Abstract
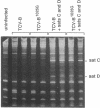
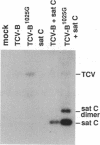
The virulent satellite [satellite C (sat C)] of turnip crinkle virus (TCV) is a small pathogenic RNA that intensifies symptoms in TCV-infected turnip plants (Brassica campestris). The virulence of sat C is determined by properties of the satellite itself and is influenced by the helper virus. Symptoms produced in infections with sat C differ in severity depending on the helper virus. The TCV-JI helper virus produces more severe symptoms than the TCV-B helper virus when inoculated with sat C. To find determinants in the TCV helper virus genome that affect satellite virulence, the TCV-JI genome was cloned and the sequence compared to the TCV-B genome. The genomes were found to differ by only five base changes, and only one of the base changes, at nucleotide position 1025, produced an amino acid change, an aspartic acid----glycine in the putative viral replicase. A chimeric TCV genome (TCV-B/JI) containing four of the five base changes (including the base change at position 1025) and a mutant TCV-B genome (TCV-B1025G) containing a single base substitution at position 1025 converted the TCV-B genome into a form that produces severe symptoms with sat C. The base change a position 1025 is located in the helicase of the putative viral replicase, and symptom intensification appears to result from differences in the rate of replication of the satellite supported by the two helper viruses.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Carrington J. C., Heaton L. A., Zuidema D., Hillman B. I., Morris T. J. The genome structure of turnip crinkle virus. Virology. 1989 May;170(1):219–226. doi: 10.1016/0042-6822(89)90369-3. [DOI] [PubMed] [Google Scholar]
- Collmer C. W., Stenzler L., Fay N., Howell S. H. Nonmutant forms of the avirulent satellite D of turnip crinkle virus are produced following inoculation of plants with mutant forms synthesized in vitro. Virology. 1991 Jul;183(1):251–259. doi: 10.1016/0042-6822(91)90137-z. [DOI] [PubMed] [Google Scholar]
- Habili N., Symons R. H. Evolutionary relationship between luteoviruses and other RNA plant viruses based on sequence motifs in their putative RNA polymerases and nucleic acid helicases. Nucleic Acids Res. 1989 Dec 11;17(23):9543–9555. doi: 10.1093/nar/17.23.9543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heaton L. A., Carrington J. C., Morris T. J. Turnip crinkle virus infection from RNA synthesized in vitro. Virology. 1989 May;170(1):214–218. doi: 10.1016/0042-6822(89)90368-1. [DOI] [PubMed] [Google Scholar]
- Jones R. W., Jackson A. O., Morris T. J. Defective-interfering RNAs and elevated temperatures inhibit replication of tomato bushy stunt virus in inoculated protoplasts. Virology. 1990 Jun;176(2):539–545. doi: 10.1016/0042-6822(90)90024-l. [DOI] [PubMed] [Google Scholar]
- Kaper J. M., Tousignant M. E., Geletka L. M. Cucumber-mosaic-virus-associated RNA-5. XII. Symptom-modulating effect is codetermined by the helper virus satellite replication support function. Res Virol. 1990 Sep-Oct;141(5):487–503. doi: 10.1016/0923-2516(90)90082-t. [DOI] [PubMed] [Google Scholar]
- Laín S., Riechmann J. L., García J. A. RNA helicase: a novel activity associated with a protein encoded by a positive strand RNA virus. Nucleic Acids Res. 1990 Dec 11;18(23):7003–7006. doi: 10.1093/nar/18.23.7003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X. H., Heaton L. A., Morris T. J., Simon A. E. Turnip crinkle virus defective interfering RNAs intensify viral symptoms and are generated de novo. Proc Natl Acad Sci U S A. 1989 Dec;86(23):9173–9177. doi: 10.1073/pnas.86.23.9173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mossop D. W., Francki R. I. Survival of a satellite RNA in vivo and its dependence on cucumber mosaic virus for replication. Virology. 1978 May 15;86(2):562–566. doi: 10.1016/0042-6822(78)90095-8. [DOI] [PubMed] [Google Scholar]
- Palukaitis P. Pathogenicity regulation by satellite RNAs of cucumber mosaic virus: minor nucleotide sequence changes alter host responses. Mol Plant Microbe Interact. 1988 Apr;1(4):175–181. doi: 10.1094/mpmi-1-175. [DOI] [PubMed] [Google Scholar]
- Petty I. T. A plasmid vector for cloning directly at the transcription initiation site of a bacteriophage T7 promoter. Nucleic Acids Res. 1988 Sep 12;16(17):8738–8738. doi: 10.1093/nar/16.17.8738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simon A. E., Engel H., Johnson R. P., Howell S. H. Identification of regions affecting virulence, RNA processing and infectivity in the virulent satellite of turnip crinkle virus. EMBO J. 1988 Sep;7(9):2645–2651. doi: 10.1002/j.1460-2075.1988.tb03117.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simon A. E., Howell S. H. The virulent satellite RNA of turnip crinkle virus has a major domain homologous to the 3' end of the helper virus genome. EMBO J. 1986 Dec 20;5(13):3423–3428. doi: 10.1002/j.1460-2075.1986.tb04664.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterworth H. E., Kaper J. M., Tousignant M. E. CARNA 5, the Small Cucumber Mosaic Virus--Dependent Replicating RNA, Regulates Disease Expression. Science. 1979 May 25;204(4395):845–847. doi: 10.1126/science.204.4395.845. [DOI] [PubMed] [Google Scholar]