affinity sequences (aptamers) can be isolated.
Multiple cycles of selection and amplification winnow an initial pool containing upwards of 1015 molecules to only those few species that have the highest affinities and specificities for a target. Previously selected aptamers have typically bound their targets with Kd values in the nanomolar to picomolar range and can discriminate against very closely related targets (e.g., proteins that differ by only a few amino acids19, 20 or targets that differ by slight modifications.21 By including a negative selection step into the selection cycle in which pool sequences are first exposed to non-desired targets to remove non-specific binders, aptamers with very discrete specificities can be isolated (Figure 1B). This very important negative selection step has been crucial in identifying aptamers which can discriminate between closely related protein targets, and more recently, in the development of selections against complex targets such as whole cells.
Whole cell SELEX
Instead of the traditional positive selection for a known target and a negative selection step against a non-target such as a support resin or the filter used for immobilization of the target, selections can be used to parse out a target by subtracting the ‘background’ in the negative step. This approach has been successfully applied to known targets on the cell surface. For example, by combining positive rounds of selection against a cell line engineered to express the RET receptor tyrosine kinase with negative selections against the non-RET expressing parental cell line, Cerchia et al. were able to elicit aptamers which bound that specific receptor.22, 23
This type of ‘toggle’ selection can also be taken to the extreme in which the experiment is performed not by knowing what the target is, but by knowing what one doesn’t want the target to be. In one of the earliest examples of using this subtraction selection technique to parse out aptamers that recognize differences between cells, Ulrich et al. raised aptamers against infective stages (trypomastigotes) of the parasite Trypanosoma cruzi. By including negative selections against the cells of the non-infective stage (epimastigotes), the authors aimed to find aptamers against epitopes specific to the infectious form. The aptamers, which proved to inhibit cell invasion by the parasite, were further used to identify host factors that T. cruzi utilizes for infection.24
Using this or similar approaches, aptamers have been selected which can distinguish rat glioblastoma from microgial cells25 and recognize populations of T cells,26, 27 liver cells,28 lung cells,29 and dendritic cells.30 In one example of whole cell selection, the Tan group performed selections to distinguish between B-lymphocytes and T-lymphocytes. By performing positive and negative selections, respectively, against CEM and Ramos cells, aptamers were selected that could mostly distinguish the target cell type in mixed cultures of cells.27 In a follow-up study, the targets of the aptamers were identified. One aptamer:target pair, sdc8 and its target, protein tyrosine kinase 7, was shown to be internalized, a trait that could prove useful for a number of applications, as will be further described below.31
To illustrate one application of whole cell selections, the Krylov group demonstrated how these selections could funnel directly into biomarker discovery. The group selected ssDNA aptamers against immature and mature dendritic cells and produced two sets of aptamers: one with preference for immature dendritic cells and the other for mature cells. The aptamers were tethered to beads and used to bind cells. These cells were lysed, washed, and the aptamer-bound targets were eluted and analyzed by mass spectrometry. The analysis revealed markers known to be expressed on these cells as well as several markers not previously associated with these cells.30
These works and others like them illustrate how these target-naïve selections could be used for biomarker discovery. Importantly, these and other selections have also shown that aptamers can be selected against different states of a cell (age, differentiation, health, progression of disease, etc.). For example, aptamers have been selected that can distinguish differentiated from non-differentiated neuronal cells32 and cells infected by the vaccinia virus from non-infected cells.33
Targeting cells with aptamers
A toolbox of aptamers that can recognize different cells and different disease or health states of a cell could be readily adapted for in vivo diagnostic as well as therapeutic purposes. While these may ultimately be adapted from aptamers identified ‘blindly’ from whole cell SELEX, the current advances have been made in adapting aptamers that target specific proteins and/or cell surface receptors. A number of these aptamers have been adapted for diagnostic and imaging purposes. For example, Borbas et al. and Hicke et al. both used radiolabeled forms of anti-MUC-1 and anti-tenascin C aptamers, respectively, to image tumors in mice.34, 35
Although aptamers identified by whole cell SELEX have not yet been applied to in vivo imaging and diagnostic applications, they have been adapted for in vitro diagnostic assays. Shangguan et al. screened aptamers generated against T-cell acute lymphoblastic leukemia cancer cells for their ability to type cancer patient samples.27 Although there was some cross-binding between the different types of samples, for the most part, the aptamers bound to their intended cell types. This suggests that a larger subset of aptamers could more definitively type a given cancer sample, as each type of cancer would produce a specific binding profile.36 However, care must be taken in interpreting these types of results, and the Ellington group has recently published a summary of factors that might be considered in using aptamers for cell typing.37
Aptamer-mediated delivery to cells
In addition to biomarker discovery and diagnostics, aptamers can also be selected that not only recognize a cell, but also be endocytosed, potentiating the escorted delivery of appended cargoes into cells.38 How to specifically deliver drugs to certain sites and not to others is a conundrum that is tantamount to the success of a therapeutic. By slightly augmenting the cell-specific aptamer selections described above, aptamers can be selected that can specifically enter and deliver agents into cells.
For the most part, targeted delivery using aptamers have been performed using aptamers selected against receptors known to be internalized, and a variety of different cargoes have now been successfully delivery both in vitro as well as in vivo (Figure 2). For example, the Farokhzad and Langer labs have used an aptamer selected against the prostate specific membrane antigen (PSMA) to deliver nanoparticles to PSMA-expressing LnCap cells for imaging39 and drug delivery.40, 41 Both the Ellington and Sullenger labs also made use of anti-PSMA aptamers, but for delivery of siRNA to LnCap cells. In a simple and elegant design, McNamara et al. delivered anti-plk and anti-bcl2 siRNAs to LnCap cells by appending the siRNA sense strand directly to the anti-PSMA aptamer. This chimera was hybridized to the antisense strand, incubated on cells and effectively knocked down gene expression. The group went on to demonstrate that this delivery system was effective in vivo.42
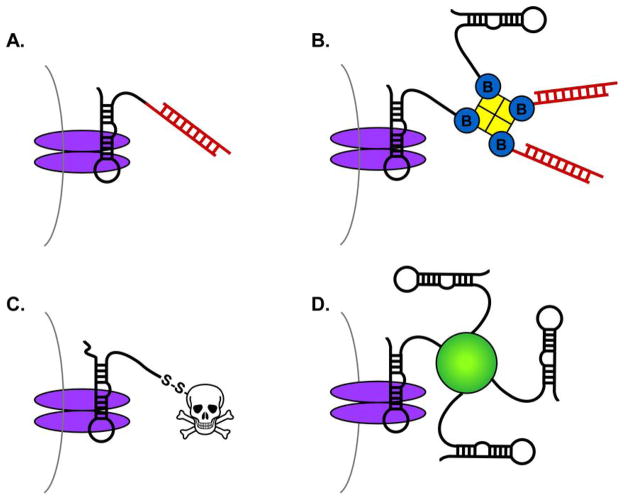
Figure 2.
Aptamer-mediated delivery strategies. A). One strand of siRNA (red) is directly linked to the aptamer (black), and the complement siRNA is annealed via hybridization.42, 47 B). siRNA or other cargoes are biotinylated and loaded onto streptavidin. A biotinylated aptamer mediates delivery of the entire particle.43 C). Toxins or other drugs are directly conjugated to the aptamer for delivery.48, 50 Finally, D). Aptamers are directly bound to nanoparticles.58, 59
In a similar manner, Chu et al. delivered siRNA by loading the four binding pockets of streptavidin with biotinylated siRNA and biotinylated aptamer.43 These constructs also showed knockdown on par with controls in which siRNA was directly transfected into cells. By using a streptavidin scaffold, the group created a more modular complex, in which different siRNA or different aptamers could eventually be co-delivered, or varying ratios of each component could be used. The multivalent construct also has the added potential benefit of garnering avidity effects, a property that was recently exploited by another group who demonstrated siRNA delivery using bivalent aptamer chimeras.44
The Rossi Lab also used aptamer-mediated delivery to inhibit HIV-1. Zhou et al. recognized that gp120 on the surface of an infected cell might allow for targeting siRNA. Indeed, this viral antigen had previously been targeted for siRNA delivery utilizing an anti-GP120 single chain antibody protamine conjugate.45 They created chimeras of anti-gp120 aptamers46 with anti-Tat siRNA. The aptamers, in of themselves, were capable of inhibiting viral replication to some extent, as they could block HIV-1:cell interactions. 46,47 However, the combination of aptamer with aptamer-delivered siRNA essentially obliterated infection in culture.47
Proteins, too, have been targeted to cells using aptamers. For example, the Ellington lab demonstrated almost a 1000-fold increase in the toxicity of the ribosomal toxin gelonin when conjugated to an anti-PSMA aptamer. The toxin normally has difficulty entering the cell, and is, therefore, relatively non-toxic. Enhanced toxicity was only observed for PSMA expressing LNCaP cells. A non-expressing PSMA line, PC3 actually showed a decreased response to the toxin-conjugate when compared to the toxin alone.48
More recently, the Neufeld lab, was able to deliver the enzyme α-L-iduronidase to the lysosomes of cells deficient in this enzyme using an anti-mouse transferrin receptor aptamer. The aptamer conjugates were able to restore enzyme activity in the cells suggesting a potential therapeutic approach for the treatment of lysosomal storage diseases.49
This ability to deliver siRNA or other gene modulators to specific cells via receptor-targeted aptamers should be a great tool for researchers and clinicians alike, perhaps allowing for site-specific control of disease or development genes. Additionally, aptamers identified from selections against cells may also be capable of internalizing and similarly adapted for delivery. For example, Xiao et al. demonstrated that an aptamer previously selected naively against T-cells bound to protein kinase 7 and could be internalized.31 They subsequently adapted this aptamer for targeted delivery of the drug doxorubicin using an acid labile drug conjugate.50
The next 20 years…
Aptamer selections have progressed from targeting and inhibiting the function of isolated proteins or molecules and simple in vitro proof-of-principle concepts to targeting epitopes on whole cell surfaces. Because aptamers can now be selected without prior knowledge of specific cell surface markers or cell surface architecture, the process of in vitro selection can be used to distinguish between different cell or disease types. Indeed a problem with treatment and diagnosis of many diseases is precisely that the specific disease marker may not be known. With cancerous cells, for example, the type of cancer, degree of progression or severity and location may all affect marker types and expression levels. Into the future, selections against whole cells, or diseased tissues should provide us with panels of aptamers that can specifically target phenotypes. These in turn can be adapted to serve as valuable diagnostics or further adapted for the targeted delivery of siRNA or other therapeutics. Further still, when one considers how peptides that home to tumors, tumor vasculature and tissues can be identified from phage display libraries in whole animals,51, 52 it seems that similar approaches for aptamers are likely on the horizon.
In addition to advances in aptamer selections and aptamer-mediated delivery, new mechanisms for function are also beginning to emerge. For example, while aptamers have historically been used as antagonists to their targets, the Sullenger lab has recently demonstrated the use of an aptamer as an agonist. Using an aptamer targeting the cell surface receptor OX40, a TNF family receptor that functions in maintaining T-cell proliferation, the group showed that while one aptamer was inadequate for stimulating the receptor and maintaining T cell activation, aptamer dimers were capable of enhancing T cell activation both in tissue culture and when injected into mice.53 The aptamer dimers were generated via molecular assembly using DNA-based scaffolds that would hybridize to portions of the anti-OX40 aptamer. In this manner, these affinity agents were readily transformed into activators. Similar effects have previously been seen when comparing the function of antibody fragments which bind a single receptor with whole antibodies which can crosslink receptors. The ease with which aptamers can readily and modularly be assembled presents a new paradigm for combinatorial drug design and aptamer engineering.
The progress in aptamer development since its inception in the Gold and Szostak labs has been rapid and impressive. With one aptamer drug already FDA approved (Macugen) and many more in clinical trials, aptamers have already demonstrated therapeutic use. 1–5 Additionally, new methods of generating aptamers as well as refinements in the traditional selections are currently ongoing. Some of these include robotically performed selections,54 FACS-based selections,26 CE-based selections,55 and single bead selections.56, 57 The newest applications of aptamers, only some of which have been reviewed here, and leading edge technologies that are being created will play significant roles in RNA-mediated biotechnology.
Contributor Information
Amy C. Yan, Email: ayan@aecom.yu.edu.
Matthew Levy, Email: mattlevy@aecom.yu.edu.
References
- 1.Anti-vascular endothelial growth factor therapy for subfoveal choroidal neovascularization secondary to age-related macular degeneration: phase II study results. Ophthalmology. 2003;110:979–86. doi: 10.1016/S0161-6420(03)00085-X. [DOI] [PubMed] [Google Scholar]
- 2.Chan MY, Cohen MG, Dyke CK, Myles SK, Aberle LG, Lin M, et al. Phase 1b randomized study of antidote-controlled modulation of factor IXa activity in patients with stable coronary artery disease. Circulation. 2008;117:2865–74. doi: 10.1161/CIRCULATIONAHA.107.745687. [DOI] [PubMed] [Google Scholar]
- 3.Alexander JH, Hafley G, Harrington RA, Peterson ED, Ferguson TB, Jr, Lorenz TJ, et al. Efficacy and safety of edifoligide, an E2F transcription factor decoy, for prevention of vein graft failure following coronary artery bypass graft surgery: PREVENT IV: a randomized controlled trial. Jama. 2005;294:2446–54. doi: 10.1001/jama.294.19.2446. [DOI] [PubMed] [Google Scholar]
- 4.Gilbert JC, DeFeo-Fraulini T, Hutabarat RM, Horvath CJ, Merlino PG, Marsh HN, et al. First-in-human evaluation of anti von Willebrand factor therapeutic aptamer ARC1779 in healthy volunteers. Circulation. 2007;116:2678–86. doi: 10.1161/CIRCULATIONAHA.107.724864. [DOI] [PubMed] [Google Scholar]
- 5.Apte RS, Modi M, Masonson H, Patel M, Whitfield L, Adamis AP. Pegaptanib 1-year systemic safety results from a safety-pharmacokinetic trial in patients with neovascular age-related macular degeneration. Ophthalmology. 2007;114:1702–12. doi: 10.1016/j.ophtha.2007.02.021. [DOI] [PubMed] [Google Scholar]
- 6.Ellington AD, Szostak JW. In vitro selection of RNA molecules that bind specific ligands. Nature. 1990;346:818–22. doi: 10.1038/346818a0. [DOI] [PubMed] [Google Scholar]
- 7.Sassanfar M, Szostak JW. An RNA motif that binds ATP. Nature. 1993;364:550–3. doi: 10.1038/364550a0. [DOI] [PubMed] [Google Scholar]
- 8.Bock LC, Griffin LC, Latham JA, Vermaas EH, Toole JJ. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature. 1992;355:564–6. doi: 10.1038/355564a0. [DOI] [PubMed] [Google Scholar]
- 9.Tuerk C, Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. 1990;249:505–10. doi: 10.1126/science.2200121. [DOI] [PubMed] [Google Scholar]
- 10.Tuerk C, MacDougal S, Gold L. RNA pseudoknots that inhibit human immunodeficiency virus type 1 reverse transcriptase. Proc Natl Acad Sci U S A. 1992;89:6988–92. doi: 10.1073/pnas.89.15.6988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lee JF, Hesselberth JR, Meyers LA, Ellington AD. Aptamer database. Nucleic Acids Res. 2003 doi: 10.1093/nar/gkh094. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yan AC, Bell KM, Breeden MM, Ellington AD. Aptamers: prospects in therapeutics and biomedicine. Front Biosci. 2005;10:1802–27. doi: 10.2741/1663. [DOI] [PubMed] [Google Scholar]
- 13.Famulok M, Mayer G, Blind M. Nucleic acid aptamers-from selection in vitro to applications in vivo. Acc Chem Res. 2000;33:591–9. doi: 10.1021/ar960167q. [DOI] [PubMed] [Google Scholar]
- 14.Famulok M, Hartig JS, Mayer G. Functional aptamers and aptazymes in biotechnology, diagnostics, and therapy. Chem Rev. 2007;107:3715–43. doi: 10.1021/cr0306743. [DOI] [PubMed] [Google Scholar]
- 15.Mairal T, Ozalp VC, Lozano Sanchez P, Mir M, Katakis I, O’Sullivan CK. Aptamers: molecular tools for analytical applications. Anal Bioanal Chem. 2008;390:989–1007. doi: 10.1007/s00216-007-1346-4. [DOI] [PubMed] [Google Scholar]
- 16.Jayasena SD. Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin Chem. 1999;45:1628–50. [PubMed] [Google Scholar]
- 17.Keefe AD, Cload ST. SELEX with modified nucleotides. Curr Opin Chem Biol. 2008;12:448–56. doi: 10.1016/j.cbpa.2008.06.028. [DOI] [PubMed] [Google Scholar]
- 18.Dollins CM, Nair S, Sullenger BA. Aptamers in immunotherapy. Hum Gene Ther. 2008;19:443–50. doi: 10.1089/hum.2008.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Li N, Wang Y, Pothukuchy A, Syrett A, Husain N, Gopalakrisha S, et al. Aptamers that recognize drug-resistant HIV-1 reverse transcriptase. Nucleic Acids Res. 2008;36:6739–51. doi: 10.1093/nar/gkn775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Misono TS, Kumar PK. Selection of RNA aptamers against human influenza virus hemagglutinin using surface plasmon resonance. Anal Biochem. 2005 doi: 10.1016/j.ab.2005.04.013. [DOI] [PubMed] [Google Scholar]
- 21.Bianchini M, Radrizzani M, Brocardo MG, Reyes GB, Gonzalez Solveyra C, Santa-Coloma TA. Specific oligobodies against ERK-2 that recognize both the native and the denatured state of the protein. J Immunol Methods. 2001;252:191–7. doi: 10.1016/s0022-1759(01)00350-7. [DOI] [PubMed] [Google Scholar]
- 22.Cerchia L, Duconge F, Pestourie C, Boulay J, Aissouni Y, Gombert K, et al. Neutralizing aptamers from whole-cell SELEX inhibit the RET receptor tyrosine kinase. PLoS Biol. 2005;3:e123. doi: 10.1371/journal.pbio.0030123. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 23.Pestourie C, Cerchia L, Gombert K, Aissouni Y, Boulay J, De Franciscis V, et al. Comparison of different strategies to select aptamers against a transmembrane protein target. Oligonucleotides. 2006;16:323–35. doi: 10.1089/oli.2006.16.323. [DOI] [PubMed] [Google Scholar]
- 24.Ulrich H, Magdesian MH, Alves MJ, Colli W. In vitro selection of RNA aptamers that bind to cell adhesion receptors of Trypanosoma cruzi and inhibit cell invasion. J Biol Chem. 2002;277:20756–62. doi: 10.1074/jbc.M111859200. [DOI] [PubMed] [Google Scholar]
- 25.Blank M, Weinschenk T, Priemer M, Schluesener H. Systematic evolution of a DNA aptamer binding to rat brain tumor microvessels. selective targeting of endothelial regulatory protein pigpen. J Biol Chem. 2001;276:16464–8. doi: 10.1074/jbc.M100347200. [DOI] [PubMed] [Google Scholar]
- 26.Raddatz MS, Dolf A, Endl E, Knolle P, Famulok M, Mayer G. Enrichment of cell-targeting and population-specific aptamers by fluorescence-activated cell sorting. Angew Chem Int Ed Engl. 2008;47:5190–3. doi: 10.1002/anie.200800216. [DOI] [PubMed] [Google Scholar]
- 27.Shangguan D, Li Y, Tang Z, Cao ZC, Chen HW, Mallikaratchy P, et al. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc Natl Acad Sci U S A. 2006;103:11838–43. doi: 10.1073/pnas.0602615103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Shangguan D, Meng L, Cao ZC, Xiao Z, Fang X, Li Y, et al. Identification of liver cancer-specific aptamers using whole live cells. Anal Chem. 2008;80:721–8. doi: 10.1021/ac701962v. [DOI] [PubMed] [Google Scholar]
- 29.Chen HW, Medley CD, Sefah K, Shangguan D, Tang Z, Meng L, et al. Molecular Recognition of Small-Cell Lung Cancer Cells Using Aptamers. Chem Med Chem. 2008 doi: 10.1002/cmdc.200800030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Berezovski MV, Lechmann M, Musheev MU, Mak TW, Krylov SN. Aptamer-facilitated biomarker discovery (AptaBiD) J Am Chem Soc. 2008;130:9137–43. doi: 10.1021/ja801951p. [DOI] [PubMed] [Google Scholar]
- 31.Xiao Z, Shangguan D, Cao Z, Fang X, Tan W. Cell-Specific Internalization Study of an Aptamer from Whole Cell Selection. Chemistry. 2008;14:1769–75. doi: 10.1002/chem.200701330. [DOI] [PubMed] [Google Scholar]
- 32.Wang C, Zhang M, Yang G, Zhang D, Ding H, Wang H, et al. Single-stranded DNA aptamers that bind differentiated but not parental cells: subtractive systematic evolution of ligands by exponential enrichment. J Biotechnol. 2003;102:15–22. doi: 10.1016/s0168-1656(02)00360-7. [DOI] [PubMed] [Google Scholar]
- 33.Tang Z, Parekh P, Turner P, Moyer RW, Tan W. Generating Aptamers for Recognition of Virus-Infected Cells. Clin Chem. 2009 doi: 10.1373/clinchem.2008.113514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Borbas KE, Ferreira CS, Perkins A, Bruce JI, Missailidis S. Design and synthesis of mono- and multimeric targeted radiopharmaceuticals based on novel cyclen ligands coupled to anti-MUC1 aptamers for the diagnostic imaging and targeted radiotherapy of cancer. Bioconjug Chem. 2007;18:1205–12. doi: 10.1021/bc0700741. [DOI] [PubMed] [Google Scholar]
- 35.Hicke BJ, Stephens AW, Gould T, Chang YF, Lynott CK, Heil J, et al. Tumor targeting by an aptamer. J Nucl Med. 2006;47:668–78. [PubMed] [Google Scholar]
- 36.Shangguan D, Cao ZC, Li Y, Tan W. Aptamers evolved from cultured cancer cells reveal molecular differences of cancer cells in patient samples. Clin Chem. 2007;53:1153–5. doi: 10.1373/clinchem.2006.083246. [DOI] [PubMed] [Google Scholar]
- 37.Li N, Ebright J, Stovall G, Chen X, Nguyen H, Singh A, et al. Technical and Biological Issues Relevant to Cell Typing by Aptamers. J Proteome Res. 2009 doi: 10.1021/pr801048z. [DOI] [PubMed] [Google Scholar]
- 38.Hicke BJ, Stephens AW. Escort aptamers: a delivery service for diagnosis and therapy. J Clin Invest. 2000;106:923–8. doi: 10.1172/JCI11324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Bagalkot V, Zhang L, Levy-Nissenbaum E, Jon S, Kantoff PW, Langer R, et al. Quantum dot-aptamer conjugates for synchronous cancer imaging, therapy, and sensing of drug delivery based on bi-fluorescence resonance energy transfer. Nano Lett. 2007;7:3065–70. doi: 10.1021/nl071546n. [DOI] [PubMed] [Google Scholar]
- 40.Gu F, Zhang L, Teply BA, Mann N, Wang A, Radovic-Moreno AF, et al. Precise engineering of targeted nanoparticles by using self-assembled biointegrated block copolymers. Proc Natl Acad Sci U S A. 2008;105:2586–91. doi: 10.1073/pnas.0711714105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zhang L, Radovic-Moreno AF, Alexis F, Gu FX, Basto PA, Bagalkot V, et al. Co-delivery of hydrophobic and hydrophilic drugs from nanoparticle-aptamer bioconjugates. ChemMedChem. 2007;2:1268–71. doi: 10.1002/cmdc.200700121. [DOI] [PubMed] [Google Scholar]
- 42.McNamara JO, 2nd, Andrechek ER, Wang Y, Viles KD, Rempel RE, Gilboa E, et al. Cell type-specific delivery of siRNAs with aptamer-siRNA chimeras. Nat Biotechnol. 2006;24:1005–15. doi: 10.1038/nbt1223. [DOI] [PubMed] [Google Scholar]
- 43.Chu TC, Twu KY, Ellington AD, Levy M. Aptamer mediated siRNA delivery. Nucleic Acids Res. 2006;34:e73. doi: 10.1093/nar/gkl388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wullner U, Neef I, Eller A, Kleines M, Tur MK, Barth S. Cell-specific induction of apoptosis by rationally designed bivalent aptamer-siRNA transcripts silencing eukaryotic elongation factor 2. Curr Cancer Drug Targets. 2008;8:554–65. doi: 10.2174/156800908786241078. [DOI] [PubMed] [Google Scholar]
- 45.Song E, Zhu P, Lee SK, Chowdhury D, Kussman S, Dykxhoorn DM, et al. Antibody mediated in vivo delivery of small interfering RNAs via cell-surface receptors. Nat Biotechnol. 2005;23:709–17. doi: 10.1038/nbt1101. [DOI] [PubMed] [Google Scholar]
- 46.Dey AK, Khati M, Tang M, Wyatt R, Lea SM, James W. An aptamer that neutralizes R5 strains of human immunodeficiency virus type 1 blocks gp120-CCR5 interaction. J Virol. 2005;79:13806–10. doi: 10.1128/JVI.79.21.13806-13810.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhou J, Li H, Li S, Zaia J, Rossi JJ. Novel Dual Inhibitory Function Aptamer-siRNA Delivery System for HIV-1 Therapy. Mol Ther. 2008 doi: 10.1038/mt.2008.92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chu TC, Marks JW, 3rd, Lavery LA, Faulkner S, Rosenblum MG, Ellington AD, et al. Aptamer:toxin conjugates that specifically target prostate tumor cells. Cancer Res. 2006;66:5989–92. doi: 10.1158/0008-5472.CAN-05-4583. [DOI] [PubMed] [Google Scholar]
- 49.Chen CH, Dellamaggiore KR, Ouellette CP, Sedano CD, Lizadjohry M, Chernis GA, et al. Aptamer-based endocytosis of a lysosomal enzyme. Proc Natl Acad Sci U S A. 2008;105:15908–13. doi: 10.1073/pnas.0808360105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Huang YF, Shangguan D, Liu H, Phillips JA, Zhang X, Chen Y, et al. Molecular Assembly of an Aptamer-Drug Conjugate for Targeted Drug Delivery to Tumor Cells. Chembiochem. 2009;10:862–8. doi: 10.1002/cbic.200800805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Arap W, Pasqualini R, Ruoslahti E. Cancer treatment by targeted drug delivery to tumor vasculature in a mouse model. Science. 1998;279:377–80. doi: 10.1126/science.279.5349.377. [DOI] [PubMed] [Google Scholar]
- 52.Trepel M, Pasqualini R, Arap W. Chapter 4 Screening Phage-Display Peptide Libraries for Vascular Targeted Peptides. Methods Enzymol. 2008;445C:83–106. doi: 10.1016/S0076-6879(08)03004-8. [DOI] [PubMed] [Google Scholar]
- 53.Dollins CM, Nair S, Boczkowski D, Lee J, Layzer JM, Gilboa E, et al. Assembling OX40 aptamers on a molecular scaffold to create a receptor-activating aptamer. Chem Biol. 2008;15:675–82. doi: 10.1016/j.chembiol.2008.05.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Cox JC, Hayhurst A, Hesselberth J, Bayer TS, Georgiou G, Ellington AD. Automated selection of aptamers against protein targets translated in vitro: from gene to aptamer. Nucleic Acids Res. 2002;30:e108. doi: 10.1093/nar/gnf107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Mendonsa SD, Bowser MT. In vitro selection of high-affinity DNA ligands for human IgE using capillary electrophoresis. Anal Chem. 2004;76:5387–92. doi: 10.1021/ac049857v. [DOI] [PubMed] [Google Scholar]
- 56.Tok JB, Fischer NO. Single microbead SELEX for efficient ssDNA aptamer generation against botulinum neurotoxin. Chem Commun (Camb) 2008:1883–5. doi: 10.1039/b717936g. [DOI] [PubMed] [Google Scholar]
- 57.Lou X, Qian J, Xiao Y, Viel L, Gerdon AE, Lagally ET, et al. Micromagnetic selection of aptamers in microfluidic channels. Proc Natl Acad Sci U S A. 2009;106:2989–94. doi: 10.1073/pnas.0813135106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Farokhzad OC, Cheng J, Teply BA, Sherifi I, Jon S, Kantoff PW, et al. Targeted nanoparticle-aptamer bioconjugates for cancer chemotherapy in vivo. Proc Natl Acad Sci U S A. 2006;103:6315–20. doi: 10.1073/pnas.0601755103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Farokhzad OC, Jon S, Khademhosseini A, Tran TN, Lavan DA, Langer R. Nanoparticle-aptamer bioconjugates: a new approach for targeting prostate cancer cells. Cancer Res. 2004;64:7668–72. doi: 10.1158/0008-5472.CAN-04-2550. [DOI] [PubMed] [Google Scholar]