Abstract
A defining feature of Trypanosoma brucei cell shape is the lateral attachment of the flagellum to the cell body, mediated by the flagellum attachment zone (FAZ). The FAZ is a complex cytoskeletal structure that connects the flagellum skeleton through two membranes to the cytoskeleton. The FAZ acts as a ‘cellular ruler’ of morphology by regulating cell length and organelle position and is therefore critical for both cell division and life cycle differentiations. Here we provide an overview of the advances in our understanding of the composition, assembly, and function of the FAZ.
Keywords: Trypanosoma brucei, flagellum, FAZ, morphology, cytoskeleton
Trends
The flagellum attachment zone (FAZ) is a large cytoskeletal structure that connects the flagellum skeleton to the cell body cytoskeleton through the membrane of both the flagellum and the cell body. The structure can be divided into eight zones.
The FAZ is a key morphogenetic structure regulating both cell length and organelle positioning.
Recent studies have identified numerous FAZ proteins. The function of a subset of these proteins has been studied by RNAi, revealing a range of different phenotypes from flagellum detachment to organelle positioning effects.
The assembly of the FAZ occurs at its proximal end – the opposite polarity to that of the flagellar axoneme and paraflagellar rod.
FAZ: A Key Regulator of Morphogenesis in Trypanosomes
Trypanosoma brucei is a protozoan parasite that causes human African trypanosomiasis and nagana in cattle. The T. brucei trypomastigote is long and slender with a single flagellum that emerges near the posterior of the cell and is subsequently attached to the cell body for most of its length as it follows a defined helical path towards the anterior of the cell. A complex structure termed the FAZ mediates this lateral attachment 1, 2, 3. The FAZ is a key regulator of cell morphogenesis; disruption of FAZ assembly has been linked to morphogenetic changes such as flagellum detachment and alterations in cell shape and size 4, 5, 6, 7, 8, 9, 10.
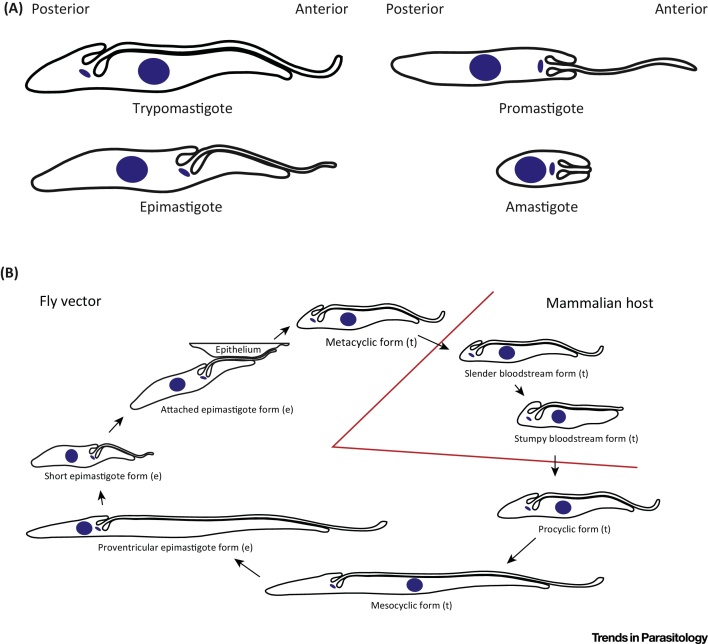
The early classifications of trypanosomes relied on their morphological characteristics, with the terminology for trypanosome shape defined by the relative position of the kinetoplast and the nucleus coupled with the length of attached flagellum (Figure 1A) [11]. During the T. brucei life cycle there are substantial changes in cell morphology that appear adapted for pathogenicity and proliferation in both the mammalian host and the insect vector; these changes have been linked to modulations of the protein composition of the FAZ (Figure 1B) 12, 13.
Figure 1.
The Morphologies of Kinetoplastid Parasites and the Life Cycle of Trypanosoma brucei. (A) Cartoons showing the common morphologies of kinetoplastid parasites with the anterior and posterior of the cell indicated. The nucleus is the large blue circle and the kinetoplast (mitochondrial DNA) is the small blue oval located at the base of the flagellum. The trypomastigote morphology has the kinetoplast posterior to the nucleus with a long attached flagellum. In the epimastigote and promastigote, the kinetoplast is anterior to the nucleus, but whereas the epimastigote has a long attached flagellum, in the promastigote the flagellum emerges from the anterior pole of the cell. In the amastigote, the kinetoplast is anterior to the nucleus with a small cell body and the flagellum does not emerge beyond the cell body. (B) A simplified cartoon showing the life cycle of T. brucei. The metacyclic form infects the mammalian host where it differentiates first into the slender bloodstream form followed by the stumpy bloodstream form. The fly vector then takes up the stumpy bloodstream form where it differentiates through several stages before differentiating into the mammalian infective metacyclic form in the salivary gland of the fly. For most of its life cycle T. brucei exists as a trypomastigote [indicated by (t)] except for the proventricular epimastigote, the short epimastigote, and the attached epimastigote forms, where it has epimastigote morphology [indicated by (e)].
The structure of the FAZ has been described in detail using various microscopy techniques but until recently only a few molecular components were known 1, 2, 14. Here we discuss the nature and functions of the FAZ proteins and their assembly in the context of the key roles the FAZ plays in defining and maintaining the shape and form of T. brucei and ensuring that this shape is faithfully inherited at division. This linear feature of the trypanosome can be thought of as a ‘cellular ruler’ whose length and constitution facilitates changes in cell form.
Cell Morphology and FAZ Structure
The distinctive shape of a trypomastigote is maintained by a helical, crosslinked, subpellicular array of microtubules arranged in parallel with their plus ends facing towards the posterior end of the cell [15]. The mitochondrial genome (kinetoplast) is linked to the flagellum basal body 16, 17, 18. The flagellum emerges from the cell and is attached to the cell body along most of its length. Flagellum attachment is mediated by the FAZ, which is a large, interconnected set of fibres, filaments, and junctional complexes linking the flagellum skeleton through both the flagellum and cell body membranes to a specialised FAZ filament and associated microtubule quartet (MtQ) 1, 2, 14, 15. The MtQ is antiparallel to the other subpellicular microtubules in the cell body. The FAZ and adjacent MtQ generate an asymmetric seam within the microtubule corset. In transverse sections the organisation of the corset microtubules (polarity: plus end at posterior end of cell) is interrupted by the MtQ (polarity: minus end at posterior end of cell) and the FAZ [15]. Five different linkage types might therefore exist to orchestrate this internal organisation of the FAZ, besides those that link components to the cell membrane (Figure 2). The identification of components of these linkages will be important for our understanding of FAZ biogenesis and function. The definition of trypanosome shape and polarity is therefore derived from the internal cytoskeleton and the flagellum position; this arrangement then coordinates the position of the single-copy organelles 19, 20.
Figure 2.
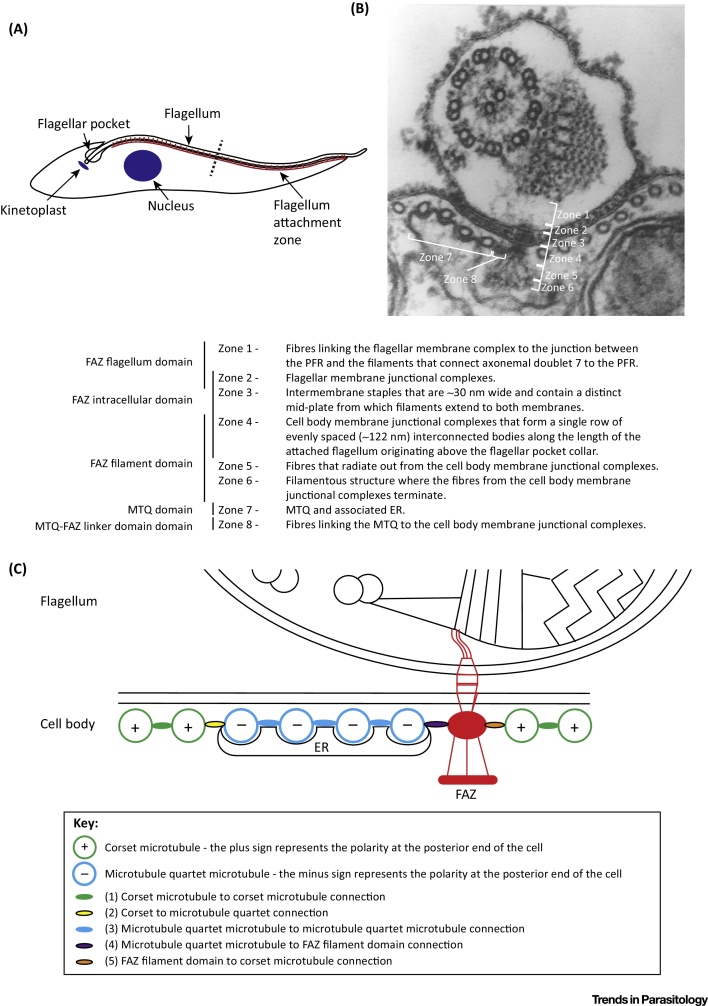
Organisation of the Flagellum Attachment Zone (FAZ) in Trypanosoma brucei. (A) Simplified cartoon of the key components of a T. brucei trypomastigote cell. Arrows indicate the flagellum, FAZ, flagellar pocket, nucleus, and kinetoplast, with the FAZ highlighted in red and the nucleus and kinetoplast highlighted in blue. The dashed line across the flagellum and FAZ indicates the region where the transverse section in (B) and (C) was taken. (B) Transmission electron microscopy image showing a transverse section of the FAZ with the different zones and structures within them. Abbreviations: ER, endoplasmic reticulum; MTQ, microtubule quartet; PFR, paraflagellar rod. Reproduced from [2]. (C) Cartoon of the transverse section shown in (B) highlighting the asymmetric seam generated in the corset microtubules by the FAZ and MTQ. The five different connections between the various components are shown.
Moving towards a molecular description of the FAZ will require definition of precise protein locations within this complex structure; therefore, in Figure 2 we have defined the major structural zones of the FAZ. Combinations of these zones can now be used to describe different domains of the FAZ, creating a hierarchical descriptive terminology (Figure 2). There is some overlap between the domains but given the present state of knowledge we believe it is useful to have a level of ambiguity.
-
(i)
Zones 1 and 2 constitute the FAZ flagellum domain.
-
(ii)
Zones 2, 3, and 4 constitute the FAZ intracellular domain.
-
(iii)
Zones 4, 5, and 6 constitute the FAZ filament domain.
-
(iv)
Zone 7 constitutes the MtQ domain.
-
(v)
Zone 8 constitutes the MtQ–FAZ linker domain.
There is an invagination of the cell membrane around the base of the flagellum called the flagellar pocket (FP), around which the MtQ wraps helping to define its overall shape. The FP is the only known site for exo- and endocytosis in the trypanosome and is the location of many receptors such as the transferrin receptor 21, 22, 23. The FP forms an interface where signals to/from the host are integrated and processed and hence has a central role in trypanosome pathogenicity. The FAZ also has a potential role in pathogenesis, as the bloodstream form (BSF) in all trypanosomatids examined is a trypomastigote (Figure 1A), and therefore the evolution of extended flagellum attachment appears advantageous in this ecological niche [24]. One possible advantage of flagellum attachment is that it allows host immune system factors bound to the flagellum to be moved towards the FP by hydrodynamic flow or other mechanisms 24, 25.
The overall FAZ structure has similarities to cell–cell adhesive junctions such as desmosomes in multicellular organisms and hence was described in trypanosomes using the term macula adherens [1]. Desmosomes are large, complex cytoskeletal structures spanning two plasma membranes normally connecting intermediate filament bundles in separate adjacent cells [26]; however, autocellular adherens junctions between regions of the same cell like the FAZ are found in certain cells of the trachea of Drosophila [27]. Desmosomes comprise two ‘half units’ positioned on adjacent cells; a half unit contains a cytoplasmic plaque that connects intermediate filaments to transmembrane proteins called cadherins that bind to the cadherins of another half unit [28]. The analogy of the desmosome and protein interaction domains within it is potentially useful for informing molecular models for the organisation of the FAZ and flagellum adhesion (FLA) proteins, as discussed below.
FAZ Protein Location Often Predicts RNAi Phenotype
FAZ molecular components were first revealed using monoclonal antibodies and these were then used to identify the first FAZ proteins and their encoding genes 3, 4, 29. Subsequent FAZ proteins have been identified using various techniques (Table 1). An important advance in understanding FAZ protein function will be the positioning of the FAZ proteins in Table 1 into the specific FAZ zones, although we recognise the difficulties. As a start, it is now possible to locate some known FAZ proteins to their respective FAZ domains. We have made an initial attempt to integrate some of these proteins into a model of their potential interactions (Figure 3). This is the first step towards eventually allowing us to understand the interaction details in the same depth as those of the desmosome (Figure 3).
Table 1.
Known FAZ Proteins
| Gene ID | Name | Localisation | Localisation Method | Discovery Route | Molecular Weight (kDa) | Known Domains | Coiled Coils | Repeats | BSF or PCF Expression | Refs |
|---|---|---|---|---|---|---|---|---|---|---|
| Tb927.8.4010 | FLA1 | Intracellular domain | Immunofluorescence | Monoclonal antibodies | 59 | NHL domain | No | No | PCF | [29] |
| Tb927.8.4060 | FLA2 | Intracellular domain | Not done | Bioinformatic | 64 | NHL domain | No | No | BSF | [6] |
| Tb927.8.4110 | FLA3 | Intracellular domain | Not done | Bioinformatic | 64 | NHL domain | No | No | BSF | [8] |
| Tb927.8.4050 | FLA1BP | Intracellular domain | eYFP tagging and immunofluorescence | Immunoprecipitation | 83 | NHL domain | No | No | PCF | [8] |
| Tb927.8.4100 | FLA1BP | Intracellular domain | eYFP tagging and immunofluorescence | Immunoprecipitation | 83 | NHL domain | No | No | PCF | [8] |
| Tb927.5.4570 | FLA3 | Intracellular domain | Immunofluorescence | Lectin binding | 89 | NHL domain | No | No | BSF | [43] |
| Tb927.5.4580 | FLA3 | Intracellular domain | Immunofluorescence | Lectin binding | 89 | NHL domain | No | No | BSF | [43] |
| Tb927.10.2880 | Ca2+ channel | Intracellular domain | Immunofluorescence | Proteome validation | 304 | Ion transporter | Yes | No | Both | [62] |
| Tb927.10.8830 | FAZ5 | Intracellular domain | eYFP tagging | Immunoprecipitation | 67 | Chaperone J domain | Yes | No | Both | [30] |
| Tb927.10.14320 | FAZ9 | FAZ filament domain | eYFP tagging | Bioinformatic | 122 | ARM repeats | Yes | No | Both | [30] |
| Tb927.11.1090 | ClpGM6 | FAZ flagellum domain | Immunofluorescence | RNAi screen | >660 | Calpain | Yes | Yes | Both | [9] |
| Tb927.8.4780 | FLAM3 | FAZ flagellum domain | eYFP tagging | Proteome validation | 468 | Clu domain | Yes | Yes | Both | [13] |
| Tb927.1.4310 | FAZ2 | FAZ filament domain | eYFP tagging | Immunoprecipitation | 184 | No domains | Yes | Yes | Both | [30] |
| Tb927.11.12530 | FAZ3 | FAZ filament domain | eYFP tagging | Immunoprecipitation | 91 | No domains | Yes | No | Both | [30] |
| Tb927.7.3330 | FAZ10 | Unknown | Immunofluorescence | BioID | 503 | PFAMB506 | Yes | Yes | Both | [63] |
| Tb927.4.2060 | FAZ8 | FAZ filament domain | eYFP tagging | Bioinformatic | 67 | PFAMB506 | Yes | No | Both | [30] |
| Tb927.4.3740 | FAZ1 | FAZ filament domain | Immunofluorescence and immunoelectron microscopy | Monoclonal antibodies | 193 | PFAMB506 | Yes | Yes | Both | [4] |
| Tb927.4.2080 | CC2D | FAZ filament domain | Immunofluorescence and eYFP tagging | Proteome validation | 105 | C2 domain/PFAMB14968 | Yes | No | Both | [5] |
| Tb927.9.10530 | FAZ4 | FAZ filament domain | eYFP tagging | Immunoprecipitation | 119 | PFAMB506 | Yes | No | Both | [30] |
| Tb927.10.840 | FAZ6 | FAZ filament domain | eYFP tagging | Immunoprecipitation | 195 | WD domain/PFAMB506 | Yes | No | Both | [30] |
| Tb927.4.5340 | FAZ11 | Unknown | Immunofluorescence | BioID | 95 | No domains | Yes | No | Both | [63] |
| Tb927.10.15390 | FAZ7 | FAZ filament domain | eYFP tagging | Immunoprecipitation | 124 | Kinesin | Yes | No | Both | [30] |
| Tb927.11.13230 | TbVAP | FAZ filament domain (ER) | Immunofluorescence | Proteome validation | 24 | MSP domain | No | No | Both | [20] |
| Tb927.9.13820 | KMP11 | Cell body and flagellum | Immunofluorescence | Immunoprecipitation | 11 | KMP11 domain | Yes | No | Both | [47] |
| Tb927.9.13880 | KMP11 | Cell body and flagellum | Immunofluorescence | Immunoprecipitation | 11 | KMP11 domain | Yes | No | Both | [47] |
| Tb927.9.13920 | KMP11 | Cell body and flagellum | Immunofluorescence | Immunoprecipitation | 11 | KMP11 domain | Yes | No | Both | [47] |
| Tb927.11.2590 | FAZ12 | FAZ filament domain | Immunofluorescence | BioID | 121 | No domains | Yes | No | Both | [48] |
| Tb927.3.1020 | FAZ13 | FAZ filament domain | Immunofluorescence | BioID | 54 | No domains | Yes | No | Both | [48] |
| Tb927.8.6980 | FAZ14 | FAZ filament domain | Immunofluorescence | BioID | 95 | no domains | Yes | No | Both | [48] |
| Tb927.11.3300 | TbSAS4 | Anterior tip of FAZ filament domain | Immunofluorescence | BioID, bioinformatics | 108 | T-complex protein 10 C terminus | Yes | No | Both | 31, 48 |
| Tb927.8.7070 | FAZ15 | Unknown | Immunofluorescence | Phosphoproteomics | 35 | No domains | No | No | Both | [31] |
| Tb927.5.3460 | FAZ16 | Unknown | Immunofluorescence | Phosphoproteomics | 57 | LysM domain | No | No | Both | [31] |
| Tb927.10.7210 | FAZ17 | Unknown | Immunofluorescence | Phosphoproteomics | 26 | No domains | No | No | Both | [31] |
| Tb927.11.15800 | TOEFAZ1 | Anterior tip of FAZ filament domain | Immunofluorescence | BioID, phosphoproteomics | 90 | No domains | Yes | No | Both | [31] |
Figure 3.
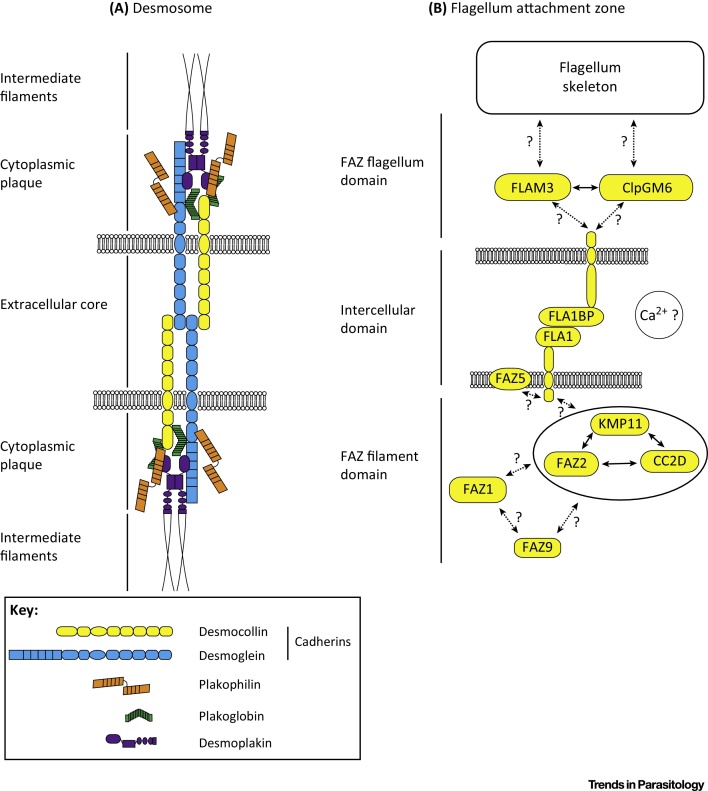
Comparison of the Molecular Organisation of a Desmosome with a Flagellum Attachment Zone (FAZ). (A) Desmosomes are found in multicellular organisms and are large structures that connect together the cytoskeleton of two cells through two plasma membranes; hence, they can be considered analogous to the FAZ. The desmosome model was adapted from [28] and shows the key protein components. The cadherins desmocollin and desmoglein constitute the extracellular core of the desmosome and connect the two halves of the desmosome together. The cadherins are connected into the cytoplasmic plaque by various proteins including plakophilin, plakoglobin, and desmoplakin that in turn bind to the intermediate filaments. (B) Potential interactions and localisations of a selected group of FAZ proteins are shown. Interactions with experimental evidence are shown with a solid line with other possible interactions shown with a dotted line and question mark. Calcium ions are included in the model, as Vickerman noted that Trypanosoma brucei harvested from an animal and then grown in blood containing citrate (a calcium chelator) had detached flagella [1]. This is a highly speculative model; however, it will provide a framework on which to integrate new insights into FAZ protein function and helps to highlight the paucity of knowledge about the FAZ compared with the desmosome.
In addition to the FAZ domains’ transverse heterogeneity, there is also heterogeneity in protein localisation longitudinally along the FAZ length 5, 30. We now discuss FAZ proteins and their function and demonstrate that a pattern has emerged linking the RNAi phenotype observed and protein localisation. In general, results so far indicate that if the FAZ protein is located within the FAZ flagellum domain, RNAi leads to shortening of the FAZ and epimastigote-like morphology, whereas if the protein is present in the FAZ intracellular domain RNAi results in flagellum detachment 6, 8, 9, 10. Finally, if the protein is present in the FAZ filament domain RNAi results in either flagellum detachment or kinetoplast or nucleus positioning effects 4, 5, 31.
FAZ Flagellum Domain (Zones 1 and 2)
We showed that calpain-like protein GM6 (ClpGM6) localised to the FAZ flagellum domain and on depletion of this protein the length of flagellum attachment to the cell body was shortened [9]. The cells underwent a remarkable morphological transition, changing from a trypomastigote to an epimastigote-like configuration with the kinetoplast juxtaposed or anterior to the nucleus with a much shorter FAZ. There was also a coordinate relocation of associated organelles such as the Golgi and FP. Remarkably, ClpGM6 RNAi was not lethal and cells with the epimastigote-like morphology continued to proliferate in culture; however, the ability of this cell line to infect tsetse flies has not been assessed. The phenotype resulting from the depletion of FLAgellar Member 3 (FLAM3), another FAZ flagellum domain protein recently examined 7, 10, showed similarity to ClpGM6; depletion of FLAM3 resulted in the cells transitioning from a trypomastigote to an epimastigote-like morphology [10]. We defined a dependency of expression between FLAM3 and ClpGM6, with loss of one resulting in the reduction of expression of the other [10]. It is possible that ClpGM6 and FLAM3 are part of the same complex, suggesting that individual FAZ proteins group into large-scale molecular complexes and facilitating an understanding of the FAZ assembly hierarchy.
During the T. brucei life cycle, trypanosomes adopt different morphologies with different FAZ lengths (Figure 1B). Rotureau and colleagues [13] examined the FAZ during the life cycle and showed that there was a drop in labelling with monoclonal antibodies specific to the FAZ between mesocyclic and epimastigote cells coinciding with a switch from trypomastigote to epimastigote morphology. In trypanosomes, protein expression regulation occurs at the post-transcriptional level and the RNA-binding proteins ALBA3/4 have been implicated in the transition to the epimastigote form [32]. On knock down of ALBA3/4, cells begin to adopt an epimastigote morphology and RNA-seq analysis of these cells showed that the ClpGM6 transcript was the one most highly downregulated [32]. The FAZ has previously been implicated in nuclear positioning during cell division [4]; does this drop in FAZ protein expression allow the migration of the nucleus that occurs between the mesocyclic and epimastigote forms?
FAZ Intracellular Domain (Zones 2, 3, and 4)
FLA1 was identified in T. brucei due to its similarity to the FAZ protein GP72 from Trypanosoma cruzi [29]. Interestingly, GP72 was identified by an anticarbohydrate monoclonal antibody, WIC29.26, and subsequent publications have revealed that the glycoprotein has an unusual sugar composition 33, 34, 35, 36, 37. Several studies revealed the involvement of specific carbohydrates and glycoproteins in flagellum attachment, including one publication that showed that loss of GDP-fucose production resulted in flagellum detachment 6, 29, 38, 39, 40, 41, 42, 43. However, overall the role of specific carbohydrates has not been much explored; this seems odd given the significance of the early finding, as the presence of glycoconjugates adds another layer of complexity and another potential regulatory step to the FAZ structure.
Both FLA1 and GP72 are transmembrane proteins with a small intracellular domain and a glycosylated extracellular region containing NHL repeats, which is a widespread protein interaction domain [44]. Immunoprecipitation of FLA1 identified FLA1-binding protein (FLA1BP), which is predicted to be a glycosylated transmembrane protein with a large extracellular region containing NHL repeats and a small intracellular domain 8, 45. Using a combination of tagged FLA1 and FLA1BP and RNAi, Sun and colleagues demonstrated that FLA1 is located on the cell body membrane and FLA1BP on the flagellum membrane [8]. Given that these proteins interact, we suggest they may be functionally analogous to the desmosomal cadherins by connecting the FAZ flagellum and FAZ filament domains (Figure 3).
The interaction between FLA1 and FLA1BP was shown in procyclic forms (PCFs) but the expression of these proteins is downregulated in BSFs [46]. The BSF surface coat (variant surface glycoprotein) is structurally distinct from the PCF surface (procyclins), suggesting a reason why BSF equivalents of FLA1 and FLA1BP have evolved. T. brucei contains two proteins (FLA2 and FLA3) that have ∼60% identity to FLA1 with the same domain structure and are upregulated in BSFs 6, 8, and therefore are likely to be able to substitute for FLA1. In addition, another BSF-specific protein that localises to the FAZ was identified encoded by near-identical genes (Tb927.5.4570 and Tb927.5.4780) that has ∼40% identity to FLA1BP with the same domain structure; unfortunately, this was also named FLA3 43, 45. Currently there is no evidence of interaction between FLA3 43, 45 and FLA2/3 6, 8 proteins and so this confusing nomenclature will best be refined when more is known about the biochemistry of these proteins.
FLA1 depletion led to flagellum detachment and eventually cell death, showing that FLA1 is required for flagellum attachment [6]. Interestingly, the T. cruzi knockout cell line of the FLA1 orthologue GP72 was viable despite having detached flagella 38, 39. The expression of the T. cruzi GP72 protein in T. brucei led to flagellum detachment and the cells were viable, albeit with a much reduced growth rate. The sequence divergence between FLA1 and GP72 appears to have resulted in incompatibilities between the binding interfaces required to ensure flagellum attachment, providing a potential route to dissect the function of the different domains of FLA1 [6]. As with FLA1 RNAi, FLA1BP knock down resulted in full-length detachment of the flagellum and also a reduction in both FAZ length and cell body length, resulting in the production of cells with an epimastigote-like morphology. The appearance of cells with an epimastigote-like morphology fits with the RNAi phenotype pattern described above, as FLA1BP is located within the FAZ flagellum domain in addition to being in the FAZ intracellular domain. However, unusually for a flagellum detachment phenotype, the cells continued to proliferate [8]. In the literature there is no clear correlation between growth and the amount of flagellum detachment. Our experience suggests that how cells are handled and prepared for microscopy has a major effect on the apparent level of detachment. It will be of interest to make quantitative comparisons over long periods of culture with careful handling to ensure that the images of the cells on the slide are representative of the live cells in the culture flask. Experience shows that, in some protocols, during the washing and settling of the cells onto a slide the flagellum tends to become detached if the FAZ structure has been weakened; therefore, the speed and duration of the centrifugation washing steps, resuspension protocols, and the slide adherence method all have an effect. Comparative studies of different mutant phenotypes using one protocol will be useful in the future, as will careful comparison of live cells or cells fixed directly in culture with those imaged as fully processed, labelled cells or cytoskeletons.
FAZ Filament Domain (Zones 4, 5, and 6)
Both coiled-coil C2 domain-containing protein (CC2D) and FAZ2 localise to the FAZ filament domain and depletion of either leads to inhibition of FAZ assembly causing full-length flagellum detachment and cell death 5, 30, 47. The reduction in FAZ length led to a decrease in cell body length during the knock down of both CC2D and FAZ2. A FAZ1 stub was still able to form near the FP in these knock down cells, suggesting that FAZ nucleation can occur without a full complement of FAZ proteins 5, 47.
FAZ1 is also found in the FAZ filament domain; however, the phenotype observed during FAZ1 knock down was different. There was no full-length flagellum detachment; instead, there were flagellum attachment defects. There was also no change in cell body length, as there was no effect on overall FAZ length [4]. However, the FAZ architecture was disorganised, with changes in the labelling patterns of anti-FAZ antibodies [4].
The function of FAZ9, a FAZ filament domain protein, has been analysed by RNAi 30, 31. On FAZ9 depletion the flagellum remained attached to the cell body; however, the kinetoplast was positioned anterior to the nucleus. This is similar to the ClpGM6 and FLAM3 RNAi phenotypes except that there was no change in the overall length of the FAZ on FAZ9 RNAi 9, 10, 31. The phenotypes observed for the knock down of these FAZ filament domain proteins may be dependent on the zone in which the proteins are found. FAZ9 may be located more towards zone 6, hence affecting kinetoplast and nucleus positioning (Figure 2), whereas CC2D and FAZ2 may be in zone 4, resulting in flagellum detachment with FAZ1 in zone 5, where knock down has a less dramatic effect on flagellum attachment.
Zoids are anucleate cytoplasts that are commonly observed in FAZ protein RNAi phenotypes 4, 5. They have a kinetoplast, basal body, and flagellum, suggesting that the positioning of the cytokinesis furrow facilitates inclusion of such organelles via the distal end of the FAZ marking the site of cytokinesis ingression [15]. Also, a clear correlation between FAZ length and cell body length exists and hence coordination between FAZ assembly and the subpellicular microtubule assembly 5, 8, 47. Overall, these data on the function of the FAZ and its constituent proteins demonstrate that the FAZ acts as a cellular ruler for trypanosome morphology, with changes in FAZ protein composition determining both FAZ length and cell body size and also organelle layout.
Besides the dynamic nature of the FAZ as a cellular ruler of cell architecture, it is also likely that it operates to provide a physical platform for spatial biochemistry enabling the specific localisation of regulatory components like polo-like kinase 31, 48, 49, 50, 51.
FAZ Proximal-End Assembly and Implications
We have elucidated the site of FAZ assembly, using cells where the expression of FAZ proteins present in each of the three domains of the FAZ was under the control of an inducible promoter [30]. For every protein examined, the newly synthesised FAZ proteins localised to the proximal end of the FAZ (the FP end), which is at the opposite end to the addition site for flagellar components, showing that the overall flagellum/FAZ structure has two distinct assembly sites (Figure 4A). Others have subsequently confirmed this result and reiterated our original model [47].
Figure 4.
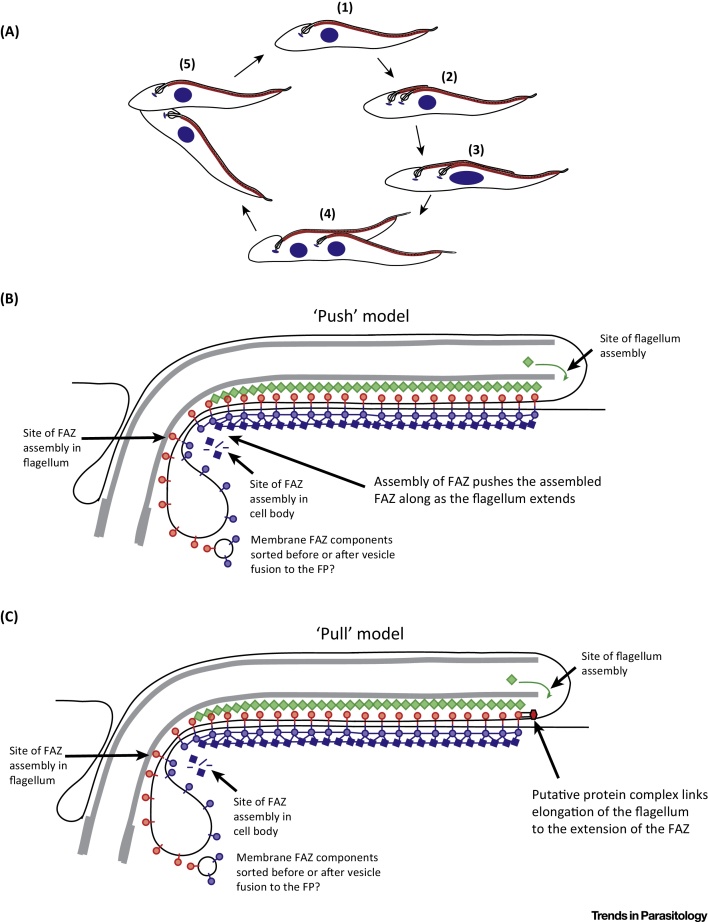
The Flagellum Attachment Zone (FAZ) Assembly ‘Pull’ Model. (A) A simplified schematic of the Trypanosoma brucei cell cycle. The cell begins to assemble a new flagellum and FAZ (highlighted in red) alongside the old one, followed by kinetoplast division (2). The new flagellum continues to extend alongside the old flagellum until the ‘stop point’ is reached after which the new flagellum does not extend along the old flagellum any further (3). After the stop point is reached the two kinetoplasts separate further, followed by mitosis (4) and finally cytokinesis (5). (B) The FAZ assembly ‘push’ model with the assembling FAZ pushing out the assembled FAZ as the flagellum elongates. FAZ proteins are transported to the proximal site of assembly; however, the exact mechanisms that target these components remain to be elucidated. (C) The FAZ assembly ‘pull’ model with a putative protein complex in the flagellum ‘pulling’ out the assembling FAZ.
Our FAZ proximal addition assembly model was confirmed by examining RNAi knock down phenotypes. The model predicts that in a cell where the construction of a new FAZ had begun before initiation of FAZ protein knock down, the distal end of the FAZ structure would be stably assembled; however, the proximal end would be depleted in the targeted FAZ protein. After short RNAi inductions of FLA1 or FAZ5 RNAi, a loop of detached flagellum was observed at the proximal end near the FP, demonstrating that the distal end of the FAZ was stable and the proximal end unstable [30].
The opposite-polarity assembly of the FAZ and flagellum has many implications for the construction of this complex cytoskeletal interface. We focus on:
-
(i)
coordinating assembly between the flagellum and FAZ;
-
(ii)
FAZ protein targeting;
-
(iii)
FAZ and flagellum connections;
-
(iv)
hierarchy and temporal order of assembly;
-
(v)
maintenance of attachment.
Coordination
The addition of LiCl has been shown to lengthen the flagellum in several organisms including trypanosomes and is therefore a useful method for manipulating the flagellum and the FAZ structure. In the presence of LiCl, trypanosomes assembled both a longer flagellum and a longer FAZ [8]. However, disruption of intraflagellum transport (IFT) by RNAi resulted in cells that were unable to assemble a flagellum yet maintained their old flagellum [52]. There was a clear FAZ1 signal associated with the expected position of the new flagellum, although the length of the FAZ was much shorter [52]. Moreover, IFT RNAi also resulted in the generation of a flagellar sleeve – a thin tube of membrane pulled out by the flagella connector – and this sleeve was laterally attached to the cell body via a FAZ [53]. Together these findings suggest that there is coordination between flagellum and FAZ extension and that initial nucleation and extension of the FAZ can occur without an assembling flagellum skeleton but for full-length FAZ assembly an intact flagellum structure is required.
How is flagellum assembly linked to that of the FAZ? As addition of FAZ proteins to the assembling FAZ occurs at the proximal end, two simple models can be envisaged for the forces that coordinate the elongation of the FAZ with the flagellum. First, FAZ subunit addition at the proximal end could ‘push’ the FAZ along as the flagellum extends (Figure 4A,B). Second, a structure associated with the assembling end of the flagellum and connected to the distal end of the FAZ could ‘pull’ the FAZ, allowing new subunits to be added at the proximal end (Figure 4C). In T. brucei the new flagellum extends alongside the old flagellum until it reaches the stop point and then does not extend any further along the old flagellum [53]. It should be noted that the pull force is required only until the stop point has been reached and after this the FAZ remains fixed in position relative to the cell body and the extension of the cell posterior allows the proximal-end assembly of the FAZ to continue (Figure 4A).
Targeting
The FAZ structure is complex, with proteins present in the flagellum, cell body, membranes, and intermembrane spaces, and thus will require targeting or retention mechanisms to ensure correct localisation; it is likely that the components located in the flagellum will have targeting or retention motifs different to those in the cell body.
Moreover, there are proteins associated with the membrane of either the flagellum or the cell body, which will be trafficked through the FP. However, the stage at which these proteins are sorted is not known: does it occur before fusion to the FP, giving rise to two different vesicle populations that then fuse in distinct regions of the FP, or does the sorting occur within the FP? Currently, no targeting signal for any FAZ protein has been fully delineated; however, the short cytoplasmic domain of both FLA1 and FLA1BP was shown to play a role in their localisation to the FAZ [8].
Further to this, how and where is the intracellular domain formed that connects the flagellum and cell body membranes, given that proteins present in zones 2 and 4 will be trafficked differently either to the flagellum or to the cell body? Do the proteins on opposing membranes snap together in the neck region of the FP while being assembled into the FAZ structure?
We suggest that FAZ substructures may be preassembled and then transported to the FP region where they are integrated into the FAZ but accumulate if FAZ or flagellum assembly is disrupted. This would be analogous to desmosome assembly, where preassembled desmosomal complexes are stored in vesicles and then transported to the cell surface when required [54]. Perhaps an accumulation of electron-dense structures at the base of the flagellum in the T. cruzi GP72 knockout cell line represents evidence of this [55].
Connections
The FAZ and flagellum most likely slide over each other as they are assembled and so understanding when the connections between the two structures are made and when they are consolidated is important. The sliding is potentially comparable with the actin/myosin sliding filament model in muscle where overall connectivity is maintained but individual connections are made and broken consecutively. Initially weak connections may be established that are easily broken as the two structures slide over each other, being consolidated only once the structure is mature. Increasing FLAM3 content may be an indication of the strengthening and maturing of the FAZ–flagellum connections [10].
Hierarchy
We know little about the hierarchy and temporal order of FAZ assembly. RNAi depletion of an intracellular domain protein (FAZ5) for a short period of time resulted in detachment of the new flagellum at its proximal end and a FAZ flagellum domain protein (ClpGM6) signal was lost from the detached portion of the flagellum; however, a cell body FAZ protein (FAZ1) signal remained present below the detached section [30]. This suggests that the flagellum FAZ proteins require the membrane FAZ proteins for assembly and that filament assembly is independent of flagellum and membrane FAZ components. In addition we have demonstrated the interdependency of FLAM3/ClpGM6 expression and localisation [10].
Maintenance
The mature flagellum and FAZ must exhibit strong connections between the structures to ensure that the flagellum remains attached despite beating. Flagellum detachment and attachment errors were observed on the knock down of certain flagellar proteins such as calmodulin, KIF9B, and PF16. Depletion of calmodulin or KIF9B results in severe defects in the assembly of an extra-axonemal structure called the paraflagellar rod (PFR), whereas loss of PF16 results in aberrant orientation of the central-pair microtubules 56, 57, 58. KIF9B depletion causes loops of flagellum to become detached, whereas PF16 or calmodulin knock down results in full-length flagellum detachment. KIF9B depletion did not affect FLA1 or FLAM3 addition [7], demonstrating that FAZ assembly had occurred. Cells with detached flagella after calmodulin knock down showed a distinct line of blobs along an indent in the cell surface on scanning electron microscopy. These blobs appear to be flagellum attachment structures, suggesting that the flagellum was initially attached but then became detached [56]. Overall these data suggest that the maintenance of flagellum attachment requires an intact axoneme or PFR skeleton but these structures are not required for FAZ assembly.
FAZ in Other Kinetoplastids
Many kinetoplastids have an attached flagellum and all FAZ proteins identified so far are kinetoplastid specific. FAZ function has mainly been studied using trypanosomes, so the FAZ function in species such as Leishmania that exist as either promastigotes or amastigotes is rather more enigmatic. It is possible that the FAZ functions, as in T. brucei, to define flagellum axial positioning and cytokinesis events [15]. The kinetoplastids are able to adopt a wide variety of different shapes; however, the genomes of these organisms have well-maintained chromosomal synteny and sequence conservation between genes 59, 60, 61. We have shown that the morphological changes that occur during ClpGM6 RNAi indicate that relatively small changes in gene expression can cause major changes in cell shape [9]. Therefore, different trypanosome morphologies are likely to be assembled using the same basic set of building blocks [9].
Concluding Remarks
Here we have argued that the FAZ acts as a cellular ruler for trypanosome morphology and is crucial in determining the overall length of the cell and the positioning of the organelles within it. Furthermore, it is likely that the wide range of kinetoplastid morphologies can be achieved through small modulations in expression of a conserved FAZ gene set rather than requiring a different gene set for each specific morphology. This conclusion has implications for understanding kinetoplastid evolution and cell-type gene expression.
An ever-expanding number of FAZ proteins are being revealed; however, currently this is little more than a list. Looking forwards we need to define subcomplexes and the assembly interdependencies of complexes; this will be greatly aided by localising these proteins into the FAZ zone system defined here and elucidating their targeting or retention mechanisms, and we have highlighted these issues in the Outstanding Questions. This area of trypanosome cell biology is at last providing much-needed insight into how these parasites acquire and morph cell shape and form.
Outstanding Questions.
Where are the known FAZ proteins specifically located within the FAZ zonal structure?
What are the targeting signals that ensure the FAZ proteins are correctly localised?
Which FAZ proteins interact with each other and in what temporal order to orchestrate assembly?
What is the function of the FAZ in kinetoplastid cell types with little flagellum attachment?
How are the FAZ assembly processes coordinated within two different cellular compartments?
Acknowledgments
The authors thank members of the Gull laboratory past and present for fruitful discussions. The Wellcome Trust supports work in the Gull laboratory (WT066839MA, 104627/Z/14/Z). The authors thank Richard Wheeler and Vladimir Varga for help with the cartoons.
References
- 1.Vickerman K. On the surface coat and flagellar adhesion in trypanosomes. J. Cell Sci. 1969;5:163–193. doi: 10.1242/jcs.5.1.163. [DOI] [PubMed] [Google Scholar]
- 2.Sherwin T., Gull K. The cell division cycle of Trypanosoma brucei brucei: timing of event markers and cytoskeletal modulations. Philos. Trans. R. Soc. Lond. B Biol. Sci. 1989;323:573–588. doi: 10.1098/rstb.1989.0037. [DOI] [PubMed] [Google Scholar]
- 3.Woods A. Definition of individual components within the cytoskeleton of Trypanosoma brucei by a library of monoclonal antibodies. J. Cell Sci. 1989;93:491–500. doi: 10.1242/jcs.93.3.491. [DOI] [PubMed] [Google Scholar]
- 4.Vaughan S. A repetitive protein essential for the flagellum attachment zone filament structure and function in Trypanosoma brucei. Protist. 2008;159:127–136. doi: 10.1016/j.protis.2007.08.005. [DOI] [PubMed] [Google Scholar]
- 5.Zhou Q. A coiled-coil- and C2-domain-containing protein is required for FAZ assembly and cell morphology in Trypanosoma brucei. J. Cell Sci. 2011;124:3848–3858. doi: 10.1242/jcs.087676. [DOI] [PubMed] [Google Scholar]
- 6.LaCount D.J. Trypanosoma brucei FLA1 is required for flagellum attachment and cytokinesis. J. Biol. Chem. 2002;277:17580–17588. doi: 10.1074/jbc.M200873200. [DOI] [PubMed] [Google Scholar]
- 7.Rotureau B. Flagellar adhesion in Trypanosoma brucei relies on interactions between different skeletal structures in the flagellum and cell body. J. Cell Sci. 2014;127:204–215. doi: 10.1242/jcs.136424. [DOI] [PubMed] [Google Scholar]
- 8.Sun S.Y. An intracellular membrane junction consisting of flagellum adhesion glycoproteins links flagellum biogenesis to cell morphogenesis in Trypanosoma brucei. J. Cell Sci. 2013;126:520–531. doi: 10.1242/jcs.113621. [DOI] [PubMed] [Google Scholar]
- 9.Hayes P. Modulation of a cytoskeletal calpain-like protein induces major transitions in trypanosome morphology. J. Cell Biol. 2014;206:377–384. doi: 10.1083/jcb.201312067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sunter J.D. Flagellum attachment zone protein modulation and regulation of cell shape in Trypanosoma brucei life cycle transitions. J. Cell Sci. 2015;128:3117–3130. doi: 10.1242/jcs.171645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Hoare C.A., Wallace F.G. Developmental stages of trypanosomatid flagellates: a new terminology. Nature. 1966;212:1385–1386. [Google Scholar]
- 12.Sharma R. The heart of darkness: growth and form of Trypanosoma brucei in the tsetse fly. Trends Parasitol. 2009;25:517–524. doi: 10.1016/j.pt.2009.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Rotureau B. Molecular bases of cytoskeleton plasticity during the Trypanosoma brucei parasite cycle. Cell. Microbiol. 2011;13:705–716. doi: 10.1111/j.1462-5822.2010.01566.x. [DOI] [PubMed] [Google Scholar]
- 14.Lacomble S. Three-dimensional cellular architecture of the flagellar pocket and associated cytoskeleton in trypanosomes revealed by electron microscope tomography. J. Cell Sci. 2009;122:1081–1090. doi: 10.1242/jcs.045740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Robinson D.R. Microtubule polarity and dynamics in the control of organelle positioning, segregation, and cytokinesis in the trypanosome cell cycle. J. Cell Biol. 1995;128:1163–1172. doi: 10.1083/jcb.128.6.1163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gluenz E. The kinetoplast duplication cycle in Trypanosoma brucei is orchestrated by cytoskeleton-mediated cell morphogenesis. Mol. Cell. Biol. 2011;31:1012–1021. doi: 10.1128/MCB.01176-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Robinson D.R., Gull K. Basal body movements as a mechanism for mitochondrial genome segregation in the trypanosome cell cycle. Nature. 1991;352:731–733. doi: 10.1038/352731a0. [DOI] [PubMed] [Google Scholar]
- 18.Ogbadoyi E.O. A high-order trans-membrane structural linkage is responsible for mitochondrial genome positioning and segregation by flagellar basal bodies in trypanosomes. Mol. Biol. Cell. 2003;14:1769–1779. doi: 10.1091/mbc.E02-08-0525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.He C.Y. Golgi duplication in Trypanosoma brucei requires Centrin2. Science. 2005;310:1196–1198. doi: 10.1126/science.1119969. [DOI] [PubMed] [Google Scholar]
- 20.Lacomble S. A Trypanosoma brucei protein required for maintenance of the flagellum attachment zone and flagellar pocket ER domains. Protist. 2012;163:602–615. doi: 10.1016/j.protis.2011.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Field M.C., Carrington M. The trypanosome flagellar pocket. Nat. Rev. Microbiol. 2009;7:775–786. doi: 10.1038/nrmicro2221. [DOI] [PubMed] [Google Scholar]
- 22.Gadelha C. Membrane domains and flagellar pocket boundaries are influenced by the cytoskeleton in African trypanosomes. Proc. Natl. Acad. Sci. U.S.A. 2009;106:17425–17430. doi: 10.1073/pnas.0909289106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Grab D.J. The transferrin receptor in African trypanosomes: identification, partial characterization and subcellular localization. Eur. J. Cell Biol. 1993;62:114–126. [PubMed] [Google Scholar]
- 24.Wheeler R.J. The limits on trypanosomatid morphological diversity. PLoS ONE. 2013;8:e79581. doi: 10.1371/journal.pone.0079581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Engstler M. Hydrodynamic flow-mediated protein sorting on the cell surface of trypanosomes. Cell. 2007;131:505–515. doi: 10.1016/j.cell.2007.08.046. [DOI] [PubMed] [Google Scholar]
- 26.Nekrasova O., Green K.J. Desmosome assembly and dynamics. Trends Cell Biol. 2013;23:537–546. doi: 10.1016/j.tcb.2013.06.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Neumann M., Affolter M. Remodelling epithelial tubes through cell rearrangements: from cells to molecules. EMBO Rep. 2006;7:36–40. doi: 10.1038/sj.embor.7400597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nekrasova O.E. Desmosomal cadherins utilize distinct kinesins for assembly into desmosomes. J. Cell Biol. 2011;195:1185–1203. doi: 10.1083/jcb.201106057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nozaki T. Characterization of the Trypanosoma brucei homologue of a Trypanosoma cruzi flagellum-adhesion glycoprotein. Mol. Biochem. Parasitol. 1996;82:245–255. doi: 10.1016/0166-6851(96)02741-7. [DOI] [PubMed] [Google Scholar]
- 30.Sunter J.D. A dynamic coordination of flagellum and cytoplasmic cytoskeleton assembly specifies cell morphogenesis in trypanosomes. J. Cell Sci. 2015;128:1580–1594. doi: 10.1242/jcs.166447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McAllaster M.R. Proteomic identification of novel cytoskeletal proteins associated with TbPLK, an essential regulator of cell morphogenesis in T. brucei. Mol. Biol. Cell. 2015;26:3013–3039. doi: 10.1091/mbc.E15-04-0219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Subota I. ALBA proteins are stage regulated during trypanosome development in the tsetse fly and participate in differentiation. Mol. Biol. Cell. 2011;22:4205–4219. doi: 10.1091/mbc.E11-06-0511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Haynes P.A. Structural characterization of novel oligosaccharides of cell-surface glycoproteins of Trypanosoma cruzi. Glycobiology. 1996;6:869–878. doi: 10.1093/glycob/6.8.869. [DOI] [PubMed] [Google Scholar]
- 34.Snary D. Cell surface antigens of Trypanosoma cruzi: use of monoclonal antibodies to identify and isolate an epimastigote specific glycoprotein. Mol. Biochem. Parasitol. 1981;3:343–356. doi: 10.1016/0166-6851(81)90035-9. [DOI] [PubMed] [Google Scholar]
- 35.Ferguson M.A. Studies on the structure of a phosphoglycoprotein from the parasitic protozoan Trypanosoma cruzi. Biochem. J. 1983;213:313–319. doi: 10.1042/bj2130313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ferguson M.A. Comparative compositions of cell surface glycoconjugates isolated from Trypanosoma cruzi epimastigotes. Biochim. Biophys. Acta. 1985;842:39–44. doi: 10.1016/0304-4165(85)90290-9. [DOI] [PubMed] [Google Scholar]
- 37.Allen S. Structure of a complex phosphoglycan epitope from gp72 of Trypanosoma cruzi. J. Biol. Chem. 2013;288:11093–11105. doi: 10.1074/jbc.M113.452763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.de Jesus A.R. Gene deletion suggests a role for Trypanosoma cruzi surface glycoprotein GP72 in the insect and mammalian stages of the life cycle. J. Cell Sci. 1993;106:1023–1033. doi: 10.1242/jcs.106.4.1023. [DOI] [PubMed] [Google Scholar]
- 39.Cooper R. Deletion of an immunodominant Trypanosoma cruzi surface glycoprotein disrupts flagellum–cell adhesion. J. Cell Biol. 1993;122:149–156. doi: 10.1083/jcb.122.1.149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Haynes P.A. Subcellular localization of Trypanosoma cruzi glycoprotein Gp72. J. Cell Sci. 1996;109:2979–2988. doi: 10.1242/jcs.109.13.2979. [DOI] [PubMed] [Google Scholar]
- 41.Woods A. Evidence for a Mr 88,000 glycoprotein with a transmembrane association to a unique flagellum attachment region in Trypanosoma brucei. J. Cell Sci. 1989;93:501–508. doi: 10.1242/jcs.93.3.501. [DOI] [PubMed] [Google Scholar]
- 42.Turnock D.C. The de novo synthesis of GDP-fucose is essential for flagellar adhesion and cell growth in Trypanosoma brucei. J. Biol. Chem. 2007;282:28853–28863. doi: 10.1074/jbc.M704742200. [DOI] [PubMed] [Google Scholar]
- 43.Woods K. Identification and characterization of a stage specific membrane protein involved in flagellar attachment in Trypanosoma brucei. PLoS ONE. 2013;8:e52846. doi: 10.1371/journal.pone.0052846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Slack F.J., Ruvkun G. A novel repeat domain that is often associated with RING finger and B-box motifs. Trends Biochem. Sci. 1998;23:474–475. doi: 10.1016/s0968-0004(98)01299-7. [DOI] [PubMed] [Google Scholar]
- 45.Portman N., Gull K. Oxford University Press; 2011. Deconstructing the Trypanosome Cytoskeleton: From Structures to Functions via Components and Complexes. [Google Scholar]
- 46.Urbaniak M.D. Comparative SILAC proteomic analysis of Trypanosoma brucei bloodstream and procyclic lifecycle stages. PLoS ONE. 2012;7:e36619. doi: 10.1371/journal.pone.0036619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhou Q. Assembly and maintenance of the flagellum attachment zone filament in Trypanosoma brucei. J. Cell Sci. 2015;128:2361–2372. doi: 10.1242/jcs.168377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hu H. SAS-4 in Trypanosoma brucei controls life cycle transitions by modulating the length of the flagellum attachment zone filament. J. Biol. Chem. 2015 doi: 10.1074/jbc.M115.694109. Published online October 26, 2015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Sun L., Wang C.C. The structural basis of localizing polo-like kinase to the flagellum attachment zone in Trypanosoma brucei. PLoS ONE. 2011;6:e27303. doi: 10.1371/journal.pone.0027303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Ikeda K.N., de Graffenried C.L. Polo-like kinase is necessary for flagellum inheritance in Trypanosoma brucei. J. Cell Sci. 2012;125:3173–3184. doi: 10.1242/jcs.101162. [DOI] [PubMed] [Google Scholar]
- 51.Li Z. Identification of a novel chromosomal passenger complex and its unique localization during cytokinesis in Trypanosoma brucei. PLoS ONE. 2008;3:e2354. doi: 10.1371/journal.pone.0002354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kohl L. Novel roles for the flagellum in cell morphogenesis and cytokinesis of trypanosomes. EMBO J. 2003;22:5336–5346. doi: 10.1093/emboj/cdg518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Davidge J.A. Trypanosome IFT mutants provide insight into the motor location for mobility of the flagella connector and flagellar membrane formation. J. Cell Sci. 2006;119:3935–3943. doi: 10.1242/jcs.03203. [DOI] [PubMed] [Google Scholar]
- 54.Demlehner M.P. Continual assembly of half-desmosomal structures in the absence of cell contacts and their frustrated endocytosis: a coordinated Sisyphus cycle. J. Cell Biol. 1995;131:745–760. doi: 10.1083/jcb.131.3.745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Rocha G.M. The flagellar attachment zone of Trypanosoma cruzi epimastigote forms. J. Struct. Biol. 2006;154:89–99. doi: 10.1016/j.jsb.2005.11.008. [DOI] [PubMed] [Google Scholar]
- 56.Ginger M.L. Calmodulin is required for paraflagellar rod assembly and flagellum–cell body attachment in trypanosomes. Protist. 2013;164:528–540. doi: 10.1016/j.protis.2013.05.002. [DOI] [PubMed] [Google Scholar]
- 57.Demonchy R. Kinesin 9 family members perform separate functions in the trypanosome flagellum. J. Cell Biol. 2009;187:615–622. doi: 10.1083/jcb.200903139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Branche C. Conserved and specific functions of axoneme components in trypanosome motility. J. Cell Sci. 2006;119:3443–3455. doi: 10.1242/jcs.03078. [DOI] [PubMed] [Google Scholar]
- 59.Berriman M. The genome of the African trypanosome Trypanosoma brucei. Science. 2005;309:416–422. doi: 10.1126/science.1112642. [DOI] [PubMed] [Google Scholar]
- 60.El-Sayed N.M. The genome sequence of Trypanosoma cruzi, etiologic agent of Chagas disease. Science. 2005;309:409–415. doi: 10.1126/science.1112631. [DOI] [PubMed] [Google Scholar]
- 61.Ivens A.C. The genome of the kinetoplastid parasite, Leishmania major. Science. 2005;309:436–442. doi: 10.1126/science.1112680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Oberholzer M. Independent analysis of the flagellum surface and matrix proteomes provides insight into flagellum signaling in mammalian-infectious Trypanosoma brucei. Mol. Cell. Proteomics. 2011;10 doi: 10.1074/mcp.M111.010538. M111.010538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Morriswood B. Novel bilobe components in Trypanosoma brucei identified using proximity-dependent biotinylation. Eukaryot. Cell. 2013;12:356–367. doi: 10.1128/EC.00326-12. [DOI] [PMC free article] [PubMed] [Google Scholar]