Abstract
Laboratory strains of mice, both conventional and genetically engineered, have been introduced as critical components of a broad range of studies investigating normal and disease biology. Currently, the genetic identity of laboratory mice is primarily confirmed by surveying polymorphisms in selected sets of “conventional” genes and/or microsatellites in the absence of a single completely sequenced mouse genome. First, we examined variations in the genomic landscapes of transposable repetitive elements, named the TREome, in conventional and genetically engineered mouse strains using murine leukemia virus-type endogenous retroviruses (MLV-ERVs) as a probe. A survey of the genomes from 56 conventional strains revealed strain-specific TREome landscapes, and certain families (e.g., C57BL) of strains were discernible with defined patterns. Interestingly, the TREome landscapes of C3H/HeJ (toll-like receptor-4 [TLR4] mutant) inbred mice were different from its control C3H/HeOuJ (TLR4 wild-type) strain. In addition, a CD14 knock-out strain had a distinct TREome landscape compared to its control/backcross C57BL/6J strain. Second, an examination of superantigen (SAg, a “TREome gene”) coding sequences of mouse mammary tumor virus-type ERVs in the genomes of the 46 conventional strains revealed a high diversity, suggesting a potential role of SAgs in strain-specific immune phenotypes. The findings from this study indicate that unexplored and intricate genomic variations exist in laboratory mouse strains, both conventional and genetically engineered. The TREome-based high-resolution genetics surveillance system for laboratory mice would contribute to efficient study design with quality control and accurate data interpretation. This genetics system can be easily adapted to other species ranging from plants to humans.
Keywords: Transposable repetitive elements (TREome), TREome landscape, TREome gene profile, Genome complexity, Genetic surveillance
1. Introduction
For more than a century, a variety of laboratory mouse strains have been identified, developed, and created all around the world. Some laboratory mouse strains (“conventional”) were developed through defined breeding schemes to stabilize, isolate, and/or acquire specific traits and the others (“genetically engineered”) were created by molecular and genetic manipulation of embryos, primarily to study functions of “conventional” genes (Austin et al., 2004; Rossant, 2013).
The majority of the conventional laboratory mouse strains, which are critical components of modern biomedical research efforts, were developed as inbred strains via initial cross-breeding followed by a well-defined set of contiguous brother–sister matings. In general, biomedical researchers accept the notion that homozygosity at all genomic loci in inbred strains is established by up to 40 sequential sibling matings (Green, 1981; Taft et al., 2006). However, it is not uncommon to witness visible phenotypic variations in a mouse population of a single inbred strain which are maintained at a single institution, not to mention multiple institutions (Phelan and Austad, 1994; Weichman and Chaillet, 1997, personal communication). Interestingly, a recent report that the genomic configurations of C57BL/6J inbred mice are altered temporally and spatially, resulting in numerous variant genomes in an individual mouse, contradicts the given properties of genetic homozygosity and/or uniformity within a population of an inbred strain (Lee et al., 2015). It is possible that the dynamic nature of the genomes, which has not been fully explored or understood, is responsible, at least in part, for the establishment of enough genetic heterozygosity sufficient for developing variable phenotypes in an inbred strain.
The genetically engineered mouse strains are mainly categorized into two types: 1) transgenic strain in which an unknown number of copies of an exogenous gene is introduced during embryonic development and 2) knock-out strain in which a specific gene locus, occasionally including unintended targets, is partially manipulated using a variety of protocols to disturb its presumed coding potentials (Li et al., 2013; Thomas and Capecchi, 1987). Similar to the conventional laboratory mouse strains, incoherent, often visible, phenotypes are manifested within a population of a transgenic or knock-out strain (Doetschman, 2009; Montagutelli, 2000; Sigmund, 2000). This phenotypic inconsistency among the population of a genetically engineered strain might be linked to dissimilarity outside of the target locus or loci which are not thoroughly interrogated during genotyping processes. In fact, not all genetically engineered strains are subjected to a defined series of brother–sister matings to establish a recombinant inbred strain (Eisener-Dorman et al., 2009). It is recommended that if the backcross strain is an inbred, the specific inbred strain or its congenic strain serves as a control for the genetically engineered strain (Lambert, 2007). As such, it would be logical to question the genetic uniformity of the areas outside the target locus or loci between a genetically engineered strain and its inbred backcross-control strain.
The genetic integrity of conventional inbred as well as genetically engineered mouse strains is primarily monitored by examining: 1) pathophysiologic characteristics (e.g., coat color, tumor, susceptibility, lifespan) and 2) molecular profiles of the genome focusing on specific marker sets (Davisson, 1990; Fox et al., 1997; Silver, 1995). In particular, the molecular monitoring protocols typically survey for single nucleotide polymorphisms (SNPs) of conventional genes, microsatellite variations, and engineered mutant gene loci (Petkov et al., 2004; http://jaxmice.jax.org/genetichealth/monitoring). In consideration of the absence of a single complete mouse genome sequence in any genome database and the estimated size of the mouse genome of over 2.5 Gb, the information gathered from a survey of the small sets of gene SNPs and microsatellites may be insufficient for confirming the genetic integrity and uniformity of any mouse strain, either conventional or genetically engineered.
Exons of conventional genes consist of a fraction (~ 1.2%) of the mouse and human genomes, and the vast majority of the residual genome is occupied by a plethora of repetitive elements (REs) which include transposable repetitive elements (TREs), named the TREome (Mouse Genome Sequencing Consortium, 2002; Lander et al., 2001). TREs have the potential to dynamically shape the genomic landscape of an individual through “copy and paste” functions. Accumulation of acquired TRE activity in an individual's genomes due to both acute and chronic stress signals from the environment (e.g., onset of puberty, injury, infection) could conceivably be one genetic mechanism shaping the uncharacterized variations in a population of mice (Bohne et al., 2008; Cho et al., 2008b; Lee et al., 2012). As such, polymorphisms of conventional genes and a minute and biased set of microsatellites may play somewhat limited roles in defining the variable phenotypes of an inbred strain. Uncharacterized and/or unexplored genetic elements (e.g., TREs) and their variations, which are inherently and adaptively embedded in the individual mouse's genomes, are likely to contribute to the processes determining specific and dynamic phenotypes.
Our recent finding of highly polymorphic TRE profiles in the mouse population is inconsistent with the current assumption of high genome identity with regard to their conventional gene sequences (Frazer et al., 2007; Kao et al., 2012; Kirby et al., 2010; Lee et al., 2011; Mekada et al., 2009). In addition, recent findings of age- and organ-specific variations in genomic configurations due to TRE activity in C57BL/6J inbred mice contradicts the perceived genetic staticity and uniformity of a given inbred mouse population (Lee et al., 2012). The genetic integrity, stability, and/or uniformity of laboratory mouse strains are typically confirmed by a set of polymorphism data (e.g., morphophysiologic phenotypes, conventional genes, microsatellites). From a logical and efficient research perspective, the current systems for monitoring mouse genetics need to be re-evaluated to account for the uncharacterized properties and effects of the inherent diversity and acquired activity of TREs and other explored elements, on a dynamic genome platform.
2. Materials and methods
2.1. Animal experiments
The following mouse strains were purchased from the Jackson Laboratory: female C57BL/6J, C3H/HeJ, C3H/HeOuJ, and CD14 knock-out (B6.129S4-Cd14tm/frm/J). All animals were provided with water and food ad libitum at a University of California Davis facility and some of the mice were aged for a period of time. The experimental protocol was approved by the Animal Use and Care Administrative Advisory Committee of the University of California Davis. Animals were sacrificed to collect tissues followed by snap-freezing in liquid nitrogen.
2.2. Genomic DNA of various laboratory mouse strains
Snap-frozen tissue samples were subjected to genomic DNA isolation using a DNeasy Tissue kit (Qiagen, Valencia, CA) and the DNA samples were normalized to 20 ng/μl. In addition, genomic DNA from 63 laboratory mouse strains, which include nine 129 substrains, were purchased from the Jackson Laboratory (Bar Harbor, ME). In addition, genomic DNA from a C57BL/6J × 129S1/SvImJF2/J (B6129SF2/J) mouse, a F2 hybrid from F1 × F1 whose parents were C57BL/6J (female) and 129S1/SvImJ (male), was obtained from the Jackson Laboratory. According to the information from the Jackson Laboratory's website, the genomic DNA was isolated from either the brain or spleen of respective mouse strains (http://jaxmice.jax.org/jaxnotes/archive/499b.html). Gender identity of each DNA sample was confirmed by amplifying a region specific for mouse chromosome Y by PCR using a pair of primers (Supplementary Table 1) followed by agarose gel electrophoresis.
2.3. Polymorphism analysis of genomic TREome (murine leukemia virus-type endogenous retrovirus [MLV-ERV]) long terminal repeats (LTRs)
The polymorphic regions of the MLV-ERV LTRs were identified from the genomic DNAs of 12 laboratory mouse strains (Jackson Laboratory) by PCR using a set of primer pairs (Supplementary Table 1) which were designed from a well-conserved region. Following ligation into a TA vector (Promega, Madison, WI), 24 colonies were picked from the MLV-ERV amplicons of each strain and plasmid DNAs were prepared using a QIAprep Spin Miniprep kit (Qiagen) before sequencing (Molecular Cloning Laboratories, South San Francisco, CA). A set of unique MLV-ERV LTR sequences was compiled for each mouse strain by multiple alignment analysis using the Vector NTI program (Invitrogen, Carlsbad, CA). Within a set of unique MLV-ERV LTR sequences for each mouse strain, the occurrence frequency of 256 four-nucleotide “word” (a nucleotide sequence of specific length) combinations at all four possible reading frames were counted using a program developed in our laboratory (unpublished). Within each strain, the occurrence frequency data for the individual words were normalized and converted into probability distribution function (PDF) values. For each word, the average and standard deviation of the PDF values from all 12 strains were calculated using Excel (Microsoft, Redmond, WA). Based on an assumption that the higher the standard deviation in a word, the more variation in the word, the extent of variations in each four-nucleotide word within the 12 strain-population was visualized with a schedule of gray shades (white-lowest variation; black-highest variation) on a 16 × 16 (= 256) matrix. To examine/simulate diversity in conventional gene sequences in comparison to the MLV-ERV LTR sequences, the single nucleotide polymorphism (SNP) data for the GAPDH gene (~4.7 kb) among 19 laboratory mouse strains (A/J, C57BL/6J, 129X1/SvJ, AKR/J, BALB/cByJ, C3H/HeJ, CAST/EiJ, DBA/2J, FVB/NJ, MOLF/EiJ, NOD/ShiLtJ, SM/J, BTBR T+Itpr3tf/J, KK/HlJ, LG/J, NZW/LacJ, PWD/PhJ, WSB/EiJ, and 129S1/SvImJ; http://www.informatics.jax.org/javawi2/servlet/WIFetch?page=snpQF) was subjected to the same PDF analysis as above.
2.4. TREome landscaping of mouse genomes using MLV-ERV sequences as a probe
Genomic DNA (20 ng) was cut with Nco-I (New England Biolab, Ipswich, MA) at 37 °C for 4 h followed by self-ligation of the cut fragments using T4 ligase (Promega) overnight at 4 °C. The TREome landscape data was collected by I-PCR amplification of the junctions spanning putative MLV-ERV integration loci using 2 μl of the ligation products, Taq polymerase (Qiagen), and a pair of inverse primers designed from the conserved MLV-ERV sequences. The primer sequences and PCR conditions are listed in Supplementary Table 1. I-PCR amplicons were resolved in a 7.5% polyacrylamide gel for visualization.
2.5. Polymorphism analysis of genomic TREome (mouse mammary tumor virus-type endogenous retrovirus [MMTV-ERV]) superantigen (SAg) genes
The MMTV-ERV SAg coding sequences were PCR amplified from the genomic DNA (57 mouse strains) obtained from the Jackson Laboratory using a set of primers (Supplementary Table 1). Following cloning of the SAg amplicons using a pGEM-T Easy kit from Promega, plasmid DNAs were prepared for 12 colonies picked from each strain using a QIAprep Spin Miniprep kit and sequenced (Molecular Cloning Laboratories). Eleven mouse strains had no visible SAg coding sequences amplified (C57L/J, CASA/Rk, CAST/EiJ, CZECHII/EiJ, Mus caroli/EiJ, Mus Pahari/Ei, PANCEVO/Ei, PERA/EiJ, PERC/Ei, SKIVE/Ei, and TIRANO/Ei). Within each mouse strain, following identification of a set of unique MMTV-LTR sequences by multiple alignment analyses using Vector NTI (Invitrogen), MMTV-ERVs' SAg open reading frames were examined and translated in silico. Polymorphisms in the putative SAg proteins were visualized using a function in the Excel program (Microsoft).
3. Results
3.1. Diversity in TREome (MLV-ERV) profiles among 12 laboratory mouse strains
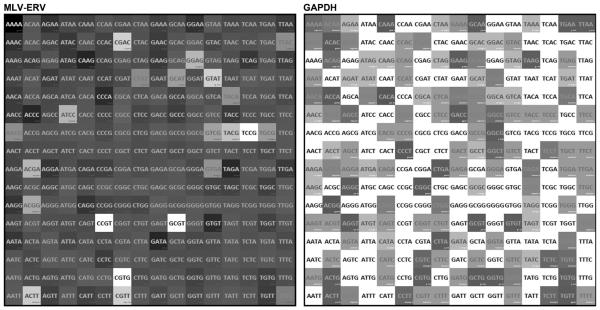
Similarity among the genomes of a range of laboratory mouse strains has been examined primarily based on SNP polymorphism (Frazer et al., 2007; Kirby et al., 2010; Mekada et al., 2009; http://mus.well.ox.ac.uk/mouse/INBREDS/). However, it needs to be noted that the genome similarity data was derived mostly from the sequences of conventional genes. To evaluate the extent of TREome diversity among different laboratory mouse strains in comparison to conventional genes, genomic profiles of MLV-ERVs, a mouse TREome family, were examined in 12 laboratory mouse strains (129P1/ReJ, 129X1/SvJ, A/HeJ, A/J, AKR/J, ALR/Lt, BALB/cJ, BDP/J, BPH/2J, BUB/BnJ, C3H/HeJ, and C3H/HeOuJ). The MLV-ERV LTR sequences were isolated from the genomic DNA of each strain and subjected to a probability distribution function analysis for the entire set of all possible four-nucleotide words in order to compute and visualize the variation levels of individual words on a 16 × 16 (=256) matrix. In contrast to the overall low variation matrix of GAPDH genes derived from 20 laboratory mouse strains including one reference, there were relatively high variations in the vast majority of the words from the MLV-ERV LTR sequences from the 12 inbred mouse strains (Fig. 1). Interestingly, about a dozen words were highly conserved among the pool of MLV-ERV LTR sequences from 12 mouse strains. Although the genome-wide survey for MLV-ERVs in this study was not comprehensive in nature, the findings from this study provide some evidence that the laboratory mouse population is highly diverse with regard to TREome/MLV-ERV profiles in their genomes in contrast to the relatively high similarity among conventional gene sequences (Frazer et al., 2007; Kirby et al., 2010). Presumably, there would be significant differences in TREome/MLV-ERV profiles between males and females since at least reference mouse chromosome Y is densely populated with both characterized and uncharacterized REs.
Fig. 1.
High-level diversity in MLV-ERV (TREome) LTR sequences among 12 laboratory mouse strains. The extent of variations among the population of MLV-ERV LTR sequences from the genomic DNA of 12 mouse strains were examined by a probability distribution function analysis for 4-nucleotide word sets. The extent of variations for the individual words were visualized on a 16 × 16 (= 256) matrix. In contrast to the overall low variation matrix of GAPDH genes (a conventional gene) from 20 laboratory mouse strains (right panel), the vast majority of the words from the MLV-ERV sequences derived from 12 mouse strains (left panel) had a high-level of variability. Variability of each word within individual sets (12 [MLV-ERV LTR] strains or 20 [GAPDH gene] strains) is coded on a gray scale, ranging from white (low = 0) to black (high = 0.001261).
3.2. Polymorphic TREome landscapes among 56 laboratory mouse strains
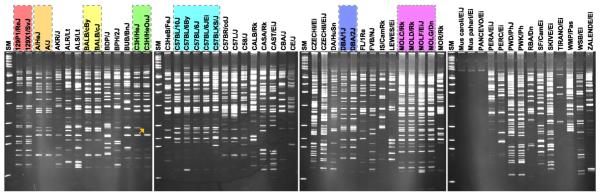
To further study genomic diversity with regard to TREome profiles among laboratory mouse strains, TREome landscapes were visualized from the genomic DNA of 56 laboratory mouse strains using MLV-ERVs as a probe. At first glance, none of the 56 strains share the same TREome landscape patterns (Fig. 2). However, it was apparent that distinct TREome landscape patterns were shared within certain individual families of strains, such as 129PI-XI/, A/, BALB/, C3H/, C57BL/, DBA/, and MOL, which presumably reflect their common TREome/MLV-ERV ancestries. Interestingly, the TREome landscapes of the C3H/HeJ and its toll-like receptor 4 (TLR4) wildtype control strain (C3H/HeOuJ) were different from each other (http://jaxmice.jax.org/jaxnotes/archive/430e.html; http://www.informatics.jax.org/reference/allele/MGI: 2386635). In addition, the C57BLKS/J strain's TREome landscape pattern was substantially different from the four other C57BL strains examined. We confirmed that these two sets of strains were the same gender (data not shown), excluding the possibility of chromosome Y effects on the TREome landscape patterns. Furthermore, the TREome landscape patterns of three strains (Mus caroli/Ei, Mus Pahari/Ei, and PANCEVO/Ei) had only a couple of visible amplicon bands while the other 53 strains displayed numerous bands. This finding may coincide with previous reports in which these three strains are placed upstream of the evolutionary pathway of mice compared to the other strains examined in this study (Boursot et al., 1993; Ideraabdullah et al., 2004; Suzuki et al., 2004).
Fig. 2.
Polymorphic TREome landscapes among the genomes of 56 laboratory mouse strains. Using MLV-ERVs as a probe, TREome landscapes of the genomes of 56 laboratory mouse strains were visualized. None of the 56 mouse strains share the same TREome landscape patterns. Distinct TREome landscape patterns were found within certain families of strains (highlighted with dotted lines and colors), such as 129PI-XI/, A/, BALB/, C3H/, C57BL/, DBA/, and MOL. The TREome landscapes of the C3H/HeJ and C3H/HeOuJ strains were different (one of the different bands is indicated with an arrow). In addition, three strains (Mus caroli/Ei, Mus Pahari/Ei, and PANCEVO/Ei) had only a couple of visible bands. SM (size marker).
3.3. Dissimilar TREome landscapes between the C3H/HeJ strain and its TLR4 wildtype control, C3H/HeOuJ
To confirm the initial finding (Fig. 2) that the C3H/HeJ strain (TLR4−/−) is different from its wildtype control strain, C3H/HeOuJ (TLR4+/+), with respect to TREome landscape patterns, we repeated the experiment using six different organs of two additional mice (12-week old females) from each strain. The pair of C3H/HeJ and C3H/HeOuJ strains has been employed widely for the studies focusing on the roles of the TLR4 gene in innate immune functions (Beutler, 2000; Beutler et al., 2001; Poltorak et al., 1998; Smirnova et al., 2000). From all six organs of both strains examined, one distinct TREome band was found only in C3H/HeJ mice whereas another band was only present in C3H/HeOuJ mice (Fig. 3). This difference in banding pattern is consistent with the initial finding described earlier (Fig. 2). In addition, within each mouse, regardless of the strain, there were substantial variations in TREome landscapes among the different organs. The findings from this experiment confirm that in addition to the difference in the TLR4 locus, the genomes of the C3H/HeJ and C3H/HeOuJ strains are different in their TREome landscapes. It is reasonable to assume that application of additional landscaping probes would reveal more TREome loci which are uniquely embedded in the genomes of either the C3H/HeJ or C3H/HeOuJ strain.
Fig. 3.
Un-identical TREome landscapes between the C3H/HeJ strain and its wildtype control, C3H/HeOuJ. With respect to the TREome landscape patterns, the C3H/HeJ strain (TLR4−/−) is markedly different from its presumed wildtype control, C3H/HeOuJ (TLR4+/+), in the genomes of all six organs (kidney [KI], liver [LI], lung [LU], lymph node [LN], spleen [SP], and thymus [TH]) examined. In particular, one distinct TREome band (enlarged in a separate window) was found only in the C3H/HeJ strain while another band was present only in the C3H/HeOuJ strain. SM (size marker).
3.4. Un-identical TREome landscapes between the CD14 knock-out strain (CD14−/−) and its backcross-control, C57BL/6J (CD14+/+) strain
Genetically engineered mouse strains (transgenic or knock-out for a specific target gene) have served as critical and popular components of modern biomedical research efforts (Fox et al., 2006; Houdebine, 2007; Pearson et al., 2008). Typically, the inbred mouse strain, which is introduced during the backcrossing process of generating a genetically engineered strain, is chosen to control the modified/mutated target gene (http://research.jax.org/grs/type/gemm/; Seong et al., 2004). In this study, we examined whether there are distinct variations in the TREome landscapes between the genomes of a pair of CD14 knock-out (12-week old female) and its backcross-control (C57BL/6J; 12-week old female) strains (Haziot et al., 1996; Poltorak et al., 1998). Within the genomes of all six organs from each strain, two unique TREome/MLV-ERV amplicon bands were visible only in the CD14 knock-out strain, while two other TREome/MLV-ERV amplicon bands were found only in C57BL/6J backcross-control strain (Fig. 4). Similar to the findings from the C3H/HeJ-C3H/HeOuJ pair, the TREome landscapes were variable depending on organs within each strain. The findings from this study indicate that the TREome landscapes of the CD14 knock-out strain are visibly discernible from its backcross-control, C57BL/6J strain, in addition to the difference at the genetically targeted locus and its flanking region on chromosome 18 (Cho et al., 2008a; Yee et al., 2008).
Fig. 4.
Apparent variations in TREome landscapes between the CD14 knock-out strain and its backcross-control, C57BL/6J strain. There were apparent differences in the TREome landscapes between the genomes of CD14 knock-out and its backcross-control (C57BL/6J) strains in all six organs (kidney [KI], liver [LI], lung [LU], lymph node [LN], spleen [SP], and thymus [TH]) examined. Two unique TREome bands were visible only in the CD14 knock-out strain, whereas two other bands were found only in the C57BL/6J control strain. SM (size marker).
3.5. Variations in TREome landscapes in 129 mouse substrains
For genetic engineering of transgenic and knock-out mouse models, such as the CD14 knock-out strain described above, embryos of various 129 mouse substrains have been extensively used as a target for the initial manipulation of the genome (Threadgill et al., 1997). Substantial levels of genetic variations among the 129 mouse substrains were reported to be linked to either accidental or intentional outcrossing(s) (Simpson et al., 1997). In this study, genomic DNA from nine 129 substrains (129S1/SvImJ, 129/Sv-Lyntm1Sor/J, 129S1/Sv-Oca2+ Tyr+ KitlSl-J/J, 129S4/SvJae-Inhbbtm1Jae/J, 129S4/SvJae-Pparatm1Gonz/J, 129S4/SvJaeSor-Gt(ROSA)26Sortm1(FLP1)Dym/J, 129S6/SvEv-Mostm1Ev/J, 129P1/ReJ, and 129X1/SvJ) were examined to survey variations in the TREome landscapes using an MLV-ERV probe. Overall, all nine 129 substrains shared a common TREome landscape pattern (Fig. 5). However, both of the 129/Sv-Lyntm1Sor/J and the 129P1/ReJ substrains lacked a unique band in its TREome landscape which is present in the other seven strains. It is interesting to note that the 129S1/SvImJ substrain has a distinct MLV-ERV amplicon band in comparison to the 129/Sv-Lyntm1Sor/J substrain, for which it often serves as a control. A comprehensive survey of the entire collection of 129 substrains, which employs additional landscaping probes allowing for high-resolution identification of substrain-specific TREome banding patterns, would provide valuable information for phenotypic interrogation of 129-derived genetically engineered strains.
Fig. 5.
TREome landscapes of 129 mouse substrains. Genomic DNAs of nine 129 substrains (129S1/SvImJ [A], 129/Sv-Lyntm1Sor/J [B], 129S1/Sv-Oca2+ Tyr+ KitlSl-J/J [C], 129S4/SvJae-Inhbbtm1Jae/J [D], 129S4/SvJae-Pparatm1Gonz/J [E], 129S4/SvJaeSor-Gt(ROSA)26Sortm1(FLP1)Dym/J [F], 129S6/SvEv-Mostm1Ev/J [G], 129P1/ReJ [H], and 129X1/SvJ [I]) were examined for variations in TREome landscapes. Overall, all substrains shared a similar TREome landscape pattern; however, both of the 129/Sv-Lyntm1Sor/J and the 129P1/ReJ substrains lacked a unique band in its TREome landscape compared to the other seven strains. SM (size marker).
3.6. TREome landscape-based surveillance of genome-crossing between two mouse strains
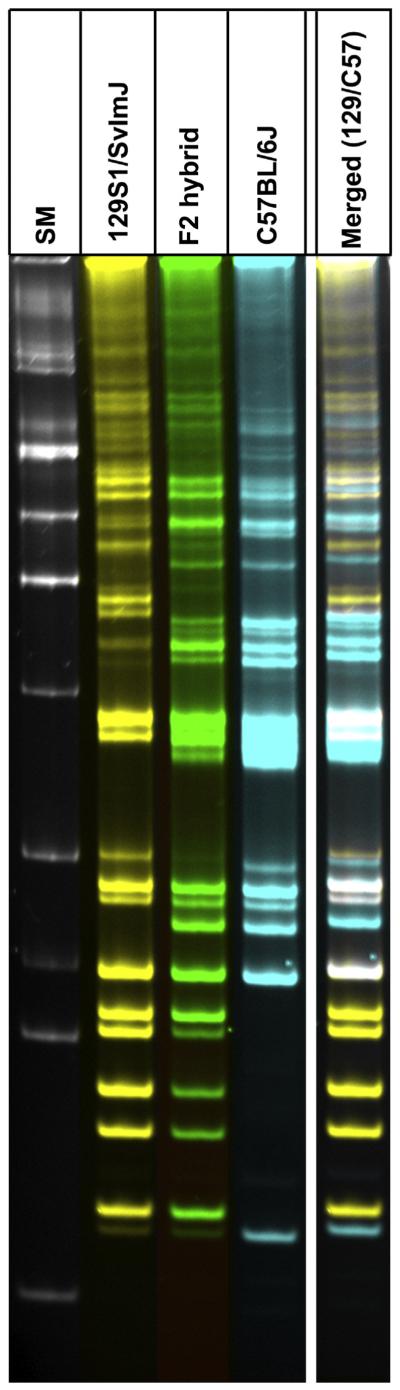
It has been a common practice to backcross chimera mice, which were derived from genetically targeted embryonic genomes of a 129 substrain, with the C57BL/6J strain to establish a stable strain (Hedrich, 2004; Threadgill et al., 1997). In this study, we examined whether the genome-crossing events between two mouse strains (C57BL/6J × 129S1/SvImJ) are reflected in the TREome landscapes of the hybrid offspring. With regard to the TREome landscape, an F2 hybrid mouse (male), which was derived from an initial crossing of C57BL/6J (female) and 129S1/SvImJ (male), was compared to a C57BL/6J mouse (male) and a 129S1/SvImJ mouse (male). Although for the most part, the pattern of the F2 hybrid TREome landscape displayed the bands from both C57BL/6J and 129S1/SvImJ genomes, it lacked certain bands which were specific only for the individual parental strains (Fig. 6). This incompletely merged hybrid banding pattern within the TREome landscape of the F2 hybrid provides insightful visual information which reveals the static status of genetic crossing between two mouse strains. The F2 hybrid's TREome landscape may be closely linked to the chromosomal cross-over events. High-resolution details of the TREome landscapes can be accomplished by implementing an optimal set of probes derived from MLV-ERVs as well as MMTV-ERVs, LINES, and other TREs/REs. The findings from this study introduce a novel idea that by using the high-resolution TREome landscaping technology, during the courses of backcrossing and/or crossing of mouse strains, the genomes of the offsprings at a series of successive generations could be monitored for their “crossing configuration” and uniformity.
Fig. 6.
TREome landscape-based monitoring of genome-crossing between two mouse strains: an example. We examined whether the genome-crossing events between two mouse strains (C57BL/6J x 129S1/SvImJ) are reflected in the TREome landscapes of a hybrid offspring. For the most part, the pattern of the F2 hybrid's TREome landscape included the bands from both the C57BL/6J and 129S1/SvImJ genomes; however, certain bands specific for the individual parental strains were not present. This F2 hybrid's TREome banding pattern provides valuable information which visualizes the genomic status of crossing events between two strains. SM (size marker).
3.7. Polymorphisms in a “TREome gene” (MMTV-ERV SAg) among laboratory mouse strains
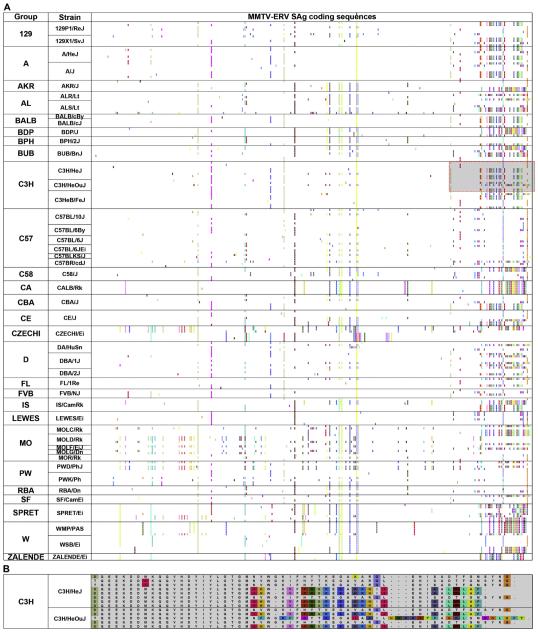
Much of the focus of recent advancements in the genomics and bioinformatics fields hinges on the notion that sequence polymorphisms, including small RNAs, and relevant functions of conventional genes are responsible for phenotypic variations in both normal and disease biology (Mardis, 2008; Mu and Zhang, 2012). To evaluate whether polymorphisms in “TREome genes” cast potential impacts on variable phenotypes among the mouse population, we examined sequence diversity in MMTV-ERV SAg genes, a well-studied immune-regulatory TREome gene (Lee et al., 2011; Peters et al., 1983), among 57 laboratory mouse strains. Only 46 of the 57 mouse strains yielded visible MMTV-ERV LTR amplicons which are presumed to harbor the SAg gene open reading frame (ORF). A high-level of SAg gene polymorphism was indicated by the finding that at least one (up to eight) unique SAg coding sequence was identified within the genomes of the 46 individual mouse strains (Fig. 7A). Altogether, 183 isoforms of the MMTV-ERV SAg gene were derived from the 46 mouse strains. As expected, the C-terminus regions of the SAg ORFs, which are reported to confer the Vβ-specificity of B lymphocytes during interactions with T lymphocytes (Acha-Orbea and MacDonald, 1995), had the highest variations in conjunction with a few polymorphic clusters in the other regions. Interestingly, the genomic profiles of the MMTV-ERV SAg ORFs were different between the C3H/HeJ inbred strain and its TLR4 wildtype control strain, C3H/HeOuJ; eight and five unique putative SAg isoforms were identified in the genomes of C3H/HeJ and C3H/HeOuJ strains, respectively (Fig. 7B). A well-defined and comprehensive survey would be necessary to confirm the extent of temporal and spatial variations in the MMTV-ERV SAg ORFs which exist in the genomes of these two strains. Polymorphisms in the MMTV-ERV SAg, a TREome gene, may differentially shape the immune profiles (e.g., negatively selected T lymphocytes) of the C3H/HeJ and C3H/HeOuJ as well as other laboratory mouse strains (Tomonari et al., 1993). It is uncertain how the potentially differential systemic immune profiles between the C3H/HeJ and C3H/HeOuJ strains, due to the MMTV-ERV SAg polymorphisms, would be networked with the function of TLR4 with regard to its role in innate immunity.
Fig. 7.
Highly polymorphic TREome gene (MMTV-ERV SAg) isoforms among 46 laboratory mouse strains. Polymorphisms in the MMTV-ERV SAg gene coding regions among 46 laboratory mouse strains were examined. A. A high-level of MMTV-ERV SAg gene polymorphism was apparent among the genomes of the 46 mouse strains; altogether, 183 MMTV-ERV SAg gene isoforms were identified. In addition, the C-terminus regions of the MMTV-ERV SAg gene coding sequences had relatively high levels of variations compared to the other regions. B. MMTV-ERV SAg isoform profiles were different between the C3H/HeJ inbred strain (eight unique isoforms) and its wildtype TLR4 control strain, C3H/HeOuJ (five unique isoforms).
4. Discussion
Currently, the normal and disease biology of laboratory mice and humans is explained primarily in the context of the function and polymorphism of conventional genes. Thus far, the majority of conventional gene-based attempts to decode the tangible mechanisms of normal and disease states and to identify diagnostic markers have been inconclusive or unsuccessful (Padyukov, 2013; Seok et al., 2013; Takao and Miyakawa, 2015a; Takao and Miyakawa, 2015b). Reportedly, laboratory mice and humans share ~ 80% of conventional gene sequences (Mouse Genome Sequencing Consortium, 2002; Guenet, 2005); this is inconsistent with the notion that phenotypic details are primarily determined by conventional genes when dramatic phenotypic distances exist between the two species. Considering the limited understanding of the complete genome information system of mice and humans, conventional gene-focused approaches would be insufficient for decoding the enormous scope of phenotypic details and their variations. The inherent TREome diversity of individual laboratory mouse strains is directly associated with the polymorphic protein coding potentials for TREome genes. Whereas the acquired activity of the inherently diverse TREome may play a role in fine-tuning the function of TREome genes as well as conventional genes, which reside near the new TREome positions, through their networks of transcription regulatory elements, contributing to strain-specific phenotypic details (Amid et al., 2009; Giardine et al., 2007).
The intricate and unexplored variations in the genomes of laboratory mouse strains are directly associated with two distinct, but interrelated, characteristics of the TREome. First, the information embedded in the TREome landscape of a mouse strain, which is defined by TRE information regarding type, copy number, and position, can be examined to understand how conventional gene(s) neighboring a genomic position of a specific TRE type is regulated. The finding from this study that TREome landscape patterns are different between the C3H/HeJ strain and its TLR4 wildtype control strain (C3H/HeOuJ) needs to be investigated further to determine whether the differences are linked to the expression of certain gene(s) other than TLR4 (Kamath et al., 2003). Similarly, confirmation of the impact of differences in the TREome landscapes between the CD14 knock-out and its backcross-control strain (C57BL/6J) on the expression of genes outside of the knock-out locus is deemed to be necessary. It is likely that some other knock-out and/or transgenic mouse strains need to be subjected to similar scrutiny of their genomic configurations with regard to the TREome landscape, in comparison to their control strains. Second, there are numerous and highly polymorphic TREome genes in the genomes of laboratory mouse strains which have not been fully identified, accounted for, or understood. Tangible coding potentials of TREome genes could be either presumed full-length or variable in length due to introduction of mutations over time. It has been demonstrated that certain TREome genes, such as the envelope genes of MLV-ERVs and MMTV-ERV SAg genes, play functional roles in biological processes (Bentvelzen, 1992; Huber et al., 1994; Kotzin et al., 1993). In addition, TREome (human endogenous retroviruses) gene isoforms isolated from a human burn patient's genomic DNA demonstrated differential potentials for regulating inflammatory mediators, such as IL-6 and IL-1β (Lee et al., 2014). Despite the previous studies which reported a range of functionality of TREome genes in both mice and humans, unfortunately, they are often called non-coding long RNAs in current literature (Geisler and Coller, 2013; Gibb et al., 2015). In this study, polymorphisms in TREome genes in laboratory mouse strains are reflected in the identification of 183 isoforms of the MMTV-ERV SAg gene which was reported to play a critical role in shaping the systemic immune cell profile (Acha-Orbea and MacDonald, 1995; Kotzin et al., 1993; Tomonari et al., 1993). The differences in the profile of MMTV-ERV SAg gene isoforms between the C3H/HeJ strain and its TLR4 control (C3H/HeOuJ) suggest that a specific immune system would be developed within each strain due to the activity of a unique set of MMTV-ERV SAg genes, especially during T lymphocyte selection events. In order to confirm the data obtained from the TLR4 studies using C3H/HeJ and its control (C3H/HeOuJ) strains, the potential impacts of the differential MMTV-ERV SAg activities on immune function should be examined.
Despite the absence of a single reference mouse genome, which is completely sequenced, it is often stated that the population of laboratory mouse strains share a high level of genome sequences (Frazer et al., 2007; Kirby et al., 2010; Mekada et al., 2009; http://mus.well.ox.ac.uk/mouse/INBREDS/). In addition, a recent change in the putative size of the NCBI's reference mouse chromosome Y from ~ 16 Mb (Build 37.2) to ~ 92 Mb (Annotation Release 105) suggests that more time is needed to confirm the size of each chromosome within individual laboratory mouse strains (Lee et al., 2013). In spite of the lack of the full sequence information from a single reference strain, current genetic monitoring systems for laboratory mice rely primarily on polymorphism data derived from limited sets of conventional genes and microsatellites to determine the genetic uniformity/status/identity of a specific strain/substrain. The finding that the TREome (MLV-ERVs) sequences are more variable among 12 laboratory mouse strains, in comparison to conventional gene sequences, suggests that the TREome landscape contributes to the formation of unique phenotypic characteristics embedded in each strain. Furthermore, the discrepancy in TREome landscapes between the CD14 knock-out and its backcross-control strain (C57BL/6J) informs that all genetically engineered mouse strains may need to be examined to confirm the genetic uniformity with their matching controls, outside of the individual targeted loci. On the other hand, the unexplained/unexpected phenotypic variations, which are frequently encountered in genetically engineered mouse strains such as runt or normal weight STAT-1−/− mice (Bona and Revillard, 2001; Kim et al., 2003), could be explained by checking the genomic configuration of the engineered mouse population. In certain circumstances, confirmation of uniformity within the entire genomes (minus targeted loci) may be necessary to validate the results collected from studies involving genetically engineered mouse strains.
To the scientific community, we propose a collaborative project, which focuses on developing a high-resolution mouse genetics surveillance system (MGSS) using the platform of TREome landscapes and TREome genes in conjunction with the current conventional gene-based monitoring systems. Genetic uniformity of established laboratory mouse strains, both conventional and genetically engineered, could be evaluated by the high-resolution MGSS for initial confirmation and maintenance of each strain. Depending on the MGSS data collected, additional breeding designs and/or surveillance protocols may be implemented to obtain acceptable levels of genetic uniformity for each strain. When new mouse strains are developed, the MGSS serves as a high-resolution tool for dynamic monitoring of genetic crossing/backcrossing. The newly developed genetics surveillance system (MGSS) would be ready for application into different species (e.g., humans, animals, plants) and fields (e.g., agriculture, forensics). We believe that identifying and accounting for variations in the TREome landscapes and TREome genes are crucial in establishing a genetically uniform laboratory mouse strain as well as decoding normal and disease biology in the up-and-coming fields of precision biology and medicine.
Supplementary Material
Acknowledgments
This study was supported, in part, by grants from the Shriners of North America (No. 86800 to KC, No. 84302 to KHL [postdoctoral fellowship]) and the National Institutes of Health (R01 GM071360 to KC).
Footnotes
Supplementary data to this article can be found online at http://dx.doi.org/10.1016/j.yexmp.2016.01.005.
References
- Acha-Orbea H, MacDonald HR. Superantigens of mouse mammary tumor virus. Annu. Rev. Immunol. 1995;13:459–486. doi: 10.1146/annurev.iy.13.040195.002331. [DOI] [PubMed] [Google Scholar]
- Amid C, et al. Manual annotation and analysis of the defensin gene cluster in the C57BL/6J mouse reference genome. BMC Genomics. 2009;10:606. doi: 10.1186/1471-2164-10-606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Austin CP, et al. The knockout mouse project. Nat. Genet. 2004;36:921–924. doi: 10.1038/ng0904-921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentvelzen P. Immunosuppression by the MMTV superantigen? Immunol. Today. 1992;13:77. doi: 10.1016/0167-5699(92)90139-x. [DOI] [PubMed] [Google Scholar]
- Beutler B. Tlr4: central component of the sole mammalian LPS sensor. Curr. Opin. Immunol. 2000;12:20–26. doi: 10.1016/s0952-7915(99)00046-1. [DOI] [PubMed] [Google Scholar]
- Beutler B, et al. Identification of toll-like receptor 4 (Tlr4) as the sole conduit for LPS signal transduction: genetic and evolutionary studies. J. Endotoxin Res. 2001;7:277–280. [PubMed] [Google Scholar]
- Bohne A, et al. Transposable elements as drivers of genomic and biological diversity in vertebrates. Chromosom. Res. 2008;16:203–215. doi: 10.1007/s10577-007-1202-6. [DOI] [PubMed] [Google Scholar]
- Bona CA, Revillard J-P. Cytokines and Cytokine Receptors: Physiology and Pathological Disorders. CRC Press; 2001. [Google Scholar]
- Boursot P, et al. The evolution of house mice. Annu. Rev. Ecol. Syst. 1993:119–152. [Google Scholar]
- Cho K, et al. Cosegregation of CD14 locus and polymorphic alleles of glucocorticoid receptor and protocadherins into CD14 knockout mouse genome. Shock. 2008a;29:724–732. doi: 10.1097/shk.0b013e31815d0b80. [DOI] [PubMed] [Google Scholar]
- Cho K, et al. Endogenous retroviruses in systemic response to stress signals. Shock. 2008b;30:105–116. doi: 10.1097/SHK.0b013e31816a363f. [DOI] [PubMed] [Google Scholar]
- Davisson M. The Jackson Laboratory Mouse Mutant Resource. 1990 [Google Scholar]
- Doetschman T. Influence of genetic background on genetically engineered mouse phenotypes. Methods Mol. Biol. 2009;530:423–433. doi: 10.1007/978-1-59745-471-1_23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisener-Dorman AF, et al. Cautionary insights on knockout mouse studies: the gene or not the gene? Brain Behav. Immun. 2009;23:318–324. doi: 10.1016/j.bbi.2008.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fox RR, et al. Handbook on Genetically Standardized JAX Mice. Jackson Laboratory; 1997. [Google Scholar]
- Fox JG, et al. The Mouse in Biomedical Research: Normative Biology, Husbandry, and Models. Academic Press; 2006. [Google Scholar]
- Frazer KA, et al. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature. 2007;448:1050–1053. doi: 10.1038/nature06067. [DOI] [PubMed] [Google Scholar]
- Geisler S, Coller J. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat. Rev. Mol. Cell Biol. 2013;14:699–712. doi: 10.1038/nrm3679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giardine B, et al. PhenCode: connecting ENCODE data with mutations and phenotype. Hum. Mutat. 2007;28:554–562. doi: 10.1002/humu.20484. [DOI] [PubMed] [Google Scholar]
- Gibb EA, et al. Activation of an endogenous retrovirus-associated long non-coding RNA in human adenocarcinoma. Genome Med. 2015;7:22. doi: 10.1186/s13073-015-0142-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green EL. Genetics and Probability in Animal Breeding Experiments. Macmillan Publishers Ltd; 1981. [Google Scholar]
- Guenet JL. The mouse genome. Genome Res. 2005;15:1729–1740. doi: 10.1101/gr.3728305. [DOI] [PubMed] [Google Scholar]
- Haziot A, et al. Resistance to endotoxin shock and reduced dissemination of Gramnegative bacteria in CD14-deficient mice. Immunity. 1996;4:407–414. doi: 10.1016/s1074-7613(00)80254-x. [DOI] [PubMed] [Google Scholar]
- Hedrich H. The Laboratory Mouse. Academic Press; 2004. [Google Scholar]
- Houdebine L-M. Target Discovery and Validation Reviews and Protocols. Springer; 2007. Transgenic Animal Models in Biomedical Research; pp. 163–202. [DOI] [PubMed] [Google Scholar]
- Huber BT, et al. The role of superantigens in the immunobiology of retroviruses. CIBA Found. Symp. 1994;187:132–140. doi: 10.1002/9780470514672.ch9. discussion 140-3. [DOI] [PubMed] [Google Scholar]
- Ideraabdullah FY, et al. Genetic and haplotype diversity among wild-derived mouse inbred strains. Genome Res. 2004;14:1880–1887. doi: 10.1101/gr.2519704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamath AB, et al. Toll-like receptor 4-defective C3H/HeJ mice are not more susceptible than other C3H substrains to infection with Mycobacterium tuberculosis. Infect. Immun. 2003;71:4112–4118. doi: 10.1128/IAI.71.7.4112-4118.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kao D, et al. ERE database: a database of genomic maps and biological properties of endogenous retroviral elements in the C57BL/6J mouse genome. Genomics. 2012;100:157–161. doi: 10.1016/j.ygeno.2012.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S, et al. Stat1 functions as a cytoplasmic attenuator of Runx2 in the transcriptional program of osteoblast differentiation. Genes Dev. 2003;17:1979–1991. doi: 10.1101/gad.1119303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirby A, et al. Fine mapping in 94 inbred mouse strains using a high-density haplotype resource. Genetics. 2010;185:1081–1095. doi: 10.1534/genetics.110.115014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotzin BL, et al. Superantigens and their potential role in human disease. Adv. Immunol. 1993;54:99–166. doi: 10.1016/s0065-2776(08)60534-9. [DOI] [PubMed] [Google Scholar]
- Lambert R. A Jackson Laboratory Resource Manual. The Jackson Laboratory; 2007. Breeding strategies for maintaining colonies of laboratory mice. [Google Scholar]
- Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- Lee YK, et al. Prevalent de novo somatic mutations in superantigen genes of mouse mammary tumor viruses in the genome of C57BL/6J mice and its potential implication in the immune system. BMC Immunol. 2011;12:5. doi: 10.1186/1471-2172-12-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee KH, et al. Age-dependent and tissue-specific structural changes in the C57BL/6J mouse genome. Exp. Mol. Pathol. 2012;93:167–172. doi: 10.1016/j.yexmp.2012.04.013. [DOI] [PubMed] [Google Scholar]
- Lee KH, et al. Large interrelated clusters of repetitive elements (REs) and RE arrays predominantly represent reference mouse chromosome Y. Chromosom. Res. 2013;21:15–26. doi: 10.1007/s10577-012-9334-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee KH, et al. Divergent and dynamic activity of endogenous retroviruses in burn patients and their inflammatory potential. Exp. Mol. Pathol. 2014;96:178–187. doi: 10.1016/j.yexmp.2014.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee KH, et al. Temporal and spatial rearrangements of a repetitive element array on C57BL/6J mouse genome. Exp. Mol. Pathol. 2015;98:439–445. doi: 10.1016/j.yexmp.2015.03.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D, et al. Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nat. Biotechnol. 2013;31:681–683. doi: 10.1038/nbt.2661. [DOI] [PubMed] [Google Scholar]
- Mardis ER. The impact of next-generation sequencing technology on genetics. Trends Genet. 2008;24:133–141. doi: 10.1016/j.tig.2007.12.007. [DOI] [PubMed] [Google Scholar]
- Mekada K, et al. Genetic differences among C57BL/6 substrains. Exp. Anim. 2009;58:141–149. doi: 10.1538/expanim.58.141. [DOI] [PubMed] [Google Scholar]
- Montagutelli X. Effect of the genetic background on the phenotype of mouse mutations. J. Am. Soc. Nephrol. 2000;11(Suppl. 16):S101–S105. [PubMed] [Google Scholar]
- Mouse Genome Sequencing Consortium Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420:520–562. doi: 10.1038/nature01262. [DOI] [PubMed] [Google Scholar]
- Mu W, Zhang W. Bioinformatic resources of microRNA sequences, gene targets, and genetic variation. Front. Genet. 2012;3:31. doi: 10.3389/fgene.2012.00031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Padyukov L. Between the Lines of Genetic Code: Genetic Interactions in Understanding Disease and Complex Phenotypes. Academic Press; 2013. [Google Scholar]
- Pearson T, et al. Humanized Mice. Springer; 2008. Humanized SCID Mouse Models for Biomedical Research; pp. 25–51. [DOI] [PubMed] [Google Scholar]
- Peters G, et al. Tumorigenesis by mouse mammary tumor virus: evidence for a common region for provirus integration in mammary tumors. Cell. 1983;33:369–377. doi: 10.1016/0092-8674(83)90418-x. [DOI] [PubMed] [Google Scholar]
- Petkov PM, et al. Development of a SNP genotyping panel for genetic monitoring of the laboratory mouse. Genomics. 2004;83:902–911. doi: 10.1016/j.ygeno.2003.11.007. [DOI] [PubMed] [Google Scholar]
- Phelan JP, Austad SN. Selecting animal models of human aging: inbred strains often exhibit less biological uniformity than F1 hybrids. J. Gerontol. 1994;49:B1–B11. doi: 10.1093/geronj/49.1.b1. [DOI] [PubMed] [Google Scholar]
- Poltorak A, et al. Defective LPS signaling in C3H/HeJ and C57BL/10ScCr mice: mutations in Tlr4 gene. Science. 1998;282:2085–2088. doi: 10.1126/science.282.5396.2085. [DOI] [PubMed] [Google Scholar]
- Rossant J. Making a knockout mouse: from stem cells to embryos. Nat. Cell Biol. 2013;15:1133. doi: 10.1038/ncb2850. [DOI] [PubMed] [Google Scholar]
- Seok J, et al. Genomic responses in mouse models poorly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. U. S. A. 2013;110:3507–3512. doi: 10.1073/pnas.1222878110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seong E, et al. To knockout in 129 or in C57BL/6: that is the question. Trends Genet. 2004;20:59–62. doi: 10.1016/j.tig.2003.12.006. [DOI] [PubMed] [Google Scholar]
- Sigmund CD. Viewpoint: are studies in genetically altered mice out of control? Arterioscler. Thromb. Vasc. Biol. 2000;20:1425–1429. doi: 10.1161/01.atv.20.6.1425. [DOI] [PubMed] [Google Scholar]
- Silver LM. Mouse Genetics: Concepts and Applications. Oxford University Press; 1995. [Google Scholar]
- Simpson EM, et al. Genetic variation among 129 substrains and its importance for targeted mutagenesis in mice. Nat. Genet. 1997;16:19–27. doi: 10.1038/ng0597-19. [DOI] [PubMed] [Google Scholar]
- Smirnova I, et al. Phylogenetic variation and polymorphism at the toll-like receptor 4 locus (TLR4) Genome Biol. 2000;1:1–002.10. doi: 10.1186/gb-2000-1-1-research002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki H, et al. Temporal, spatial, and ecological modes of evolution of Eurasian Mus based on mitochondrial and nuclear gene sequences. Mol. Phylogenet. Evol. 2004;33:626–646. doi: 10.1016/j.ympev.2004.08.003. [DOI] [PubMed] [Google Scholar]
- Taft RA, et al. Know thy mouse. Trends Genet. 2006;22:649–653. doi: 10.1016/j.tig.2006.09.010. [DOI] [PubMed] [Google Scholar]
- Takao K, Miyakawa T. Correction for Takao and Miyakawa, Genomic responses in mouse models greatly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. U. S. A. 2015a;112:E1163–E1167. doi: 10.1073/pnas.1502188112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takao K, Miyakawa T. Genomic responses in mouse models greatly mimic human inflammatory diseases. Proc. Natl. Acad. Sci. U. S. A. 2015b;112:1167–1172. doi: 10.1073/pnas.1401965111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas KR, Capecchi MR. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell. 1987;51:503–512. doi: 10.1016/0092-8674(87)90646-5. [DOI] [PubMed] [Google Scholar]
- Threadgill DW, et al. Genealogy of the 129 inbred strains: 129/SvJ is a contaminated inbred strain. Mamm. Genome. 1997;8:390–393. doi: 10.1007/s003359900453. [DOI] [PubMed] [Google Scholar]
- Tomonari K, et al. Influence of viral superantigens on V beta- and V alpha-specific positive and negative selection. Immunol. Rev. 1993;131:131–168. doi: 10.1111/j.1600-065x.1993.tb01534.x. [DOI] [PubMed] [Google Scholar]
- Weichman K, Chaillet JR. Phenotypic variation in a genetically identical population of mice. Mol. Cell. Biol. 1997;17:5269–5274. doi: 10.1128/mcb.17.9.5269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yee KS, et al. The effect of CAG repeat length polymorphism in the murine glucocorticoid receptor on transactivation potential. Exp. Mol. Pathol. 2008;84:200–205. doi: 10.1016/j.yexmp.2008.04.003. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.