Abstract
A method for designing the operating parameters (surface light intensity, operating temperature and agitation rate) was proposed for microalgal protein production. Furthermore, quadratic model was established and validated (R2 > 0.90) with experimental data. It was recorded that temperature and agitation rate were slightly interdependent. The microalgal protein performance could be estimated using the simulated experimental setup and procedure developed in this study. The results also showed a holistic approach for opening a new avenue on simulation design for microalgal protein optimization.
Keywords: design, microalgae, model, optimization, protein
Introduction
Microalgae are very diverse and represent a rich source of phytochemicals, which can be used in food, animal feed, aquaculture, cosmetics, pharmaceutical and bio-fuel industries.1,2 Most successful microalgal cultivations are limited to the production of high value products, high revenues of which offset the high capital and operational expenditures incurred in generating and processing the biomass.3 Tetraselmis is a green unicellular organism, 10 to 20 µm in size, usually motile, ellipsoid to ovoid, with 4 flagella of equal length.4 This marine genus also has a large spectrum of antimicrobial activity and its members show probiotic properties. Several Tetraselmis species are economically important as they are ideal for mass cultivation because of their euryhaline and eurythermal nature.5
Response Surface Methodology (RSM) is a useful model for studying the effect of several factors influencing the process of seeking the optimal conditions. This approach reduces the number of experiments, improves statistical interpretation possibilities, and indicates the interaction between multiple variables.6,7 RSM has been successfully applied in many researches, which has become more and more attractive in process optimization.8
There is still lack of guidance on designing the microalgal protein production. To fill this gap, central composite design (CCD) and response surface methodology (RSM) were applied to design the experiments and optimize the cultivation process for Tetraselmis striata selected as model microalgae. This simulation design was performed for obtaining the maximum information in minimum time. In addition, this method allows making decision for the estimation of microalgal performance before the large-scale productions.
Materials and Methods
Maintenance and growth conditions of microalgal strain
The green microalgae Tetraselmis striata EgeMacc-042 was obtained from Ege University Microalgae Culture Collection, Izmir, Turkey. Maintenance of microalgal strain was performed according to the method described in Demirel et al.9
Tetraselmis striata cells were cultured in 250 mL flasks containing 150 mL of F/2 medium in an orbital shaking incubator (IKA® KS 4000ic Thermoshake, Werke GmbH & Co. KG, Germany) with a 20 mm shaking diameter at different light intensities, temperatures and agitation rates for 12 d Illumination was provided by LED downlight lamp (Cata 10 W CT-5254) from the top of the orbital shaking incubator. Irradiance was measured in the center of the flask with a quantum meter (Lambda L1–185).
Measurement of microalgal growth
Samples were taken at indicated times, and the cell concentration was determined by counting triplicate samples in a Neubauer hemocytometer. The amount of protein was determined using Bradford method with Brilliant Blue G 250 dye.10
The growth rate (µ) of the cells was calculated from the initial logarithmic phase of growth for at least 48 h, as µ = (ln C2 - lnC1)/dt, where C2 is final cell concentration, C1 is initial cell concentration and dt is the time required for the increase in concentration from C1 to C2.
Experimental design and data analysis
The experimental simulation design was carried out using 23 full-factorial experiments design with 6 axial points (α = 1.682) and 4 replicates at the central point (55 μmolphotons m−2s−1, 25°C, 150 rpm), according to the Central Composite Design (CCD) by using the Design Expert software (version 7.0.0, Stat-Ease Inc., Minneapolis, MN). The range and the levels of the process variables are given in Table 1. A total of 18 runs were used to optimize the range and the levels of the chosen variables. Each run was completed in 12 d Protein amount (Y, µg/100 µL) was selected as a response of the system.
Table 1.
Experimental range and levels of the independent variables
| Coded Levels |
||||||
|---|---|---|---|---|---|---|
| Independent Variables | Symbol Coded | −α | −1 | 0 | +1 | +α |
| Light intensity (μmol photons m−2s−1) | X1 | 30 | 40 | 55 | 70 | 80 |
| Temperature (°C) | X2 | 20 | 22 | 25 | 28 | 30 |
| Agitation rate (rpm) | X3 | 100 | 120 | 150 | 180 | 200 |
Results and Discussion
A set of experiments was designed by central composite design using response surface methodology and evaluated the influence of physical process variables (light intensity, temperature and agitation rate) for protein amount of Tetraselmis striata. The experimental simulation design matrix, the experimental values and the predicted values for Tetraselmis striata are given in Table 2. As shown in Table 2, 5 different light intensities; X1-μmolphotons m−2s−1 (30, 40, 55, 70, 80), 5 different temperatures; X2-°C (20, 22, 25, 28, 30) and 5 different agitation rates; X3-rpm (100, 120, 150, 180, 200) were tested as physical variables. Also, the protein amount for Tetraselmis striata ranged from 1.44 to 20.28 µg/100 µL, depending on the physical conditions of experiments.
Table 2.
Experimental simulation design matrix and the results for Tetraselmis striata
| Standard order | Variables in non-coded levels |
Tetraselmis striata Protein (μg/ 100 μL) |
|||
|---|---|---|---|---|---|
| Runs | X1 | X2 | X3 | Actual value | Predicted value |
| 1 | 30 | 25 | 150 | 3.532±0.196 | 9.558 |
| 2 | 55 | 25 | 150 | 20.282±0.501 | 24.252 |
| 3 | 40 | 22 | 120 | 1.444±0.112 | 7.385 |
| 4 | 70 | 22 | 120 | 5.932±0.224 | 8.275 |
| 5 | 55 | 25 | 150 | 19.337±0.446 | 24.254 |
| 6 | 40 | 28 | 180 | 7.944±0.243 | 15.991 |
| 7 | 55 | 20 | 150 | 18.063±0.425 | 23.309 |
| 8 | 70 | 28 | 120 | 10.357±0.459 | 12.543 |
| 9 | 40 | 22 | 180 | 12.500±0.386 | 20.811 |
| 10 | 70 | 22 | 180 | 12.341±0.652 | 18.744 |
| 11 | 55 | 25 | 200 | 2.325±0.135 | 17.374 |
| 12 | 70 | 28 | 180 | 9.968±0.682 | 14.373 |
| 13 | 55 | 25 | 100 | 4.901±0.248 | 4.662 |
| 14 | 80 | 25 | 150 | 2.214±0.168 | 8.951 |
| 15 | 55 | 30 | 150 | 14.802±0.701 | 22.850 |
| 16 | 55 | 25 | 150 | 19.008±0.813 | 24.254 |
| 17 | 40 | 28 | 120 | 7.171±0.682 | 11.205 |
| 18 | 55 | 25 | 150 | 15.393±0.436 | 24.254 |
Optimization of physical conditions for protein amount of Tetraselmis striata
Analysis of variance (ANOVA) was used to analyze the responses defined by the design (Table 3). Regression analysis revealed a coefficient of determination (R2) value of 0.9034, indicating that the sample variation of only 9.66% of the total variation was not explained by the model. Meanwhile, the adjusted correlation coefficient (Adj. R2 = 0.7947) and the predicted correlation coefficient (Pred. R2 = 0.3795) values also confirmed that the model was good. The importance of each coefficient was determined by the values of F and p.11 The model F-value was 8.31 with a low p-value (0.0033) for protein amount, implying that the model was adequate for the response variables that were tested. The associated p-value was used to judge whether F was large enough to indicate statistical significance or not.7 Moreover, lack of fit F-value (2.30) implied that the lack of fit was not significant relative to pure error. A quadratic polynomial equation in terms of actual factors was found for physical process conditions of Tetraselmis striata, as following:
| (1) |
where Y is the predicted response; protein amount (µg/100 µL), and X1, X2 and X3 are the values of the test variables; light intensity (μmol photons m−2s−1), temperature (°C) and agitation rate (rpm) respectively. Figure 1 shows the regression plot of the protein amount of actual values against those predicted by Eqn. (1), revealing a linear mathematical relationship among them. The relationship between the actual values and predicted values demonstrated that the model covered the experimental range of studies sufficiently.
Table 3.
Analysis of variance (ANOVA) of the model for protein amount of Tetraselmis striata
| Source | Sum of squares | Degree of freedom | Mean square | F-value | p > F |
|---|---|---|---|---|---|
| Model | 622.68 | 9 | 69.19 | 8.31 | 0.0033 |
| X1: Light intensity | 3.93 | 1 | 3.93 | 0.47 | 0.5116 |
| X2:Temperature | 0.37 | 1 | 0.37 | 0.045 | 0.8373 |
| X3: Agitation | 13.38 | 1 | 13.38 | 1.61 | 0.2405 |
| X1X2 | 0.097 | 1 | 0.097 | 0.012 | 0.9167 |
| X1X3 | 4.22 | 1 | 4.22 | 0.51 | 0.4968 |
| X2X3 | 36.47 | 1 | 36.47 | 4.38 | 0.0697 |
| 342.96 | 1 | 342.96 | 41.21 | 0.0002 | |
| 2.16 | 1 | 2.16 | 0.26 | 0.6244 | |
| 309.36 | 1 | 309.36 | 37.17 | 0.0003 | |
| Residual | 66.58 | 8 | 8.32 | ||
| Lack of fit | 52.80 | 5 | 10.56 | 2.30 | 0.2626 |
| Pure error | 13.79 | 3 | 4.60 | ||
| Cor. total | 689.27 | 17 | |||
| Std. dev. | 2.88 | R-squared | 0.9034 | ||
| Mean | 10.42 | Adj. R-squared | 0.7947 | ||
| C.V.% | 27.69 | Pred. R-squared | 0.3795 | ||
| Press | 427.72 | Adeq. precision | 7.279 |
Figure 1.
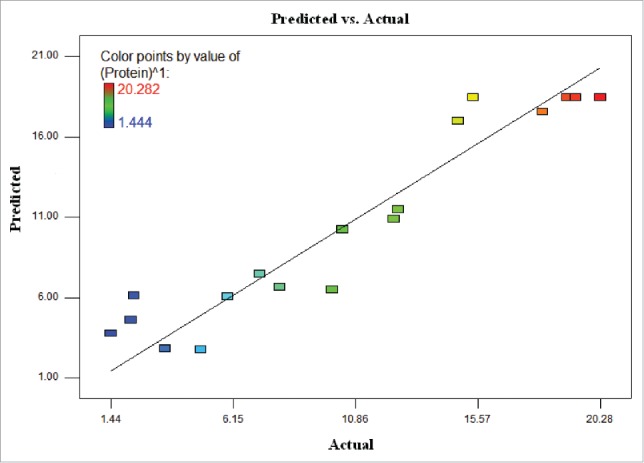
The relationship between the predicted values and actual values for protein amount of T. striata.
With the help of Design Expert 7.0.0, the model graph of the response for T. striata was established in Figure 2 and cubic response surface was found. The shape of the response surface also showed a moderate interaction between 2 factors. At the lowest and the highest light intensities within the studied range of temperature, the protein amount was found at low level. The protein amount increased with increasing the light intensity from 40 to 55 μmol photons m−2 s−1 within the studied range of temperature. Furthermore, more than 11.5 µg/100µL protein amounts occurred when the light intensity was greater than 40 μmol photons m−2 s−1.
Figure 2.
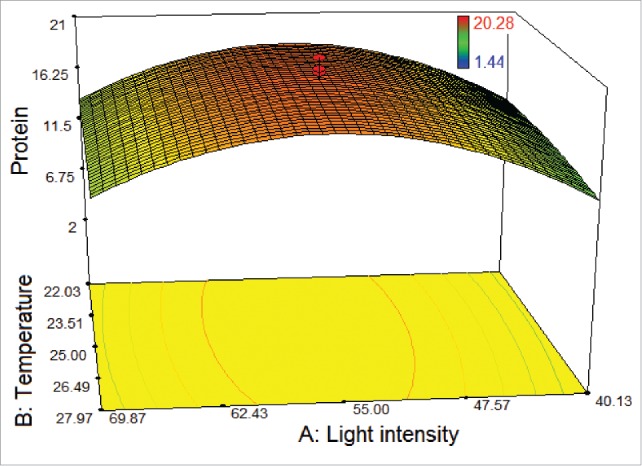
3D response surface plot of central composite design showing the mutual effects of light intensity and temperature on protein amount (μg/100 μL) of T. striata.
As shown in Figure 3, a weak effect on the response was observed for the agitation rate of 180 rpm at the maximum and the minimum levels of the light intensity. In addition, higher and lower levels of both agitation rate and light intensity did not result in higher protein amount. In fact, it was observed that the lowest protein amount was found at 120 rpm under the light intensity of 40 μmol photons m−2 s−1, as the cells could not proliferate. Protein amount was found to extremely increase with decreasing the light intensity (from 70 to 55 μmol photons m−2 s−1) and increasing the agitation rate (from 120 to 150 rpm), while the temperature was kept at 25°C.
Figure 3.
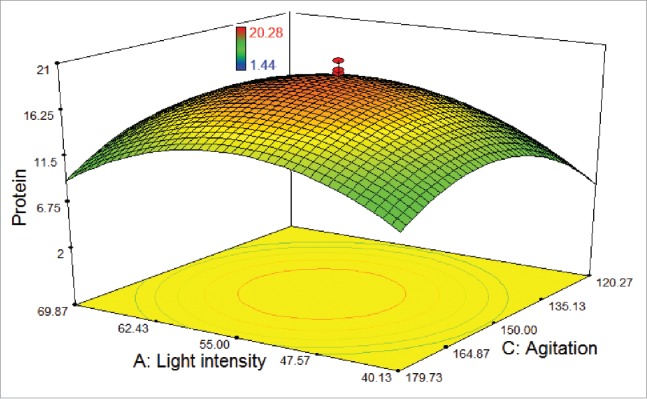
3D response surface plot of central composite design showing the mutual effects of light intensity and agitation rate on protein amount (μg/100 μL) of T. striata.
Figure 4 shows the interaction between the temperature and agitation rate. The highest protein amount could be obtained in the temperature levels and at the agitation rates, ranging from 23.5°C to 26.5°C and from 135 rpm to 165 rpm, respectively, while the light intensity was set at 55 μmol photons m−2s−1. Nevertheless, below the level of agitation rate of 135 rpm showed a declining trend on protein amount due to the metabolic change affected the physiological state of the cells. It was recorded that temperature and agitation rate were slightly interdependent.
Figure 4.
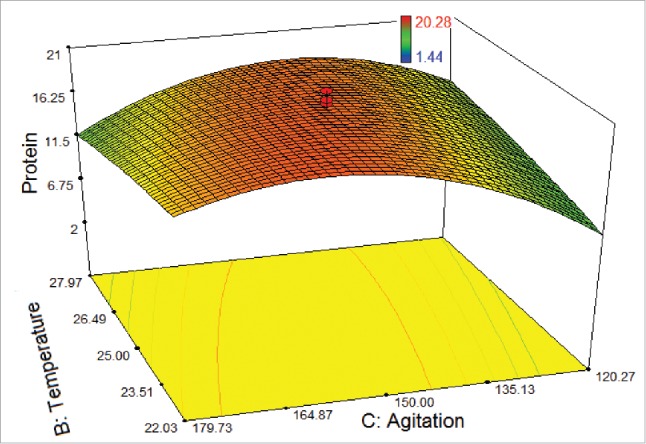
3D response surface plot of central composite design showing the mutual effects of temperature and agitation rate on protein amount (μg/100 μL) of T. striata.
It is also important to underline that light has profound quantitative and qualitative effects on protein formation. The photosynthetic process needs light to take place and hence microalgal cultivation must be carried out with artificial light or under sunlight. Light intensity not only affects biomass productivity, but also controls the profile of biochemical compositions of photosynthetic microalgae.12 Furthermore, temperature impacts the cell physiology by changing the rate of chemical reactions and the stability of cellular components.13 It should be noted here that high light intensity stimulated faster growth than high temperature for the protein production of T. striata.
Validation of the model for protein amount of T. striata
The optimization of the process for the response was generated by the numerical optimization technique following desirability function. In the optimization stage, the physical process variables were set within the range between low (−1) and high (+1) and the response was set to the maximum value. The optimization solution of T. striata (approximately under the light intensity of 55 μmol photons m−2s−1, in 22°C and at the agitation rate of 160 rpm) was selected because it resulted in the highest predicted response with the highest desirability of 0.915.
To verify the predicted results, validation experiment was performed in triplicate tests. Validation under the optimized conditions was performed in a 250-mL Erlenmeyer flask containing 150 mL F/2 medium by measuring the cell count and the protein amount. As shown in Figure 5, the experimental protein result was found to be 22.65 ± 0.56 µg/100 µL, while the predicted maximum protein amount was 20.28 µg/100 µL, indicating the accuracy of the optimization result. Furthermore, the maximum cell concentration of 19 ± 0.44 × 105 cells mL−1, which corresponded to the growth rate of 0.211 day−1, was obtained for T. striata under the optimized conditions. As reported by Laws et al.,14 the marine prasinophyte Tetraselmis suecica was grown in a chemostat culture system over a series of phosphate-limited growth rates ranging from 0.164 to 0.755 d−1.
Figure 5.
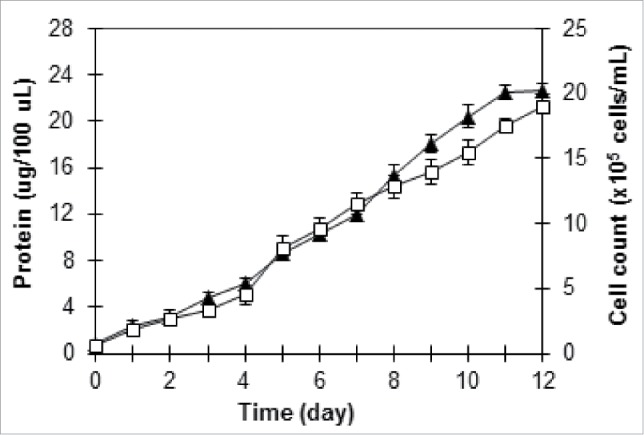
Growth profile of T. striata under optimized conditions: protein (▴), cell count (□).
In conclusion, the good agreement between the predicted values and the experimental values confirmed the validity of the model and the developed quadratic model was proven to be statistically adequate for microalgal protein production. It was reported that the experimental protein amount was 22.65 ± 0.56 µg/100 µL, which was closer to the predicted protein amount (20.28 µg/100 µL) at 160 rpm in 22°C under the light intensity of 55 μmol photons m−2s−1. In this paper, the noteworthy result was that both the light intensity and the agitation rate were dominant factors for obtaining high protein amount. The results also showed a holistic approach for opening a new avenue on simulation design of microalgal protein optimization.
Disclosure of Potential Conflicts of Interest
No potential conflicts of interest were disclosed.
Acknowledgments
The author would like to thank Prof. Dr. Meltem Conk Dalay and Dr. Zeliha Demirel for their supports and comments during the study.
References
- 1.Brennan L, Owende P. Biofuels from microalgae-a review of technologies for production, processing, and extractions of biofuels and co-products. Renew Sust Energy Rev 2010; 14(2):557-77; http://dx.doi.org/ 10.1016/j.rser.2009.10.009 [DOI] [Google Scholar]
- 2.Pulz O, Gross W. Valuable products from biotechnology of microalgae. Appl Microbiol Biotechnol 2004; 65(6):635-48; PMID:15300417; http://dx.doi.org/ 10.1007/s00253-004-1647-x [DOI] [PubMed] [Google Scholar]
- 3.Fon Sing S, Isdepsky A, Borowitzka MA, Lewis DM. Pilot-scale continuous recycling of growth medium for the mass culture of a halotolerant Tetraselmis sp. in raceway ponds under increasing salinity: A novel protocol for commercial microalgal biomass production. Bioresour Technol 2014; 161:47-54; PMID:24681683; http://dx.doi.org/ 10.1016/j.biortech.2014.03.010 [DOI] [PubMed] [Google Scholar]
- 4.Regan DL. Other micro-algae In (Eds. Borowitzka MA, Borowitzka LJ) Microalgal Biotechnology 1988; pp. 135-50, Cambridge University Press, Cambridge. [Google Scholar]
- 5.Fabregas J, Abalde J, Herrero C, Cabezas B, Veiga M. Growth of the marine microalga Tetraselmis suecica in batch cultures with different salinities and nutrient concentrations. Aquacult 1984; 42:207-15; http://dx.doi.org/ 10.1016/0044-8486(84)90101-7 [DOI] [Google Scholar]
- 6.Vishwanatha KS, Rao AGA, Singh SA. Acid protease production by solid state fermentation using Aspergillus oryzae MTCC 5341: optimization of process parameters. J Ind Microbiol Biotechnol 2010; 37:129-38; PMID:19937364; http://dx.doi.org/ 10.1007/s10295-009-0654-4 [DOI] [PubMed] [Google Scholar]
- 7.Zhou X, Xin ZJ, Lu XH, Yang XP, Zhao MR, Wang L, Liang JP. High efficiency degradation crude oil by a novel mutant irradiated from Dietzia strain by 12C6+ heavy ion using response surface methodology. Bioresour Technol 2013; 137:386-93; PMID:23603188; http://dx.doi.org/ 10.1016/j.biortech.2013.03.097 [DOI] [PubMed] [Google Scholar]
- 8.Guo J, Zhuang Y, Chen L, Liu J, Li D, Ye N. Process optimization for microwave-assisted direct liquefaction of Sargassum polycystum C. Agardh using response surface methodology. Bioresour Technol 2012; 120:19-25; PMID:22776261; http://dx.doi.org/ 10.1016/j.biortech.2012.06.013 [DOI] [PubMed] [Google Scholar]
- 9.Demirel Z, Imamoglu E, Dalay MC. Fatty acid profile and lipid content of Cylindrotheca closterium cultivated in air-lift photobioreactor. J Chem Technol Biotechnol 2015; http://dx.doi.org/ 10.1002/jctb.4687 [DOI] [Google Scholar]
- 10.Bradford MM. A rapid and sensitive for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 1976; 72:248-54; PMID:942051; http://dx.doi.org/ 10.1016/0003-2697(76)90527-3 [DOI] [PubMed] [Google Scholar]
- 11.Liu Y, Wang J, Zheng Y, Wang A. Adsorption of methylene blue by kapok fiber treated by sodium chlorite optimized with response surface methodology. Chem Eng J 2012; 184:248-55; http://dx.doi.org/ 10.1016/j.cej.2012.01.049 [DOI] [Google Scholar]
- 12.Hu Q. Environmental effects on cell composition In (Ed. Richmond A.) Handbook of Microalgal Culture: Biotechnology and Applied Phycology 2004; pp. 83-93, Blackwell Science Ltd, Oxford. [Google Scholar]
- 13.Sandnes JM, Källqvist T, Wenner D, Gislerød HR. Combined influence of light and temperature on growth rates of Nannochloropsis oceanica: linking cellular responses to large-scale biomass production. J Appl Phycol 2005; 17:515-25; http://dx.doi.org/ 10.1007/s10811-005-9002-x [DOI] [Google Scholar]
- 14.Laws EA, Pei S, Bienfang P, Grant S. Phosphate-limited growth and uptake kinetics of the marine prasinophyte Tetraselmis suecica (Kylin) Butcher Aquacult 2011; 322:117-21; http://dx.doi.org/ 10.1016/j.aquaculture.2011.09.041 [DOI] [Google Scholar]


