Significance
The virulence and host range of viruses is controlled by the interaction of the host innate immune system with viral molecules. This interaction is an important driver for the evolution of both the host and the virus. The attenuation of myxoma virus, a rabbit-specific poxvirus, after its deliberate release to control European rabbit populations, and the increased resistance of the rabbits, is one of the best-known examples for host–virus coevolution on the population level. We show that the myxoma virus protein M156 specifically inhibited the antiviral protein kinase R (PKR) from rabbits but not PKR from other mammals, that PKR inhibition correlated with virus replication during infection, and that M156 contains a loss-of-function mutation in Australian field isolates.
Keywords: poxvirus, myxoma virus, PKR, translational regulation, host–pathogen interaction
Abstract
Myxoma virus (MYXV) is a rabbit-specific poxvirus, which is highly virulent in European rabbits. The attenuation of MYXV and the increased resistance of rabbits following the release of MYXV in Australia is one of the best-documented examples of host–pathogen coevolution. To elucidate the molecular mechanisms that contribute to the restriction of MYXV infection to rabbits and MYXV attenuation in the field, we have studied the interaction of the MYXV protein M156 with the host antiviral protein kinase R (PKR). In yeast and cell-culture transfection assays, M156 only inhibited rabbit PKR but not PKR from other tested mammalian species. Infection assays with human HeLa PKR knock-down cells, which were stably transfected with human or rabbit PKR, revealed that only human but not rabbit PKR was able to restrict MYXV infection, whereas both PKRs were able to restrict replication of a vaccinia virus (VACV) strain that lacks the PKR inhibitors E3 and K3. Inactivation of M156R led to MYXV virus attenuation in rabbit cells, which was rescued by the ectopic expression of VACV E3 and K3. We further show that a mutation in the M156 encoding gene that was identified in more than 50% of MYXV field isolates from Australia resulted in an M156 variant that lost its ability to inhibit rabbit PKR and led to virus attenuation. The species-specific inhibition of rabbit PKR by M156 and the M156 loss-of-function in Australian MYXV field isolates might thus contribute to the species specificity of MYXV and to the attenuation in the field, respectively.
Poxviruses are large double-stranded DNA viruses that exclusively replicate in the cytoplasm of infected cells. Members of the Poxviridae family can productively infect a wide variety of animal hosts. Interestingly, the binding and entry of poxviruses into cells is largely independent of host species after which virus replication is initiated. The successful completion of virus replication, however, depends on the effective subversion of the host cell’s innate immune responses (1). Even closely related poxviruses can exhibit drastic differences in their host ranges. Whereas some poxviruses have only one host species, such as variola virus, the causative agent of smallpox, which is restricted to humans, others, such as cowpox and monkeypox viruses, can infect many different species and thus display very broad host ranges (2). A number of poxvirus genes have been discovered that influence the host range and cell tropism of poxviruses and have therefore been termed “host range genes” (3, 4). Although the molecular mechanisms responsible for their host range functions have not been elucidated in detail, it is clear that most poxviral host range proteins interact with components of the host immune system and that host species-specific interactions likely play a major role.
Myxoma virus (MYXV) is a poxvirus that belongs to the genus leporipoxvirus and shows a restricted host range infecting only leporids (rabbits and hares). MYXV is highly lethal to European (E.) rabbits causing case fatality rates (CFRs) of close to 100%. Since 1950, MYXV was repeatedly introduced into Australia to combat the invasive feral E. rabbit population, which has caused ecological and economical havoc. Shortly after the release of the MYXV standard laboratory strain (SLS), which caused a CFR of 99.8% in laboratory rabbits (grade 1 virulence), attenuated virus strains began to appear in the wild and started to outcompete the more virulent parental strain. The predominant strains found in the field are of grade 3 and grade 4 virulence and exhibit a CFR in laboratory rabbits between 70–95% and 50–70%, respectively. Concomitantly, rabbits evolved increased resistance to MYXV infection. Evolution of attenuated MYXV and increased resistance of rabbits to infection were also observed after the illegal release of MYXV in Europe (reviewed in ref. 5). The molecular mechanisms of the attenuation of MYXV and the increased resistance of rabbits to infection are unknown. Recently, the complete genomes of 24 MYXV strains that were collected in the field in Australia were reported. Although a number of mutations were discovered, it was not immediately clear which mutations led to changes in MYXV virulence (6, 7). Two candidate genes that might contribute to changes in virulence are M029L and M156R, the MYXV orthologs of vaccinia virus (VACV) E3L and K3L, respectively. VACV E3L and K3L are virulence and host range genes, and their protein products E3 and K3 inhibit the activation and activity of PKR (8, 9). PKR is an antiviral protein that is found in most vertebrates. It is constitutively expressed at moderate levels and can be induced by type I interferons. PKR is composed of two N-terminal double-stranded RNA (dsRNA) binding domains that sense viral dsRNA, and a C-terminal kinase domain. Upon binding to dsRNA, two inactive PKR monomers dimerize and undergo autophosphorylation. Activated PKR subsequently phosphorylates the alpha subunit of eukaryotic translation initiation factor 2 (eIF2), which leads to the general suppression of protein translation and inhibition of virus replication. During vertebrate evolution PKR has evolved rapidly, likely as a consequence of positive selective pressure exerted by viral PKR antagonists. We and others previously showed that VACV K3 inhibits PKR in a species-specific manner; e.g., whereas mouse PKR was sensitive to K3 inhibition, human PKR was largely resistant (10, 11). MYXV 156 is a homolog of eIF2α and was previously tested for its ability to inhibit human PKR in a heterologous yeast assay in which it showed no inhibition of human PKR activity (12). Here we explored the hypothesis that M156R evolved to inhibit rabbit PKR and that species-specific inhibition of PKR contributes to the restricted host range of MYXV to rabbits. We further tested whether variations found in MYXV field isolates affected the inhibitory potential of M156 and M029 against rabbit PKR.
Results
Predominant Expression of a Short M156 Form.
M156R is located at the 3′ end of the genome and partly overlaps with the inverted terminal repeat (ITR) region in the reference Lausanne (Lu) strain (13). Among all known poxvirus K3 orthologs, M156 is unique, because it contains a predicted N-terminal extension, based on an elongated ORF (open reading frame) (Fig. 1A and Fig. S1A). In MYXV strains that descended from South America, M156R is annotated to encode a 102-amino acid protein with a predicted molecular mass of 12 kDa (13). The M156 orthologs of the closest relatives, rabbit fibroma virus and the Californian MYXV MSW strain, lack this putative extension, but contain a putative start codon that encodes for 78- and 77-aa-long ORFs, respectively, with predicted molecular masses of 9 kDa (Fig. S1A) (14, 15). A putative start codon at the corresponding position is also found in the South American-derived MYXV strains. In the solution structure of M156, for which the long M156R ORF was used, the first 32 amino acids were unstructured (12). A predicted poxvirus early promoter motif is absent in the 500 bp 5′ to the first start codon, but is present at nucleotide position −42 relative to the second predicted start codon (Fig. S1B). To determine the authentic ORF of M156R, we performed rapid amplification of cDNA ends (RACE) PCR to determine the transcriptional start sites (TSSs) using total RNA from MYXV-Lu–infected RK13 cells 4, 12, or 24 h postinfection (hpi). Prominent bands were observed at about 120 bp at all time points but not in the uninfected control (Fig. S1C). PCR products were randomly cloned and sequenced. TSSs of 17 out of 18 clones started within the extended ORF, with the majority (82%) of sequences starting at positions −15 to −7 relative to the second start codon (Figs. S1B and S2). None of the discovered TSSs are predicted to lead to the translation of the long M156 isoform. To analyze the expression pattern of M156 and compare its molecular mass to that of transfected M156, we performed Western blot analyses of RK13 cells infected with Lu MYXV at various time points (Fig. S1D). M156 expression was detected as early as 1 hpi and was most strongly expressed after 4–24 hpi and thus showed comparable expression to M-T7, an early expressed MYXV protein. In contrast, expression of the late protein M130 was not observed until 8 hpi. We also transfected RK13 cells with plasmids encoding the short or long M156 isoforms and analyzed their expression using anti-M156 serum. The short M156 isoform (Fig. S1D, lane 7) was detected at a comparable size as M156 in MYXV-infected cells. We were unable to detect the expression of the long M156 isoform under any conditions using our M156 antiserum. In conclusion, our data indicate that the short form of M156 is the predominant isoform in the tested cells, and we therefore used this version in subsequent experiments. To avoid confusion with M156R annotations of MYXV genomes, we kept the numbering of the long isoform when referring to specific amino acids.
Fig. 1.
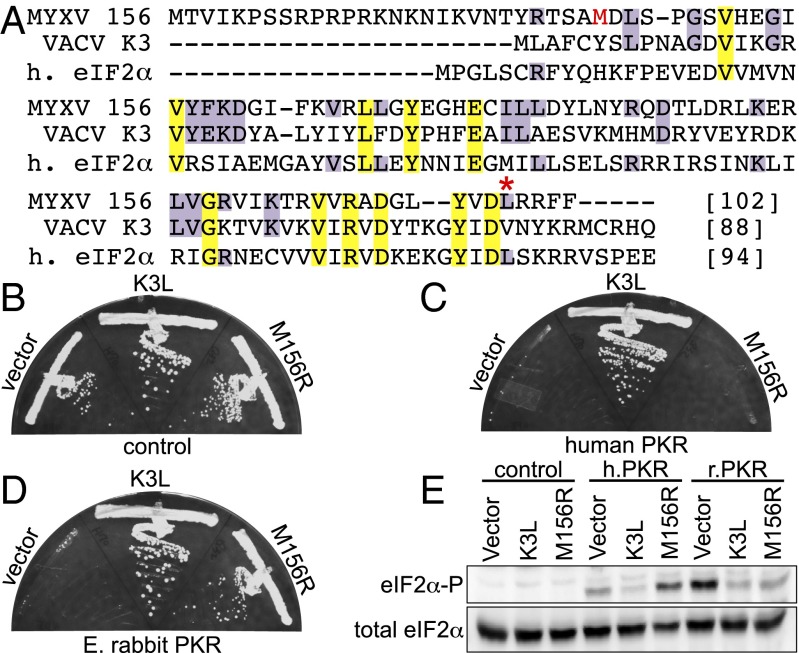
M156 inhibits rabbit but not human PKR in yeast. (A) Multiple sequence alignment of MYXV-Lu M156, VACV K3, and human (h) eIF2α. Conserved residues are highlighted in yellow (100% conservation) or purple (identical with M156). The methionine encoded by the putative start codon of the predominant M156 isoform is shown in red. An asterisk indicates L98 in MYXV-Lu M156. Plasmids encoding VACV K3, MYXV M156, or empty vector under the control of a yeast GAL-CYC1 hybrid promoter were transformed into isogenic yeast strains, which have either empty vector (control) (B), human PKR (C), or E. rabbit PKR (D) stably integrated at the LEU2 locus under the control of the GAL-CYC1 promoter. Transformants were colony purified and grown under inducing conditions at 30 °C for 4 d. Results shown are representative of four independent transformants for each plasmid. (E) Transformants described above were grown in liquid SC−Gal medium for 4 h to induce expression. Whole cell protein extracts were obtained from equal numbers of cells and subjected to Western blot analyses. The blots were probed with phospho-specific antibodies against Ser51 eIF2α (eIF2α-P), then stripped and probed with polyclonal antiserum against total eIF2α.
Fig. S1.
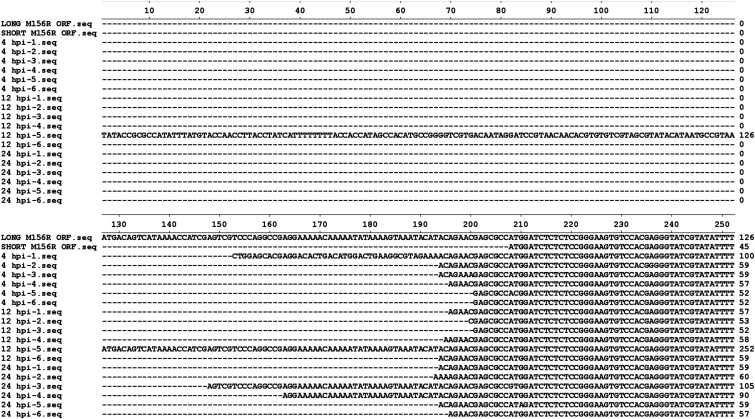
Identification of the major M156 transcription start sites (TSSs). (A) Multiple sequences alignment of human eIF2α and viral eIF2α homologs from MYXV-Lu, rabbit fibroma virus, MYXV-MSW, VACV, and Rana catesbeiana virus Z. Conserved residues are highlighted in yellow (100% conservation) or purple (identical with MYXV-Lu). An asterisk indicates L98 in MYXV-Lu M156. (B) Illustration of the observed TSSs for M156R. The genomic composition of the 5′ end of M156R locus is indicated with red arrows denoting the presence of ATG codons. Numbers indicate relative nucleotide positions with respect to the identified start codon. Blue bars represent observed TSSs at any of the three time points. The height of each bar is proportional to the number of times each particular TSS was observed. A predicted poxvirus early promoter motif is highlighted in the light green box. (C) The 5′-RACE PCR was performed with RNA collected from MYXV-infected RK13 cells at 4, 12, and 24 hpi. Total PCR products were cloned and sequenced. (D) M156 is an early protein. RK13 cells were left uninfected (mock, lane 1) or infected with MYXV with a MOI of 10. Total protein was collected at indicated time points (lanes 2–6). Short and long forms of M156R were transiently transfected and total protein was collected 24 h later. All protein samples were separated on SDS/PAGE and subjected to Western blotting analyses. The membrane was first probed with an anti-M156 antibody, then consecutively stripped and probed for M-T7 (early gene), M130 (late gene), and β-actin (loading control).
Fig. S2.
Sequence alignment of the 5′ RACE PCR products. European rabbit RK13 cells were infected with MYXV-Lu and the total RNA was collected at indicated time points. The 5′ RACE PCRs were performed and the PCR products were cloned and sequenced. The 5′ ends of the predicted long (126 bp) and short (45 bp) forms of M156R were aligned with sequences of the 5′ RACE PCR clones by the program MegAlign (DNASTAR).
Host-Specific PKR Inhibition by M156.
Yeast-based assays have been previously used to analyze interactions of PKR and viral inhibitors (10, 11, 16–18). In one of these assays, M156 failed to inhibit human PKR, whereas its VACV ortholog K3 showed PKR inhibition (12). We hypothesized that M156R evolved to inhibit rabbit PKR. To test this hypothesis, we generated isogenic yeast strains that were stably transformed with empty vector (control), human PKR, or E. rabbit PKR under the control of a galactose-inducible promoter (18). These strains were subsequently transformed with plasmids encoding VACV K3, MYXV M156, or empty vector (control), which are also under the control of the galactose-inducible promoter. Under inducing conditions (galactose-containing agar plates), all transformants of the control strain expressing no PKR showed comparable growth, indicating that K3 or M156 alone had no effect on yeast growth (Fig. 1B). Induction of both human and E. rabbit PKR expression was toxic in the vector-transformed strains, whereas this toxicity was suppressed by K3 (Fig. 1 C and D). Expression of M156 had no effect on human PKR toxicity, whereas it reduced toxicity of E. rabbit PKR, as indicated by no growth and growth, respectively (Fig. 1 C and D). The transformants were also grown in liquid media to measure eIF2α phosphorylation levels 4 h after galactose induction by Western blot analysis. No eIF2α phosphorylation was observed in the absence of PKR (Fig. 1E). Both human and E. rabbit PKR induced eIF2α phosphorylation, which was reduced by the expression of K3 for both PKR. Expression of M156 only inhibited eIF2α phosphorylation induced by E. rabbit PKR, but not that mediated by human PKR. These results demonstrate that M156 is an inhibitor of E. rabbit PKR but not of human PKR in this yeast assay and also explain why no inhibition of human PKR was observed previously (12).
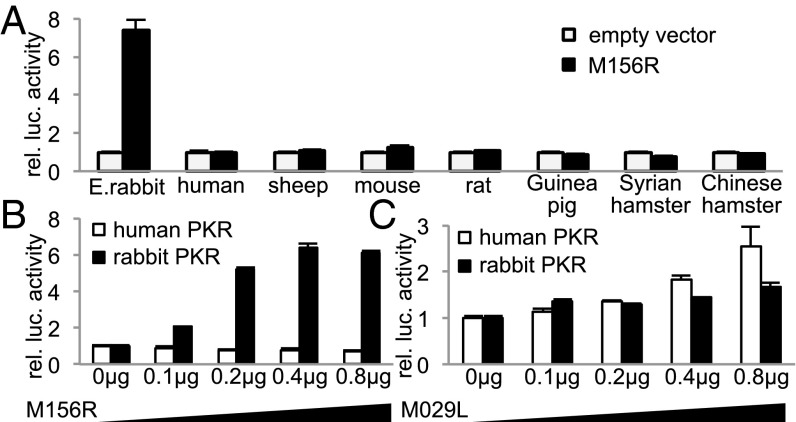
To extend the analysis of host-specific PKR inhibition by M156, we used a previously described assay in HeLa cells, in which the endogenous PKR was stably knocked down by shRNA, designated as HeLa-PKRkd cells (10). To avoid the effect of RNA interference on transfected human PKR, we used knock-down resistant human PKR (PKRkd-res) for subsequent experiments. HeLa-PKRkd cells were cotransfected with E. rabbit, human, sheep, mouse, rat, Guinea pig, Syrian hamster or Chinese hamster PKR, M156R or a control plasmid (empty vector), and firefly luciferase. Our previous studies showed that the reduction of luciferase activity is a more sensitive indicator of PKR activity than phosphorylation of eIF2α in transient transfection assays (10). Luciferase activities were normalized to PKR-only transfected cells. Increases in luciferase activity indicate inhibition of PKR activity. Transfection of M156R resulted in strongly increased luciferase expression only in cells cotransfected with E. rabbit but not in cells cotransfected with the other species’ PKRs, which indicates that M156 inhibited only E. rabbit PKR, whereas the other species’ PKRs were resistant (Fig. 2A). This result therefore confirms and extends the results of species-specific M156 activity obtained in the yeast assays. Because MYXV contains a second PKR inhibitor encoded by M029L, which belongs to the poxvirus E3L family of dsRNA-binding proteins, we analyzed whether M029 also exhibits species-specific PKR inhibitory activity. HeLa-PKRkd cells were cotransfected with human or E. rabbit PKR and increasing amounts of M029L or M156R. Whereas E. rabbit PKR but not human PKR was inhibited in a dose-dependent manner by M156, both PKRs showed comparable sensitivity to M029, indicating that only M156 exhibits pronounced species specificity (Fig. 2 B and C). Myxoma virus proteins M-T5, M011, M013, and M130, used as controls, had no effect on PKR activity (Fig. S3).
Fig. 2.
Species-specific inhibition of rabbit PKR by M156. (A) Human HeLa-PKRkd cells were transfected with expression vectors for luciferase (0.05 µg), MYXV M156R (0.4 µg), and PKR (0.2 µg) from the indicated mammalian species. Luciferase light units were normalized to PKR-only transfected cells to obtain relative luciferase activities. Constant amounts (0.1 µg) of human PKR or E. rabbit PKR were cotransfected with increasing amounts of M156R (B) or M029L (C), and relative luciferase activities are shown. Experiments were performed in triplicate and the results are representative of three independent experiments. Error bars indicate SD.
Fig. S3.
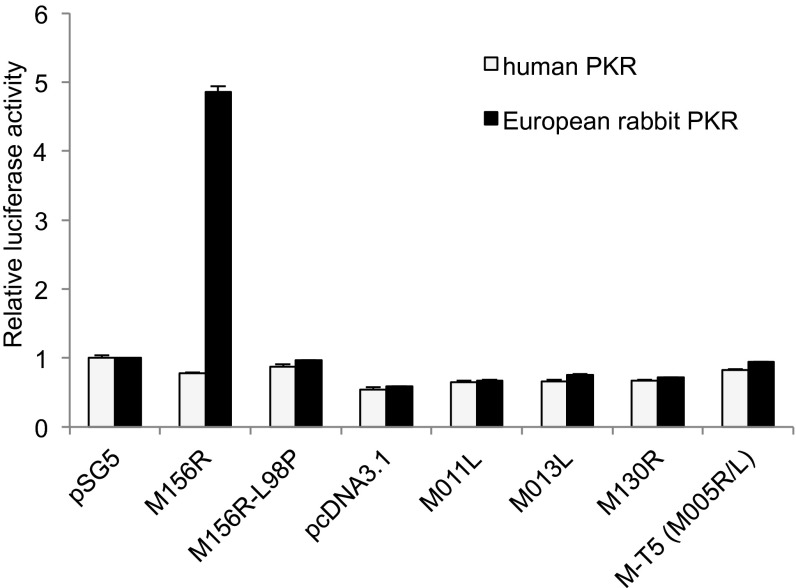
M156 but not M011, M013, M130, and M-T5 inhibits E. rabbit PKR. Human HeLa-PKRkd cells were transfected with expression vectors for luciferase (0.05 µg), human, or E. rabbit PKR (0.2 µg) and myxoma virus M156, M156-L98P, M011, M013, M130, or MT-5 (0.4 µg each). pSG5 and pcDNA3.1 serve as empty vector controls. Luciferase light units were normalized to cells transfected only with PKR and pSG5 to obtain relative luciferase activities. Error bars indicate SD.
Myxoma Virus Can Overcome the Antiviral Effects of Rabbit PKR.
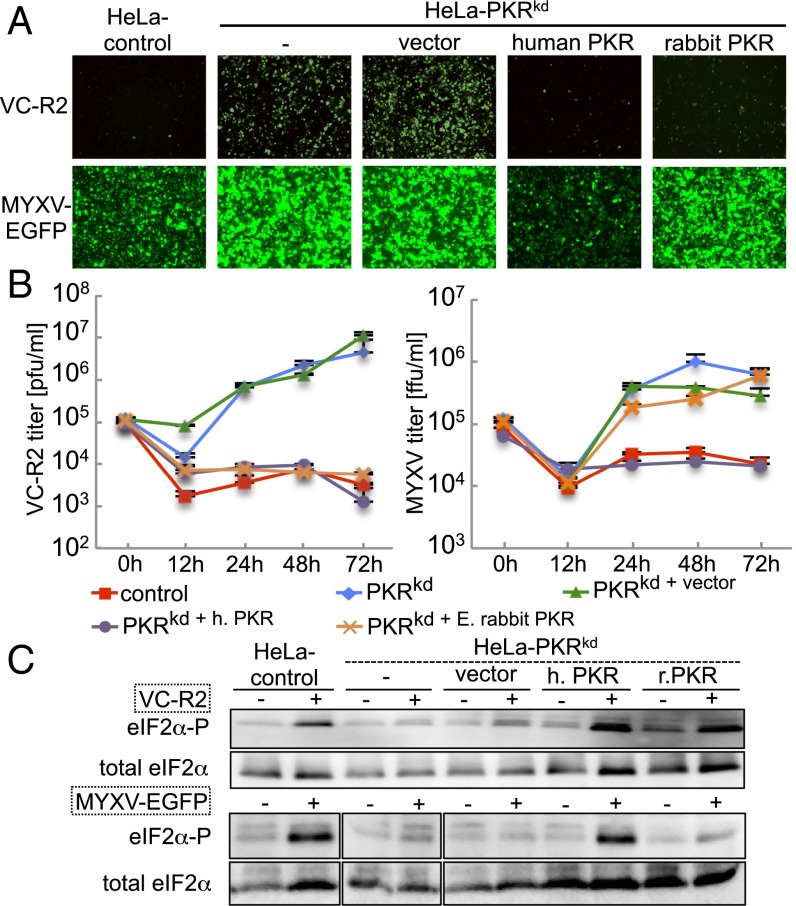
To characterize the antiviral effects of E. rabbit and human PKR during poxvirus infection, we established an infection assay using HeLa-PKRkd cells in which we stably expressed PKR. To mimic natural PKR expression, we constructed a plasmid that contains the human PKR promoter as characterized (19), followed by the rabbit β-globin intron and a multiple cloning site, into which we cloned human PKRkd-res or E. rabbit PKR, followed by two FLAG tags. HeLa-PKRkd cells were stably transfected with the PKR-expressing plasmids or empty vector and clones derived from single colonies were analyzed for PKR expression. For subsequent experiments, a clone (h14) that expressed transgenic human PKR (HeLa-PKRkd+humanPKR) at levels comparable to that of control HeLa cells was chosen, as determined by Western blot using an anti-human PKR antibody (Fig. S4). An E. rabbit PKR-expressing clone (r15, HeLa-PKRkd+E. rabbit PKR) was chosen that showed comparable PKR expression to that of HeLa-PKRkd+humanPKR, using an anti-FLAG tag antibody (Fig. S4). HeLa control cells, as well as HeLa-PKRkd and its derivatives stably transfected with vector, human, or rabbit PKR were infected with VC-R2, a VACV strain that lacks both PKR inhibitors K3 and E3, to analyze if VACV replication is suppressed when PKR is expressed. In VC-R2, the E3L gene is replaced with a destabilized EGFP driven by the native E3L promoter and its expression can be used as readout for VACV protein expression. EGFP was expressed in HeLa-PKRkd and HeLa-PKRkd+vector cells, whereas it was strongly suppressed in HeLa control cells and HeLa-PKRkd cells expressing human or E. rabbit PKR (Fig. 3A). We also determined VC-R2 replication in the infected cells by performing plaque assays on rabbit RK13 cells expressing E3 and K3, which are permissive for VC-R2. VC-R2 could only replicate in HeLa-PKRkd and HeLa-PKRkd+vector, whereas endogenous levels of PKR expression completely suppressed VC-R2 replication (Fig. 3B). These results demonstrate that transgenic expression of either human or E. rabbit PKR can compensate for the loss of PKR in HeLa cells with respect to the suppression of VACV replication. We next infected the same cell lines with MYXV that expresses EGFP under the control of a synthetic early/late poxvirus promoter (20). In HeLa control and HeLa-PKRkd+humanPKR cells, relatively weak EGFP expression was observed, whereas strong EGFP was observed in PKR-deficient and E. rabbit PKR-expressing cells (Fig. 3A). This observation correlated with MYXV replication in these cells, which was only suppressed in cells expressing human PKR but not E. rabbit PKR (Fig. 3B). We also monitored eIF2α phosphorylation levels in mock- and virus-infected congenic HeLa cells by Western blot analyses to assess whether the effects of transgene expression on virus replication correlated with eIF2α phosphorylation. In HeLa-PKRkd and HeLa-PKRkd+vector cells, basal eIF2α phosphorylation levels were not increased by VC-R2 or MYXV infection (Fig. 3C). In HeLa-PKRkd+humanPKR and HeLa-PKRkd+E. rabbit PKR cells, VC-R2 infection induced strong eIF2α phosphorylation to levels comparable to that in the HeLa control cells. MYXV infection only led to increased eIF2α phosphorylation in HeLa control and HeLa-PKRkd+humanPKR cells, which express human PKR, but not in HeLa-PKRkd, HeLa-PKRkd+vector, and cells expressing E. rabbit PKR (Fig. 3C). Thus, the effects of PKR on virus replication correlated well with eIF2α phosphorylation levels. The combined results show that MYXV was resistant to the antiviral effects of E. rabbit PKR but sensitive to those of human PKR.
Fig. S4.
PKR expression in HeLa cells and derived cell lines. HeLa-PKRkd cells were stably transfected with vector, human (h), or E. rabbit (r) PKR under the control of the human PKR promoter. The human PKR-expressing clone h14 (lane 3) showed PKR expression comparable to that in HeLa control cells (lane 2) using an anti-human PKR antibody. Untransfected HeLa-PKRkd cells are shown in lanes 1 and 4. Clone h14 was used to identify rabbit PKR-expressing clone r15, which showed comparable expression levels using an anti-FLAG tag antibody (lanes 5 and 6). After stripping, membranes were reprobed with anti–β-actin antibodies.
Fig. 3.
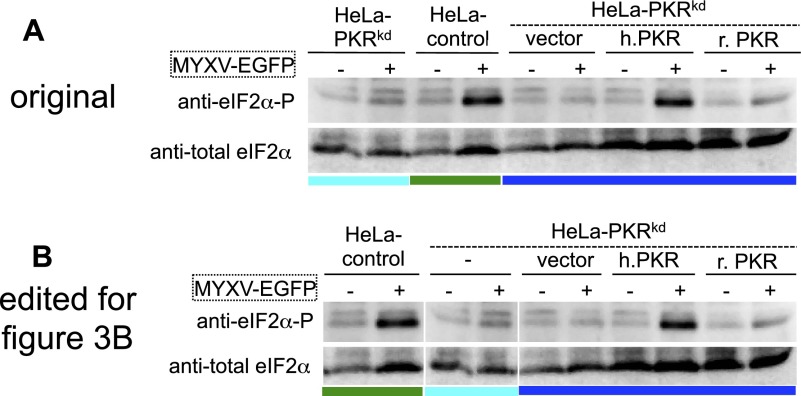
Human PKR but not rabbit PKR suppresses MYXV replication. (A) Control and congenic HeLa-PKRkd HeLa cell lines that were stably transfected with vector, human PKR, or E. rabbit PKR were infected with VC-R2 [VACV∆E3L(EGFP-d2)/∆K3L] or MYXV-EGFP at a MOI of 0.1 or 1, respectively. Representative images taken 48 hpi at 100× magnification are shown. EGFP levels in the two viruses cannot be compared directly, because in VC-R2, a destabilized EGFPd2 is driven by the natural E3L promoter, whereas in MYXV, unmodified EGFP is driven by a synthetic early–late poxvirus promoter. (B) Indicated HeLa cell lines were infected with VC-R2 or MYXV-EGFP at a MOI of 0.1 or 1, respectively. Viruses were collected at 0, 12, 24, 48, and 72 hpi and titered on RK13 cells, which stably express VACV E3 and K3. (C) Total protein lysates were collected from mock (−), VC-R2, or MYXV-EGFP–infected HeLa cells at 8 hpi to measure phosphorylated and total eIF2α by Western blot analyses. Lysates shown were run on the same gel in a different order and lanes were reordered for clarity (Fig. S7 shows original).
Fig. S7.
Original Western blot modified for Fig. 3B. (A) Original Western blots for phosphorylated and total eIF2a in total protein lysates collected from mock (−) or MYXV-EGFP (+) infected HeLa cells at 8 hpi. For clarity, the lanes that show eIF2α from HeLa-PKRkd (light blue bar) and HeLa control cells (green bar) were switched to generate the figure shown in B and Fig. 3B.
M156 Deficiency Leads to MYXV Attenuation in Rabbit Cells.
We inactivated M156R in MYXV-M029LKO, which lacks the other PKR inhibitor M029 (21), to generate MYXV-M029LKOM156RKO, which is devoid of both PKR inhibitors (Fig. S5). Deletion of M029L alone resulted in smaller plaque sizes and approximtely 10- to 50-fold titer reduction in infected RK13 cells (21) (Fig. 4 A and B). Inactivation of M156R in this strain resulted in further attenuation as indicated by the reduced EGFP expression and an additional 10- to 17-fold titer reduction (Fig. 4 A and B). Viral protein synthesis was abolished in MYXV-M029LKOM156RKO as indicated by the absence of M-T7 and M130 expression (Fig. 4C). In RK13+E3L+K3L, replication and protein synthesis of MYXV-M029LKOM156RKO were not impaired, indicating that the attenuation of this strain is caused by its inability to inhibit PKR. Interestingly, we consistently observed higher expression of the early proteins M156 and M-T7 in both RK13 and RK13+E3L+K3L cells infected with MYXV-M029LKO, which indicates that M029 might be a negative regulator of some MYXV proteins’ expression.
Fig. S5.
Construction of recombinant myxoma virus strains. The M156 coding sequence was partially replaced by a DNA cassette containing a monomeric red fluorescent protein (mRFP) encoding gene. The genomic region surrounding the M156R locus is shown in A. The region duplicated in the inverted terminal repeat is indicated. (B) The recombination plasmid to delete M156R contains a part of the M154L gene, the intergenic region between M154L and M156R, part of the M156R sequence, a poxvirus synthetic early/late (sE/L) promoter followed by the mRFP gene, followed by a part of the M156R sequence, the intergenic region between M156R and M008.1L and a part of M008.1L. To generate revertant M156R- (MYXV-M029LKO/M156RKOM156Rrev) and M156R-L98P- (MYXV-M029LKO/M156RKOM156R-L98P) containing strains, the constructs shown in C and D, respectively, were used.
Fig. 4.
Deletion of M156R further attenuates MYXV that lacks M029L. RK13 cells (RK13-WT) and RK13 cells stably expressing E3 and K3 (RK13+E3L+K3L) were infected with MYXV-EGFP (WT), MYXV-M029LKO, or MYXV-M029LKOM156RKO at a MOI of 5. (A) Fluorescent images were taken at 48 hpi at 100× magnification. (B) Viruses were collected at 0, 12, 24, and 48 hpi and titered on RK13+E3L+K3L cells. Error bars indicate SD (n = 2). (C) Total protein lysates were collected at 4 and 24 hpi to monitor expression of MYXV early (M156 and M-T7) and late (M130) proteins. β-Actin was included as loading control.
Loss-of-Function Mutation in M156R of Australian MYXV Isolates.
Despite the availability of the full genome sequences of 24 MYXV isolates from Australia, the molecular mechanisms that underlie the attenuation of MYXV strains in the field are currently unclear. One of the identified mutations leads to a predicted leucine-to-proline amino acid substitution at position 98 (L98P) in M156R in 13 of 24 isolates (7). The corresponding leucine residue in the costructure of eIF2α with human PKR (22) is shown in Fig. 5A. We introduced this mutation into M156-encoding plasmids using site-directed mutagenesis and tested its effect on E. rabbit PKR in yeast and cell culture assays. The mutation had no effect on yeast growth in a control strain, but abolished the M156-mediated rescue of yeast growth in a strain expressing rabbit PKR (Fig. 5 B and C). In correlation with this result, M156-L98P was unable to inhibit M156-mediated inhibition of eIF2α phosphorylation in yeast grown in liquid medium (Fig. 5D). The inability of M156-L98P to inhibit PKR was also confirmed in the cell-culture–based transfection assays, in which even the highest amounts of M156-L98P had no effect on PKR activity (Fig. 5E). In transfected cells, both M156 and M156-L98P were expressed at comparable levels as determined by Western blot analysis and the loss of PKR inhibition was therefore not due to altered M156 expression (Fig. 5E). A predicted alanine-to-valine substitution at position 17 in M029, which was present in 11 of 24 isolates (7), did not result in altered PKR inhibition (Fig. S6). To test the effect of M156-L98P in MYXV infection, we inserted it or wild-type M156R (revertant) into MYXV-M029LKOM156RKO and infected RK13 cells with the viruses. M156 was able to restore EGFP expression, whereas M156-L98P could not (Fig. 5F). EGFP expression correlated well with virus replication, which was restored by M156 but not by M156-L98P (Fig. 5G), confirming that the latter is also a loss-of-function mutation in the context of MYXV infection. The loss of M156 function in some MYXV isolates is thus a good candidate for a contributing factor to the virulence attenuation of MYXV in Australia.
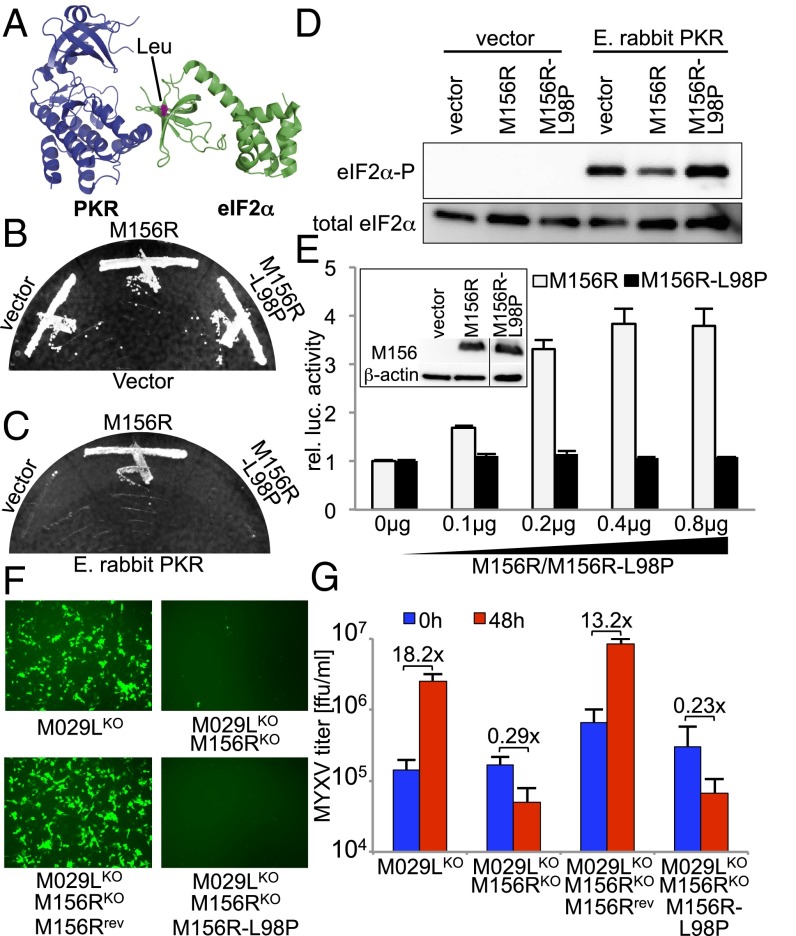
Fig. 5.
A naturally occurring M156R mutant lost its ability to inhibit PKR. (A) Crystal structure of the PKR-eIF2α complex. The residue highlighted in pink in the structure of eIF2α corresponds to Leu-98 in M156. Plasmids encoding empty vector, M156, or M156-L98P under the control of a yeast GAL-CYC1 hybrid promoter were transformed into isogenic vector (B) or E. rabbit PKR (C) containing yeast strains. Colony-purified transformants were grown under inducing conditions at 30 °C for 4 d. (D) Transformants were grown in liquid SC −Gal medium for 4 h to induce expression. Lysates were subjected to Western blot analysis for phosphorylated and total eIF2α. (E) HeLa-PKRkd cells were transfected with expression vectors for luciferase (0.05 µg), E. rabbit PKR (0.1 µg), and increasing amounts of M156R or M156R-L98P. After 48 h, luciferase activities were determined and normalized to control transfections lacking PKR inhibitors to obtain relative luciferase activities. (Inset) HeLa-PKRkd cells were transiently transfected with 3 µg of vector, M156R, or M156R-L98P plasmids. Total protein lysates were collected 48 hours after transfection to monitor M156 and β-actin expression. All samples were run on the same gel and lanes were spliced together (line) for clarity. (F and G) RK13 cells were infected with MYXV-M029LKO, MYXV-M029LKOM156RKO, MYXV-M029LKOM156RKOM156Rrevertant(rev), and MYXV-M029LKOM156RKOM156R-L98P (MOI of 5). Fluorescent images were taken at 24 hpi (F) and virus titers were measured at 0 and 48 hpi (G).
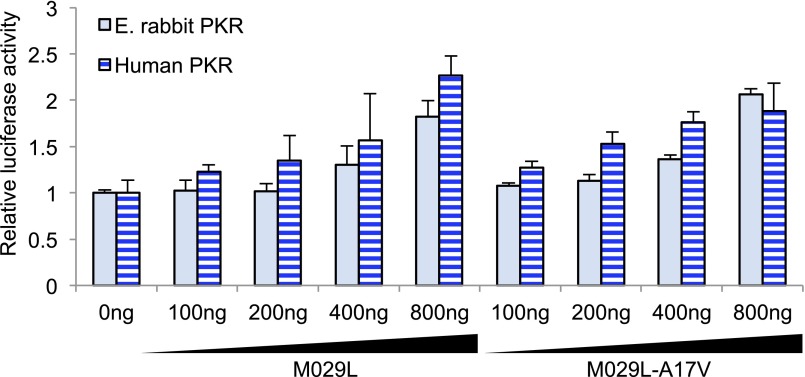
Fig. S6.
M029-A17V does not show altered PKR inhibition. Constant amounts of human or E. rabbit PKR (0.1 µg) and firefly luciferase (0.05 µg) were cotransfected with increasing amounts of M029L or M029L-A17V. Relative luciferase activities are shown. Experiments were performed in triplicate and the results are representative of three independent experiments. Error bars indicate SD.
Discussion
We have shown here that M156 of MYXV is a species-specific inhibitor of rabbit PKR, that human PKR but not rabbit PKR can restrict MYXV replication in congenic HeLa cells, and that M156 deficiency leads to virus attenuation in RK13 cells. The extent of species-specific PKR inhibition by M156 is remarkable, considering its likely mode of action as a pseudosubstrate inhibitor (23). Poxvirus K3 homologs share the common S1 fold with eIF2α, the substrate of PKR and they all likely bind to the same region of PKR. In contrast to the poxvirus K3 members, which can share as little as 20% sequence identity with each other (10), eIF2α is highly conserved, e.g., 100% sequence identity between human and rabbits in the S1 domain. The finding that PKR has been evolving rapidly can be explained by the high selective pressure exerted by pathogen-derived antagonists, such as poxviral K3 orthologs, which has caused PKR to diversify. Still, PKR faces the challenge to maintain its interaction with eIF2α, while evading inhibition by K3-family pseudosubstrate inhibitors. The best-studied PKR pseudosubstrate inhibitor is VACV K3. It was previously shown that K3 inhibited mouse PKR much better than human PKR in cell transfection assays and that hominoid PKR were more resistant to K3 inhibition than PKR from Old World and New World monkeys in yeast assays, indicating the potential for species-specific inhibition (10, 11). A limitation of studying the interaction of VACV with host proteins is that its “natural” host is unknown. Studying the interaction of MYXV proteins with innate immune proteins of a naturally infected rabbit host can therefore provide unique insights into this host–pathogen relationship. Potential species specificity of M156 was not previously considered as an explanation for its inability to inhibit human PKR in a yeast assay (12). Our results that only rabbit PKR but not human PKR was inhibited by M156 in yeast demonstrate that this assay system is informative and that species-specific activity can explain the earlier negative results obtained in yeast. This finding was confirmed and extended by the transfection assays, which showed that of the eight tested mammalian PKRs, only rabbit PKR was strongly inhibited by M156. Because MYXV has evolved in rabbits, it can be assumed that its gene products adapted to inhibit the immune response of its hosts. However, it is remarkable that PKR from other mammals were not considerably affected as “off-species targets” because M156 is predicted to bind to the PKR kinase domain at the same interaction surface as the PKR target eIF2α. The species-specific PKR inhibition by M156 is the first example where a species-specific interaction correlates with the host species restriction of MYXV virus.
The host range function of many poxvirus genes was described in cultured cells that originated from different species. Because it cannot be ruled out that cell-specific characteristics that are species independent contribute to the observed differences, we established a system that allows the comparison of PKR from different species in congenic HeLa cells. Infection of these cells showed that human PKR restricted MYXV infection, but that MYXV replicated as well in cells expressing rabbit PKR as in cells in which endogenous PKR expression was knocked down. Importantly, rabbit PKR was as efficient as human PKR in inhibiting the replication of a VACV strain that lacks the PKR inhibitors E3 and K3. These results demonstrate that MYXV can overcome the antiviral activities of only rabbit PKR, and this correlates with the ability of M156 to inhibit rabbit PKR but not human PKR. Congenic cells expressing PKR from different species might also prove useful for studying species-specific effects of other viral inhibitors.
The coevolution of both MYXV and the feral European rabbit host after the deliberate release of MYXV into Australia and Europe constitutes one of the best-studied examples of host–virus evolution on the population level. The introduction of MYXV into Australia initially led to a dramatic reduction of the rabbit population; however, within a few years, rabbit numbers rebounded. This could be attributed to the increased resistance of rabbits against MYXV and to the attenuation of MYXV in the field, the latter of which allowed for more efficient virus transmission (5). The molecular mechanisms behind these two phenomena are, however, poorly understood. The full genome sequencing of 24 MYXV that were isolated between 1951 and 1999 constituted an important step in understanding the molecular basis for MYXV evolution in Australia. Whereas many mutations were present in the isolates, no common mutations were identified that could explain differences in virulence (6, 7). The L98P mutation in M156R was identified in 13 of these isolates. Because M156-L98P was unable to inhibit rabbit PKR and led to MYXV attenuation in our assays (Fig. 5), this mutation can help explain and likely contributes to the attenuation of MYXV in Australia. The corresponding residue in eIF2α and viral homologs is highly conserved, being either a leucine in eIF2α and vIF2α, a PKR inhibitor from ranaviruses and leporipoxvirus 156, or a valine in K3/M156 orthologs from other poxviruses (18). This residue is part of a β-sheet (β5), which also comprises residues corresponding to M156 Y95 and D97, which are 100% conserved in all eIF2α homologs and are involved in the binding of eIF2α to PKR (12, 18, 22). A proline in this β-sheet would likely disrupt the β-sheet and abolish binding to PKR. Indeed, it would be instructive to know whether the rabbit PKR locus has also undergone any virus-induced selection pressures over the half century since the first field releases of MYXV.
In conclusion, the results presented here show that rabbit PKR was specifically inhibited by M156, that human PKR plays an essential role in suppressing MYXV replication in HeLa cells, and that a naturally occurring mutation in M156 abolished rabbit PKR inhibition. These phenomena might contribute to the strict host restriction of MYXV infection to only rabbits, the prevention of productive infection in nonrabbit species (particularly humans), and the attenuation of MYXV strains currently extant in Australia. Moreover, this study shows that the choice of biologically meaningful host–virus systems is important for studying host–virus interactions. Often, studies of viral molecules or viruses are performed with molecules, cell lines, or animals of nonhost species. The results of such experiments could be misleading, if the virus molecules act in a species-specific manner as shown here for M156 and rabbit PKR.
Materials and Methods
Plasmids, Yeast Strains, and Cell Lines.
PKR and viral genes were cloned into pSG5 for expression in mammalian cells. Generation of yeast strains, stably transformed with empty vector (J673) or human PKR (J983) under the control of a yeast GAL-CYC1 hybrid promoter, was described (18). A yeast strain stably transformed with E. rabbit PKR (O8) was generated using the same methods. VACV K3L and M156R were cloned into the vector pYX113, which contains the GAL-CYC1 hybrid promoter. HeLa-PKRkd cells (24), kindly provided by Charles Samuel, were stably transfected with knock-down resistant human PKR and E. rabbit PKR under the control of the human PKR promoter (19). The generation of RK13+E3L+K3L cells was described (21). See SI Materials and Methods for details.
RACE PCR.
E. rabbit RK13 cells were infected with MYXV-EGFP at an multiplicity of infection (MOI) of 1. Total RNA was collected with TRIzol reagent at 4, 12, and 24 h postinfection. RACE PCR was performed with the GeneRacer Core Kit (Invitrogen) according to the manufacturer’s instruction. See SI Materials and Methods for details.
Yeast Growth and eIF2α Phosphorylation Assays.
Experiments were performed as previously described (18, 25). See SI Materials and Methods for details.
Luciferase Assay.
The 5 × 104 HeLa-PKRkd cells were seeded in 24-well plates 1 day before transfection. For each transfection, firefly luciferase (pGL3 promoter, 0.05 μg, Promega), and pSG5 plasmids encoding PKR (0.2 μg), M156, or M029 (0.4 μg) were transfected using GenJet-HeLa (Signagen). For titration experiments, amounts of transfected plasmids are indicated in the figures. For controls, empty pSG5 vector was transfected using the same amount. Each transfection was conducted in triplicate. After 48 h, cell lysates were harvested using mammalian lysis buffer (Goldbio) and luciferase activity was determined using luciferase detection reagents (Promega) in a luminometer (Berthold).
Construction of Recombinant Viruses.
The construction of VC-R2 and MYXV-EGFP were described previously (20, 26) and the construction of M156R mutant viruses is illustrated in Fig. S5 and described in SI Materials and Methods.
Infection Assays.
HeLa-control, HeLa-PKRkd, HeLa-PKRkd+vector, HeLa-PKRkd+hPKR, and HeLa-PKRkd+Erabbit PKR cells were seeded into six-well plates and confluent monolayers (1 × 106 cells) were mock infected with PBS or infected with VC-R2 and MYXV-EGFP at an MOI of 0.1 and 1, respectively. RK13 cells were infected with MYXV-EGFP or MYXV-EGFP–derived recombinant viruses at an MOI of 5. Fluorescent pictures were taken with an inverted fluorescent microscope (Leica) and viruses were collected at 0, 24, 48, and 72 hpi for titration in RK13 cells that stably express E3 and K3. See SI Materials and Methods for details.
Western Blot Analyses.
Protein lysates from RK13 cells that were infected with MYXV-EGFP (MOI = 10) were blotted on PVDF membranes and incubated with anti-M156 (27), and after stripping, reprobed with antiserum against M-T7 (28), M130 (29), and anti–β-actin. For eIF2α phosphorylation assays, PKR-expressing cells were infected with VC-R2 or MYXV-EGFP at an MOI of 5. Total protein lysates were collected 8 hpi and eIF2α was detected with phospho-specific eIF2α and total eIF2α. See SI Materials and Methods for details.
SI Materials and Methods
Plasmids, Yeast Strains, and Cell Lines.
PKR and viral genes were cloned into the pSG5 vector (Stratagene) for transient expression in mammalian cells. European rabbit (Oryctolagus cuniculus) PKR was cloned from spleen total RNA from a New Zealand white rabbit. The obtained PKR (accession number: KT272867) differed from NP_001075682 at eight amino acid positions but was identical to ENSOCUT00000007225 (Ensmbl database). Sheep (Ovis aries) PKR (KT272868) was cloned from MDOK cells (ATCC CRL-1633). Three identical cDNA were obtained that differ from the predicted sheep PKR (XM_004007300.1) at six synonymous nucleotide positions. Rat (Rattus norvegicus) PKR was cloned from rat-1 cells and is identical to XM_008764426.1. Syrian hamster (Mesocricetus auratus) PKR was cloned from BHK-21 cells (ATCC CCL-10) and is identical to NM_001281945.1. Chinese hamster (Cricetulus griseus) PKR (KT272869) was cloned from CHO cells. Guinea pig (Cavia porcellus) PKR (KT272870) was cloned from 104C1 cells (ATCC CRL-1405) and is identical to the predicted GenBank sequence XM_003472904.2. Cloning of knock-down resistant human PKR and mouse PKR was described (10). VACV K3L, M029L, and M156R short and long forms were cloned into pSG5. The M156R-L98P mutation was generated by site-directed mutagenesis PCR using the PfuUltra High-Fidelity DNA polymerase (Invitrogen). PCR was performed using the pSG5-M156R short construct as the template with primers containing the leucine-to-proline mutation (CTG to CCG) flanked by 15 nucleotides identical to the template up- and downstream of the mutation. All ORFs were sequenced to confirm correct sequences.
Generation of yeast strains stably transformed with empty vector (J673) or human PKR (J983) under the control of a yeast GAL-CYC1 hybrid promoter was described (18). A yeast strain stably transformed with E. rabbit PKR (O8) was generated using the same methods. VACV K3L and M156R were cloned into the vector pYX113 (R&D Systems), which contains the GAL-CYC1 hybrid promoter and the selectable marker URA3. HeLa-PKRkd cells and control HeLa cells were kindly provided by Charles Samuel (24). RK13 cells were kindly provided by Bernard Moss. To generate HeLa-PKRkd cells that express PKR, a plasmid was constructed that contains the human PKR promoter as described (19) (nucleotide positions −412 to +97 relative to the transcriptional start site followed by the intron of the rabbit β-globin gene from pSG5 and a multiple cloning site followed by a sequence encoding two FLAG tags and inserted into the pEGFP-N1 (Clontech) in place of the CMV promoter/enhancer, the multiple cloning site, and the EGFP ORF. Knock-down resistant human PKR and E. rabbit PKR were cloned into this vector with the FLAG tags at their C termini. HeLa-PKRkd cells were transfected using GenJet-HeLa (SignaGen Laboratories) according to the manufacturer’s instructions. Cells were trypsinized 48 h after transfection and cultured in the presence of 1 µg/mL geneticin (Invitrogen) for 14 d. To select clones, cells were seeded on 96-well plates at a density of 0.3 and one cell per well until single colonies appeared. Clones were then amplified and proteins were isolated to detect the expression of transfected PKR with an anti-FLAG tag antibody (ABM) and the absence of endogenous human PKR with the monoclonal anti-human PKR antibody 71/10. A human PKR reconstituted clone (HeLa-h14) that showed a comparable PKR expression level to the endogenous human PKR in HeLa control cells was selected and amplified. A E. rabbit PKR-expressing clone that showed PKR expression levels comparable to that of human PKR in HeLa-h14, as determined with the anti-FLAG tag antibody, was used for subsequent experiments. For transfections, plasmids were prepared using NucleoBond Xtra Midi Endotoxin Free (Macherey-Nagel).
5′ RACE PCR.
E. rabbit RK13 cells were infected with MYXV-EGFP at a MOI of 1. Total RNA was collected with TRIzol reagent (Invitrogen) at 4, 12, and 24 h postinfection. RACE PCR was performed with the GeneRacer Core Kit (Invitrogen) according to the manufacturer’s instructions. Briefly, two rounds of PCR were conducted to amplify the 5′ end of M156R cDNA using 5′ adaptor primers provided in the kit and M156R-specific reverse primers. The second-round PCR products were cloned into the TOPO-TA vector (Invitrogen) and six colonies from each time point were randomly chosen for sequencing.
Yeast Growth and eIF2α Phosphorylation Assays.
Yeast transformation was performed using the lithium acetate/PEG method. J673, J983, and O8 strains were transformed with empty vector pYX113, K3, or M156 expressing plasmids. For each transformation, four independent clones were picked and colony purified on SD plates. Purified colonies were then streaked on either glucose (noninducing conditions) or galactose containing medium [synthetic complete medium containing 2% (wt/vol) galactose and all amino acids except uracil, inducing conditions], and grown for 4 d at 30 °C. For Western blot analyses of yeast proteins, all transformants were grown to saturation in SD medium overnight. The starting culture was then diluted 1:40 in SD medium on the second day and grown to OD600 = 0.6 and subsequently shifted to SC−Gal medium to induce expression. After 4 h, ODs of the cultures were obtained and cultures were diluted accordingly in water to reach the same ODs. Equal amounts of cells for each sample were lysed using the trichloroacetic acid (TCA) method. Samples were mixed with loading buffer with reducing agent and neutralized with 1 M Tris base. Proteins were fractionated on 10% (vol/vol) SDS/PAGE gels and then transferred to PVDF membranes (Millipore). Membranes were incubated with antiphospho-S51 eIF2α (Bio-Source International) and then stripped and reprobed with antitotal eIF2α antibodies (25). Enhanced chemiluminescence was detected with a Kodak image system.
Luciferase Assay.
The 5 × 104 HeLa-PKRkd cells were seeded in 24-well plates 1 day before transfection. For each transfection, firefly luciferase (pGL3 promoter, 0.05 μg, Promega), and pSG5 plasmids encoding PKR (0.5 μg), M156, M156-L98P, or M029 (0.4 μg) were transfected using GenJet-HeLa. For titration experiments, amounts of transfected plasmids are indicated in the figures. For controls, empty pSG5 vector was transfected using the same amount. Each transfection was conducted in triplicate. After 48 h, cell lysates were harvested using mammalian lysis buffer (Goldbio) and luciferase activity was determined using luciferase detection reagents (Promega) in a luminometer (Berthold). Each experiment was performed at least three times and representative experiments are shown in the results. Error bars represent the SD of three independent transfections.
Infection Assays and Western Blot Analyses.
RK13 cells were infected with mock (PBS) or MYXV-EGFP (Lu strain) at a MOI of 10. Proteins were collected at the indicated time points using Nonidet P-40 lysis buffer (50 mM Tris, 150 mM NaCl, 1% Nonidet P-40, and proteinase inhibitor mixture; Roche). Protein concentrations were determined by the Bradford assay (BioRad). Protein samples were separated on 10% (vol/vol) SDS/PAGE gels and transferred to PVDF membranes (GE Healthcare) using a methanol-based wet transfer apparatus (BioRad). Membranes were blocked with 5% (wt/vol) nonfat milk or 5% (wt/vol) BSA when detecting eIF2α-P in TBST buffer (20 mM Tris, 150 mM NaCl, and 0.1% Tween-20 pH 7.6) for 3 h at 4 °C and then incubated overnight with antiserum raised against M156 (27) at 4 °C with rocking, and then stripped and reprobed with antiserum against M-T7 (28), M130 (29), and an antibody against β-actin (Sigma-Aldrich). After washing, membranes were incubated with goat anti-rabbit or goat anti-mouse secondary antibodies (1:30,000) in TBST containing 5% (wt/vol) nonfat milk for 1 h at room temperature with agitation followed by TBST washing four times for 20 min each. Proteins were detected with Proto-Glo ECL (National Diagnostics) and images were taken with a Kodak image system.
HeLa-control, HeLa-PKRkd, HeLa-PKRkd+vector, HeLa-PKRkd+h. PKR, and HeLa-PKRkd+E. rabbit PKR cells were seeded into six-well plates and confluent monolayers (1 × 106 cells) were mock infected with PBS or infected with VC-R2 and MYXV-EGFP at a MOI of 0.1 and 1, respectively. Fluorescent pictures were taken with an inverted fluorescent microscope (Leica) at 48 hpi and viruses were collected at 0, 24, 48, and 72 hpi and were titered in RK13 cells that stably express E3 and K3 (21). For eIF2α phosphorylation assays, the previously described PKR-expressing cells were infected with VC-R2 and MYXV-EGFP at a MOI of 5. Total protein lysates were collected at 8 hpi with 1% SDS in PBS and fractionated on 10% (vol/vol) SDS/PAGE gels. Western blot analyses were performed as described above. Primary antibodies detecting S51 phospho-eIF2α (Bio-Source International) and total eIF2α (Santa Cruz) were diluted 1:10,000 and 1:1,000, respectively, in TBST.
Construction of Recombinant Viruses.
The generation of VC-R2, a vaccinia virus Copenhagen strain that lacks E3L and K3L (VACVΔK3L/ΔE3L) and carries a destabilized EGFP gene in the E3L locus, and MYXV-EGFP in which a synthetic vaccinia virus early/late promoter-driven EGFP cassette was inserted into the intergenic region in the M135–136 locus, were described previously (20, 26). The strategy to generate recombinant myxoma viruses is illustrated in Fig. S5. To create a M156R deficient virus, a recombinant plasmid was first constructed in which a poxvirus synthetic early/late promoter-driven mRFP is flanked by sequences 5′ and 3′ of the M156R locus, which contain partial sequences of M154L (5′), M008.1L (3′) and intergenic sequences (both 5′ and 3′). MYXV-M029LKOM156RKO was generated by infecting RK13+E3L+K3L cells with MYXV-EGFP (Lu), followed by transfection of the recombinant plasmid. Multiple rounds of foci purification were performed with the same cell line using mRFP as selection marker. The recombinant locus was then PCR amplified from the viral clone and sequenced to confirm successful recombination. To generate MYXV-M029LKOM156RKOM156Rrev and MYXV-M029LKOM156RKOM156R-L98P recombinant viruses, RK13+E3L+K3L cells were infected with MYXV-M029LKOM156RKO followed by transfection of recombinant plasmids in which the original M156 or the M156-L98P coding sequences were flanked by 5′ and 3′ arms described above. Multiple rounds of foci purification were performed in RK13+E3L+K3L cells based on the absence of mRFP expression. In all recombinant MYXV, the M156R locus as well as the region comprising the partially duplicated M156R copy in the 5′ region was PCR amplified and sequenced to confirm successful integration and exclude unwanted mutations.
Acknowledgments
We thank Thomas E. Dever for helpful discussions. This work was supported, in part, by Grant R01 AI114851 (to S.R.) from the National Institute of Allergy and Infectious Diseases, National Institutes of Health and grants from the Johnson Cancer Research Center (to C.P., S.L.H., and S.R.).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
Data deposition: The sequences reported in this paper have been deposited in the GenBank database (accession nos. KT272867–KT272870).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1515613113/-/DCSupplemental.
References
- 1.Seet BT, et al. Poxviruses and immune evasion. Annu Rev Immunol. 2003;21:377–423. doi: 10.1146/annurev.immunol.21.120601.141049. [DOI] [PubMed] [Google Scholar]
- 2.Haller SL, Peng C, McFadden G, Rothenburg S. Poxviruses and the evolution of host range and virulence. Infect Genet Evol. 2014;21:15–40. doi: 10.1016/j.meegid.2013.10.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Werden SJ, Rahman MM, McFadden G. Poxvirus host range genes. Adv Virus Res. 2008;71:135–171. doi: 10.1016/S0065-3527(08)00003-1. [DOI] [PubMed] [Google Scholar]
- 4.Bratke KA, McLysaght A, Rothenburg S. A survey of host range genes in poxvirus genomes. Infect Genet Evol. 2013;14:406–425. doi: 10.1016/j.meegid.2012.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kerr PJ. Myxomatosis in Australia and Europe: A model for emerging infectious diseases. Antiviral Res. 2012;93(3):387–415. doi: 10.1016/j.antiviral.2012.01.009. [DOI] [PubMed] [Google Scholar]
- 6.Kerr PJ, et al. Evolutionary history and attenuation of myxoma virus on two continents. PLoS Pathog. 2012;8(10):e1002950. doi: 10.1371/journal.ppat.1002950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kerr PJ, et al. Genome scale evolution of myxoma virus reveals host-pathogen adaptation and rapid geographic spread. J Virol. 2013;87(23):12900–12915. doi: 10.1128/JVI.02060-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Davies MV, Chang HW, Jacobs BL, Kaufman RJ. The E3L and K3L vaccinia virus gene products stimulate translation through inhibition of the double-stranded RNA-dependent protein kinase by different mechanisms. J Virol. 1993;67(3):1688–1692. doi: 10.1128/jvi.67.3.1688-1692.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Langland JO, Jacobs BL. The role of the PKR-inhibitory genes, E3L and K3L, in determining vaccinia virus host range. Virology. 2002;299(1):133–141. doi: 10.1006/viro.2002.1479. [DOI] [PubMed] [Google Scholar]
- 10.Rothenburg S, Seo EJ, Gibbs JS, Dever TE, Dittmar K. Rapid evolution of protein kinase PKR alters sensitivity to viral inhibitors. Nat Struct Mol Biol. 2009;16(1):63–70. doi: 10.1038/nsmb.1529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Elde NC, Child SJ, Geballe AP, Malik HS. Protein kinase R reveals an evolutionary model for defeating viral mimicry. Nature. 2009;457(7228):485–489. doi: 10.1038/nature07529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ramelot TA, et al. Myxoma virus immunomodulatory protein M156R is a structural mimic of eukaryotic translation initiation factor eIF2alpha. J Mol Biol. 2002;322(5):943–954. doi: 10.1016/s0022-2836(02)00858-6. [DOI] [PubMed] [Google Scholar]
- 13.Cameron C, et al. The complete DNA sequence of myxoma virus. Virology. 1999;264(2):298–318. doi: 10.1006/viro.1999.0001. [DOI] [PubMed] [Google Scholar]
- 14.Willer DO, McFadden G, Evans DH. The complete genome sequence of shope (rabbit) fibroma virus. Virology. 1999;264(2):319–343. doi: 10.1006/viro.1999.0002. [DOI] [PubMed] [Google Scholar]
- 15.Kerr PJ, et al. Comparative analysis of the complete genome sequence of the California MSW strain of myxoma virus reveals potential host adaptations. J Virol. 2013;87(22):12080–12089. doi: 10.1128/JVI.01923-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kawagishi-Kobayashi M, Silverman JB, Ung TL, Dever TE. Regulation of the protein kinase PKR by the vaccinia virus pseudosubstrate inhibitor K3L is dependent on residues conserved between the K3L protein and the PKR substrate eIF2alpha. Mol Cell Biol. 1997;17(7):4146–4158. doi: 10.1128/mcb.17.7.4146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kawagishi-Kobayashi M, Cao C, Lu J, Ozato K, Dever TE. Pseudosubstrate inhibition of protein kinase PKR by swine pox virus C8L gene product. Virology. 2000;276(2):424–434. doi: 10.1006/viro.2000.0561. [DOI] [PubMed] [Google Scholar]
- 18.Rothenburg S, Chinchar VG, Dever TE. Characterization of a ranavirus inhibitor of the antiviral protein kinase PKR. BMC Microbiol. 2011;11:56. doi: 10.1186/1471-2180-11-56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kuhen KL, Samuel CE. Isolation of the interferon-inducible RNA-dependent protein kinase Pkr promoter and identification of a novel DNA element within the 5′-flanking region of human and mouse Pkr genes. Virology. 1997;227(1):119–130. doi: 10.1006/viro.1996.8306. [DOI] [PubMed] [Google Scholar]
- 20.Johnston JB, et al. Role of the serine-threonine kinase PAK-1 in myxoma virus replication. J Virol. 2003;77(10):5877–5888. doi: 10.1128/JVI.77.10.5877-5888.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rahman MM, Liu J, Chan WM, Rothenburg S, McFadden G. Myxoma virus protein M029 is a dual function immunomodulator that inhibits PKR and also conscripts RHA/DHX9 to promote expanded host tropism and viral replication. PLoS Pathog. 2013;9(7):e1003465. doi: 10.1371/journal.ppat.1003465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Dar AC, Dever TE, Sicheri F. Higher-order substrate recognition of eIF2alpha by the RNA-dependent protein kinase PKR. Cell. 2005;122(6):887–900. doi: 10.1016/j.cell.2005.06.044. [DOI] [PubMed] [Google Scholar]
- 23.Dar AC, Sicheri F. X-ray crystal structure and functional analysis of vaccinia virus K3L reveals molecular determinants for PKR subversion and substrate recognition. Mol Cell. 2002;10(2):295–305. doi: 10.1016/s1097-2765(02)00590-7. [DOI] [PubMed] [Google Scholar]
- 24.Zhang P, Samuel CE. Protein kinase PKR plays a stimulus- and virus-dependent role in apoptotic death and virus multiplication in human cells. J Virol. 2007;81(15):8192–8200. doi: 10.1128/JVI.00426-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Dever TE, et al. Phosphorylation of initiation factor 2 alpha by protein kinase GCN2 mediates gene-specific translational control of GCN4 in yeast. Cell. 1992;68(3):585–596. doi: 10.1016/0092-8674(92)90193-g. [DOI] [PubMed] [Google Scholar]
- 26.Brennan G, Kitzman JO, Rothenburg S, Shendure J, Geballe AP. Adaptive gene amplification as an intermediate step in the expansion of virus host range. PLoS Pathog. 2014;10(3):e1004002. doi: 10.1371/journal.ppat.1004002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Liu J, et al. Myxoma virus M064 is a novel member of the poxvirus C7L superfamily of host range factors that controls the kinetics of myxomatosis in European rabbits. J Virol. 2012;86(9):5371–5375. doi: 10.1128/JVI.06933-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Mossman K, et al. Myxoma virus M-T7, a secreted homolog of the interferon-gamma receptor, is a critical virulence factor for the development of myxomatosis in European rabbits. Virology. 1996;215(1):17–30. doi: 10.1006/viro.1996.0003. [DOI] [PubMed] [Google Scholar]
- 29.Barrett JW, et al. Myxoma virus M130R is a novel virulence factor required for lethal myxomatosis in rabbits. Virus Res. 2009;144(1-2):258–265. doi: 10.1016/j.virusres.2009.05.009. [DOI] [PubMed] [Google Scholar]