Abstract
Background
Interleukin-10 (IL-10) is a multifunctional regulatory cytokine that might be associated with increased risk of type 2 diabetes mellitus (T2DM). IL-10 gene polymorphisms have been reported to be associated with T2DM in several ethnic populations with controversial results.
Objectives
This work is an updated meta-analysis aiming at the evaluation of the association between IL-10 gene polymorphisms: rs1800872 (− 592 C > A), rs1800896 (− 1082 A > G) and rs1800871 (− 819 C > T) with the risk of T2DM.
Methods
All available full text studies published up to July 2015 were included in this meta-analysis. Mainly Pubmed and Science Direct databases were searched for all eligible studies pertinent to testing the association between IL-10 gene polymorphisms with the susceptibility to T2DM. Further analyses of the pooled and stratified data in terms of individual polymorphic types and subject ethnicity were done and assessed using varied genetic models.
Results
Fifteen case-control studies with a total of 26 comparisons (10 for IL-10 − 592 C > A rs1800872, 11 for IL-10 − 1082 A > G rs1800896 and 5 for IL-10 − 819 C > T rs1800871 polymorphisms) met our inclusion criteria. IL-10 − 1082 A > G polymorphism was the only one to show an association with T2DM in all pooled sample particularly among Asian and European (high frequency of the G allele) ethnic groups. On the other hand, IL-10 − 592 C > A and − 819 C > T were significantly associated with T2DM only among African subjects.
Conclusions
This meta-analysis demonstrated that IL-10 − 1082 A > G polymorphism was associated with increased risk of development of T2DM in total subjects no matter was their ethnic background, while both IL-10 − 592 C > A and − 819 C > T polymorphisms were associated with that risk only among African subjects.
Keywords: Meta-analysis, T2DM, Gene, Polymorphism, IL-10
1. Introduction
Type 2 diabetes mellitus (T2DM) is a complex disease characterized by progressive β-cell dysfunction with insulin resistance and impaired insulin secretion (Steyn, NP, et al., 2009, Lin, JD, et al., 2010). The basic etiologic pathology of T2DM is speculated to involve multiple factors including genetic, environmental and immune ones (Pickup, 2004).One of the important cytokines with a probable major role is the interleukin-10 (IL-10) known of its multifunctional role in the inflammatory response of immune disorders (van Exel, E, et al., 2002, Del Prete, G, et al., 1993). The IL-10 gene is located on chromosome 1 (1q31–1q32) which is composed of five exons (Eskdale et al., 1997). Polymorphisms implicated to affect IL-10 transcription and secretion include rs1800872 (− 592 C > A), rs1800896 (− 1082 A > G) and rs1800871 (− 819 C > T) (Eskdale, J, et al., 1997, Kilpinen, S, et al., 2002, Li, J, et al., 2013).
Although several studies and few meta-analyses have evaluated the association between IL-10 gene polymorphisms with the risk of development of T2DM in different ethnic populations, their results showed controversies and inconsistencies that might be due relatively small sample size, selection bias and ethnic difference (Bai, H, et al., 2014, Helaly, MA and Hatata, EZ, 2013, Kung, WJ, et al., 2010, Yin, YW, et al., 2012, Li, J, et al., 2013, Zhang, F, et al., 2013, Mtiraoui, N, et al., 2009). Another issue of concern is the under-representation of African-based studies. For this reason, we planned this meta-analysis to have an updated evidence based testing for the association of IL-10 gene polymorphisms with the increased risk of T2DM in a pooled sample of subjects of European, Asian as well African origins.
2. Methods
2.1. Study identification and selection
This meta-analysis followed the preferred reporting items for systematic reviews and meta-analyses (PRISMA) criteria (Moher et al., 2010). A literature search was conducted using Pubmed and Science Direct citation databases to identify articles (up to July 2015) that examined the association between IL-10 gene (− 592 C > A, − 1082 A > G and − 819 C > T) polymorphisms and risk of T2DM. Combinations of keywords such as: [“interleukin-10”, “IL-10”, “− 1082 A > G”, “− 592 C > A”, “− 819 C > T”, “polymorphism”, “variant”, “type 2 diabetes mellitus” and “T2DM”] were entered as both Medical Subject Headings (MeSHs) and text words without any restrictions on language or country.
2.2. Inclusion and exclusion criteria
Inclusion criteria were defined as follows: (a) full text articles evaluating the association between genetic polymorphisms of IL-10 (− 592 C > A, − 1082 A > G and − 819 C > T) with the risk of T2DM, (b) the design is a case–controlled study based on unrelated individuals, and (c) sufficient data (genotype distributions for cases and controls) available to estimate an odds ratio (OR) with its 95% confidence interval (CI). Studies were excluded if one of the following existed: (a) reviews and abstracts of meetings or conferences; (b) studies containing overlapping data; (c) studies in which the genotype frequencies or numbers could not be ascertained and (d) studies in which family members were studied, because their analysis is based on linkage considerations. If more than one article were published by the same authors using the same sample series, studies with the recently published ones were included.
2.3. Data extraction
The following information was carefully extracted from all eligible publications independently by two investigators: (1) name of the first author; (2) year of publication; (3) country of origin; (4) ethnicity of the studied population; (5) number of cases and controls assigned to certain polymorphism of IL-10 (− 592 C > A, − 1082 A > G and − 819 C > T); (6) genotyping method and (7) Hardy–Weinberg equilibrium (HWE) for controls.
2.4. Statistical analysis
The meta-analysis examined the comparisons between T2DM patients and healthy controls. The Hardy–Weinberg equilibrium (HWE) was evaluated in control groups by chi square test. If the study was found not to be in HWE with p value less than 0.05, it was considered to be in disequilibrium. Allele frequencies of the IL-10 gene (− 592 C > A, − 1082 A > G and − 819 C > T) polymorphisms in each of the studies were determined using the allelic counting method. The strength of association was evaluated by examining the pooled odd ratios (ORs) and their 95% confidence intervals (CIs) for each study while, within- and between-study heterogeneity were assessed using Cochran's Q statistic. The genetic models evaluated for pooled ORs of these three polymorphisms have included allelic contrast, recessive model, dominant model, overdominant model, homozygote contrast and heterozygote contrast. The heterogeneity test was used to assess the probability of the null hypothesis that all studies were evaluating the same effect. The random-effects model was used for meta-analysis when a significant Q statistic (p < 0.10) indicated heterogeneity across studies (DerSimonian and Laird, 1986), while the fixed-effect model was used when heterogeneity was not significant (Mantel and Haenszel, 1959). We quantified the effect of heterogeneity by using the recently developed I2measure, where I2 = 100% × (Q − df)/Q (Higgins and Thompson, 2002). The I2measure ranges between 0 and 100%, and it represents the proportion of inter-study variability attributable to heterogeneity rather than chance. I2 values of 25, 50, and 75% were defined as low, moderate, and high estimates, respectively. Funnel plot and Egger's linear regression test were used to evaluate publication bias (Egger et al., 1997), and a P value < 0.1 was considered statistically significant. Statistical manipulations were performed using the comprehensive meta-analysis computer program (Biosta, Englewood, NJ, USA).
3. Results
3.1. Meta-analysis of − 592 C > A and T2DM susceptibility
In total, 10 studies including 2921 T2DM cases and 3316 controls investigating the relationship between the IL-10 − 592 C > A polymorphism and the development of T2DM were available and met the criteria for this meta-analysis (Fig. 1A). The detailed characteristics of these studies were shown in (Table 1). Among these, six studies were performed among Asians (Wang, JD, et al., 2010, Kung, WJ, et al., 2010, Chang, YH, et al., 2005, Arababadi, MK, et al., 2012, Saxena, M, et al., 2013, Bai, H, et al., 2014); two studies among Europeans (Tsiavou, A, et al., 2004, Scarpelli, D, et al., 2006); one study among Africans (Mtiraoui et al., 2009), and one among Mexicans (García-Elorriaga et al., 2013). The overall frequencies of IL-10 − 592 C > A gene polymorphism in control subjects was consistent with Hardy–Weinberg equilibrium (HWE) in spite of the presence of 4 non-consistent studies (Wang, JD, et al., 2010, Kung, WJ, et al., 2010, García-Elorriaga, G, et al., 2013, Saxena, M, et al., 2013). The results of Eggers test suggested no publication bias for all comparisons (Fig.5A and Table 2).
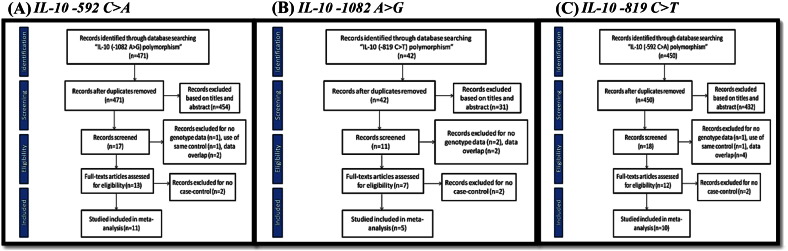
Fig. 1.
Flow chart of studies of (A) IL-10 − 592 C > A, (B) IL-10 − 1082 A > G and (C) IL-10 − 819 C > T polymorphisms in the meta-analysis.
Table 1.
Characteristics of studies of IL-10 gene polymorphisms included in the meta-analysis.
| First author [Ref] | Year | Country | Ethnicity | Sample size |
Genotyping methods | Cases/controls (genotype) | Case/control (allele) | P (HWE) controls | ||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cases | Controls | |||||||||||
| − 592 A/C | CC | AC | AA | C | A | |||||||
| Bai et al. | 2014 | China | Asian | 364 | 677 | Mass array | 153/313 | 162/299 | 49/65 | 468/925 | 260/429 | 0.59 |
| Saxena et al. | 2013 | India | Asian | 213 | 140 | PCR-RFLP | 116/62 | 87/72 | 10/6 | 319/196 | 107/84 | 0.008⁎ |
| García-Elorriaga et al. | 2013 | Mexico | Mexican | 21 | 47 | PCR-RFLP | 11/13 | 9/34 | 1/0 | 31/60 | 11/34 | < 0.001⁎ |
| Arababadi et al. | 2012 | Iran | Asian | 200 | 100 | PCR-RFLP | 107/22 | 83/55 | 10/23 | 297/99 | 103/101 | 0.32 |
| Wang et al. | 2010 | China | Asian | 224 | 275 | PCR-RFLP | 66/113 | 122/138 | 36/24 | 254/364 | 194/186 | 0.04⁎ |
| Kung et al. | 2010 | Taiwan | Asian | 47 | 25 | PCR-RFLP | 4/1 | 36/24 | 7/0 | 44/26 | 50/24 | < 0.001⁎ |
| Mtiraoui et al. | 2009 | Tunisia | African | 917 | 748 | PCR-ASA | 425/403 | 395/298 | 97/47 | 1245/1104 | 589/392 | 0.41 |
| Scarpelli et al. | 2006 | Italy | European | 551 | 1131 | Sequencing | 301/615 | 226/449 | 24/67 | 828/1679 | 274/583 | 0.21 |
| Chang et al. | 2005 | China | Asian | 353 | 134 | PCR-RFLP | 42/10 | 158/52 | 153/72 | 242/72 | 464/196 | 0.88 |
| Tsiavou et al. | 2004 | Greece | European | 31 | 39 | PCR-SSP | 17/19 | 13/18 | 1/2 | 47/56 | 15/22 | 0.38 |
| − 1082G/A | AA | GA | GG | A | G | |||||||
| Bai et al. | 2014 | China | Asian | 364 | 677 | Mass array | 252/495 | 72/129 | 40/53 | 576/1119 | 152/235 | < 0.001⁎ |
| García-Elorriaga et al. | 2013 | Mexico | Mexican | 21 | 47 | PCR-RFLP | 6/4 | 9/18 | 6/25 | 21/26 | 21/68 | 0.77 |
| Helaly et al. | 2013 | Egypt | African | 69 | 98 | ARMS-PCR | 2/8 | 41/85 | 26/5 | 45/101 | 93/95 | < 0.001⁎ |
| Erdogan et al. | 2012 | Turkey | European | 91 | 112 | PCR-RFLP | 22/44 | 69/54 | 0/14 | 113/142 | 69/82 | 0.68 |
| Forte et al. | 2010 | Italy | European | 490 | 349 | PCR | 161/118 | 228/176 | 101/55 | 550/412 | 430/286 | 0.43 |
| Kung et al. | 2010 | Taiwan | Asian | 47 | 25 | PCR-RFLP | 2/0 | 45/25 | 0/0 | 49/25 | 45/25 | < 0.001⁎ |
| Mtiraouli et al. | 2009 | Tunisia | African | 917 | 748 | PCR-ASA | 121/106 | 426/326 | 370/316 | 668/538 | 1166/958 | 0.14 |
| Kolla et al. | 2009 | India | Asian | 198 | 202 | PCR | 124/148 | 42/41 | 32/13 | 290/337 | 106/67 | 0.002⁎ |
| Scarpelli et al. | 2006 | Italy | European | 551 | 1131 | sequencing | 219/485 | 264/516 | 68/130 | 702/1486 | 400/776 | 0.68 |
| Babel et al. | 2006 | Germany | European | 44 | 114 | PCR-SSP | 12/30 | 8/42 | 24/42 | 32/126 | 56/46 | 0.006⁎ |
| Tsiavou et al. | 2004 | Greece | European | 31 | 39 | PCR-SSP | 10/17 | 13/17 | 8/5 | 33/51 | 29/27 | 0.82 |
| − 819 T/C | CC | TC | TT | C | T | |||||||
| Bai et al. | 2014 | China | Asian | 364 | 677 | Mass array | 151/295 | 183/336 | 30/46 | 485/926 | 243/428 | < 0.001⁎ |
| Kung et al. | 2010 | Taiwan | Asian | 47 | 25 | PCR-RFLP | 0/1 | 47/24 | 0/0 | 47/26 | 47/24 | < 0.001⁎ |
| Mtiraouli et al. | 2009 | Tunisia | African | 402 | 748 | PCR-ASA | 199/488 | 173/228 | 30/32 | 571/1204 | 233/292 | 0.41 |
| Chang et al. | 2005 | Taiwan | Asian | 370 | 175 | PCR-RFLP | 41/13 | 159/71 | 170/91 | 241/97 | 499/253 | 0.87 |
| Tsiavou et al. | 2004 | Greece | European | 31 | 39 | PCR-SSP | 17/19 | 13/18 | 1/2 | 47/56 | 15/22 | 0.38 |
Asterisk indicate significant P of HWE, and it is deviation from this equation.
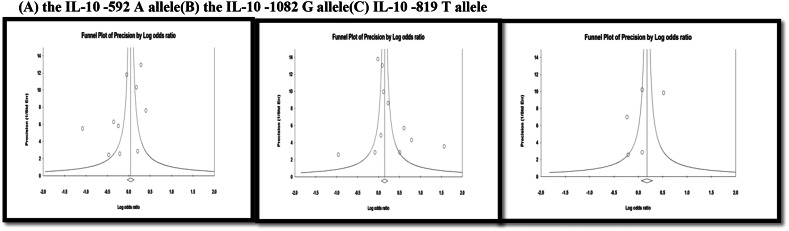
Fig. 5.
Funnel plot for the association between (A) the IL-10 − 592 A allele, (B) the IL-10 − 1082 G allele and (C) IL-10 − 819 T allele with T2DM in all study subjects (Egger's regression P value = 0.188, 0.277 and 0.544, respectively).
Table 2.
Meta-analysis of the association between IL-10-592 C > A polymorphism and T2DM.
| Comparison | Population | Sample size |
No. of studies | Test of association |
Test of heterogeneity |
Publication bias |
||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T2DM | Control | OR | 95% CI | P-value | Model | Q test | P-value | I2 (%) | P-value (Egger's) | |||
| A allele versus C allele (allelic contrast) |
Overall | 5842 | 6632 | 10 | 0.901 | 0.700–1.161 | 0.421 | R | 69.82 | < 0.001 | 87.11 | 0.188 |
| Asian | 2802 | 2702 | 6 | 0.859 | 0.565–1.304 | 0.475 | R | 54.90 | < 0.001 | 90.89 | 0.398 | |
| European | 1164 | 2340 | 2 | 0.946 | 0.805–1.113 | 0.504 | F | 0.161 | 0.688 | 0.0 | NA | |
| African | 1834 | 1496 | 1 | 1.332 | 1.145–1.550 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
| AC + AA versus CC (dominant model) |
Overall | 2921 | 3316 | 10 | 1.106 | 0.967–1.265 | 0.140 | R | 52.37 | < 0.001 | 82.81 | 0.08 |
| Asian | 1401 | 1351 | 6 | 0.725 | 0.410–1.283 | 0.270 | R | 39.67 | < 0.001 | 87.40 | 0.289 | |
| European | 582 | 1170 | 2 | 0.980 | 0.802–1.196 | 0.840 | F | 0.227 | 0.634 | 0.0 | NA | |
| African | 917 | 748 | 1 | 1.352 | 1.114–1.641 | 0.002 | F | 0.0 | 1.0 | 0.0 | NA | |
| AA versus CC + AC (recessive model) |
Overall | 2921 | 3316 | 10 | 1.225 | 0.940–1.596 | 0.134 | R | 46.68 | < 0.001 | 80.72 | 0.935 |
| Asian | 1401 | 1351 | 6 | 0.945 | 0.466–1.917 | 0.876 | R | 34.85 | < 0.001 | 85.65 | 0.998 | |
| European | 582 | 1170 | 2 | 0.719 | 0.450–1.149 | 0.168 | F | 0.016 | 0.900 | 0.0 | NA | |
| African | 917 | 748 | 1 | 1.764 | 1.228–2.536 | 0.002 | F | 0.0 | 1.0 | 0.0 | NA | |
| AC versus CC (heterozygote contrast) |
Overall | 2533 | 3010 | 10 | 1.074 | 0.936–1.232 | 0.310 | R | 35.47 | < 0.001 | 74.62 | 0.053 |
| Asian | 1136 | 1161 | 6 | 0.760 | 0.471–1.227 | 0.261 | R | 25.47 | < 0.001 | 80.37 | 0.291 | |
| European | 557 | 1101 | 2 | 1.017 | 0.828–1.250 | 0.871 | F | 0.230 | 0.632 | 0.0 | NA | |
| African | 820 | 701 | 1 | 1.257 | 1.026–1.540 | 0.027 | F | 0.0 | 1.0 | 0.0 | NA | |
| AA versus CC (homozygote contrast) |
Overall | 1630 | 1877 | 10 | 1.297 | 0.979–1.718 | 0.070 | R | 59.05 | < 0.001 | 84.76 | 0.462 |
| Asian | 753 | 711 | 6 | 0.806 | 0.298–2.181 | 0.671 | R | 47.01 | < 0.001 | 89.36 | 0.556 | |
| European | 343 | 703 | 2 | 0.725 | 0.450–1.168 | 0.186 | F | 0.044 | 0.835 | 0.0 | NA | |
| African | 522 | 450 | 1 | 1.957 | 1.346–2.845 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
| AC versus CC + AA (overdominant model) |
Overall | 2921 | 3316 | 10 | 1.035 | 0.912–1.174 | 0.598 | R | 21.66 | 0.010 | 58.45 | 0.018 |
| Asian | 1401 | 1351 | 6 | 0.888 | 0.659–1.196 | 0.433 | R | 13.99 | 0.016 | 64.28 | 0.170 | |
| European | 582 | 1170 | 2 | 1.045 | 0.854–1.280 | 0.667 | F | 0.207 | 0.649 | 0.0 | NA | |
| African | 917 | 748 | 1 | 1.143 | 0.939–1.390 | 0.183 | F | 0.0 | 1.0 | 0.0 | NA | |
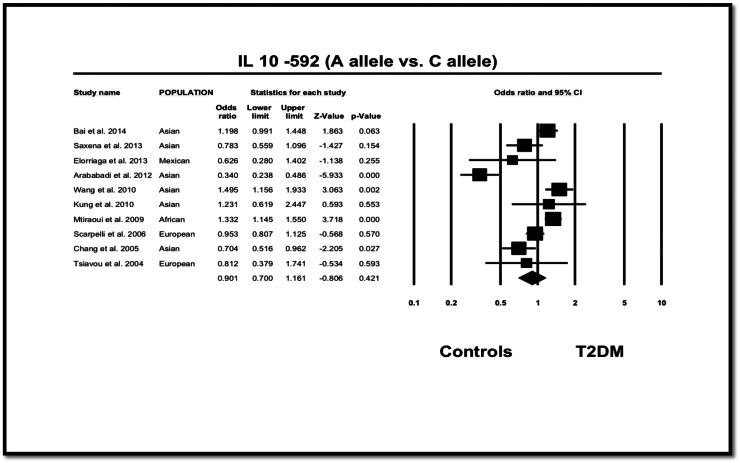
Analysis showed no significant differences between cases and controls regarding the allelic (OR = 0.901, 95% CI = 0.700–0.161, p = 0.421), dominant (OR = 1.106, 95% CI = 0.967–1.265, p = 0.140) and recessive (OR = 1.225, 95% CI = 0.940–1.596, p = 0.134) (Fig. 2) models. However, analysis based on the ethnic distribution of subjects, it was found that the IL-10 − 592 A allele was only positively associated with T2DM among African population (OR = 1.332, 95% CI = 1.145–1.550, p < 0.001), which was also noted in the dominant, recessive, heterozygote and homozygote models (Table 2, Fig. 2).
Fig. 2.
OR and 95% CI for individual studies and pooled data for the association of IL-10-592 A versus C allele.
3.2. Meta-analysis of IL 10 − 1082 A > G and T2DM susceptibility
In total, 11 studies including 2823 T2DM cases and 3542 controls examined the relationship between the IL-10 − 1082 A > G polymorphism and the development of T2DM were available and met the criteria for this meta-analysis (Fig. 1B). The detailed characteristics of the studies included were shown in (Table 1). Among these, five studies were performed among Europeans (Tsiavou, A, et al., 2004, Babel, N, et al., 2006, Scarpelli, D, et al., 2006, Forte, GI, et al., 2010, Erdogan, M, et al., 2012), three studies were performed among Asians (Kolla, VK, et al., 2009, Kung, WJ, et al., 2010 and Bai et al., 2014), two studies were performed among Africans (Mtiraoui et al., 2009 and Helaly and Hatata, 2013), and one among Mexicans (García-Elorriaga et al., 2013). The overall frequencies of IL-10 − 1082 A > G gene polymorphism in control subjects was consistent with Hardy–Weinberg equilibrium (HWE) in spite of the presence of 5 non-consistent studies (Bai, H, et al., 2014, Helaly, MA and Hatata, EZ, 2013, Kung, WJ, et al., 2010, Kolla, VK, et al., 2009, Babel, N, et al., 2006). The results of Eggers test suggested no publication bias for all comparisons (Fig. 5B, Table 3).
Table 3.
Meta-analysis of the association between IL-10 − 1082 A > G polymorphism and T2D.
| Comparison | Population | Sample size |
No. of studies | Test of association |
Test of heterogeneity |
Publication bias |
||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T2DM | Control | OR | 95% CI | P-value | Model | Q test | P-value | I2 (%) | P-value (Egger's) | |||
| G allele versus A allele (allelic contrast) |
Overall | 5646 | 7028 | 11 | 1.309 | 1.055–1.623 | 0.015 | R | 56.66 | < 0.001 | 82.35 | 0.227 |
| European | 2414 | 3434 | 5 | 1.474 | 1.030–2.108 | 0.034 | R | 27.44 | < 0.001 | 85.42 | 0.213 | |
| Asian | 1218 | 1808 | 3 | 1.370 | 0.986–1.904 | 0.061 | R | 4.67 | 0.097 | 57.18 | 0.914 | |
| African | 1972 | 1692 | 2 | 1.424 | 0.647–3.135 | 0.379 | R | 11.10 | 0.001 | 90.99 | NA | |
| AG + GG versus AA (dominant model) |
Overall | 2823 | 3542 | 11 | 1.174 | 1.044–1.320 | 0.008 | F | 14.24 | 0.162 | 29.79 | 0.963 |
| European | 1207 | 1745 | 5 | 1.155 | 0.987–1.352 | 0.072 | F | 4.45 | 0.350 | 9.86 | 0.407 | |
| Asian | 609 | 904 | 3 | 1.315 | 1.042–1.660 | 0.021 | F | 2.055 | 0.358 | 2.655 | 0.801 | |
| African | 986 | 846 | 2 | 1.120 | 0.850–1.477 | 0.421 | F | 1.515 | 0.218 | 33.97 | NA | |
| GG versus AA + AG (recessive model) |
Overall | 2776 | 3517 | 10 | 1.472 | 1.001–2.165 | 0.049 | R | 46.69 | < 0.001 | 80.72 | 0.337 |
| European | 1207 | 1745 | 5 | 1.338 | 0.857–2.088 | 0.200 | R | 9.96 | 0.041 | 59.82 | 0.792 | |
| Asian | 562 | 879 | 2 | 1.757 | 1.221–2.529 | 0.002 | F | 2.56 | 0.109 | 61.02 | NA | |
| African | 986 | 846 | 2 | 3.060 | 0.265–35.32 | 0.370 | R | 22.09 | < 0.001 | 95.47 | NA | |
| AG versus AA (heterozygote contrast) |
Overall | 2148 | 2884 | 11 | 1.118 | 0.984–1.270 | 0.086 | F | 14.32 | 0.159 | 30.19 | 0.699 |
| European | 1006 | 1499 | 5 | 1.158 | 0.810–1.653 | 0.421 | R | 10.67 | 0.031 | 62.51 | 0.898 | |
| Asian | 537 | 838 | 3 | 1.123 | 0.857–1.472 | 0.400 | F | 0.670 | 0.715 | 0.0 | 0.490 | |
| African | 590 | 525 | 2 | 1.165 | 0.869–1.562 | 0.306 | F | 0.398 | 0.528 | 0.0 | NA | |
| GG versus AA (homozygote contrast) |
Overall | 1604 | 2113 | 10 | 1.387 | 0.957–2.011 | 0.084 | R | 29.72 | < 0.001 | 69.72 | 0.716 |
| European | 625 | 940 | 5 | 1.252 | 0.983–1.594 | 0.069 | F | 5.63 | 0.228 | 29.01 | 0.710 | |
| Asian | 448 | 709 | 2 | 1.806 | 1.248–2.612 | 0.002 | F | 2.71 | 0.100 | 63.05 | NA | |
| African | 519 | 435 | 2 | 4.017 | 0.213–75.73 | 0.353 | R | 10.21 | 0.001 | 90.21 | NA | |
| AG versus AA + GG (overdominant model) |
Overall | 2823 | 3542 | 11 | 0.964 | 0.739–1.256 | 0.784 | R | 38.97 | < 0.001 | 74.34 | 0.439 |
| European | 1207 | 1745 | 5 | 1.072 | 0.677–1.689 | 0.767 | R | 21.96 | < 0.001 | 81.79 | 0.961 | |
| Asian | 609 | 904 | 3 | 1.042 | 0.798–1.360 | 0.763 | F | 0.471 | 0.790 | 0.0 | 0.225 | |
| African | 986 | 846 | 2 | 0.524 | 0.108–2.537 | 0.422 | R | 16.39 | < 0.001 | 93.90 | NA | |
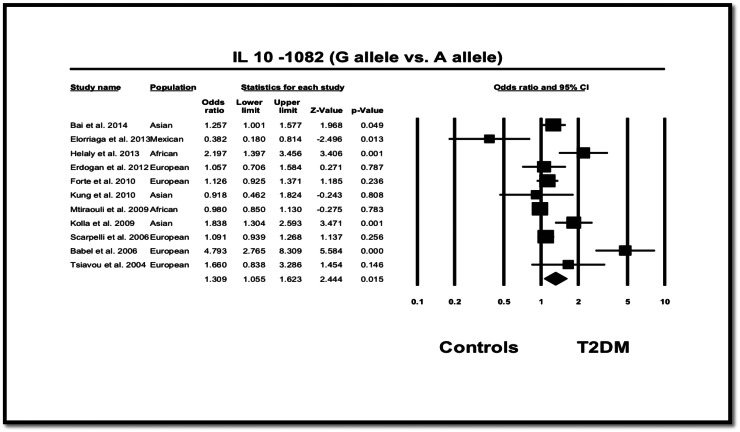
Overall, a significant association between the − 1082 A > G polymorphism and T2DM was found regarding the allelic contrast model (G vs. A allele, OR = 1.309, 95% CI: 1.055–1.623, p = 0.015), dominant model (OR = 1.174, 95% CI: 1.044–1.320, p = 0.008) and recessive model (OR = 1.472, 95% CI: 1.001–2.165, p = 0.049) (Fig. 3). Stratified analyses showed that significant associations were found in European population for the allelic contrast model (ORs of 1.474 and 95% CI = 1.030–2.108, p = 0.034); and in Asian populations for the dominant model (OR = 1.315 and 95% CI = 1.042–1.660, p = 0.021), recessive model (OR = 1.757 and 95% CI = 1.221–2.529, p = 0.002) and homozygote contrast model (OR = 1.806 and 95%CI = 1.248–2.612, p = 0.002) (Table 3, Fig. 3). On the other hand, African cases showed no significant differences in genotype distribution of − 1082 A > G from controls (G vs. A: OR = 1.424 and 95% CI = 0.647–3.135, p = 0.379) (Table 3, Fig. 3).
Fig. 3.
OR and 95% CI for individual studies and pooled data for the association of IL-10-1082 G versus A allele.
3.3. Meta-analysis of − 819 C > T and T2DM susceptibility
For − 819 C > T, there were five studies available involving 1214 cases and 1664 controls and met the criteria for this meta-analysis (Fig. 1C). The detailed characteristics of the studies included were shown in (Table 1). Among these, three studies were performed on Asians (Chang, YH, et al., 2005, Kung, WJ, et al., 2010 and Bai et al., 2014), one study on Africans (Mtiraoui et al., 2009), and one study was performed on Europeans (Tsiavou et al., 2004). The distribution of genotype of the IL-10-819 C > T gene polymorphism in control groups was not consistent with the HWE in two studies but the overall result was coping with the genetic equilibrium (Kung et al., 2010 and Bai et al., 2014). The results of Egger's test suggested no publication bias for all comparisons (Fig. 5C and Table 4).
Table 4.
Meta-analysis of the association between IL-10 − 819 C > T polymorphism and T2D.
| Comparison | Population | Sample size |
No. of studies | Test of association |
Test of heterogeneity |
Publication bias |
||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| T2DM | Control | OR | 95% CI | P-value | Model | Q test | P-value | I2 (%) | P-value (Egger's) | |||
| T allele versus C allele (allelic contrast) |
Overall | 2428 | 3328 | 5 | 1.092 | 0.786–1.518 | 0.600 | R | 21.62 | < 0.001 | 81.49 | 0.544 |
| Asian | 1562 | 1754 | 3 | 0.986 | 0.845–1.151 | 0.860 | F | 3.31 | 0.191 | 39.58 | 0.859 | |
| European | 62 | 78 | 1 | 0.812 | 0.379–1.741 | 0.593 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 804 | 1496 | 1 | 1.683 | 1.379–2.053 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
| CT + TT versus CC (dominant model) |
Overall | 1214 | 1664 | 5 | 1.150 | 0.717–1.843 | 0.562 | R | 17.37 | 0.002 | 76.97 | 0.646 |
| Asian | 781 | 877 | 3 | 1.024 | 0.806–1.301 | 0.845 | F | 3.27 | 0.195 | 38.90 | 0.959 | |
| European | 31 | 39 | 1 | 0.782 | 0.304–2.015 | 0.611 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 402 | 748 | 1 | 1.915 | 1.496–2.450 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
| TT versus CC + CT (recessive model) |
Overall | 1167 | 1639 | 4 | 1.144 | 0.729–1.794 | 0.558 | R | 7.35 | 0.062 | 59.19 | 0.854 |
| Asian | 734 | 852 | 2 | 0.924 | 0.693–1.232 | 0.589 | F | 2.18 | 0.140 | 54.12 | NA | |
| European | 31 | 39 | 1 | 0.617 | 0.053–7.134 | 0.699 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 402 | 748 | 1 | 1.804 | 1.080–3.016 | 0.024 | F | 0.0 | 1.0 | 0.0 | NA | |
| CT versus CC (heterozygote contrast) |
Overall | 983 | 1493 | 5 | 1.172 | 0.752–1.827 | 0.483 | R | 14.29 | 0.006 | 72.10 | 0.708 |
| Asian | 581 | 740 | 3 | 1.019 | 0.796–1.305 | 0.880 | F | 2.29 | 0.319 | 12.49 | 0.866 | |
| European | 30 | 37 | 1 | 0.807 | 0.307–2.125 | 0.665 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 372 | 716 | 1 | 1.861 | 1.439–2.407 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
| TT versus CC (homozygote contrast) |
Overall | 639 | 986 | 4 | 1.178 | 0.606–2.292 | 0.629 | R | 10.21 | 0.017 | 70.61 | 0.594 |
| Asian | 392 | 445 | 2 | 0.899 | 0.426–1.899 | 0.781 | R | 3.20 | 0.074 | 68.76 | NA | |
| European | 18 | 21 | 1 | 0.559 | 0.046–6.727 | 0.647 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 229 | 520 | 1 | 2.299 | 1.360–3.885 | 0.002 | F | 0.0 | 1.0 | 0.0 | NA | |
| CT versus CC + TT (overdominant model) |
Overall | 1214 | 1664 | 5 | 1.235 | 0.900–1.693 | 0.191 | R | 10.47 | 0.033 | 61.80 | 0.985 |
| Asian | 781 | 877 | 3 | 1.058 | 0.859–1.304 | 0.594 | F | 1.17 | 0.557 | 0.0 | 0.022 | |
| European | 31 | 39 | 1 | 0.843 | 0.325–2.182 | 0.724 | F | 0.0 | 1.0 | 0.0 | NA | |
| African | 402 | 748 | 1 | 1.723 | 1.340–2.216 | < 0.001 | F | 0.0 | 1.0 | 0.0 | NA | |
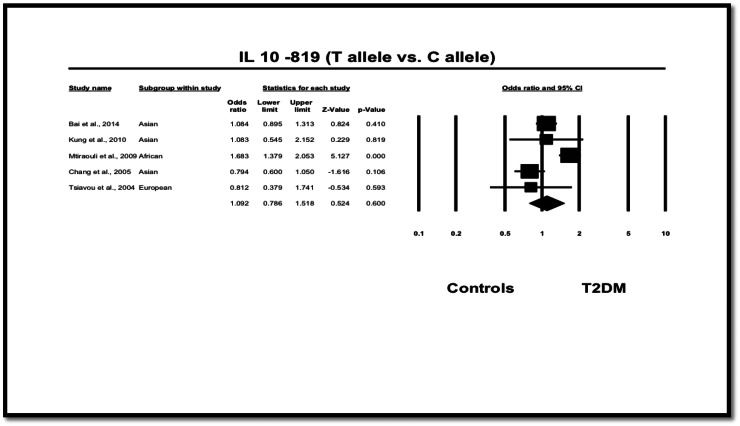
We detected no association between − 819 T allele and risk of developing T2DM in overall pooled subjects regarding the allele contrast method (T versus C allele; OR = 1.092, 95%CI = 0.786–1.518, p = 0.600), dominant model (OR = 1.150, 95% CI = 0.717–1.843, p = 0.562) (Fig. 4) neither for the recessive model (OR = 1.144, 95% CI = 0.729–1.794, p = 0.558). Regarding the ethnic distribution of subjects, although Asians and Europeans showed no association of IL-10 − 819 C > T polymorphism and T2DM, interestingly, the T allele was positively associated with T2DM among only the African population (T vs. C: OR = 1.683, 95% CI = 1.379–2.053, p < 0.001), which was also noted in the dominant, recessive, heterozygote and homozygote models (Table 4, Fig. 4).
Fig. 4.
OR and 95% CI for individual studies and pooled data for the association of IL-10 − 819 T versus C allele.
3.4. Heterogeneity and publication bias
Between-study heterogeneity concerning the IL-10 − 592 C > A and − 1082 A > G polymorphisms were significant among all subjects and thus, the random effect model was used (Table 2, Table 3). On the other hand between-study heterogeneity of the other IL-10 − 819 C > T polymorphism was not significant among all subjects and thus the fixed effect model was used (Table 4). Funnel plots were performed to assess the possibility of publication bias. The results of Egger's regression test suggested no publication bias for the three studied meta-analyses (Egger's regression test P value was > 0.1) (Fig. 5).
4. Discussion
T2DM is a complex heterogeneous status of metabolic disorders including hyperglycemia and impaired insulin function and secretion (Kramer et al., 2015). Several reports have demonstrated the association between IL-10 − 592 C > A, − 1082 A > G and − 819 C > T gene polymorphisms and the susceptibility of T2DM with relatively inconclusive conclusions (Bai, H, et al., 2014, Arababadi, MK, et al., 2012, Scarpelli, D, et al., 2006, Zhang, F, et al., 2013, Helaly, MA and Hatata, EZ, 2013, Erdogan, M, et al., 2012, Mtiraoui, N, et al., 2009, Chang, YH, et al., 2005 and Li et al., 2013). Knowing that the meta-analysis is a suitable method for evaluating small effects in human genetic association studies, we have designed this study to investigate and update the results recently given for the association of IL-10 − 592 C > A, − 1082 A > G and − 819 C > T gene polymorphisms with T2DM susceptibility in different ethnic subjects. In this meta-analysis, we updated the data of IL-10 gene in which we added two studies for − 592 C > A (Bai, H, et al., 2014, García-Elorriaga, G, et al., 2013), three studies for − 1082 A > G (Bai, H, et al., 2014, García-Elorriaga, G, et al., 2013 and Helaly and Hatata, 2013) in addition to one study for − 819 C > T (Bai et al., 2014) polymorphisms.
In this meta-analysis, in spite of the negative association of IL-10 − 592 C > A polymorphism with T2DM susceptibility in total subjects, stratified analysis indicated that the IL-10 − 592 A allele was significantly associated with T2DM among African but not European and Asian subjects. Interestingly, the IL-10 − 819 T allele was also significantly associated with T2DM among African but not in European and Asian subjects. On contrast, the IL-10 − 1082 G allele was significantly associated with T2DM in total subjects, particularly in European and Asian but not in African subjects.
Comparing our updated meta-analysis to the previously done studies, a partial agreement was found. A nearly similar conclusion was made by Hau et al., in their meta-analysis study reporting a significant association of IL-10 − 592 CA + AA and − 819 CT + TT polymorphisms in African subjects; and IL-10 − 1082 AG + GG polymorphism among Asian subjects with a negative association in European subjects (Hua et al., 2013). On the other hand, Yin, YW, et al., 2012, Yin, YW, et al., 2013 reported a positive association of IL-10 − 1082 G allele with the risk of T2DM in European subjects with no significant association of IL-10 − 592 C > A polymorphism in all ethnicity-subgroups (Yin, YW, et al., 2012, Yin, YW, et al., 2013). In agreement with our results, Li et al., revealed a significant association of IL-10 − 1082 (AG + GG vs. AA) with T2DM in overall subjects with no significant associations of IL-10 − 592 A and − 819 T alleles with T2DM in total studied subjects Asian, African and European ethnicities (Li et al., 2013). On contrast, another meta-analysis done by Zhang et al. suggested that variations of the − 1082 G/A and − 819 C/T have a protective effects with decreased risk of T2DM in Caucasian subjects for (the allelic contrast A vs. G; C vs. T, homozygote contrast AA vs. GG; CC vs. TT, dominant model AA + AG vs.GG; CC + CT vs.TT), and in Asian subjects for the homozygote contrast and recessive models of − 1082 G/A (AA vs. GG and AA vs. AG + GG) (Zhang et al., 2013). In addition, several individual studies have evaluated the association between IL-10 − 592 C > A, − 1082 A > G and − 819 C > T gene polymorphisms and the risk of development of T2DM (Bai, H, et al., 2014, Helaly, MA and Hatata, EZ, 2013, Li, J, et al., 2013 and Yin et al., 2012). Similar to our results, some reported positive association with IL-10 − 1082 G allele carriage (AG + GG) genotypes and the risk of development of T2DM in subjects of various ethnic origins as Egyptians (Helaly and Hatata, 2013), Chinese (Bai et al., 2014), Turkish (Erdogan et al., 2012), Mexican (García-Elorriaga et al., 2013) and Indian (Kolla et al., 2009) subjects. On the contrary, several other studies have revealed no association between IL-10 − 1082 A > G polymorphism and the risk of development of T2DM in some other populations like that done among Taiwanese (Kung et al., 2010), Greek (Tsiavou et al., 2004), Tunisian (Mtiraoui et al., 2009), Italian (Scarpelli et al., 2006) and Germen (Babel et al., 2006) subjects.
Similar to our findings, some studies revealed no significant association with IL-10 − 592 C allele carriage (AC + CC) genotypes and the risk of development of T2DM like that done among Indian (Saxena et al., 2013), Mexican (García-Elorriaga et al., 2013), Taiwanese (Kung et al., 2010), Italian (Scarpelli et al., 2006) and Greek (Tsiavou et al., 2004) subjects. However, other reports showed significant association of this polymorphism with the risk of development of T2DM like that done among Iranian (Arababadi et al., 2012), Chinese (Bai et al., 2014) and Tunisian (Mtiraoui et al., 2009) subjects. Few reports have studied the association of IL-10 − 819 C > T polymorphism and the risk of development of T2DM and found no association for this polymorphism like those done among Chinese (Bai et al., 2014), Taiwanese (Kung et al., 2010 and Chang et al., 2005) and Greek (Tsiavou et al., 2004) subjects. Contrastingly, one study reported a positive association between IL-10 − 819 T allele and TT genotype and the risk of development of T2DM among Tunisian subjects (Mtiraoui et al., 2009). This discrepancy in results pertinent to individual genotype association with T2DM might obviously be due to innate genetic diversity among ethnicities in addition to multiple interactive environmental factors as climate, diet, lifestyle and economic status. Factors related to the study design or sample size should also be put into consideration (Brooks et al., 2012 and Li et al., 2013).
In fact, studies concerning IL-10 − 592 C > A and − 819 C > T polymorphisms among African ethnicities were somewhat lacking. This meta-analysis involved the results of only two studies investigating these polymorphisms among African subjects. In that respect, we recommend undertaking wider scale studies with appropriate sample size of Africans to further evaluate this association.
Despite our efforts in performing a comprehensive analysis, some limitations of this meta-analysis have to be considered including the relatively insufficient raw data in spite of laying down strict search strategy in the study; noting that some relevant studies could not be included in the meta-analysis. In addition, between-studies heterogeneity and publication bias might influence the net results of the meta-analysis. Moreover, some genetic polymorphisms of included studies were deviant from HWE that might imply a potential bias during control selection or genotyping errors. Haplotype analysis was also tried, but due to the lack of enough data it was not presented here, so we also recommend a linkage and haplotype analysis on a larger scale sample in various populations in order to have a better look into the situation of this cytokine in the pathogenesis of T2DM. In spite of these limitations, this meta-analysis demonstrated that IL-10 − 1082 A > G polymorphism was associated with increased risk of development of T2DM in all subjects while both IL-10 − 592 C > A and − 819 C > T polymorphisms were associated with that risk only among African subjects.
Conflict of interest
The authors declare that they have no conflict of interest related to this work.
References
- Arababadi M.K., Reza Mirzaei M., Ali Sajadi S.M., Hassanshahi G., Ahmadabadi B.N., Salehabadi V.A., Derakhshan R., Kennedy D. Interleukin (IL)-10 gene polymorphisms are associated with type 2 diabetes with and without nephropathy: a study of patients from the southeast region of Iran. Inflammation. 2012, Jun;35(3):797–802. doi: 10.1007/s10753-011-9376-7. [DOI] [PubMed] [Google Scholar]
- Babel N., Gabdrakhmanova L., Hammer M.H., Schoenemann C., Skrypnikov V., Poliak N., Volk H.D., Reinke P. Predictive value of cytokine gene polymorphisms for the development of end-stage renal disease. J. Nephrol. 2006, Nov–Dec;19(6):802–807. [PubMed] [Google Scholar]
- Bai H., Jing D., Guo A., Yin S. Association between interleukin 10 gene polymorphisms and risk of type 2 diabetes mellitus in a Chinese population. J. Int. Med. Res. 2014, Jun;42(3):702–710. doi: 10.1177/0300060513505813. [DOI] [PubMed] [Google Scholar]
- Brooks M.V., Leake A., Parsons C., Pham V. A review of the literature: evaluating dietary intake of Filipino Americans at risk for type 2 diabetes. Nurs. Forum. 2012;47:27–33. doi: 10.1111/j.1744-6198.2011.00226.x. [DOI] [PubMed] [Google Scholar]
- Chang Y.H., Huang C.N., Wu C.Y., Shiau M.Y. Association of interleukin-10 A-592C and T-819C polymorphisms with type 2 diabetes mellitus. Hum. Immunol. 2005, Dec;66(12):1258–1263. doi: 10.1016/j.humimm.2005.05.001. [DOI] [PubMed] [Google Scholar]
- Del Prete G., De Carli M., Almerigogna F., Giudizi M.G., Biagiotti R., Romagnani S. Human IL-10 is produced by both type 1 helper (Th1) and type 2 helper (Th2) T cell clones and inhibits their antigen-specific proliferation and cytokine production. J. Immunol. 1993, Jan 15;150(2):353–360. [PubMed] [Google Scholar]
- DerSimonian R., Laird N. Meta-analysis in clinical trials. Control. Clin. Trials. 1986, Sep;7(3):177–188. doi: 10.1016/0197-2456(86)90046-2. [DOI] [PubMed] [Google Scholar]
- Egger M., Smith G.D., Phillips A.N. Meta-analysis: principles and procedures. BMJ. 1997, Dec 6;315(7121):1533–1537. doi: 10.1136/bmj.315.7121.1533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erdogan M., Cetinkalp S., Ozgen A.G., Saygili F., Berdeli A., Yilmaz C. Interleukin-10 (− 1082G/A) gene polymorphism in patients with type 2 diabetes with and without nephropathy. Genet. Test. Mol. Biomarkers. 2012, Feb;16(2):91–94. doi: 10.1089/gtmb.2011.0075. [DOI] [PubMed] [Google Scholar]
- Eskdale J., Kube D., Tesch H., Gallagher G. Mapping of the human IL10 gene and further characterization of the 5′ flanking sequence. Immunogenetics. 1997;46:120–128. doi: 10.1007/s002510050250. [DOI] [PubMed] [Google Scholar]
- Forte G.I., Pilato G., Vaccarino L., Sanacore M., Candore G., Romano G.C., Testa R., Franceschi C., Capri M., Marra M., Bonfigli A.R., Caruso C., Scola L., Lio D. Risk profiles in type 2 diabetes (metabolic syndrome): integration of IL-10 polymorphisms and laboratory parameters to identify vascular damages related complications. Curr. Pharm. Des. 2010;16(7):898–903. doi: 10.2174/138161210790883642. [DOI] [PubMed] [Google Scholar]
- García-Elorriaga G., Vera-Ramírez L., del Rey-Pineda G., González-Bonilla C. 592 and − 1082 interleukin-10 polymorphisms in pulmonary tuberculosis with type 2 diabetes. Asian Pac. J. Trop. Med. 2013, Jul;6(7):505–509. doi: 10.1016/S1995-7645(13)60086-3. [DOI] [PubMed] [Google Scholar]
- Helaly M.A., Hatata E.Z., El-magd M.A., Ibrahem E.F., Al Said A., Abd el-aal I.A., Settin A.S. Association of IL-10 and IL-6 gene polymorphisms with type 2 diabetes mellitus among Egyptian patients. Eur. J. Gen. Med. 2013;10:158–162. [Google Scholar]
- Higgins J.P., Thompson S.G. Quantifying heterogeneity in a meta-analysis. Stat. Med. 2002, Jun 15;21(11):1539–1558. doi: 10.1002/sim.1186. [DOI] [PubMed] [Google Scholar]
- Hua Y., Shen J., Song Y., Xing Y., Ye X. Interleukin-10 − 592C/A, − 819C/T and − 1082A/G polymorphisms with risk of type 2 diabetes mellitus: a HuGE review and meta-analysis. PLoS ONE. 2013, Jun 21;8(6) doi: 10.1371/journal.pone.0066568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kilpinen S., Huhtala H., Hurme M. The combination of the interleukin-1alpha (IL-1alpha-889) genotype and the interleukin-10 (IL-10 ATA) haplotype is associated with increased interleukin-10 (IL-10) plasma levels in healthy individuals. Eur. Cytokine Netw. 2002, Jan-Mar;13(1):66–71. [PubMed] [Google Scholar]
- Kolla V.K., Madhavi G., Pulla Reddy B., SrikanthBabu B.M., Yashovanthi J., Valluri V.L., Ramesh J., Akka J. Association of tumor necrosis factor alpha, interferon gamma and interleukin 10 gene polymorphisms with peripheral neuropathy in South Indian patients with type 2 diabetes. Cytokine. 2009, Sep;47(3):173–177. doi: 10.1016/j.cyto.2009.06.007. [DOI] [PubMed] [Google Scholar]
- Kramer C.K., Zinman B., Choi H., Retnakaran R. Effect of short-term intensive insulin therapy on post-challenge hyperglucagonemia in early type 2 diabetes. J. Clin. Endocrinol. Metab. 2015, Aug;100(8):2987–2995. doi: 10.1210/jc.2015-1947. [DOI] [PubMed] [Google Scholar]
- Kung W.J., Lin C.C., Liu S.H., Chaung H.C. Association of interleukin-10 polymorphisms with cytokines in type 2 diabetic nephropathy. Diabetes Technol. Ther. 2010;12:809–813. doi: 10.1089/dia.2010.0085. [DOI] [PubMed] [Google Scholar]
- Li J., Wu S., Wang M.R., Wang T.T., Zhu J.M. Association of the interleukin-10 − 592A/C, − 1082G/A and − 819T/C gene polymorphisms with type 2 diabetes: a meta-analysis. Gene. 2013, Jun 1;521(2):211–216. doi: 10.1016/j.gene.2013.03.072. [DOI] [PubMed] [Google Scholar]
- Lin J.D., Hsia T.L., Wu C.Z., Su C.C., Ma W.Y., Hsieh A.T., Hsieh C.H., Wang K., Chu Y.M., Pei D. The first and second phase of insulin secretion in naive Chinese type 2 diabetes mellitus. Metabolism. 2010, Jun;59(6):780–786. doi: 10.1016/j.metabol.2009.09.024. [DOI] [PubMed] [Google Scholar]
- Mantel N., Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 1959, Apr;22(4):719–748. [PubMed] [Google Scholar]
- Moher D., Liberati A., Tetzlaff J., DG Altman, PRISMA Group Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. Int. J. Surg. 2010;8(5):336–341. doi: 10.1016/j.ijsu.2010.02.007. [DOI] [PubMed] [Google Scholar]
- Mtiraoui N., Ezzidi I., Kacem M., Hadj Ben, Mohamed M., chaieb M., AB Haj Jilani, mahjoub T., WY. almawi. Predictive value of interleukin-10 promoter genotypes and haplotypes in determining the susceptibility to nephropathy in type 2 diabetes patients. Diabetes Metab. Res. Rev. 2009, Jan;25(1):57–63. doi: 10.1002/dmrr.892. [DOI] [PubMed] [Google Scholar]
- Pickup J.C. Inflammation and activated innate immunity in the pathogenesis of type 2 diabetes. Diabetes Care. 2004, Mar;27(3):813–823. doi: 10.2337/diacare.27.3.813. [DOI] [PubMed] [Google Scholar]
- Saxena M., Srivastava N., Banerjee M. Association of IL-6, TNF-α and IL-10 gene polymorphisms with type 2 diabetes mellitus. Mol. Biol. Rep. 2013, Nov;40(11):6271–6279. doi: 10.1007/s11033-013-2739-4. [DOI] [PubMed] [Google Scholar]
- Scarpelli D., Cardellini M., Andreozzi F., Laratta E., Hribal M.L., Marini M.A., Tassi V., Lauro R., Perticone F., Sesti G. Variants of the interleukin-10 promoter gene are associated with obesity and insulin resistance but not type 2 diabetes in Caucasian Italian subjects. Diabetes. 2006, May;55(5):1529–1533. doi: 10.2337/db06-0047. [DOI] [PubMed] [Google Scholar]
- Steyn N.P., Lambert E.V., Tabana H. Conference on “Multidisciplinary approaches to nutritional problems”. Symposium on “Diabetes and health”. Nutrition interventions for the prevention of type 2 diabetes. Proc. Nutr. Soc. 2009, Feb;68(1):70–‐55. doi: 10.1017/S0029665108008823. [DOI] [PubMed] [Google Scholar]
- Tsiavou A., Hatziagelaki E., Chaidaroglou A., Manginas A., Koniavitou K., Degiannis D., Raptis S.A. TNF-alpha, TGF-beta1, IL-10, IL-6, gene polymorphisms in latent autoimmune diabetes of adults (LADA) and type 2 diabetes mellitus. J. Clin. Immunol. 2004, Nov;24(6):591–599. doi: 10.1007/s10875-004-6239-0. [DOI] [PubMed] [Google Scholar]
- van Exel E., Gussekloo J., de Craen A.J., Frölich M., Der Wiel A Bootsma-Van, RG westendorp. Leiden 85 plus study. low production capacity of interleukin-10 associates with the metabolic syndrome and type 2 diabetes: the Leiden 85-plus study. Diabetes. 2002, Apr;51(4):1088–1092. doi: 10.2337/diabetes.51.4.1088. [DOI] [PubMed] [Google Scholar]
- Wang J.D., Fang H., Yan Q.Z., Zhou D.H., Yao H.J. Relationship of serum interleukin-10 level and its gene promoter 592 polymorphism to type 2 diabetes mellitus. Chin. Gen. Pract. 2010;13:1185–1188. [Google Scholar]
- Yin Y.W., Hu A.M., Sun Q.Q., Zhang B.B., Liu H.L., Wang Q., Zeng Y.H., Xu R.J., Zhang S.J., Shi L.B. Association between interleukin 10 gene − 1082 A/G polymorphism and the risk of type 2 diabetes mellitus: a meta-analysis of 4250 subjects. Cytokine. 2013, May;62(2):226–231. doi: 10.1016/j.cyto.2013.02.025. [DOI] [PubMed] [Google Scholar]
- Yin Y.W., Sun Q.Q., Zhang B.B., Hu A.M., Liu H.L., Wang Q., Zeng Y.H., Xu R.J., Ma J.B., Shi L.B. Association between interleukin-10 gene − 592 C/A polymorphism and the risk of type 2 diabetes mellitus: a meta-analysis of 5320 subjects. Hum. Immunol. 2012;73:960–965. doi: 10.1016/j.humimm.2012.06.006. [DOI] [PubMed] [Google Scholar]
- Zhang F., Yang Y., Lei H., Qiu J., Wang Y., Hu D., Skrip L., Chen F. A meta-analysis about the association between − 1082G/A and − 819C/T polymorphisms of IL-10 gene and risk of type 2 diabetes. Hum. Immunol. 2013, May;74(5):618–626. doi: 10.1016/j.humimm.2013.01.021. [DOI] [PubMed] [Google Scholar]