Abstract
Sequencing of exomes and genomes has revealed abundant genetic variation affecting the coding sequences of human transcription factors (TFs), but the consequences of such variation remain largely unexplored. We developed a computational, structure-based approach to evaluate TF variants for their impact on DNA-binding activity and used universal protein binding microarrays to assay sequence-specific DNA-binding activity across 41 reference and 117 variant alleles found in individuals of diverse ancestries and families with Mendelian diseases. We found 77 variants in 28 genes that affect DNA-binding affinity or specificity and identified thousands of rare alleles likely to alter the DNA-binding activity of human sequence-specific TFs. Our results suggest that most individuals have unique repertoires of TF DNA-binding activities, which may contribute to phenotypic variation.
Exome sequencing studies have identified many nonsynonymous single nucleotide polymorphisms (nsSNPs) in transcription factors (TFs) (1). Genetic variants that alter transcript expression levels have been associated with human disease risk and are widespread in human populations (2, 3). Numerous Mendelian diseases are attributable to mutations in TFs (4). Missense SNPs that change the amino acid sequence of TF DNA binding domains (DBDs) might disrupt their DNA binding activities and thus have detrimental consequences on their gene regulatory functions. Despite their medical importance, the consequences of coding variation in DBDs on TF function have remained largely unexplored.
We identified 53,384 unique DBD polymorphisms (DBDPs) (Table S1) (here, defined as missense variants) in a curated, high-confidence set of 1,254 sequence-specific human TFs (5, 6) (Table S2) from genotype data for 64,706 individuals encompassing African, Asian and European ancestries (Fig. 1A) (1, 2, 7). We also identified 4,552 unique nonsense mutations that result in partial or full DBD truncation (Table S3).
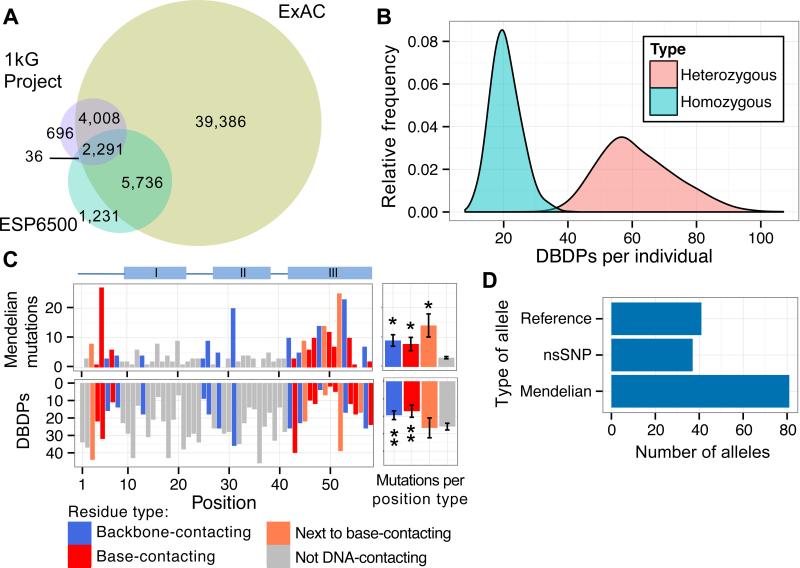
Figure 1. Evaluation of coding variation in TF DBDs.
(A) Number of unique DBDPs in 1kG (Phase 3), ESP 6500, or ExAC v0.2 individuals. (B) Histograms of unique DBDPs per individual in either homozygous or heterozygous states. (C) Number of Mendelian mutations, and nsSNPs found in ExAC, across all homeodomain TFs annotated by their position and type of DNA contact associated with each position. “I”, “II, “III” refer to α-helices; III is the DNA-recognition helix. Adjacent bar graphs depict mean number of variants for each position type; enrichment (*) or depletion (**) relative to non-DNA-contacting residues (P < 0.05, permutation test), error bars = 1 standard error of the mean, N = 332 Mendelian mutations, 1,300 nsSNPs, and 11 base-contacting, 12 phosphate-backbone-contacting, 5 neighboring-DNA-contacting, and 30 non-DNA-contacting positions. (D) Allele types assayed by PBMs.
We found a median of 60 heterozygous and 20 homozygous DBDPs (Fig. 1B) per genome. We found a significant depletion (odds ratio = 3.7, P = 0.005, Fisher's exact test) of DBDPs among TFs with known Mendelian disease mutations (6, 8), suggesting that DBDPs in disease-associated TFs have phenotypic consequences.
We developed a computational approach (6) to evaluate missense substitutions in TF DBDs for their impact on DNA-binding activity. Existing methods for predicting the impact of missense mutations (9, 10) do not adequately consider the roles of residues in protein-DNA interactions, which we reasoned should improve predictions. We first focused on homeodomain DBDs, as most known Mendelian disease mutations in TFs occur in homeodomain proteins. We analyzed homeodomain-DNA co-crystal structures in the Protein Data Bank to assemble a composite protein-DNA ‘contact map’ (Fig. S1). As anticipated, residues that contact DNA bases or phosphate backbone, or that immediately neighbor base-contacting residues, are enriched among Mendelian disease mutations (P < 0.005, permutation test). In contrast, individuals in the population are depleted for variants at base- or backbone- contacting positions (P = 0.0134 or 0.0312, respectively, permutation test) (Fig. 1C). This highlights the value of considering protein-DNA contacts in predicting the impact of variants.
Based on these results, we expanded our approach to other TF families. For each variant we considered multiple criteria, including: (a) position of the residue relative to the protein-DNA interface in homologous co-crystal structures (Fig. S1); (b) DNA-binding specificity-determining residues for particular DBD classes (Fig. S2); (c) scores from tools that predict mutation pathogenicity (9, 10); (d) minor allele frequencies; and (e) phenotypic associations from genome-wide association studies (11) or known Mendelian disease mutations (8).
Using these criteria, we selected 36 TF DBDPs (6) to assay for direct, sequence-specific DNA-binding activity (Fig. S3). These DBDPs were obtained from 1000 Genomes Project (1kG) Phase 2, the Exome Sequencing Project (ESP 6500), and the Exome Aggregation Consortium (ExAC). To calibrate the effects of these nsSNPs, we selected 81 Mendelian disease mutations, which are known or believed to be pathogenic (Fig. 1D) (8, 12). The 117 variant DBD alleles span six major structural classes, representing 41 distinct TF allelic series (Fig. S4). We assayed these 158 DBD alleles using universal protein binding microarrays (PBMs) (6), on which each non-palindromic 8-bp sequence occurs on at least 32 spots (13) (Table S4).
We identified variant-induced changes in DNA-binding specificity (14) (Fig. 2A) or affinity (Fig. 2B) by comparing the enrichment (E) scores of each of 32,768 nonredundant, ungapped 8-mers represented on PBMs to those of the corresponding reference allele (6, 13). DNA-binding changes were reproducible across replicate PBM experiments and support previously reported DNA-binding affinity differences (Table S5, Fig. S5). We categorized all 117 variant alleles as having altered DNA-binding specificity, affinity, both, or neither (Table S6). Three nsSNPs completely abrogated sequence-specific DNA-binding (Fig. 2C, Fig. S6). In total, 77 variants altered DNA-binding affinity and/or specificity (Fig. 2D). Several nsSNPs predicted to be damaging but not scored here as having altered DNA-binding might cause subtle changes beyond the sensitivity of our approach or alternatively affect protein-protein interactions.
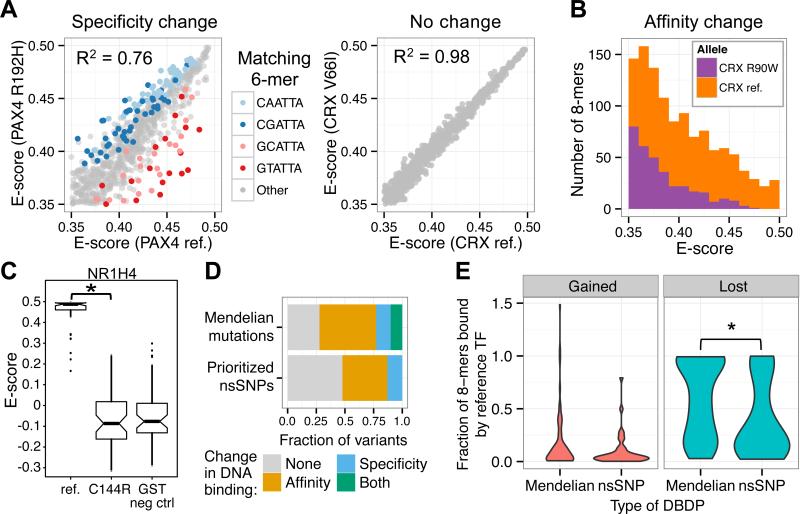
Figure 2. Perturbed DNA-binding caused by nsSNPs or Mendelian disease mutations.
(A) Specificity change in PAX4 R192H allele compared to no change in CRX V66I allele. Colored 6-mers are allele-preferred (Q < 0.05, intersection-union test with Benjamini-Hochberg correction). (B) Altered E-score distribution of CRX R90W allele relative to the reference allele indicates altered DNA binding affinity. (C) Box plots depict E-scores of NR1H4 reference and C144R alleles and GST negative controls (6) for the top 50 8-mers bound by NR1H4 reference allele. C144R abolished binding specificity (* P < 2.2 × 10−16, Wilcoxon rank-sum test), resulting in E-scores indistinguishable from GST negative controls (Table S7). (D) Fraction of alleles with observed changes in DNA-binding affinity, specificity, both, or neither as determined from PBM binding profiles. Prioritized nsSNPs exclude those predicted as benign by both PolyPhen-2 and SIFT. (E) Violin plots depicting fraction of 8-mer binding sites gained or lost by variants relative to the number of 8-mers bound by the reference allele. Gains or losses were defined as E ≥ 0.4 for one allele and E < 0.4 for the other allele. * P = 0.0044, Wilcoxon rank-sum test.
Compared to DBDPs, Mendelian disease mutants lost a larger fraction of 8-mers bound by the corresponding reference alleles (P = 0.0044, Wilcoxon rank-sum test), consistent with more extreme phenotypes being associated with more drastic in vitro binding changes. The overall difference in gained 8-mers was not significantly different between these two sets of variants (P = 0.32, Wilcoxon rank-sum test; Fig. 2E).
PBM binding profiles within an allelic series differed for variants associated with distinct disease phenotypes (Fig. S7), supporting results from a yeast one-hybrid screen of Mendelian disease TF mutants (15). They also provided molecular insights into the molecular basis of clinical heterogeneity of disease mutations affecting the same genes. For example, CRX is associated with Mendelian diseases of retinal degeneration (16). The R90W allele, associated with the severe disease Leber congenital amaurosis 7 (17), lost the ability to bind most 8-mers bound by wildtype CRX. In contrast, the R41W allele, associated with cone-rod dystrophy 2 (18), resulted in a moderate specificity change (Fig. S7B).
The 8-mer binding profiles of HOXD13 alleles displayed a range of effects; several of these alleles are associated with various limb malformations (19) (Fig. 3A). The I297V and N298S variants, predicted to be benign, did not alter DNA-binding activity. The Q325K and Q325R alleles gained recognition of novel motifs, consistent with those learned from chromatin immunoprecipitation with high-throughput sequencing (ChIP-Seq) data (12). Allele-preferred 8-mers (Fig. 3B, Fig. S8A) are enriched within ChIP-Seq peaks bound exclusively by the respective allele (Fig. 3C, Figs. S8B, S9) (P < 0.01, Wilcoxon signed-rank test). Putative target genes, associated with ChIP-Seq peaks enriched (P < 2.2 × 10−16, one-tailed Wilcoxon signed-rank test) for Q325K- or Q325R-preferred versus reference-preferred 8-mers (Fig. S10) (6), are over-represented among genes up-regulated by the corresponding allele (P < 0.01, permutation test) (Fig. 3D, Figs. S8C, S11), consistent with HOXD13 acting as a transcriptional activator (20). These results suggest that these variants’ changes in binding specificity alter genomic occupancy, leading to inappropriate gene expression through gained binding sites.
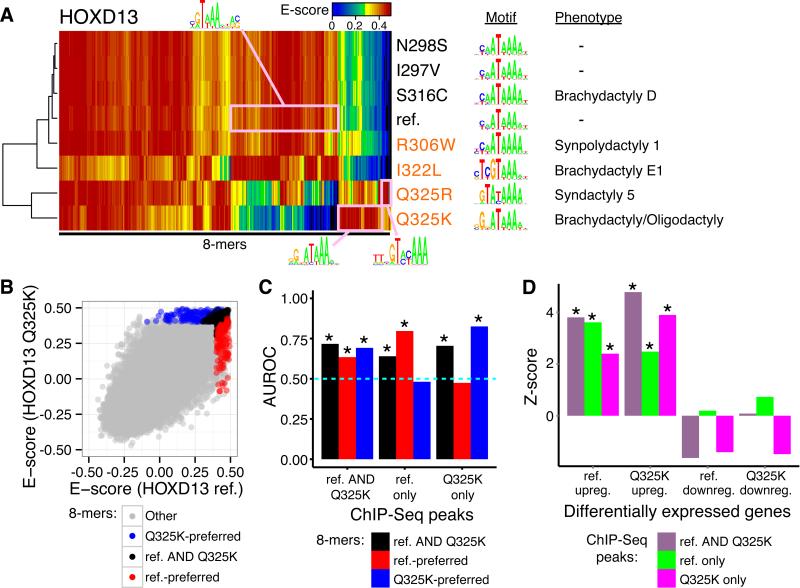
Figure 3. Perturbations in TF DNA-binding and gene expression associated with HOXD13 genetic variants.
(A) Heatmap depicting PBM E-scores of DBD alleles (rows) for all 8-mers (columns) bound strongly (E > 0.45) by at least one allele, with corresponding motifs (13) and phenotypes. Rows and columns were clustered hierarchically. Pink boxes highlight allele-preferred sequences with corresponding motifs, generated by alignment of the indicated 8-mers (14). Variants in orange font exhibited altered specificity. “–” indicates no known phenotype. (B) Scatter plot comparing 8-mer E-scores of HOXD13 reference versus Q325K alleles. Allele-preferred and allele-common 8-mers (6) are colored. (C) PBM-derived allele-preferred 8-mers are enriched (* P < 0.01, Wilcoxon signed-rank test) within genomic regions bound in vivo exclusively by the respective allele. Dashed horizontal line indicates AUROC = 0.5 (no enrichment or depletion). (D) Genes associated with ChIP-Seq peaks enriched for reference-preferred versus Q325K-preferred 8-mers are over-represented (* P < 0.01, permutation test) among genes up-regulated by the same allele. Z-scores were calculated using 100 random background gene sets (6).
As expected, mutations in residues that either contact DNA or neighbor a base-contacting residue were enriched (odds ratio = 4.3, P = 0.003, Fisher's exact test) among DBDPs with altered DNA binding affinity or specificity (Fig. 4A). Interestingly, we also found variants at non-DNA-contacting positions that altered DNA binding, potentially by affecting protein conformation or stability. We identified 3,833 unique missense variants that are predicted to be damaging by both Polyphen-2 and SIFT and occur at DNA-contacting residues (Fig. 4B). These values are likely an underestimate of damaged DBDPs across all human TFs (Fig. 4C). These damaging nsSNPs occur at lower frequencies in the ExAC population than nsSNPs for which no change in DNA-binding is predicted (P < 0.05, permutation test) (Fig. 4D), suggesting that they are more likely to be deleterious.
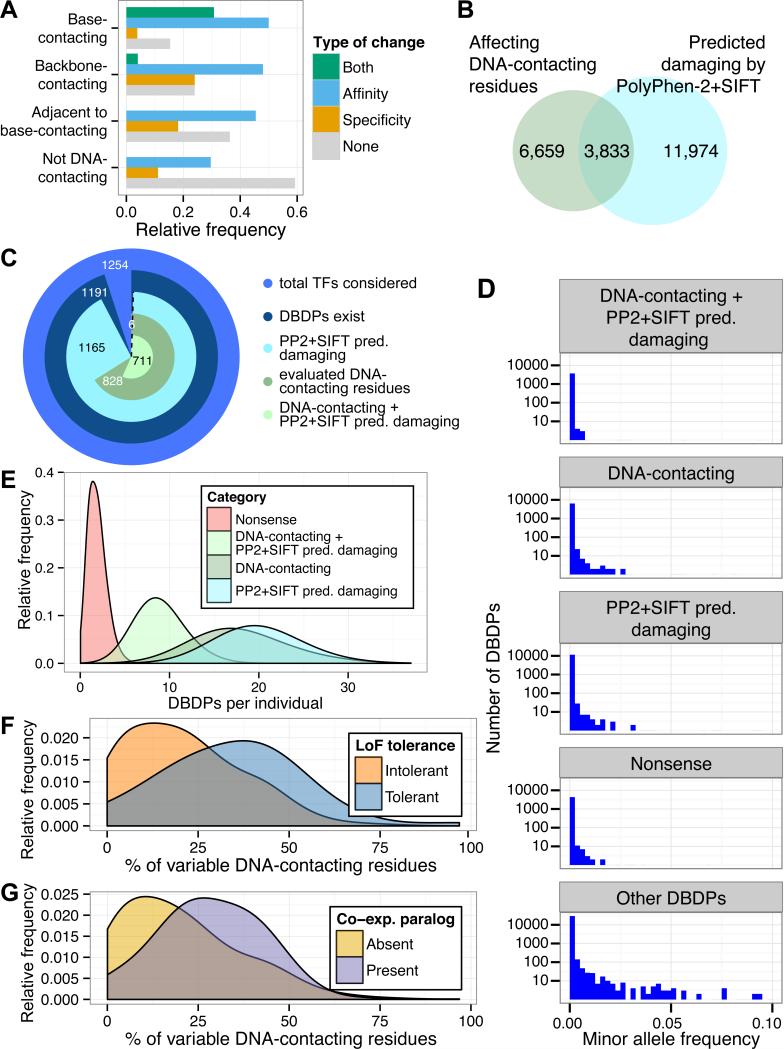
Figure 4. Properties of ExAC DBDPs predicted to alter DNA-binding activity.
(A) Relative frequency of DNA-binding changes observed for variants at DNA-contacting residues. “Both” comprises residues at which variants changed DNA binding affinity and specificity either simultaneously in one protein or separately across different proteins. (B) Overlap between DBDPs affecting DNA-contacting residues in zf-C2H2, Fork_head, HLH, and Homeobox Pfam domains (blue) or predicted as “probably damaging” by PolyPhen-2 and “damaging” by SIFT (green). (C) Number of sequence-specific TFs for which DBDPs were identified and their evaluation, as in (B). (D) Minor allele frequencies (ExAC v0.2) of nsSNPs. (E) Histogram of DBD variants per individual (1000 Genomes Project Phase 3), annotated as in C. (F) Fraction of DNA-contacting residues per TF altered by at least one nsSNP (ExAC), for genes tolerant of homozygous or compound heterozygous LoF mutations versus genes for which LoF-tolerance was not observed (21). (G) Fraction of variable DNA-contacting residues (ExAC) in TFs with versus without at least one co-expressed paralog.
Per individual, there were very few (median = 2) nonsense DBD variants, but a wide range in the number of putatively damaging missense variants (median = 9, DBDPs at DNA-contacting residues and predicted as damaging by Polyphen-2 and SIFT) (Fig. 4E, Fig. S12). Hence, we investigated what mechanisms might allow damaged DBDPs to be tolerated. TFs reported to tolerate homozygous LoF mutations in Icelanders (21) had a significantly higher fraction of DNA-contacting residues altered by our identified nsSNPs (P = 6.63 × 10−8, permutation test) (Fig. 4F). TFs with a co-expressed paralog (22) had a significantly higher fraction of variable DNA-contacting residues (P = 6.11 × 10−8, permutation test) (Fig. 4G); this enrichment was significant independent of LoF-tolerance status (P < 0.005, t-test) (6). Additional compensation could arise from epistasis with cis-regulatory variants (23). Damaged DBDPs might be associated with undiagnosed or subclinical phenotypes, variably penetrant phenotypes due to epistatic or gene-environment interactions, or phenotypes that present in later life.
Our results highlight the utility of PBM profiling to reveal changes in the DNA binding activities of DBD variants. PBM profiling of DBDPs identified through additional sequencing studies may elucidate disease pathologies by revealing alterations in DNA binding that result in transcriptional dysregulation.
Our analyses suggest that most unrelated individuals have a unique repertoire of TF alleles with a distinct landscape of DNA binding activities. Variants with subtle changes in DNA-binding activities may confer reduced deleteriousness and thus have greater potential for giving rise to phenotypic variation. Analysis of genetic interactions among TFs, TF variants, and noncoding regulatory variation likely will provide insights into the structure of genetic variation that leads to phenotypic differences among people.
Supplementary Material
One Sentence Summary.
Comprehensive analysis of Mendelian disease mutations and single nucleotide polymorphisms (SNPs) in human transcription factors reveals a continuum of alterations in DNA binding activity.
Acknowledgments
We thank Max Hume, Yu-Han Hsu, Yun Shen and Dawit Balcha for technical assistance, and Alexander Gimelbrant for helpful discussions. This work was supported by the National Institutes of Health (grants NHGRI R01 HG003985 to M.L.B. and T.H., and P50 HG004233 to M.V. and D.E.H.), an A*STAR National Science Scholarship to K.H.K., and National Science Foundation Graduate Research Fellowships to L.A.B. and J.M.R. TF PBM data have been deposited into UniPROBE (publication dataset accession BAR15A). GST negative control PBM 8-mer data are provided in Table S7. M.L.B. is a co-inventor on U.S. patents # 6,548,021 and #8,530,638 on PBM technology and corresponding universal sequence designs, respectively. Universal PBM array designs used in this study are available via a Materials Transfer Agreement with The Brigham & Women's Hospital, Inc. A.V., J.V.K., J.M.R., N.S., T.H., and S.Y. performed experiments, L.A.B., J.V.K., J.M.R., S.S.G., E.J.R., J.W., L.M., K.H.K., S.I., T.S., L.S., R.G., and C.C. performed data analysis, M.K., M.J.D., M.V., D.E.H., and M.L.B. supervised research, L.A.B., L.M., K.H.K, D.E.H., and M.L.B. designed the study and wrote the manuscript, L.A.B., J.V.K., J.M.R., S.S.G., L.M., K.H.K, S.I., and M.L.B. prepared figures and tables.
Footnotes
Supplementary Materials:
Materials and Methods
Figures S1-S12
Tables S1-S7
References (24-55)
References and Notes
- 1.Consortium EA. bioRxiv. 2015 [Google Scholar]
- 2.Abecasis GR, et al. Nature. 2010;467:1061–1073. doi: 10.1038/nature09534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Westra H-J, et al. Nat Genet. 2013;45:1238–1243. doi: 10.1038/ng.2756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Veraksa A, Del Campo M, McGinnis W. Mol Genet Metab. 2000;69:85–100. doi: 10.1006/mgme.2000.2963. [DOI] [PubMed] [Google Scholar]
- 5.Vaquerizas JM, Kummerfeld SK, Teichmann SA, Luscombe NM. Nat Rev Genet. 2009;10:252–263. doi: 10.1038/nrg2538. [DOI] [PubMed] [Google Scholar]
- 6.Materials and methods are available as supplementary materials on Science Online.
- 7.Fu W, et al. Nature. 2013;493:216–220. doi: 10.1038/nature11690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Consortium U. Nucleic Acids Res. 2015;43:D204–212. [Google Scholar]
- 9.Adzhubei IA, et al. Nat Meth. 2010;7:248–249. doi: 10.1038/nmeth0410-248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ng PC, Henikoff S. Nucleic Acids Res. 2003;31:3812–3814. doi: 10.1093/nar/gkg509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Welter D, et al. Nucleic Acids Res. 2014;42:D1001–1006. doi: 10.1093/nar/gkt1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ibrahim DM, et al. Genome Res. 2013;23:2091–2102. doi: 10.1101/gr.157610.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Berger MF, et al. Nat Biotech. 2006;24:1429–1435. doi: 10.1038/nbt1246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jiang B, Liu JS, Bulyk ML. Bioinformatics. 2013;29:1390–1398. doi: 10.1093/bioinformatics/btt152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fuxman Bass JI, et al. Cell. 2015;161:661–673. doi: 10.1016/j.cell.2015.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Freund CL, et al. Cell. 1997;91:543–553. doi: 10.1016/s0092-8674(00)80440-7. [DOI] [PubMed] [Google Scholar]
- 17.Swaroop A, et al. Hum. Mol. Genet. 1999;8:299–305. doi: 10.1093/hmg/8.2.299. [DOI] [PubMed] [Google Scholar]
- 18.Swain PK, et al. Neuron. 1997;19:1329–1336. doi: 10.1016/s0896-6273(00)80423-7. [DOI] [PubMed] [Google Scholar]
- 19.Brison N, Debeer P, Tylzanowski P. Dev Dyn. 2014;243:37–48. doi: 10.1002/dvdy.24037. [DOI] [PubMed] [Google Scholar]
- 20.Salsi V, Vigano MA, Cocchiarella F, Mantovani R, Zappavigna V. Dev Biol. 2008;317:497–507. doi: 10.1016/j.ydbio.2008.02.048. [DOI] [PubMed] [Google Scholar]
- 21.Sulem P, et al. Nat Genet. 2015;47:448–452. doi: 10.1038/ng.3243. [DOI] [PubMed] [Google Scholar]
- 22.Ouedraogo M, et al. PLoS One. 2012;7:e50653. doi: 10.1371/journal.pone.0050653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lappalainen T, Montgomery SB, Nica AC, Dermitzakis ET. Am J Hum Genet. 2011;89:459–463. doi: 10.1016/j.ajhg.2011.08.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.